1. Introduction
Over the past few decades, gene therapy has been prevalent and considered a promising strategy to prevent and treat various diseases such as cancer, infectious diseases, cardiovascular diseases, etc. [
1,
2,
3]. To this end, it is an urgency to develop gene delivery vectors with low cytotoxicity and high transfection efficiency by overcoming several barriers in the eukaryotic cells, including cellular uptake, endosome escape and nuclear transport. Current gene delivery systems mainly consist of viral and non-viral vectors. Viral vectors are highly effective and dominant in gene delivery, accounting for more than 70% of the vectors used in clinical trials [
4], while it is subject to safety problems, such as immunogenicity, inability for fast clearance, and high cost of manufacturing. On the other hand, non-viral vectors are advantageous for their safety, flexibility in packaging nucleic acids and ease production [
5]. Common non-viral vectors include inorganic nanoparticles, lipids, polymers, peptides, hybrid systems, etc.
Cell-penetrating peptides (CPPs), consisted of around 5–30 amino acids, are known as highly active non-viral delivery vectors. They are degradable, easy to synthesize and modify, and enable simple formation of non-covalent complex by mixing CPP and nucleic acid. CPPs can penetrate cell membranes and convey various bioactive cargo into the cells via energy-dependent or -independent mechanisms [
6]. The most common CPP is the HIV-1 Tat peptide, which has been successfully applied for the delivery of drugs, protein and DNA into cells [
7]. RALA is a superior gene delivery CPP developed by McCarthy et al. [
6]. It was derived from fusogenic peptides, including glutamic acid-rich GALA and lysine-rich KALA. It has been applied as a vector to develop DNA vaccination for cervical cancer [
8], as well as a vector for both FKBPL DNA and RNAi delivery [
9]. Under a collaboration with McCarthy, Yan et al. developed collagen/GAG scaffolds activated by RALA-siMMP-9 complexes for improved diabetic wound healing application [
10]. It has been reported that histidine possesses strong endosomolytic ability and histidine enriched CPP demonstrated enhanced cellular uptake and endosomal escape [
11]. While RALA has only one histidine on its backbone. In order to improve the transfection efficiency of RALA, our group designed a series of peptides named HALA by replacing arginine with histidine within RALA. Among these HALA peptides, HALA2 (WEARLARALARALARHLARALAHALHACEA) showed improved pDNA and siRNA transfection efficiency [
12].
Development of safe and high-efficient CPPs is crucial for the clinical application of gene therapy. Enhancement of cellular interaction of CPPs by introduction of cell binding sequences (such as GAG binding sequences) had demonstrated superior efficacy in cargo delivery [
13,
14]. Dixon et al. developed CPPs with glycosaminoglycan-binding domain to enhance cell interaction and thus improve intracellular translocation [
13,
14]. Therefore, it would be very interesting to explore if the gene delivery efficacy of RALA can be further improved by using this strategy. VGVAPG is a common peptide sequence derived from an extracellular matrix protein called elastin [
15]. During degradation
in vivo, elastin releases bioactive peptide fragments known as elastin-derived peptides (EDPs) or elastokines [
16]. These elastokines mainly bind to the elastin receptor complex on the cell surface and subsequently trigger a variety of cellular functions, including cancer progress [
16]. As a common elastokine, VGVAPG can specifically bind to a 67-kDa elastin-binding protein which is a central part of the elastin receptor complex [
17]. The VGVAPG sequence has been reported to play important role in the proliferation and differentiation of cells [
18,
19], cytokine production [
20], immune cell modulation [
21,
22] and angiogenesis [
23]. Currently, there is no reported study on exploring this sequence for gene delivery.
Therefore, we aimed to explore if the introduction of cell binding sequence VGVAPG onto the C-terminal of RALA can improve its gene delivery efficiency and the effect of the modified peptide (RALA-E) on cellular behavior.
