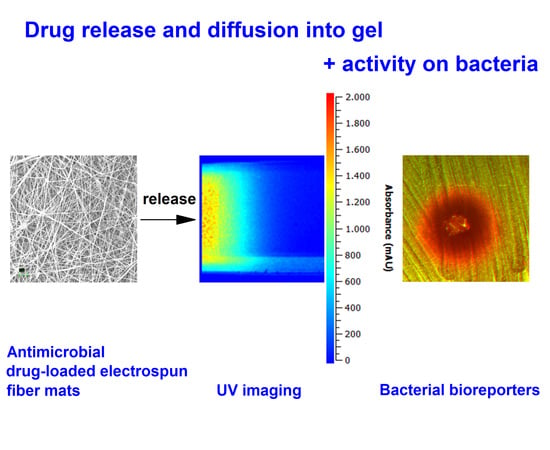
Monitoring of Antimicrobial Drug Chloramphenicol Release from Electrospun Nano- and Microfiber Mats Using UV Imaging and Bacterial Bioreporters
Abstract
1. Introduction
2. Materials and Methods
2.1. Materials, Bacteria and Release Media
2.2. Preparation and Characterization Methods
2.3. Drug Release Studies
2.3.1. Drug Release Testing into Buffer Solution and Agar Hydrogel by UV-VIS Spectrophotometry and HPLC
2.3.2. UV Imaging for Drug Release Monitoring in Hydrogel
2.4. Antibacterial Activity Studies—Drug Release and Effect on Bacterial Growth
2.4.1. Antibacterial Activity Testing
2.4.2. Bioreporter Plasmid and Strain Preparation
2.4.3. Bioreporter Disc Diffusion Assay
2.5. Data Analysis
3. Results and Discussion
3.1. Preparation and Characterization of Electrospun Fiber Mats
3.2. Drug Release into Buffer Measured by UV-VIS Spectrophotometry
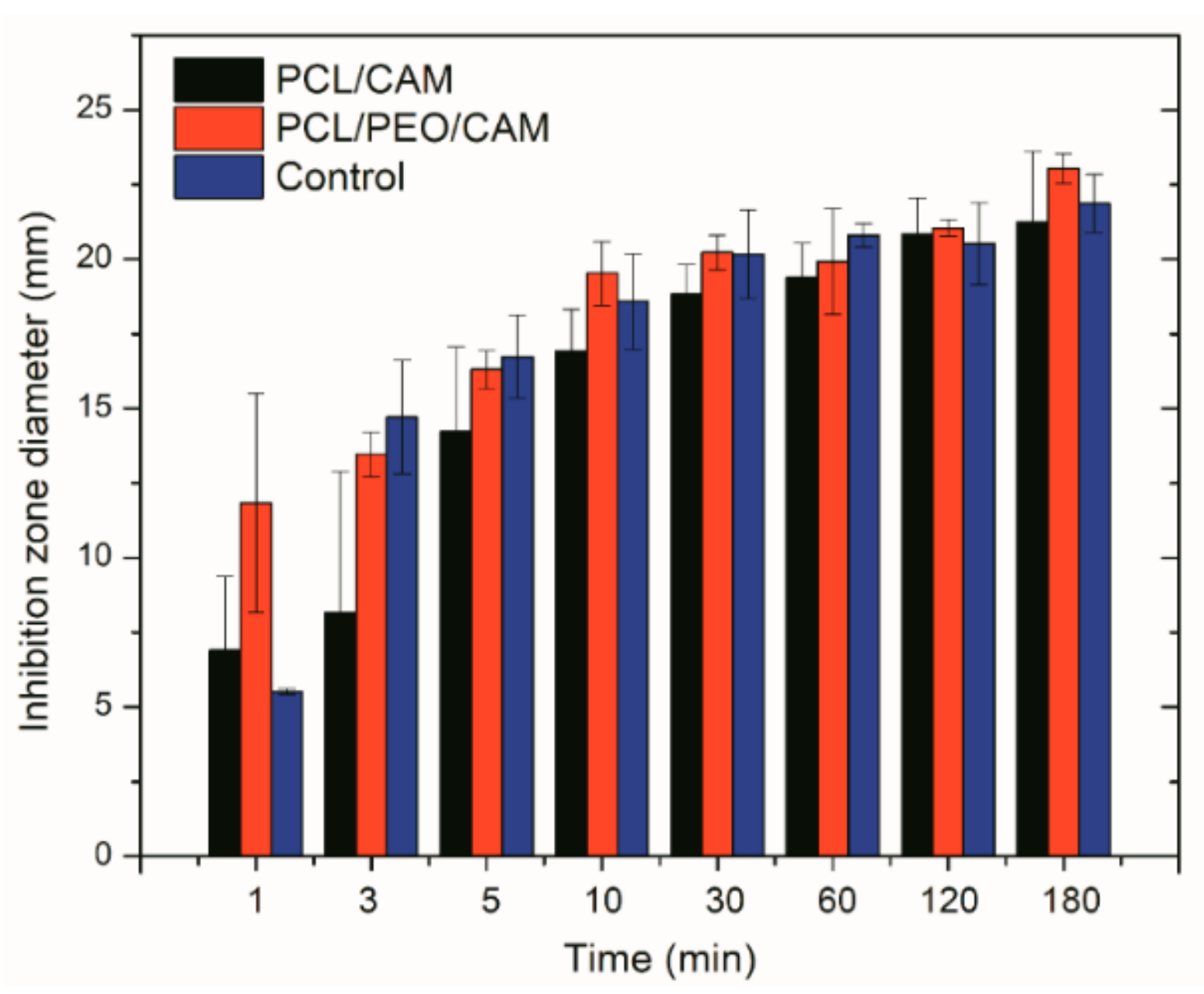
3.3. Antibacterial Activity of Electrospun Fiber Mats
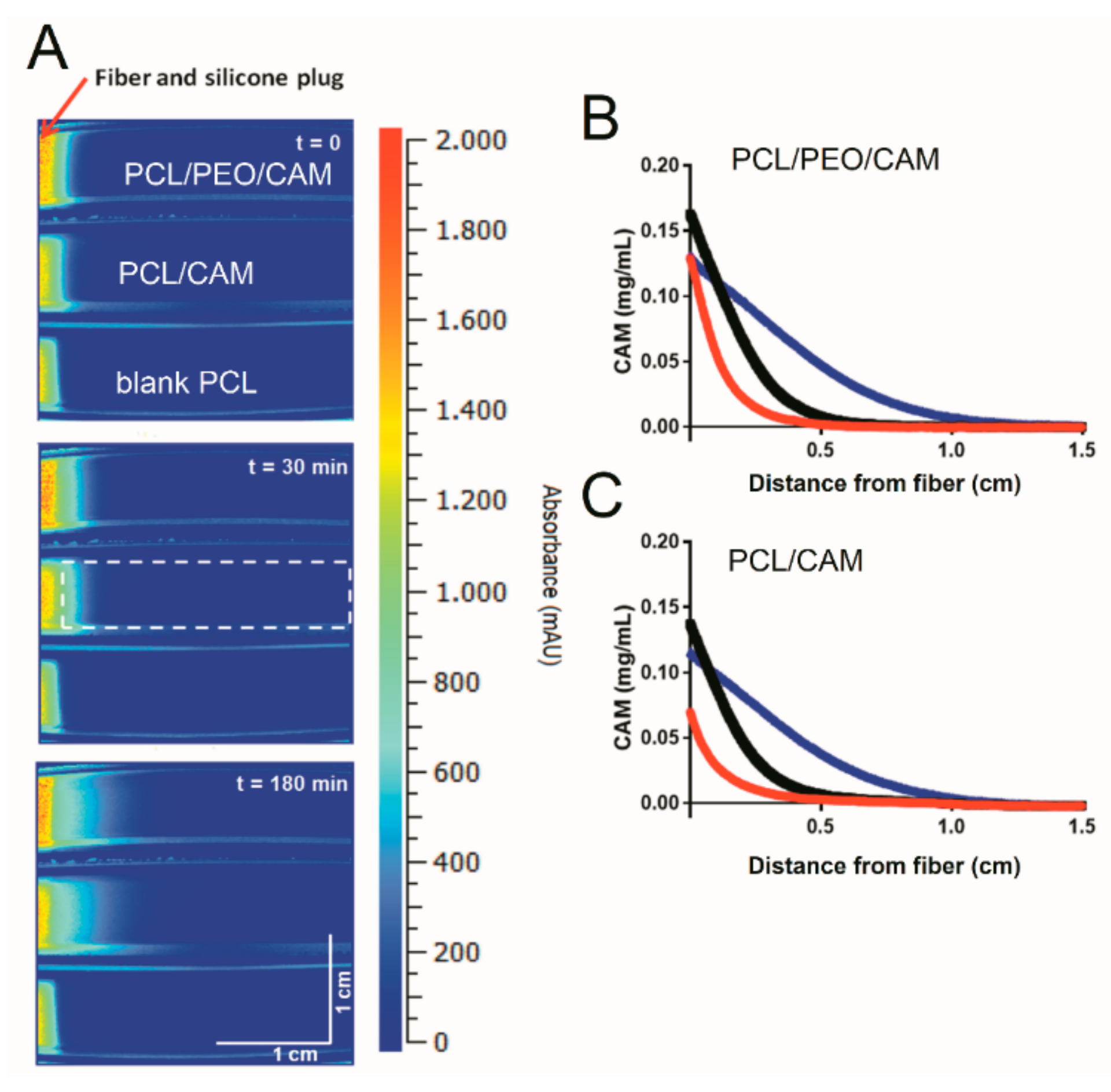
3.4. Drug Release into Agar Hydrogel Measured by HPLC
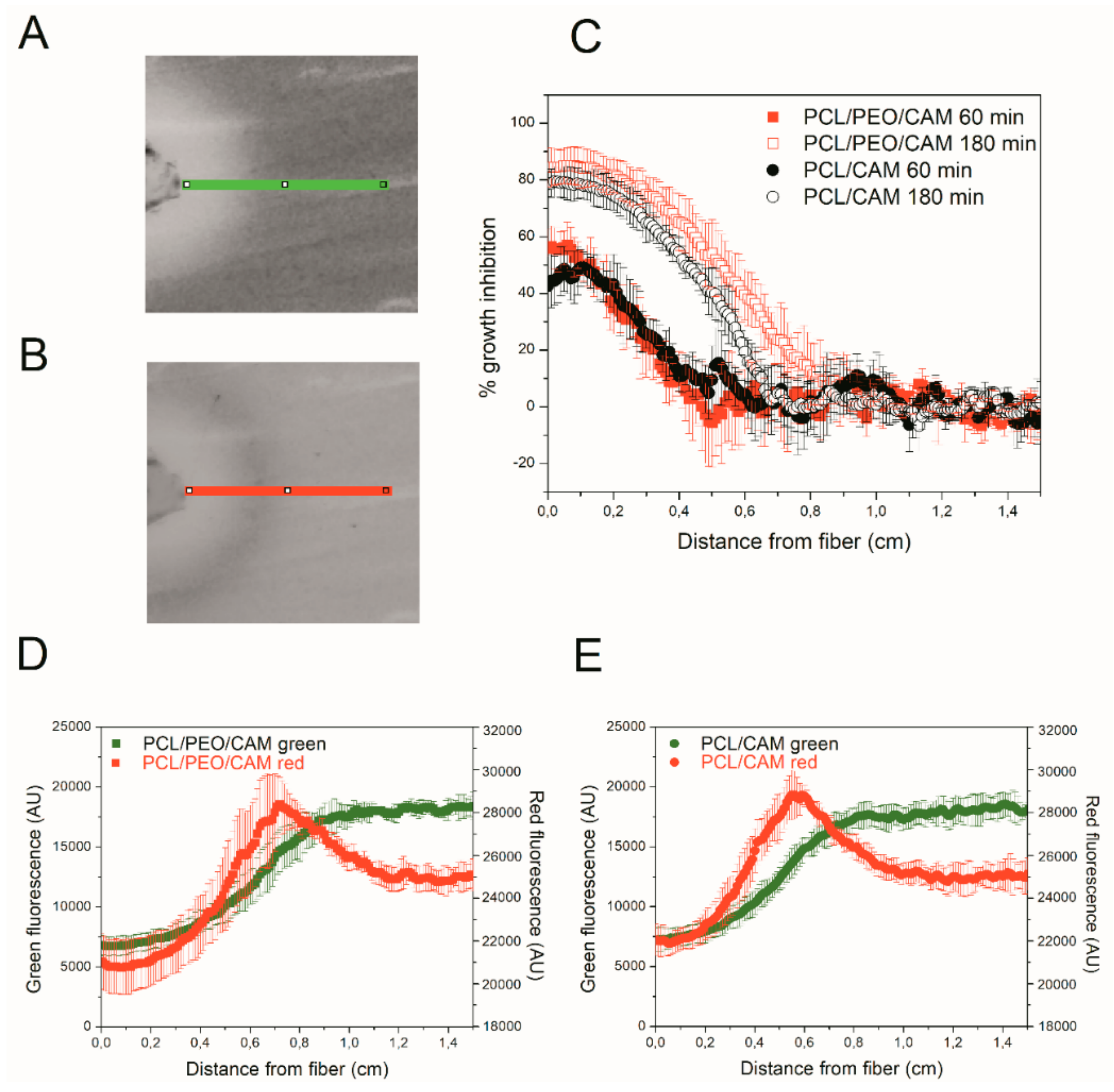
3.5. Detecting CAM Release into Agar Hydrogel with Bioreporter Strain
3.6. Drug Release into Agarose Hydrogel Using UV Imaging
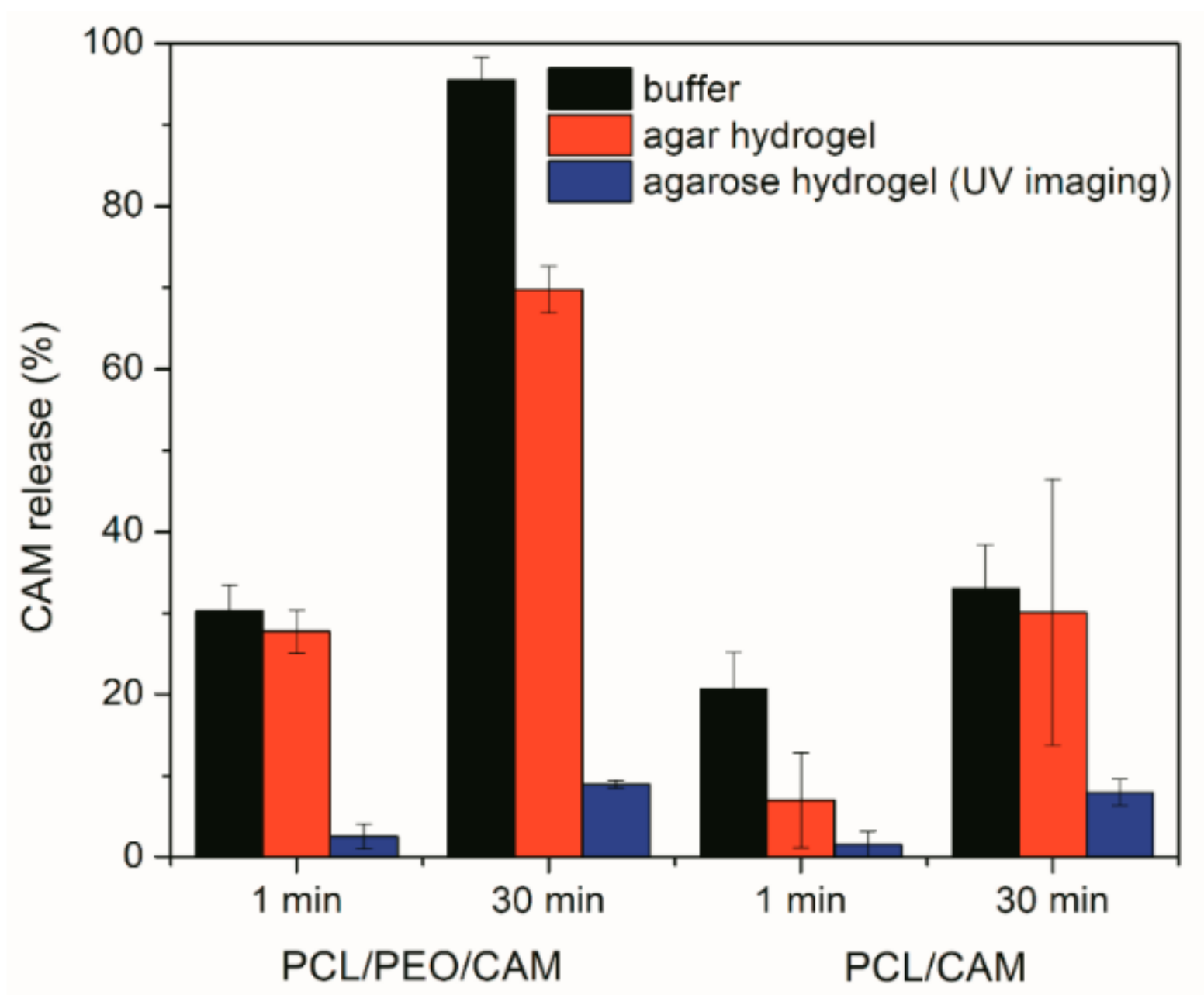
3.7. Comparison between Different Drug Release Model Systems
4. Summary and Conclusions
Supplementary Materials
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
Appendix A. Drug release Into Agar Hydrogel
| mg/Samples | % Extracted | ||
|---|---|---|---|
| 0.119 | 66.6% | E1 | 1x extraction |
| 0.122 | 68.2% | E2 | |
| 0.119 | 66.6% | E3 | |
| 67.1% | |||
| 0.186 | 103.9% | E1 | 2x extraction |
| 0.191 | 106.8% | E2 | |
| 0.177 | 98.9% | E3 | |
| 103.2% | |||
| Zone | 5 min | 10 min | 15 min | 30 min | 60 min | 120 min | 1440 min |
|---|---|---|---|---|---|---|---|
| 1 | 86.8 | 80.6 | 78.1 | 88.5 | 66.6 | 62.6 | 11.8 |
