Chlamydiaphage φCPG1 Capsid Protein Vp1 Inhibits Chlamydia trachomatis Growth via the Mitogen-Activated Protein Kinase Pathway
Abstract
:1. Introduction
2. Materials and Methods
2.1. McCoy Cell Culture
2.2. C. trachomatis Infection
2.3. Vp1 Expression, Identification, and Purification
2.4. Treatments and Experimental Design
2.5. Western Blotting
2.6. Real-Time PCR
2.7. IL-1 and IL-8 Enzyme-Linked Immunosorbent Assay
2.8. Data Analysis
3. Results
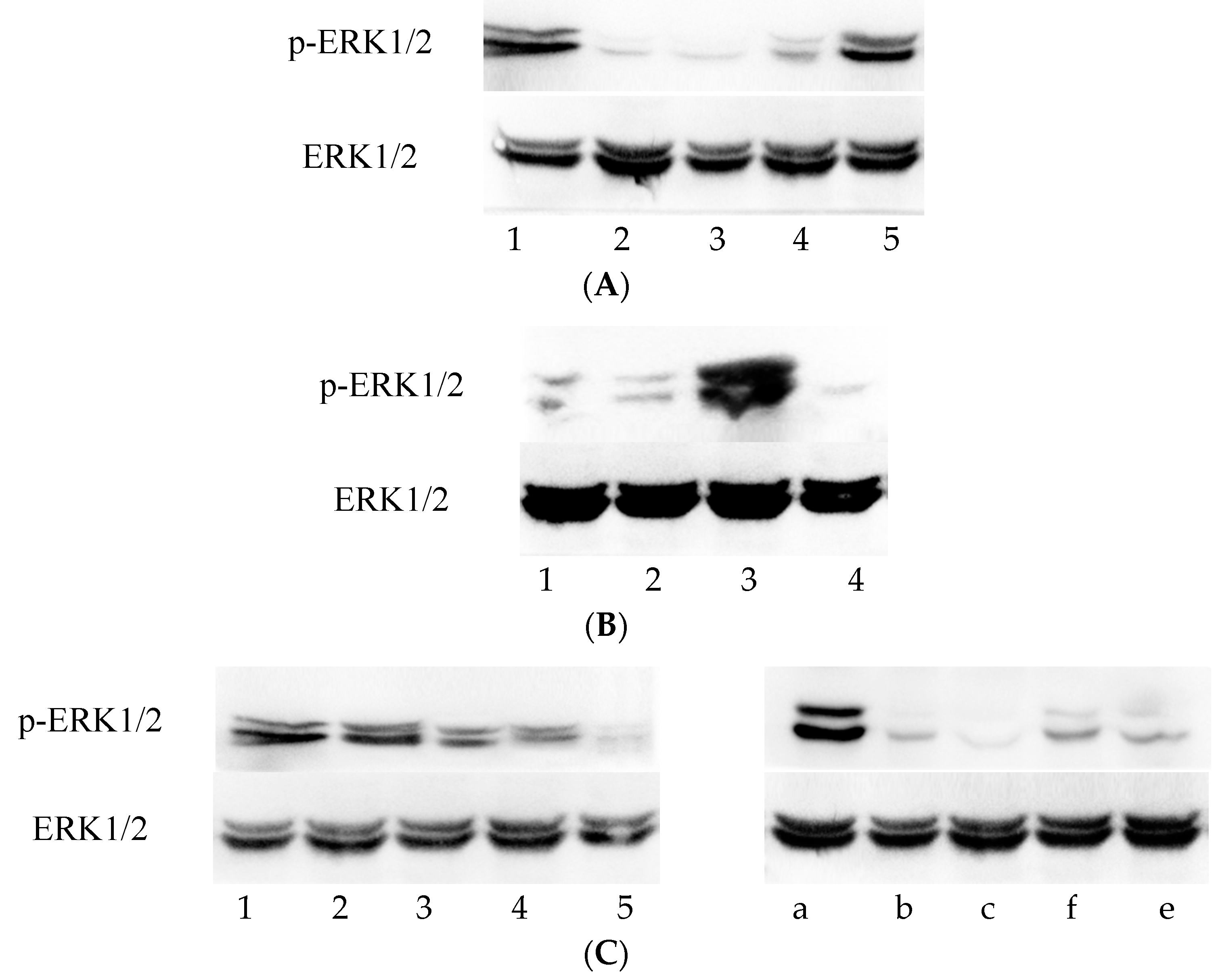
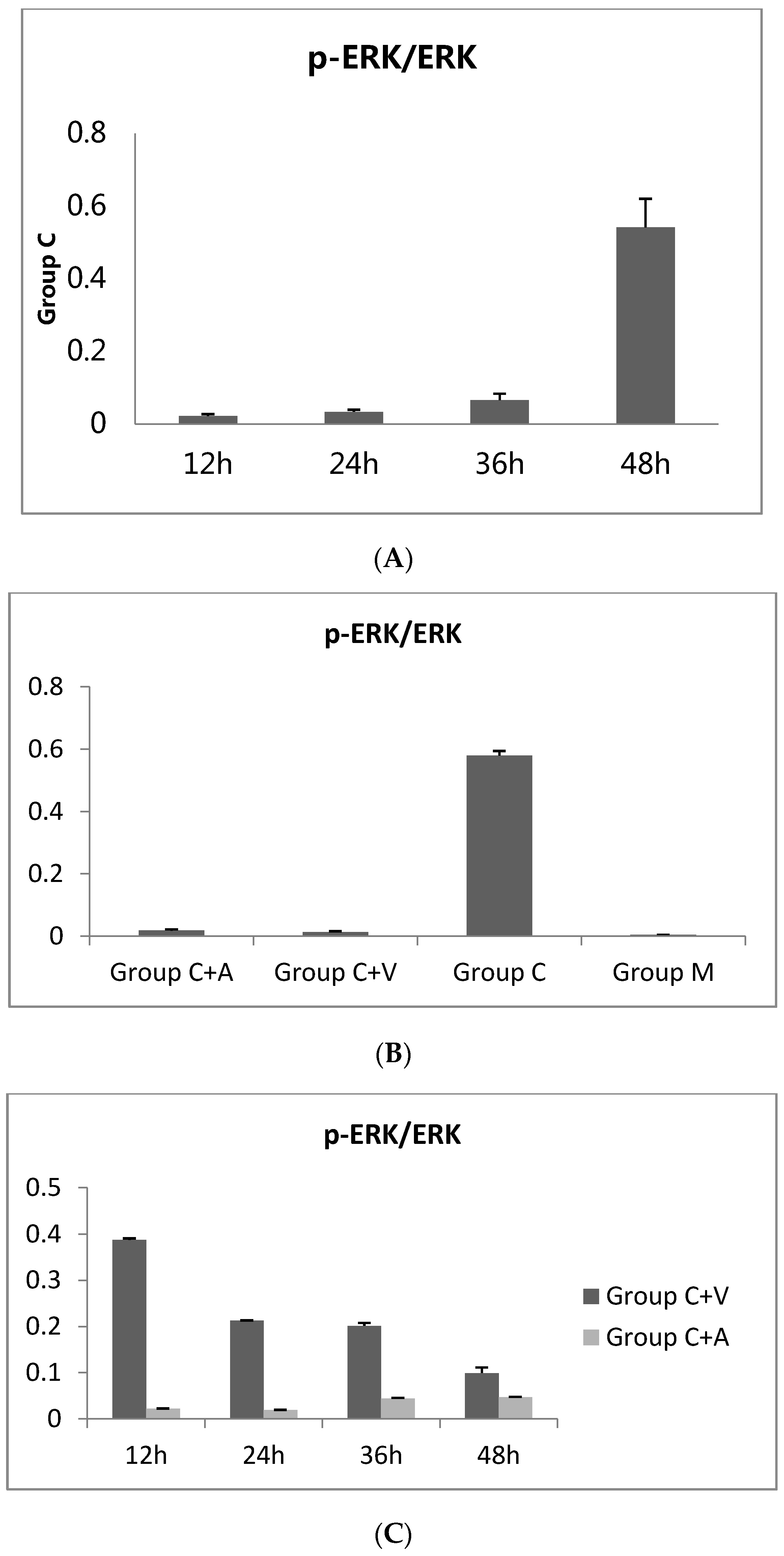
3.1. ERK1/2 Phosphorylation Increased throughout C. trachomatis Growth
3.2. p-ERK1/2 Expression Was Reduced in the Vp1- and Azithromycin-Treated Cells as the Infection Rate Decreased
