Nucleic Acid Hybrids as Advanced Antibacterial Nanocarriers
Abstract
1. Introduction
2. DNA Nanocarriers
2.1. Principles for Fabrication
2.2. From Test Tube to Biological Applications: Current Challenges
3. Nucleic Acid Hybrid Nanocarriers
3.1. Fabrication with Organic Materials
3.2. Fabrication with Inorganic Materials
4. Nucleic Acid Hybrid Nanocarriers as Advanced Systems against Bacterial Infections
4.1. Nucleic Acid Hybrids with Enhanced Cellular Uptake
4.2. Nucleic Acid Hybrids to Control or Sustain Delivery of Antimicrobials
4.3. Nucleic Acid Hybrid with Enhanced Binding to Bacteria Cells
5. Outlook
- Non-specificity: encouraging specific interactions, such as aptamer-driven interactions with the hybrids, can be pursued.
- Rapid drug release: strategies that sustain the release of antimicrobial cargos, such as the use of liposomal vesicles and other coating methods, can be used.
- Stability concerns: towards clinical translation, future investigation of the antibacterial stability of the hybrids under conditions relevant to infections and translating these using in vivo conditions is important.
- Long-term storage: perceiving the integrity of the hybrids as solution or in their lyophilized form is a challenge, yet necessary to better understand the long-term storage of the hybrids.
- Complex formation: many antimicrobial agents are cationic/amphiphilic in nature, so complex formation may cause structural distortion of the nanomaterial. Co-delivery systems using multiple loading approaches can be pursued.
- Limited on loading higher concentrations of cationic cargos: high drug loading is challenged directly by the formation of complexes, which may significantly reduce the potency of the bound drug. Expanding to incorporate multiple antimicrobial agents or loading antimicrobial drugs into hybrid nanostructures with inherent antimicrobial activity provides room to achieve enhanced potency by addressing multiple targets.
Funding
Conflicts of Interest
References
- Ruddaraju, L.K.; Pammi, S.V.N.; Guntuku, G.S.; Padavala, V.S.; Kolapalli, V.R.M. A review on anti-bacterials to combat resistance: From ancient era of plants and metals to present and future perspectives of green nano technological combinations. Asian J. Pharm. Sci. 2020, 15, 42–59. [Google Scholar] [CrossRef] [PubMed]
- WHO. Antimicrobial Resistance, Global Report on Surveillance 2014; WHO: Geneva, Switzerland, 2014; p. 257. [Google Scholar]
- Interagency Coordination Group on Antimicrobial Resistance (IAGC). No Time to Wait: Securing the Future from Drug Resistant Infections. Available online: https://www.who.int/antimicrobial-resistance/interagency-coordination-group/IACG_final_report_EN.pdf?ua=1 (accessed on 19 May 2020).
- Cassini, A.; Hogberg, L.D.; Plachouras, D.; Quattrocchi, A.; Hoxha, A.; Simonsen, G.S.; Colomb-Cotinat, M.; Kretzschmar, M.E.; Devleesschauwer, B.; Cecchini, M.; et al. Attributable deaths and disability-adjusted life-years caused by infections with antibiotic-resistant bacteria in the EU and the European Economic Area in 2015: A population-level modelling analysis. Lancet Infect. Dis. 2019, 19, 56–66. [Google Scholar] [CrossRef]
- European Commission. A European One Health Action Plan against Antimicrobial Resistance (AMR). 2017. Available online: https://ec.europa.eu/health/amr/ (accessed on 19 May 2020).
- Podolsky, S.H. The evolving response to antibiotic resistance (1945–2018). Palgrave Commun. 2018, 4, 1–8. [Google Scholar] [CrossRef]
- Michael, C.A.; Dominey-Howes, D.; Labbate, M. The antimicrobial resistance crisis: Causes, consequences, and management. Front. Public Health 2014, 2, 145. [Google Scholar] [CrossRef]
- He, Y.; Yuan, Q.; Mathieu, J.; Stadler, L.; Senehi, N.; Sun, R.; Alvarez, P.J. Alvarez Antibiotic resistance genes from livestock waste: Occurrence, dissemination, and treatment. npj Clean Water 2020, 3, 1–11. [Google Scholar] [CrossRef]
- Santajit, S.; Indrawattana, N. Mechanisms of Antimicrobial Resistance in ESKAPE Pathogens. Biomed. Res. Int. 2016, 2016, 2475067. [Google Scholar] [CrossRef]
- Marturano, J.E.; Lowery, T.J. ESKAPE Pathogens in Bloodstream Infections Are Associated With Higher Cost and Mortality but Can Be Predicted Using Diagnoses Upon Admission. Open Forum Infect. Dis. 2019, 6, ofz503. [Google Scholar] [CrossRef]
- Anon. WHO Publishes List of Bacteria for Which New Antibiotics are Urgently Needed. 2017. Available online: https://www.who.int/news-room/detail/27-02-2017-who-publishes-list-of-bacteria-for-which-new-antibiotics-are-urgently-needed (accessed on 19 May 2020).
- WHO. Prioritization of Pathogens to Guide Discovery, Research and Development of New Antibiotics for Drug-Resistant Bacterial Infections, Including Tuberculosis. 2017. Available online: https://www.who.int/medicines/areas/rational_use/PPLreport_2017_09_19.pdf?ua=1 (accessed on 19 May 2020).
- 2019 Antibacterial Agents in Clinical Development an Analysis of the Antibacterial Clinical Development Pipeline. 2019. Available online: https://apps.who.int/iris/bitstream/handle/10665/330420/9789240000193-eng.pdf (accessed on 19 May 2020).
- WHO. Antibacterial Agents in Preclinical Development an Open Access Database. 2019. Available online: https://apps.who.int/iris/bitstream/handle/10665/330290/WHO-EMP-IAU-2019.12-eng.pdf (accessed on 19 May 2020).
- Anon. Lack of New Antibiotics Threatens Global Efforts to Contain Drug-Resistant Infections. 2020. Available online: https://www.who.int/news-room/detail/17-01-2020-lack-of-new-antibiotics-threatens-global-efforts-to-contain-drug-resistant-infections (accessed on 19 May 2020).