| 2 | 13.2 | 19.4 | 21.9 | 11.5 | 33.4 | 37.4 | 54.6 |
| 3 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | 26.9 |
| 4 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | 6.3 |
| 5 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | 0.4 |
Appendix B
Appendix C
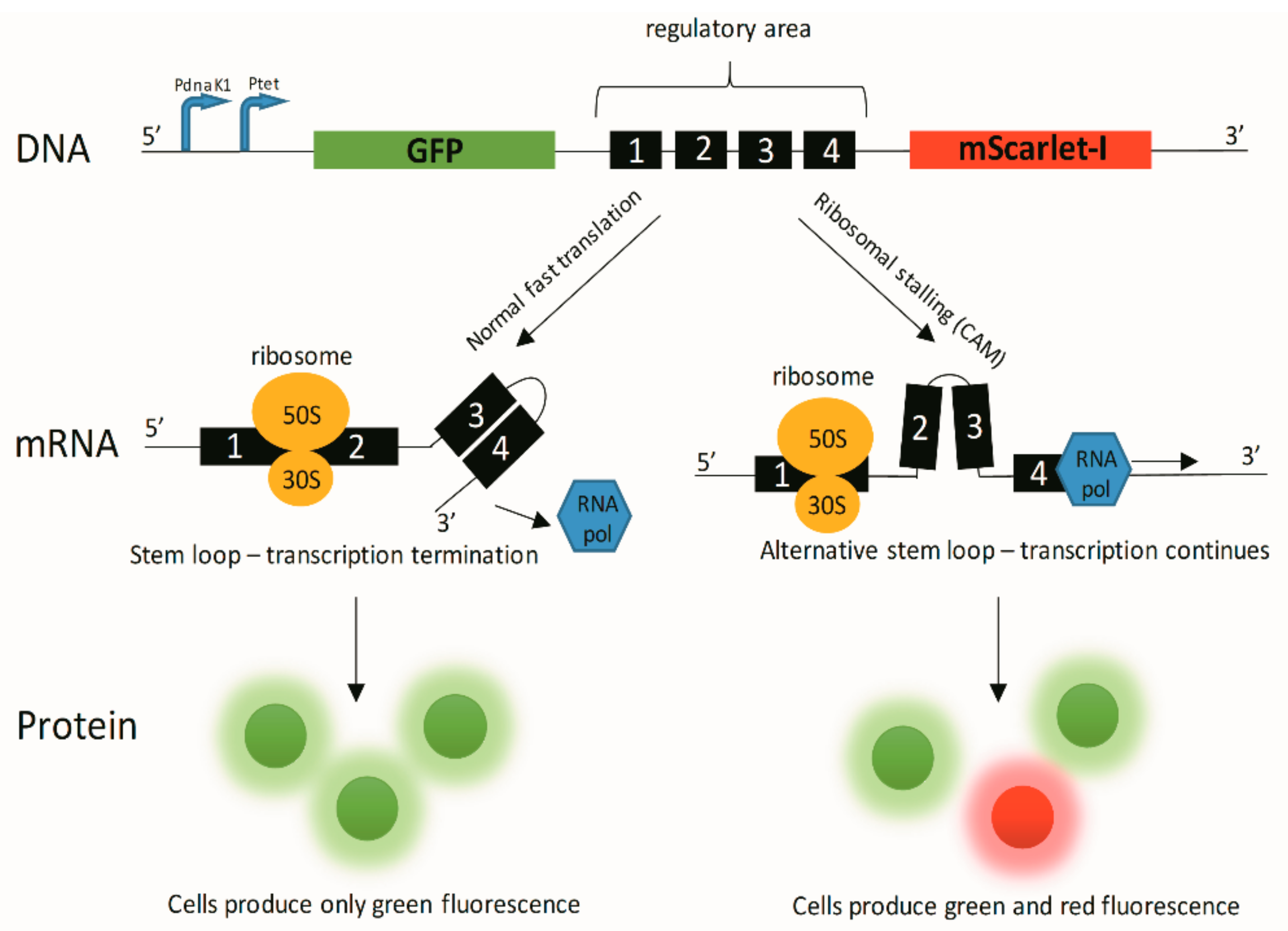
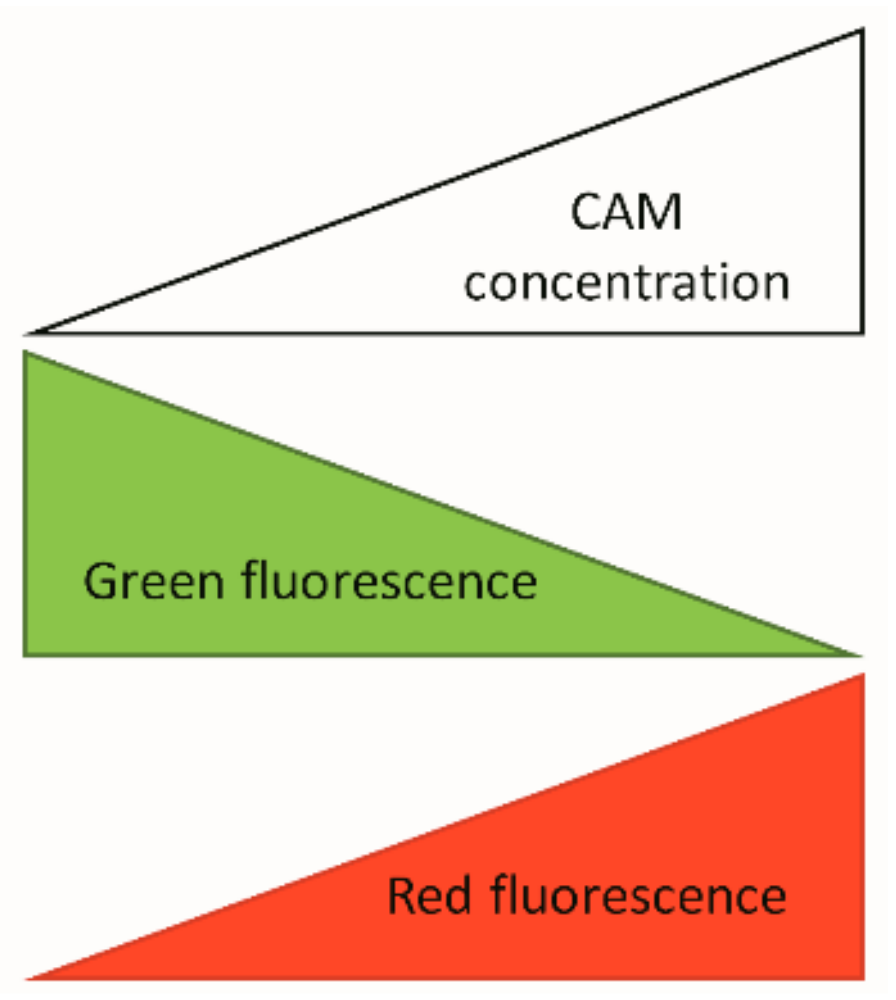
References
- Weng, L.; Xie, J. Smart electrospun nanofibers for controlled drug release: Recent advances and new perspectives. Curr. Pharm. Des. 2015, 21, 1944–1959. [Google Scholar] [CrossRef] [PubMed]
- Torres-Martinez, E.J.; Cornejo Bravo, J.M.; Serrano Medina, A.; Pérez González, G.L.; Villarreal Gómez, L.J. A summary of electrospun nanofibers as drug delivery system: Drugs loaded and biopolymers used as matrices. Curr. Drug Deliv. 2018, 15, 1360–1374. [Google Scholar] [CrossRef] [PubMed]
- Sill, T.J.; von Recum, H.A. Electrospinning: Applications in drug delivery and tissue engineering. Biomaterials 2008, 29, 1989–2006. [Google Scholar] [CrossRef] [PubMed]
- Ye, K.; Kuang, H.; You, Z.; Morsi, Y.; Mo, X. Electrospun nanofibers for tissue engineering with drug loading and release. Pharmaceutics 2019, 11, 182. [Google Scholar] [CrossRef] [PubMed]
- Liu, Y.; He, J.-H. Bubble electrospinning for mass production of nanofibers. Int. J. Nonlinear Sci. Numer. Simul. 2007, 8, 393–396. [Google Scholar] [CrossRef]
- Sultanova, Z.; Kaleli, G.; Kabay, G.; Mutlu, M. Controlled release of a hydrophilic drug from coaxially electrospun polycaprolactone nanofibers. Int. J. Pharm. 2016, 505, 133–138. [Google Scholar] [CrossRef] [PubMed]
- Thakkar, S.; Misra, M. Electrospun polymeric nanofibers: New horizons in drug delivery. Eur. J. Pharm. Sci. 2017, 107, 148–167. [Google Scholar] [CrossRef]
- Gao, S.; Tang, G.; Hua, D.; Xiong, R.; Han, J.; Jiang, S.; Zhang, Q.; Huang, C. Stimuli-responsive bio-based polymeric systems and their applications. J. Mater. Chem. B 2019, 7, 709–729. [Google Scholar] [CrossRef]
