Anti-Inflammatory Polymeric Nanoparticles Based on Ketoprofen and Dexamethasone
Abstract
1. Introduction
2. Materials and Methods
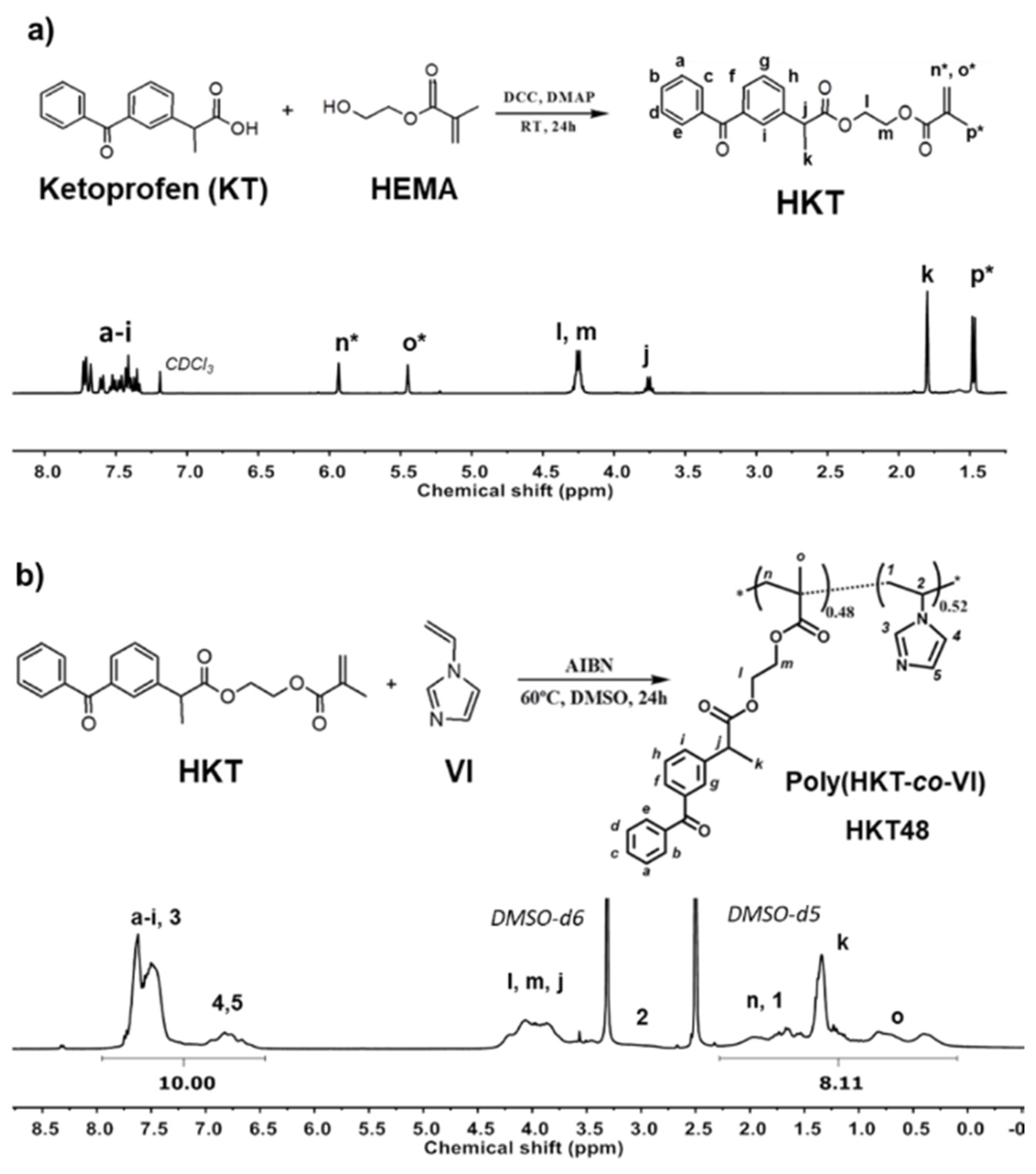
2.1. Synthesis and Characterization of Ketoprofen-Derived Monomer (HKT)
2.2. Synthesis and Characterization of Copolymer of Ketoprofen-Based Methacrylic Monomer and 1-vinyl imidazole, poly(HKT-co-VI)
2.2.1. Proton Nuclear Magnetic Resonance (1H-NMR)
2.2.2. Size Exclusion Chromatography (SEC)
2.2.3. Differential Scanning Calorimetry (DSC)
2.3. Preparation and Characterization of Self-Assembled Nanoparticles
2.3.1. Nanoprecipitation Method
2.3.2. Characterization of NPs
2.3.3. Dexamethasone and Coumarin-6 Encapsulation
2.3.4. Encapsulation Efficiency (%EE) and Loading Capacity (%LC)
2.4. Cell Culture
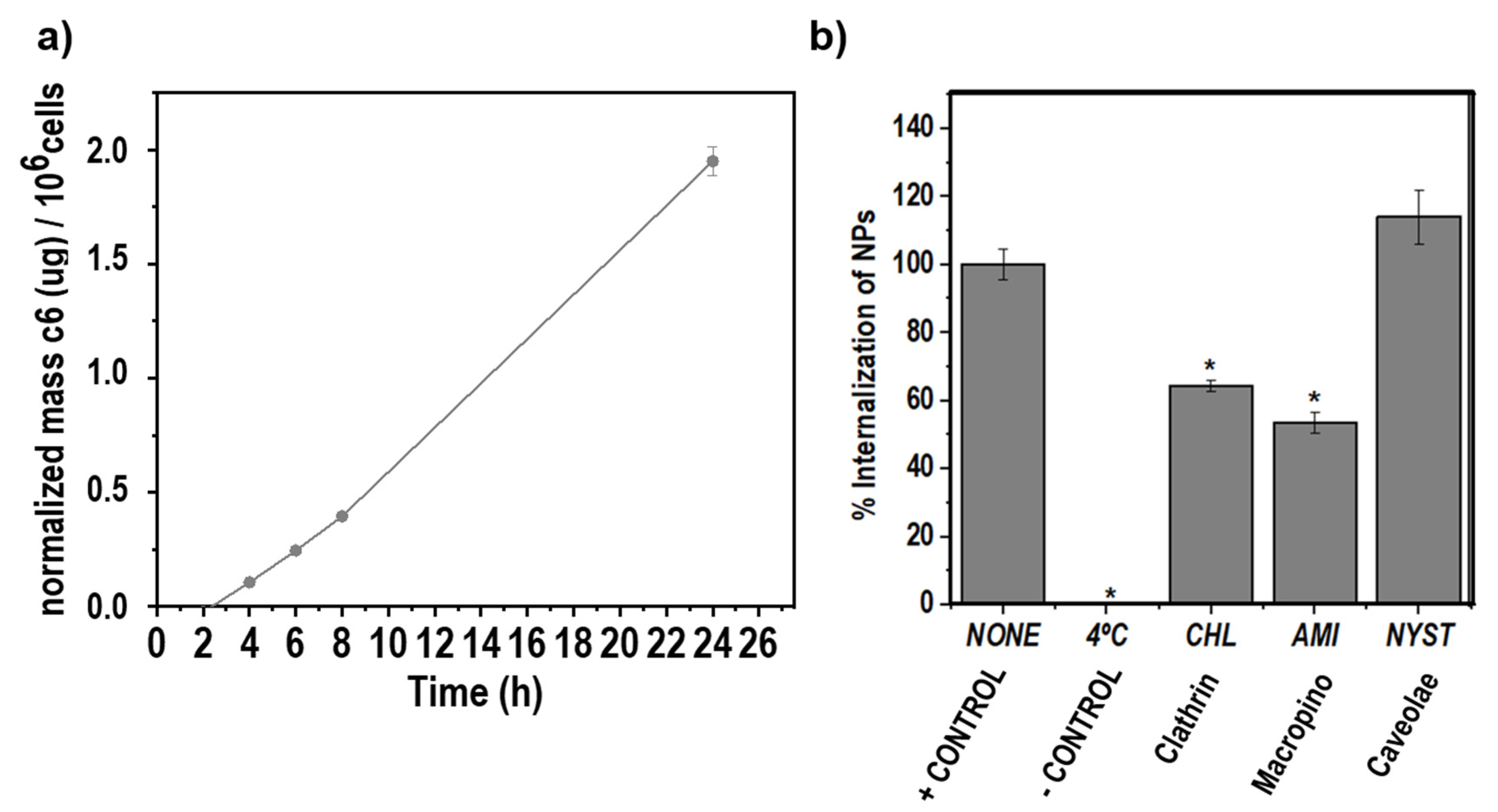
2.4.1. Uptake Rate of c6-Loaded KT NPs by RAW264.7 Macrophages
2.4.2. Route of Nanoparticle Internalization
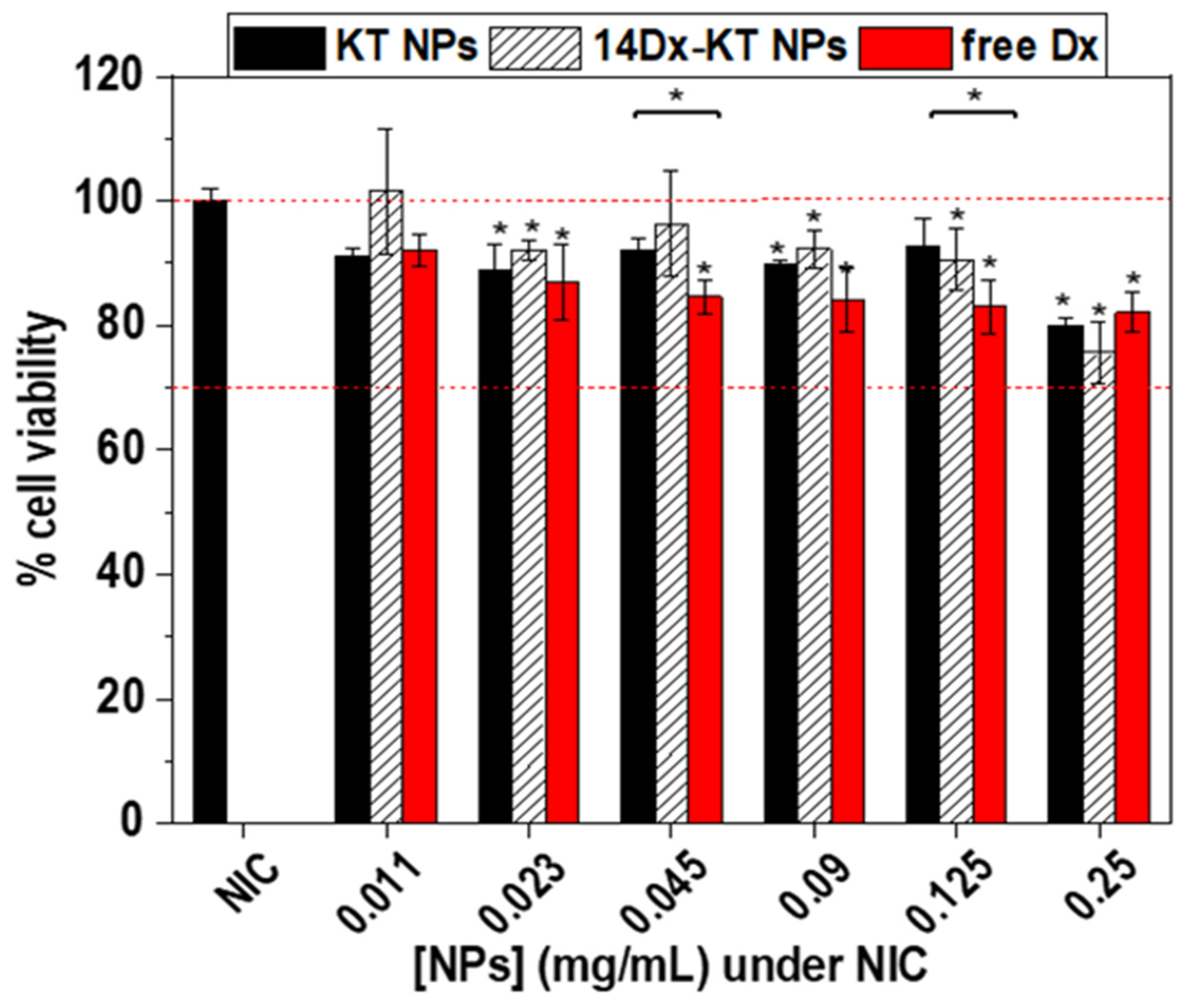
2.4.3. In Vitro Cytotoxicity Assay of NPs
2.4.4. Nitric Oxide (NO) Assay
2.4.5. RNA Extraction, Reverse Transcription, Real-Time Quantitative PCR (RT-qPCR)
3. Results and Discussion
3.1. Methacrylic Derivative of Ketoprofen Monomer (HKT)
3.2. Synthesis and Characterization of Copolymer of Ketoprofen-Based Methacrylic Monomer and 1-Vinyl Imidazole, Poly(HKT-co-VI)
3.3. Preparation and Characterization of Self-Assembled Nanoparticles
3.4. Dexamethasone and Coumarin-6 Encapsulation
3.5. Uptake Rate of c6-Loaded NPs by RAW264.7 Macrophages
3.6. In Vitro Cytotoxicity Assay of NPs
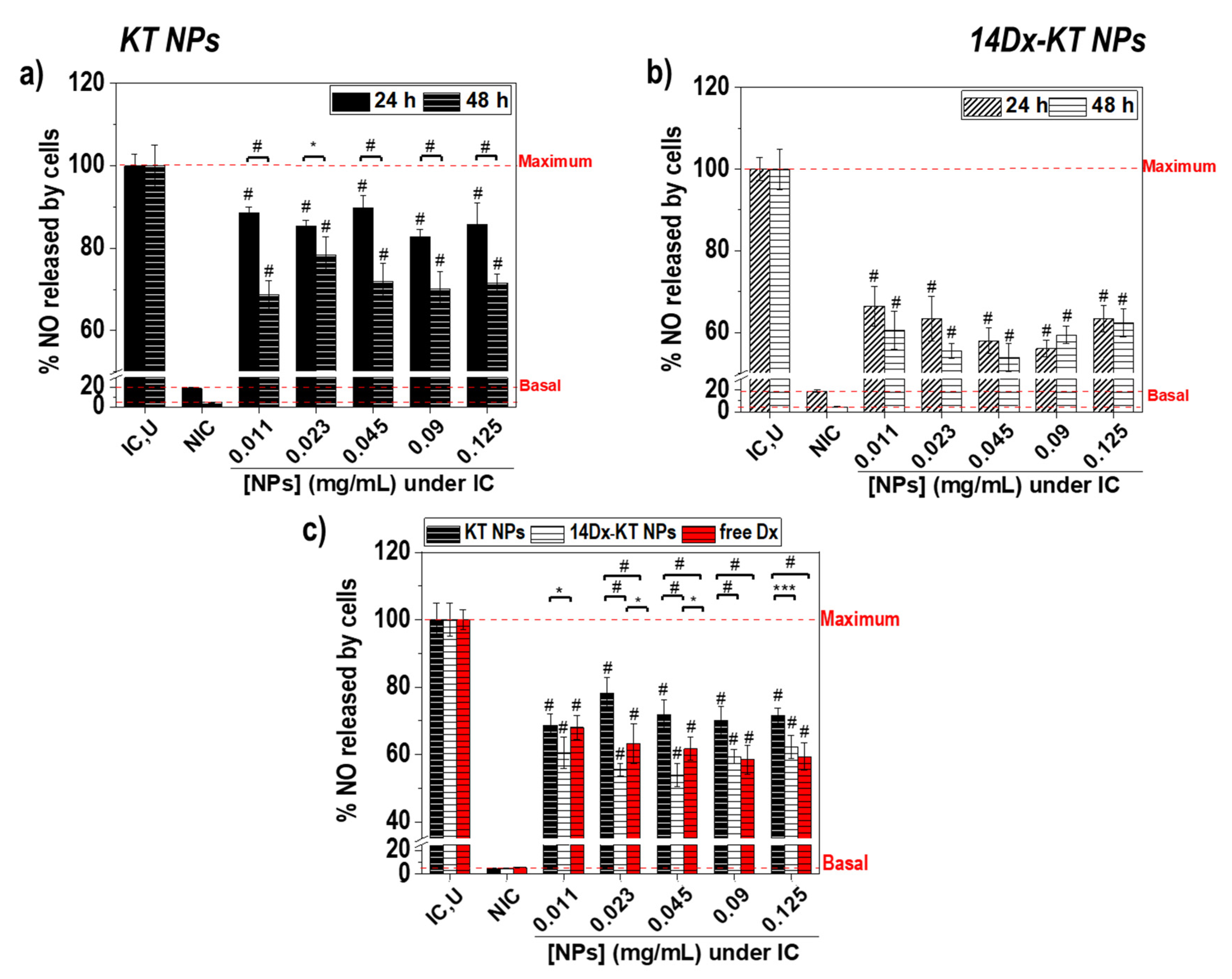
3.7. Effect of Polymeric Nanoparticles Based on Ketoprofen and Dexamethasone on Macrophage NO Levels