- Abed, N.; Couvreur, P. Nanocarriers for antibiotics: A promising solution to treat intracellular bacterial infections. Int. J. Antimicrob. Agents 2014, 43, 485–496. [Google Scholar] [CrossRef]
- Chen, H.; Jin, Y.; Wang, J.; Wang, Y.; Jiang, W.; Dai, H.; Pang, S.; Lei, L.; Ji, J.; Wang, B. Design of smart targeted and responsive drug delivery systems with enhanced antibacterial properties. Nanoscale 2018, 10, 20946–20962. [Google Scholar] [CrossRef]
- Hu, Q.; Li, H.; Wang, L.; Gu, H.; Fan, C. DNA Nanotechnology-Enabled Drug Delivery Systems. Chem. Rev. 2019, 119, 6459–6506. [Google Scholar] [CrossRef]
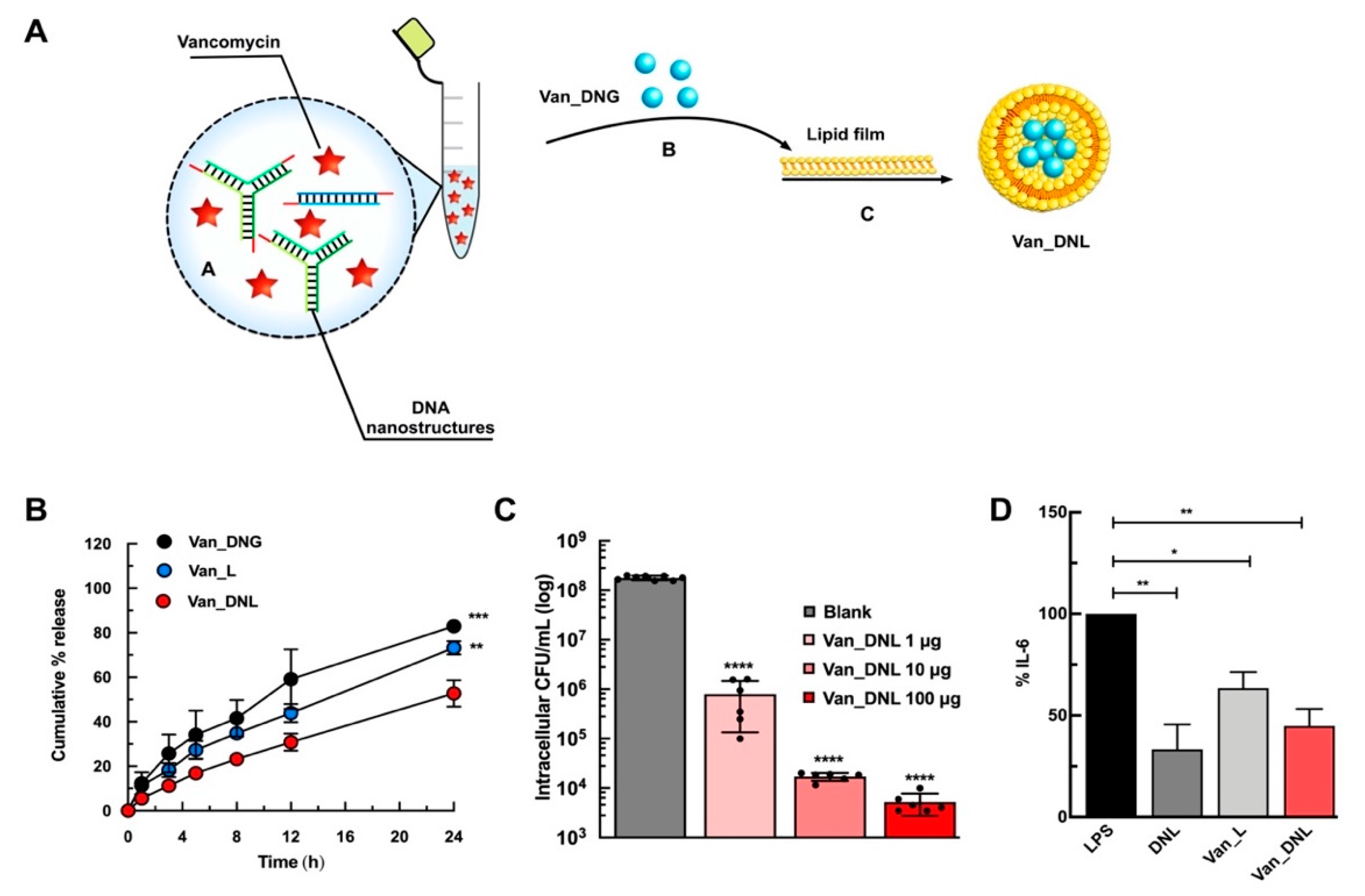
- Obuobi, S.; Julin, K.; Fredheim, E.G.; Johannessen, M.; Škalko-Basnet, N. Liposomal delivery of antibiotic loaded nucleic acid nanogels with enhanced drug loading and synergistic anti-inflammatory activity against S. aureus intracellular infections. J. Control. Release 2020. [Google Scholar] [CrossRef] [PubMed]
- Schlomann, B.H.; Wiles, T.J.; Wall, E.S.; Guillemin, K.; Parthasarathy, R. Sublethal antibiotics collapse gut bacterial populations by enhancing aggregation and expulsion. Proc. Natl. Acad. Sci. USA 2019, 116, 21392–21400. [Google Scholar] [CrossRef]
- Andersson, D.I.; Hughes, D. Microbiological effects of sublethal levels of antibiotics. Nat. Rev. Microbiol. 2014, 12, 465–478. [Google Scholar] [CrossRef] [PubMed]
- Gonzalez Gomez, A.; Syed, S.; Marshall, K.; Hosseinidoust, Z. Liposomal Nanovesicles for Efficient Encapsulation of Staphylococcal Antibiotics. ACS Omega 2019, 4, 10866–10876. [Google Scholar] [CrossRef] [PubMed]
- Chen, M.; Xie, S.; Wei, J.; Song, X.; Ding, Z.; Li, X. Antibacterial Micelles with Vancomycin-Mediated Targeting and pH/Lipase-Triggered Release of Antibiotics. ACS Appl. Mater. Interfaces 2018, 10, 36814–36823. [Google Scholar] [CrossRef] [PubMed]
- Lee, B.; Park, J.; Ryu, M.; Kim, S.; Joo, M.; Yeom, J.H.; Kim, S.; Park, Y.; Lee, K.; Bae, J. Antimicrobial peptide-loaded gold nanoparticle-DNA aptamer conjugates as highly effective antibacterial therapeutics against Vibrio vulnificus. Sci. Rep. 2017, 7, 13572. [Google Scholar] [CrossRef]
- Ghaeini-Hesaroeiye, S.; Boddohi, S.; Vasheghani-Farahani, E. Dual responsive chondroitin sulfate based nanogel for antimicrobial peptide delivery. Int. J. Biol. Macromol. 2020, 143, 297–304. [Google Scholar] [CrossRef]
- Sercombe, L.; Veerati, T.; Moheimani, F.; Wu, S.Y.; Sood, A.K.; Hua, S. Advances and Challenges of Liposome Assisted Drug Delivery. Front. Pharm. 2015, 6, 286. [Google Scholar] [CrossRef]
- Santos, R.S.; Figueiredo, C.; Azevedo, N.F.; Braeckmans, K.; De Smedt, S.C. Nanomaterials and molecular transporters to overcome the bacterial envelope barrier: Towards advanced delivery of antibiotics. Adv. Drug Deliv. Rev. 2018, 136, 28–48. [Google Scholar] [CrossRef]
- Anderson, S.D.; Gwenin, V.V.; Gwenin, C.D. Magnetic Functionalized Nanoparticles for Biomedical, Drug Delivery and Imaging Applications. Nanoscale Res. Lett. 2019, 14, 188. [Google Scholar] [CrossRef]
- Belteky, P.; Ronavari, A.; Igaz, N.; Szerencses, B.; Toth, I.Y.; Pfeiffer, I.; Kiricsi, M.; Konya, Z. Silver nanoparticles: Aggregation behavior in biorelevant conditions and its impact on biological activity. Int. J. Nanomed. 2019, 14, 667–687. [Google Scholar] [CrossRef] [PubMed]
- Deng, J.; Gao, C. Recent advances in interactions of designed nanoparticles and cells with respect to cellular uptake, intracellular fate, degradation and cytotoxicity. Nanotechnology 2016, 27, 412002. [Google Scholar] [CrossRef] [PubMed]
- Grabowski, N.; Hillaireau, H.; Vergnaud-Gauduchon, J.; Nicolas, V.; Tsapis, N.; Kerdine-Romer, S.; Fattal, E. Surface-Modified Biodegradable Nanoparticles’ Impact on Cytotoxicity and Inflammation Response on a Co-Culture of Lung Epithelial Cells and Human-Like Macrophages. J. Biomed. Nanotechnol. 2016, 12, 135–146. [Google Scholar] [CrossRef] [PubMed]
- Obuobi, S.; Wang, Y.; Khara, J.S.; Riegger, A.; Kuan, S.L.; Ee, P.L.R. Antimicrobial and Anti-Biofilm Activities of Surface Engineered Polycationic Albumin Nanoparticles with Reduced Hemolytic Activity. Macromol. Biosci. 2018, 18, e1800196. [Google Scholar] [CrossRef] [PubMed]
