Antigenic and Biological Characterization of ORF2–6 Variants at Early Times Following PRRSV Infection
Abstract
:1. Introduction
2. Materials and Methods
2.1. Cells, Virus, and Pigs
2.2. Cloning and Sequencing PRRSV Variants
2.3. Identification of Viral Haplotypes
2.4. Construction of Chimeric Virus
2.5. Virus Neutralization Assays
2.6. PRRSV Binding and Entry Assays
2.7. Virus Replication Assays
2.8. Statistical Analyses
2.9. Nucleotide Sequence Accession Numbers
3. Results
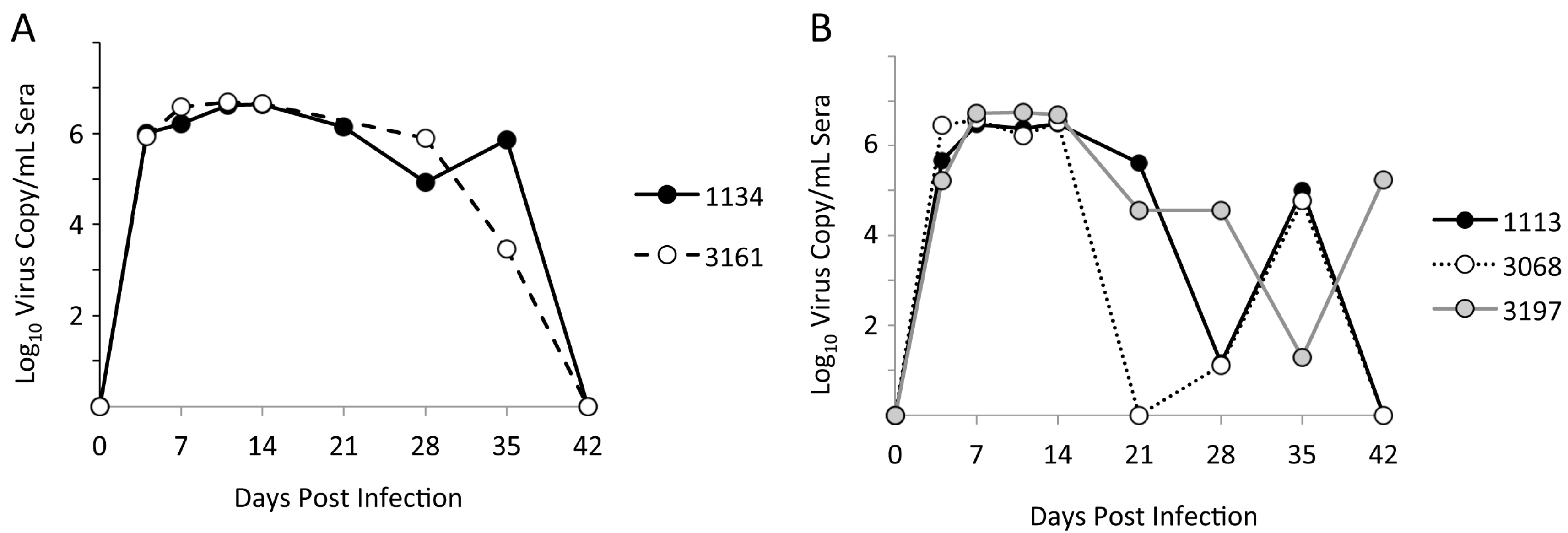
3.1. Limited Genetic Variation at Early Times after Experimental PRRSV Infection
3.2. Location and Patterns of Genetic Variation in ORF2–6 at Early Times after Infection
3.3. Identification of Pig-Specific ORF2–6 Haplotypes within Late Day Virus Populations
3.4. Variation in ORF2–6 Increases Resistance to Neutralizing Antibody in Some, but Not All, Pigs
3.5. Variation in Either Major or Minor Envelope Glycoproteins Can Mediate Escape from Autologous Neutralizing Antibody
3.6. The Effects of Neutralizing Antibody at Early Steps in the PRRSV Replication Cycle
3.7. The Effect of ORF2–6 Variation on PRRSV Replication Phenotype
4. Discussion
Supplementary Materials
Acknowledgments
Author Contributions
Conflicts of Interest
References
- Wensvoort, G.; Terpstra, C.; Pol, J.M.A.; ter Laak, E.A.; Bloemraad, M.; de Kluyver, E.P.; Kragten, C.; van Buiten, L.; den Besten, A.; Wagenaar, F.; et al. Mystery swine disease in The Netherlands: The isolation of Lelystad virus. Vet. Q. 1991, 13, 121–130. [Google Scholar] [CrossRef] [PubMed]
- Loula, T. Mystery Pig-Disease. Agri-Practice 1991, 12, 23–34. [Google Scholar]
- Collins, J.E.; Benfield, D.A.; Christianson, W.T.; Harris, L.; Hennings, J.C.; Shaw, D.P.; Goyal, S.M.; McCullough, S.; Morrison, R.B.; Joo, H.S.; et al. Isolation of swine infertility and respiratory syndrome virus (Isolate ATCC VR-2332) in North America and experimental reproduction of the disease in gnotobiotic pigs. J. Vet. Diagn. Investig. 1992, 4, 117–126. [Google Scholar] [CrossRef] [PubMed]
- Holtkamp, D.J.; Kliebenstein, J.B.; Zimmerman, J.J.; Neumann, E.; Rotto, H.; Yoder, T.K.; Wang, C.; Yeske, P.; Mowrer, C.L.; Haley, C. Economic Impact of Porcine Reproductive and Respiratory Syndrome Virus on U.S. Pork Producers; Animal Industry Report 2012; AS 658; Iowa State University: Ames, IA, USA, 2012. [Google Scholar]
- Kimman, T.G.; Cornelissen, L.A.; Moormann, R.J.; Rebel, J.M.; Stockhofe-Zurwieden, N. Challenges for porcine reproductive and respiratory syndrome virus (PRRSV) vaccinology. Vaccine 2009, 27, 3704–3718. [Google Scholar] [CrossRef] [PubMed]
- Ansari, I.H.; Kwon, B.; Osorio, F.A.; Pattnaik, A.K. Influence of N-linked glycosylation of porcine reproductive and respiratory syndrome virus GP5 on virus infectivity, antigenicity, and ability to induce neutralizing antibodies. J. Virol. 2006, 80, 3994–4004. [Google Scholar] [CrossRef] [PubMed]
