Plant Translation Factors and Virus Resistance
Abstract
:1. Introduction
2. Biological Functions of Translation Factors in Plants
2.1. Translation of Cellular mRNAs
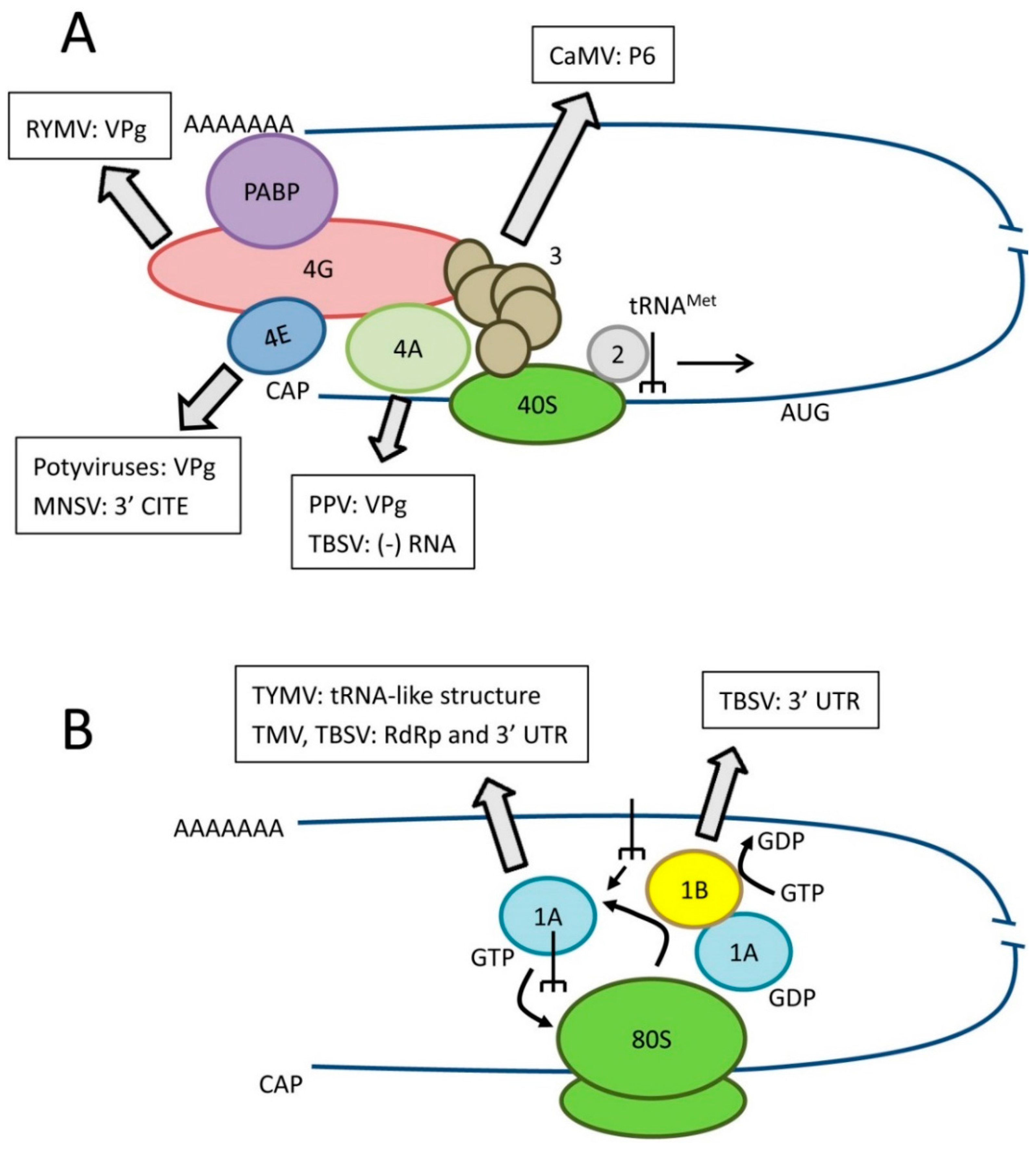
2.2. Additional Biological Functions of Translation Factors
3. Characterization of the Interactions between Cellular Translation Factors and Plant Viruses and Identification of Potential Sources of Viral Resistance
3.1. Translation Factors Facilitating Non-Canonical Viral RNA Translation
3.1.1. Core eIF4F/iso4F Components Recruited by Viral RNAs or Viral VPg Proteins
3.1.2. eIF4A-Like DEAD-Box Helicases Assisting the Translation of Bromovirus and Tombusvirus RNAs
3.1.3. Recruitment of eEF1A by tRNA-Like Structure at the 3′ End of Viral RNAs
3.1.4. Recruitment of an eIF3 Subunit and Associated Factors by Cauliflower Mosaic Virus for Translation Re-Initiation
3.2. Translation factors Regulating Viral RNA Replication
3.2.1 Recruitment of DEAD-Box RNA Helicases to Tombusvirus and Potyvirus VRCs
3.2.2. Regulation of Tymovirus, Tobamovirus and Tombusvirus Replication by eEF1A
3.3. Other Roles for Translation Factors in Enhancing the Virus Infection Cycle
5. Concluding Remarks
Acknowledgments
Conflicts of Interest
References
- Sanfacon, H.; Jovel, J. Interactions Between plant and Virus Proteomes in Susceptible Hosts: Identification of New Targets for Antiviral Strategies. In Biotechnology and Plant Disease Management; Punja, Z.K., De Boer, S.H., Sanfacon, H., Eds.; CAB International: Wallingford, UK, 2007; pp. 87–108. [Google Scholar]
- Nagy, P.D.; Pogany, J. The dependence of viral RNA replication on co-opted host factors. Nat. Rev. Microbiol. 2012, 10, 137–149. [Google Scholar] [CrossRef] [PubMed]
- Wang, A. Dissecting the Molecular Network of Virus-Plant Interactions: The Complex Roles of Host Factors. Annu. Rev. Phytopathol. 2015, 53. [Google Scholar] [CrossRef] [PubMed]
- Zhang, J.; Roberts, R.; Rakotondrafara, A.M. The role of the 5′ untranslated regions of Potyviridae in translation. Virus Res. 2015. [Google Scholar] [CrossRef] [PubMed]
- Simon, A.E.; Miller, W.A. 3′ cap-independent translation enhancers of plant viruses. Annu. Rev. Microbiol. 2013, 67, 21–42. [Google Scholar] [CrossRef] [PubMed]
- Wang, A.; Krishnaswamy, S. Eukaryotic translation initiation factor 4E-mediated recessive resistance to plant viruses and its utility in crop improvement. Mol. Plant Pathol. 2012, 13, 795–803. [Google Scholar] [CrossRef] [PubMed]
- Le Gall, O.; Aranda, M.A.; Caranta, C. Plant resistance to viruses mediated by translation initiation factors. In Recent Advances in Plant Virology; Caranta, C., Aranda, M.A., Tepfer, M., Lopez-Moya, J., Eds.; Caister Academic Press: Wymondham, Norfolk, VA, USA, 2011; pp. 177–194. [Google Scholar]
- Robaglia, C.; Caranta, C. Translation initiation factors: a weak link in plant RNA virus infection. Trends Plant Sci. 2006, 11, 40–45. [Google Scholar] [CrossRef] [PubMed]
- Maule, A.J.; Caranta, C.; Boulton, M.I. Sources of natural resistance to plant viruses: Status and prospects. Mol. Plant Pathol. 2007, 8, 223–231. [Google Scholar] [CrossRef] [PubMed]
- Calvo, M.; Martinez-Turino, S.; Garcia, J.A. Resistance to Plum pox virus Strain C in Arabidopsis thaliana and Chenopodium foetidum involves genome-linked viral protein and other viral determinants and might depend on compatibility with host translation initiation factors. Mol. Plant Microbe Interact. 2014, 27, 1291–1301. [Google Scholar] [CrossRef] [PubMed]
- Nieto, C.; Rodriguez-Moreno, L.; Rodriguez-Hernandez, A.M.; Aranda, M.A.; Truniger, V. Nicotiana benthamiana resistance to non-adapted Melon necrotic spot virus results from an incompatible interaction between virus RNA and translation initiation factor 4E. The Plant J. 2011, 66, 492–501. [Google Scholar] [CrossRef] [PubMed]
- Ghoshal, B.; Sanfacon, H. Symptom recovery in virus-infected plants: Revisiting the role of RNA silencing mechanisms. Virology 2015, 479–480, 167–179. [Google Scholar] [CrossRef] [PubMed]
- Jackson, R.J.; Hellen, C.U.; Pestova, T.V. The mechanism of eukaryotic translation initiation and principles of its regulation. Nat. Rev. Mol. Cell. Biol. 2010, 11, 113–127. [Google Scholar] [CrossRef] [PubMed]
- Valasek, L.S. “Ribozoomin”—Translation initiation from the perspective of the ribosome-bound eukaryotic initiation factors (eIFs). Curr. Protein Pept. Sci. 2012, 13, 305–330. [Google Scholar] [CrossRef] [PubMed]
- Browning, K.S. Plant translation initiation factors: it is not easy to be green. Biochem. Soc. Trans. 2004, 32, 589–591. [Google Scholar] [CrossRef] [PubMed]
- Kawaguchi, R.; Bailey-Serres, J. Regulation of translational initiation in plants. Curr. Opin. Plant Biol. 2002, 5, 460–465. [Google Scholar] [CrossRef]
- Patrick, R.M.; Browning, K.S. The eIF4F and eIFiso4F Complexes of Plants: An Evolutionary Perspective. Comp. Funct. Genomics 2012, 2012. [Google Scholar] [CrossRef] [PubMed]
- Roy, B.; von Arnim, A.G. Translational regulation of cytoplasmic mRNAs. Arabidopsis Book 2013, 11. [Google Scholar] [CrossRef] [PubMed]
- Bush, M.S.; Hutchins, A.P.; Jones, A.M.; Naldrett, M.J.; Jarmolowski, A.; Lloyd, C.W.; Doonan, J.H. Selective recruitment of proteins to 5′ cap complexes during the growth cycle in Arabidopsis. Plant J. 2009, 59, 400–412. [Google Scholar] [CrossRef] [PubMed]
- Browning, K.S.; Humphreys, J.; Hobbs, W.; Smith, G.B.; Ravel, J.M. Determination of the amounts of the protein synthesis initiation and elongation factors in wheat germ. J. Biol. Chem. 1990, 265, 17967–17973. [Google Scholar] [PubMed]
- Duprat, A.; Caranta, C.; Revers, F.; Menand, B.; Browning, K.S.; Robaglia, C. The Arabidopsis eukaryotic initiation factor ISO4E is dispensable for plant growth but required for susceptibility to potyviruses. Plant J. 2002, 32, 927–934. [Google Scholar] [CrossRef] [PubMed]