- Afrasiabi, S.; Pourhajibagher, M.; Raoofian, R.; Tabarzad, M.; Bahador, A. Therapeutic applications of nucleic acid aptamers in microbial infections. J. Biomed. Sci. 2020, 27, 6. [Google Scholar] [CrossRef]
- Navya, P.N.; Kaphle, A.; Srinivas, S.P.; Bhargava, S.K.; Rotello, V.M.; Daima, H.K. Current trends and challenges in cancer management and therapy using designer nanomaterials. Nano Converg. 2019, 6, 23. [Google Scholar] [CrossRef]
- Zhang, Q.; Jiang, Q.; Li, N.; Dai, L.; Liu, Q.; Song, L.; Wang, J.; Li, Y.; Tian, J.; Ding, B.; et al. DNA origami as an in vivo drug delivery vehicle for cancer therapy. ACS Nano 2014, 8, 6633–6643. [Google Scholar] [CrossRef]
- Chang, M.; Yang, C.S.; Huang, D.M. Aptamer-conjugated DNA icosahedral nanoparticles as a carrier of doxorubicin for cancer therapy. ACS Nano 2011, 5, 6156–6163. [Google Scholar] [CrossRef]
- Jackson, T.L. Intracellular accumulation and mechanism of action of doxorubicin in a spatio-temporal tumor model. J. Biol. 2003, 220, 201–213. [Google Scholar] [CrossRef]
- Mathur, D.; Medintz, I.L. The Growing Development of DNA Nanostructures for Potential Healthcare-Related Applications. Adv. Healthc Mater. 2019, 8, e1801546. [Google Scholar] [CrossRef]
- Seeman, N.C. Nucleic acid junctions and lattices. J. Biol. 1982, 99, 237–247. [Google Scholar] [CrossRef]
- Seeman, N.C.; Sleiman, H.F. DNA nanotechnology. Nat. Rev. Mater. 2018, 3. [Google Scholar] [CrossRef]
- Linko, V.; Ora, A.; Kostiainen, M.A. DNA Nanostructures as Smart Drug-Delivery Vehicles and Molecular Devices. Trends Biotechnol. 2015, 33, 586–594. [Google Scholar] [CrossRef]
- Rothemund, P.W. Folding DNA to create nanoscale shapes and patterns. Nature 2006, 440, 297–302. [Google Scholar] [CrossRef] [PubMed]
- Wang, W.; Chen, S.; An, B.; Huang, K.; Bai, T.; Xu, M.; Bellot, G.; Ke, Y.; Xiang, Y.; Wei, B. Complex wireframe DNA nanostructures from simple building blocks. Nat. Commun. 2019, 10, 1067. [Google Scholar] [CrossRef]
- Bujold, K.E.; Lacroix, A.; Sleiman, H.F. DNA Nanostructures at the Interface with Biology. Chem.-Us 2018, 4, 495–521. [Google Scholar] [CrossRef]
- Green, C.M.; Mathur, D.; Medintz, I.L. Understanding the fate of DNA nanostructures inside the cell. J. Mater. Chem. B 2020. [Google Scholar] [CrossRef] [PubMed]
- Walsh, A.S.; Yin, H.; Erben, C.M.; Wood, M.J.; Turberfield, A.J. DNA cage delivery to mammalian cells. ACS Nano 2011, 5, 5427–5432. [Google Scholar] [CrossRef]
- Rosi, N.L.; Giljohann, D.A.; Thaxton, C.S.; Lytton-Jean, A.K.; Han, M.S.; Mirkin, C.A. Oligonucleotide-modified gold nanoparticles for intracellular gene regulation. Science 2006, 312, 1027–1030. [Google Scholar] [CrossRef]
- Lacroix, A.; Vengut-Climent, E.; de Rochambeau, D.; Sleiman, H.F. Uptake and Fate of Fluorescently Labeled DNA Nanostructures in Cellular Environments: A Cautionary Tale. ACS Cent. Sci. 2019, 5, 882–891. [Google Scholar] [CrossRef]
- Rhee, W.J.; Bao, G. Slow non-specific accumulation of 2’-deoxy and 2’-O-methyl oligonucleotide probes at mitochondria in live cells. Nucleic Acids Res. 2010, 38, e109. [Google Scholar] [CrossRef] [PubMed]
- Kim, K.R.; Kang, S.J.; Lee, A.Y.; Hwang, D.; Park, M.; Park, H.; Kim, S.; Hur, K.; Chung, H.S.; Mao, C.; et al. Highly tumor-specific DNA nanostructures discovered by in vivo screening of a nucleic acid cage library and their applications in tumor-targeted drug delivery. Biomaterials 2019, 195, 1–12. [Google Scholar] [CrossRef] [PubMed]
- Anastassacos, F.M.; Zhao, Z.; Zeng, Y.; Shih, W.M. Glutaraldehyde Cross-Linking of Oligolysines Coating DNA Origami Greatly Reduces Susceptibility to Nuclease Degradation. J. Am. Chem. Soc. 2020, 142, 3311–3315. [Google Scholar] [CrossRef] [PubMed]
- Stephanopoulos, N. Strategies for Stabilizing DNA Nanostructures to Biological Conditions. Chembiochem 2019, 20, 2191–2197. [Google Scholar] [CrossRef]
- Ramakrishnan, S.; Ijas, H.; Linko, V.; Keller, A. Structural stability of DNA origami nanostructures under application-specific conditions. Comput. Struct. Biotechnol. J. 2018, 16, 342–349. [Google Scholar] [CrossRef]
- Chien, M.P.; Thompson, M.P.; Gianneschi, N.C. DNA-nanoparticle micelles as supramolecular fluorogenic substrates enabling catalytic signal amplification and detection by DNAzyme probes. Chem. Commun. (Camb.) 2011, 47, 167–169. [Google Scholar] [CrossRef]
