PH-Responsive, Cell-Penetrating, Core/Shell Magnetite/Silver Nanoparticles for the Delivery of Plasmids: Preparation, Characterization, and Preliminary In Vitro Evaluation
Abstract
1. Introduction
2. Materials and Methods
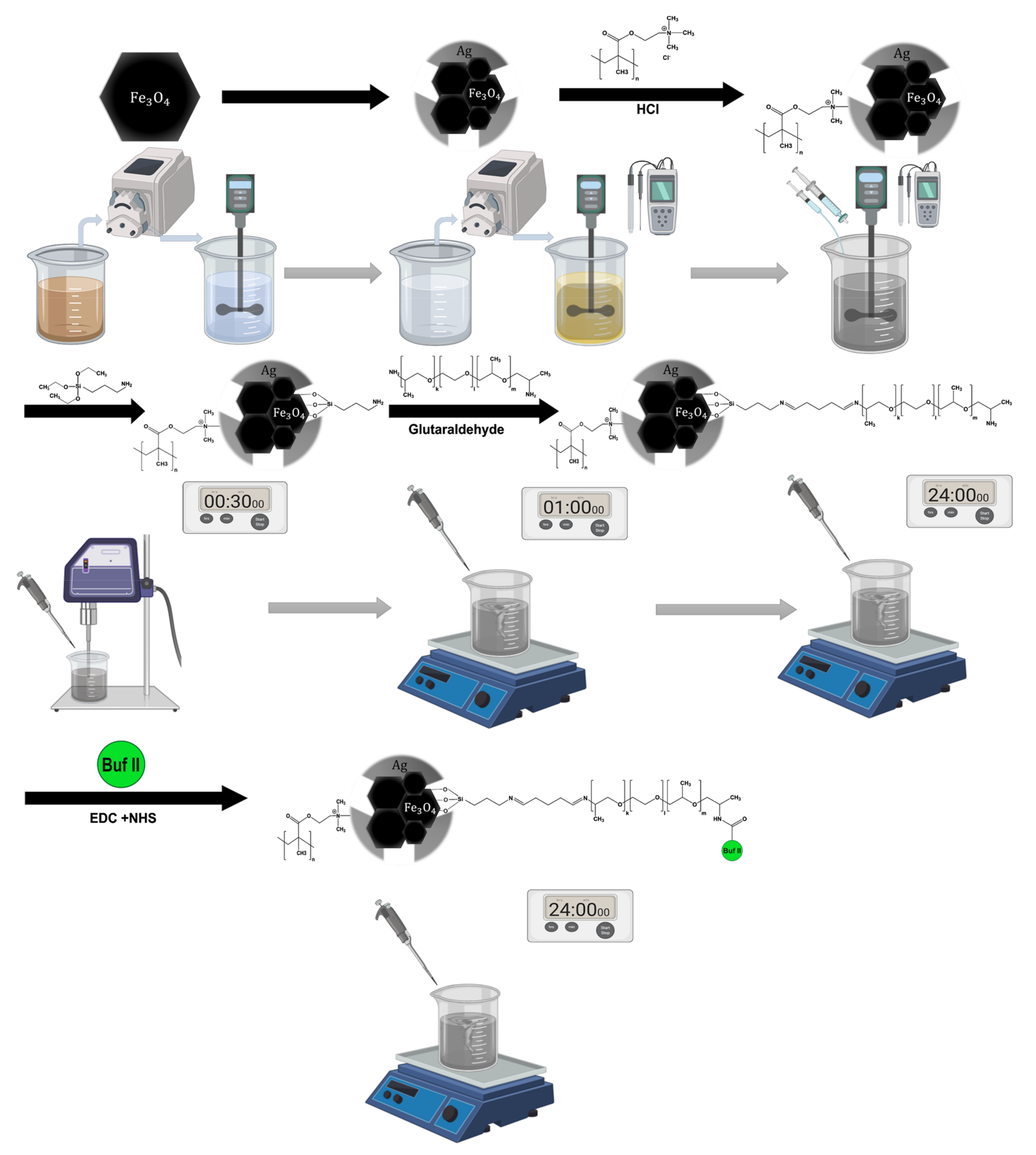
2.1. Magnetite Core and Silver Shell Synthesis
2.2. Conjugation of the PH-Responsive Polymer DMAEMA on Magnetite/Silver Nanoparticles
2.3. Conjugation of Buforin II (BUF-II) on Magnetite/Silver Nanoparticles
2.4. Labeling of Magnetite/Silver-pDMAEMA-PEA-BUF-II Nanobioconjugates with Rhodamine B
2.5. Synthesis of Liposomes and Magnetoliposomes
2.6. Characterization of Magnetite/Silver and Surface Conjugations
2.7. Loading and Delivery of Circular DNA
2.8. Hemolysis Assay
2.9. Platelet Aggregation Assay
2.10. LDH Cytotoxicity Assay
2.11. Cell Translocation and Endosome Escape
3. Results
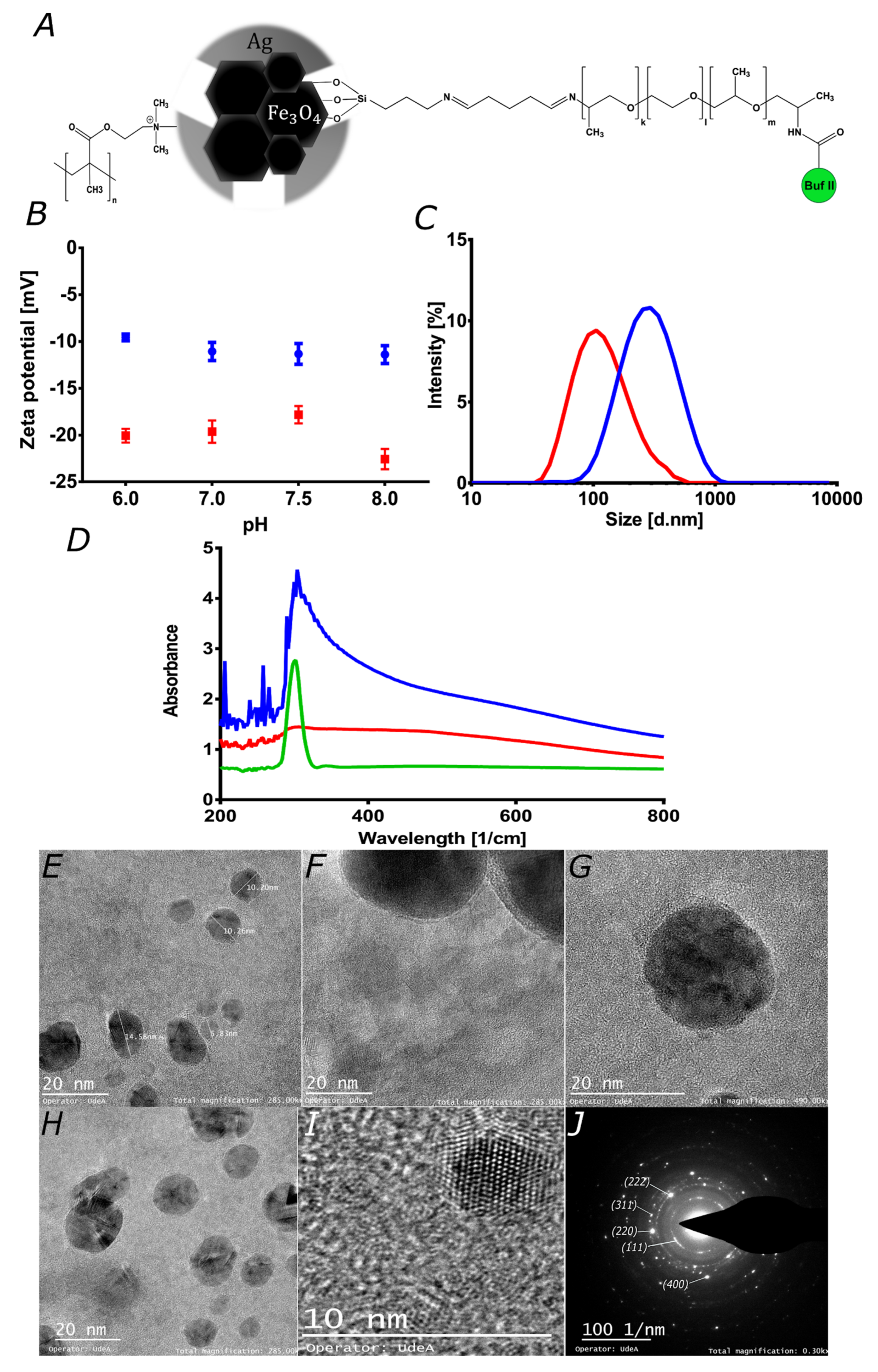
3.1. Characterization Magnetite/Silver Nanoparticles
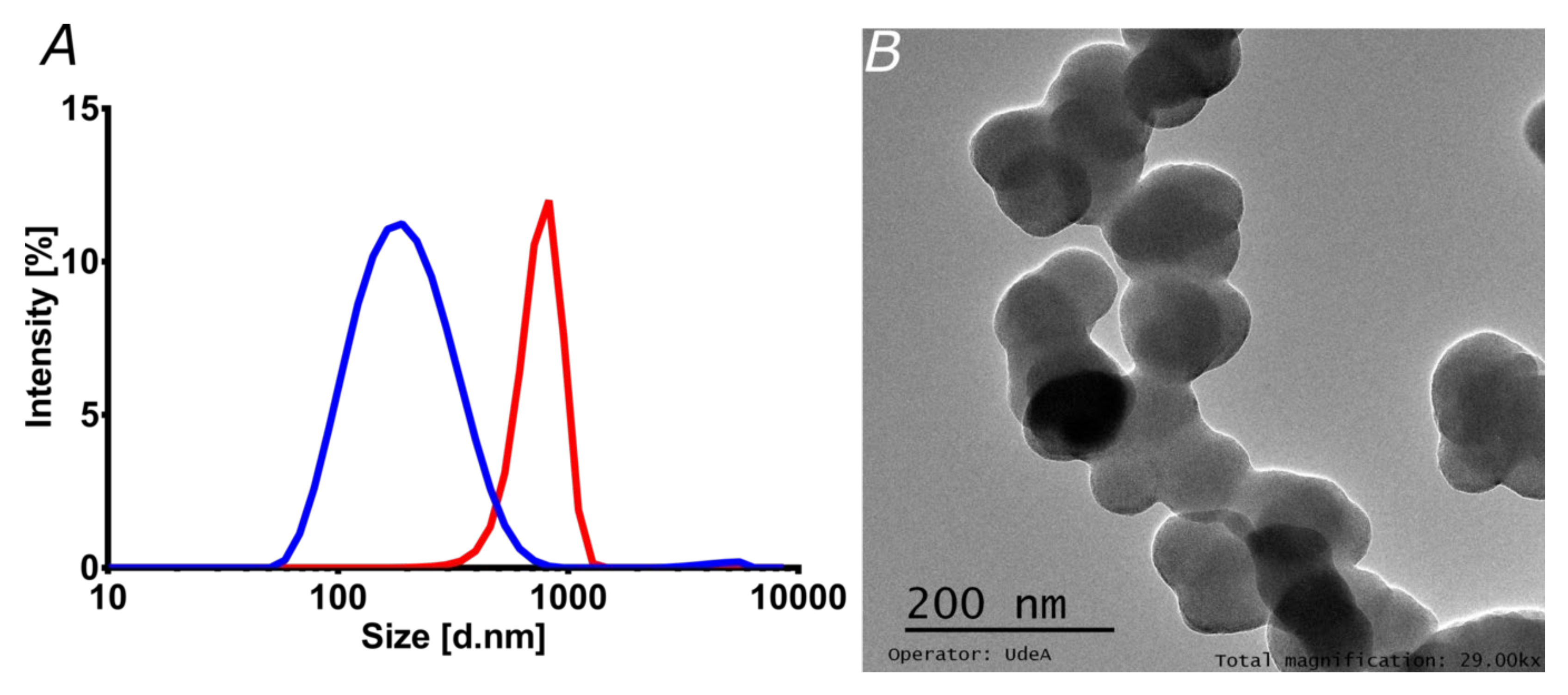
3.2. Characterization of Liposomes and Magnetoliposomes
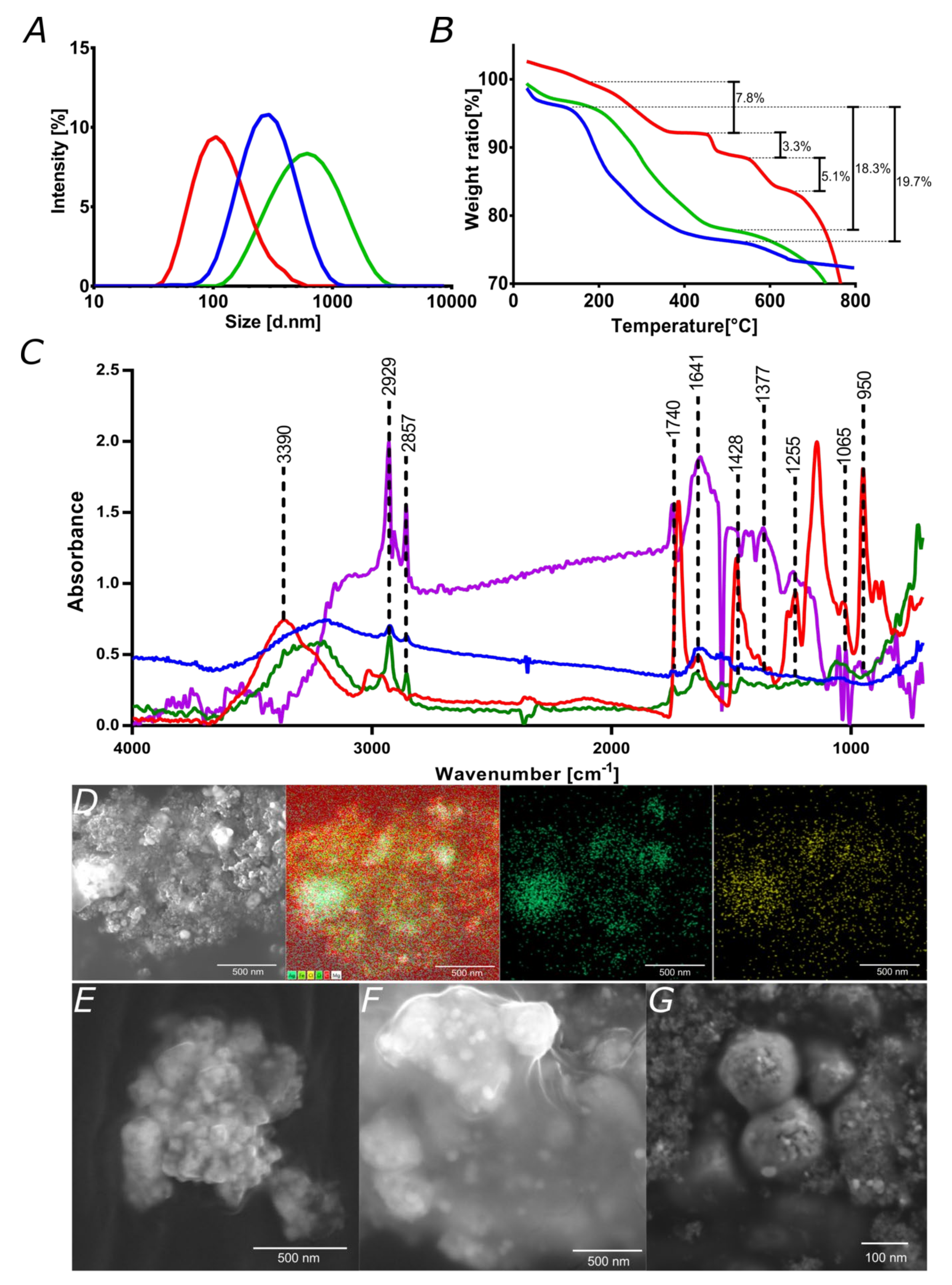
3.3. Characterization of Nanoconjugates and Nanobioconjugates
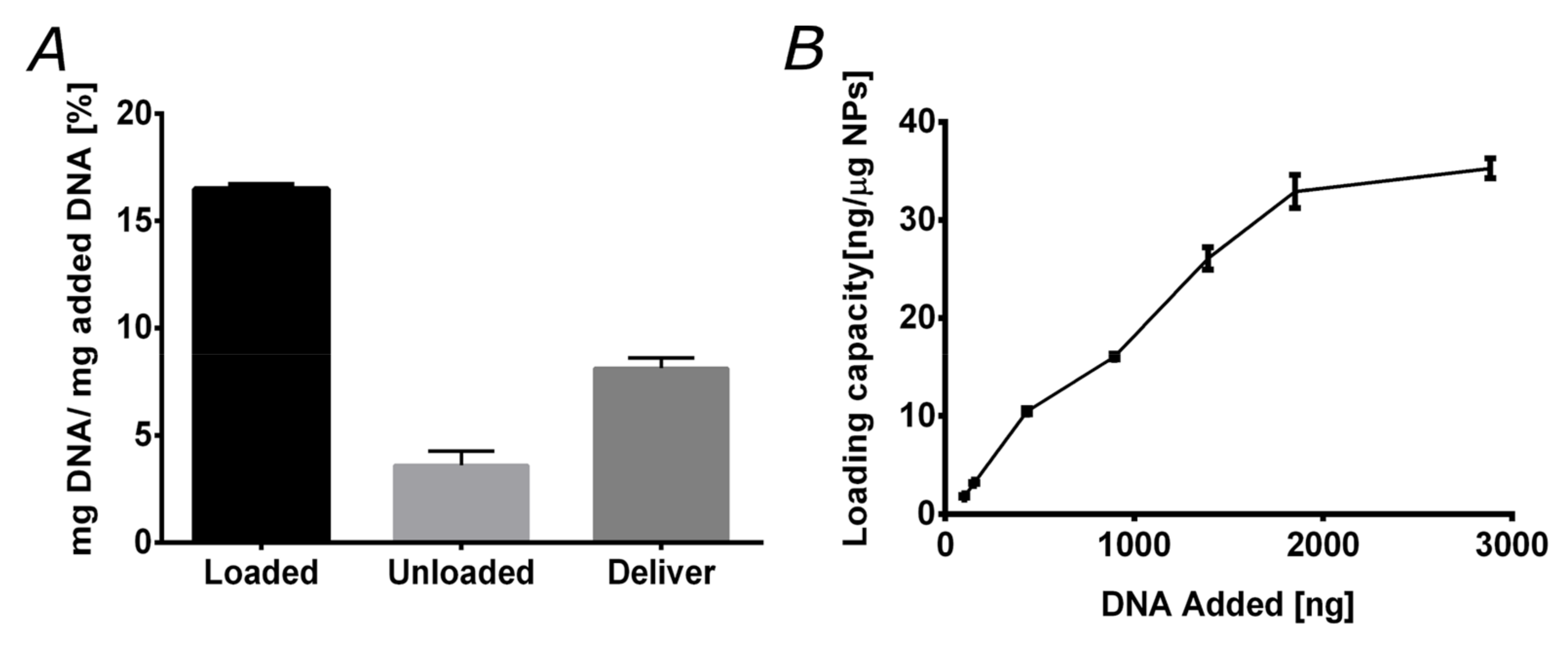
3.4. DNA Loading and Delivery
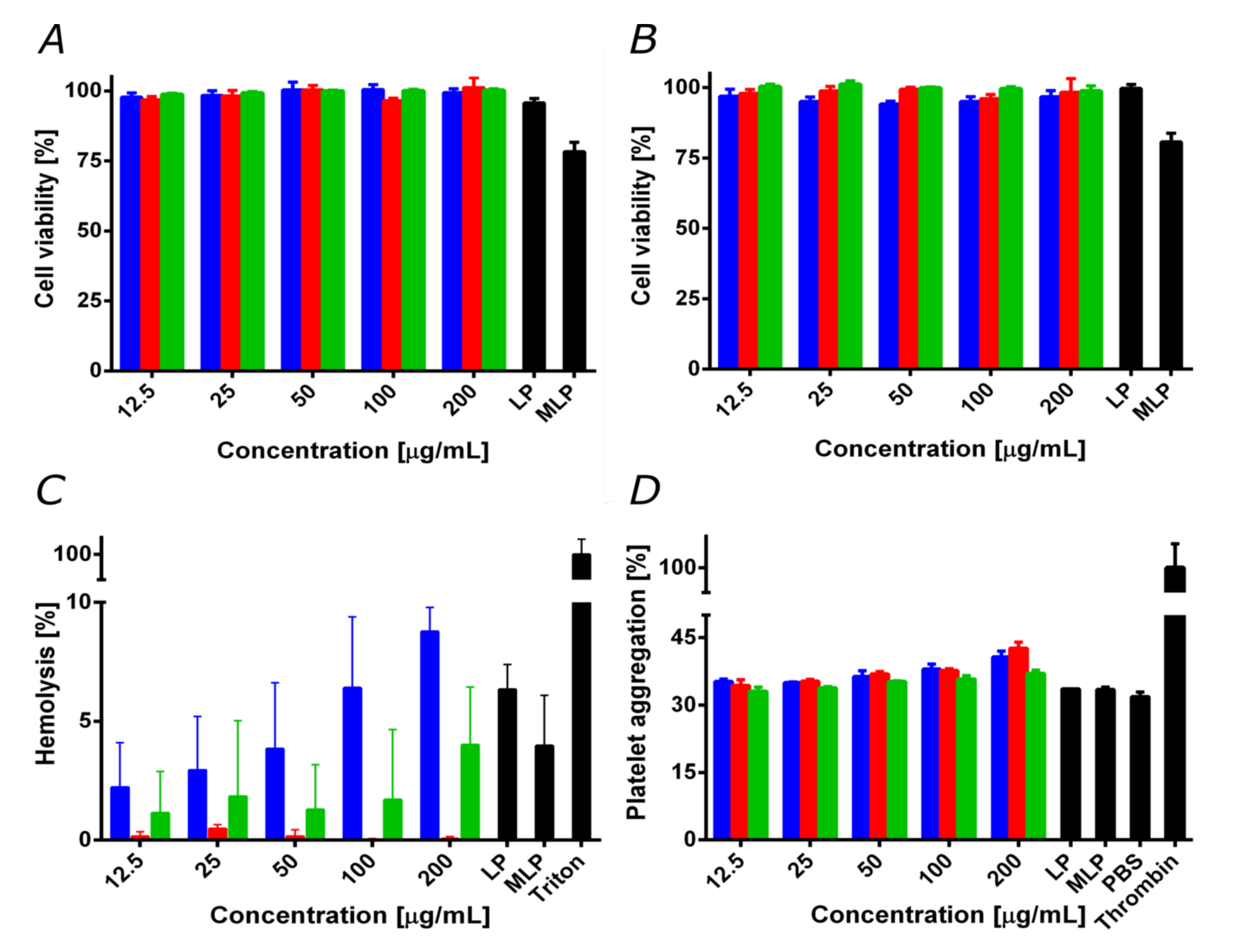
3.5. Biocompatibility
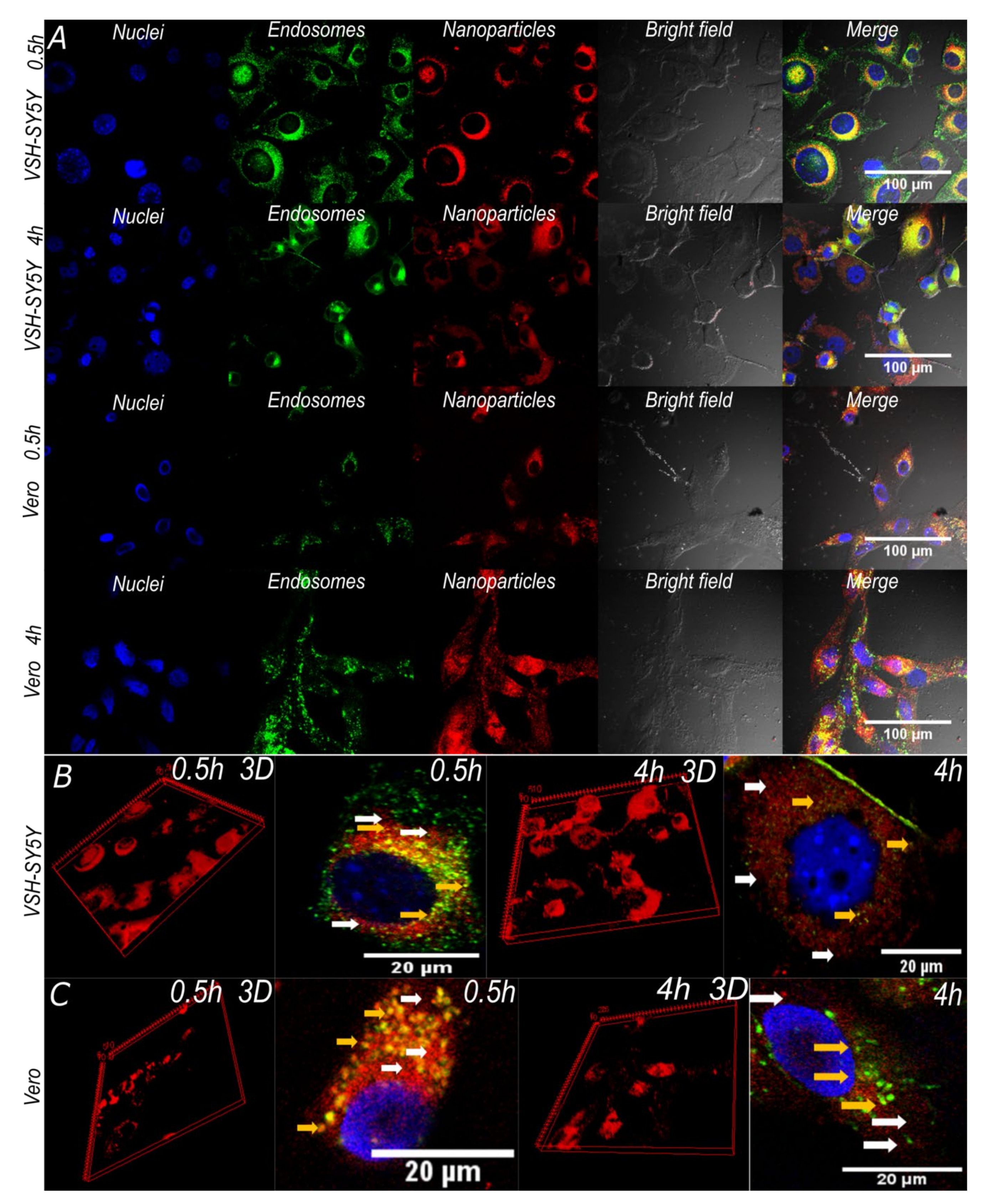
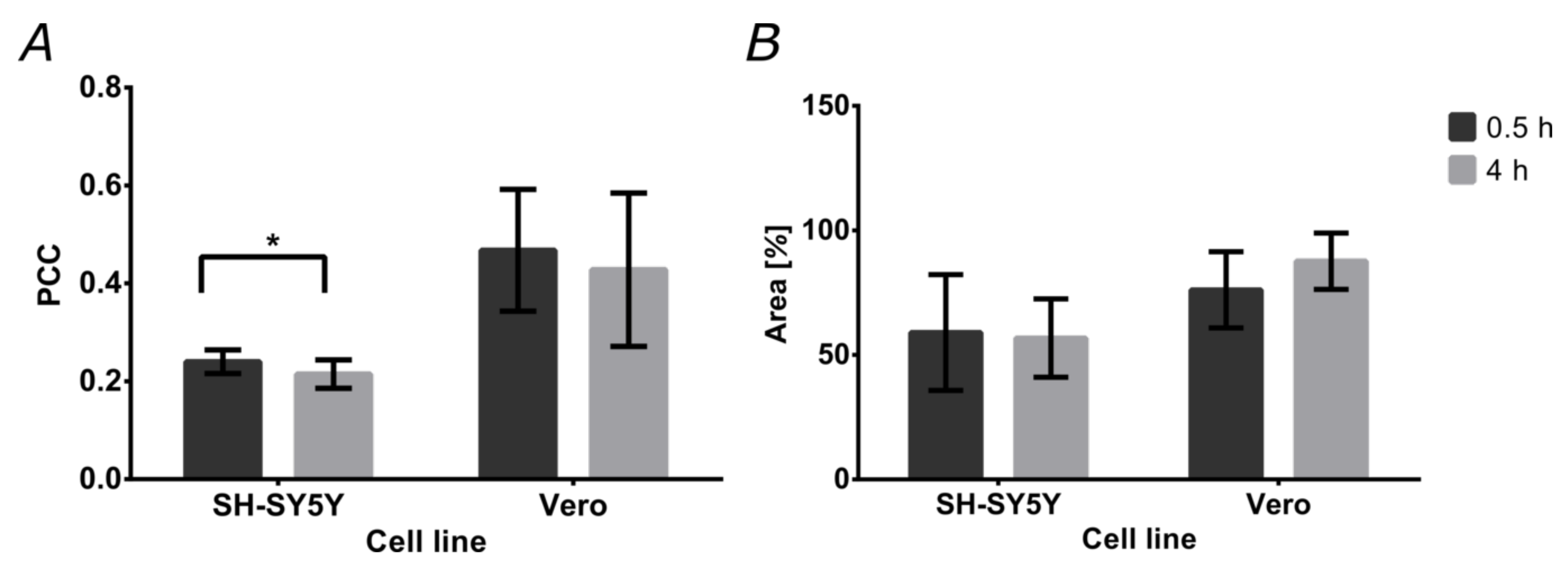
3.6. Cell Translocation and Endosome Escape
4. Conclusions
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
References
- Owen, A.; Rannard, S. Strengths weaknesses, opportunities and challenges for long acting injectable therapies: Insights for applications in {HIV} therapy. Adv. Drug Deliv. Rev. 2016, 103, 144–156. [Google Scholar] [CrossRef] [PubMed]
- Alexis, F.; Pridgen, E.; Molnar, L.K.; Farokhzad, O.C. Factors Affecting the Clearance and Biodistribution of Polymeric Nanoparticles. Mol. Pharm. 2008, 5, 505–515. [Google Scholar] [CrossRef] [PubMed]
- Wen, H.; Jung, H.; Li, X. Drug Delivery Approaches in Addressing Clinical Pharmacology-Related Issues: Opportunities and Challenges. AAPS J. 2015, 17, 1327–1340. [Google Scholar] [CrossRef] [PubMed]
- Jin, S.; Ye, K. Nanoparticle-Mediated Drug Delivery and Gene Therapy. Biotechnol. Prog. 2007, 23, 32–41. [Google Scholar] [CrossRef] [PubMed]
- Kay, M.A.; Glorioso, J.C.; Naldini, L. Viral vectors for gene therapy: The art of turning infectious agents into vehicles of therapeutics. Nat. Med. 2001, 7, 33–40. [Google Scholar] [CrossRef] [PubMed]
- Caffery, B.; Lee, J.; Alexander-Bryant, A. Vectors for Glioblastoma Gene Therapy: Viral & Non-Viral Delivery Strategies. Nanomaterials 2019, 9, 105. [Google Scholar] [CrossRef]
- Patra, J.K.; Das, G.; Fraceto, L.F.; Campos, E.V.R.; del Pilar Rodriguez-Torres, M.; Acosta-Torres, L.S.; Diaz-Torres, L.A.; Grillo, R.; Swamy, M.K.; Sharma, S.; et al. Nano based drug delivery systems: Recent developments and future prospects. J. Nanobiotechnol. 2018, 16. [Google Scholar] [CrossRef]