- Combe, J.P.; Petracek, M.E.; van Eldik, G.; Meulewaeter, F.; Twell, D. Translation initiation factors eIF4E and eIFiso4E are required for polysome formation and regulate plant growth in tobacco. Plant Mol. Biol. 2005, 57, 749–760. [Google Scholar] [CrossRef] [PubMed]
- Mayberry, L.K.; Allen, M.L.; Nitka, K.R.; Campbell, L.; Murphy, P.A.; Browning, K.S. Plant cap-binding complexes eukaryotic initiation factors eIF4F and eIFISO4F: molecular specificity of subunit binding. J. Biol. Chem. 2011, 286, 42566–42574. [Google Scholar] [CrossRef] [PubMed]
- Mayberry, L.K.; Allen, M.L.; Dennis, M.D.; Browning, K.S. Evidence for Variation in the Optimal Translation Initiation Complex: Plant eIF4B, eIF4F, and eIFISO4F Differentially Promote Translation of mRNAs. Plant Physiol. 2009, 150, 1844–1854. [Google Scholar] [CrossRef] [PubMed]
- Andersen, G.R.; Nissen, P.; Nyborg, J. Elongation factors in protein biosynthesis. Trends Biochem. Sci. 2003, 28, 434–441. [Google Scholar] [CrossRef]
- Goodfellow, I.G.; Roberts, L.O. Eukaryotic initiation factor 4E. Int. J. Biochem. Cell. Biol. 2008, 40, 2675–2680. [Google Scholar] [CrossRef] [PubMed]
- Culjkovic, B.; Topisirovic, I.; Borden, K.L. Controlling gene expression through RNA regulons: the role of the eukaryotic translation initiation factor eIF4E. Cell. Cycle 2007, 6, 65–69. [Google Scholar] [CrossRef] [PubMed]
- Bokros, C.L.; Hugdahl, J.D.; Kim, H.H.; Hanesworth, V.R.; van Heerden, A.; Browning, K.S.; Morejohn, L.C. Function of the p86 subunit of eukaryotic initiation factor ISO4F as a microtubule-associated protein in plant cells. Proc. Natl. Acad. Sci. USA 1995, 92, 7120–7124. [Google Scholar] [CrossRef] [PubMed]
- Hugdahl, J.D.; Bokros, C.L.; Morejohn, L.C. End-to-end annealing of plant microtubules by the p86 subunit of eukaryotic initiation factor-ISO4F. Plant Cell. 1995, 7, 2129–2138. [Google Scholar] [CrossRef] [PubMed]
- Mateyak, M.K.; Kinzy, T.G. eEF1A: solid thinking outside the ribosome. The Journal of biological chemistry 2010, 285, 21209–21213. [Google Scholar] [CrossRef] [PubMed]
- Au, H.H.; Jan, E. Novel viral translation strategies. Wiley Interdiscip. Rev. RNA 2014, 5, 779–801. [Google Scholar] [CrossRef] [PubMed]
- Firth, A.E.; Brierley, I. Non-canonical translation in RNA viruses. J. Gen. Virol. 2012, 93, 1385–1409. [Google Scholar] [CrossRef] [PubMed]
- Thompson, S.R. Tricks an IRES uses to enslave ribosomes. Trends Microbiol. 2012, 20, 558–66. [Google Scholar] [CrossRef] [PubMed]
- Miller, W.A.; Jackson, J.; Feng, Y. Cis- and trans-regulation of luteovirus gene expression by the 3′ end of the viral genome. Virus Res. 2015. [Google Scholar] [CrossRef] [PubMed]
- Simon, A.E. 3′UTRs of carmoviruses. Virus Res. 2015. [Google Scholar] [CrossRef] [PubMed]
- Chujo, T.; Ishibashi, K.; Miyashita, S.; Ishikawa, M. Functions of the 5′- and 3′-untranslated regions of tobamovirus RNA. Virus Res. 2015. [Google Scholar] [CrossRef] [PubMed]
- Truniger, V.; Nieto, C.; Gonzalez-Ibeas, D.; Aranda, M. Mechanism of plant eIF4E-mediated resistance against a Carmovirus (Tombusviridae): Cap-independent translation of a viral RNA controlled in cis by an (a)virulence determinant. Plant J. 2008, 56, 716–727. [Google Scholar] [CrossRef] [PubMed]
- Jiang, J.; Laliberte, J.F. The genome-linked protein VPg of plant viruses-A protein with many partners. Curr. Opin. Virol. 2011, 1, 347–354. [Google Scholar] [CrossRef] [PubMed]
- Rantalainen, K.I.; Eskelin, K.; Tompa, P.; Makinen, K. Structural flexibility allows the functional diversity of potyvirus genome-linked protein VPg. J. Virol. 2011, 85, 2449–2457. [Google Scholar] [CrossRef] [PubMed]
- Hebrard, E.; Bessin, Y.; Michon, T.; Longhi, S.; Uversky, V.N.; Delalande, F.; Van Dorsselaer, A.; Romero, P.; Walter, J.; Declerck, N.; Fargette, D. Intrinsic disorder in Viral Proteins Genome-Linked: Experimental and predictive analyses. Virology J. 2009, 6. [Google Scholar] [CrossRef] [PubMed]
- Grzela, R.; Szolajska, E.; Ebel, C.; Madern, D.; Favier, A.; Wojtal, I.; Zagorski, W.; Chroboczek, J. Virulence factor of potato virus Y, genome-attached terminal protein VPg, is a highly disordered protein. J. Biol. Chem. 2008, 283, 213–221. [Google Scholar] [CrossRef] [PubMed]
- Rantalainen, K.I.; Uversky, V.N.; Permi, P.; Kalkkinen, N.; Dunker, A.K.; Makinen, K. Potato virus A genome-linked protein VPg is an intrinsically disordered molten globule-like protein with a hydrophobic core. Virology 2008, 377, 280–288. [Google Scholar] [CrossRef] [PubMed]
- Wittmann, S.; Chatel, H.; Fortin, M.G.; Laliberte, J.F. Interaction of the viral protein genome linked of turnip mosaic potyvirus with the translational eukaryotic initiation factor ISO4E of Arabidopsis thaliana using the yeast two-hybrid system. Virology 1997, 234, 84–92. [Google Scholar] [CrossRef] [PubMed]
- Leonard, S.; Plante, D.; Wittmann, S.; Daigneault, N.; Fortin, M.G.; Laliberte, J.F. Complex formation between potyvirus VPg and translation eukaryotic initiation factor 4E correlates with virus infectivity. J. Virol. 2000, 74, 7730–7737. [Google Scholar] [CrossRef] [PubMed]
- Gallie, D.R. Cap-independent translation conferred by the 5′ leader of tobacco etch virus is eukaryotic initiation factor 4G dependent. J. Virol. 2001, 75, 12141–12152. [Google Scholar] [CrossRef] [PubMed]
- Khan, M.A.; Miyoshi, H.; Gallie, D.R.; Goss, D.J. Potyvirus genome-linked protein, VPg, directly affects wheat germ in vitro translation: interactions with translation initiation factors eIF4F and eIFiso4F. J. Biol. Chem. 2008, 283, 1340–1349. [Google Scholar] [CrossRef] [PubMed]
- Plante, D.; Viel, C.; Leonard, S.; Tampo, H.; Laliberte, J.F.; Fortin, M.G. Turnip mosaic virus VPg does not disrupt the translation initiation complex but interfere with cap binding. Physiol. Molec. Plant Pathol. 2004, 64, 219–226. [Google Scholar] [CrossRef]