- Li, Z.; Zhang, Y.; Fullhart, P.; Mirkin, C.A. Reversible and Chemically Programmable Micelle Assembly with DNA Block-Copolymer Amphiphiles. Nano Lett. 2004, 4, 1055–1058. [Google Scholar] [CrossRef]
- Alemdaroglu, F.E.; Alemdaroglu, N.C.; Langguth, P.; Herrmann, A. DNA Block Copolymer Micelles—A Combinatorial Tool for Cancer Nanotechnology. Adv. Mater. 2008, 20, 899–902. [Google Scholar] [CrossRef]
- Willem de Vries, J.; Schnichels, S.; Hurst, J.; Strudel, L.; Gruszka, A.; Kwak, M.; Bartz-Schmidt, K.U.; Spitzer, M.S.; Herrmann, A. DNA nanoparticles for ophthalmic drug delivery. Biomaterials 2018, 157, 98–106. [Google Scholar] [CrossRef]
- Huo, S.; Li, H.; Boersma, A.J.; Herrmann, A. DNA Nanotechnology Enters Cell Membranes. Adv. Sci. (Weinh.) 2019, 6, 1900043. [Google Scholar] [CrossRef]
- Liu, K.; Zheng, L.; Liu, Q.; de Vries, J.W.; Gerasimov, J.Y.; Herrmann, A. Nucleic acid chemistry in the organic phase: From functionalized oligonucleotides to DNA side chain polymers. J. Am. Chem. Soc. 2014, 136, 14255–14262. [Google Scholar] [CrossRef] [PubMed]
- Hurley, J.H.; Hanson, P.I. Membrane budding and scission by the ESCRT machinery: It’s all in the neck. Nat. Rev. Mol. Cell Biol. 2010, 11, 556–566. [Google Scholar] [CrossRef]
- Langecker, M.; Arnaut, V.; List, J.; Simmel, F.C. DNA nanostructures interacting with lipid bilayer membranes. Acc. Chem. Res. 2014, 47, 1807–1815. [Google Scholar] [CrossRef]
- Anaya, M.; Kwak, M.; Musser, A.J.; Mullen, K.; Herrmann, A. Tunable hydrophobicity in DNA micelles: Design, synthesis, and characterization of a new family of DNA amphiphiles. Chemistry 2010, 16, 12852–12859. [Google Scholar] [CrossRef] [PubMed]
- Ohmann, A.; Gopfrich, K.; Joshi, H.; Thompson, R.F.; Sobota, D.; Ranson, N.A.; Aksimentiev, A.; Keyser, U.F. Controlling aggregation of cholesterol-modified DNA nanostructures. Nucleic Acids Res. 2019, 47, 11441–11451. [Google Scholar] [CrossRef] [PubMed]
- Zhang, Y.; Peng, R.; Xu, F.; Ke, Y. Hierarchical Self-Assembly of Cholesterol-DNA Nanorods. Bioconjugate Chem. 2019, 30, 1845–1849. [Google Scholar] [CrossRef]
- Choi, K.M.; Kwon, I.C.; Ahn, H.J. Self-assembled amphiphilic DNA-cholesterol/DNA-peptide hybrid duplexes with liposome-like structure for doxorubicin delivery. Biomaterials 2013, 34, 4183–4190. [Google Scholar] [CrossRef] [PubMed]
- Jorge, A.F.; Aviñó, A.; Pais, A.A.; Eritja, R.; Fàbrega, C. DNA-based nanoscaffolds as vehicles for 5-fluoro- 2’-deoxyuridine oligomers in colorectal cancer therapy. Nanoscale 2018, 10, 7238–7249. [Google Scholar] [CrossRef] [PubMed]
- Thelu, H.V.P.; Atchimnaidu, S.; Perumal, D.; Harikrishnan, K.S.; Vijayan, S.; Varghese, R. Self-Assembly of an aptamer-decorated, dna−protein hybrid nanogel: A biocompatible nanocarrier for targeted cancer therapy. ACS Appl. Bio Mater. 2019, 2, 5227–5234. [Google Scholar] [CrossRef]
- Wan, L.; Chen, Q.; Liu, J.; Yang, X.; Huang, J.; Li, L.; Guo, X.; Zhang, J.; Wang, K. Programmable Self-Assembly of DNA-Protein Hybrid Hydrogel for Enzyme Encapsulation with Enhanced Biological Stability. Biomacromolecules 2016, 17, 1543–1550. [Google Scholar] [CrossRef]
- Wu, Y.; Li, C.; Boldt, F.; Wang, Y.; Kuan, S.L.; Tran, T.T.; Mikhalevich, V.; Fortsch, C.; Barth, H.; Yang, Z.; et al. Programmable protein-DNA hybrid hydrogels for the immobilization and release of functional proteins. Chem. Commun. (Camb.) 2014, 50, 14620–14622. [Google Scholar] [CrossRef]
- Xu, Y.; Jiang, S.; Simmons, C.R.; Narayanan, R.P.; Zhang, F.; Aziz, A.M.; Yan, H.; Stephanopoulos, N. Tunable Nanoscale Cages from Self-Assembling DNA and Protein Building Blocks. ACS Nano 2019, 13, 3545–3554. [Google Scholar] [CrossRef]
- Julin, S.; Korpi, A.; Nonappa; Shen, B.; Liljestrom, V.; Ikkala, O.; Keller, A.; Linko, V.; Kostiainen, M.A. DNA origami directed 3D nanoparticle superlattice via electrostatic assembly. Nanoscale 2019, 11, 4546–4551. [Google Scholar] [CrossRef] [PubMed]
- Zhou, T.; Wang, Y.; Dong, Y.; Chen, C.; Liu, D.; Yang, Z. Tetrahedron DNA dendrimers and their encapsulation of gold nanoparticles. Bioorganic Med. Chem. 2014, 22, 4391–4394. [Google Scholar] [CrossRef]
- Huo, S.D.; Gong, N.Q.; Jiang, Y.; Chen, F.; Guo, H.B.; Gan, Y.L.; Wang, Z.S.; Herrmann, A.; Liang, X.J. Gold-DNA nanosunflowers for efficient gene silencing with controllable transformation. Sci. Adv. 2019, 5. [Google Scholar] [CrossRef] [PubMed]