- Chou, S.F.; Carson, D.; Woodrow, K.A. Current strategies for sustaining drug release from electrospun nanofibers. J. Control. Release 2015, 220, 584–591. [Google Scholar] [CrossRef]
- Singh, A.; Rath, G.; Singh, R.; Goyal, A.K. Nanofibers: An effective tool for controlled and sustained drug delivery. Curr. Drug Deliv. 2018, 15, 155–166. [Google Scholar] [CrossRef]
- Blanchemain, N.; Laurent, T.; Chai, F.; Neut, C.; Haulon, S.; Krump-konvalinkova, V.; Morcellet, M.; Martel, B.; Kirkpatrick, C.J.; Hildebrand, H.F. Polyester vascular prostheses coated with a cyclodextrin polymer and activated with antibiotics: Cytotoxicity and microbiological evaluation. Acta Biomater. 2008, 4, 1725–1733. [Google Scholar] [CrossRef]
- Liu, X.; Shao, W.; Luo, M.; Bian, J.; Yu, D.-G. Electrospun blank nanocoating for improved sustained release profiles from medicated gliadin nanofibers. Nanomaterials 2018, 8, 184. [Google Scholar]
- Chew, S.Y.; Wen, J.; Yim, E.K.; Leong, K.W. Sustained release of proteins from electrospun biodegradable fibers. Biomacromolecules 2005, 6, 2017–2024. [Google Scholar] [CrossRef] [PubMed]
- Gaharwar, A.K.; Mihaila, S.M.; Kulkarni, A.A.; Patel, A.; Di Luca, A.; Reis, R.L.; Gomes, M.E.; Van Blitterswijk, C.; Moroni, L.; Khademhosseini, A. Amphiphilic beads as depots for sustained drug release integrated into fibrillar scaffolds. J. Control. Release 2014, 187, 66–73. [Google Scholar] [CrossRef] [PubMed]
- Zupančič, Š.; Preem, L.; Kristl, J.; Putrinš, M.; Tenson, T.; Kocbek, P.; Kogermann, K. Impact of PCL nanofiber mat structural properties on hydrophilic drug release and antibacterial activity on periodontal pathogens. Eur. J. Pharm. Sci. 2018, 122, 347–358. [Google Scholar] [CrossRef]
- Fu, Y.; Kao, W.J. Drug release kinetics and transport mechanisms of non-degradable and degradable polymeric delivery systems. Expert Opin. Drug Deliv. 2010, 7, 429–444. [Google Scholar] [CrossRef] [PubMed]
