The Emerging Role of miRNAs in HTLV-1 Infection and ATLL Pathogenesis
Abstract
:1. Introduction
2. MiRNA Biogenesis
3. MiRNA Profile in HTLV-1-Transformed Cell Lines and ATLL Patients
4. HTLV-1 Interferes with Cellular miRNA Machinery
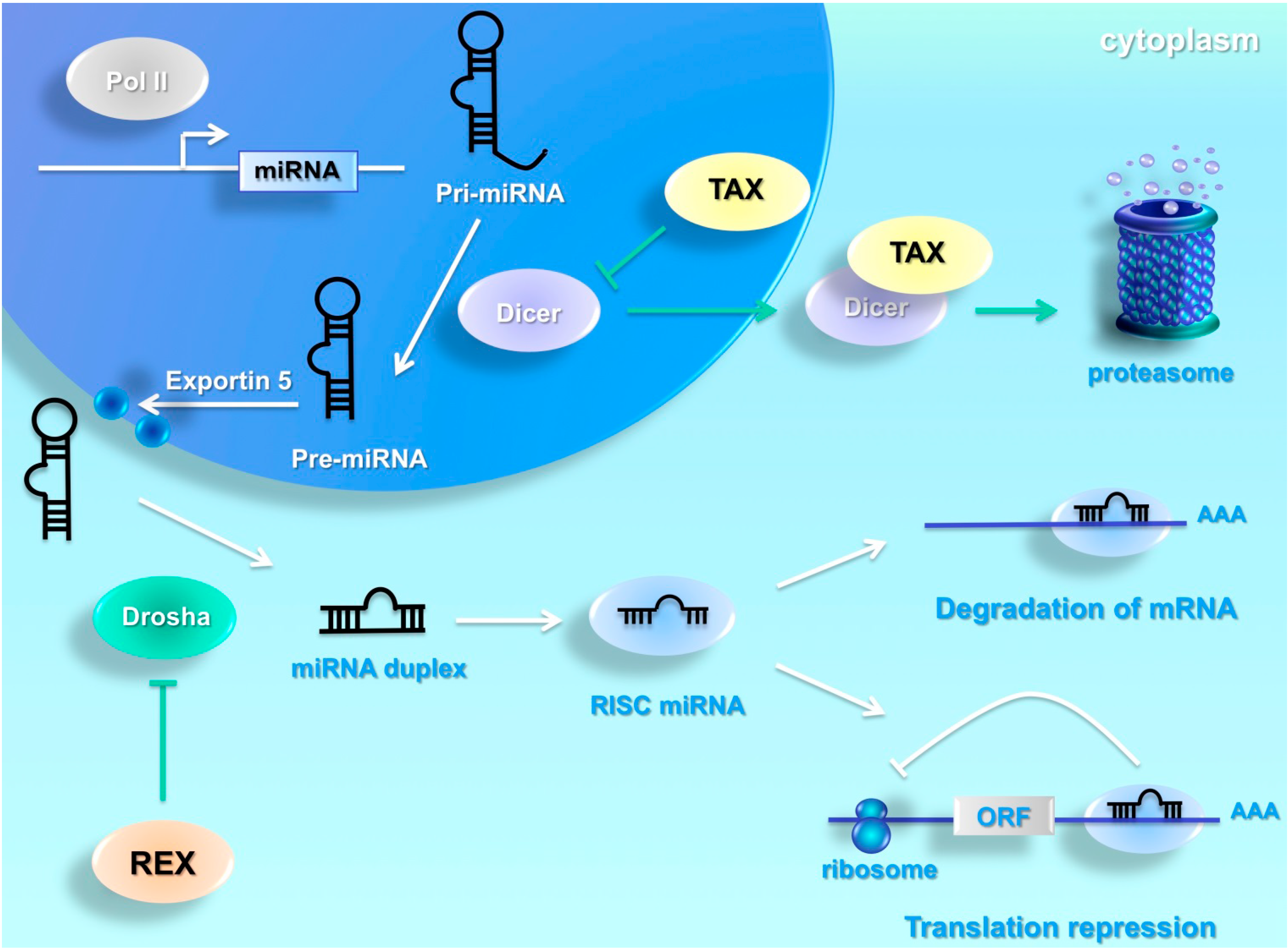
5. MiRNAs Target the HTLV-1 Genome
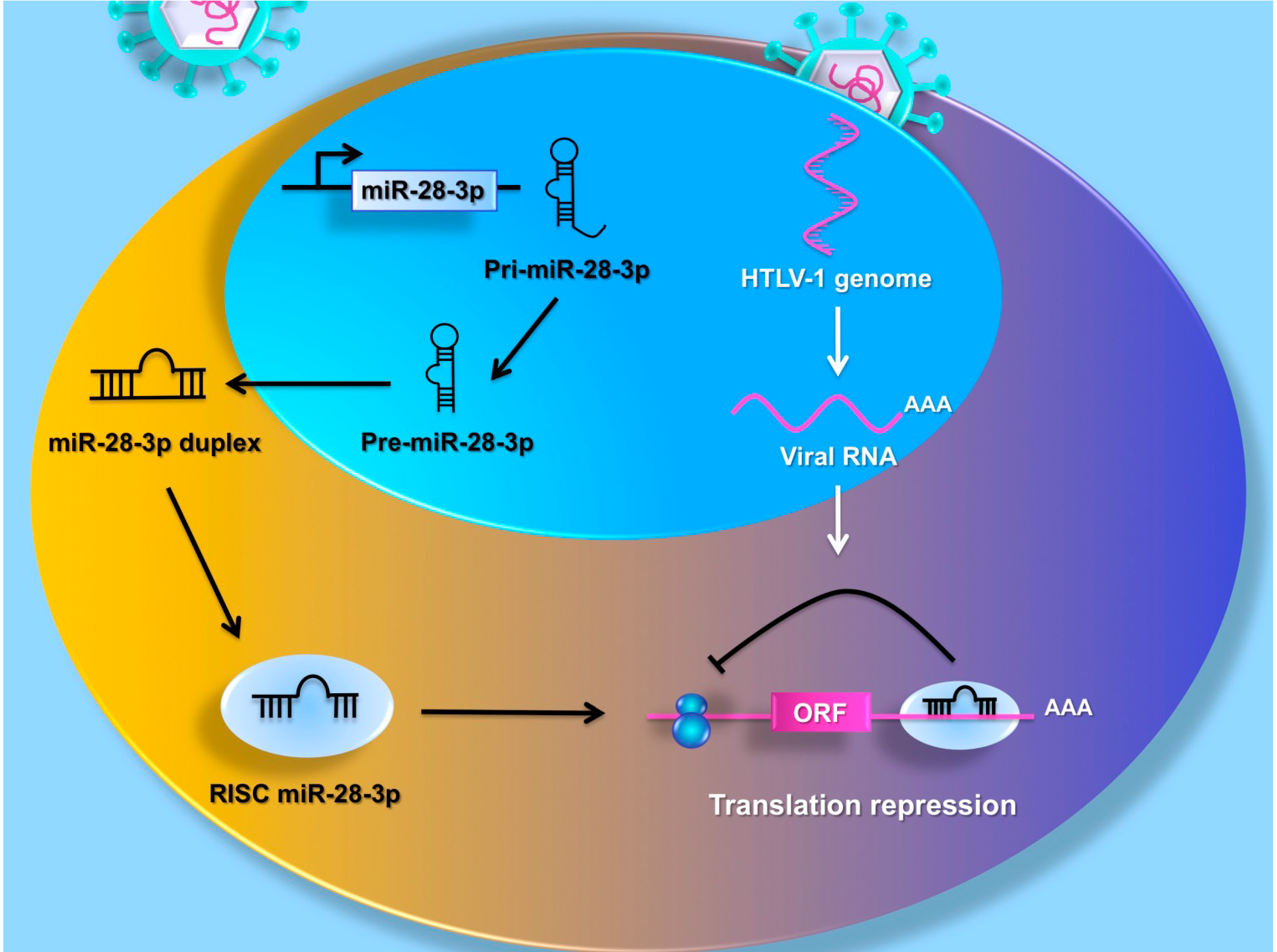
6. MiRNAs Promote Cell Proliferation
6.1. MiR-146a
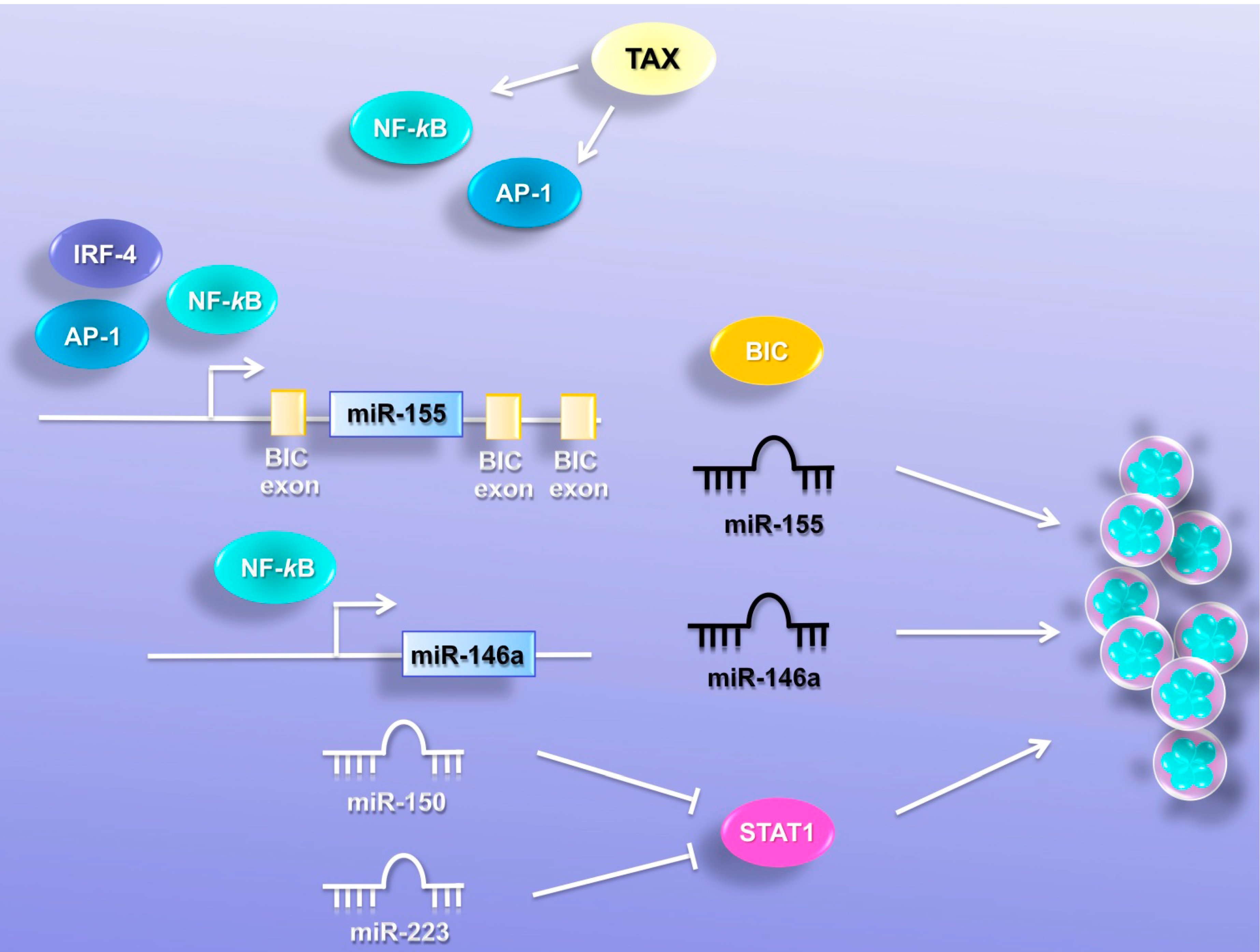
6.2. MiR-155
6.3. MiR-150 and MiR-223
7. MiRNAs Induce Resistance to Apoptosis
7.1. MiR-31
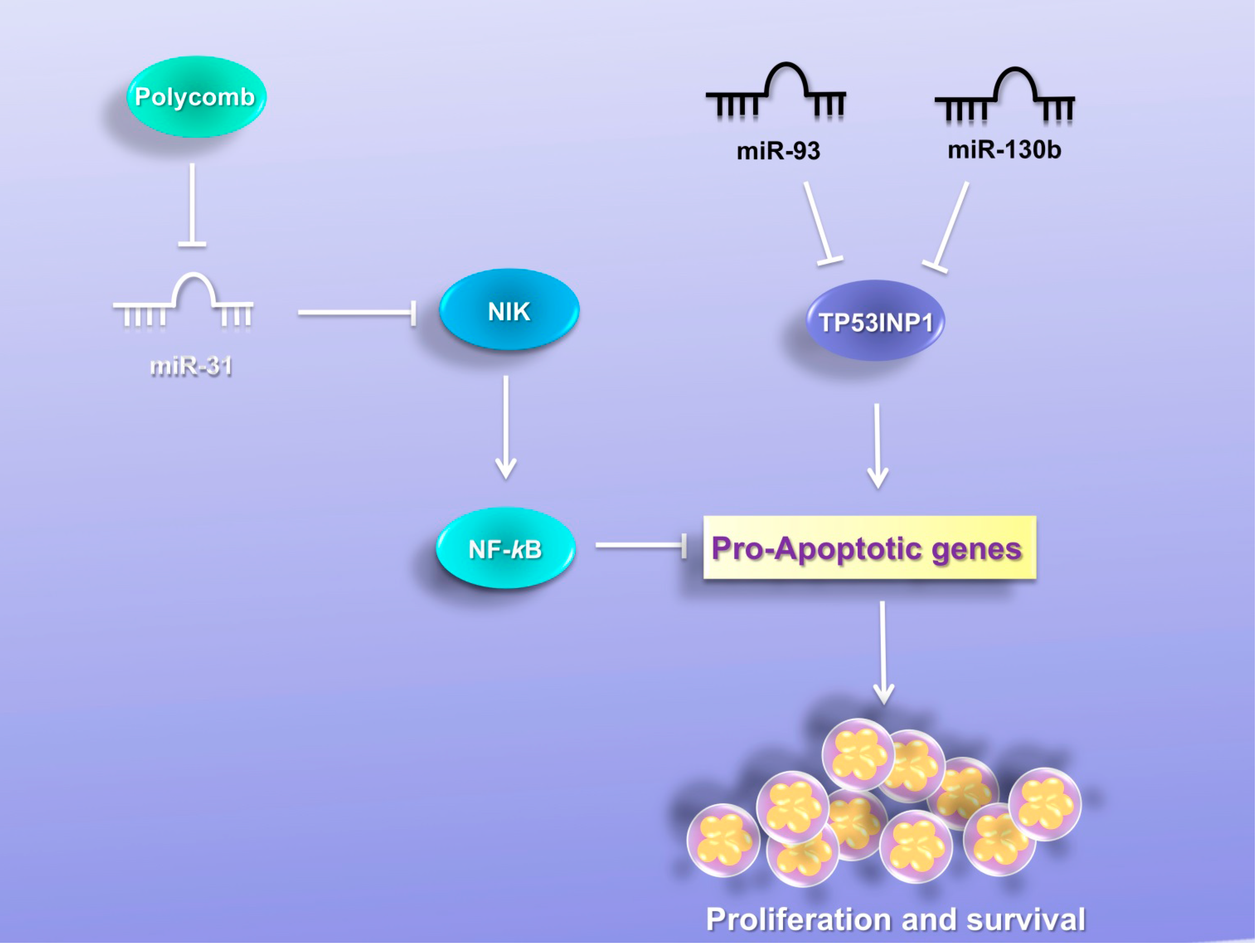
7.2. MiR-130b and MiR-93
8. MiRNAs Promote Chromatin Remodeling
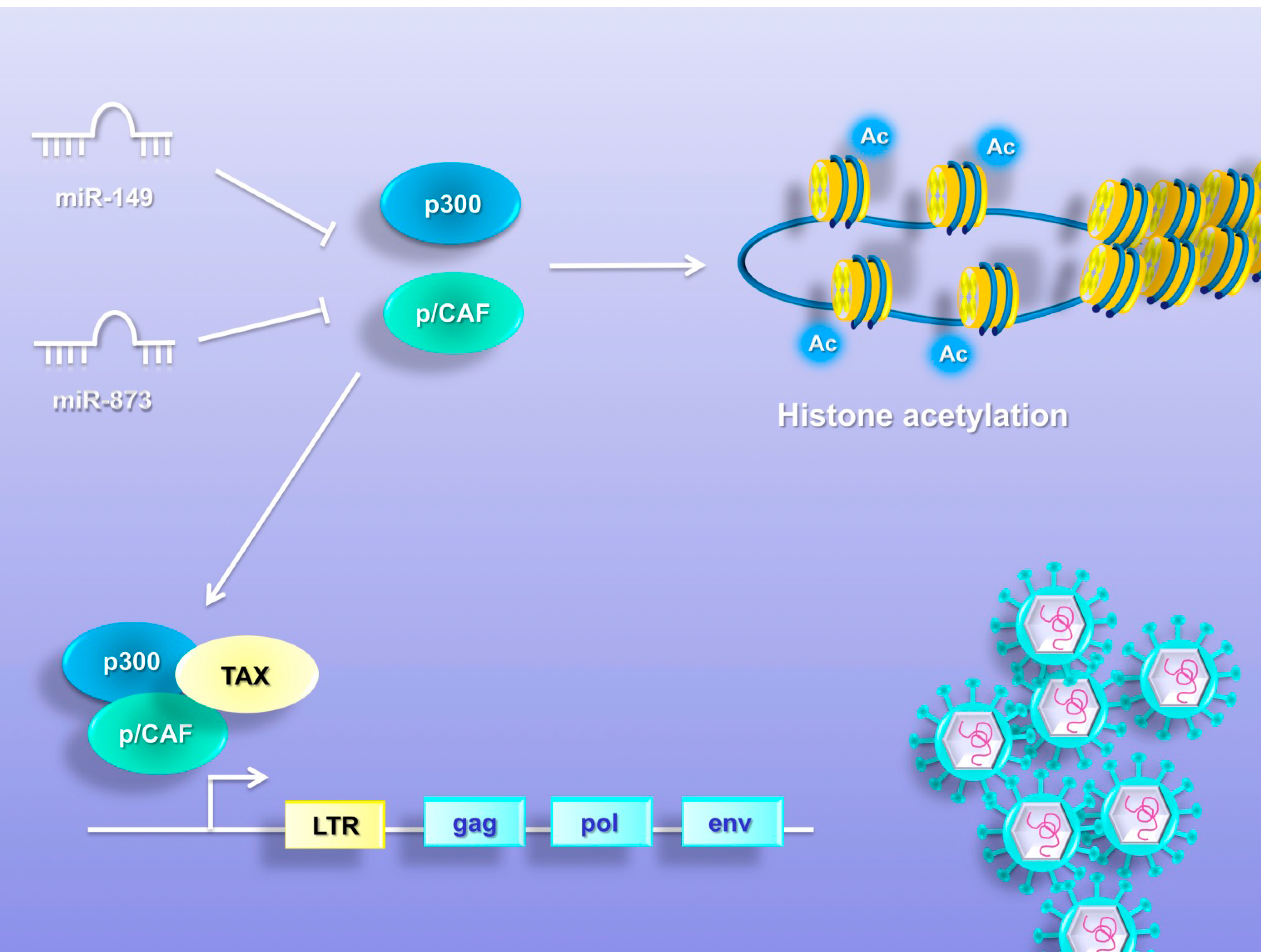
9. MiRNAs Induce Genetic Instability
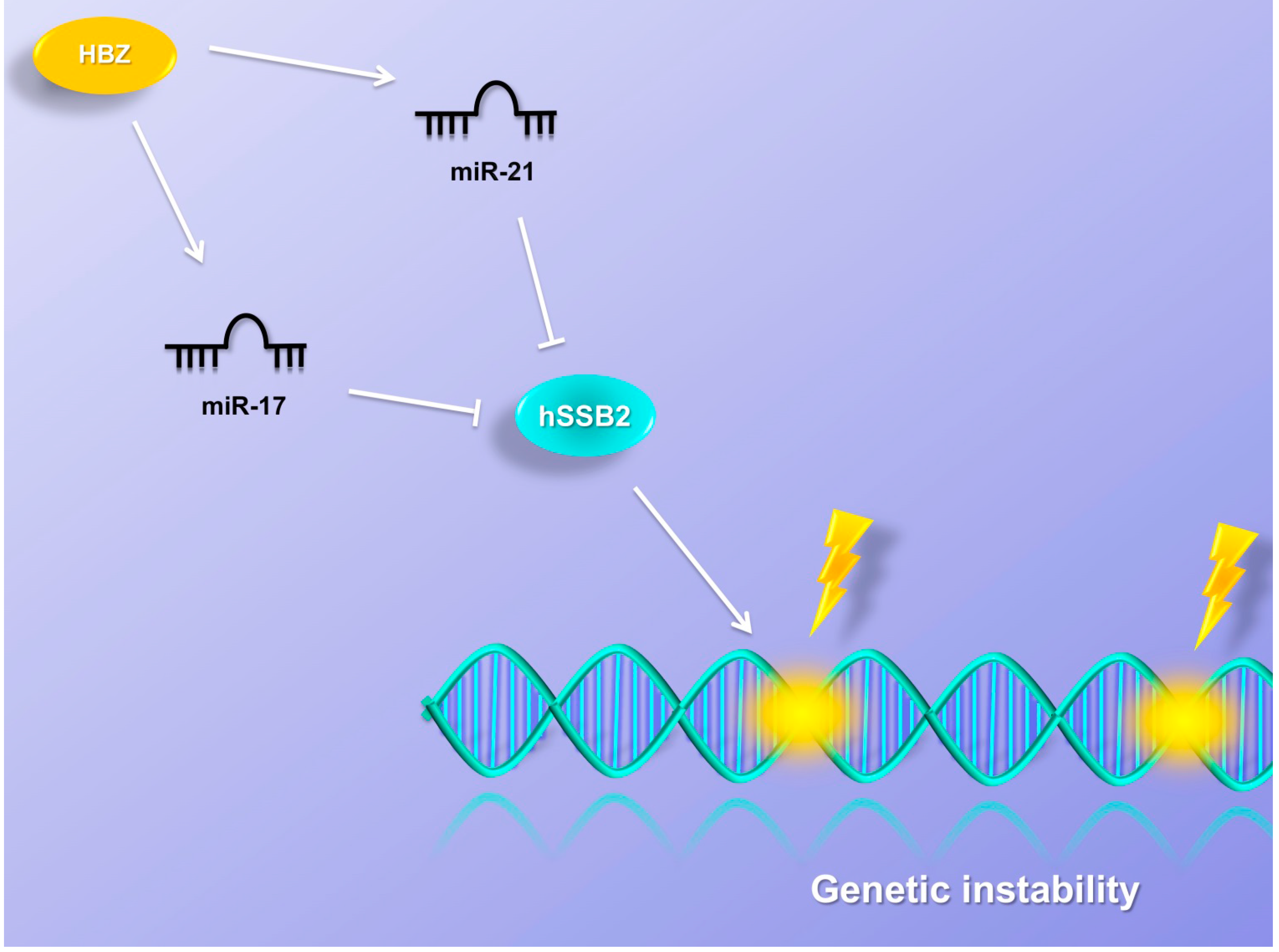
10. Conclusions and Prospective
Acknowledgments
Author Contributions
Conflicts of Interest
References
- Yoshida, M.; Miyoshi, I.; Hinuma, Y. Isolation and characterization of retrovirus from cell lines of human adult T-cell leukemia and its implication in the disease. Proc. Natl. Acad. Sci. USA 1982, 79, 2031–2035. [Google Scholar] [CrossRef] [PubMed]
- Nicot, C. Current views in HTLV-I-associated adult T-cell leukemia/lymphoma. Am. J. Hematol. 2005, 78, 232–239. [Google Scholar] [CrossRef] [PubMed]
- Gessain, A.; Barin, F.; Vernant, J.C.; Gout, O.; Maurs, L.; Calender, A.; de The, G. Antibodies to human T-lymphotropic virus type-I in patients with tropical spastic paraparesis. Lancet 1985, 2, 407–410. [Google Scholar] [CrossRef]
- Shimoyama, M. Diagnostic criteria and classification of clinical subtypes of adult T-cell leukaemia-lymphoma. A report from the lymphoma study group (1984–1987). Br. J. Haematol. 1991, 79, 428–437. [Google Scholar] [CrossRef] [PubMed]
- Bazarbachi, A.; Ghez, D.; Lepelletier, Y.; Nasr, R.; de The, H.; el-Sabban, M.E.; Hermine, O. New therapeutic approaches for adult T-cell leukaemia. Lancet Oncol. 2004, 5, 664–672. [Google Scholar] [CrossRef]
- Franchini, G.; Nicot, C.; Johnson, J.M. Seizing of T cells by human T-cell leukemia/lymphoma virus type 1. Adv. Cancer Res. 2003, 89, 69–132. [Google Scholar] [PubMed]
- Matsuoka, M.; Jeang, K.T. Human T-cell leukemia virus type 1 (HTLV-1) and leukemic transformation: Viral infectivity, tax, HBZ and therapy. Oncogene 2011, 30, 1379–1389. [Google Scholar] [CrossRef] [PubMed]
- Kashanchi, F.; Brady, J.N. Transcriptional and post-transcriptional gene regulation of HTLV-1. Oncogene 2005, 24, 5938–5951. [Google Scholar] [CrossRef] [PubMed]
- Giam, C.Z.; Jeang, K.T. HTLV-1 tax and adult T-cell leukemia. Front. Biosci. 2007, 12, 1496–1507. [Google Scholar] [CrossRef] [PubMed]
- Boxus, M.; Twizere, J.C.; Legros, S.; Dewulf, J.F.; Kettmann, R.; Willems, L. The HTLV-1 tax interactome. Retrovirology 2008, 5, e76. [Google Scholar] [CrossRef] [PubMed]
- Satou, Y.; Yasunaga, J.; Yoshida, M.; Matsuoka, M. HTLV-I basic leucine zipper factor gene mRNA supports proliferation of adult T cell leukemia cells. Proc. Natl. Acad. Sci. USA 2006, 103, 720–725. [Google Scholar] [CrossRef] [PubMed]
- Zhao, T.; Matsuoka, M. HBZ and its roles in HTLV-1 oncogenesis. Front. Microbial. 2012, 3, e247. [Google Scholar] [CrossRef] [PubMed]