2. Materials and Methods
2.1. Preparation of Peptide/Nucleic Acid Complexes
RALA and RALA-E peptides (
Table 1) with 98% purity were synthesized by Jiangsu Jitai Peptide industry science and technology Co., Ltd. (Yancheng, Jiangsu, China), supplied as a desalted lyophilized powder. The pDNA (pCMV-C-EGFP) with length of 5010 bp was purchased from the Beyotime (D2626, Shanghai, China). The miR-146a with length of 22 bp (Sequence: UGAGAACUGAAUUCCAUGGGUU) was provided by GenePharma (Suzhou, China).
The complexes were prepared at different N:P ratios ranging from 1 to 12 by adding appropriate amount of peptide solution to the pDNA or miR-146a solutions. Then the peptide/pDNA or peptide/miRNA solutions were mixed by using pipette to form the peptide/pDNA or peptide/miRNA complexes. The complexes were incubated at room temperature for 20 min for stabilization. The peptide, pDNA and miR-146a were dissolved in ultrapure water and the complexes were also prepared in ultrapure water. The final concentrations of pDNA and miR-146a in the above prepared complexes solutions were fixed to 100 nmol/L and 800 nmol/L, respectively. These used concentrations were already optimized from previous studies [
10,
13]. The N:P ratios of peptide to nucleic acid were calculated based on the molar ratio of the positively charged arginine in the peptide to the negatively charged phosphates in the pDNA/miRNA backbone [
9,
10]. All complexes were used immediately following their preparation unless otherwise stated.
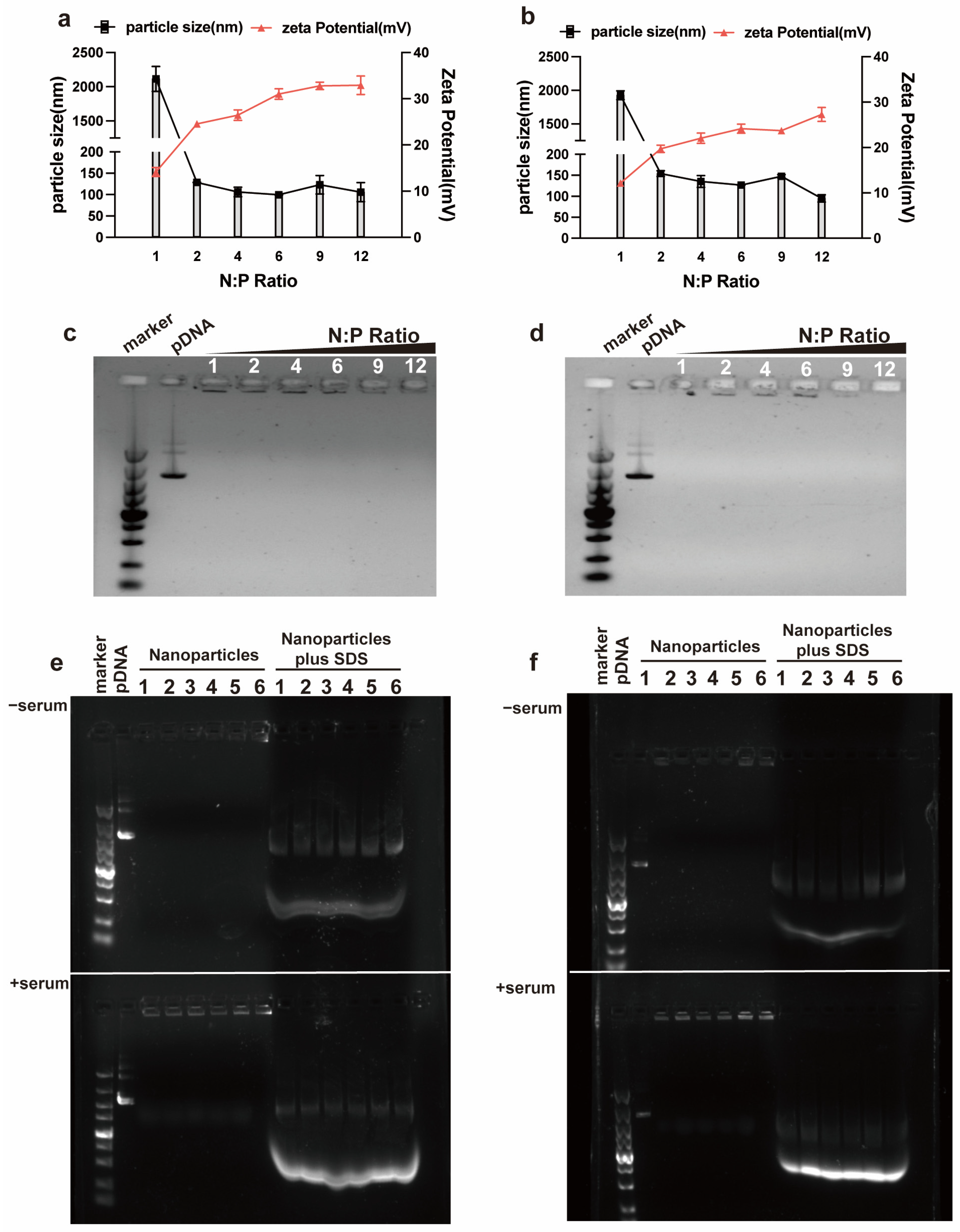
2.2. Agarose Gel Retardation Analysis