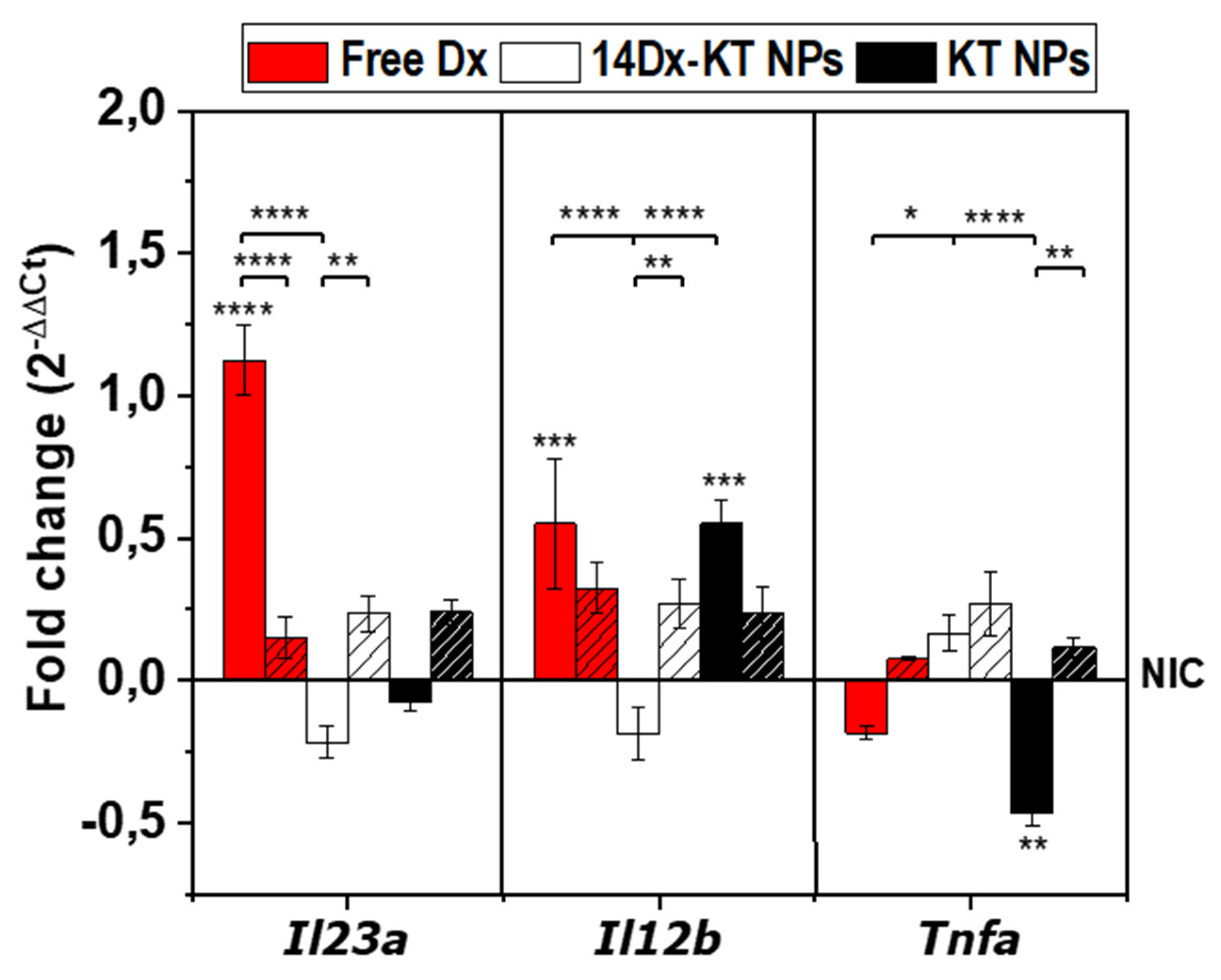
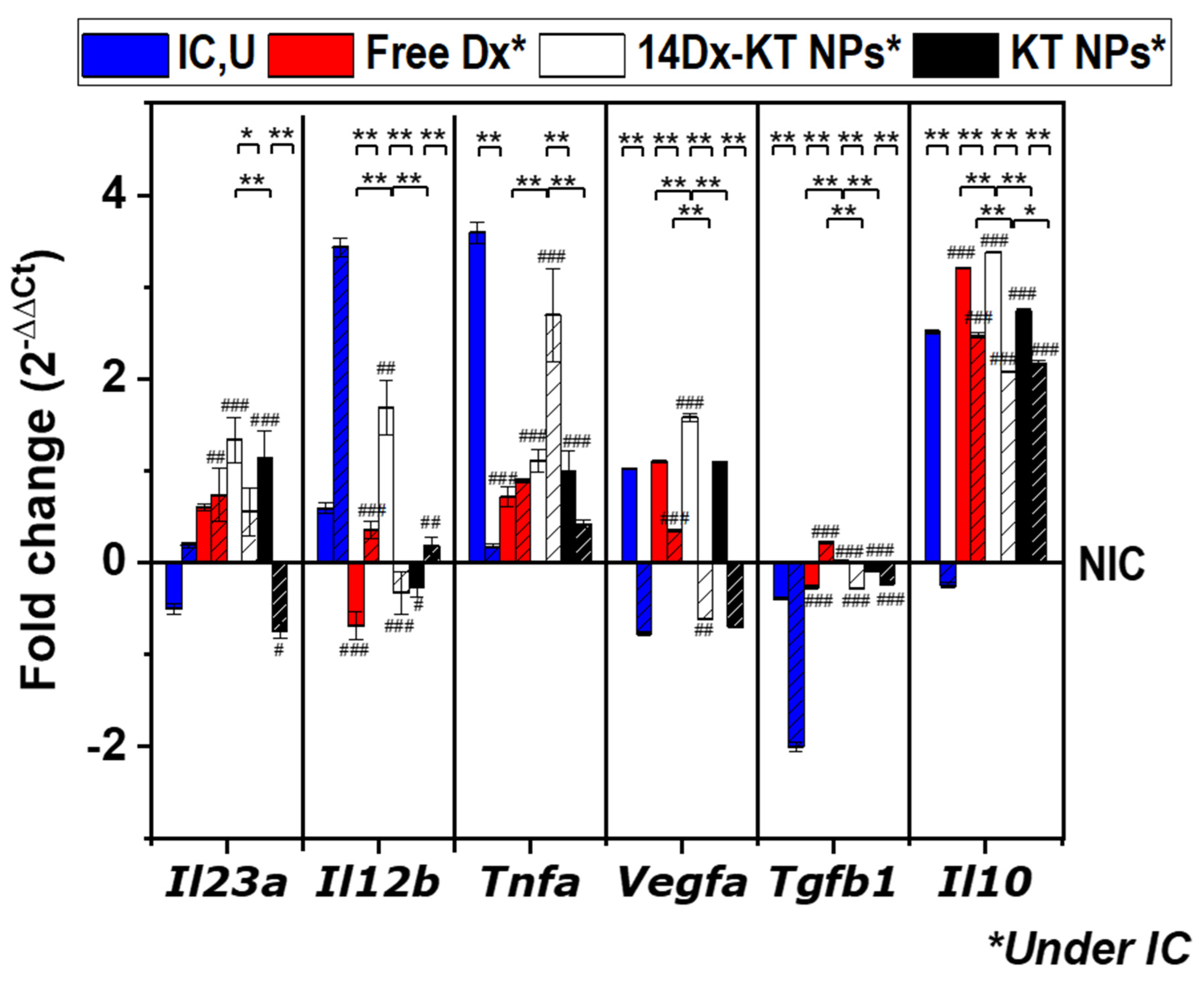
3.8. Real-Time PCR Analysis of the Expression of M1-M2 Specific Reference Genes after NPs Treatment in Non-Stimulated Macrophages and LPS-Stimulated Macrophages
4. Conclusions
Supplementary Materials
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
References
- Dinarello, C.A. Anti-inflammatory agents: Present and future. Cell 2010, 140, 935–950. [Google Scholar] [CrossRef] [PubMed]
- Lai, Y.; Dong, C. Therapeutic antibodies that target inflammatory cytokines in autoimmune diseases. Int. Immunol. 2016, 28, 181–188. [Google Scholar] [CrossRef] [PubMed]
- Toussirot, E. The IL23/Th17 pathway as a therapeutic target in chronic inflammatory diseases. Inflamm. Allergy Drug Targets 2012, 11, 159–168. [Google Scholar] [CrossRef] [PubMed]
- Pfeifle, R. Analysis of the Role of the IL-23/Th17 Axis on B Cell-Mediated Autoimmunity. Ph.D. Thesis, Friedrich–Alexander University Erlangen–Nürnberg, Erlangen, Germany, 2017. [Google Scholar]
- Ergen, E.N.; Yusuf, N. Inhibition of interleukin-12 and/or interleukin-23 for the treatment of psoriasis: What is the evidence for an effect on malignancy? Exp. Dermatol. 2018, 27, 737–747. [Google Scholar] [CrossRef] [PubMed]
- Vom Berg, J.; Prokop, S.; Miller, K.R.; Obst, J.; Kälin, R.E.; Lopategui-Cabezas, I.; Wegner, A.; Mair, F.; Schipke, C.G.; Peters, O.; et al. Inhibition of IL-12/IL-23 signaling reduces Alzheimer‘s disease-like pathology and cognitive decline. Nat. Med. 2012, 18, 1812–1819. [Google Scholar] [CrossRef] [PubMed]
- Mortezavi, M.; Ritchlin, C. IL12/IL23 Inhibition in the treatment of psoriatic arthritis. Curr. Treat. Opt. Rheumatol. 2015, 1, 197–209. [Google Scholar] [CrossRef]