- Nakano, K.; Watanabe, T. HTLV-1 rex: The courier of viral messages making use of the host vehicle. Front. Microbiol. 2012, 3, e330. [Google Scholar] [CrossRef] [PubMed]
- Ivey, K.N.; Srivastava, D. Micrornas as regulators of differentiation and cell fate decisions. Cell Stem Cell 2010, 7, 36–41. [Google Scholar] [CrossRef] [PubMed]
- O’Connell, R.M.; Rao, D.S.; Chaudhuri, A.A.; Baltimore, D. Physiological and pathological roles for micrornas in the immune system. Nat. Rev. Immunol. 2010, 10, 111–122. [Google Scholar] [CrossRef] [PubMed]
- Bouyssou, J.M.; Manier, S.; Huynh, D.; Issa, S.; Roccaro, A.M.; Ghobrial, I.M. Regulation of micrornas in cancer metastasis. Biochim. Biophys. Acta 2014, 1845, 255–265. [Google Scholar] [CrossRef] [PubMed]
- Lee, R.C.; Feinbaum, R.L.; Ambros, V. The c. Elegans heterochronic gene lin-4 encodes small RNAs with antisense complementarity to lin-14. Cell 1993, 75, 843–854. [Google Scholar] [CrossRef]
- Wightman, B.; Ha, I.; Ruvkun, G. Posttranscriptional regulation of the heterochronic gene lin-14 by lin-4 mediates temporal pattern formation in C. Elegans. Cell 1993, 75, 855–862. [Google Scholar] [CrossRef]
- Ha, M.; Kim, V.N. Regulation of microRNA biogenesis. Nat. Rev. Mol. Cell Biol. 2014, 15, 509–524. [Google Scholar] [CrossRef] [PubMed]
- Lee, Y.; Jeon, K.; Lee, J.T.; Kim, S.; Kim, V.N. Microrna maturation: Stepwise processing and subcellular localization. EMBO J. 2002, 21, 4663–4670. [Google Scholar] [CrossRef] [PubMed]
- Holley, C.L.; Topkara, V.K. An introduction to small non-coding RNAs: MiRNA and snoRNA. Cardiovasc. Drugs Ther. 2011, 25, 151–159. [Google Scholar] [CrossRef] [PubMed]
- Havens, M.A.; Reich, A.A.; Duelli, D.M.; Hastings, M.L. Biogenesis of mammalian micrornas by a non-canonical processing pathway. Nucleic Acids Res. 2012, 40, 4626–4640. [Google Scholar] [CrossRef] [PubMed]
- Xia, J.; Zhang, W. Noncanonical micrornas and endogenous siRNAs in lytic infection of murine gammaherpesvirus. PLoS ONE 2012, 7, e47863. [Google Scholar] [CrossRef] [PubMed]
- Valencia-Sanchez, M.A.; Liu, J.; Hannon, G.J.; Parker, R. Control of translation and mRNA degradation by mirnas and sirnas. Genes Dev. 2006, 20, 515–524. [Google Scholar] [CrossRef] [PubMed]
- Brennecke, J.; Hipfner, D.R.; Stark, A.; Russell, R.B.; Cohen, S.M. Bantam encodes a developmentally regulated microRNA that controls cell proliferation and regulates the proapoptotic gene hid in drosophila. Cell 2003, 113, 25–36. [Google Scholar] [CrossRef]
- Cimmino, A.; Calin, G.A.; Fabbri, M.; Iorio, M.V.; Ferracin, M.; Shimizu, M.; Wojcik, S.E.; Aqeilan, R.I.; Zupo, S.; Dono, M.; et al. Mir-15 and mir-16 induce apoptosis by targeting bcl2. Proc. Natl. Acad. Sci. USA 2005, 102, 13944–13949. [Google Scholar] [CrossRef] [PubMed]
- Pichler, K.; Schneider, G.; Grassmann, R. Microrna mir-146a and further oncogenesis-related cellular micrornas are dysregulated in HTLV-1-transformed T lymphocytes. Retrovirology 2008, 5, e100. [Google Scholar] [CrossRef] [PubMed]
- Bellon, M.; Lepelletier, Y.; Hermine, O.; Nicot, C. Deregulation of microrna involved in hematopoiesis and the immune response in HTLV-I adult T-cell leukemia. Blood 2009, 113, 4914–4917. [Google Scholar] [CrossRef] [PubMed]
