Cedratvirus, a Double-Cork Structured Giant Virus, is a Distant Relative of Pithoviruses
Abstract
:1. Introduction
2. Materials and Methods
2.1. Culture and Isolation Procedures
2.2. End-Point Dilution and Centrifugation Methods
2.3. Cytometry Applications
2.4. Electron Microscopy and Infectious Cycle Description
2.5. Genome Sequencing
2.6. Genome Assembly
2.7. Study of the Genome Organization and Genome Annotation
2.8. Phylogenetic Analyses
3. Results
3.1. Isolation of Cedratvirus
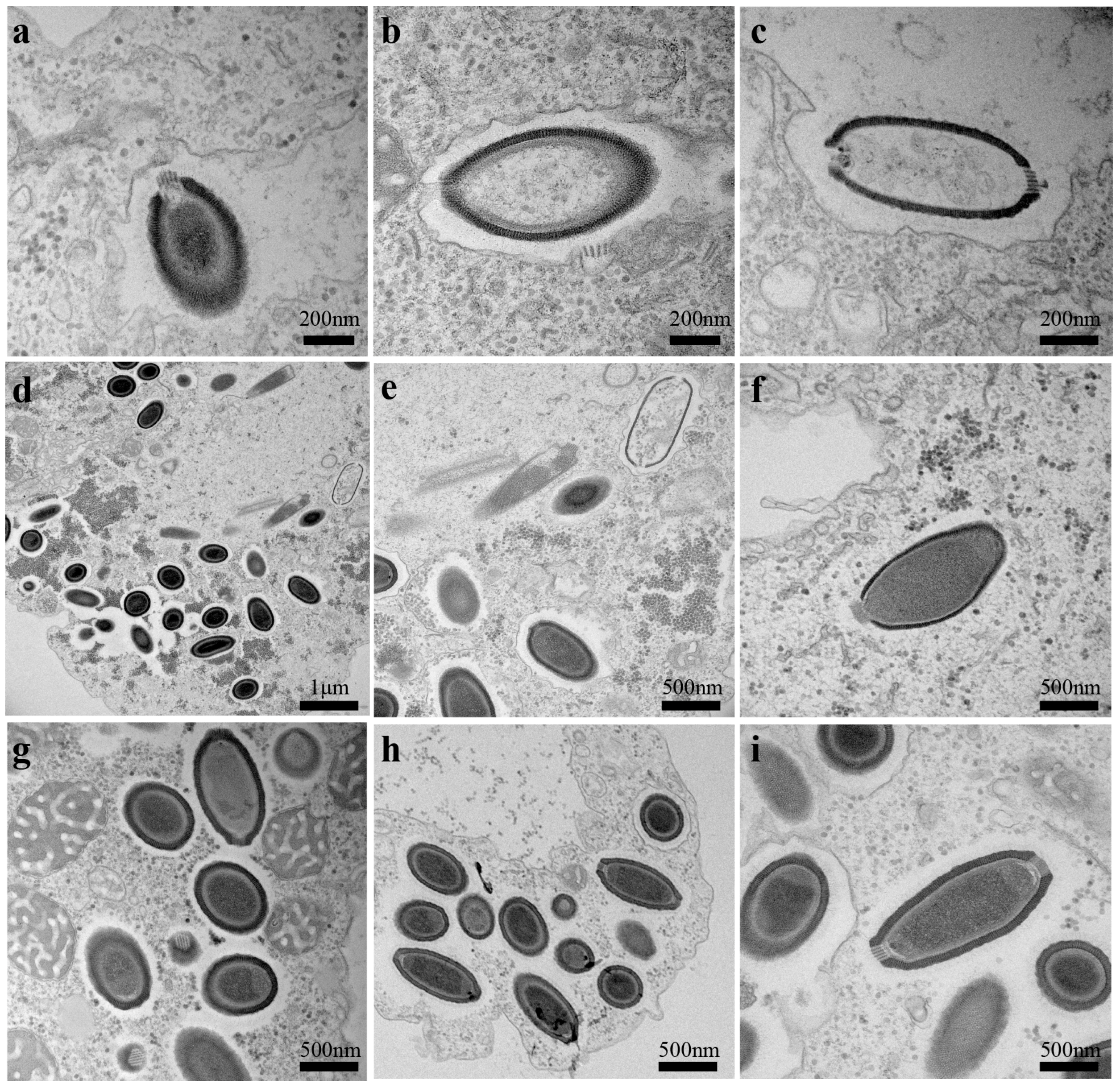
3.2. Ultrastructure and Replicative Cycle of Cedratvirus
3.3. Genome Analysis
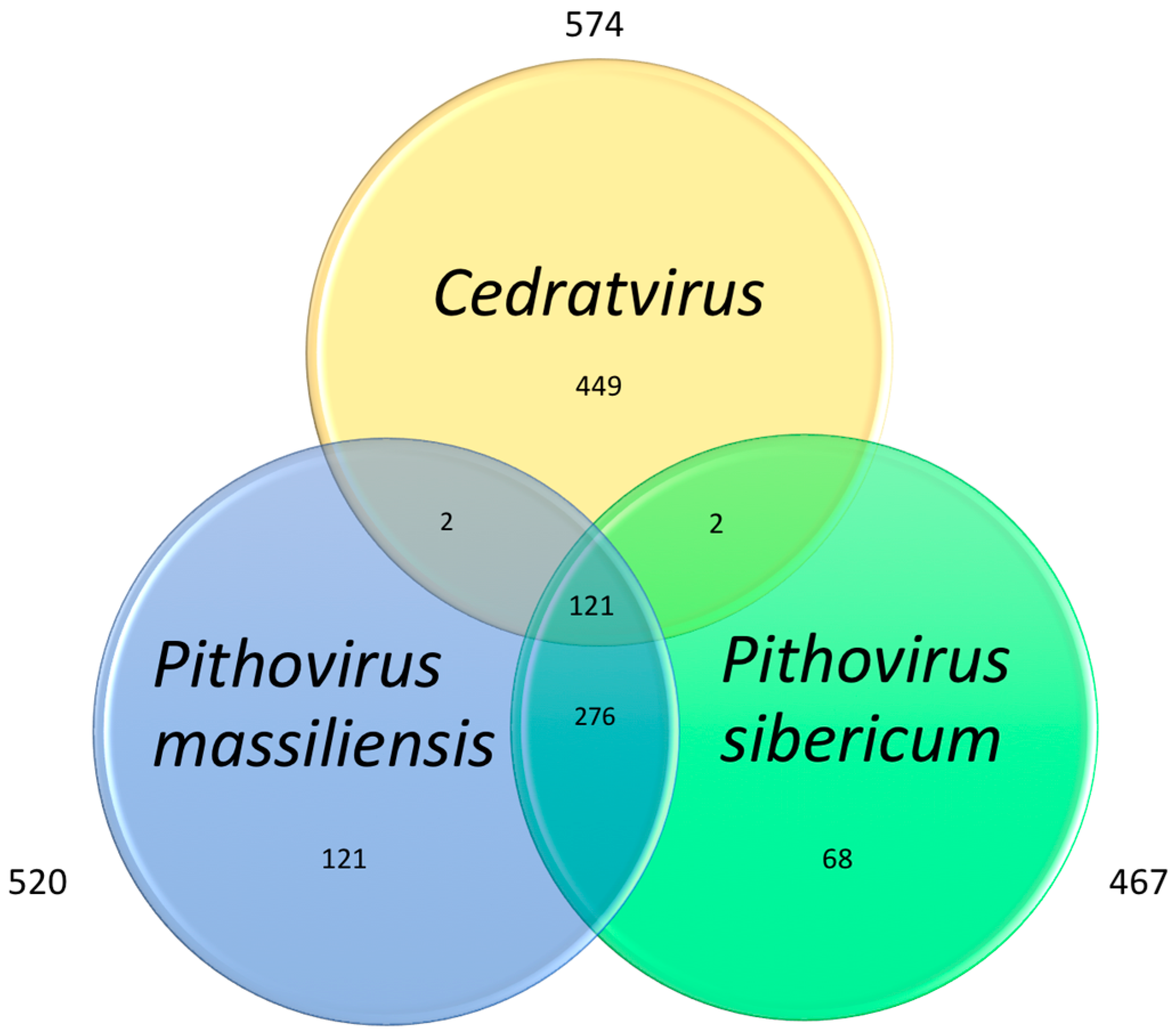
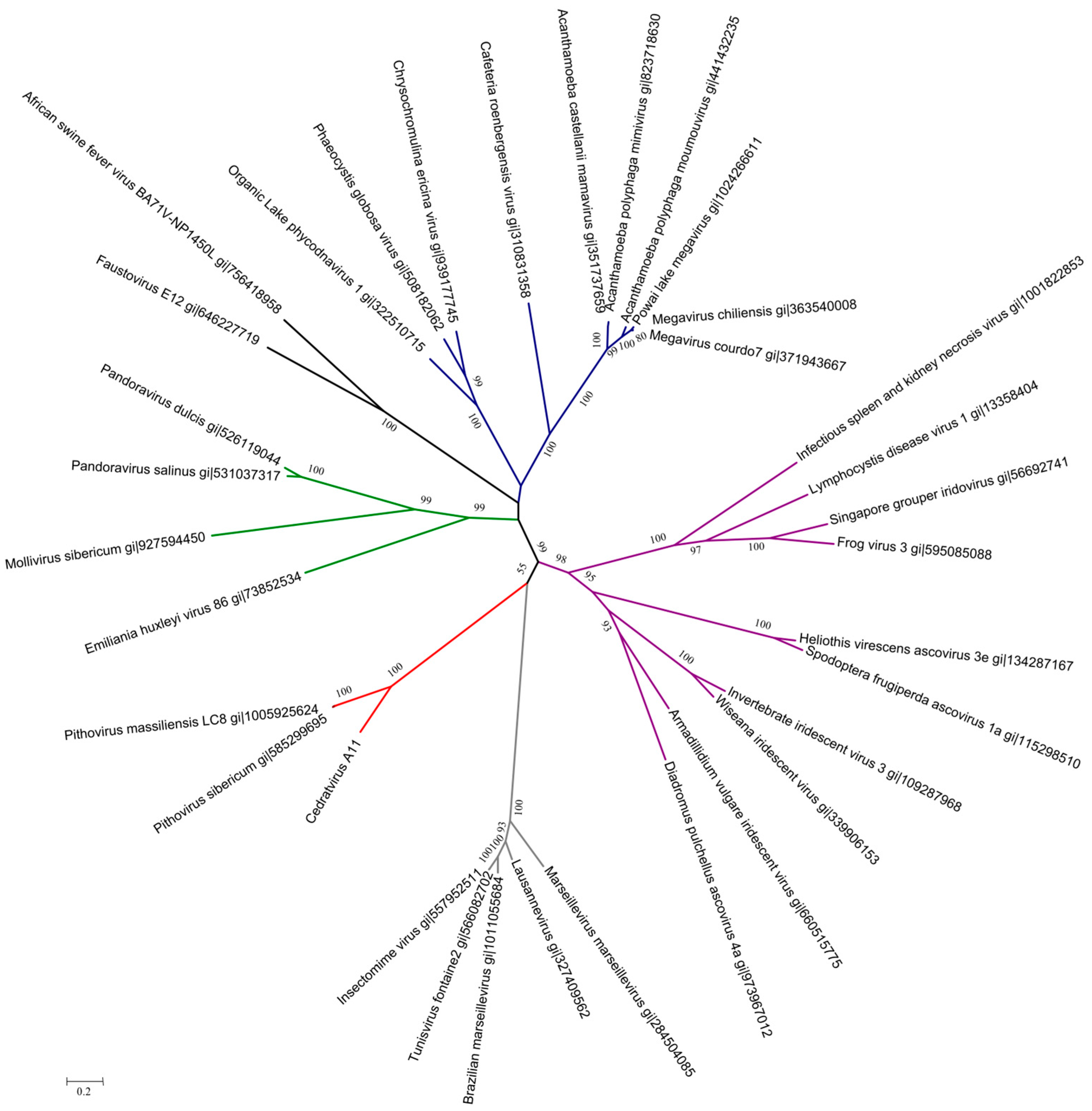
3.4. Cedratvirus, Pithovirus sibericum and Pithovirus massiliensis: Three Members of a Putative Extended Family
4. Discussion
Supplementary Materials
Author Contributions
Conflicts of Interest
References
- Forterre, P. To be or not to be alive: How recent discoveries challenge the traditional definitions of viruses and life. Stud. Hist. Philos. Sci. C Stud. Hist. Philos. Biol. Biomed. Sci. 2016, 59, 100–108. [Google Scholar] [CrossRef] [PubMed]
- La Scola, B. A Giant Virus in Amoebae. Science 2003, 299, 2033. [Google Scholar] [CrossRef] [PubMed]
- Boyer, M.; Yutin, N.; Pagnier, I.; Barrassi, L.; Fournous, G.; Espinosa, L.; Robert, C.; Azza, S.; Sun, S.; Rossmann, M.G.; et al. Giant Marseillevirus highlights the role of amoebae as a melting pot in emergence of chimeric microorganisms. Proc. Natl. Acad. Sci. USA 2009, 106, 21848–21853. [Google Scholar] [CrossRef] [PubMed]
- Reteno, D.G.; Benamar, S.; Khalil, J.B.; Andreani, J.; Armstrong, N.; Klose, T.; Rossmann, M.; Colson, P.; Raoult, D.; La Scola, B. Faustovirus, an asfarvirus-related new lineage of giant viruses infecting amoebae. J. Virol. 2015, 89, 6585–6594. [Google Scholar] [CrossRef] [PubMed]