- Pal, S.; Deng, Z.; Ding, B.; Yan, H.; Liu, Y. DNA-origami-directed self-assembly of discrete silver-nanoparticle architectures. Angew. Chem. Int. Ed. Engl. 2010, 49, 2700–2704. [Google Scholar] [CrossRef] [PubMed]
- Eskelinen, A.P.; Moerland, R.J.; Kostiainen, M.A.; Torma, P. Self-assembled silver nanoparticles in a bow-tie antenna configuration. Small 2014, 10, 1057–1062. [Google Scholar] [CrossRef]
- Pal, S.; Varghese, R.; Deng, Z.; Zhao, Z.; Kumar, A.; Yan, H.; Liu, Y. Site-specific synthesis and in situ immobilization of fluorescent silver nanoclusters on DNA nanoscaffolds by use of the Tollens reaction. Angew. Chem. Int. Ed. Engl. 2011, 50, 4176–4179. [Google Scholar] [CrossRef] [PubMed]
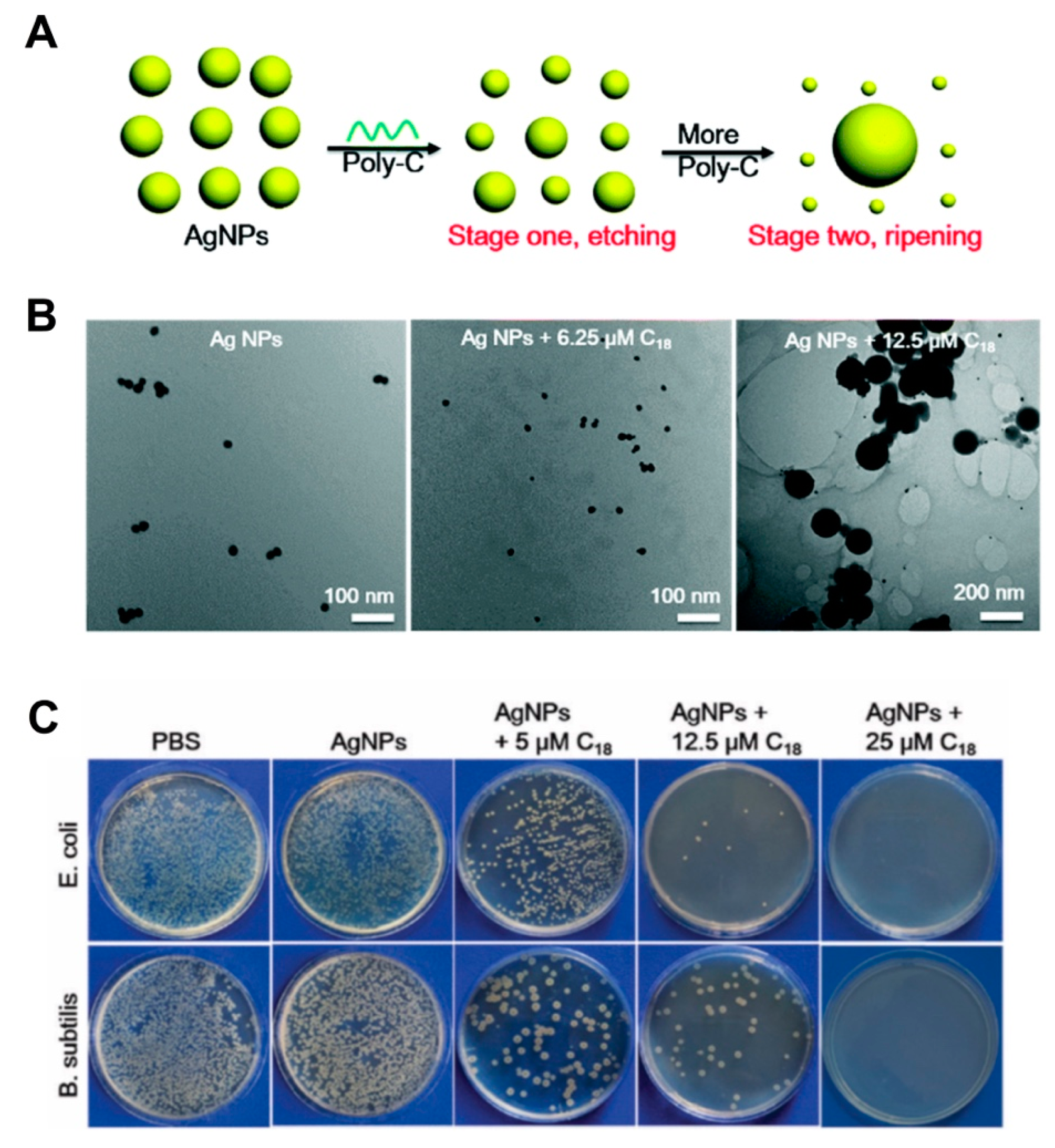
- Hu, S.; Yi, T.; Huang, Z.; Liu, B.; Wang, J.; Yi, X.; Liu, J. Etching silver nanoparticles using DNA. Mater. Horiz. 2019, 6, 155. [Google Scholar] [CrossRef]
- Deng, Z.; Samanta, A.; Nangreave, J.; Yan, H.; Liu, Y. Robust DNA-functionalized core/shell quantum dots with fluorescent emission spanning from UV-vis to near-IR and compatible with DNA-directed self-assembly. J. Am. Chem. Soc. 2012, 134, 17424–17427. [Google Scholar] [CrossRef]
- Zhang, L.; Jean, S.R.; Ahmed, S.; Aldridge, P.M.; Li, X.; Fan, F.; Sargent, E.H.; Kelley, S.O. Multifunctional quantum dot DNA hydrogels. Nat. Commun. 2017, 8, 381. [Google Scholar] [CrossRef]
- Trads, J.B.; Torring, T.; Gothelf, K.V. Site-Selective Conjugation of Native Proteins with DNA. Acc. Chem. Res. 2017, 50, 1367–1374. [Google Scholar] [CrossRef]
- Sagredo, S.; Pirzer, T.; Aghebat Rafat, A.; Goetzfried, M.A.; Moncalian, G.; Simmel, F.C.; de la Cruz, F. Orthogonal Protein Assembly on DNA Nanostructures Using Relaxases. Angew. Chem. Int. Ed. Engl. 2016, 55, 4348–4352. [Google Scholar] [CrossRef] [PubMed]
- Zhang, F.; Nangreave, J.; Liu, Y.; Yan, H. Structural DNA nanotechnology: State of the art and future perspective. J. Am. Chem. Soc. 2014, 136, 11198–11211. [Google Scholar] [CrossRef] [PubMed]
- Abu-Salah, K.M.; Zourob, M.M.; Mouffouk, F.; Alrokayan, S.A.; Alaamery, M.A.; Ansari, A.A. DNA-Based Nanobiosensors as an Emerging Platform for Detection of Disease. Sensors 2015, 15, 14539–14568. [Google Scholar] [CrossRef] [PubMed]
- Li, S.; Jiang, Q.; Liu, S.; Zhang, Y.; Tian, Y.; Song, C.; Wang, J.; Zou, Y.; Anderson, G.J.; Han, J.Y.; et al. A DNA nanorobot functions as a cancer therapeutic in response to a molecular trigger in vivo. Nat. Biotechnol. 2018, 36, 258–264. [Google Scholar] [CrossRef] [PubMed]
- Liu, X.; Xu, Y.; Yu, T.; Clifford, C.; Liu, Y.; Yan, H.; Chang, Y. A DNA nanostructure platform for directed assembly of synthetic vaccines. Nano Lett. 2012, 12, 4254–4259. [Google Scholar] [CrossRef]
- Auvinen, H.; Zhang, H.; Nonappa; Kopilow, A.; Niemela, E.H.; Nummelin, S.; Correia, A.; Santos, H.A.; Linko, V.; Kostiainen, M.A. Protein Coating of DNA Nanostructures for Enhanced Stability and Immunocompatibility. Adv. Healthc. Mater. 2017, 6. [Google Scholar] [CrossRef]
- Xu, X.; Fang, S.; Zhuang, Y.; Wu, S.; Pan, Q.; Li, L.; Wang, X.; Sun, X.; Liu, B.; Wu, Y. Cationic Albumin Encapsulated DNA Origami for Enhanced Cellular Transfection and Stability. Materials 2019, 12, 949. [Google Scholar] [CrossRef]
- Jain, A.; Singh, S.K.; Arya, S.K.; Kundu, S.C.; Kapoor, S. Protein nanoparticles: Promising platforms for drug delivery applications. ACS Biomater. Sci. Eng. 2018, 4, 3939–3961. [Google Scholar] [CrossRef]
- Ng, D.Y.W.; Wu, Y.; Kuan, S.L.; Weil, T. Programming Supramolecular Biohybrids as Precision Therapeutics. Acc. Chem. Res. 2014, 47, 3471–3480. [Google Scholar] [CrossRef] [PubMed]
- Niemeyer, C.M. Semisynthetic DNA-protein conjugates for biosensing and nanofabrication. Angew. Chem. Int. Ed. Engl. 2010, 49, 1200–1216. [Google Scholar] [CrossRef]
- Koussa, M.A.; Sotomayor, M.; Wong, W.P. Protocol for sortase-mediated construction of DNA-protein hybrids and functional nanostructures. Methods 2014, 67, 134–141. [Google Scholar] [CrossRef] [PubMed]
- Kazane, S.A.; Sok, D.; Cho, E.H.; Uson, M.L.; Kuhn, P.; Schultz, P.G.; Smider, V.V. Site-specific DNA-antibody conjugates for specific and sensitive immuno-PCR. Proc. Natl. Acad. Sci. USA 2012, 109, 3731–3736. [Google Scholar] [CrossRef]