- Yeung, M.L.; Yasunaga, J.; Bennasser, Y.; Dusetti, N.; Harris, D.; Ahmad, N.; Matsuoka, M.; Jeang, K.T. Roles for micrornas, miR-93 and miR-130b, and tumor protein 53-induced nuclear protein 1 tumor suppressor in cell growth dysregulation by human T-cell lymphotrophic virus 1. Cancer Res. 2008, 68, 8976–8985. [Google Scholar] [CrossRef] [PubMed]
- Yamagishi, M.; Nakano, K.; Miyake, A.; Yamochi, T.; Kagami, Y.; Tsutsumi, A.; Matsuda, Y.; Sato-Otsubo, A.; Muto, S.; Utsunomiya, A.; et al. Polycomb-mediated loss of miR-31 activates nik-dependent NF- κb pathway in adult T cell leukemia and other cancers. Cancer Cell 2012, 21, 121–135. [Google Scholar] [CrossRef] [PubMed]
- Bennasser, Y.; Yeung, M.L.; Jeang, K.T. HIV-1 tar RNA subverts RNA interference in transfected cells through sequestration of tar RNA-binding protein, TRBP. J. Biol. Chem. 2006, 281, 27674–27678. [Google Scholar] [CrossRef] [PubMed]
- Haasnoot, J.; de Vries, W.; Geutjes, E.J.; Prins, M.; de Haan, P.; Berkhout, B. The ebola virus vp35 protein is a suppressor of RNA silencing. PLoS Pathog. 2007, 3, e86. [Google Scholar] [CrossRef] [PubMed]
- Godshalk, S.E.; Bhaduri-McIntosh, S.; Slack, F.J. Epstein-barr virus-mediated dysregulation of human microRNA expression. Cell Cycle 2008, 7, 3595–3600. [Google Scholar] [CrossRef] [PubMed]
- De Vries, W.; Haasnoot, J.; Fouchier, R.; de Haan, P.; Berkhout, B. Differential RNA silencing suppression activity of NS 1 proteins from different influenza a virus strains. J. Gen. Virol. 2009, 90, 1916–1922. [Google Scholar] [CrossRef] [PubMed]
- Xie, K.L.; Zhang, Y.G.; Liu, J.; Zeng, Y.; Wu, H. Micrornas associated with HBV infection and HBV-related HCC. Theranostics 2014, 4, 1176–1192. [Google Scholar] [CrossRef] [PubMed]
- Wang, Y.; Kato, N.; Jazag, A.; Dharel, N.; Otsuka, M.; Taniguchi, H.; Kawabe, T.; Omata, M. Hepatitis C virus core protein is a potent inhibitor of RNA silencing-based antiviral response. Gastroenterology 2006, 130, 883–892. [Google Scholar] [CrossRef] [PubMed]
- Lu, S.; Cullen, B.R. Adenovirus va1 noncoding RNA can inhibit small interfering RNA and microRNA biogenesis. J. Virol. 2004, 78, 12868–12876. [Google Scholar] [CrossRef] [PubMed]
- Van Duyne, R.; Guendel, I.; Klase, Z.; Narayanan, A.; Coley, W.; Jaworski, E.; Roman, J.; Popratiloff, A.; Mahieux, R.; Kehn-Hall, K.; et al. Localization and sub-cellular shuttling of HTLV-1 tax with the mirna machinery. PLoS ONE 2012, 7, e40662. [Google Scholar] [CrossRef] [PubMed]
- Abe, M.; Suzuki, H.; Nishitsuji, H.; Shida, H.; Takaku, H. Interaction of human T-cell lymphotropic virus type I rex protein with dicer suppresses RNAi silencing. FEBS Lett. 2010, 584, 4313–4318. [Google Scholar] [CrossRef] [PubMed]
- Gottwein, E. Roles of micrornas in the life cycles of mammalian viruses. Curr. Top. Microbiol. Immunol. 2013, 371, 201–227. [Google Scholar] [PubMed]
- Sedger, L.M. MicroRNA control of interferons and interferon induced anti-viral activity. Mol. Immunol. 2013, 56, 781–793. [Google Scholar] [CrossRef] [PubMed]
- Lei, X.; Bai, Z.; Ye, F.; Huang, Y.; Gao, S.J. MicroRNAs control herpesviral dormancy. Cell Cycle 2010, 9, 1225–1226. [Google Scholar] [CrossRef] [PubMed]
- Swaminathan, S. Noncoding RNAs produced by oncogenic human herpesviruses. J. Cell. Physiol. 2008, 216, 321–326. [Google Scholar] [CrossRef] [PubMed]