- Scheid, P.; Zöller, L.; Pressmar, S.; Richard, G.; Michel, R. An extraordinary endocytobiont in Acanthamoeba sp. isolated from a patient with keratitis. Parasitol. Res. 2008, 102, 945–950. [Google Scholar] [CrossRef] [PubMed]
- Philippe, N.; Legendre, M.; Doutre, G.; Couté, Y.; Poirot, O.; Lescot, M.; Arslan, D.; Seltzer, V.; Bertaux, L.; Bruley, C.; et al. Pandoraviruses: Amoeba viruses with genomes up to 2.5 Mb reaching that of parasitic eukaryotes. Science 2013, 341, 281–286. [Google Scholar] [CrossRef] [PubMed]
- Legendre, M.; Bartoli, J.; Shmakova, L.; Jeudy, S.; Labadie, K.; Adrait, A.; Lescot, M.; Poirot, O.; Bertaux, L.; Bruley, C.; et al. Thirty-thousand-year-old distant relative of giant icosahedral DNA viruses with a pandoravirus morphology. Proc. Natl. Acad. Sci. USA 2014, 111, 4274–4279. [Google Scholar] [CrossRef] [PubMed]
- Legendre, M.; Lartigue, A.; Bertaux, L.; Jeudy, S.; Bartoli, J.; Lescot, M.; Alempic, J.-M.; Ramus, C.; Bruley, C.; Labadie, K.; et al. In-depth study of Mollivirus sibericum, a new 30,000-y-old giant virus infecting Acanthamoeba. Proc. Natl. Acad. Sci. USA 2015, 112, E5327–E5335. [Google Scholar] [CrossRef] [PubMed]
- Levasseur, A.; Andreani, J.; Delerce, J.; Bou Khalil, J.; Robert, C.; La Scola, B.; Raoult, D. Comparison of a modern and fossil Pithovirus reveals its genetic conservation and evolution. Genome Biol. Evol. 2016, 8, 2333–2339. [Google Scholar] [CrossRef] [PubMed]
- Bou Khalil, J.Y.; Andreani, J.; Raoult, D.; La Scola, B. A Rapid strategy for the isolation of new faustoviruses from environmental samples using Vermamoeba vermiformis. J. Vis. Exp. 2016, e54104. [Google Scholar] [CrossRef] [PubMed]