- Faaberg, K.S.; Hocker, J.D.; Erdman, M.M.; Harris, D.L.; Nelson, E.A.; Torremorell, M.; Plagemann, P.G. Neutralizing antibody responses of pigs infected with natural GP5 N-glycan mutants of porcine reproductive and respiratory syndrome virus. Viral Immunol. 2006, 19, 294–304. [Google Scholar] [CrossRef] [PubMed]
- Kim, W.I.; Yoon, K.J. Molecular assessment of the role of envelope-associated structural proteins in cross neutralization among different PRRS viruses. Virus Genes 2008, 37, 380–391. [Google Scholar] [CrossRef] [PubMed]
- Costers, S.; Lefebvre, D.J.; Van Doorsselaere, J.; Vanhee, M.; Delputte, P.L.; Nauwynck, H.J. GP4 of porcine reproductive and respiratory syndrome virus contains a neutralizing epitope that is susceptible to immunoselection in vitro. Arch. Virol. 2010, 155, 371–378. [Google Scholar] [CrossRef] [PubMed]
- Meulenberg, J.J.; Petersen-den Besten, A.; De Kluyver, E.P.; Moormann, R.J.; Schaaper, W.M.; Wensvoort, G. Characterization of proteins encoded by ORFs 2 to 7 of Lelystad virus. Virology 1995, 206, 155–163. [Google Scholar] [CrossRef]
- Mardassi, H.; Massie, B.; Dea, S. Intracellular synthesis, processing, and transport of proteins encoded by ORFs 5 to 7 of porcine reproductive and respiratory syndrome virus. Virology 1996, 221, 98–112. [Google Scholar] [CrossRef] [PubMed]
- Delputte, P.L.; Vanderheijden, N.; Nauwynck, H.J.; Pensaert, M.B. Involvement of the matrix protein in attachment of porcine reproductive and respiratory syndrome virus to a heparinlike receptor on porcine alveolar macrophages. J. Virol. 2002, 76, 4312–4320. [Google Scholar] [CrossRef] [PubMed]
- Snijder, E.J.; Dobbe, J.C.; Spaan, W.J.M. Heterodimerization of the two major envelope proteins is essential for arterivirus infectivity. J. Virol. 2003, 77, 97–104. [Google Scholar] [CrossRef] [PubMed]
- Delputte, P.L.; Costers, S.; Nauwynck, H.J. Analysis of porcine reproductive and respiratory syndrome virus attachment and internalization: Distinctive roles for heparan sulphate and sialoadhesin. J. Gen. Virol. 2005, 86, 1441–1445. [Google Scholar] [CrossRef] [PubMed]
- Van Breedam, W.; Delputte, P.L.; Van Gorp, H.; Misinzo, G.; Vanderheijden, N.; Duan, X.; Nauwynck, H.J. Porcine reproductive and respiratory syndrome virus entry into the porcine macrophage. J. Gen. Virol. 2010, 91, 1659–1667. [Google Scholar] [CrossRef] [PubMed]
- Ostrowski, M.; Galeota, J.A.; Jar, A.M.; Platt, K.B.; Osorio, F.A.; Lopez, O.J. Identification of neutralizing and nonneutralizing epitopes in the porcine reproductive and respiratory syndrome virus GP5 ectodomain. J. Virol. 2002, 76, 4241–4250. [Google Scholar] [CrossRef] [PubMed]
- Wissink, E.H.J. The major envelope protein, GP5, of a European porcine reproductive and respiratory syndrome virus contains a neutralization epitope in its N-terminal ectodomain. J. Gen. Virol. 2003, 84, 1535–1543. [Google Scholar] [CrossRef] [PubMed]