- Kotze, P.G.; Ma, C.; Almutairdi, A.; Panaccione, R. Clinical utility of ustekinumab in Crohn‘s disease. J. Inflamm. Res. 2018, 11, 35–47. [Google Scholar] [CrossRef]
- Oppmann, B.; Lesley, R.; Blom, B.; Timans, J.C.; Xu, Y.; Hunte, B.; Vega, F.; Yu, N.; Wang, J.; Singh, K.; et al. Novel p19 protein engages IL-12p40 to form a cytokine, IL-23, with biological activities similar as well as distinct from IL-12. Immunity 2000, 13, 715–725. [Google Scholar] [CrossRef]
- Henrickson, S.E.; Ruffner, M.A.; Kwan, M. Unintended immunological consequences of biologic therapy. Curr. Allergy Asthma Rep. 2016, 16, 46. [Google Scholar] [CrossRef]
- Espinosa-Cano, E.; Aguilar, M.R.; Vázquez, B.; Román, J.S. Chapter 7-Inflammation-responsive polymers. In Smart Polymers and Their Applications, 2nd ed.; Aguilar, M.R., Román, J.S., Eds.; Woodhead Publishing: Cambridge, UK, 2019; pp. 219–254. [Google Scholar]
- Paik, J.H.; Ju, J.H.; Lee, J.Y.; Boudreau, M.D.; Hwang, D.H. Two opposing effects of non-steroidal anti-inflammatory drugs on the expression of the inducible cyclooxygenase. Mediation through different signaling pathways. J. Biol. Chem. 2000, 275, 28173–28179. [Google Scholar] [CrossRef]
- Alaseem, A.M.; Madiraju, P.; Aldebeyan, S.A.; Noorwali, H.; Antoniou, J.; Mwale, F. Naproxen induces type X collagen expression in human bone-marrow-derived mesenchymal stem cells through the upregulation of 5-lipoxygenase. Tissue. Eng. Part A 2015, 21, 234–245. [Google Scholar] [CrossRef] [PubMed]
- Whittaker, D.S.; Bahjat, K.S.; Moldawer, L.L.; Clare-Salzler, M.J. Autoregulation of human monocyte-derived dendritic cell maturation and IL-12 production by cyclooxygenase-2-mediated prostanoid production. J. Immunol. 2000, 165, 4298–4304. [Google Scholar] [CrossRef] [PubMed]