- Khalil, J.Y.B.; Andreani, J.; La Scola, B. Updating strategies for isolating and discovering giant viruses. Curr. Opin. Microbiol. 2016, 31, 80–87. [Google Scholar] [CrossRef] [PubMed]
- Khalil, J.Y.B.; Robert, S.; Reteno, D.G.; Andreani, J.; Raoult, D.; La Scola, B. High-throughput isolation of giant viruses in liquid medium using automated flow cytometry and fluorescence staining. Front. Microbiol. 2016, 7, 26. [Google Scholar] [CrossRef] [PubMed]
- Bou Khalil, J.Y.; Langlois, T.; Andreani, J.; Soraing, J.M.; Raoult, D.; Camoin, L.; La Scola, B. Flow cytometry sorting to separate viable giant viruses from amoeba co-culture supernatants. Front Cell Infect Microbiol. 2016. under review. [Google Scholar]
- Bou Khalil, J.Y.; Benamar, S.; Baudoin, J.-P.; Croce, O.; Blanc-Tailleur, C.; Pagnier, I.; Raoult, D.; La Scola, B. Developmental cycle and genome analysis of “Rubidus massiliensis”, a new Vermamoeba vermiformis pathogen. Front. Cell Infect. Microbiol. 2016, 6, 31. [Google Scholar] [CrossRef] [PubMed]
- Ye, J.; Coulouris, G.; Zaretskaya, I.; Cutcutache, I.; Rozen, S.; Madden, T.L. Primer-BLAST: A tool to design target-specific primers for polymerase chain reaction. BMC Bioinform. 2012, 13, 134. [Google Scholar] [CrossRef] [PubMed]
- Simpson, J.T.; Wong, K.; Jackman, S.D.; Schein, J.E.; Jones, S.J.M.; Birol, I. ABySS: A parallel assembler for short read sequence data. Genome Res. 2009, 19, 1117–1123. [Google Scholar] [CrossRef] [PubMed]
- Boetzer, M.; Henkel, C.V.; Jansen, H.J.; Butler, D.; Pirovano, W. Scaffolding pre-assembled contigs using SSPACE. Bioinformatics 2011, 27, 578–579. [Google Scholar] [CrossRef] [PubMed]