- Nakanishi, T.; Fukushima, S.; Okamoto, K.; Suzuki, M.; Matsumura, Y.; Yokoyama, M.; Okano, T.; Sakurai, Y.; Kataoka, K. Development of the polymer micelle carrier system for doxorubicin. J. Control. Release 2001, 74, 295–302. [Google Scholar] [CrossRef]
- Ngernyuang, N.; Seubwai, W.; Daduang, S.; Boonsiri, P.; Limpaiboon, T.; Daduang, J. Targeted delivery of 5-fluorouracil to cholangiocarcinoma cells using folic acid as a targeting agent. Mater. Sci. Eng. C 2016, 60, 411–415. [Google Scholar] [CrossRef]
- Chen, B.; Wang, J.; Chen, B.; Cheng, J.; Cai; Guohua, X.; Ran, D.L.; Wang, X. Apoptotic mechanism of human leukemia K562/A02 cells induced by magnetic iron oxide nanoparticles co-loaded with daunorubicin and 5-bromotetrandrin. Int. J. Nanomed. 2011, 1027. [Google Scholar] [CrossRef][Green Version]
- Gurumurthy, C.B.; Quadros, R.M.; Sato, M.; Mashimo, T.; Lloyd, K.C.K.; Ohtsuka, M. {CRISPR}/Cas9 and the Paradigm Shift in Mouse Genome Manipulation Technologies. In Genome Editing; Springer International Publishing: Cham, Switzerland, 2016; pp. 65–77. [Google Scholar]
- Seruggia, D.; Montoliu, L. CRISPR/Cas9 Approaches to Investigate the Noncoding Genome. In Genome Editing; Springer International Publishing: Cham, Switzerland, 2016; pp. 31–43. [Google Scholar]
- Dai, W.-J.; Zhu, L.-Y.; Yan, Z.-Y.; Xu, Y.; Wang, Q.-L.; Lu, X.-J. {CRISPR}-Cas9 for in vivo Gene Therapy: Promise and Hurdles. Mol. Ther. Nucleic Acids 2016, 5, e349. [Google Scholar] [CrossRef] [PubMed]
- Leal, A.F.; Espejo-Mojica, A.J.; Sánchez, O.F.; Ramírez, C.M.; Reyes, L.H.; Cruz, J.C.; Alméciga-Díaz, C.J. Lysosomal storage diseases: Current therapies and future alternatives. J. Mol. Med. 2020. [Google Scholar] [CrossRef] [PubMed]
- Ho, B.; Loh, S.; Chan, W.; Soh, B. In Vivo Genome Editing as a Therapeutic Approach. Int. J. Mol. Sci. 2018, 19, 2721. [Google Scholar] [CrossRef]
- Benskey, F.P.M.J. Viral Vectors for Gene Therapy; Manfredsson, F.P., Benskey, M.J., Eds.; Springer: New York, NY, USA, 2019. [Google Scholar]
- Martín, F.; Tristán-Manzano, M.; Maldonado-Pérez, N.; Sánchez-Hernández, S.; Benabdellah, K.; Cobo, M. Stable Genetic Modification of Mesenchymal Stromal Cells Using Lentiviral Vectors. In Methods in Molecular Biology; Springer: New York, NY, USA, 2019; pp. 267–280. [Google Scholar]
- Sandro, Q.; Relizani, K.; Benchaouir, R. AAV Production Using Baculovirus Expression Vector System. In Methods in Molecular Biology; Springer: New York, NY, USA, 2019; pp. 91–99. [Google Scholar]
- Hidai, C.; Kitano, H. Nonviral Gene Therapy for Cancer: A Review. Diseases 2018, 6, 57. [Google Scholar] [CrossRef] [PubMed]
- Agarwal, S.; Zhang, Y.; Maji, S.; Greiner, A. PDMAEMA based gene delivery materials. Mater. Today 2012, 15, 388–393. [Google Scholar] [CrossRef]