- Mitsuhashi, M.; Liu, J.; Cao, S.; Shi, X.; Ma, X. Regulation of interleukin-12 gene expression and its anti-tumor activities by prostaglandin E2 derived from mammary carcinomas. J. Leukoc. Biol. 2004, 76, 322–332. [Google Scholar] [CrossRef] [PubMed]
- Bedognetti, D.; Roelands, J.; Decock, J.; Wang, E.; Hendrickx, W. The MAPK hypothesis: Immune-regulatory effects of MAPK-pathway genetic dysregulations and implications for breast cancer immunotherapy. Emerg. Top. Life Sci. 2017, 1, 429–445. [Google Scholar]
- Boucher, J.G.; Parato, K.A.; Frappier, F.; Fairman, P.; Busca, A.; Saxena, M.; Blahoianu, M.A.; Ma, W.; Gajanayaka, N.; Parks, R.J.; et al. Disparate regulation of LPS-induced MAPK signaling and IL-12p40 expression between different myeloid cell types with and without HIV infection. Viral Immunol. 2010, 23, 17–28. [Google Scholar] [CrossRef]
- Burnett, B.P.; Levy, R.M. 5-Lipoxygenase metabolic contributions to NSAID-induced organ toxicity. Adv. Ther. 2012, 29, 79–98. [Google Scholar] [CrossRef]
- Brenner, R.; Ayala, S.; Garda, H. Effect of dexamethasone on the fatty acid composition of total liver microsomal lipids and phosphatidylcholine molecular species. Lipids 2002, 36, 1337–1345. [Google Scholar] [CrossRef]
- Ma, W.; Gee, K.; Lim, W.; Chambers, K.; Angel, J.B.; Kozlowski, M.; Kumar, A. Dexamethasone inhibits IL-12p40 production in lipopolysaccharide-stimulated human monocytic cells by down-regulating the activity of c-Jun N-Terminal Kinase, the activation Protein-1, and NF-κB transcription factors. J. Immunol. 2004, 172, 318–330. [Google Scholar] [CrossRef]