- Sun, L.; Li, Q. The miRNAs of herpes simplex virus (HSV). Virol. Sin. 2012, 27, 333–338. [Google Scholar] [CrossRef] [PubMed]
- Kincaid, R.P.; Burke, J.M.; Sullivan, C.S. RNA virus microrna that mimics a B-cell oncomir. Proc. Natl. Acad. Sci. USA 2012, 109, 3077–3082. [Google Scholar] [CrossRef] [PubMed]
- Rosewick, N.; Momont, M.; Durkin, K.; Takeda, H.; Caiment, F.; Cleuter, Y.; Vernin, C.; Mortreux, F.; Wattel, E.; Burny, A.; et al. Deep sequencing reveals abundant noncanonical retroviral micrornas in B-cell leukemia/lymphoma. Proc. Natl. Acad. Sci. USA 2013, 110, 2306–2311. [Google Scholar] [CrossRef] [PubMed]
- Huang, J.; Wang, F.; Argyris, E.; Chen, K.; Liang, Z.; Tian, H.; Huang, W.; Squires, K.; Verlinghieri, G.; Zhang, H. Cellular micrornas contribute to HIV-1 latency in resting primary CD4+ T lymphocytes. Nat. Med. 2007, 13, 1241–1247. [Google Scholar] [CrossRef] [PubMed]
- Bai, X.T.; Nicot, C. MiR-28–3p is a cellular restriction factor that inhibits human T cell leukemia virus, type 1 (HTLV-1) replication and virus infection. J. Biol. Chem. 2015, 290, 5381–5390. [Google Scholar] [CrossRef] [PubMed]
- He, H.; Jazdzewski, K.; Li, W.; Liyanarachchi, S.; Nagy, R.; Volinia, S.; Calin, G.A.; Liu, C.G.; Franssila, K.; Suster, S.; et al. The role of microrna genes in papillary thyroid carcinoma. Proc. Natl. Acad. Sci. USA 2005, 102, 19075–19080. [Google Scholar] [CrossRef] [PubMed]
- Pacifico, F.; Crescenzi, E.; Mellone, S.; Iannetti, A.; Porrino, N.; Liguoro, D.; Moscato, F.; Grieco, M.; Formisano, S.; Leonardi, A. Nuclear factor-κb contributes to anaplastic thyroid carcinomas through up-regulation of mir-146a. J. Clin. Endocrinol. Metab. 2010, 95, 1421–1430. [Google Scholar] [CrossRef] [PubMed]
- Bhaumik, D.; Scott, G.K.; Schokrpur, S.; Patil, C.K.; Campisi, J.; Benz, C.C. Expression of microRNA-146 suppresses NF-κb activity with reduction of metastatic potential in breast cancer cells. Oncogene 2008, 27, 5643–5647. [Google Scholar] [CrossRef] [PubMed]
- Lavon, I.; Zrihan, D.; Granit, A.; Einstein, O.; Fainstein, N.; Cohen, M.A.; Cohen, M.A.; Zelikovitch, B.; Shoshan, Y.; Spektor, S.; et al. Gliomas display a microrna expression profile reminiscent of neural precursor cells. Neuro Oncol. 2010, 12, 422–433. [Google Scholar] [PubMed]
- Wang, X.; Tang, S.; Le, S.Y.; Lu, R.; Rader, J.S.; Meyers, C.; Zheng, Z.M. Aberrant expression of oncogenic and tumor-suppressive micrornas in cervical cancer is required for cancer cell growth. PLoS ONE 2008, 3, e2557. [Google Scholar] [CrossRef] [PubMed]
- Li, L.; Chen, X.P.; Li, Y.J. Microrna-146a and human disease. Scand. J. Immunol. 2010, 71, 227–231. [Google Scholar] [CrossRef] [PubMed]
- Kogo, R.; Mimori, K.; Tanaka, F.; Komune, S.; Mori, M. Clinical significance of miR-146a in gastric cancer cases. Clin. Cancer Res. 2011, 17, 4277–4284. [Google Scholar] [CrossRef] [PubMed]
- Lin, S.L.; Chiang, A.; Chang, D.; Ying, S.Y. Loss of miR-146a function in hormone-refractory prostate cancer. RNA 2008, 14, 417–424. [Google Scholar] [CrossRef] [PubMed]
- Garzon, R.; Volinia, S.; Liu, C.G.; Fernandez-Cymering, C.; Palumbo, T.; Pichiorri, F.; Fabbri, M.; Coombes, K.; Alder, H.; Nakamura, T.; et al. Microrna signatures associated with cytogenetics and prognosis in acute myeloid leukemia. Blood 2008, 111, 3183–3189. [Google Scholar] [CrossRef] [PubMed]