- Natu, M.V.; de Sousa, H.C.; Gil, M.H. Effects of drug solubility, state and loading on controlled release in bicomponent electrospun fibers. Int. J. Pharm. 2010, 397, 50–58. [Google Scholar] [CrossRef] [PubMed]
- Chou, S.-F.; Woodrow, K.A. Relationships between mechanical properties and drug release from electrospun fibers of PCL and PLGA blends. J. Mech. Behav. Biomed. Mater. 2017, 65, 724–733. [Google Scholar] [CrossRef] [PubMed]
- Brettmann, B.K.; Cheng, K.; Myerson, A.S.; Trout, B.L. Electrospun formulations containing crystalline active pharmaceutical ingredients. Pharm. Res. 2013, 30, 238–246. [Google Scholar] [CrossRef]
- Radacsi, N.; Giapis, K.P.; Ovari, G.; Szabó-Révész, P.; Ambrus, R. Electrospun nanofiber-based niflumic acid capsules with superior physicochemical properties. J. Pharm. Biomed. Anal. 2019, 166, 371–378. [Google Scholar] [CrossRef]
- Paskiabi, F.A.; Bonakdar, S.; Shokrgozar, M.A.; Imani, M.; Jahanshiri, Z.; Shams-Ghahfarokhi, M.; Razzaghi-Abyaneh, M. Terbinafine-loaded wound dressing for chronic superficial fungal infections. Mater. Sci. Eng. C 2017, 73, 130–136. [Google Scholar] [CrossRef] [PubMed]
- Preem, L.; Mahmoudzadeh, M.; Putrinš, M.; Meos, A.; Laidmäe, I.; Romann, T.; Aruväli, J.; Härmas, R.; Koivuniemi, A.; Bunker, A.; et al. Interactions between chloramphenicol, carrier polymers, and bacteria–implications for designing electrospun drug delivery systems countering wound infection. Mol. Pharm. 2017, 14, 4417–4430. [Google Scholar] [CrossRef] [PubMed]
- Hu, J.; Prabhakaran, M.P.; Tian, L.; Ding, X.; Ramakrishna, S. Drug-loaded emulsion electrospun nanofibers: Characterization, drug release and in vitro biocompatibility. RSC Adv. 2015, 5, 100256–100267. [Google Scholar] [CrossRef]
- Fu, Y.; Kao, W.J. Drug release kinetics and transport mechanisms from semi-interpenetrating networks of gelatin and poly (ethylene glycol) diacrylate. Pharm. Res. 2009, 26, 2115–2124. [Google Scholar] [CrossRef] [PubMed]