- Grissa, I.; Vergnaud, G.; Pourcel, C. CRISPRFinder: A web tool to identify clustered regularly interspaced short palindromic repeats. Nucleic Acids Res. 2007, 35, W52–W57. [Google Scholar] [CrossRef] [PubMed]
- Besemer, J.; Lomsadze, A.; Borodovsky, M. GeneMarkS: A self-training method for prediction of gene starts in microbial genomes. Implications for finding sequence motifs in regulatory regions. Nucleic Acids Res. 2001, 29, 2607–2618. [Google Scholar] [CrossRef] [PubMed]
- Schattner, P.; Brooks, A.N.; Lowe, T.M. The tRNAscan-SE, snoscan and snoGPS web servers for the detection of tRNAs and snoRNAs. Nucleic Acids Res. 2005, 33, W686–W689. [Google Scholar] [CrossRef] [PubMed]
- Laslett, D.; Canback, B. ARAGORN, a program to detect tRNA genes and tmRNA genes in nucleotide sequences. Nucleic Acids Res. 2004, 32, 11–16. [Google Scholar] [CrossRef] [PubMed]
- Yutin, N.; Wolf, Y.I.; Raoult, D.; Koonin, E.V. Eukaryotic large nucleo-cytoplasmic DNA viruses: Clusters of orthologous genes and reconstruction of viral genome evolution. Virol. J. 2009, 6, 223. [Google Scholar] [CrossRef] [PubMed]
- Howe, E.; Holton, K.; Nair, S.; Schlauch, D.; Sinha, R.; Quackenbush, J. MeV: MultiExperiment Viewer. In Biomedical Informatics for Cancer Research; Springer: Boston, MA, USA, 2010; pp. 267–277. [Google Scholar]
- Boratyn, G.M.; Schäffer, A.A.; Agarwala, R.; Altschul, S.F. Domain enhanced lookup time accelerated BLAST. Biol. Direct 2012. [Google Scholar] [CrossRef] [PubMed]
- Lechner, M.; Findeiß, S.; Steiner, L.; Marz, M.; Stadler, P.F.; Prohaska, S.J. Proteinortho: Detection of (Co-)orthologs in large-scale analysis. BMC Bioinform. 2011, 12, 124. [Google Scholar] [CrossRef] [PubMed]