RALA/pDNA or RALA-E/pDNA complexes at N:P ratios of 1, 2, 4, 6, 9, 12 were prepared using procedure as mentioned above (The final concentrations of pDNA and miR-146a were fixed to 100 nmol/L). The 1% agarose gel was prepared by adding 0.5 g Agarose (Biosharp, Hefei, China) into 50 mL TAE buffer (Biosharp, Hefei, China), followed by heating until boiling in a microwave oven. Afterwards, electrophoresis of the peptide/pDNA complexes was performed by adding the complexes onto the agarose gel containing Gel Red Nucleic Acid Gel Stain (#11918, Accurate Biology, Changsha, China), under 120 V for 30 min and within the TAE running buffer. Afterward, the gel was visualized by using a ChemiDoc Touch (Bio-rad, Hercules, CA, USA).
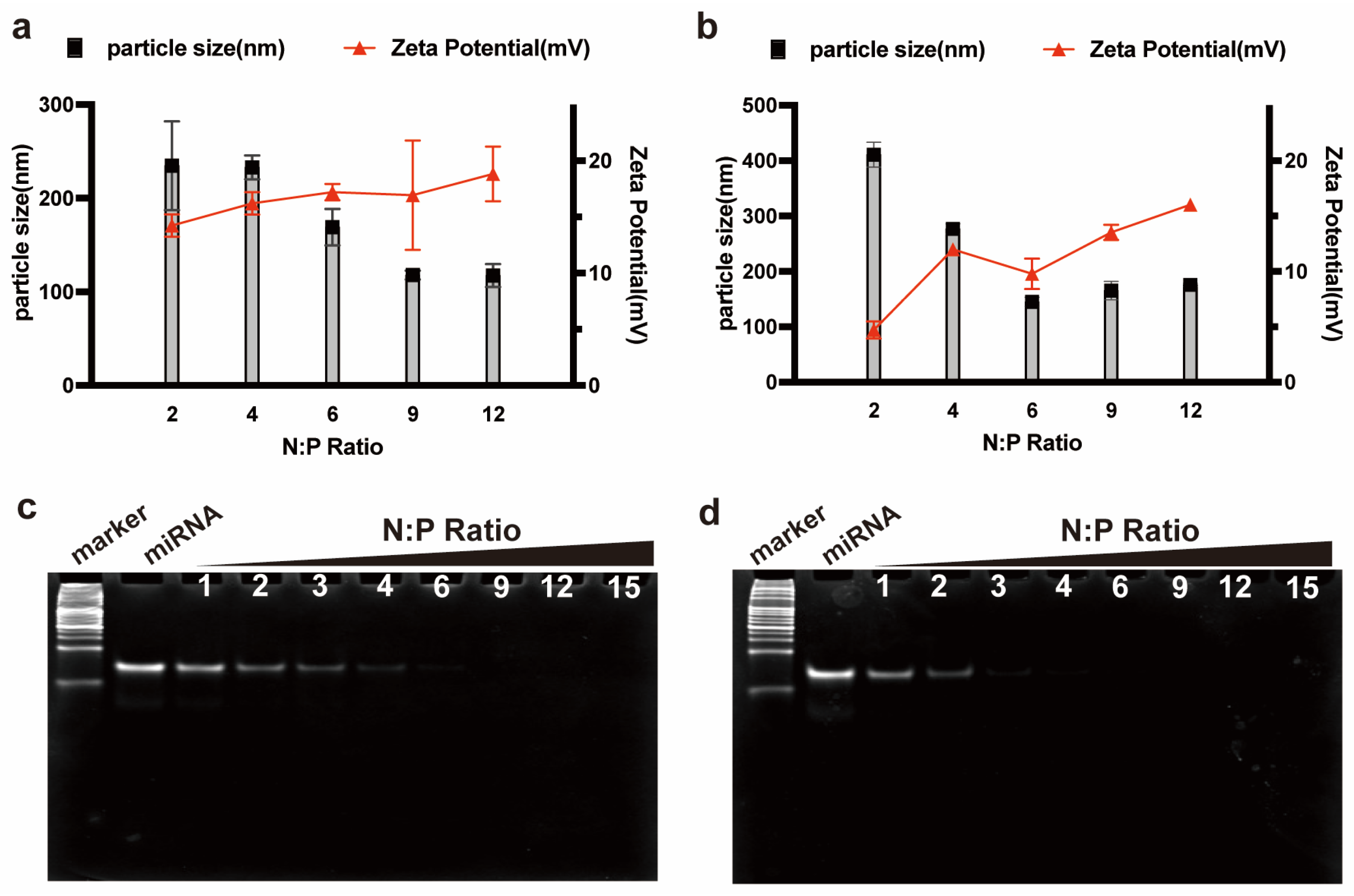
2.3. PAGE Gel Retardation Assay
Peptide/miRNA complexes at N:P ratios of 1–15 were prepared. The PAGE gel was prepared by using components as mentioned in
Table 2. Samples and 5 µL 20 bp DNA ladder (Takara, Kyoto, Japan) was added to the gel. The electrophoresis was run within tris-acetate buffer at 80 V for 2 h. Then the gel was stained in the TBE buffer with 2 µL loading dye (Thermo Fisher Scientific, Waltham, MA, USA). Images were taken using a ChemiDoc Touch (Bio-rad, Hercules, CA, USA) after staining the gel in Gel Red Nucleic Acid Gel Stain solution for 1 h.
2.4. Particle Size and Zeta Potential Analysis
Peptide/pDNA and peptide/miRNA complexes at N:P ratios of 1, 2, 4, 6, 9, 12 were formulated as described in
Section 2.1 and diluted 20 times (50 µL to 1 mL) by ddH
2O for the measurement of the particle size, Zeta potential by using a Malvern Zetasizer ZS 3000 (Malvern Instruments, Malvern, Worcestershire, UK).