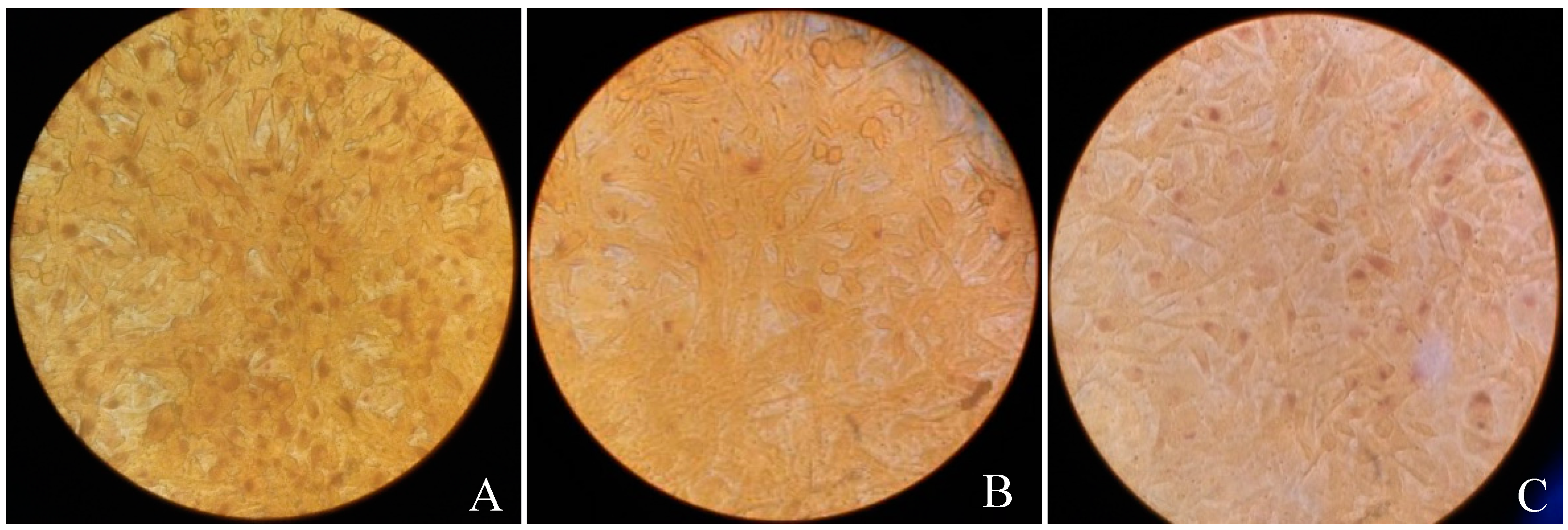
3.3. Vp1 Inhibits C. trachomatis in the Late Infection Period, Whereas Azithromycin Inhibits Growth in the Early Period
3.4. The ERK1 Gene Was Up-Regulated and the ERK2 Gene Was Downregulated after C. trachomatis Infection
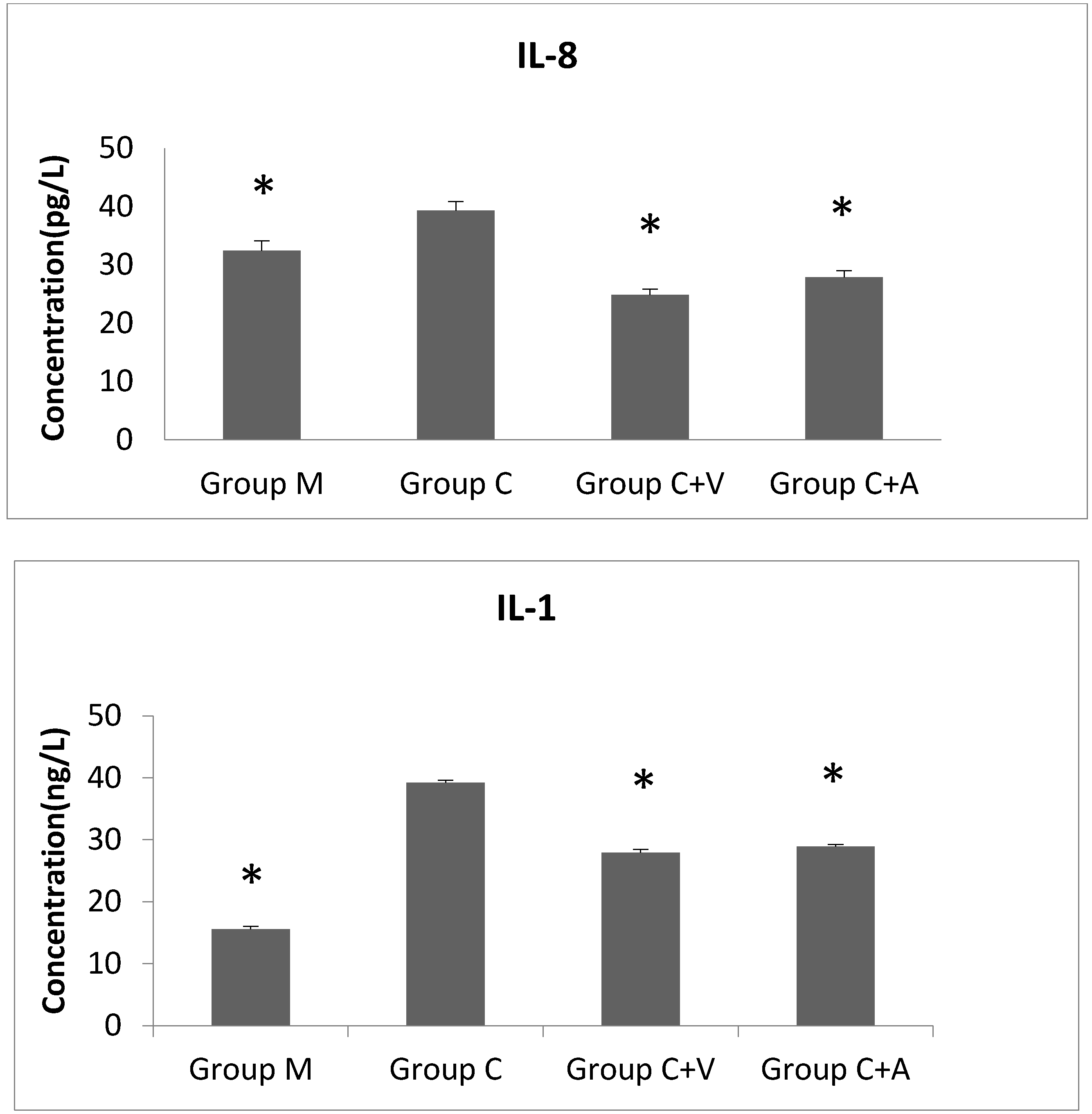
3.5. IL-8 and IL-1 Levels Decreased after Vp1 and Azithromycin Treatments
4. Discussion
5. Conclusions
Acknowledgments
Author Contributions
Conflicts of Interest
References
- Centers for Disease Control and Prevention. 2014 Sexually Transmitted Diseases Surveillance. Available online: http://www.cdc.gov/std/stats13/ (accessed on 10 February 2016).