- Eskelin, K.; Hafren, A.; Rantalainen, K.I.; Makinen, K. Potyviral VPg enhances viral RNA Translation and inhibits reporter mRNA translation in planta. J. Virol. 2011, 85, 9210–9221. [Google Scholar] [CrossRef] [PubMed]
- Hafren, A.; Eskelin, K.; Makinen, K. Ribosomal protein P0 promotes Potato virus A infection and functions in viral translation together with VPg and eIFISO4E. J. Virol. 2013, 87, 4302–4312. [Google Scholar] [CrossRef] [PubMed]
- Nicaise, V.; Gallois, J.L.; Chafiai, F.; Allen, L.M.; Schurdi-Levraud, V.; Browning, K.S.; Candresse, T.; Caranta, C.; Le Gall, O.; German-Retana, S. Coordinated and selective recruitment of eIF4E and eIF4G factors for potyvirus infection in Arabidopsis thaliana. FEBS Lett. 2007, 581, 1041–1046. [Google Scholar] [CrossRef] [PubMed]
- Contreras-Paredes, C.A.; Silva-Rosales, L.; Daros, J.A.; Alejandri-Ramirez, N.D.; Dinkova, T.D. The absence of eukaryotic initiation factor eIFISO4E affects the systemic spread of a Tobacco etch virus isolate in Arabidopsis thaliana. Mol. Plant Microbe Interact. 2013, 26, 461–470. [Google Scholar] [CrossRef] [PubMed]
- Hebrard, E.; Poulicard, N.; Gerard, C.; Traore, O.; Wu, H.C.; Albar, L.; Fargette, D.; Bessin, Y.; Vignols, F. Direct interaction between the Rice yellow mottle virus (RYMV) VPg and the central domain of the rice eIFISO4G1 factor correlates with rice susceptibility and RYMV virulence. Mol. Plant Microbe Interact. 2010, 23, 1506–1513. [Google Scholar] [CrossRef] [PubMed]
- Leonard, S.; Chisholm, J.; Laliberte, J.F.; Sanfacon, H. Interaction in vitro between the proteinase of Tomato ringspot virus (genus Nepovirus) and the eukaryotic translation initiation factor iso4E from Arabidopsis thaliana. J. Gen. Virol. 2002, 83, 2085–2089. [Google Scholar] [PubMed]
- Noueiry, A.O.; Chen, J.; Ahlquist, P. A mutant allele of essential, general translation initiation factor DED1 selectively inhibits translation of a viral mRNA. Proc. Natl. Acad. Sci. USA 2000, 97, 12985–12990. [Google Scholar] [CrossRef] [PubMed]
- Kovalev, N.; Pogany, J.; Nagy, P.D. A Co-Opted DEAD-Box RNA helicase enhances tombusvirus plus-strand synthesis. PLoS Pathog. 2012, 8, e1002537. [Google Scholar] [CrossRef] [PubMed]
- Dreher, T.W. Role of tRNA-like structures in controlling plant virus replication. Virus Res. 2009, 139, 217–229. [Google Scholar] [CrossRef] [PubMed]
- Dreher, T.W.; Uhlenbeck, O.C.; Browning, K.S. Quantitative assessment of EF-1α.GTP binding to aminoacyl-tRNAs, aminoacyl-viral RNA, and tRNA shows close correspondence to the RNA binding properties of EF-Tu. J. Biol. Chem. 1999, 274, 666–672. [Google Scholar] [CrossRef] [PubMed]
- Colussi, T.M.; Costantino, D.A.; Hammond, J.A.; Ruehle, G.M.; Nix, J.C.; Kieft, J.S. The structural basis of transfer RNA mimicry and conformational plasticity by a viral RNA. Nature 2014, 511, 366–369. [Google Scholar] [CrossRef] [PubMed]
- Matsuda, D.; Dreher, T.W. The tRNA-like structure of Turnip yellow mosaic virus RNA is a 3′-translational enhancer. Virology 2004, 321, 36–46. [Google Scholar] [CrossRef] [PubMed]
- Ryabova, L.A.; Pooggin, M.M.; Hohn, T. Translation reinitiation and leaky scanning in plant viruses. Virus Res. 2006, 119, 52–62. [Google Scholar] [CrossRef] [PubMed]
- Bonneville, J.M.; Sanfacon, H.; Futterer, J.; Hohn, T. Posttranscriptional trans-activation in cauliflower mosaic virus. Cell. 1989, 59, 1135–1143. [Google Scholar] [CrossRef]
- Park, H.S.; Himmelbach, A.; Browning, K.S.; Hohn, T.; Ryabova, L.A. A plant viral "reinitiation" factor interacts with the host translational machinery. Cell. 2001, 106, 723–733. [Google Scholar] [CrossRef]
- Thiebeauld, O.; Schepetilnikov, M.; Park, H.S.; Geldreich, A.; Kobayashi, K.; Keller, M.; Hohn, T.; Ryabova, L.A. A new plant protein interacts with eIF3 and 60S to enhance virus-activated translation re-initiation. EMBO J. 2009, 28, 3171–3184. [Google Scholar] [CrossRef] [PubMed]
- Bureau, M.; Leh, V.; Haas, M.; Geldreich, A.; Ryabova, L.; Yot, P.; Keller, M. P6 protein of Cauliflower mosaic virus, a translation reinitiator, interacts with ribosomal protein L13 from Arabidopsis thaliana. J. Gen. Virol. 2004, 85, 3765–3775. [Google Scholar] [CrossRef] [PubMed]
- Schepetilnikov, M.; Kobayashi, K.; Geldreich, A.; Caranta, C.; Robaglia, C.; Keller, M.; Ryabova, L.A. Viral factor TAV recruits TOR/S6K1 signalling to activate reinitiation after long ORF translation. EMBO J. 2011, 30, 1343–1356. [Google Scholar] [CrossRef] [PubMed]
- Ouibrahim, L.; Giner Rubio, A.; Moretti, A.; Montane, M.H.; Menand, B.; Meyer, C.; Robaglia, C.; Caranta, C. Potyviruses differ in their requirement for TOR signaling. J. Gen. Virol. 2015, in press. [Google Scholar] [CrossRef] [PubMed]
- Cotton, S.; Grangeon, R.; Thivierge, K.; Mathieu, I.; Ide, C.; Wei, T.; Wang, A.; Laliberte, J.F. Turnip mosaic virus RNA replication complex vesicles are mobile, align with microfilaments, and are each derived from a single viral genome. J. Virol. 2009, 83, 10460–10471. [Google Scholar] [CrossRef] [PubMed]
- Kawamura-Nagaya, K.; Ishibashi, K.; Huang, Y.P.; Miyashita, S.; Ishikawa, M. Replication protein of tobacco mosaic virus cotranslationally binds the 5′ untranslated region of genomic RNA to enable viral replication. Proc. Natl. Acad. Sci. USA 2014, 111, E1620–E1628. [Google Scholar] [CrossRef] [PubMed]
- Chen, J.; Noueiry, A.; Ahlquist, P. An alternate pathway for recruiting template RNA to the brome mosaic virus RNA replication complex. J. Virol. 2003, 77, 2568–2577. [Google Scholar] [CrossRef] [PubMed]
- Quadt, R.; Kao, C.C.; Browning, K.S.; Hershberger, R.P.; Ahlquist, P. Characterization of a host protein associated with brome mosaic virus RNA-dependent RNA polymerase. Proc. Natl. Acad. Sci. USA 1993, 90, 1498–1502. [Google Scholar] [CrossRef] [PubMed]
- Osman, T.A.; Buck, K.W. The tobacco mosaic virus RNA polymerase complex contains a plant protein related to the RNA-binding subunit of yeast eIF-3. J. Virol. 1997, 71, 6075–6082. [Google Scholar] [PubMed]