- Starczynowski, D.T.; Kuchenbauer, F.; Argiropoulos, B.; Sung, S.; Morin, R.; Muranyi, A.; Hirst, M.; Hogge, D.; Marra, M.; Wells, R.A.; et al. Identification of miR-145 and miR-146a as mediators of the 5q- syndrome phenotype. Nat. Med. 2010, 16, 49–58. [Google Scholar] [CrossRef] [PubMed]
- Visone, R.; Rassenti, L.Z.; Veronese, A.; Taccioli, C.; Costinean, S.; Aguda, B.D.; Volinia, S.; Ferracin, M.; Palatini, J.; Balatti, V.; et al. Karyotype-specific microRNA signature in chronic lymphocytic leukemia. Blood 2009, 114, 3872–3879. [Google Scholar] [CrossRef] [PubMed]
- Tomita, M.; Tanaka, Y.; Mori, N. Microrna mir-146a is induced by HTLV-1 tax and increases the growth of HTLV-1-infected T-cells. Int. J. Cancer 2012, 130, 2300–2309. [Google Scholar] [CrossRef] [PubMed]
- Sandhu, R.; Rein, J.; D'Arcy, M.; Herschkowitz, J.I.; Hoadley, K.A.; Troester, M.A. Overexpression of miR-146a in basal-like breast cancer cells confers enhanced tumorigenic potential in association with altered p53 status. Carcinogenesis 2014, 35, 2567–2575. [Google Scholar] [CrossRef] [PubMed]
- Xiao, B.; Zhu, E.D.; Li, N.; Lu, D.S.; Li, W.; Li, B.S.; Zhao, Y.L.; Mao, X.H.; Guo, G.; Yu, P.W.; et al. Increased miR-146a in gastric cancer directly targets SMAD4 and is involved in modulating cell proliferation and apoptosis. Oncol. Rep. 2012, 27, 559–566. [Google Scholar] [PubMed]
- Hsieh, J.Y.; Huang, T.S.; Cheng, S.M.; Lin, W.S.; Tsai, T.N.; Lee, O.K.; Wang, H.W. MiR-146a-5p circuitry uncouples cell proliferation and migration, but not differentiation, in human mesenchymal stem cells. Nucleic Acids Res. 2013, 41, 9753–9763. [Google Scholar] [CrossRef] [PubMed]
- Wang, Y.; Li, Z.; He, C.; Wang, D.; Yuan, X.; Chen, J.; Jin, J. Micrornas expression signatures are associated with lineage and survival in acute leukemias. Blood Cells Mol. Dis. 2010, 44, 191–197. [Google Scholar] [CrossRef] [PubMed]
- Vasilatou, D.; Papageorgiou, S.; Pappa, V.; Papageorgiou, E.; Dervenoulas, J. The role of micro RNAs in normal and malignant hematopoiesis. Eur. J. Haematol. 2010, 84, 1–16. [Google Scholar] [CrossRef] [PubMed]
- Baltimore, D.; Boldin, M.P.; O’Connell, R.M.; Rao, D.S.; Taganov, K.D. Micrornas: New regulators of immune cell development and function. Nat. Immunol. 2008, 9, 839–845. [Google Scholar] [CrossRef] [PubMed]
- Kluiver, J.; Poppema, S.; de Jong, D.; Blokzijl, T.; Harms, G.; Jacobs, S.; Kroesen, B.J.; van den Berg, A. Bic and mir-155 are highly expressed in hodgkin, primary mediastinal and diffuse large B cell lymphomas. J. Pathol. 2005, 207, 243–249. [Google Scholar] [CrossRef] [PubMed]
- Tili, E.; Croce, C.M.; Michaille, J.J. Mir-155: On the crosstalk between inflammation and cancer. Int. Rev. Immunol. 2009, 28, 264–284. [Google Scholar] [CrossRef] [PubMed]
- Costinean, S.; Zanesi, N.; Pekarsky, Y.; Tili, E.; Volinia, S.; Heerema, N.; Croce, C.M. Pre-B cell proliferation and lymphoblastic leukemia/high-grade lymphoma in e(mu)-miR-155 transgenic mice. Proc. Natl. Acad. Sci. USA 2006, 103, 7024–7029. [Google Scholar] [CrossRef] [PubMed]
- Chen, Z.; Ma, T.; Huang, C.; Hu, T.; Li, J. The pivotal role of microRNA-155 in the control of cancer. J. Cell. Physiol. 2014, 229, 545–550. [Google Scholar] [CrossRef] [PubMed]