- Parish, W.L.; Laumann, E.O.; Cohen, M.S.; Suiming, P.; Heyi, Z.; Irving, H.; Tianfu, W.; Kwai Hang, N. Population-based study of chlamydial infection in China: A hidden epidemic. JAMA 2003, 289, 1265–1273. [Google Scholar] [CrossRef] [PubMed]
- Hsia, R.C.; Ting, L.M.; Bavoil, P.M. Microvirus of Chlamydia psittaci strain guinea pig inclusion conjunctivitis: Isolation and molecular characterization. Microbiology 2000, 146, 1651–1660. [Google Scholar] [CrossRef] [PubMed]
- Liu, B.L.; Everson, J.S.; Fane, B.; Giannikopoulou, P.; Vretou, E.; Lambden, P.R.; Clarke, I.N. Molecular characterization of a bacteriophage (Chp2) from Chlamydia psittaci. J. Virol. 2000, 74, 3464–3469. [Google Scholar] [CrossRef] [PubMed]
- Read, T.D.; Brunham, R.C.; Shen, C.; Gill, S.R.; Heidelberg, J.F.; White, O.; Hickey, E.K.; Peterson, J.; Utterback, T.; Berry, K.; et al. Genome sequences of Chlamydia trachomatis MoPn and Chlamydia pneumoniae AR39. Nucleic Acids Res. 2000, 28, 1397–1406. [Google Scholar] [CrossRef] [PubMed]
- Sliwa-Dominiak, J.; Suszynska, E.; Pawlikowska, M.; Deptula, W. Chlamydia bacteriophages. Arch. Microbiol. 2013, 195, 765–771. [Google Scholar] [CrossRef] [PubMed]
- Liu, Y.-J.; Hou, S.-P.; Wei, J.-R.; Li, Y.; Qi, M.-L.; Wang, H.-P.; Liu, Q.-Z. The effect of Chlamydiaphage phiCPG1 capsid protein Vp1 on the Chlamydia trachomatis. Chin. J. Microbiol. Immunol. 2012, 32, 403–408. (In Chinese) [Google Scholar]
- Jürgen, R.D.; Christina, G.; Hangxing, Y.; Katharina, W.; Otto, G.P.; Elisabeth, L.T.; Vera, F.B.; Eberhard, S. Persistent Chlamydia trachomatis infection of HeLa cells mediates apoptosis resistance through a Chlamydia protease-like activity factor-independent mechanism and induces high mobility group box 1 release. Infect. Immunity 2012, 80, 195–205. [Google Scholar]
- Wen, C.; Pooja, S.; Youmin, Z.; Ding, C.; Zhongyu, L.; Guangming, Z. Intracellular interleukin-1α mediates interleukin-8 production induced by Chlamydia trachomatis infection via a mechanism independent of type I interleukin-1 receptor. Infect. Immun. 2008, 76, 942–951. [Google Scholar]
- Du, K.; Wang, F.-Y.; Huo, Z. Development of Chlamydia trachomatis is dependent on the MEK/ERK survival pathways. Life Sci. Res. 2010, 14, 424–426. (In Chinese) [Google Scholar]
- Kavathas, P.B.; Boeras, C.M.; Mulla, M.J.; Abrahams, V.M. Nod1, but not the ASC inflammasome, contributes to induction of IL-1β secretion in human trophoblasts after sensing of Chlamydia trachomatis. Mucosal Immunol. 2013, 6, 235–243. [Google Scholar] [CrossRef] [PubMed]
- Chen, F.; Cheng, W.; Zhang, S.; Zhong, G.; Ping, Y.U. Induction of IL-8 by Chlamydia trachomatis through MAPK pathway rather than NF-κB pathway. J. Cent. South Univ. 2010, 35, 307–313. [Google Scholar]
- Wang, Y.X.; Li, L.-T. Detection of IL-2,IL-6 and IL-8 in Chlamydia trachomatis-infected reproductive tract in pregnant women. J. Shanxi Med. Univ. 2009, 05, 472–474. (In Chinese) [Google Scholar]
- Patton, D.L.; Sweeney, Y.T.; Stamm, W.E. Significant reduction in inflammatory response in the macaque model of Chlamydial pelvic inflammatory disease with azithromycin treatment. J. Infect. Dis. 2005, 192, 129–135. [Google Scholar] [CrossRef] [PubMed]
- Srivastava, P.; Vardhan, H.; Bhengraj, A.R.; Jha, R.; Singh, L.C.; Salhan, S.; Mittal, A. Azithromycin treatment modulates the extracellular signal-regulated kinase mediated pathway and inhibits inflammatory cytokines and chemokines in epithelial cells from infertile women with recurrent chlamydia trachomatis infection. DNA Cell Biol. 2011, 30, 545–554. [Google Scholar] [CrossRef] [PubMed]
- Liu, Q.-Z.; Yaom, W.-F.; Qi, M.-L.; Li, Y.; Wang, H.-P.; Tian, J.-Q. Cloning, expression and identification of chlamydial GPIC capsid Vp1 protein. Chin. J. Dermatol. 2006, 39, 714–716. (In Chinese) [Google Scholar]
- Wang, X.; Seed, B. A PCR primer bank for quantitative gene expression analysis. Nucleic Acids Res. 2003, 31. [Google Scholar] [CrossRef]
- Fremin, C.; Ezan, F.; Boisselier, P.; Bessard, A.; Pages, G.; Pouyssegur, J.; Baffet, G. ERK2 but not ERK1 plays a key role in hepatocyte replication: An RNAi-mediated ERK2 knockdown approach in wild-type and ERK1 null hepatocytes. Hepatology 2007, 45, 1035–1045. [Google Scholar] [CrossRef] [PubMed]
- Potterat, J.J. 2010 European guideline for Chlamydia trachomatis infections: Recommending partner notification look-back periods. Int. J. STD AIDS 2011, 22. [Google Scholar] [CrossRef] [PubMed]
- Hafner, L.M.; Wilson, D.P.; Timms, P. Development status and future prospects for a vaccine against Chlamydia trachomatis infection. Vaccine 2014, 32, 1563–1571. [Google Scholar] [CrossRef] [PubMed]
- Golkar, Z.; Bagasra, O.; Pace, D.G. Bacteriophage therapy: A potential solution for the antibiotic resistance crisis. J. Infect. Dev. Ctries 2014, 8, 129–136. [Google Scholar] [CrossRef] [PubMed]
- Elizabeth, K.; Daniel, D.V.; Guram, G.; Zemphira, A.; Lasha, G.; Sarah, K.; Abedon, S.T. Phage therapy in clinical practice: Treatment of human infections. Curr. Pharm. Biotechnol. 2010, 11, 69–86. [Google Scholar]
| Group C | Group C + V | Group C + A | Group V | Group A | Group M | |
|---|---|---|---|---|---|---|
| C. trachomatis | √ | √ | √ | |||
| Vp1 | √ | √ | ||||
| Azithromycin | √ | √ | ||||
| McCoy cell | √ | √ | √ | √ | √ | √ |
| Gene | Primer Sequences (5′ to 3′) | Products Size (bp) |
|---|---|---|
| Mapk1 (ERK2) | F GTATTCTTGGATCTCCATCACAGG | 246 |
| R TGGGCTCATCACTTGGGTCA | ||
| Mapk3 (ERK1) | F CAAACAAGCGCATCACAGTAGA | 113 |
| R CAGCTCCATGTCGAAGGTGAAT | ||
| Actin | F GCCTTCCTTCTTGGGTAT | 97 |
| R GGCATAGAGGTCTTTACGG |
© 2016 by the authors; licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons by Attribution (CC-BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Guo, Y.; Guo, R.; Zhou, Q.; Sun, C.; Zhang, X.; Liu, Y.; Liu, Q. Chlamydiaphage φCPG1 Capsid Protein Vp1 Inhibits Chlamydia trachomatis Growth via the Mitogen-Activated Protein Kinase Pathway. Viruses 2016, 8, 99. https://doi.org/10.3390/v8040099
Guo Y, Guo R, Zhou Q, Sun C, Zhang X, Liu Y, Liu Q. Chlamydiaphage φCPG1 Capsid Protein Vp1 Inhibits Chlamydia trachomatis Growth via the Mitogen-Activated Protein Kinase Pathway. Viruses. 2016; 8(4):99. https://doi.org/10.3390/v8040099
Chicago/Turabian StyleGuo, Yuanli, Rui Guo, Quan Zhou, Changgui Sun, Xinmei Zhang, Yuanjun Liu, and Quanzhong Liu. 2016. "Chlamydiaphage φCPG1 Capsid Protein Vp1 Inhibits Chlamydia trachomatis Growth via the Mitogen-Activated Protein Kinase Pathway" Viruses 8, no. 4: 99. https://doi.org/10.3390/v8040099
APA StyleGuo, Y., Guo, R., Zhou, Q., Sun, C., Zhang, X., Liu, Y., & Liu, Q. (2016). Chlamydiaphage φCPG1 Capsid Protein Vp1 Inhibits Chlamydia trachomatis Growth via the Mitogen-Activated Protein Kinase Pathway. Viruses, 8(4), 99. https://doi.org/10.3390/v8040099