- Botting, R.M. Inhibitors of cyclooxygenases: Mechanisms, selectivity and uses. J. Physiol. Pharmacol. 2006, 57, 113–124. [Google Scholar]
- Rao, P.; Knaus, E.E. Evolution of nonsteroidal anti-inflammatory drugs (NSAIDs): Cyclooxygenase (COX) inhibition and beyond. J. Pharm. Pharm. Sci. 2008, 11, 81–110. [Google Scholar] [CrossRef]
- Bareford, L.M.; Swaan, P.W. Endocytic mechanisms for targeted drug delivery. Adv. Drug Deliv. Rev. 2007, 59, 748–758. [Google Scholar] [CrossRef] [PubMed]
- Al-Khoury, H.; Espinosa-Cano, E.; Aguilar, M.R.; Roman, J.S.; Syrowatka, F.; Schmidt, G.; Groth, T. Anti-inflammatory surface coatings based on polyelectrolyte multilayers of heparin and polycationic nanoparticles of naproxen-bearing polymeric drugs. Biomacromolecules 2019, 20, 4015–4025. [Google Scholar] [CrossRef] [PubMed]
- Suarez, P.; Rojo, L.; González-Gómez, Á.; Román, J.S. Self-Assembling gradient copolymers of vinylimidazol and (Acrylic)ibuprofen with anti-inflammatory and zinc chelating properties. Macromol. Biosci. 2013, 13, 1174–1184. [Google Scholar] [CrossRef]
- Srivastava, A.; Waite, J.H.; Stucky, G.D.; Mikhailovsky, A. Fluorescence investigations into complex coacervation between polyvinylimidazole and sodium alginate. Macromolecules 2009, 42, 2168–2176. [Google Scholar] [CrossRef] [PubMed][Green Version]
- Livak, K.J.; Schmittgen, T.D. Analysis of relative gene expression data using real-time quantitative PCR and the 2(-Delta Delta C(T)) Method. Methods 2001, 25, 402–408. [Google Scholar] [CrossRef]
- DiCiccio, J.E.; Steinberg, B.E. Lysosomal pH and analysis of the counter ion pathways that support acidification. J. Gen. Physiol. 2011, 137, 385–390. [Google Scholar] [CrossRef]