2.5. Serum Stability Evaluation
Peptide/pDNA complexes were prepared at N:P ratio of 9. Then, the complexes were incubated for 1, 2, 3, 4, 5 and 6 h at room temperature with or without 10% fetal calf serum (Gibco, Thermo Fisher Scientific, Waltham, MA, USA) [
6]. Subsequently, 10% sodium dodecyl sulfate (SDS, Sigma-Aldrich, St. Louis, MO, USA) was added to decomplex the peptide/pDNA complexes. After incubation for 10 min, samples were electrophoresed within a 1% agarose gel containing GelRed with Tris-acetate running buffer, under 120 V for 30 min. Afterwards, the gel was visualized using a ChemiDoc Touch (Bio-rad, Hercules, CA, USA).
2.6. Cell Culture
Human embryonic kidney-293T (HEK-293T), HeLa and THP-1 cell lines were provided by Stem Cell Bank, Chinese Academy of Sciences (Shanghai, China). Real Time-PCR was utilized to detect mycoplasma of these cell lines cells and all the cells used in this study were proofed to be free of mycoplasma. HEK-293T and HeLa cells were cultured in high glucose Dulbecco’s Modified Eagle’s Medium (DMEM, Thermo Fisher Scientific, Waltham, MA, USA) supplemented with 1% penicillin/streptomycin and 10% fetal bovine serum (Gibco, Thermo Fisher Scientific, Waltham, MA, USA) and maintained in an incubator under standard cell culture conditions (37 °C, 90% humidity and 5% CO2). THP-1 cells were cultured in Roswell Park Memorial Institute (RPMI) 1640 (Gibco, Thermo Fisher Scientific, Waltham, MA, USA) with 1% penicillin/streptomycin and 10% fetal bovine serum (Gibco, Thermo Fisher Scientific, Waltham, MA, USA) in standard culture conditions.