- Lee, J.; Lee, S.; Kwon, Y.-E.; Kim, Y.-J.; Choi, J.S. Gene Delivery by {PAMAM} Dendrimer Conjugated with the Nuclear Localization Signal Peptide Derived from Influenza B Virus Nucleoprotein. Macromol. Res. 2019, 27, 360–368. [Google Scholar] [CrossRef]
- Lee, J.; Oh, J.; Lee, E.; Kim, Y.-P.; Lee, M. Conjugation of prostate cancer-specific aptamers to polyethylene glycol-grafted polyethylenimine for enhanced gene delivery to prostate cancer cells. J. Ind. Eng. Chem. 2019, 73, 182–191. [Google Scholar] [CrossRef]
- You, Y.-Z.; Manickam, D.S.; Zhou, Q.-H.; Oupický, D. Reducible poly(2-dimethylaminoethyl methacrylate): Synthesis cytotoxicity, and gene delivery activity. J. Control. Release 2007, 122, 217–225. [Google Scholar] [CrossRef]
- Zhao, X.; Deng, L.; Deng, H.; Dong, A.; Wang, W.; Zhang, J. In Situ Template Polymerization to Prepare Liposome-Coated {PDMAEMA} Nanogels with Controlled Size High Stability, Low Cytotoxicity, and Responsive Drug Release for Intracellular {DOX} Release. Macromol. Chem. Phys. 2018, 219, 1800071. [Google Scholar] [CrossRef]
- Reis, C.P.; Neufeld, R.J.; Ribeiro, A.J.; Veiga, F. Nanoencapsulation I. Methods for preparation of drug-loaded polymeric nanoparticles. Nanomed. Nanotechnol. Biol. Med. 2006, 2, 8–21. [Google Scholar] [CrossRef]
- Majewski, A.P.; Schallon, A.; Jérôme, V.; Freitag, R.; Müller, A.H.E.; Schmalz, H. Dual-Responsive Magnetic Core{\textendash}Shell Nanoparticles for Nonviral Gene Delivery and Cell Separation. Biomacromolecules 2012, 13, 857–866. [Google Scholar] [CrossRef] [PubMed]
- Gass, S.; Cohen, J.M.; Pyrgiotakis, G.; Sotiriou, G.A.; Pratsinis, S.E.; Demokritou, P. Safer Formulation Concept for Flame-Generated Engineered Nanomaterials. ACS Sustain. Chem. Eng. 2013, 1, 843–857. [Google Scholar] [CrossRef] [PubMed][Green Version]
- Dai, L.; Song, L.; Huang, Y.; Zhang, L.; Lu, X.; Zhang, J.; Chen, T. Bimetallic Au/Ag Core{\textendash}Shell Superstructures with Tunable Surface Plasmon Resonance in the Near-Infrared Region and High Performance Surface-Enhanced Raman Scattering. Langmuir 2017, 33, 5378–5384. [Google Scholar] [CrossRef]
- Chaudhuri, R.G.; Paria, S. Core/Shell Nanoparticles: Classes Properties, Synthesis Mechanisms, Characterization, and Applications. Chem. Rev. 2011, 112, 2373–2433. [Google Scholar] [CrossRef] [PubMed]
- Varadan, V.K.; Chen, L.; Xie, J. Nanomedicine: Design and Applications of Magnetic Nanomaterials Nanosensors and Nanosystems; Wiley: Chichester, UK, 2008; p. 467. [Google Scholar]
- Cao, W.-P.; Ren, Q.-B.; Luo, M.-B. Translocation of polymers into crowded media with dynamic attractive nanoparticles. Phys. Rev. E 2015, 92. [Google Scholar] [CrossRef]
- Kumar, M.N.V.R.; Sameti, M.; Mohapatra, S.S.; Kong, X.; Lockey, R.F.; Bakowsky, U.; Lindenblatt, G.; Schmidt, C.H.; Lehr, C.-M. Cationic Silica Nanoparticles as Gene Carriers: Synthesis Characterization and Transfection Efficiency in vitro and in vivo. J. Nanosci. Nanotechnol. 2004, 4, 876–881. [Google Scholar] [CrossRef]