- d’Arcy, R.; Tirelli, N. Fishing for fire: Strategies for biological targeting and criteria for material design in anti-inflammatory therapies. Polym. Adv. Technol. 2014, 25, 478–498. [Google Scholar] [CrossRef]
- Bhattacharjee, S. DLS and zeta potential–What they are and what they are not? J. Control. Release 2016, 235, 337–351. [Google Scholar] [CrossRef]
- Cheng, C.J.; Tietjen, G.T.; Saucier-Sawyer, J.K.; Saltzman, W.M. A holistic approach to targeting disease with polymeric nanoparticles. Nat. Rev. Drug Discov. 2015, 14, 239–247. [Google Scholar] [CrossRef]
- Sadat, S.M.A.; Jahan, T.; Haddadi, A. effects of size and surface charge of polymeric nanoparticles on “in vitro” and “in vivo” applications. J. Biomater. Nanobiotechnol. 2016, 7, 91–108. [Google Scholar] [CrossRef]
- Pillay, V.; Murugan, K.; Choonara, Y.E.; Kumar, P.; Bijukumar, D.; Du Toit, L.C. Parameters and characteristics governing cellular internalization and trans-barrier trafficking of nanostructures. Int. J. Nanomed. 2015, 10, 2191–2206. [Google Scholar] [CrossRef] [PubMed]
- Banik, B.L.; Fattahi, P.; Brown, J.L. Polymeric nanoparticles: The future of nanomedicine. Wiley Interdiscip. Rev. Nanomed. Nanobiotechnol. 2016, 8, 271–299. [Google Scholar] [CrossRef] [PubMed]
- Kou, L.; Sun, J.; Zhai, Y.; He, Z. The endocytosis and intracellular fate of nanomedicines: Implication for rational design. Asian J. Pharm. Sci. 2013, 8, 1–10. [Google Scholar] [CrossRef]
- Vangara, K.K.; Liu, J.L.; Palakurthi, S. Hyaluronic acid-decorated PLGA-PEG nanoparticles for targeted delivery of SN-38 to ovarian cancer. Anticancer. Res. 2013, 33, 2425–2434. [Google Scholar]