- Srikar, R.; Yarin, A.L.; Megaridis, C.M.; Bazilevsky, A.V.; Kelley, E. Desorption-limited mechanism of release from polymer nanofibers. Langmuir 2008, 24, 965–974. [Google Scholar] [CrossRef] [PubMed]
- Nakielski, P.; Kowalczyk, T.; Zembrzycki, K.; Kowalewski, T.A. Experimental and numerical evaluation of drug release from nanofiber mats to brain tissue. J. Biomed. Mater. Res. Part B Appl. Biomater. 2015, 103, 282–291. [Google Scholar] [CrossRef] [PubMed]
- Seif, S.; Graef, F.; Gordon, S.F.; Windbergs, M. Monitoring drug release from electrospun fibers using an in situ fiber-optic system Diss. Technol. 2016, 23, 6–12. [Google Scholar]
- Østergaard, J. UV imaging in pharmaceutical analysis. J. Pharm. Biomed. Anal. 2018, 147, 140–148. [Google Scholar] [CrossRef]
- Asare-Addo, K.; Walton, K.; Ward, A.; Totea, A.-M.; Taheri, S.; Alshafiee, M.; Mawla, N.; Bondi, A.; Evans, W.; Adebisi, A.; et al. Direct imaging of the dissolution of salt forms of a carboxylic acid drug. Int. J. Pharm. 2018, 551, 290–299. [Google Scholar] [CrossRef]
- Østergaard, J.; Meng-Lund, E.; Larsen, S.W.; Larsen, C.; Petersson, K.; Lenke, J.; Jensen, H. Real-time UV imaging of nicotine release from transdermal patch. Pharm. Res. 2010, 27, 2614–2623. [Google Scholar] [CrossRef]
- Ye, F.; Yaghmur, A.; Jensen, H.; Larsen, S.W.; Larsen, C.; Østergaard, J. Real-time UV imaging of drug diffusion and release from Pluronic F127 hydrogels. Eur. J. Pharm. Sci. 2011, 43, 236–243. [Google Scholar] [CrossRef] [PubMed]
- Ye, F.; Larsen, S.W.; Yaghmur, A.; Jensen, H.; Larsen, C.; Østergaard, J. Real-time UV imaging of piroxicam diffusion and distribution from oil solutions into gels mimicking the subcutaneous matrix. Eur. J. Pharm. Sci. 2012, 46, 72–78. [Google Scholar] [CrossRef] [PubMed]