- Thivierge, K.; Cotton, S.; Dufresne, P.J.; Mathieu, I.; Beauchemin, C.; Ide, C.; Fortin, M.G.; Laliberte, J.F. Eukaryotic elongation factor 1A interacts with Turnip mosaic virus RNA-dependent RNA polymerase and VPg-Pro in virus-induced vesicles. Virology 2008, 377, 216–225. [Google Scholar] [CrossRef] [PubMed]
- Yamaji, Y.; Kobayashi, T.; Hamada, K.; Sakurai, K.; Yoshii, A.; Suzuki, M.; Namba, S.; Hibi, T. In vivo interaction between Tobacco mosaic virus RNA-dependent RNA polymerase and host translation elongation factor 1A. Virology 2006, 347, 100–108. [Google Scholar] [CrossRef] [PubMed]
- Beauchemin, C.; Boutet, N.; Laliberte, J.F. Visualisation of the interaction between the precursors of the viral protein linked to the genome (VPg) of Turnip mosaic virus and the translation eukaryotic initiation factor iso 4E in planta. J. Virol. 2007, 81, 775–782. [Google Scholar] [CrossRef] [PubMed]
- Huang, T.S.; Wei, T.; Laliberte, J.F.; Wang, A. A host RNA helicase-like protein, AtRH8, interacts with the potyviral genome-linked protein, VPg, associates with the virus accumulation complex, and is essential for infection. Plant Physiol. 2010, 152, 255–266. [Google Scholar] [CrossRef] [PubMed]
- Beauchemin, C.; Laliberte, J.F. The poly(A) binding protein is internalized in virus-induced vesicles or redistributed to the nucleolus during Turnip mosaic virus infection. J. Virol. 2007, 81, 10905–10913. [Google Scholar] [CrossRef] [PubMed]
- Dufresne, P.J.; Ubalijoro, E.; Fortin, M.G.; Laliberte, J.F. Arabidopsis thaliana class II poly(A)-binding proteins are required for efficient multiplication of turnip mosaic virus. J. Gen. Virol. 2008, 89, 2339–2348. [Google Scholar] [CrossRef] [PubMed]
- Kovalev, N.; Barajas, D.; Nagy, P.D. Similar roles for yeast Dbp2 and Arabidopsis RH20 DEAD-box RNA helicases to Ded1 helicase in tombusvirus plus-strand synthesis. Virology 2012, 432, 470–484. [Google Scholar] [CrossRef] [PubMed]
- Kovalev, N.; Nagy, P.D. The expanding functions of cellular helicases: the tombusvirus RNA replication enhancer co-opts the plant eIF4AIII-like AtRH2 and the DDX5-like AtRH5 DEAD-box RNA helicases to promote viral asymmetric RNA replication. PLoS Pathog. 2014, 10, e1004051. [Google Scholar] [CrossRef] [PubMed]
- Nishikiori, M.; Dohi, K.; Mori, M.; Meshi, T.; Naito, S.; Ishikawa, M. Membrane-bound tomato mosaic virus replication proteins participate in RNA synthesis and are associated with host proteins in a pattern distinct from those that are not membrane bound. J. Virol. 2006, 80, 8459–8468. [Google Scholar] [CrossRef] [PubMed]
- Li, Z.; Pogany, J.; Panavas, T.; Xu, K.; Esposito, A.M.; Kinzy, T.G.; Nagy, P.D. Translation elongation factor 1A is a component of the tombusvirus replicase complex and affects the stability of the p33 replication co-factor. Virology 2009, 385, 245–260. [Google Scholar] [CrossRef] [PubMed]
- Matsuda, D.; Yoshinari, S.; Dreher, T.W. eEF1A binding to aminoacylated viral RNA represses minus strand synthesis by TYMV RNA-dependent RNA polymerase. Virology 2004, 321, 47–56. [Google Scholar] [CrossRef] [PubMed]
- Li, Z.; Pogany, J.; Tupman, S.; Esposito, A.M.; Kinzy, T.G.; Nagy, P.D. Translation elongation factor 1A facilitates the assembly of the tombusvirus replicase and stimulates minus-strand synthesis. PLOS Pathog. 2010, 6, e1001175. [Google Scholar] [CrossRef] [PubMed]
- Sasvari, Z.; Izotova, L.; Kinzy, T.G.; Nagy, P.D. Synergistic roles of eukaryotic translation elongation factors 1Bgamma and 1A in stimulation of tombusvirus minus-strand synthesis. PLoS Pathog. 2011, 7, e1002438. [Google Scholar] [CrossRef] [PubMed]
- Zeenko, V.V.; Ryabova, L.A.; Spirin, A.S.; Rothnie, H.M.; Hess, D.; Browning, K.S.; Hohn, T. Eukaryotic elongation factor 1A interacts with the upstream pseudoknot domain in the 3′ untranslated region of tobacco mosaic virus RNA. J. Virol. 2002, 76, 5678–5691. [Google Scholar] [CrossRef] [PubMed]
- Yamaji, Y.; Sakurai, K.; Hamada, K.; Komatsu, K.; Ozeki, J.; Yoshida, A.; Yoshii, A.; Shimizu, T.; Namba, S.; Hibi, T. Significance of eukaryotic translation elongation factor 1A in tobacco mosaic virus infection. Arch. Virol. 2010, 155, 263–268. [Google Scholar] [CrossRef] [PubMed]
- Hwang, J.; Oh, C.S.; Kang, B.C. Translation elongation factor 1B (eEF1B) is an essential host factor for Tobacco mosaic virus infection in plants. Virology 2013, 439, 105–114. [Google Scholar] [CrossRef] [PubMed]
- Gao, Z.; Johansen, E.; Eyers, S.; Thomas, C.L.; Noel Ellis, T.H.; Maule, A.J. The potyvirus recessive resistance gene, sbm1, identifies a novel role for translation initiation factor eIF4E in cell-to-cell trafficking. Plant J. 2004, 40, 376–385. [Google Scholar] [CrossRef] [PubMed]
- Arroyo, R.; Soto, M.J.; Martinez-Zapater, J.; Ponz, F. Impaired cell-to-cell movement of Potato virus Y in pepper plants carrying the ya (pr21) resistance gene. Mol. Plant Microbe Interact. 1996, 9, 314–318. [Google Scholar] [CrossRef]
- Csorba, T.; Kontra, L.; Burgyan, J. Viral silencing suppressors: Tools forged to fine-tune host-pathogen coexistence. Virology 2015, in press. [Google Scholar] [CrossRef] [PubMed]
- Pumplin, N.; Voinnet, O. RNA silencing suppression by plant pathogens: Defence, counter-defence and counter-counter-defence. Nat. Rev. Microbiol. 2013, 11, 745–760. [Google Scholar] [CrossRef] [PubMed]
- Lanet, E.; Delannoy, E.; Sormani, R.; Floris, M.; Brodersen, P.; Crete, P.; Voinnet, O.; Robaglia, C. Biochemical evidence for translational repression by Arabidopsis microRNAs. Plant Cell. 2009, 21, 1762–1768. [Google Scholar] [CrossRef] [PubMed]
- Brodersen, P.; Sakvarelidze-Achard, L.; Bruun-Rasmussen, M.; Dunoyer, P.; Yamamoto, Y.Y.; Sieburth, L.; Voinnet, O. Widespread translational inhibition by plant miRNAs and siRNAs. Science 2008, 320, 1185–1190. [Google Scholar] [CrossRef] [PubMed]
- Iwakawa, H.O.; Tomari, Y. Molecular insights into microRNA-mediated translational repression in plants. Mol. Cell. 2013, 52, 591–601. [Google Scholar] [CrossRef] [PubMed]