- Lee, K.; Conboy, M.; Park, H.M.; Jiang, F.; Kim, H.J.; Dewitt, M.A.; Mackley, V.A.; Chang, K.; Rao, A.; Skinner, C.; et al. Nanoparticle delivery of Cas9 ribonucleoprotein and donor DNA in vivo induces homology-directed {DNA} repair. Nat. Biomed. Eng. 2017, 1, 889–901. [Google Scholar] [CrossRef]
- Liu, Z.; Winters, M.; Holodniy, M.; Dai, H. {siRNA} Delivery into Human T{\hspace{0.25em}}Cells and Primary Cells with Carbon-Nanotube Transporters. Angew. Chemie Int. Ed. 2007, 46, 2023–2027. [Google Scholar] [CrossRef]
- Cuellar, M.; Cifuentes, J.; Perez, J.; Suarez-Arnedo, A.; Serna, J.A.; Groot, H.; Muñoz-Camargo, C.; Cruz, J.C. Novel BUF2-magnetite nanobioconjugates with cell-penetrating abilities. Int. J. Nanomed. 2018, 13, 8087–8094. [Google Scholar] [CrossRef]
- Patil, Y.P.; Jadhav, S. Novel methods for liposome preparation. Chem. Phys. Lipids 2014, 177, 8–18. [Google Scholar] [CrossRef]
- Philip, D. Honey mediated green synthesis of silver nanoparticles. Spectrochim. Acta Part A Mol. Biomol. Spectrosc. 2010, 75, 1078–1081. [Google Scholar] [CrossRef]
- Schindelin, J.; Arganda-Carreras, I.; Frise, E.; Kaynig, V.; Longair, M.; Pietzsch, T.; Preibisch, S.; Rueden, C.; Saalfeld, S.; Schmid, B.; et al. Fiji: An open-source platform for biological-image analysis. Nat. Methods 2012, 9, 676–682. [Google Scholar] [CrossRef] [PubMed]
- Cheng, Z.; Tan, A.L.K.; Tao, Y.; Shan, D.; Ting, K.E.; Yin, X.J. Synthesis and Characterization of Iron Oxide Nanoparticles and Applications in the Removal of Heavy Metals from Industrial Wastewater. Int. J. Photoenergy 2012, 2012, 1–5. [Google Scholar] [CrossRef]
- Koo, K.N.; Ismail, A.F.; Othman, M.H.D.; Bidin, N.; Rahman, M.A. Preparation and characterization of superparamagnetic magnetite (Fe3O4) nanoparticles: A short review. Malaysian J. Fundam. Appl. Sci. 2019, 15, 23–31. [Google Scholar] [CrossRef]
- Matahum, J.S.; Su, C.-M.; Wang, W.-J.; Lou, S.-L.; Ger, T.-R. Effect of Surface Charge on the Uptake of Magnetic Nanoparticles in Mouse Fibroblast Cells. IEEE Magn. Lett. 2017, 8, 1–5. [Google Scholar] [CrossRef]
- Cherng, J.-Y.; van de Wetering, P.; Talsma, H.; Crommelin, D.J.A.; Hennink, W.E. Effect of size and serum proteins on transfection efficiency of poly ((2-dimethylamino)ethyl methacrylate)-plasmid nanoparticles. Pharm. Res. 1996, 13, 1038–1042. [Google Scholar] [CrossRef]
- Perez, J.; Cifuentes, J.; Cuellar, M.; Suarez-Arnedo, A.; Cruz, J.C.; Muñoz-Camargo, C. Cell-Penetrating And Antibacterial BUF-II Nanobioconjugates: Enhanced Potency Via Immobilization On Polyetheramine-Modified Magnetite Nanoparticles. Int. J. Nanomed. 2019, 14, 8483–8497. [Google Scholar] [CrossRef] [PubMed]