- Ryu, Y.; Hong, C.A.; Song, Y.; Beak, J.; Seo, B.A.; Lee, J.J.; Kim, H.S. Modular protein-DNA hybrid nanostructures as a drug delivery platform. Nanoscale 2020, 12, 4975–4981. [Google Scholar] [CrossRef] [PubMed]
- Yang, X.Y.; Yang, M.; Pang, B.; Vara, M.; Xia, Y. Gold nanomaterials at work in biomedicine. Chem. Rev. 2015, 115, 10410–10488. [Google Scholar] [CrossRef]
- Ghosh, P.; Han, G.; De, M.; Kim, C.K.; Rotello, V.M. Gold nanoparticles in delivery applications. Adv. Drug Deliv. Rev. 2008, 60, 1307–1315. [Google Scholar] [CrossRef]
- Hurst, S.J.; Lytton-Jean, A.K.; Mirkin, C.A. Maximizing DNA loading on a range of gold nanoparticle sizes. Anal. Chem. 2006, 78, 8313–8318. [Google Scholar] [CrossRef]
- Kapadia, C.H.; Melamed, J.R.; Day, E.S. Spherical Nucleic Acid Nanoparticles: Therapeutic Potential. BioDrugs 2018, 32, 297–309. [Google Scholar] [CrossRef] [PubMed]
- Sun, T.; Zhang, Y.S.; Pang, B.; Hyun, D.C.; Yang, M.; Xia, Y. Engineered nanoparticles for drug delivery in cancer therapy. Angew. Chem. Int. Ed. Engl. 2014, 53, 12320–12364. [Google Scholar] [CrossRef]
- Zhou, C.; Long, M.; Qin, Y.P.; Sun, X.K.; Zheng, J. Luminescent Gold Nanoparticles with Efficient Renal Clearance. Angew. Chem. Int. Ed. 2011, 50, 3168–3172. [Google Scholar] [CrossRef] [PubMed]
- Sim, W.; Barnard, R.T.; Blaskovich, M.A.T.; Ziora, Z.M. Antimicrobial Silver in Medicinal and Consumer Applications: A Patent Review of the Past Decade (2007–2017). Antibiotics 2018, 7, 93. [Google Scholar] [CrossRef] [PubMed]
- Turner, R.J. Metal-based antimicrobial strategies. Microb. Biotechnol. 2017, 10, 1062–1065. [Google Scholar] [CrossRef] [PubMed]
- Vaidya, M.Y.; McBain, A.J.; Butler, J.A.; Banks, C.E.; Whitehead, K.A. Antimicrobial Efficacy and Synergy of Metal Ions against Enterococcus faecium, Klebsiella pneumoniae and Acinetobacter baumannii in Planktonic and Biofilm Phenotypes. Sci. Rep. 2017, 7, 5911. [Google Scholar] [CrossRef]
- Shariq Qayyum, A.U.K. Nanoparticles vs. biofilms: A battle against another paradigm of antibiotic resistance. MedChemComm 2016, 7, 1479–1498. [Google Scholar] [CrossRef]
- Bhatia, E.; Banerjee, R. Hybrid silver-gold nanoparticles suppress drug resistant polymicrobial biofilm formation and intracellular infection. J. Mater. Chem. B 2020. [Google Scholar] [CrossRef]
- Flores, C.Y.; Minan, A.G.; Grillo, C.A.; Salvarezza, R.C.; Vericat, C.; Schilardi, P.L. Citrate-capped silver nanoparticles showing good bactericidal effect against both planktonic and sessile bacteria and a low cytotoxicity to osteoblastic cells. ACS Appl. Mater. Interfaces 2013, 5, 3149–3159. [Google Scholar] [CrossRef]
- Mu, H.; Tang, J.; Liu, Q.; Sun, C.; Wang, T.; Duan, J. Potent Antibacterial Nanoparticles against Biofilm and Intracellular Bacteria. Sci. Rep. 2016, 6, 18877. [Google Scholar] [CrossRef]
- Schneider, G. Antimicrobial silver nanoparticles—Regulatory situation in the European Union. Mater. Today Proc. 2017, 4, S200–S207. [Google Scholar] [CrossRef]
- Calderon-Jimenez, B.; Johnson, M.E.; Montoro Bustos, A.R.; Murphy, K.E.; Winchester, M.R.; Vega Baudrit, J.R. Silver Nanoparticles: Technological Advances, Societal Impacts, and Metrological Challenges. Front. Chem. 2017, 5, 6. [Google Scholar] [CrossRef]
- Anjum, S.; Singh, S.; Benedicte, L.; Roger, P.; Panigrahi, M.; Gupta, B. Biomodification Strategies for the Development of Antimicrobial Urinary Catheters: Overview and Advances. Glob. Chall. 2018, 2, 1700068. [Google Scholar] [CrossRef] [PubMed]
- Lara, H.H.; Garza-Trevino, E.N.; Ixtepan-Turrent, L.; Singh, D.K. Silver nanoparticles are broad-spectrum bactericidal and virucidal compounds. J. Nanobiotechnol. 2011, 9, 30. [Google Scholar] [CrossRef] [PubMed]
- Rudakiya, D.M.; Pawar, K. Bactericidal potential of silver nanoparticles synthesized using cell-free extract of Comamonas acidovorans: In vitro and in silico approaches. 3 Biotech. 2017, 7, 92. [Google Scholar] [CrossRef] [PubMed]
- Rai, M.; Yadav, A.; Gade, A. Silver nanoparticles as a new generation of antimicrobials. Biotechnol. Adv. 2009, 27, 76–83. [Google Scholar] [CrossRef] [PubMed]
- Liu, Q.; Song, C.; Wang, Z.G.; Li, N.; Ding, B. Precise organization of metal nanoparticles on DNA origami template. Methods 2014, 67, 205–214. [Google Scholar] [CrossRef] [PubMed]
- Julin, S.; Nummelin, S.; Kostiainen, M.A.; Linko, V. DNA nanostructure-directed assembly of metal nanoparticle superlattices. J. Nanopartical Res. 2018, 20, 119. [Google Scholar] [CrossRef]
- Khalandi, B.; Asadi, N.; Milani, M.; Davaran, S.; Abadi, A.J.; Abasi, E.; Akbarzadeh, A. A Review on Potential Role of Silver Nanoparticles and Possible Mechanisms of their Actions on Bacteria. Drug Res. (Stuttg.) 2017, 67, 70–76. [Google Scholar] [CrossRef]
- Qiao, Z.; Yao, Y.; Song, S.; Yin, M.; Luo, J. Silver nanoparticles with pH induced surface charge switchable properties for antibacterial and antibiofilm applications. J. Mater. Chem. B 2019, 7, 830–840. [Google Scholar] [CrossRef]
- Ghai, I.; Ghai, S. Understanding antibiotic resistance via outer membrane permeability. Infect. Drug Resist. 2018, 11, 523–530. [Google Scholar] [CrossRef]
- Soares, D.S.; Reis, A.D.F.; Silva Junior, G.B.D.; Leite, T.T.; Parente Filho, S.L.A.; Rocha, C.V.O.; Daher, E.F. Polymyxin-B and vancomycin-associated acute kidney injury in critically ill patients. Pathog. Glob. Health 2017, 111, 137–142. [Google Scholar] [CrossRef][Green Version]
- Malanovic, N.; Lohner, K. Gram-positive bacterial cell envelopes: The impact on the activity of antimicrobial peptides. Biochim. Biophys. Acta 2016, 1858, 936–946. [Google Scholar] [CrossRef] [PubMed]
- Li, X.Z.; Plésiat, P.; Nikaido, H. The challenge of efflux-mediated antibiotic resistance in Gram-negative bacteria. Clin. Microbiol. Rev. 2015, 28, 337–418. [Google Scholar] [CrossRef] [PubMed]
- Masi, M.; Réfregiers, M.; Pos, K.M.; Pagès, J.M. Mechanisms of envelope permeability and antibiotic influx and efflux in Gram-negative bacteria. Nat. Microbiol. 2017, 2, 17001. [Google Scholar] [CrossRef] [PubMed]
- Reygaert, W.C. An overview of the antimicrobial resistance mechanisms of bacteria. Aims Microbiol. 2018, 4, 482–501. [Google Scholar] [CrossRef] [PubMed]
- Huang, Y.W.; Lee, H.J.; Tolliver, L.M.; Aronstam, R.S. Delivery of nucleic acids and nanomaterials by cell-penetrating peptides: Opportunities and challenges. Biomed. Res. Int. 2015, 2015, 834079. [Google Scholar] [CrossRef]
- Puckett, S.E.; Reese, K.A.; Mitev, G.M.; Mullen, V.; Johnson, R.C.; Pomraning, K.R.; Mellbye, B.L.; Tilley, L.D.; Iversen, P.L.; Freitag, M.; et al. Bacterial resistance to antisense peptide phosphorodiamidate morpholino oligomers. Antimicrob. Agents Chemother. 2012, 56, 6147–6153. [Google Scholar] [CrossRef]
- Zhang, Y.; Ma, W.; Zhu, Y.; Shi, S.; Li, Q.; Mao, C.; Zhao, D.; Zhan, Y.; Shi, J.; Li, W.; et al. Inhibiting Methicillin-Resistant Staphylococcus aureus by Tetrahedral DNA Nanostructure-Enabled Antisense Peptide Nucleic Acid Delivery. Nano Lett. 2018, 18, 5652–5659. [Google Scholar] [CrossRef]
- Ghosal, A.; Vitali, A.; Stach, J.E.; Nielsen, P.E. Role of SbmA in the uptake of peptide nucleic acid (PNA)-peptide conjugates in E. coli. ACS Chem. Biol. 2013, 8, 360–367. [Google Scholar] [CrossRef]
- Setyawati, M.I.; Kutty, R.V.; Tay, C.Y.; Yuan, X.; Xie, J.; Leong, D.T. Novel theranostic DNA nanoscaffolds for the simultaneous detection and killing of Escherichia coli and Staphylococcus aureus. ACS Appl. Mater. Interfaces 2014, 6, 21822–21831. [Google Scholar] [CrossRef]
- Liu, Y.; Sun, Y.; Li, S.; Liu, M.; Qin, X.; Chen, X.; Lin, Y. Tetrahedral Framework Nucleic Acids Deliver Antimicrobial Peptides with Improved Effects and Less Susceptibility to Bacterial Degradation. Nano Lett. 2020, 20, 3602–3610. [Google Scholar] [CrossRef]
- Readman, J.B.; Dickson, G.; Coldham, N.G. Tetrahedral DNA Nanoparticle Vector for Intracellular Delivery of Targeted Peptide Nucleic Acid Antisense Agents to Restore Antibiotic Sensitivity in Cefotaxime-Resistant Escherichia coli. Nucleic Acid 2017, 27, 176–181. [Google Scholar] [CrossRef] [PubMed]
- Debacker, A.J.; Sharma, V.K.; Meda Krishnamurthy, P.; O’Reilly, D.; Greenhill, R.; Watts, J.K. Next-Generation Peptide Nucleic Acid Chimeras Exhibit High Affinity and Potent Gene Silencing. Biochemistry 2019, 58, 582–589. [Google Scholar] [CrossRef] [PubMed]
- Hegarty, J.P.; Stewart, D.B., Sr. Advances in therapeutic bacterial antisense biotechnology. Appl. Microbiol. Biotechnol. 2018, 102, 1055–1065. [Google Scholar] [CrossRef] [PubMed]
- Zhang, Y.; Xie, X.; Ma, W.; Zhan, Y.; Mao, C.; Shao, X.; Lin, Y. Multi-targeted Antisense Oligonucleotide Delivery by a Framework Nucleic Acid for Inhibiting Biofilm Formation and Virulence. Nano-Micro. Lett. 2020, 12, 74. [Google Scholar] [CrossRef]
- Mell, J.C.; Hall, I.M.; Redfield, R.J. Defining the DNA uptake specificity of naturally competent Haemophilus influenzae cells. Nucleic Acids Res. 2012, 40, 8536–8549. [Google Scholar] [CrossRef]
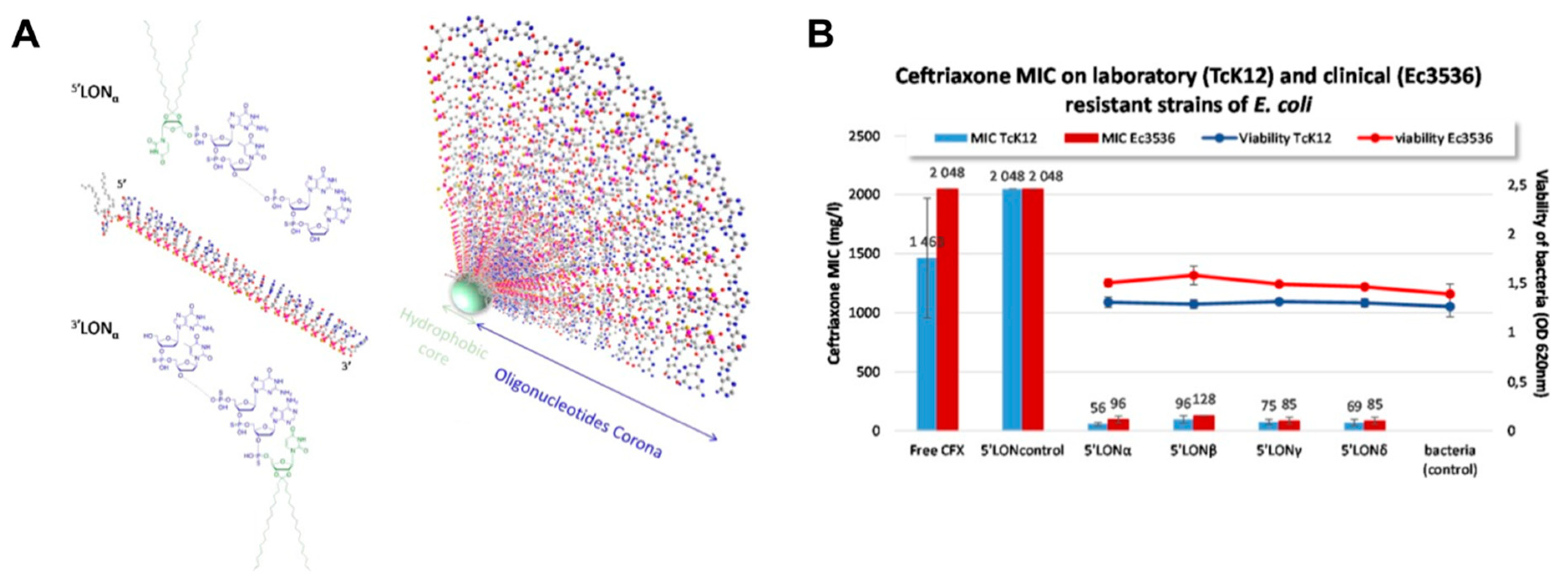
- Kauss, T.; Arpin, C.; Bientz, L.; Vinh Nguyen, P.; Vialet, B.; Benizri, S.; Barthelemy, P. Lipid oligonucleotides as a new strategy for tackling the antibiotic resistance. Sci. Rep. 2020, 10, 1054. [Google Scholar] [CrossRef]
- Khara, J.S.; Obuobi, S.; Wang, Y.; Hamilton, M.S.; Robertson, B.D.; Newton, S.M.; Yang, Y.Y.; Langford, P.R.; Ee, P.L.R. Disruption of drug-resistant biofilms using de novo designed short alpha-helical antimicrobial peptides with idealized facial amphiphilicity. Acta Biomater. 2017, 57, 103–114. [Google Scholar] [CrossRef]
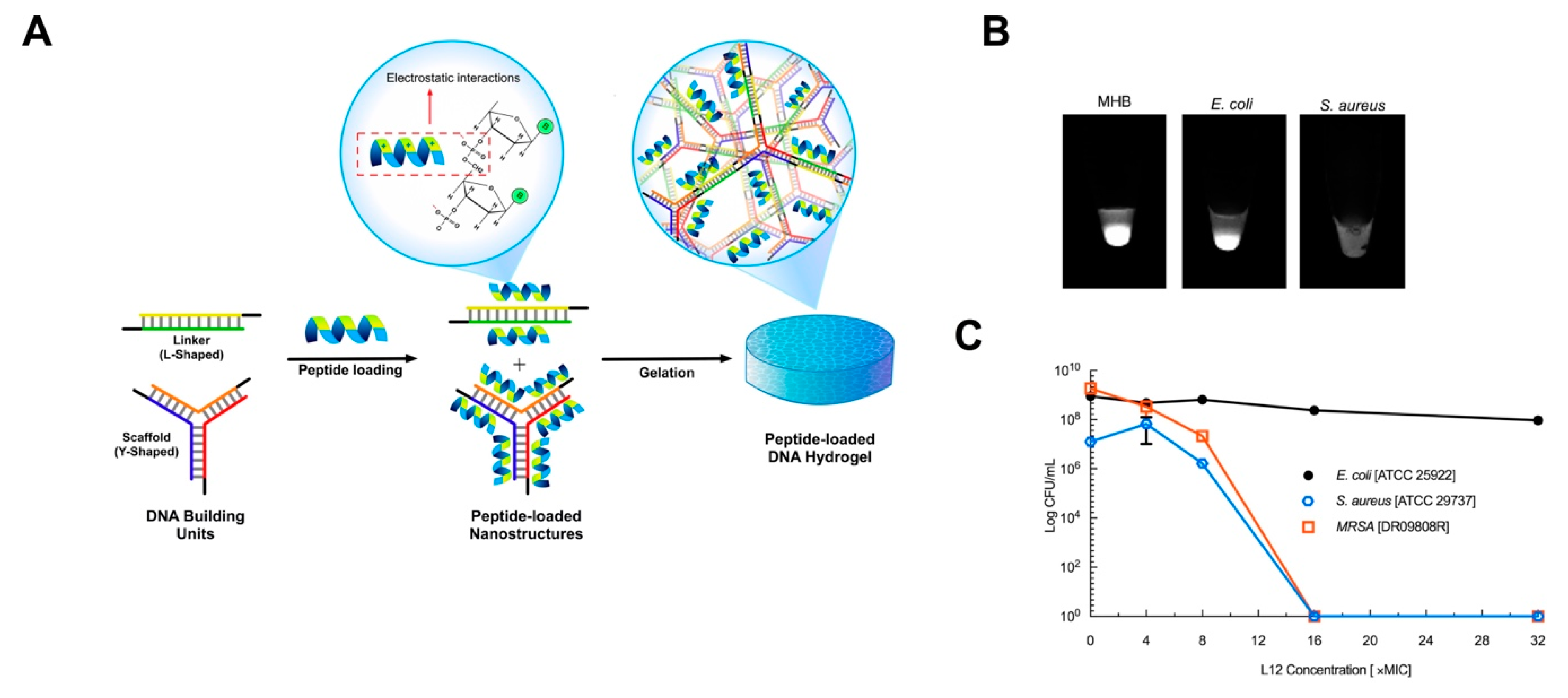
- Obuobi, S.; Tay, H.K.; Tram, N.D.T.; Selvarajan, V.; Khara, J.S.; Wang, Y.; Ee, P.L.R. Facile and efficient encapsulation of antimicrobial peptides via crosslinked DNA nanostructures and their application in wound therapy. J. Control. Release 2019, 313, 120–130. [Google Scholar] [CrossRef]
- Jiang, X.; Li, M.; Guo, X.; Chen, H.; Yang, M.; Rasooly, A. Self-Assembled DNA-THPS Hydrogel as a Topical Antibacterial Agent for Wound Healing. ACS Appl. Bio. Mater. 2019, 2, 1262–1269. [Google Scholar] [CrossRef]
- Jeon, H.; Nam, H.; Lee, J.B. Sustained Release of Minor-Groove-Binding Antibiotic Netropsin from Calcium-Coated Groove-Rich DNA Particles. Pharmaceutics 2019, 11, 387. [Google Scholar] [CrossRef]
- Kumari, S.; Rajit Prasad, S.; Mandal, D.; Das, P. Carbon dot-DNA-protoporphyrin hybrid hydrogel for sustained photoinduced antimicrobial activity. J. Colloid Interface Sci. 2019, 553, 228–238. [Google Scholar] [CrossRef] [PubMed]
- Ning, Y.; Cheng, L.; Ling, M.; Feng, X.; Chen, L.; Wu, M.; Deng, L. Efficient suppression of biofilm formation by a nucleic acid aptamer. Pathog. Dis. 2015, 73, ftv034. [Google Scholar] [CrossRef] [PubMed]
- Vivekananda, J.; Salgado, C.; Millenbaugh, N.J. DNA aptamers as a novel approach to neutralize Staphylococcus aureus α-toxin. Biochem. Biophys. Res. Commun. 2014, 444, 433–438. [Google Scholar] [CrossRef] [PubMed]
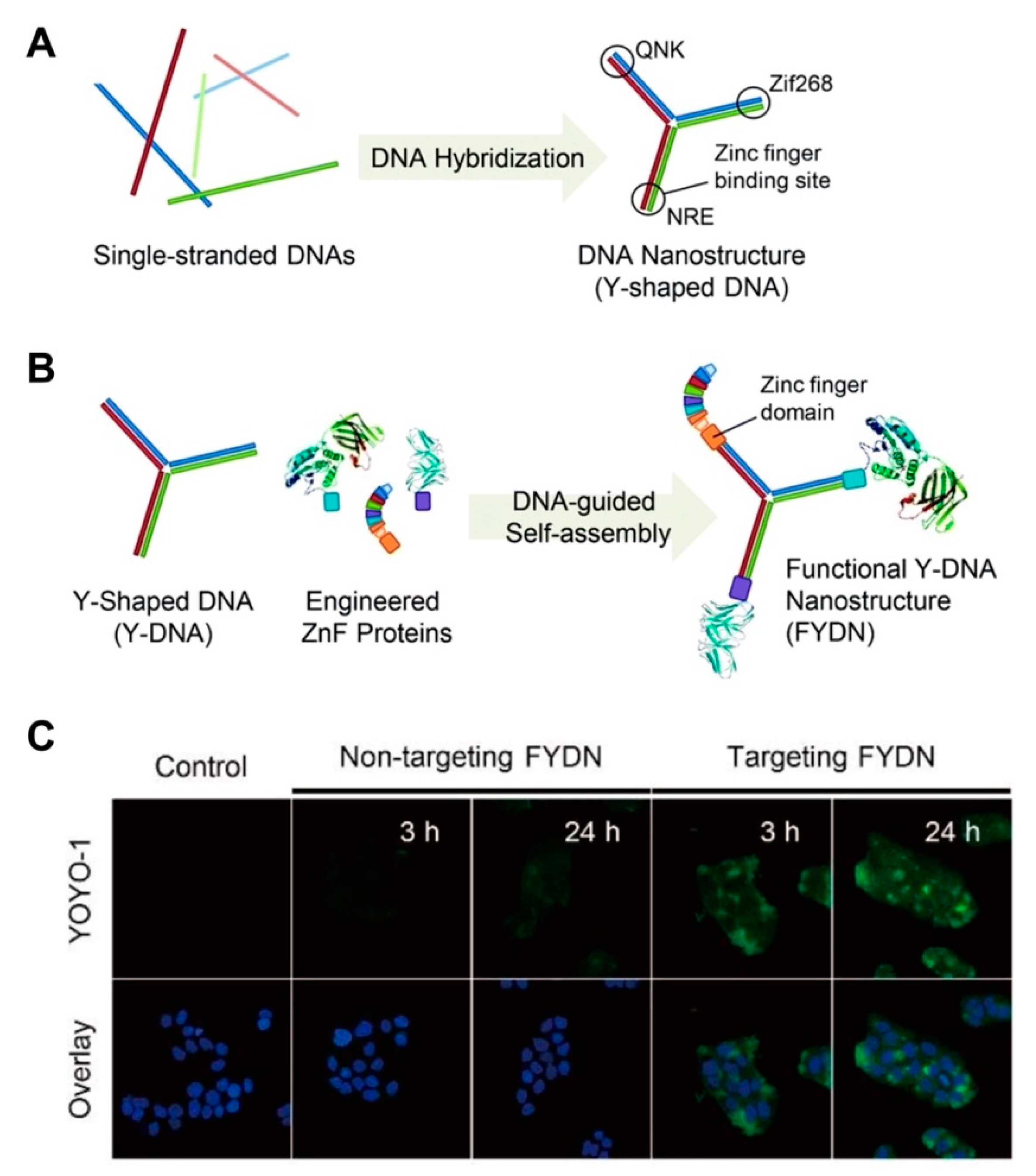
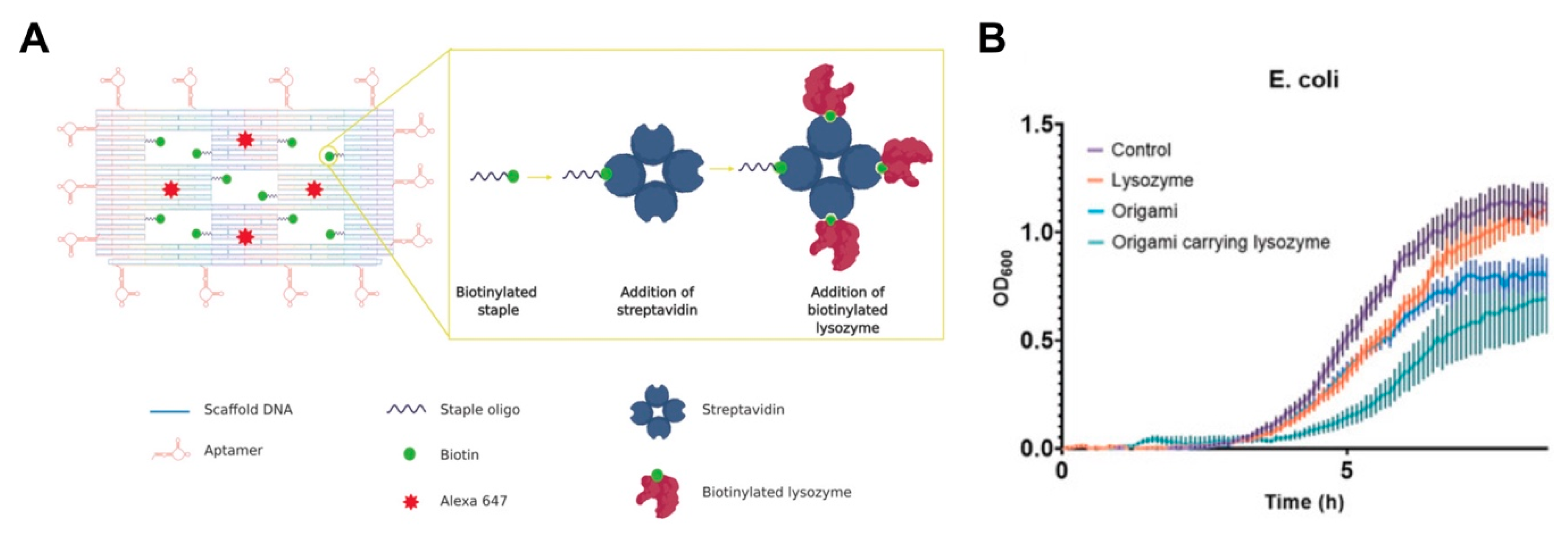
- Mela, I.; Vallejo-Ramirez, P.P.; Makarchuk, S.; Christie, G.; Bailey, D.; Henderson, R.M.; Sugiyama, H.; Endo, M.; Kaminski, C.F. DNA Nanostructures for Targeted Antimicrobial Delivery. Angew. Chem. Int. Ed. Engl. 2020. [Google Scholar] [CrossRef]
- Taskova, M.; Mantsiou, A.; Astakhova, K. Synthetic Nucleic Acid Analogues in Gene Therapy: An Update for Peptide-Oligonucleotide Conjugates. Chembiochem 2017, 18, 1671–1682. [Google Scholar] [CrossRef] [PubMed]
- Zhang, Q.; Lin, S.; Shi, S.; Zhang, T.; Ma, Q.; Tian, T.; Zhou, T.; Cai, X.; Lin, Y. Anti-inflammatory and Antioxidative Effects of Tetrahedral DNA Nanostructures via the Modulation of Macrophage Responses. ACS Appl. Mater. Interfaces 2018, 10, 3421–3430. [Google Scholar] [CrossRef]

| Group | Functionalization | Hybrid Nanostructure | Biological Application Tested | Representative References |
|---|---|---|---|---|
| Organic | Hydrophobic |
|
| |
| Protein/peptides |
|
| ||
| Inorganic | Gold |
|
| |
| Silver |
|
| ||
| Quantum dot |
|
|
© 2020 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Obuobi, S.; Škalko-Basnet, N. Nucleic Acid Hybrids as Advanced Antibacterial Nanocarriers. Pharmaceutics 2020, 12, 643. https://doi.org/10.3390/pharmaceutics12070643
Obuobi S, Škalko-Basnet N. Nucleic Acid Hybrids as Advanced Antibacterial Nanocarriers. Pharmaceutics. 2020; 12(7):643. https://doi.org/10.3390/pharmaceutics12070643
Chicago/Turabian StyleObuobi, Sybil, and Nataša Škalko-Basnet. 2020. "Nucleic Acid Hybrids as Advanced Antibacterial Nanocarriers" Pharmaceutics 12, no. 7: 643. https://doi.org/10.3390/pharmaceutics12070643
APA StyleObuobi, S., & Škalko-Basnet, N. (2020). Nucleic Acid Hybrids as Advanced Antibacterial Nanocarriers. Pharmaceutics, 12(7), 643. https://doi.org/10.3390/pharmaceutics12070643