- Alva, V.; Nam, S.-Z.; Söding, J.; Lupas, A.N. The MPI bioinformatics Toolkit as an integrative platform for advanced protein sequence and structure analysis. Nucleic Acids Res. 2016, 44, W410–W415. [Google Scholar] [CrossRef] [PubMed]
- Edgar, R.C. MUSCLE: A multiple sequence alignment method with reduced time and space complexity. BMC Bioinform. 2004, 5, 113. [Google Scholar] [CrossRef] [PubMed]
- Price, M.N.; Dehal, P.S.; Arkin, A.P. FastTree: Computing large minimum evolution trees with profiles instead of a distance matrix. Mol. Biol. Evol. 2009, 26, 1641–1650. [Google Scholar] [CrossRef] [PubMed]
- Letunic, I.; Bork, P. Interactive tree of life (iTOL) v3: An online tool for the display and annotation of phylogenetic and other trees. Nucleic Acids Res. 2016, 44, W242–W245. [Google Scholar] [CrossRef] [PubMed]
- Fischer, M.G. Giant viruses come of age. Curr. Opin. Microbiol. 2016, 31, 50–57. [Google Scholar] [CrossRef] [PubMed]
- Aherfi, S.; La Scola, B.; Pagnier, I.; Raoult, D.; Colson, P. The expanding family Marseilleviridae. Virology 2014, 466–467, 27–37. [Google Scholar] [CrossRef] [PubMed]
- Dornas, F.P.; Assis, F.L.; Aherfi, S.; Arantes, T.; Abrahão, J.S.; Colson, P.; La Scola, B. A Brazilian marseillevirus is the founding member of a lineage in family Marseilleviridae. Viruses 2016, 8, 76. [Google Scholar] [CrossRef] [PubMed]
- Santini, S.; Jeudy, S.; Bartoli, J.; Poirot, O.; Lescot, M.; Abergel, C.; Barbe, V.; Wommack, K.E.; Noordeloos, A.A.M.; Brussaard, C.P.D.; et al. Genome of Phaeocystis globosa virus PgV-16T highlights the common ancestry of the largest known DNA viruses infecting eukaryotes. Proc. Natl. Acad. Sci. USA 2013, 110, 10800–10805. [Google Scholar] [CrossRef] [PubMed]