- Griesser, J.; Hetényi, G.; Federer, C.; Steinbring, C.; Ellemunter, H.; Niedermayr, K.; Bernkop-Schnürch, A. Highly mucus permeating and zeta potential changing self-emulsifying drug delivery systems: A potent gene delivery model for causal treatment of cystic fibrosis. Int. J. Pharm. 2019, 557, 124–134. [Google Scholar] [CrossRef] [PubMed]
- Rao, Y.S.; Kotakadi, V.S.; Prasad, T.N.V.K.V.; Reddy, A.V.; Gopal, D.V.R.S. Green synthesis and spectral characterization of silver nanoparticles from Lakshmi tulasi (Ocimum sanctum) leaf extract. Spectrochim. Acta Part A Mol. Biomol. Spectrosc. 2013, 103, 156–159. [Google Scholar] [CrossRef]
- Singh, S.; Bharti, A.; Meena, V.K. Structural thermal, zeta potential and electrical properties of disaccharide reduced silver nanoparticles. J. Mater. Sci. Mater. Electron. 2014, 25, 3747–3752. [Google Scholar] [CrossRef]
- Ling, D.; Lee, N.; Hyeon, T. Chemical synthesis and assembly of uniformly sized iron oxide nanoparticles for medical applications. Acc. Chem. Res. 2015, 48, 1276–1285. [Google Scholar] [CrossRef] [PubMed]
- Garza-Navarro, M.; Torres-Castro, A.; González, V.; Ortiz, U.; la Rosa, E. De Magnetite and magnetite/silver core/shell nanoparticles with diluted magnet-like behavior. J. Solid State Chem. 2010, 183, 99–104. [Google Scholar] [CrossRef]
- Li, Y.; Bolinger, J.; Yu, Y.; Glass, Z.; Shi, N.; Yang, L.; Wang, M.; Xu, Q. Intracellular delivery and biodistribution study of CRISPR/Cas9 ribonucleoprotein loaded bioreducible lipidoid nanoparticles. Biomater. Sci. 2019, 7, 596–606. [Google Scholar] [CrossRef] [PubMed]
- Meyer, F.; Minoia, A.; Raquez, J.M.; Spasova, M.; Lazzaroni, R.; Dubois, P. Poly(amino-methacrylate) as versatile agent for carbon nanotube dispersion: An experimental theoretical and application study. J. Mater. Chem. 2010, 20, 6873. [Google Scholar] [CrossRef]
- Liu, X.; Ni, P.; He, J.; Zhang, M. Synthesis and Micellization of pH/Temperature-Responsive Double-Hydrophilic Diblock Copolymers Polyphosphoester-block-poly[2-(dimethylamino)ethyl methacrylate] Prepared via {ROP} and {ATRP}. Macromolecules 2010, 43, 4771–4781. [Google Scholar] [CrossRef]
- Liu, X.; Liang, M.; Liu, M.; Su, R.; Wang, M.; Qi, W.; He, Z. Highly Efficient Catalysis of Azo Dyes Using Recyclable Silver Nanoparticles Immobilized on Tannic Acid-Grafted Eggshell Membrane. Nanoscale Res. Lett. 2016, 11. [Google Scholar] [CrossRef]
- AshaRani, P.V.; Mun, G.L.K.; Hande, M.P.; Valiyaveettil, S. Cytotoxicity and Genotoxicity of Silver Nanoparticles in Human Cells. ACS Nano 2008, 3, 279–290. [Google Scholar] [CrossRef]
- Asharani, P.V.; Wu, Y.L.; Gong, Z.; Valiyaveettil, S. Toxicity of silver nanoparticles in zebrafish models. Nanotechnology 2008, 19, 255102. [Google Scholar] [CrossRef]
- He, X.; Wu, X.; Cai, X.; Lin, S.; Xie, M.; Zhu, X.; Yan, D. Functionalization of Magnetic Nanoparticles with Dendritic–Linear–Brush-Like Triblock Copolymers and Their Drug Release Properties. Langmuir 2012, 28, 11929–11938. [Google Scholar] [CrossRef]
- Setyawati, M.I.; Yuan, X.; Xie, J.; Leong, D.T. The influence of lysosomal stability of silver nanomaterials on their toxicity to human cells. Biomaterials 2014, 35, 6707–6715. [Google Scholar] [CrossRef]
- Sanyasi, S.; Majhi, R.K.; Kumar, S.; Mishra, M.; Ghosh, A.; Suar, M.; Satyam, P.V.; Mohapatra, H.; Goswami, C.; Goswami, L. Polysaccharide-capped silver Nanoparticles inhibit biofilm formation and eliminate multi-drug-resistant bacteria by disrupting bacterial cytoskeleton with reduced cytotoxicity towards mammalian cells. Sci. Rep. 2016, 6. [Google Scholar] [CrossRef] [PubMed]
- Maleki, H.; Rai, A.; Pinto, S.; Evangelista, M.; Cardoso, R.M.S.; Paulo, C.; Carvalheiro, T.; Paiva, A.; Imani, M.; Simchi, A.; et al. High Antimicrobial Activity and Low Human Cell Cytotoxicity of Core–Shell Magnetic Nanoparticles Functionalized with an Antimicrobial Peptide. ACS Appl. Mater. Interfaces 2016, 8, 11366–11378. [Google Scholar] [CrossRef]
- De Oliveira, J.F.A.; Saito, Â.; Bido, A.T.; Kobarg, J.; Stassen, H.K.; Cardoso, M.B. Defeating Bacterial Resistance and Preventing Mammalian Cells Toxicity Through Rational Design of Antibiotic-Functionalized Nanoparticles. Sci. Rep. 2017, 7. [Google Scholar] [CrossRef]
- Lin, J.-J.; Lin, W.-C.; Dong, R.-X.; Hsu, S. The cellular responses and antibacterial activities of silver nanoparticles stabilized by different polymers. Nanotechnology 2012, 23, 65102. [Google Scholar] [CrossRef] [PubMed]
- Lale, S.V.; Kumar, A.; Naz, F.; Bharti, A.C.; Koul, V. Multifunctional ATRP based pH responsive polymeric nanoparticles for improved doxorubicin chemotherapy in breast cancer by proton sponge effect/endo-lysosomal escape. Polym. Chem. 2015, 6, 2115–2132. [Google Scholar] [CrossRef]
- Cerda-Cristerna, B.I.; Flores, H.; Pozos-Guillén, A.; Pérez, E.; Sevrin, C.; Grandfils, C. Hemocompatibility assessment of poly(2-dimethylamino ethylmethacrylate) ({PDMAEMA})-based polymers. J. Control. Release 2011, 153, 269–277. [Google Scholar] [CrossRef]
- Kwon, T.; Woo, H.J.; Kim, Y.H.; Lee, H.J.; Park, K.H.; Park, S.; Youn, B. Optimizing Hemocompatibility of Surfactant-Coated Silver Nanoparticles in Human Erythrocytes. J. Nanosci. Nanotechnol. 2012, 12, 6168–6175. [Google Scholar] [CrossRef]
- Jun, E.-A.; Lim, K.-M.; Kim, K.; Bae, O.-N.; Noh, J.-Y.; Chung, K.-H.; Chung, J.-H. Silver nanoparticles enhance thrombus formation through increased platelet aggregation and procoagulant activity. Nanotoxicology 2010, 5, 157–167. [Google Scholar] [CrossRef]
- Vorobjev, I.A.; Barteneva, N.S. Multi-parametric imaging of cell heterogeneity in apoptosis analysis. Methods 2017, 112, 105–123. [Google Scholar] [CrossRef]
- Park, B.S.; Song, Y.S.; Yee, S.-B.; Lee, B.G.; Seo, S.Y.; Park, Y.C.; Kim, J.-M.; Kim, H.M.; Yoo, Y.H. Phospho-ser 15-p53 translocates into mitochondria and interacts with Bcl-2 and Bcl-xL in eugenol-induced apoptosis. Apoptosis 2005, 10, 193–200. [Google Scholar] [CrossRef]
- Choi, J.; Reipa, V.; Hitchins, V.M.; Goering, P.L.; Malinauskas, R.A. Physicochemical Characterization and In Vitro Hemolysis Evaluation of Silver Nanoparticles. Toxicol. Sci. 2011, 123, 133–143. [Google Scholar] [CrossRef] [PubMed]
- Nair, B.G.; Fukuda, T.; Mizuki, T.; Hanajiri, T.; Maekawa, T. Intracellular trafficking of superparamagnetic iron oxide nanoparticles conjugated with {TAT} peptide: 3-dimensional electron tomography analysis. Biochem. Biophys. Res. Commun. 2012, 421, 763–767. [Google Scholar] [CrossRef] [PubMed]
- Schweiger, C.; Hartmann, R.; Zhang, F.; Parak, W.J.; Kissel, T.H.; Rivera_Gil, P. Quantification of the internalization patterns of superparamagnetic iron oxide nanoparticles with opposite charge. J. Nanobiotechnol. 2012, 10, 28. [Google Scholar] [CrossRef] [PubMed]
- Ren, Q.-B.; Ma, S.-H.; Chen, Y.-J.; Sun, L.-Z.; Cao, W.-P. Numerical simulation on polymer translocation into crowded environment with nanoparticles. Colloid Polym. Sci. 2016, 294, 1351–1357. [Google Scholar] [CrossRef]
- Kim, H.R.; Andrieux, K.; Gil, S.; Taverna, M.; Chacun, H.; Desmaële, D.; Taran, F.; Georgin, D.; Couvreur, P. Translocation of Poly(ethylene glycol-co-hexadecyl)cyanoacrylate Nanoparticles into Rat Brain Endothelial Cells:~ Role of Apolipoproteins in Receptor-Mediated Endocytosis. Biomacromolecules 2007, 8, 793–799. [Google Scholar] [CrossRef]
- Hu, G.; Jiao, B.; Shi, X.; Valle, R.P.; Fan, Q.; Zuo, Y.Y. Physicochemical Properties of Nanoparticles Regulate Translocation across Pulmonary Surfactant Monolayer and Formation of Lipoprotein Corona. ACS Nano 2013, 7, 10525–10533. [Google Scholar] [CrossRef]
- Koren, E.; Torchilin, V.P. Cell-penetrating peptides: Breaking through to the other side. Trends Mol. Med. 2012, 18, 385–393. [Google Scholar] [CrossRef]
- Ayala, V.; Herrera, A.P.; Latorre-Esteves, M.; Torres-Lugo, M.; Rinaldi, C. Effect of surface charge on the colloidal stability and in vitro uptake of carboxymethyl dextran-coated iron oxide nanoparticles. J. Nanoparticle Res. 2013, 15. [Google Scholar] [CrossRef]
- Mayor, S.; Parton, R.G.; Donaldson, J.G. Clathrin-independent pathways of endocytosis. Cold Spring Harb. Perspect. Biol. 2014, 6, a016758. [Google Scholar] [CrossRef]
- Thapa, R.K.; Sullivan, M.O. Gene delivery by peptide-assisted transport. Curr. Opin. Biomed. Eng. 2018, 7, 71–82. [Google Scholar] [CrossRef]
- Freeman, E.C.; Weiland, L.M.; Meng, W.S. Modeling the proton sponge hypothesis: Examining proton sponge effectiveness for enhancing intracellular gene delivery through multiscale modeling. J. Biomater. Sci. Polym. Ed. 2012, 24, 398–416. [Google Scholar] [CrossRef] [PubMed]
- Wiradharma, N.; Khan, M.; Tong, Y.W.; Wang, S.; Yang, Y.-Y. Self-assembled Cationic Peptide Nanoparticles Capable of Inducing Efficient Gene Expression In Vitro. Adv. Funct. Mater. 2008, 18, 943–951. [Google Scholar] [CrossRef]
- Chen, P.; Chen, B.; Pan, R.; Askhatova, D. Effective small interfering {RNA} delivery in vitro via a new stearylated cationic peptide. Int. J. Nanomed. 2015, 3303. [Google Scholar] [CrossRef]
- Degors, I.M.S.; Wang, C.; Rehman, Z.U.; Zuhorn, I.S. Carriers Break Barriers in Drug Delivery: Endocytosis and Endosomal Escape of Gene Delivery Vectors. Acc. Chem. Res. 2019, 52, 1750–1760. [Google Scholar] [CrossRef]
- Zheng, C.; Xu, J.; Yao, X.; Xu, J.; Qiu, L. Polyphosphazene nanoparticles for cytoplasmic release of doxorubicin with improved cytotoxicity against Dox-resistant tumor cells. J. Colloid Interface Sci. 2011, 355, 374–382. [Google Scholar] [CrossRef] [PubMed]
- Garcia-Garcia, E.; Andrieux, K.; Gil, S.; Kim, H.R.; Le Doan, T.; Desmaële, D.; d’Angelo, J.; Taran, F.; Georgin, D.; Couvreur, P. A methodology to study intracellular distribution of nanoparticles in brain endothelial cells. Int. J. Pharm. 2005, 298, 310–314. [Google Scholar] [CrossRef]
- Hackenberg, S.; Scherzed, A.; Kessler, M.; Hummel, S.; Technau, A.; Froelich, K.; Ginzkey, C.; Koehler, C.; Hagen, R.; Kleinsasser, N. Silver nanoparticles: Evaluation of {DNA} damage toxicity and functional impairment in human mesenchymal stem cells. Toxicol. Lett. 2011, 201, 27–33. [Google Scholar] [CrossRef] [PubMed]
© 2020 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Ramírez-Acosta, C.M.; Cifuentes, J.; Castellanos, M.C.; Moreno, R.J.; Muñoz-Camargo, C.; Cruz, J.C.; Reyes, L.H. PH-Responsive, Cell-Penetrating, Core/Shell Magnetite/Silver Nanoparticles for the Delivery of Plasmids: Preparation, Characterization, and Preliminary In Vitro Evaluation. Pharmaceutics 2020, 12, 561. https://doi.org/10.3390/pharmaceutics12060561
Ramírez-Acosta CM, Cifuentes J, Castellanos MC, Moreno RJ, Muñoz-Camargo C, Cruz JC, Reyes LH. PH-Responsive, Cell-Penetrating, Core/Shell Magnetite/Silver Nanoparticles for the Delivery of Plasmids: Preparation, Characterization, and Preliminary In Vitro Evaluation. Pharmaceutics. 2020; 12(6):561. https://doi.org/10.3390/pharmaceutics12060561
Chicago/Turabian StyleRamírez-Acosta, Carlos M., Javier Cifuentes, Maria Claudia Castellanos, Rodolfo José Moreno, Carolina Muñoz-Camargo, Juan C. Cruz, and Luis H. Reyes. 2020. "PH-Responsive, Cell-Penetrating, Core/Shell Magnetite/Silver Nanoparticles for the Delivery of Plasmids: Preparation, Characterization, and Preliminary In Vitro Evaluation" Pharmaceutics 12, no. 6: 561. https://doi.org/10.3390/pharmaceutics12060561
APA StyleRamírez-Acosta, C. M., Cifuentes, J., Castellanos, M. C., Moreno, R. J., Muñoz-Camargo, C., Cruz, J. C., & Reyes, L. H. (2020). PH-Responsive, Cell-Penetrating, Core/Shell Magnetite/Silver Nanoparticles for the Delivery of Plasmids: Preparation, Characterization, and Preliminary In Vitro Evaluation. Pharmaceutics, 12(6), 561. https://doi.org/10.3390/pharmaceutics12060561