- Østergaard, J.; Lenke, J.; Jensen, S.S.; Sun, Y.; Ye, F. UV imaging for in vitro dissolution and release studies: Initial experiences. Diss. Technol. 2014, 21, 27–38. [Google Scholar] [CrossRef]
- Ye, F.; Larsen, S.W.; Yaghmur, A.; Jensen, H.; Larsen, C.; Ostergaard, J. Drug release into hydrogel-based subcutaneous surrogates studied by UV imaging. J. Pharm. Biomed. Anal. 2012, 71, 27–34. [Google Scholar] [CrossRef] [PubMed]
- Daunert, S.; Barrett, G.; Feliciano, J.S.; Shetty, R.S.; Shrestha, S.; Smith-Spencer, W. Genetically engineered whole-cell sensing systems: Coupling biological recognition with reporter genes. Chem. Rev. 2000, 100, 2705–2738. [Google Scholar] [CrossRef] [PubMed]
- Van der Meer, J.R.; Belkin, S. Where microbiology meets microengineering: Design and applications of reporter bacteria. Nat. Rev. Microbiol. 2010, 8, 511–522. [Google Scholar] [CrossRef]
- Blattner, F.R.; Plunkett, G.; Bloch, C.A.; Perna, N.T.; Burland, V.; Riley, M.; Collado-Vides, J.; Glasner, J.D.; Rode, C.K.; Mayhew, G.F.; et al. The complete genome sequence of Escherichia coli K-12. Science 1997, 277, 1453–1462. [Google Scholar] [CrossRef]
- Sambrook, J.; Russell, D. Molecular Cloning: A Laboratory Manual, 3rd ed.; Cold Spring Harbor Laboratory Press: New York, NY, USA, 2012. [Google Scholar]
- Neidhardt, F.C.; Bloch, P.L.; Smith, D.F. Culture medium for enterobacteria. J. Bacteriol. 1974, 119, 736–747. [Google Scholar]
- Quan, J.; Tian, J. Circular polymerase extension cloning for high-throughput cloning of complex and combinatorial DNA libraries. Nat. Protoc. 2011, 6, 242–251. [Google Scholar] [CrossRef]
- Murina, V.; Kasari, M.; Takada, H.; Hinnu, M.; Saha, C.K.; Grimshaw, J.W.; Seki, T.; Reith, M.; Putrinš, M.; Tenson, T.; et al. ABCF ATPases involved in protein synthesis, ribosome assembly and antibiotic resistance: Structural and functional diversification across the tree of life. J. Mol. Biol. 2019, 431, 3568–3590. [Google Scholar] [CrossRef]