- Reis, R.S.; Hart-Smith, G.; Eamens, A.L.; Wilkins, M.R.; Waterhouse, P.M. Gene regulation by translational inhibition is determined by Dicer partnering proteins. Nat. Plants 2015, 1. [Google Scholar] [CrossRef]
- Yang, L.; Wu, G.; Poethig, R.S. Mutations in the GW-repeat protein SUO reveal a developmental function for microRNA-mediated translational repression in Arabidopsis. Proc. Natl. Acad. Sci. USA 2012, 109, 315–320. [Google Scholar] [CrossRef] [PubMed]
- Li, S.; Liu, L.; Zhuang, X.; Yu, Y.; Liu, X.; Cui, X.; Ji, L.; Pan, Z.; Cao, X.; Mo, B.; Zhang, F.; Raikhel, N.; Jiang, L.; Chen, X. MicroRNAs inhibit the translation of target mRNAs on the endoplasmic reticulum in Arabidopsis. Cell. 2013, 153, 562–574. [Google Scholar] [CrossRef] [PubMed]
- Ma, X.; Nicole, M.C.; Meteignier, L.V.; Hong, N.; Wang, G.; Moffett, P. Different roles for RNA silencing and RNA processing components in virus recovery and virus-induced gene silencing in plants. J. Exp. Bot. 2015, in press. [Google Scholar] [CrossRef] [PubMed]
- Bhattacharjee, S.; Zamora, A.; Azhar, M.T.; Sacco, M.A.; Lambert, L.H.; Moffett, P. Virus resistance induced by NB-LRR proteins involves Argonaute4-dependent translational control. Plant J. 2009, 58, 940–951. [Google Scholar] [CrossRef] [PubMed]
- Ghoshal, B.; Sanfacon, H. Temperature-dependent symptom recovery in Nicotiana benthamiana plants infected with tomato ringspot virus is associated with reduced translation of viral RNA2 and requires ARGONAUTE 1. Virology 2014, 456–457, 188–197. [Google Scholar] [CrossRef] [PubMed]
- Karran, R.A.; Sanfacon, H. Tomato ringspot virus Coat Protein Binds to ARGONAUTE 1 and Suppresses the Translation Repression of a Reporter Gene. Mol. Plant Microbe Interact. 2014, 27, 933–943. [Google Scholar] [CrossRef] [PubMed]
- Wilczynska, A.; Bushell, M. The complexity of miRNA-mediated repression. Cell. Death Differ. 2015, 22, 22–33. [Google Scholar] [CrossRef] [PubMed]
- Fukaya, T.; Tomari, Y. MicroRNAs mediate gene silencing via multiple different pathways in Drosophila. Mol. Cell. 2012, 48, 825–836. [Google Scholar] [CrossRef] [PubMed]
- Meijer, H.A.; Kong, Y.W.; Lu, W.T.; Wilczynska, A.; Spriggs, R.V.; Robinson, S.W.; Godfrey, J.D.; Willis, A.E.; Bushell, M. Translational repression and eIF4A2 activity are critical for microRNA-mediated gene regulation. Science 2013, 340, 82–85. [Google Scholar] [CrossRef] [PubMed]
- Gibbs, A.; Ohshima, K. Potyviruses and the digital revolution. Annu. Rev. Phytopathol. 2010, 48, 205–223. [Google Scholar] [CrossRef] [PubMed]
- Charron, C.; Nicolai, M.; Gallois, J.L.; Robaglia, C.; Moury, B.; Palloix, A.; Caranta, C. Natural variation and functional analyses provide evidence for co-evolution between plant eIF4E and potyviral VPg. Plant J. 2008, 54, 56–68. [Google Scholar] [CrossRef] [PubMed]
- Moury, B.; Charron, C.; Janzac, B.; Simon, V.; Gallois, J.L.; Palloix, A.; Caranta, C. Evolution of plant eukaryotic initiation factor 4E (eIF4E) and potyvirus genome-linked protein (VPg): A game of mirrors impacting resistance spectrum and durability. Infect. Genet. Evol. 2014, 27, 472–480. [Google Scholar] [CrossRef] [PubMed]
- Truniger, V.; Aranda, M.A. Recessive resistance to plant viruses. Adv. Vir. Res. 2009, 75, 119–159. [Google Scholar]
- Kang, B.C.; Yeam, I.; Frantz, J.D.; Murphy, J.F.; Jahn, M.M. The pvr1 locus in Capsicum encodes a translation initiation factor eIF4E that interacts with Tobacco etch virus VPg. Plant J. 2005, 42, 392–405. [Google Scholar] [CrossRef] [PubMed]
- Ashby, J.A.; Stevenson, C.E.; Jarvis, G.E.; Lawson, D.M.; Maule, A.J. Structure-based mutational analysis of eIF4E in relation to sbm1 resistance to pea seed-borne mosaic virus in pea. PLoS ONE 2011, 6, e15873. [Google Scholar] [CrossRef] [PubMed]
- German-Retana, S.; Walter, J.; Doublet, B.; Roudet-Tavert, G.; Nicaise, V.; Lecampion, C.; Houvenaghel, M.C.; Robaglia, C.; Michon, T.; Le Gall, O. Mutational analysis of plant cap-binding protein eIF4E reveals key amino acids involved in biochemical functions and potyvirus infection. J. Virol. 2008, 82, 7601–7612. [Google Scholar] [CrossRef] [PubMed]
- Ibiza, V.P.; Canizares, J.; Nuez, F. EcoTILLING in Capsicum species: Searching for new virus resistances. BMC Genomics 2010, 11, e631. [Google Scholar] [CrossRef] [PubMed]
- Nieto, C.; Piron, F.; Dalmais, M.; Marco, C.F.; Moriones, E.; Gomez-Guillamon, M.L.; Truniger, V.; Gomez, P.; Garcia-Mas, J.; Aranda, M.A.; Bendahmane, A. EcoTILLING for the identification of allelic variants of melon eIF4E, a factor that controls virus susceptibility. BMC Plant Biol. 2007, 7. [Google Scholar] [CrossRef] [PubMed]
- Konecna, E.; Safarova, D.; Navratil, M.; Hanacek, P.; Coyne, C.; Flavell, A.; Vishnyakova, M.; Ambrose, M.; Redden, R.; Smykal, P. Geographical gradient of the eIF4E alleles conferring resistance to potyviruses in pea (Pisum) germplasm. PLoS ONE 2014, 9, e90394. [Google Scholar] [CrossRef] [PubMed]
- Rubio, M.; Nicolai, M.; Caranta, C.; Palloix, A. Allele mining in the pepper gene pool provided new complementation effects between pvr2-eIF4E and pvr6-eIFISO4E alleles for resistance to pepper veinal mottle virus. J. Gen. Virol. 2009, 90, 2808–2814. [Google Scholar] [CrossRef] [PubMed]
- Piron, F.; Nicolai, M.; Minoia, S.; Piednoir, E.; Moretti, A.; Salgues, A.; Zamir, D.; Caranta, C.; Bendahmane, A. An induced mutation in tomato eIF4E leads to immunity to two potyviruses. PLoS ONE 2010, 5, e11313. [Google Scholar] [CrossRef] [PubMed]
- Rodriguez-Hernandez, A.M.; Gosalvez, B.; Sempere, R.N.; Burgos, L.; Aranda, M.A.; Truniger, V. Melon RNA interference (RNAi) lines silenced for Cm-eIF4E show broad virus resistance. Mol. Plant Pathol. 2012, 3, 755–763. [Google Scholar] [CrossRef] [PubMed]
- Mazier, M.; Flamain, F.; Nicolai, M.; Sarnette, V.; Caranta, C. Knock-down of both eIF4E1 and eIF4E2 genes confers broad-spectrum resistance against potyviruses in tomato. PLoS ONE 2011, 6, e29595. [Google Scholar] [CrossRef] [PubMed]
- Wang, X.; Kohalmi, S.E.; Svircev, A.; Wang, A.; Sanfacon, H.; Tian, L. Silencing of the host factor eIFISO4E gene confers plum pox virus resistance in plum. PLoS ONE 2013, 8, e50627. [Google Scholar] [CrossRef] [PubMed]
- Kang, B.C.; Yeam, I.; Li, H.; Perez, K.W.; Jahn, M.M. Ectopic expression of a recessive resistance gene generates dominant potyvirus resistance in plants. Plant Biotechnol. J. 2007, 4, 526–536. [Google Scholar] [CrossRef] [PubMed]
- Cavatorta, J.; Perez, K.W.; Gray, S.M.; Van Eck, J.; Yeam, I.; Jahn, M. Engineering virus resistance using a modified potato gene. Plant Biotechnol. J. 2011, 9, 1014–1021. [Google Scholar] [CrossRef] [PubMed]
- Duan, H.; Richael, C.; Rommens, C.M. Overexpression of the wild potato eIF4E-1 variant Eva1 elicits Potato virus Y resistance in plants silenced for native eIF4E-1. Transgenic Res. 2012, 21, 929–938. [Google Scholar] [CrossRef] [PubMed]
- Kim, J.; Kang, W.H.; Hwang, J.; Yang, H.B.; Dosun, K.; Oh, C.S.; Kang, B.C. Transgenic Brassica rapa plants over-expressing eIFISO4E variants show broad-spectrum Turnip mosaic virus (TuMV) resistance. Mol. Plant Pathol. 2014, 15, 615–626. [Google Scholar] [CrossRef] [PubMed]
- Schaad, M.C.; Anderberg, R.J.; Carrington, J.C. Strain-specific interaction of the tobacco etch virus NIa protein with the translation initiation factor eIF4E in the yeast two-hybrid system. Virology 2000, 273, 300–306. [Google Scholar] [CrossRef] [PubMed]
- Estevan, J.; Marena, A.; Callot, C.; Lacombe, S.; Moretti, A.; Caranta, C.; Gallois, J.L. Specific requirement for translation initiation factor 4E or its isoform drives plant host susceptibility to Tobacco etch virus. BMC Plant Biol. 2014, 14. [Google Scholar] [CrossRef] [PubMed]
- Sato, M.; Nakahara, K.; Yoshii, M.; Ishikawa, M.; Uyeda, I. Selective involvement of members of the eukaryotic initiation factor 4E family in the infection of Arabidopsis thaliana by potyviruses. FEBS Lett. 2005, 579, 1167–1171. [Google Scholar] [CrossRef] [PubMed]
- Hwang, J.; Li, J.; Liu, W.Y.; An, S.J.; Cho, H.; Her, N.H.; Yeam, I.; Kim, D.; Kang, B.C. Double mutations in eIF4E and eIFiso4E confer recessive resistance to Chilli veinal mottle virus in pepper. Mol. Cells 2009, 27, 329–336. [Google Scholar] [CrossRef] [PubMed]
- Ruffel, S.; Gallois, J.L.; Moury, B.; Robaglia, C.; Palloix, A.; Caranta, C. Simultaneous mutations in translation initiation factors eIF4E and eIFISO4E are required to prevent pepper veinal mottle virus infection of pepper. J. Gen. Virol. 2006, 87, 2089–2098. [Google Scholar] [CrossRef] [PubMed]
- Jenner, C.E.; Nellist, C.F.; Barker, G.C.; Walsh, J.A. Turnip mosaic virus (TuMV) is able to use alleles of both eIF4E and eIFISO4E from multiple loci of the diploid Brassica rapa. Mol. Plant Microbe Interact. 2010, 23, 1498–1505. [Google Scholar] [CrossRef] [PubMed]
- Ayme, V.; Petit-Pierre, J.; Souche, S.; Palloix, A.; Moury, B. Molecular dissection of the potato virus Y VPg virulence factor reveals complex adaptations to the pvr2 resistance allelic series in pepper. J. Gen. Virol. 2007, 88, 1594–1601. [Google Scholar] [CrossRef] [PubMed]
- Ayme, V.; Souche, S.; Caranta, C.; Jacquemond, M.; Chadoeuf, J.; Palloix, A.; Moury, B. Different mutations in the genome-linked protein VPg of potato virus Y confer virulence on the pvr2(3) resistance in pepper. Mol. Plant Microbe Interact. 2006, 19, 557–563. [Google Scholar] [CrossRef] [PubMed]
- Moury, B.; Morel, C.; Johansen, E.; Guilbaud, L.; Souche, S.; Ayme, V.; Caranta, C.; Palloix, A.; Jacquemond, M. Mutations in potato virus Y genome-linked protein determine virulence toward recessive resistances in Capsicum annuum and Lycopersicon hirsutum. Mol. Plant Microbe Interact. 2004, 17, 322–329. [Google Scholar] [CrossRef] [PubMed]
- Gallois, J.L.; Charron, C.; Sanchez, F.; Pagny, G.; Houvenaghel, M.C.; Moretti, A.; Ponz, F.; Revers, F.; Caranta, C.; German-Retana, S. Single amino acid changes in the turnip mosaic virus viral genome-linked protein (VPg) confer virulence towards Arabidopsis thaliana mutants knocked out for eukaryotic initiation factors eIFISO4E and eIFISO4G. J. Gen. Virol. 2010, 91, 288–293. [Google Scholar] [CrossRef] [PubMed]
- Abdul-Razzak, A.; Guiraud, T.; Peypelut, M.; Walter, J.; Houvenaghel, M.C.; Candresse, T.; Le Gall, O.; German-Retana, S. Involvement of the cylindrical inclusion (CI) protein in the overcoming of an eIF4E-mediated resistance against Lettuce mosaic potyvirus. Mol. Plant Pathol. 2009, 10, 109–113. [Google Scholar] [CrossRef] [PubMed]
- Sorel, M.; Svanella-Dumas, L.; Candresse, T.; Acelin, G.; Pitarch, A.; Houvenaghel, M.C.; German-Retana, S. Key mutations in the cylindrical inclusion involved in lettuce mosaic virus adaptation to eIF4E-mediated resistance in lettuce. Mol. Plant Microbe Interact. 2014, 27, 1014–1024. [Google Scholar] [CrossRef] [PubMed]
- Tavert-Roudet, G.; Abdul-Razzak, A.; Doublet, B.; Walter, J.; Delaunay, T.; German-Retana, S.; Michon, T.; Le Gall, O.; Candresse, T. The C terminus of lettuce mosaic potyvirus cylindrical inclusion helicase interacts with the viral VPg and with lettuce translation eukaryotic initiation factor 4E. J. Gen. Virol. 2012, 93, 184–93. [Google Scholar] [CrossRef] [PubMed]
- Sorel, M.; Garcia, J.A.; German-Retana, S. The Potyviridae cylindrical inclusion helicase: a key multipartner and multifunctional protein. Mol. Plant Microbe Interact. 2014, 27, 215–26. [Google Scholar] [CrossRef] [PubMed]
- Nakahara, K.S.; Shimada, R.; Choi, S.H.; Yamamoto, H.; Shao, J.; Uyeda, I. Involvement of the P1 cistron in overcoming eIF4E-mediated recessive resistance against Clover yellow vein virus in pea. Mol. Plant Microbe Interact. 2010, 23, 1460–1469. [Google Scholar] [CrossRef] [PubMed]
- Moury, B.; Janzac, B.; Ruellan, Y.; Simon, V.; Ben Khalifa, M.; Fakhfakh, H.; Fabre, F.; Palloix, A. Interaction patterns between potato virus Y and eIF4E-mediated recessive resistance in the Solanaceae. J. Virol. 2014, 88, 9799–9807. [Google Scholar] [CrossRef] [PubMed]
- Ruffel, S.; Caranta, C.; Palloix, A.; Lefebvre, V.; Caboche, M.; Bendahmane, A. Structural analysis of the eukaryotic initiation factor 4E gene controlling potyvirus resistance in pepper: Exploitation of a BAC library. Gene 2004, 338, 209–216. [Google Scholar] [CrossRef] [PubMed]
- Ruffel, S.; Dussault, M.H.; Palloix, A.; Moury, B.; Bendahmane, A.; Robaglia, C.; Caranta, C. A natural recessive resistance gene against potato virus Y in pepper corresponds to the eukaryotic initiation factor 4E (eIF4E). Plant J. 2002, 32, 1067–1075. [Google Scholar] [CrossRef] [PubMed]
- Nicaise, V.; German-Retana, S.; Sanjuan, R.; Dubrana, M.P.; Mazier, M.; Maisonneuve, B.; Candresse, T.; Caranta, C.; LeGall, O. The eukaryotic translation initiation factor 4E controls lettuce susceptibility to the Potyvirus Lettuce mosaic virus. Plant Physiol. 2003, 132, 1272–1282. [Google Scholar] [CrossRef] [PubMed]
- Andrade, M.; Abe, Y.; Nakahara, k. S.; Uyeda, I. The cyv-2 resistance to clover yellow vein virus in pea is controlled by the eukaryotic initiation factor 4E. J. Gen. Plant Pathol. 2009, 75, 241–249. [Google Scholar] [CrossRef]
- Bruun-Rasmussen, M.; Moller, I.S.; Tulinius, G.; Hansen, J.K.; Lund, O.S.; Johansen, I.E. The same allele of translation initiation factor 4E mediates resistance against two Potyvirus spp. in Pisum sativum. Mol. Plant Microbe Interact. 2007, 20, 1075–1082. [Google Scholar] [CrossRef] [PubMed]
- Naderpour, M.; Lund, O.S.; Larsen, R.; Johansen, E. Potyviral resistance derived from cultivars of Phaseolus vulgaris carrying bc-3 is associated with the homozygotic presence of a mutated eIF4E allele. Mol. Plant Pathol. 2010, 11, 255–263. [Google Scholar] [CrossRef] [PubMed]
- Hart, J.P.; Griffiths, P.D. A series of eIF4E alleles at the Bc-3 locus are associated with recessive resistance to Clover yellow vein virus in common bean. Theor. Appl. Genet. 2013, 126, 2849–2863. [Google Scholar] [CrossRef] [PubMed]
- Ruffel, S.; Gallois, J.L.; Lesage, M.L.; Caranta, C. The recessive potyvirus resistance gene pot-1 is the tomato orthologue of the pepper pvr2-eIF4E gene. Mol. Genet. Genomics 2005, 274, 346–353. [Google Scholar] [CrossRef] [PubMed]
- Hofinger, B.J.; Russell, J.R.; Bass, C.G.; Baldwin, T.; dos Reis, M.; Hedley, P.E.; Li, Y.; Macaulay, M.; Waugh, R.; Hammond-Kosack, K.E.; Kanyuka, K. An exceptionally high nucleotide and haplotype diversity and a signature of positive selection for the eIF4E resistance gene in barley are revealed by allele mining and phylogenetic analyses of natural populations. Mol. Ecol. 2011, 20, 3653–3668. [Google Scholar] [CrossRef] [PubMed]
- Kanyuka, K.; Druka, A.; Caldwell, D.G.; Tymon, A.; McCallum, N.; Waugh, R.; Adams, M.J. Evidence that the recessive bymovirus resistance locus rym4 in barley corresponds to the eukaryotic translation initiation factor 4E gene. Mol. Plant Pathol. 2005, 6, 449–458. [Google Scholar] [CrossRef] [PubMed]
- Kanyuka, K.; McGrann, G.; Alhudaib, K.; Hariri, D.; Adams, M.J. Biological and sequence analysis of a novel European isolate of Barley mild mosaic virus that overcomes the barley rym5 resistance gene. Arch. Virol. 2004, 149, 1469–1480. [Google Scholar] [CrossRef] [PubMed]
- Kuhne, T.; Shi, N.; Proeseler, G.; Adams, M.J.; Kanyuka, K. The ability of a bymovirus to overcome the rym4-mediated resistance in barley correlates with a codon change in the VPg coding region on RNA1. J. Gen. Virol. 2003, 84, 2853–2859. [Google Scholar] [CrossRef] [PubMed]
- Stein, N.; Perovic, D.; Kumlehn, J.; Pellio, B.; Stracke, S.; Streng, S.; Ordon, F.; Graner, A. The eukaryotic translation initiation factor 4E confers multiallelic recessive Bymovirus resistance in Hordeum vulgare (L.). Plant J. 2005, 42, 912–922. [Google Scholar] [CrossRef] [PubMed]
- Perovic, D.; Kramer, I.; Habekuss, A.; Perner, K.; Pickering, R.; Proeseler, G.; Kanyuka, K.; Ordon, F. Genetic analyses of BaMMV/BaYMV resistance in barley accession HOR4224 result in the identification of an allele of the translation initiation factor 4e (Hv-eIF4E) exclusively effective against Barley mild mosaic virus (BaMMV). Theor. Appl. Genet. 2014, 127, 1061–71. [Google Scholar] [CrossRef] [PubMed]
- Ling, K.S.; Harris, K.R.; Meyer, J.D.; Levi, A.; Guner, N.; Wehner, T.C.; Bendahmane, A.; Havey, M.J. Non-synonymous single nucleotide polymorphisms in the watermelon eIF4E gene are closely associated with resistance to zucchini yellow mosaic virus. Theor. Appl. Genet. 2009, 120, 191–200. [Google Scholar] [CrossRef] [PubMed]
- Marandel, G.; Salava, J.; Abbott, A.; Candresse, T.; Decroocq, V. Quantitative trait loci meta-analysis of Plum pox virus resistance in apricot (Prunus armeniaca L.): new insights on the organization and the identification of genomic resistance factors. Mol. Plant Pathol. 2009, 10, 347–360. [Google Scholar] [CrossRef] [PubMed]
- Rusholme, R.L.; Higgins, E.E.; Walsh, J.A.; Lydiate, D.J. Genetic control of broad-spectrum resistance to turnip mosaic virus in Brassica rapa (Chinese cabbage). J. Gen. Virol. 2007, 88, 3177–3186. [Google Scholar] [CrossRef] [PubMed]
- Nellist, C.F.; Qian, W.; Jenner, C.E.; Moore, J.D.; Zhang, S.; Wang, X.; Briggs, W.H.; Barker, G.C.; Sun, R.; Walsh, J.A. Multiple copies of eukaryotic translation initiation factors in Brassica rapa facilitate redundancy, enabling diversification through variation in splicing and broad-spectrum virus resistance. Plant J. 2014, 77, 261–268. [Google Scholar] [CrossRef] [PubMed]
- Walsh, J.A.; Rusholme, R.L.; Hughes, S.L.; Jenner, C.E.; Bambridge, J.M.; Lydiate, D.J.; Green, S.K. Different classes of resistance to Turnip mosaic virus in Brassica rapa. Eur. J. Plant Pathol. 2002, 108, 15–20. [Google Scholar] [CrossRef]
- Lellis, A.D.; Kasschau, K.D.; Whitham, S.A.; Carrington, J.C. Loss-of-susceptibility mutants of Arabidopsis thaliana reveal an essential role for eIFISO4E during potyvirus infection. Curr. Biol. 2002, 12, 1046–1051. [Google Scholar] [CrossRef]
- Albar, L.; Bangratz-Reyser, M.; Hebrard, E.; Ndjiondjop, M.N.; Jones, M.; Ghesquiere, A. Mutations in the eIFISO4G translation initiation factor confer high resistance of rice to Rice yellow mottle virus. Plant J. 2006, 47, 417–426. [Google Scholar] [CrossRef] [PubMed]
- Thiemele, D.; Boisnard, A.; Ndjiondjop, M.N.; Cheron, S.; Sere, Y.; Ake, S.; Ghesquiere, A.; Albar, L. Identification of a second major resistance gene to Rice yellow mottle virus, RYMV2, in the African cultivated rice species, O. glaberrima. Theor. Appl. Genet. 2010, 121, 169–179. [Google Scholar] [CrossRef] [PubMed]
- Poulicard, N.; Pinel-Galzi, A.; Traore, O.; Vignols, F.; Ghesquiere, A.; Konate, G.; Hebrard, E.; Fargette, D. Historical contingencies modulate the adaptability of Rice yellow mottle virus. PLoS Pathog. 2012, 8, e1002482. [Google Scholar] [CrossRef] [PubMed]
- Traore, O.; Pinel-Galzi, A.; Issaka, S.; Poulicard, N.; Aribi, J.; Ake, S.; Ghesquiere, A.; Sere, Y.; Konate, G.; Hebrard, E.; Fargette, D. The adaptation of Rice yellow mottle virus to the eIFISO4G-mediated rice resistance. Virology 2010, 408, 103–108. [Google Scholar] [CrossRef] [PubMed]
- Hebrard, E.; Pinel-Galzi, A.; Fargette, D. Virulence domain of the RYMV genome-linked viral protein VPg towards rice rymv1–2-mediated resistance. Arch. Virol. 2008, 153, 1161–1164. [Google Scholar] [CrossRef] [PubMed]
- Hebrard, E.; Pinel-Galzi, A.; Bersoult, A.; Sire, C.; Fargette, D. Emergence of a resistance-breaking isolate of Rice yellow mottle virus during serial inoculations is due to a single substitution in the genome-linked viral protein VPg. J. Gen. Virol. 2006, 87, 1369–1373. [Google Scholar] [CrossRef] [PubMed]
- Poulicard, N.; Pinel-Galzi, A.; Hebrard, E.; Fargette, D. Why Rice yellow mottle virus, a rapidly evolving RNA plant virus, is not efficient at breaking rymv1–2 resistance. Mol. Plant Pathol. 2010, 11, 145–154. [Google Scholar] [CrossRef] [PubMed]
- Poulicard, N.; Pinel-Galzi, A.; Fargette, D.; Hebrard, E. Alternative mutational pathways, outside the VPg, of rice yellow mottle virus to overcome eIFISO4G-mediated rice resistance under strong genetic constraints. J. Gen. Virol. 2014, 95, 219–224. [Google Scholar] [CrossRef] [PubMed]
- Nieto, C.; Morales, M.; Orjeda, G.; Clepet, C.; Monfort, A.; Sturbois, B.; Puigdomenech, P.; Pitrat, M.; Caboche, M.; Dogimont, C.; Garcia-Mas, J.; Aranda, M.A.; Bendahmane, A. An eIF4E allele confers resistance to an uncapped and non-polyadenylated RNA virus in melon. Plant J. 2006, 48, 452–462. [Google Scholar] [CrossRef] [PubMed]
- Diaz, J.A.; Nieto, C.; Moriones, E.; Truniger, V.; Aranda, M.A. Molecular characterization of a Melon necrotic spot virus strain that overcomes the resistance in melon and nonhost plants. Mol. Plant Microbe Interact. 2004, 17, 668–675. [Google Scholar] [CrossRef] [PubMed]
- Miras, M.; Sempere, R.N.; Kraft, J.J.; Miller, W.A.; Aranda, M.A.; Truniger, V. Interfamilial recombination between viruses led to acquisition of a novel translation-enhancing RNA element that allows resistance breaking. New Phytol. 2014, 202, 233–246. [Google Scholar] [CrossRef] [PubMed]
- Nicholson, B.L.; Zaslaver, O.; Mayberry, L.K.; Browning, K.S.; White, K.A. Tombusvirus Y-shaped translational enhancer forms a complex with eIF4F and can be functionally replaced by heterologous translational enhancers. J. Virol. 2013, 87, 1872–1883. [Google Scholar] [CrossRef] [PubMed]
- Pinel-Galzi, A.; Traore, O.; Sere, Y.; Hebrard, E.; Fargette, D. The biogeography of viral emergence: Rice yellow mottle virus as a case study. Curr. Opin. Virol. 2015, 10, 7–13. [Google Scholar] [CrossRef] [PubMed]
- Pinel-Galzi, A.; Rakotomalala, M.; Sangu, E.; Sorho, F.; Kanyeka, Z.; Traore, O.; Sereme, D.; Poulicard, N.; Rabenantoandro, Y.; Sere, Y.; Konate, G.; Ghesquiere, A.; Hebrard, E.; Fargette, D. Theme and variations in the evolutionary pathways to virulence of an rna plant virus species. PLoS Pathog. 2007, 3, e180. [Google Scholar] [CrossRef] [PubMed]
- Lee, J.H.; Muhsin, M.; Atienza, G.A.; Kwak, D.Y.; Kim, S.M.; De Leon, T.B.; Angeles, E.R.; Coloquio, E.; Kondoh, H.; Satoh, K.; Cabunagan, R.C.; Cabauatan, P.Q.; Kikuchi, S.; Leung, H.; Choi, I.R. Single nucleotide polymorphisms in a gene for translation initiation factor (eIF4G) of rice (Oryza sativa) associated with resistance to Rice tungro spherical virus. Mol. Plant Microbe Interact. 2010, 23, 29–38. [Google Scholar] [CrossRef] [PubMed]
- Yoshii, M.; Nishikiori, M.; Tomita, K.; Yoshioka, N.; Kozuka, R.; Naito, S.; Ishikawa, M. The Arabidopsis cucumovirus multiplication 1 and 2 loci encode translation initiation factors 4E and 4G. J. Virol. 2004, 78, 6102–6111. [Google Scholar] [CrossRef] [PubMed]
- Belhaj, K.; Chaparro-Garcia, A.; Kamoun, S.; Patron, N.J.; Nekrasov, V. Editing plant genomes with CRISPR/Cas9. Curr. Opin. Biotechnol. 2014, 32C, 76–84. [Google Scholar] [CrossRef] [PubMed]
© 2015 by Her Majesty the Queen in Right of Canada, as represented by the Minister of Agriculture and Agri-Food Canada; licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Sanfaçon, H. Plant Translation Factors and Virus Resistance. Viruses 2015, 7, 3392-3419. https://doi.org/10.3390/v7072778
Sanfaçon H. Plant Translation Factors and Virus Resistance. Viruses. 2015; 7(7):3392-3419. https://doi.org/10.3390/v7072778
Chicago/Turabian StyleSanfaçon, Hélène. 2015. "Plant Translation Factors and Virus Resistance" Viruses 7, no. 7: 3392-3419. https://doi.org/10.3390/v7072778
APA StyleSanfaçon, H. (2015). Plant Translation Factors and Virus Resistance. Viruses, 7(7), 3392-3419. https://doi.org/10.3390/v7072778