- Habbe, N.; Koorstra, J.B.; Mendell, J.T.; Offerhaus, G.J.; Ryu, J.K.; Feldmann, G.; Mullendore, M.E.; Goggins, M.G.; Hong, S.M.; Maitra, A. Microrna miR-155 is a biomarker of early pancreatic neoplasia. Cancer Biol. Ther. 2009, 8, 340–346. [Google Scholar] [CrossRef] [PubMed]
- Zang, Y.S.; Zhong, Y.F.; Fang, Z.; Li, B.; An, J. MiR-155 inhibits the sensitivity of lung cancer cells to cisplatin via negative regulation of APAF-1 expression. Cancer Gene Ther. 2012, 19, 773–778. [Google Scholar] [CrossRef] [PubMed]
- Saito, Y.; Suzuki, H.; Tsugawa, H.; Imaeda, H.; Matsuzaki, J.; Hirata, K.; Hosoe, N.; Nakamura, M.; Mukai, M.; Saito, H.; et al. Overexpression of mir-142–5p and mir-155 in gastric mucosa-associated lymphoid tissue (malt) lymphoma resistant to helicobacter pylori eradication. PLoS ONE 2012, 7, e47396. [Google Scholar] [CrossRef] [PubMed]
- Garzon, R.; Garofalo, M.; Martelli, M.P.; Briesewitz, R.; Wang, L.; Fernandez-Cymering, C.; Volinia, S.; Liu, C.G.; Schnittger, S.; Haferlach, T.; et al. Distinctive microrna signature of acute myeloid leukemia bearing cytoplasmic mutated nucleophosmin. Proc. Natl. Acad. Sci. USA 2008, 105, 3945–3950. [Google Scholar] [CrossRef] [PubMed]
- Tomita, M. Important roles of cellular microrna mir-155 in leukemogenesis by human T-cell leukemia virus type 1 infection. ISRN Microbiol. 2012, 2012, e978607. [Google Scholar] [CrossRef] [PubMed]
- Babar, I.A.; Cheng, C.J.; Booth, C.J.; Liang, X.; Weidhaas, J.B.; Saltzman, W.M.; Slack, F.J. Nanoparticle-based therapy in an in vivo micro RNA-155 (miR-155)-dependent mouse model of lymphoma. Proc. Natl. Acad. Sci. USA 2012, 109, E1695–E1704. [Google Scholar] [CrossRef] [PubMed]
- Calin, G.A.; Ferracin, M.; Cimmino, A.; di Leva, G.; Shimizu, M.; Wojcik, S.E.; Iorio, M.V.; Visone, R.; Sever, N.I.; Fabbri, M.; et al. A micro RNA signature associated with prognosis and progression in chronic lymphocytic leukemia. N. Engl. J. Med. 2005, 353, 1793–1801. [Google Scholar] [CrossRef] [PubMed]
- Lawrie, C.H. Micrornas and lymphomagenesis: A functional review. Br. J. Haematol. 2013, 160, 571–581. [Google Scholar] [CrossRef] [PubMed]
- Wang, L.; Toomey, N.L.; Diaz, L.A.; Walker, G.; Ramos, J.C.; Barber, G.N.; Ning, S. Oncogenic IRFS provide a survival advantage for epstein-barr virus- or human T-cell leukemia virus type 1-transformed cells through induction of Bic expression. J. Virol. 2011, 85, 8328–8337. [Google Scholar] [CrossRef] [PubMed]
- Shaffer, A.L.; Emre, N.C.; Romesser, P.B.; Staudt, L.M. Irf4: Immunity. Malignancy! Therapy? Clin. Cancer Res. 2009, 15, 2954–2961. [Google Scholar] [CrossRef] [PubMed]
- Ramos, J.C.; Ruiz, P.; Ratner, L.; Reis, I.M.; Brites, C.; Pedroso, C.; Byrne, G.E.; Toomey, N.L.; Andela, V.; Harhaj, E.W.; et al. IRF -4 and c-Rel expression in antiviral-resistant adult T-cell leukemia/lymphoma. Blood 2007, 109, 3060–3068. [Google Scholar] [CrossRef] [PubMed]
- Sharma, S.; Mamane, Y.; Grandvaux, N.; Bartlett, J.; Petropoulos, L.; Lin, R.; Hiscott, J. Activation and regulation of interferon regulatory factor 4 in HTLV type 1-infected T lymphocytes. AIDS Res. Hum. Retroviruses 2000, 16, 1613–1622. [Google Scholar] [CrossRef] [PubMed]