- Yutin, N.; Colson, P.; Raoult, D.; Koonin, E.V. Mimiviridae: Clusters of orthologous genes, reconstruction of gene repertoire evolution and proposed expansion of the giant virus family. Virol. J. 2013, 10, 106. [Google Scholar] [CrossRef] [PubMed]
| Cedratvirus A11 | Pithovirus sibericum | Pithovirus massilliensis | |
|---|---|---|---|
| Genome size (bp) | 589,068 | 610,033 | 686,015 |
| GC % | 42.6 | 35.8 | 35.4 |
| Genes | 574 | 467 | 520 |
| tRNA | 0 | 0 | 0 |
| Coding capacity (%) | 78.5 | 69.0 | 64.0 |
| Global morphology | Ovoid particle with double corks | Ovoid particle with a single cork | Ovoid particle with a single cork |
© 2016 by the authors; licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC-BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Andreani, J.; Aherfi, S.; Bou Khalil, J.Y.; Di Pinto, F.; Bitam, I.; Raoult, D.; Colson, P.; La Scola, B. Cedratvirus, a Double-Cork Structured Giant Virus, is a Distant Relative of Pithoviruses. Viruses 2016, 8, 300. https://doi.org/10.3390/v8110300
Andreani J, Aherfi S, Bou Khalil JY, Di Pinto F, Bitam I, Raoult D, Colson P, La Scola B. Cedratvirus, a Double-Cork Structured Giant Virus, is a Distant Relative of Pithoviruses. Viruses. 2016; 8(11):300. https://doi.org/10.3390/v8110300
Chicago/Turabian StyleAndreani, Julien, Sarah Aherfi, Jacques Yaacoub Bou Khalil, Fabrizio Di Pinto, Idir Bitam, Didier Raoult, Philippe Colson, and Bernard La Scola. 2016. "Cedratvirus, a Double-Cork Structured Giant Virus, is a Distant Relative of Pithoviruses" Viruses 8, no. 11: 300. https://doi.org/10.3390/v8110300
APA StyleAndreani, J., Aherfi, S., Bou Khalil, J. Y., Di Pinto, F., Bitam, I., Raoult, D., Colson, P., & La Scola, B. (2016). Cedratvirus, a Double-Cork Structured Giant Virus, is a Distant Relative of Pithoviruses. Viruses, 8(11), 300. https://doi.org/10.3390/v8110300