- Cormack, B.P.; Valdivia, R.H.; Falkow, S. FACS-optimized mutants of the green fluorescent protein (GFP). Gene 1996, 173, 33–38. [Google Scholar] [CrossRef]
- Bindels, D.S.; Haarbosch, L.; van Weeren, L.; Postma, M.; Wiese, K.E.; Mastop, M.; Aumonier, S.; Gotthard, G.; Royant, A.; Hink, M.A.; et al. mScarlet: A bright monomeric red fluorescent protein for cellular imaging. Nat. Methods 2016, 14, 53–56. [Google Scholar] [CrossRef] [PubMed]
- Osterman, I.A.; Prokhorova, I.V.; Sysoev, V.O.; Boykova, Y.V.; Efremenkova, O.V.; Svetlov, M.S.; Kolb, V.A.; Bogdanov, A.A.; Sergiev, P.V.; Dontsova, O.A. Attenuation-based dual-fluorescent-protein reporter for screening translation inhibitors. Antimicrob. Agents Chemother. 2012, 56, 1774–1783. [Google Scholar] [CrossRef] [PubMed]
- Schneider, C.A.; Rasband, W.S.; Eliceiri, K.W. NIH image to ImageJ: 25 years of image analysis. Nat. Methods 2012, 9, 671–675. [Google Scholar] [CrossRef] [PubMed]
- Mekkawy, A.I.; El-Mokhtar, M.A.; Nafady, N.A.; Yousef, N.; Hamad, M.A.; El-Shanawany, S.M.; Ibrahim, E.H.; Elsabahy, M. In vitro and in vivo evaluation of biologically synthesized silver nanoparticles for topical applications: Effect of surface coating and loading into hydrogels. Int. J. Nanomed. 2017, 12, 759–777. [Google Scholar] [CrossRef] [PubMed]
- Weiser, J.R.; Saltzman, W.M. Controlled release for local delivery of drugs: Barriers and models. J. Control. Release 2014, 190, 664–673. [Google Scholar] [CrossRef] [PubMed]
- Sadri, M.; Mohammadi, A.; Hosseini, H. Drug release rate and kinetic investigation of composite polymeric nanofibers. Nanomed. Res. J. 2016, 1, 112–121. [Google Scholar]
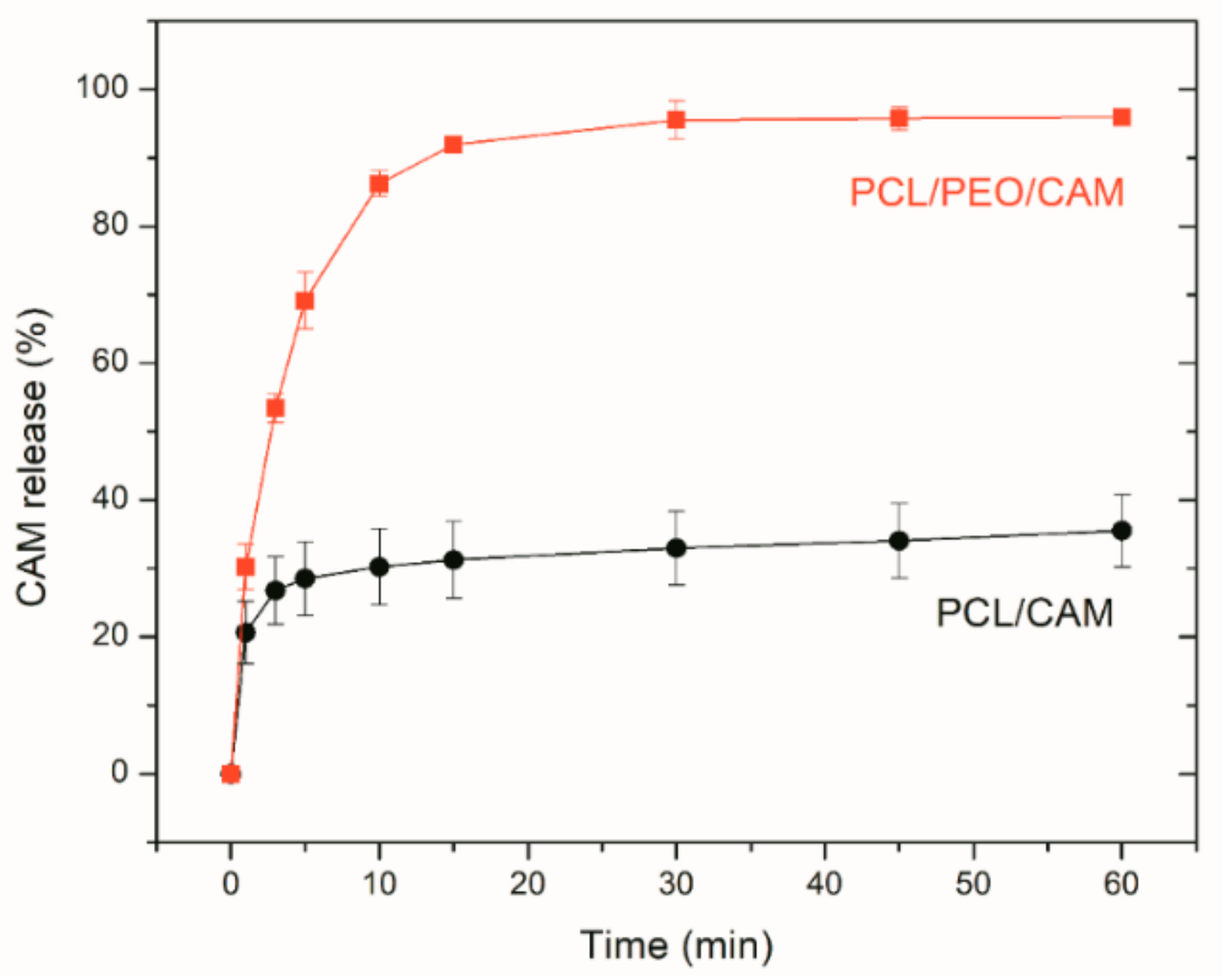


| Formulations | Materials/Polymer | Materials/Solvent | Distance (cm) | Flow Rate (ml/h) | Applied Voltage (kV) |
|---|---|---|---|---|---|
| PCL | PCL 12.5% (w/V), CONTROL | Chloroform:methanol (3:1 V/V) | 14 | 1.0 | 9 |
| PCL/CAM | PCL 12.5% (w/V) + CAM (4% w/w, solid state) | Chloroform:methanol (3:1 V/V) | 14 | 1.0 | 9 |
| PCL/PEO | PCL 10% + PEO 2% (w/V), CONTROL | Chloroform:methanol (3:1 V/V) | 17 | 2.5 | 12 |
| PCL/PEO/CAM | PCL 10% (w/V) + PEO 2% (w/V) + CAM (4% w/w, solid state) | Chloroform:methanol (3:1 V/V) | 17 | 2.5 | 12 |
| Electrospun Fiber Formulations | Theoretical CAM Content/% | Measured CAM Content/% ± SD |
|---|---|---|
| PCL/CAM | 4 | 4.0 ± 0.2 |
| PCL/PEO/CAM | 4 | 3.8 ± 0.3 |
© 2019 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Preem, L.; Bock, F.; Hinnu, M.; Putrinš, M.; Sagor, K.; Tenson, T.; Meos, A.; Østergaard, J.; Kogermann, K. Monitoring of Antimicrobial Drug Chloramphenicol Release from Electrospun Nano- and Microfiber Mats Using UV Imaging and Bacterial Bioreporters. Pharmaceutics 2019, 11, 487. https://doi.org/10.3390/pharmaceutics11090487
Preem L, Bock F, Hinnu M, Putrinš M, Sagor K, Tenson T, Meos A, Østergaard J, Kogermann K. Monitoring of Antimicrobial Drug Chloramphenicol Release from Electrospun Nano- and Microfiber Mats Using UV Imaging and Bacterial Bioreporters. Pharmaceutics. 2019; 11(9):487. https://doi.org/10.3390/pharmaceutics11090487
Chicago/Turabian StylePreem, Liis, Frederik Bock, Mariliis Hinnu, Marta Putrinš, Kadi Sagor, Tanel Tenson, Andres Meos, Jesper Østergaard, and Karin Kogermann. 2019. "Monitoring of Antimicrobial Drug Chloramphenicol Release from Electrospun Nano- and Microfiber Mats Using UV Imaging and Bacterial Bioreporters" Pharmaceutics 11, no. 9: 487. https://doi.org/10.3390/pharmaceutics11090487
APA StylePreem, L., Bock, F., Hinnu, M., Putrinš, M., Sagor, K., Tenson, T., Meos, A., Østergaard, J., & Kogermann, K. (2019). Monitoring of Antimicrobial Drug Chloramphenicol Release from Electrospun Nano- and Microfiber Mats Using UV Imaging and Bacterial Bioreporters. Pharmaceutics, 11(9), 487. https://doi.org/10.3390/pharmaceutics11090487