- Plagemann, P.G.; Rowland, R.R.; Faaberg, K.S. The primary neutralization epitope of porcine respiratory and reproductive syndrome virus strain VR-2332 is located in the middle of the GP5 ectodomain. Arch. Virol. 2002, 147, 2327–2347. [Google Scholar] [CrossRef] [PubMed]
- Plagemann, P.G. Neutralizing antibody formation in swine infected with seven strains of porcine reproductive and respiratory syndrome virus as measured by indirect ELISA with peptides containing the GP5 neutralization epitope. Viral Immunol. 2006, 19, 285–293. [Google Scholar] [CrossRef] [PubMed]
- Li, J.; Murtaugh, M.P. Dissociation of porcine reproductive and respiratory syndrome virus neutralization from antibodies specific to major envelope protein surface epitopes. Virology 2012, 433, 367–376. [Google Scholar] [CrossRef] [PubMed]
- Delisle, B.; Gagnon, C.A.; Lambert, M.E.; D’Allaire, S. Porcine reproductive and respiratory syndrome virus diversity of Eastern Canada swine herds in a large sequence dataset reveals two hypervariable regions under positive selection. Infect. Genet. Evol. 2012, 12, 1111–1119. [Google Scholar] [CrossRef] [PubMed]
- Vanhee, M.; Van Breedam, W.; Costers, S.; Geldhof, M.; Noppe, Y.; Nauwynck, H. Characterization of antigenic regions in the porcine reproductive and respiratory syndrome virus by the use of peptide-specific serum antibodies. Vaccine 2011, 29, 4794–4804. [Google Scholar] [CrossRef] [PubMed]
- Loving, C.L.; Osorio, F.A.; Murtaugh, M.P.; Zuckermann, F.A. Innate and adaptive immunity against porcine reproductive and respiratory syndrome virus. Vet. Immunol. Immunopathol. 2015, 167, 1–14. [Google Scholar] [CrossRef] [PubMed]
- Trible, B.R.; Popescu, L.N.; Monday, N.; Calvert, J.G.; Rowland, R.R. A single amino acid deletion in the matrix protein of porcine reproductive and respiratory syndrome virus confers resistance to a polyclonal swine antibody with broadly neutralizing activity. J. Virol. 2015, 89, 6515–6520. [Google Scholar] [CrossRef] [PubMed]
- Van Nieuwstadt, A.P.; Meulenberg, J.J.; van Essen-Zanbergen, A.; Petersen-den Besten, A.; Bende, R.J.; Moormann, R.J.; Wensvoort, G. Proteins encoded by open reading frames 3 and 4 of the genome of Lelystad virus (Arteriviridae) are structural proteins of the virion. J. Virol. 1996, 70, 4767–4772. [Google Scholar] [PubMed]
- Wissink, E.H.; Kroese, M.V.; van Wijk, H.A.; Rijsewijk, F.A.; Meulenberg, J.J.; Rottier, P.J. Envelope protein requirements for the assembly of infectious virions of porcine reproductive and respiratory syndrome virus. J. Virol. 2005, 79, 12495–12506. [Google Scholar] [CrossRef] [PubMed]
- De Lima, M.; Ansari, I.H.; Das, P.B.; Ku, B.J.; Martinez-Lobo, F.J.; Pattnaik, A.K.; Osorio, F.A. GP3 is a structural component of the PRRSV type II (US) virion. Virology 2009, 390, 31–36. [Google Scholar] [CrossRef] [PubMed]
- Das, P.B.; Dinh, P.X.; Ansari, I.H.; de Lima, M.; Osorio, F.A.; Pattnaik, A.K. The minor envelope glycoproteins GP2a and GP4 of porcine reproductive and respiratory syndrome virus interact with the receptor CD163. J. Virol. 2010, 84, 1731–1740. [Google Scholar] [CrossRef] [PubMed]
- Whitworth, K.M.; Rowland, R.R.; Ewen, C.L.; Trible, B.R.; Kerrigan, M.A.; Cino-Ozuna, A.G.; Samuel, M.S.; Lightner, J.E.; McLaren, D.G.; Mileham, A.J.; et al. Gene-edited pigs are protected from porcine reproductive and respiratory syndrome virus. Nat. Biotechnol. 2016, 34, 20–22. [Google Scholar] [CrossRef] [PubMed]
- Vu, H.L.; Kwon, B.; Yoon, K.J.; Laegreid, W.W.; Pattnaik, A.K.; Osorio, F.A. Immune evasion of porcine reproductive and respiratory syndrome virus through glycan shielding involves both glycoprotein 5 as well as glycoprotein 3. J. Virol. 2011, 85, 5555–5564. [Google Scholar] [CrossRef] [PubMed]
- Cancel-Tirado, S.M.; Evans, R.B.; Yoon, K.J. Monoclonal antibody analysis of porcine reproductive and respiratory syndrome virus epitopes associated with antibody-dependent enhancement and neutralization of virus infection. Vet. Immunol. Immunopathol. 2004, 102, 249–262. [Google Scholar] [CrossRef] [PubMed]
- Meulenberg, J.J.; van Nieuwstadt, A.P.; van Essen-Zandbergen, A.; Langeveld, J.P. Posttranslational processing and identification of a neutralization domain of the GP4 protein encoded by ORF4 of Lelystad virus. J. Virol. 1997, 71, 6061–6067. [Google Scholar] [PubMed]
- Wu, W.H.; Fang, Y.; Farwell, R.; Steffen-Bien, M.; Rowland, R.R.; Christopher-Hennings, J.; Nelson, E.A. A 10-kDa structural protein of porcine reproductive and respiratory syndrome virus encoded by ORF2b. Virology 2001, 287, 183–191. [Google Scholar] [CrossRef] [PubMed]
- Johnson, C.R.; Griggs, T.F.; Gnanandarajah, J.; Murtaugh, M.P. Novel structural protein in porcine reproductive and respiratory syndrome virus encoded by an alternative ORF5 present in all arteriviruses. J. Gen. Virol. 2011, 92, 1107–1116. [Google Scholar] [CrossRef] [PubMed]
- Robinson, S.R.; Figueiredo, M.C.; Abrahante, J.E.; Murtaugh, M.P. Immune response to ORF5a protein immunization is not protective against porcine reproductive and respiratory syndrome virus infection. Vet. Microbiol. 2013, 164, 281–285. [Google Scholar] [CrossRef] [PubMed]
- Lopez, O.J.; Osorio, F.A. Role of neutralizing antibodies in PRRSV protective immunity. Vet. Immunol. Immunopathol. 2004, 102, 155–163. [Google Scholar] [CrossRef] [PubMed]
- Osorio, F.A.; Galeota, J.A.; Nelson, E.; Brodersen, B.; Doster, A.; Wills, R.; Zuckermann, F.; Laegreid, W.W. Passive transfer of virus-specific antibodies confers protection against reproductive failure induced by a virulent strain of porcine reproductive and respiratory syndrome virus and establishes sterilizing immunity. Virology 2002, 302, 9–20. [Google Scholar] [CrossRef] [PubMed]
- Murtaugh, M.P.; Xiao, Z.; Zuckermann, F. Immunological responses of swine to porcine reproductive and respiratory syndrome virus infection. Viral Immunol. 2002, 15, 533–547. [Google Scholar] [CrossRef] [PubMed]
- Welch, S.K.W.; Calvert, J.G. A brief review of CD163 and its role in PRRSV infection. Virus Res. 2010, 154, 98–103. [Google Scholar] [CrossRef] [PubMed]
- Duan, X.; Nauwynck, H.J.; Favoreel, H.; Pensaert, M.B. Porcine reproductive and respiratory syndrome virus infection of alveolar macrophages can be blocked by monoclonal antibodies against cell surface antigens. Adv. Exp. Med. Biol. 1998, 440, 81–88. [Google Scholar] [PubMed]
- Vanderheijden, N.; Delputte, P.L.; Favoreel, H.W.; Vandekerckhove, J.; Van Damme, J.; van Woensel, P.A.; Nauwynck, H.J. Involvement of sialoadhesin in entry of porcine reproductive and respiratory syndrome virus into porcine alveolar macrophages. J. Virol. 2003, 77, 8207–8215. [Google Scholar] [CrossRef] [PubMed]
- Prather, R.S.; Rowland, R.R.; Ewen, C.; Trible, B.; Kerrigan, M.; Bawa, B.; Teson, J.M.; Mao, J.; Lee, K.; Samuel, M.S. An intact sialoadhesin (Sn/SIGLEC1/CD169) is not required for attachment/internalization of the porcine reproductive and respiratory syndrome virus. J. Virol. 2013, 87, 9538–9546. [Google Scholar] [CrossRef] [PubMed]
- Calvert, J.G.; Slade, D.E.; Shields, S.L.; Jolie, R.; Mannan, R.M.; Ankenbauer, R.G.; Welch, S.K. CD163 expression confers susceptibility to porcine reproductive and respiratory syndrome viruses. J. Virol. 2007, 81, 7371–7379. [Google Scholar] [CrossRef] [PubMed]
- Schaer, D.J.; Schaer, C.A.; Buehler, P.W.; Boykins, R.A.; Schoedon, G.; Alayash, A.I.; Schaffner, A. CD163 is the macrophage scavenger receptor for native and chemically modified hemoglobins in the absence of haptoglobin. Blood 2006, 107, 373–380. [Google Scholar] [CrossRef] [PubMed]
- Van Gorp, H.; Van Breedam, W.; Delputte, P.L.; Nauwynck, H.J. Sialoadhesin and CD163 join forces during entry of the porcine reproductive and respiratory syndrome virus. J. Gen. Virol. 2008, 89, 2943–2953. [Google Scholar] [CrossRef] [PubMed]
- Dokland, T. The structural biology of PRRSV. Virus Res. 2010, 154, 86–97. [Google Scholar] [CrossRef] [PubMed]
- Gao, J.M.; Xiao, S.Q.; Xiao, Y.H.; Wang, X.P.; Zhang, C.; Zhao, Q.; Nan, Y.C.; Huang, B.C.; Liu, H.L.; Liu, N.N.; et al. MYH9 is an essential factor for porcine reproductive and respiratory syndrome virus infection. Sci. Rep. 2016, 6, 25120. [Google Scholar] [CrossRef] [PubMed]
- Pierson, T.C.; Fremont, D.H.; Kuhn, R.J.; Diamond, M.S. Structural insights into the mechanisms of antibody-mediated neutralization of flavivirus infection: Implications for vaccine development. Cell Host Microbe 2008, 4, 229–238. [Google Scholar] [CrossRef] [PubMed]
- Chen, Y.; He, S.; Sun, L.; Luo, Y.; Sun, Y.; Xie, J.; Zhou, P.; Su, S.; Zhang, G. Genetic variation, pathogenicity, and immunogenicity of highly pathogenic porcine reproductive and respiratory syndrome virus strain XH-GD at different passage levels. Arch. Virol. 2016, 161, 77–86. [Google Scholar] [CrossRef] [PubMed]
- Yu, X.; Chen, N.; Deng, X.; Cao, Z.; Han, W.; Hu, D.; Wu, J.; Zhang, S.; Wang, B.; Gu, X; et al. Genomic sequencing reveals mutations potentially related to the overattenuation of a highly pathogenic porcine reproductive and respiratory syndrome virus. Clin. Vaccine Immunol. 2013, 20, 613–619. [Google Scholar] [CrossRef] [PubMed]
- Truong, H.M.; Lu, Z.; Kutish, G.F.; Galeota, J.; Osorio, F.A.; Pattnaik, A.K. A highly pathogenic porcine reproductive and respiratory syndrome virus generated from an infectious cDNA clone retains the in vivo virulence and transmissibility properties of the parental virus. Virology 2004, 325, 308–319. [Google Scholar] [CrossRef] [PubMed]
- Lunney, J.K.; Steibel, J.P.; Reecy, J.M.; Fritz, E.; Rothschild, M.F.; Kerrigan, M.; Trible, B.; Rowland, R.R. Probing genetic control of swine responses to PRRSV infection: Current progress of the PRRS host genetics consortium. BMC Proc. 2011, 5 (Suppl. 4), S30. [Google Scholar] [CrossRef] [PubMed]
- Boddicker, N.; Waide, E.H.; Rowland, R.R.; Lunney, J.K.; Garrick, D.J.; Reecy, J.M.; Dekkers, J.C. Evidence for a major QTL associated with host response to porcine reproductive and respiratory syndrome virus challenge. J. Anim. Sci. 2012, 90, 1733–1746. [Google Scholar] [CrossRef] [PubMed]
- Wu, W.; Blythe, D.C.; Loyd, H.; Mealey, R.H.; Tallmadge, R.L.; Dorman, K.S.; Carpenter, S. Decreased infectivity of a neutralization-resistant equine infectious anemia virus variant can be overcome by efficient cell-to-cell spread. J. Virol. 2011, 85, 10421–10424. [Google Scholar] [CrossRef] [PubMed]
- Van Gorp, H.; Van Breedam, W.; Delputte, P.L.; Nauwynck, H.J. The porcine reproductive and respiratory syndrome virus requires trafficking through CD163-positive early endosomes, but not late endosomes, for productive infection. Arch. Virol. 2009, 154, 1939–1943. [Google Scholar] [CrossRef] [PubMed]
- Costers, S.; Vanhee, M.; Van Breedam, W.; Van Doorsselaere, J.; Geldhof, M.; Nauwynck, H.J. GP4-specific neutralizing antibodies might be a driving force in PRRSV evolution. Virus Res. 2010, 154, 104–113. [Google Scholar] [CrossRef] [PubMed]
- Fan, B.; Liu, X.; Bai, J.; Zhang, T.; Zhang, Q.; Jiang, P. The amino acid residues at 102 and 104 in GP5 of porcine reproductive and respiratory syndrome virus regulate viral neutralization susceptibility to the porcine serum neutralizing antibody. Virus Res. 2015, 204, 21–30. [Google Scholar] [CrossRef] [PubMed]
- Murtaugh, M.P.; Stadejek, T.; Abrahante, J.E.; Lam, T.T.; Leung, F.C. The ever-expanding diversity of porcine reproductive and respiratory syndrome virus. Virus Res. 2010, 154, 18–30. [Google Scholar] [CrossRef] [PubMed]