- Suzuki, S.; Zhou, Y.; Refaat, A.; Takasaki, I.; Koizumi, K.; Yamaoka, S.; Tabuchi, Y.; Saiki, I.; Sakurai, H. Human T cell lymphotropic virus 1 manipulates interferon regulatory signals by controlling the TAK1-IRF3 and IRF4 pathways. J. Biol. Chem. 2010, 285, 4441–4446. [Google Scholar] [CrossRef] [PubMed]
- Liu, F.; Kong, X.; Lv, L.; Gao, J. MiR-155 targets tp53inp1 to regulate liver cancer stem cell acquisition and self-renewal. FEBS Lett. 2015, 589, 500–506. [Google Scholar] [CrossRef] [PubMed]
- Papakonstantinou, N.; Ntoufa, S.; Chartomatsidou, E.; Papadopoulos, G.; Hatzigeorgiou, A.; Anagnostopoulos, A.; Chlichlia, K.; Ghia, P.; Muzio, M.; Belessi, C.; et al. Differential microrna profiles and their functional implications in different immunogenetic subsets of chronic lymphocytic leukemia. Mol. Med. 2013, 19, 115–123. [Google Scholar] [CrossRef] [PubMed]
- Mraz, M.; Chen, L.; Rassenti, L.Z.; Ghia, E.M.; Li, H.; Jepsen, K.; Smith, E.N.; Messer, K.; Frazer, K.A.; Kipps, T.J. MiR-150 influences B-cell receptor signaling in chronic lymphocytic leukemia by regulating expression of GAB1 and FOXP1. Blood 2014, 124, 84–95. [Google Scholar] [CrossRef] [PubMed]
- Morris, V.A.; Zhang, A.; Yang, T.; Stirewalt, D.L.; Ramamurthy, R.; Meshinchi, S.; Oehler, V.G. MicroRNA-150 expression induces myeloid differentiation of human acute leukemia cells and normal hematopoietic progenitors. PLoS ONE 2013, 8, e75815. [Google Scholar] [CrossRef] [PubMed]
- Machova Polakova, K.; Lopotova, T.; Klamova, H.; Burda, P.; Trneny, M.; Stopka, T.; Moravcova, J. Expression patterns of microRNAs associated with cml phases and their disease related targets. Mol. Cancer 2011, 10, e41. [Google Scholar] [CrossRef] [PubMed]
- Xu, L.; Liang, Y.N.; Luo, X.Q.; Liu, X.D.; Guo, H.X. Association of mirnas expression profiles with prognosis and relapse in childhood acute lymphoblastic leukemia. Zhonghua Xue Ye Xue Za Zhi 2011, 32, 178–181. [Google Scholar] [PubMed]
- Zhao, J.J.; Lin, J.; Lwin, T.; Yang, H.; Guo, J.; Kong, W.; Dessureault, S.; Moscinski, L.C.; Rezania, D.; Dalton, W.S.; et al. Microrna expression profile and identification of miR-29 as a prognostic marker and pathogenetic factor by targeting CDK6 in mantle cell lymphoma. Blood 2010, 115, 2630–2639. [Google Scholar] [CrossRef] [PubMed]
- Cao, M.; Hou, D.; Liang, H.; Gong, F.; Wang, Y.; Yan, X.; Jiang, X.; Wang, C.; Zhang, J.; Zen, K.; et al. MiR-150 promotes the proliferation and migration of lung cancer cells by targeting SRC kinase signalling inhibitor 1. Eur. J. Cancer 2014, 50, 1013–1024. [Google Scholar] [CrossRef] [PubMed]
- Avery-Kiejda, K.A.; Braye, S.G.; Mathe, A.; Forbes, J.F.; Scott, R.J. Decreased expression of key tumour suppressor micrornas is associated with lymph node metastases in triple negative breast cancer. BMC Cancer 2014, 14, e51. [Google Scholar] [CrossRef] [PubMed]
- Huang, S.; Chen, Y.; Wu, W.; Ouyang, N.; Chen, J.; Li, H.; Liu, X.; Su, F.; Lin, L.; Yao, Y. MiR-150 promotes human breast cancer growth and malignant behavior by targeting the pro-apoptotic purinergic p2x7 receptor. PLoS ONE 2013, 8, e80707. [Google Scholar] [CrossRef] [PubMed]