2.7. Macrophage Polarization
THP-1 cells were seeded in 6-well plate with the density of 80% in the RPMI 1640 complete medium and stimulated by Phorbol-12-myristate-13-acetate (PMA, 100 ng/mL; Sigma-Aldrich, St. Louis, MO, USA) for 24 h to differentiation into M0 macrophages. Then the M0 macrophages were polarized toward M1 macrophages by treatment with LPS (1000 ng/mL; Sigma-Aldrich, St. Louis, MO, USA) and IFN-γ (20 ng/mL; Sigma-Aldrich, St. Louis, MO, USA) for 48 h. M1 macrophages were then used for the following transfection essay.
2.8. Transfection and Endocytosis Mechanism Studies
For pDNA transfection, 50,000 HEK-293T cells and HeLa cells were seeded per well in a 24-well plate in complete high glucose DMEM approximately 24 h prior to transfection. The media in the wells were removed and replaced by OptiMEM Reduced Serum Medium (GibcoTM, Thermo Fisher Scientific, Waltham, MA, USA) 1 h prior to transfection. Peptide/pDNA complexes were formulated at a N:P ratio of 9 and incubated for 20 min before adding to the cells (final concentration of the pDNA was fixed to 10 nmol/L). Complexes were then incubated with the cells in OptiMEM Reduced Serum Medium for 6 h, before the transfection media were removed and replaced with fresh high glucose DMEM media. Cells were cultured for 2 days before further assessment.
For endocytosis mechanism study, transfection was conducted on HEK-293T cells, under 4 °C or with the addition of either 5 mg/mL Methyl-β-cyclodextrin (MβCD, Aladdin, Shanghai, China), 10 mg/mL Chlorpromazine(Aladdin, Shanghai, China) or 10 mg/mL Genistein (Aladdin, Shanghai, China). Cells after transfection with the addition of various inhibitors or incubated under 4 °C were analyzed by flow cytometer (Beckman Coulter, Brea, CA, USA). Decreasing proportion ratio was calculated according to the following equations (RT: room temperature):
miRNA transfection was performed on M1 macrophages. The peptide/miRNA complexes were prepared at N:P ratios of 9. The media in the wells were removed and replaced with OptiMEM Reduced Serum Medium (Thermo Fisher Scientific, Waltham, MA, USA) 1 h prior to transfection. M1 macrophage were transfected with the peptide/miRNA complex for 6 h, then the culture medium was changed to RMPI 1640. Non-target miRNA was delivered as a negative control.
2.9. Fluorescence Imaging
Fluorescence imaging was conducted by using a Leica DMIL microscope (Leica Microsystems, Wetzlar, Germany) equipped with FITC filter with the magnification of 40 times the actual size. Images of the transfected HEK-293T cells and HeLa cells were taken 2 days after the transfection of the GFP protein-expression pDNA, and images of the M1 macrophages were taken 6 h after transfection of miRNA with FAM- fluorophore.
2.10. Flow Cytometry Analysis
For the transfection efficiency experiment, HEK-293T and HeLa cells were firstly detached by using trypsin (Sigma-Aldrich, St. Louis, MO, USA), and then the detached cells were collected by centrifuge (2000 rpm, 3 min) in FACS tubes 48 h after transfection of pDNA. The macrophages were detached by using accutase (Sigma-Aldrich, St. Louis, MO, USA) 6 h after the transfection of miRNA. The detached cells were then washed with FACS Buffer (1× PBS, 0.5% BSA, 2 mM EDTA) by centrifuging at 3500 rpm for 5 minutes under 4 °C. After discarding the supernatant, the cells in each tube were resuspended in 300 μL of FACS buffer. Transfection efficiency was quantitatively assessed by using a flow cytometry (Beckman Coulter, Brea, CA, USA). Non-transfected cells were used as control.
For cytotoxicity test, the cells were washed with FACS Buffer, then resuspended in 100 μL of FACS buffer and dyed by using 1 µL Ghost Dye™ Violet 510 (GV510, Tonbo Biosciences, San Diego, CA, USA) for 15 min in dark on ice. Afterwards, the cells were washed again and resuspended in 300 μL of FACS buffer. Cytotoxicity was detected by using CytoFLEX flow cytometry (Beckman Coulter, Brea, CA, USA).Non-transfected cells were used as control. Data were analyzed by using the FlowJo V10 software (BD Life Sciences–FlowJo, Ashland, OR, USA).
2.11. Confocal Microscope Observation
M1 macrophages cells were stained and observed by confocal microscope, after transfection by peptide/miR-146a at N:P 9 for 6 and 24 h. Briefly, the transfected M1 macrophages were washed by PBS (Gibco, Thermo Fisher Scientific, Waltham, MA, USA), then fixed for 10 min at room temperature by using 4% paraformaldehyde solution (Biosharp, Hefei, China). Afterwards, the cells were permeabilized with 0.5% Triton X-100 (Biofroxx, Hesse, Germany) solution for 5 min. In the following, the cells were stained in FITC–conjugated phalloidin solution (diluted 100× in PBS; YEASEN, 40735ES75, Shanghai, China) for 30 min, in dark and under room temperature. After being washed by PBS, cells were incubated in DAPI (1:1000, 1 min, room temperature; Invitrogen, Carlsbad, CA, USA). The stained cells were observed by confocal fluorescence microscopy (LSM 710 NLO, Zeiss, Oberkochen, Germany).
2.12. Quantitative Real-Time PCR Assay
Total RNA was isolated from M1 macrophages by using TRIzol
TM reagents (Invitrogen, Carlsbad, CA, USA) 36 h after transfection. The reverse transcription of first-strand cDNA was performed as following steps: 37 °C for 15 min, 85 °C for 5 s, and then 37 °C for 10 min by using a PrimeScript
®RT reagent kit (Takara, Kyoto, Japan) according to the manufacturer’s protocol. Quantitative real-time PCR analysis for mRNA expression of TRAF6, TNF-α, IL-1β and iNOS on macrophages were performed by using SYBR Green PCT Master Mix (Takara, Kyoto, Japan). The mRNA level of β-actin was used as an internal control. The sequences of oligonucleotides used are described in
Table 3.
2.13. Statistical Analysis
Statistical analysis was performed using Prism 5.0 (GraphPad Software, Inc., 630 La Jolla, CA, USA). Statistically significant differences of data were calculated using the one-tailed unpaired t-test with a p-value ≤ 0.05 considered as significant. Three independent experiments were performed for each analysis and all the results were presented as mean ± SD (n = 3), unless otherwise specified.
4. Discussion
In this study, we modified RALA peptide with VGVAPG fragment and generated RALA-E, in order to improve cellular interaction of the CPP/nucleic acid complexes and facilitate cell transfection. Our results showed that such modification significantly increased the transfection efficiency of pDNA in HEK-293T and HeLa cells. Additionally, both RALA and RALA-E peptides can deliver miRNA into macrophages to modulate the expressions of target genes, with good cytocompatibility. Our research illustrates for the first time that modification of CPP with VGVAPG fragment can be a promising strategy for the design of novel CPPs with improved gene transfection efficacy.
Necessary premise for CPPs as gene delivery carrier is their ability to complex the nucleic acid to form stable complexes and maintain the integrity of loaded genes. Our study showed that RALA-E had strong binding capacity to both pDNA and miR-146a. This illustrates that RALA-E is capable to complex nucleic acid of a wide range of length, from less than 30 bp to more than 5000 bp, making it a versatile gene delivery vector.
The particle size and Zeta potential of complexes are key factors affecting cellular uptake. Complexes with the Zeta potential above 10 mV are considered cationic and can produce sufficient interaction with negatively charged cell surface [
24], thereby were endocytosed successfully. The weakly positively charged complexes will gather together due to insufficient charge repulsion, which affects the uptake efficiency. RALA contains a high proportion of positively charged arginine, which can bind to negatively charged nucleic acids through non-covalent electrostatic interaction. As the peptide/nucleic acid complexes were prepared at N:P ratio 9 in current study, therefore the complexes possessed a large number of positive charges to facilitate their membrane permeability, making them easy to be taken up. The amino acids within VGVAPG are all neutral, so it has little effect on the charge of the peptide.
Transfection efficiency is one critical aspect to evaluate whether a peptide can be a promising gene delivery vector. Several studies had reported that RALA/nucleic acid complexes possessed excellent transfection efficiency [
6,
8,
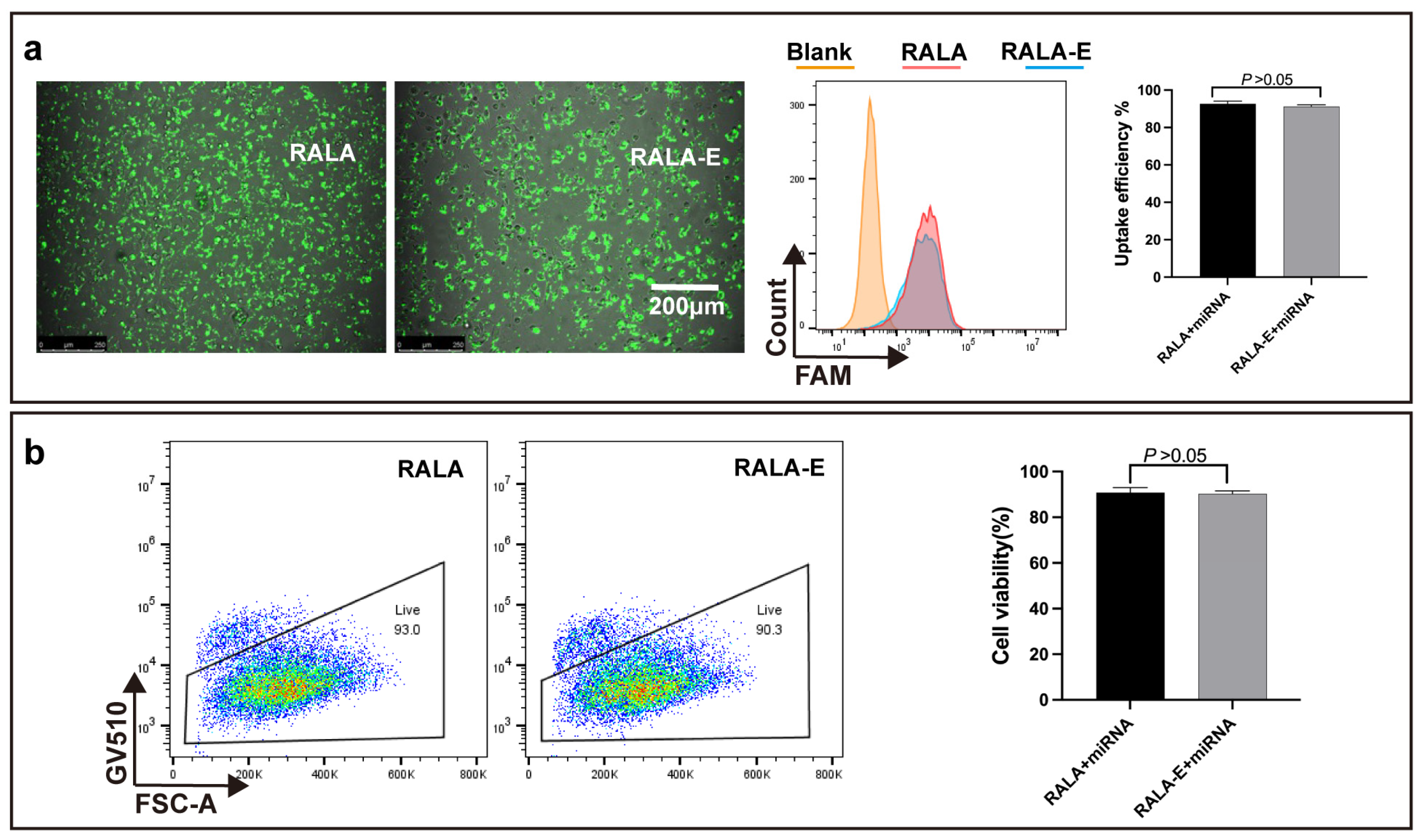
10]. In this study, RALA-E significantly improved transfection efficiency of pDNA in HEK-293T and HeLa cell lines compared to RALA, indicating that the VGVAPG modification was effective. Though the VGVAPG modification failed to enhance the uptake ability of miRNA in macrophages, this may be explained by the excellent performance of RALA for small size nucleic acid delivery [
10], leaving no much room for further improvement (up to around 90% cellular uptake%). In overall, the high transfection efficiency of RALA-E/nucleic acid complexes supported that RALA-E can be a promising gene delivery vector.
There are two main pathways for the internalization of the CPP/nucleic acid complexes into the cells, namely the endocytic or energy-dependent pathway, and the direct penetration or energy-independent pathways [
25]. By performing the transfection under 4 °C, we confirmed that the endocytosis (or energy-dependent pathway) was predominant in cellular internalization process for the complexes in both RALA and RALA-E groups. In the following, we studied the specific endocytosis mechanism of the peptide/pDNA complexes by using various inhibitors.
Clathrin-mediated, caveolae-mediated and lipid raft-mediated endocytosis are main pathways for endocytosis. Chlorpromazine is a common inhibitor for clathrin mediated endocytosis and genistein is able to inhibit caveolae mediated endocytosis [
12]. Methyl-β-cyclodextrin (mβCD) can remove cholesterol from the plasma membrane and thus inhibit several pathways, such as cholesterol-enriched microdomains, caveolae and lipid rafts. The less affected transfection efficiency of RALA-E group compared to RALA group after inhibitors treatment, is very likely related with the binding of the VGVAPG sequence to the EBD on the cell surface. The VGVAPG-EBD interaction can act as an anchor for the complex to attach to the cells, which can prolong the staying time of the complexes on the cell surface and thus may greatly facilitate the subsequent endocytosis events. Without this sequence, the attachment of the RALA/pDNA complexes on the cell surface mainly depended on electrostatic interaction and the staying time of the complexes on the cell surface would be much shorter if the complexes were not endocytosed. The mβCD showed superior inhibition effect on RALA-E group compared to other inhibitors. This is probably due to its role on the block of the interaction of VGVAPG sequence and the elastin binding protein. As discussed above, the mβCD can destroy the cholesterol in the plasma membrane. Cholesterol is the key component in the lipid raft. The removal of cholesterol would destroy the microdomains of lipid raft. Rusciani et al. reported that elastin receptor complex and the lipid rafts colocalized on the plasma membrane, and the disruption of or depletion of glycolipids in the microdomain of lipid raft can block the elastin receptor complex signaling [
26]. Therefore, the destroy of the lipid raft structure by mβCD would also affect the function or integrity of the elastin receptor complex, and subsequently hindered the binding of VGVAPG sequence on elastin binding protein which is the outer part of the elastin receptor complex.
Successful gene delivery vectors need to be able to effectively overcome the membrane barrier of cells and deliver genes to characteristic organelles to play biological functions within cells. Therefore, a desirable vector should be able to enter the nucleus in a short period of time without being degraded by lysosomal enzymes. Most of the loaded miR-146a can successfully enter the nucleus 24 h after the transfection, indicating that the modified peptide had good endosomal escape ability.
The successful modulate the target gene expression by delivered miR-146a is the ultimate goal of this study. The inhibition results on target genes showed that RALA-E/miR-146a complexes can successfully inhibit the expression of inflammatory factors of M1 macrophages, further proving the potential of RALA-E as a gene delivery vector for clinical application.
In the next step, we plan to incorporate the RALA-E/nucleic acid complexes into elastin-based hydrogels which were recently developed by our group [
27], as a biological regenerative and sustained release platform, to regulate the inflammatory microenvironment and improve the healing of chronic wounds.