- Thaa, B.; Sinhadri, B.C.; Tielesch, C.; Krause, E.; Veit, M. Signal peptide cleavage from GP5 of PRRSV: A minor fraction of molecules retains the decoy epitope, a presumed molecular cause for viral persistence. PLoS ONE 2013, 8, e65548. [Google Scholar] [CrossRef] [PubMed]
- Chen, N.; Trible, B.R.; Kerrigan, M.A.; Tian, K.; Rowland, R.R. ORF5 of porcine reproductive and respiratory syndrome virus (PRRSV) is a target of diversifying selection as infection progresses from acute infection to virus rebound. Infect. Genet. Evol. 2016, 40, 167–175. [Google Scholar] [CrossRef] [PubMed]
- Robinson, S.R.; Li, J.; Nelson, E.A.; Murtaugh, M.P. Broadly neutralizing antibodies against the rapidly evolving porcine reproductive and respiratory syndrome virus. Virus Res. 2015, 203, 56–65. [Google Scholar] [CrossRef] [PubMed]
- Tian, D.; Wei, Z.; Zevenhoven-Dobbe, J.C.; Liu, R.; Tong, G.; Snijder, E.J.; Yuan, S. Arterivirus minor envelope proteins are a major determinant of viral tropism in cell culture. J. Virol. 2012, 86, 3701–3712. [Google Scholar] [CrossRef] [PubMed]



| Sample ID | Viremia Classification | Day Post Inoculation | Number Clones Sequenced a | Pairwise Identity b |
|---|---|---|---|---|
| Inoculum | N/A | 0 | 25 | 99.71 |
| 1134 | Prolonged | 7 | 8 | 99.76 |
| 28 | 21 | 99.63 | ||
| 3161 | Prolonged | 7 | 9 | 99.71 |
| 28 | 28 | 99.72 | ||
| 1113 | Rebound | 7 | 9 | 99.71 |
| 35 | 26 | 99.62 | ||
| 3068 | Rebound | 7 | 9 | 99.40 |
| 35 | 31 | 99.78 | ||
| 3197 | Rebound | 7 | 10 | 99.49 |
| 41 | 26 | 99.76 | ||
| Combined | 202 | 99.57 c |
© 2017 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Evans, A.B.; Loyd, H.; Dunkelberger, J.R.; Van Tol, S.; Bolton, M.J.; Dorman, K.S.; Dekkers, J.C.M.; Carpenter, S. Antigenic and Biological Characterization of ORF2–6 Variants at Early Times Following PRRSV Infection. Viruses 2017, 9, 113. https://doi.org/10.3390/v9050113
Evans AB, Loyd H, Dunkelberger JR, Van Tol S, Bolton MJ, Dorman KS, Dekkers JCM, Carpenter S. Antigenic and Biological Characterization of ORF2–6 Variants at Early Times Following PRRSV Infection. Viruses. 2017; 9(5):113. https://doi.org/10.3390/v9050113
Chicago/Turabian StyleEvans, Alyssa B., Hyelee Loyd, Jenelle R. Dunkelberger, Sarah Van Tol, Marcus J. Bolton, Karin S. Dorman, Jack C. M. Dekkers, and Susan Carpenter. 2017. "Antigenic and Biological Characterization of ORF2–6 Variants at Early Times Following PRRSV Infection" Viruses 9, no. 5: 113. https://doi.org/10.3390/v9050113
APA StyleEvans, A. B., Loyd, H., Dunkelberger, J. R., Van Tol, S., Bolton, M. J., Dorman, K. S., Dekkers, J. C. M., & Carpenter, S. (2017). Antigenic and Biological Characterization of ORF2–6 Variants at Early Times Following PRRSV Infection. Viruses, 9(5), 113. https://doi.org/10.3390/v9050113