- Xiao, C.; Calado, D.P.; Galler, G.; Thai, T.H.; Patterson, H.C.; Wang, J.; Rajewsky, N.; Bender, T.P.; Rajewsky, K. MiR-150 controls B cell differentiation by targeting the transcription factor c-myb. Cell 2007, 131, 146–159. [Google Scholar] [CrossRef] [PubMed]
- Ghisi, M.; Corradin, A.; Basso, K.; Frasson, C.; Serafin, V.; Mukherjee, S.; Mussolin, L.; Ruggero, K.; Bonanno, L.; Guffanti, A.; et al. Modulation of microRNA expression in human T-cell development: Targeting of notch3 by miR-150. Blood 2011, 117, 7053–7062. [Google Scholar] [CrossRef] [PubMed]
- Bousquet, M.; Zhuang, G.; Meng, C.; Ying, W.; Cheruku, P.S.; Shie, A.T.; Wang, S.; Ge, G.; Wong, P.; Wang, G.; et al. MiR-150 blocks MLL-AF9-associated leukemia through oncogene repression. Mol. Cancer Res. 2013, 11, 912–922. [Google Scholar] [CrossRef] [PubMed]
- Wu, Q.; Jin, H.; Yang, Z.; Luo, G.; Lu, Y.; Li, K.; Ren, G.; Su, T.; Pan, Y.; Feng, B.; et al. MiR-150 promotes gastric cancer proliferation by negatively regulating the pro-apoptotic gene EGR2. Biochem. Biophys. Res. Commun. 2010, 392, 340–345. [Google Scholar] [CrossRef] [PubMed]
- Watanabe, A.; Tagawa, H.; Yamashita, J.; Teshima, K.; Nara, M.; Iwamoto, K.; Kume, M.; Kameoka, Y.; Takahashi, N.; Nakagawa, T.; et al. The role of microRNA-150 as a tumor suppressor in malignant lymphoma. Leukemia 2011, 25, 1324–1334. [Google Scholar] [CrossRef] [PubMed]
- Gessain, A.; Cassar, O. Epidemiological aspects and world distribution of HTLV-1 infection. Front. Microbial. 2012, 3, e388. [Google Scholar] [CrossRef] [PubMed]
- Stamatopoulos, B.; Meuleman, N.; Haibe-Kains, B.; Saussoy, P.; van den Neste, E.; Michaux, L.; Heimann, P.; Martiat, P.; Bron, D.; Lagneaux, L. MicroRNA-29c and microRNA-223 down-regulation has in vivo significance in chronic lymphocytic leukemia and improves disease risk stratification. Blood 2009, 113, 5237–5245. [Google Scholar] [CrossRef] [PubMed]
- Mi, S.; Lu, J.; Sun, M.; Li, Z.; Zhang, H.; Neilly, M.B.; Wang, Y.; Qian, Z.; Jin, J.; Zhang, Y.; et al. Microrna expression signatures accurately discriminate acute lymphoblastic leukemia from acute myeloid leukemia. Proc. Natl. Acad. Sci. USA 2007, 104, 19971–19976. [Google Scholar] [CrossRef] [PubMed]
- Liu, T.Y.; Chen, S.U.; Kuo, S.H.; Cheng, A.L.; Lin, C.W. E2a-positive gastric malt lymphoma has weaker plasmacytoid infiltrates and stronger expression of the memory B-cell-associated miR-223: Possible correlation with stage and treatment response. Mod. Pathol. 2010, 23, 1507–1517. [Google Scholar] [CrossRef] [PubMed]
- Laios, A.; O'Toole, S.; Flavin, R.; Martin, C.; Kelly, L.; Ring, M.; Finn, S.P.; Barrett, C.; Loda, M.; Gleeson, N.; et al. Potential role of mir-9 and mir-223 in recurrent ovarian cancer. Mol. Cancer 2008, 7, e35. [Google Scholar] [CrossRef] [PubMed]
- Kumar, V.; Palermo, R.; Talora, C.; Campese, A.F.; Checquolo, S.; Bellavia, D.; Tottone, L.; Testa, G.; Miele, E.; Indraccolo, S.; et al. Notch and NF-κb signaling pathways regulate mir-223/fbxw7 axis in T-cell acute lymphoblastic leukemia. Leukemia 2014, 28, 2324–2335. [Google Scholar] [CrossRef] [PubMed]
- Lee, J.E.; Hong, E.J.; Nam, H.Y.; Kim, J.W.; Han, B.G.; Jeon, J.P. Microrna signatures associated with immortalization of EBV-transformed lymphoblastoid cell lines and their clinical traits. Cell Prolif. 2011, 44, 59–66. [Google Scholar] [CrossRef] [PubMed]
- