Isolation and Characterization of a Double Stranded DNA Megavirus Infecting the Toxin-Producing Haptophyte Prymnesium parvum
Abstract
:1. Introduction
2. Materials and Methods
2.1. Prymnesium parvum Culture Conditions
2.2. Isolation of Lytic Virus Particles
2.3. Transmission Electron Microscopy
2.4. Host Specificity
2.5. Infection Cycle
2.6. Chloroform Sensitivity
2.7. Viral DNA Extraction, Sequencing, and Phylogenetic Analyses
3. Results
3.1. Isolation of Lytic Virus Particles
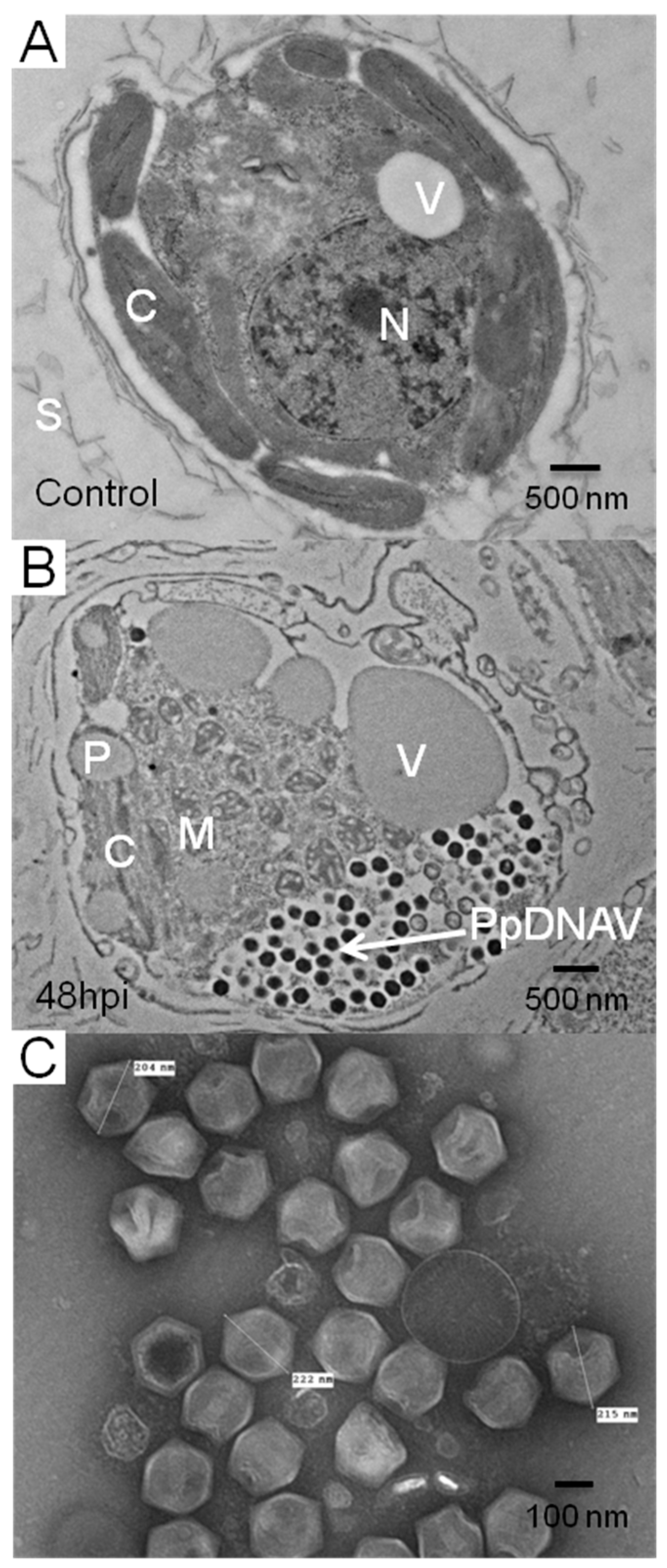
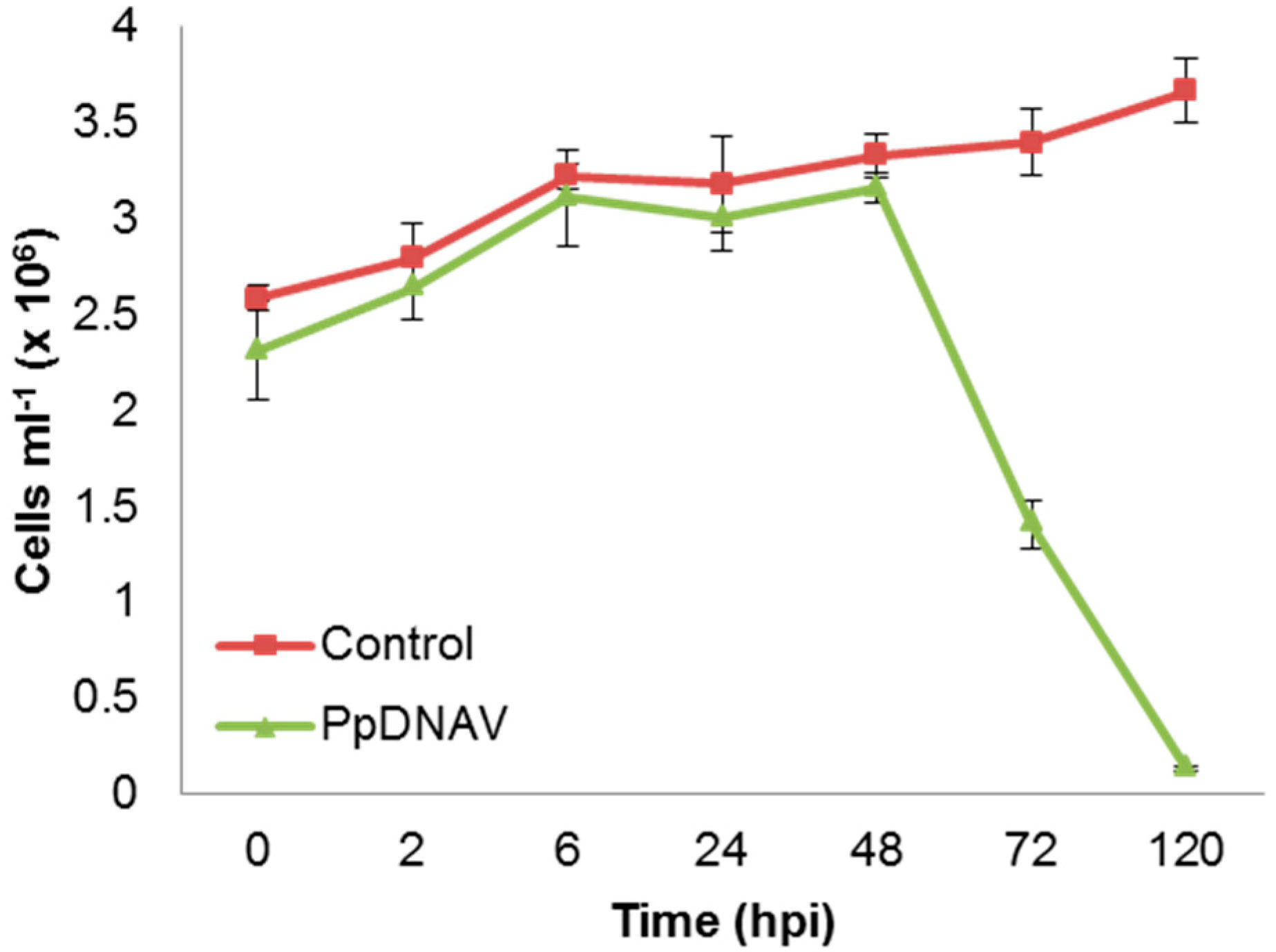
3.2. Virus Morphology, Host Range, and Infectious Properties
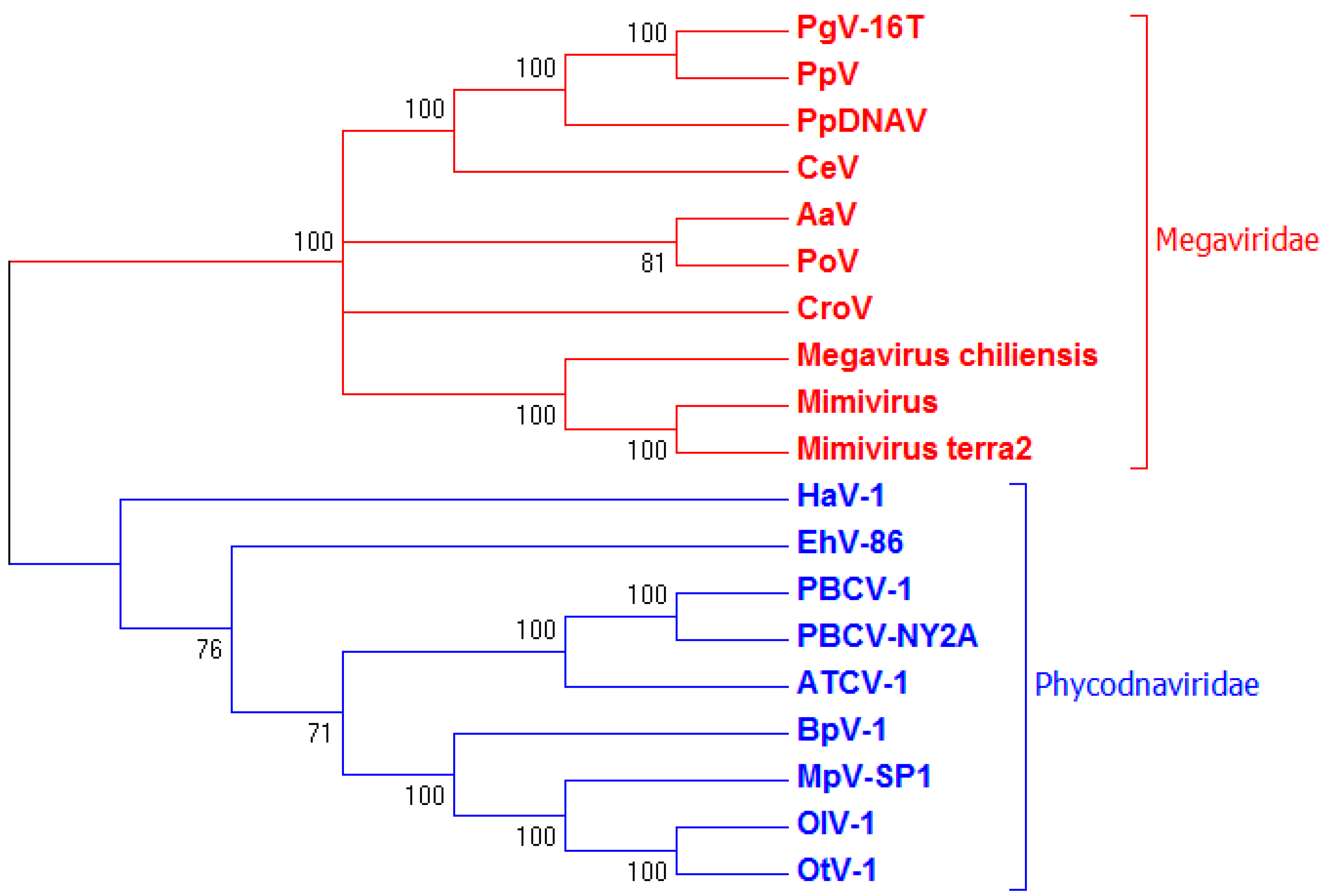
3.3. Genome Sequencing and Phylogenetic Analysis
4. Discussion
Supplementary Materials
Acknowledgments
Author Contributions
Conflicts of Interest
References
- Thingstad, T.F. Elements of a theory for the mechanisms controlling abundance, diversity, and biogeochemical role of lytic bacterial viruses in aquatic systems. Limnol. Oceanogr. 2000, 45, 1320–1328. [Google Scholar] [CrossRef]
- Fuhrman, J.A. Marine viruses and their biogeochemical and ecological effects. Nature 1999, 399, 541–548. [Google Scholar] [CrossRef] [PubMed]
- Santini, S.; Jeudy, S.; Bartoli, J.; Poirot, O.; Lescot, M.; Abergel, C.; Barbe, V.; Wommack, K.E.; Noordeloos, A.A.; Brussaard, C.P.; et al. Genome of Phaeocystis globosa virus PgV-16T highlights the common ancestry of the largest known DNA viruses infecting eukaryotes. Proc. Natl. Acad. Sci. USA 2013, 110, 10800–10805. [Google Scholar] [CrossRef] [PubMed]
- Wilson, W.H.; Tarran, G.A.; Schroeder, D.; Cox, M.; Oke, J.; Malin, G. Isolation of viruses responsible for the demise of an Emiliania huxleyi bloom in the English Channel. J. Mar. Biol. Assoc. UK 2002, 82, 369–377. [Google Scholar] [CrossRef]
- Schroeder, D.C.; Oke, J.; Malin, G.; Wilson, W.H. Coccolithovirus (Phycodnaviridae): Characterisation of a new large dsDNA algal virus that infects Emiliana huxleyi. Arch. Virol. 2002, 147, 1685–1698. [Google Scholar] [CrossRef] [PubMed]
- Brussaard, C.P.D.; Short, S.M.; Frederickson, C.M.; Suttle, C.A. Isolation and Phylogenetic Analysis of Novel Viruses Infecting the Phytoplankton Phaeocystis globosa (Prymnesiophyceae). Appl. Environ. Microbiol. 2004, 70, 3700–3705. [Google Scholar] [CrossRef] [PubMed]
- Moniruzzaman, M.; Gann, E.R.; LeCleir, G.R.; Kang, Y.; Gobler, C.J.; Wilhelm, S.W. Diversity and dynamics of algal Megaviridae members during a harmful brown tide caused by the pelagophyte, Aureococcus anophagefferens. FEMS Microbiol. Ecol. 2016, 92, fiw058. [Google Scholar] [CrossRef] [PubMed]
- Short, S.M. The ecology of viruses that infect eukaryotic algae. Environ. Microbiol. 2012, 14, 2253–2271. [Google Scholar] [CrossRef] [PubMed]
- Brussaard, C.P.D. Viral Control of Phytoplankton Populations—A Review. J. Eukaryot. Microbiol. 2004, 51, 125–138. [Google Scholar] [CrossRef] [PubMed]
- Suttle, C.A. Marine viruses—Major players in the global ecosystem. Nat. Rev. Microl. 2007, 5, 801–812. [Google Scholar] [CrossRef] [PubMed]
- Wilhelm, S.W.; Suttle, C.A. Viruses and Nutrient Cycles in the Sea: Viruses play critical roles in the structure and function of aquatic food webs. Bioscience 1999, 49, 781–788. [Google Scholar] [CrossRef]
- Nagasaki, K.; Yamaguchi, M. Isolation of a virus infectious to the harmful bloom causing microalga Heterosigma akashiwo (Raphidophyceae). Aquat. Microb. Ecol. 1997, 13, 135–140. [Google Scholar] [CrossRef]
- Keizo, N.; Mineo, Y. Intra-species host specificity of HaV (Heterosigma akashiwo virus) clones. Aquat. Microb. Ecol. 1998, 14, 109–112. [Google Scholar] [CrossRef]
- Keizo, N.; Mineo, Y. Effect of temperature on the algicidal activity and the stability of HaV (Heterosigma akashiwo virus). Aquat. Microb. Ecol. 1998, 15, 211–216. [Google Scholar] [CrossRef]
- Lawrence, J.E.; Chan, A.M.; Suttle, C.A. A novel virus (HaNIV) causes lysis of the toxic bloom-forming alga Heterosigma akashiwo (Raphidophyceae). J. Phycol. 2001, 37, 216–222. [Google Scholar] [CrossRef]
- Lawrence, J.E.; Chan, A.M.; Suttle, C.A. Viruses causing lysis of the toxic bloom-forming alga Heterosigma akashiwo (Raphidophyceae) are widespread in coastal sediments of British Columbia, Canada. Limnol. Oceanogr. 2002, 47, 545–550. [Google Scholar] [CrossRef]
- Tai, V.; Lawrence, J.E.; Lang, A.S.; Chan, A.M.; Culley, A.I.; Suttle, C.A. Characterization of HaRNAV, a single-stranded RNA virus causing lysis of Heterosigma akashiwo (Raphidophyceae). J. Phycol. 2003, 39, 343–352. [Google Scholar] [CrossRef]
- Janice, E.L.; Curtis, A.S. Effect of viral infection on sinking rates of Heterosigma akashiwo and its implications for bloom termination. Aquat. Microb. Ecol. 2004, 37, 1–7. [Google Scholar] [CrossRef]
- Lawrence, J.E.; Brussaard, C.P.D.; Suttle, C.A. Virus-Specific Responses of Heterosigma akashiwo to Infection. Appl. Environ. Microbiol. 2006, 72, 7829–7834. [Google Scholar] [CrossRef] [PubMed]
- Granéli, E.; Edvardsen, B.; Roelke, D.L.; Hagström, J.A. The ecophysiology and bloom dynamics of Prymnesium spp. Harmful Algae 2012, 14, 260–270. [Google Scholar] [CrossRef]
- Manning, S.R.; La Claire, J.W. Prymnesins: Toxic Metabolites of the Golden Alga, Prymnesium parvum Carter (Haptophyta). Mar. Drugs 2010, 8, 678–704. [Google Scholar] [CrossRef] [PubMed]
- Schoemann, V.; Becquevort, S.; Stefels, J.; Rousseau, V.; Lancelot, C. Phaeocystis blooms in the global ocean and their controlling mechanisms: A review. J. Sea Res. 2005, 53, 43–66. [Google Scholar] [CrossRef]
- Leblanc, K.; Hare, C.E.; Feng, Y.; Berg, G.M.; DiTullio, G.R.; Neeley, A.; Benner, I.; Sprengel, C.; Beck, A.; Sanudo-Wilhelmy, S.A.; et al. Distribution of calcifying and silicifying phytoplankton in relation to environmental and biogeochemical parameters during the late stages of the 2005 North East Atlantic Spring Bloom. Biogeosciences 2009, 6, 2155–2179. [Google Scholar] [CrossRef]
- Malin, G.; Wilson, W.H.; Bratbak, G.; Liss, P.S.; Mann, N.H. Elevated production of dimethylsulfide resulting from viral infection of cultures of Phaeocystis pouchetii. Limnol. Oceanogr. 1998, 43, 1389–1393. [Google Scholar] [CrossRef]
- Edvardsen, B.; Paasche, E. Bloom Dynamics and Physiology of Prymnesium and Chrysochromulina in Physiological Ecology of Harmful Algal Blooms; Springer-Verlag: Berlin/Heidelberg, Germany, 1998. [Google Scholar]
- Roelke, D.L.; Barkoh, A.; Brooks, B.W.; Grover, J.P.; Hambright, K.D.; LaClaire, J.W.; Moeller, P.D.R.; Patino, R. A chronicle of a killer alga in the west: Ecology, assessment, and management of Prymnesium parvum blooms. Hydrobiologia 2016, 764, 29–50. [Google Scholar] [CrossRef]
- Schwierzke, L.; Roelke, D.L.; Brooks, B.W.; Grover, J.P.; Valenti, T.W.; Lahousse, M.; Miller, C.J.; Pinckney, J.L. Prymnesium parvum Population Dynamics During Bloom Development: A Role Assessment of Grazers and Virus. J. Am. Water. Resour. Assoc. 2010, 46, 63–75. [Google Scholar] [CrossRef]
- Brenner, S.; Horne, R.W. A negative staining method for high resolution electron microscopy of viruses. Biochim. Biophys. Acta 1959, 34, 103–110. [Google Scholar] [CrossRef]
- Gordon, G.B.; Miller, L.R.; Bensch, K.G. Fixation of tissue culture cells for ultrastructural cytochemistry. Exp. Cell Res. 1963, 31, 440–443. [Google Scholar] [CrossRef]
- Martínez Martínez, J.; Boere, A.; Gilg, I.; van Lent, J.W.M.; Witte, H.J.; van Bleijswijk, J.D.L.; Brussaard, C.P.D. New lipid envelope-containing dsDNA virus isolates infecting Micromonas pusilla reveal a separate phylogenetic group. Aquat. Microb. Ecol. 2015, 74, 17–28. [Google Scholar] [CrossRef]
- Besemer, J.; Lomsadze, A.; Borodovsky, M. GeneMarkS: A self-training method for prediction of gene starts in microbial genomes. Implications for finding sequence motifs in regulatory regions. Nucleic Acids Res. 2001, 29, 2607–2618. [Google Scholar] [CrossRef] [PubMed]
- Altschul, S.F.; Gish, W.; Miller, W.; Myers, E.W.; Lipman, D.J. Basic local alignment search tool. J. Mol. Biol. 1990, 215, 403–410. [Google Scholar] [CrossRef]
- Katoh, K.; Toh, H. Recent developments in the MAFFT multiple sequence alignment program. Brief. Bioinform. 2008, 9, 286–298. [Google Scholar] [CrossRef] [PubMed]
- Saitou, N.; Nei, M. The neighbor-joining method: A new method for reconstructing phylogenetic trees. Mol. Biol. Evol. 1987, 4, 406–425. [Google Scholar] [CrossRef] [PubMed]
- Kumar, S.; Stecher, G.; Tamura, K. MEGA7: Molecular Evolutionary Genetics Analysis version 7.0 for bigger datasets. Mol. Biol. Evol. 2016, 33, 1870–1874. [Google Scholar] [CrossRef] [PubMed]
- Den Boon, J.A.; Diaz, A.; Ahlquist, P. Cytoplasmic Viral Replication Complexes. Cell Host Microbe 2010, 8, 77–85. [Google Scholar] [CrossRef] [PubMed]
- Holdway, P.A.; Watson, R.A.; Moss, B. Aspects of the ecology of Prymnesium parvum (Haptophyta) and water chemistry in the Norfolk Broads, England. Freshwat. Biol. 1978, 8, 295–311. [Google Scholar] [CrossRef]
- Zauberman, N.; Mutsafi, Y.; Halevy, D.B.; Shimoni, E.; Klein, E.; Xiao, C.; Sun, S.; Minsky, A. Distinct DNA Exit and Packaging Portals in the Virus Acanthamoeba polyphaga mimivirus. PLoS Biol. 2008, 6, e114. [Google Scholar] [CrossRef] [PubMed]
- Mutsafi, Y.; Fridmann-Sirkis, Y.; Milrot, E.; Hevroni, L.; Minsky, A. Infection cycles of large DNA viruses: Emerging themes and underlying questions. Virology 2014, 466–467, 3–14. [Google Scholar] [CrossRef] [PubMed]
- Feldman, H.A.; Wang, S.S. Sensitivity of various viruses to chloroform. Proc. Soc. Exp. Biol. Med. 1961, 106, 736–738. [Google Scholar] [CrossRef] [PubMed]
- Nedbalová, L.; Střížek, A.; Sigler, K.; Řezanka, T. Effect of salinity on the fatty acid and triacylglycerol composition of five haptophyte algae from the genera Coccolithophora, Isochrysis and Prymnesium determined by LC-MS/APCI. Phytochemistry 2016, 130, 64–76. [Google Scholar] [CrossRef] [PubMed]
- Johannessen, T.V.; Bratbak, G.; Larsen, A.; Ogata, H.; Egge, E.S.; Edvardsen, B.; Eikrem, W.; Sandaa, R.-A. Characterisation of three novel giant viruses reveals huge diversity among viruses infecting Prymnesiales (Haptophyta). Virology 2015, 476, 180–188. [Google Scholar] [CrossRef] [PubMed]
- Wilhelm, S.W.; Coy, S.R.; Gann, E.R.; Moniruzzaman, M.; Stough, J.M.A. Standing on the Shoulders of Giant Viruses: Five Lessons Learned about Large Viruses Infecting Small Eukaryotes and the Opportunities They Create. PLoS Pathog. 2016, 12, e1005752. [Google Scholar] [CrossRef] [PubMed]
- Gallot-Lavallée, L.; Pagarete, A.; Legendre, M.; Santini, S.; Sandaa, R.-A.; Himmelbauer, H.; Ogata, H.; Bratbak, G.; Claverie, J.-M. The 474-Kilobase-Pair Complete Genome Sequence of CeV-01B, a Virus Infecting Haptolina (Chrysochromulina) ericina (Prymnesiophyceae). Genome Announc. 2015, 3, e01413–e01415. [Google Scholar] [CrossRef]
- Sandaa, R.-A.; Heldal, M.; Castberg, T.; Thyrhaug, R.; Bratbak, G. Isolation and Characterization of Two Viruses with Large Genome Size Infecting Chrysochromulina ericina (Prymnesiophyceae) and Pyramimonas orientalis (Prasinophyceae). Virology 2001, 290, 272–280. [Google Scholar] [CrossRef] [PubMed]
- Moniruzzaman, M.; LeCleir, G.R.; Brown, C.M.; Gobler, C.J.; Bidle, K.D.; Wilson, W.H.; Wilhelm, S.W. Genome of brown tide virus (AaV), the little giant of the Megaviridae, elucidates NCLDV genome expansion and host-virus coevolution. Virology 2014, 466–467, 60–70. [Google Scholar] [CrossRef] [PubMed]
- Mirza, S.F.; Staniewski, M.A.; Short, C.M.; Long, A.M.; Chaban, Y.V.; Short, S.M. Isolation and characterization of a virus infecting the freshwater algae Chrysochromulina parva. Virology 2015, 486, 105–115. [Google Scholar] [CrossRef] [PubMed]
- Henrikson, J.C.; Gharfeh, M.S.; Easton, A.C.; Easton, J.D.; Glenn, K.L.; Shadfan, M.; Mooberry, S.L.; Hambright, K.D.; Cichewicz, R.H. Reassessing the ichthyotoxin profile of cultured Prymnesium parvum (golden algae) and comparing it to samples collected from recent freshwater bloom and fish kill events in North America. Toxicon 2010, 55, 1396–1404. [Google Scholar] [CrossRef] [PubMed]
- Kozakai, H.; Oshima, Y.; Yasumoto, T. Isolation and Structural Elucidation of Hemolysin from the Phytoflagellate Prymnesium parvum. Agric. Biol. Chem. 1982, 46, 233–236. [Google Scholar] [CrossRef]
- Igarashi, T.; Satake, M.; Yasumoto, T. Prymnesin-2: A Potent Ichthyotoxic and Hemolytic Glycoside Isolated from the Red Tide Alga Prymnesium parvum. J. Am. Chem. Soc. 1996, 118, 479–480. [Google Scholar] [CrossRef]
- Igarashi, T.; Satake, M.; Yasumoto, T. Structures and Partial Stereochemical Assignments for Prymnesin-1 and Prymnesin-2: Potent Hemolytic and Ichthyotoxic Glycosides Isolated from the Red Tide Alga Prymnesium parvum. J. Am. Chem. Soc. 1999, 121, 8499–8511. [Google Scholar] [CrossRef]
- Rasmussen, S.A.; Meier, S.; Andersen, N.G.; Blossom, H.E.; Duus, J.Ø.; Nielsen, K.F.; Hansen, P.J.; Larsen, T.O. Chemodiversity of Ladder-Frame Prymnesin Polyethers in Prymnesium parvum. J. Nat. Prod. 2016, 79, 2250–2256. [Google Scholar] [CrossRef] [PubMed]
- Roux, S.; Brum, J.R.; Dutilh, B.E.; Sunagawa, S.; Duhaime, M.B.; Loy, A.; Poulos, B.T.; Solonenko, N.; Lara, E.; Poulain, J.; et al. Ecogenomics and potential biogeochemical impacts of globally abundant ocean viruses. Nature 2016, 537, 689–693. [Google Scholar] [CrossRef] [PubMed]
- Karsenti, E.; Acinas, S.G.; Bork, P.; Bowler, C.; De Vargas, C.; Raes, J.; Sullivan, M.; Arendt, D.; Benzoni, F.; Claverie, J.-M.; et al. A Holistic Approach to Marine Eco-Systems Biology. PLoS Biol. 2011, 9, e1001177. [Google Scholar] [CrossRef] [PubMed]

| Genus/Species | Strain Code | Lysis with PpDNAV |
|---|---|---|
| Prymnesium parvum | 946/6 | + |
| Prymnesium parvum | 94A | - |
| Prymnesium parvum | 94C | + |
| Prymnesium parvum | 579 | - |
| Prymnesium patelliferum | 527A | + |
| Prymnesium patelliferum | 527C | + |
| Prymnesium patelliferum | 527D | - |
| Prymnesium sp. | 522 | - |
| Prymnesium sp. | 569 | - |
| Prymnesium sp. | 592 | + |
| Prymnesium sp. | 593 | - |
| Prymnesium sp. | 595 | - |
| Prymnesium sp. | 596 | - |
| Prymnesium sp. | 597 | - |
| Prymnesium sp. | 598 | - |
© 2017 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license ( http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Wagstaff, B.A.; Vladu, I.C.; Barclay, J.E.; Schroeder, D.C.; Malin, G.; Field, R.A. Isolation and Characterization of a Double Stranded DNA Megavirus Infecting the Toxin-Producing Haptophyte Prymnesium parvum. Viruses 2017, 9, 40. https://doi.org/10.3390/v9030040
Wagstaff BA, Vladu IC, Barclay JE, Schroeder DC, Malin G, Field RA. Isolation and Characterization of a Double Stranded DNA Megavirus Infecting the Toxin-Producing Haptophyte Prymnesium parvum. Viruses. 2017; 9(3):40. https://doi.org/10.3390/v9030040
Chicago/Turabian StyleWagstaff, Ben A., Iulia C. Vladu, J. Elaine Barclay, Declan C. Schroeder, Gill Malin, and Robert A. Field. 2017. "Isolation and Characterization of a Double Stranded DNA Megavirus Infecting the Toxin-Producing Haptophyte Prymnesium parvum" Viruses 9, no. 3: 40. https://doi.org/10.3390/v9030040
APA StyleWagstaff, B. A., Vladu, I. C., Barclay, J. E., Schroeder, D. C., Malin, G., & Field, R. A. (2017). Isolation and Characterization of a Double Stranded DNA Megavirus Infecting the Toxin-Producing Haptophyte Prymnesium parvum. Viruses, 9(3), 40. https://doi.org/10.3390/v9030040