- Yameen, B.; Choi, W.I.; Vilos, C.; Swami, A.; Shi, J.; Farokhzad, O.C. Insight into nanoparticle cellular uptake and intracellular targeting. J. Control. Release 2014, 190, 485–499. [Google Scholar] [CrossRef]
- Sahin, M.; Arslan, C.; Nazıroğlu, M.; Tunç, E.; Demirci, M.; Sutcu, R.; Yilmaz, N. Asymmetric dimethylarginine and nitric oxide levels as signs of endothelial dysfunction in behcet’s disease. Ann. Clin. Lab. Sci. 2006, 36, 449–454. [Google Scholar]
- Reichel, D.; Tripathi, M.; Perez, J.M. Biological effects of nanoparticles on macrophage polarization in the tumor microenvironment. Nanotheranostics 2019, 3, 66–88. [Google Scholar] [CrossRef]
- Mulén, B.; Rojas, J.M.; Pérez-Yagüe, S.; Morales, M.P.; Barber, D.F. Polyethylenimine-coated SPIONs trigger macrophage activation through TLR-4 signaling and ROS production and modulate podosome dynamics. Biomaterials 2015, 52, 494–506. [Google Scholar]
- Ghezzi, P.; Sacco, S.; Agnello, D.; Marullo, A.; Caselli, G.F.; Bertini, R. Lps induces IL-6 in the brain and in serum largely through TNF production. Cytokine 2000, 12, 1205–1210. [Google Scholar] [CrossRef]
- Zakharova, M.; Ziegler, H.K. Paradoxical Anti-Inflammatory actions of TNF-α: Inhibition of IL-12 and IL-23 via TNF receptor 1 in macrophages and dendritic cells. J. Immunol. 2005, 175, 5024–5033. [Google Scholar] [CrossRef]
- Shachar, I.; Karin, N. The dual roles of inflammatory cytokines and chemokines in the regulation of autoimmune diseases and their clinical implications. J. Leukoc. Biol. 2013, 93, 51–61. [Google Scholar] [CrossRef] [PubMed]
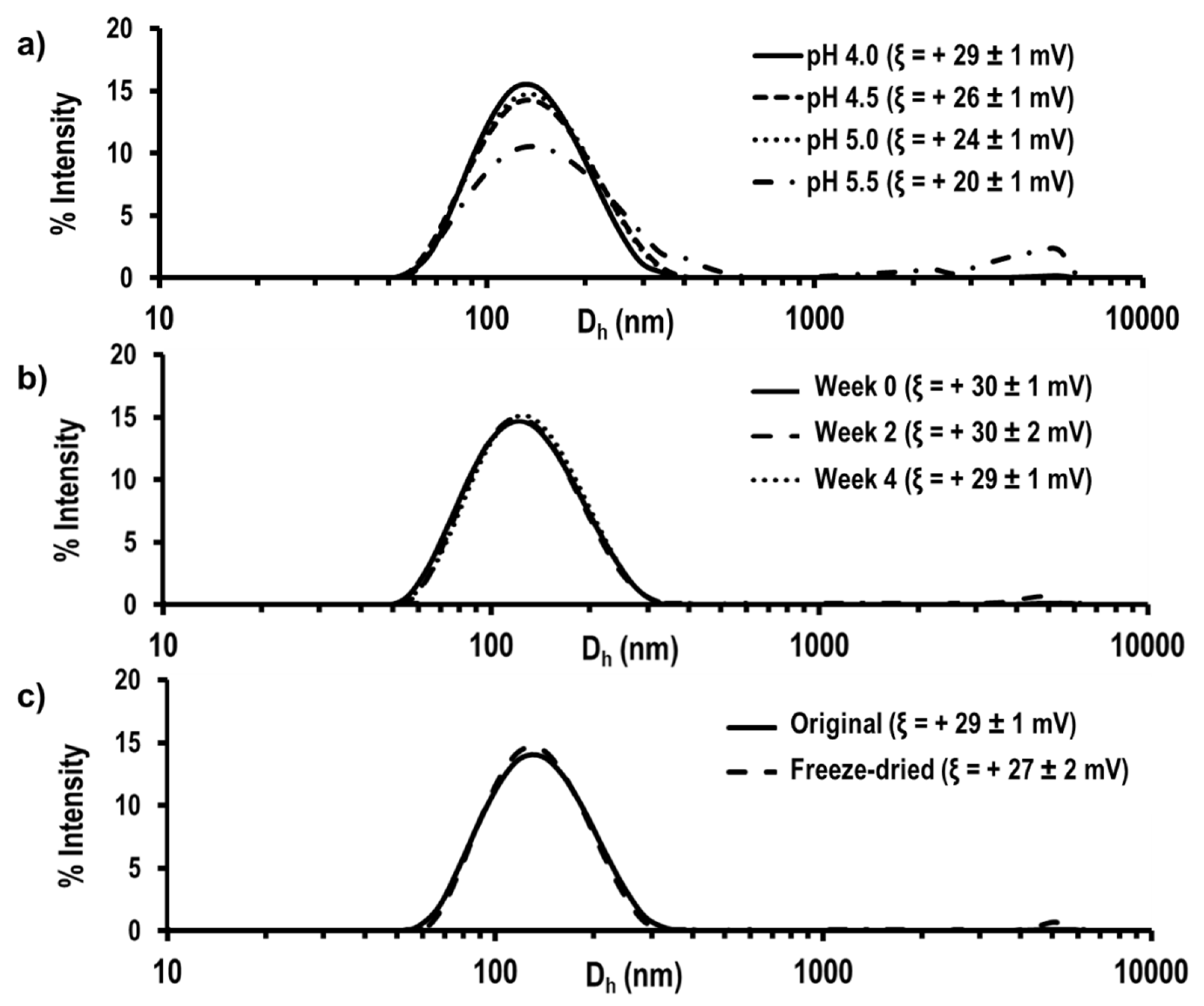
| [NPs]A.P. a | VF (mL) | [HKT48]O.P. b | Dh c (nm) | PdI d | ξ e (mV) |
|---|---|---|---|---|---|
| 1.0 mg/mL | 10 | 10 mg/mL | 117 ± 1 | 0.139 ± 0.004 | +29 ± 1 |
| 20 | 10 mg/mL | 119 ± 3 | 0.153 ± 0.018 | +29 ± 1 | |
| 30 | 10 mg/mL | 116 ± 1 | 0.142 ± 0.027 | +30 ± 1 | |
| 5.0 mg/mL | 10 | 10 mg/mL | 115 ± 1 | 0.124 ± 0.008 | +30 ± 1 |
| 20 | 10 mg/mL | 119 ± 1 | 0.112 ± 0.013 | +31 ± 1 | |
| 30 | 15 mg/mL | 163 ± 3 | 0.186 ± 0.021 | +29 ± 1 |
| Sample | % EE a | % LC b | Dh c (nm) | PdI d | ξ e (mV) |
|---|---|---|---|---|---|
| KT NPs | - | - | 117 ± 1 | 0.139 ± 0.004 | +30 ± 1 |
| 14Dx-KT NPs | 14 | 3.85 | 140 ± 1 | 0.081 ± 0.010 | +29 ± 1 |
| 47c6-KT NPs | 47 | 0.47 | 126 ± 2 | 0.154 ± 0.021 | +32 ± 2 |
© 2020 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Espinosa-Cano, E.; Aguilar, M.R.; Portilla, Y.; Barber, D.F.; San Román, J. Anti-Inflammatory Polymeric Nanoparticles Based on Ketoprofen and Dexamethasone. Pharmaceutics 2020, 12, 723. https://doi.org/10.3390/pharmaceutics12080723
Espinosa-Cano E, Aguilar MR, Portilla Y, Barber DF, San Román J. Anti-Inflammatory Polymeric Nanoparticles Based on Ketoprofen and Dexamethasone. Pharmaceutics. 2020; 12(8):723. https://doi.org/10.3390/pharmaceutics12080723
Chicago/Turabian StyleEspinosa-Cano, Eva, Maria Rosa Aguilar, Yadileiny Portilla, Domingo F. Barber, and Julio San Román. 2020. "Anti-Inflammatory Polymeric Nanoparticles Based on Ketoprofen and Dexamethasone" Pharmaceutics 12, no. 8: 723. https://doi.org/10.3390/pharmaceutics12080723
APA StyleEspinosa-Cano, E., Aguilar, M. R., Portilla, Y., Barber, D. F., & San Román, J. (2020). Anti-Inflammatory Polymeric Nanoparticles Based on Ketoprofen and Dexamethasone. Pharmaceutics, 12(8), 723. https://doi.org/10.3390/pharmaceutics12080723






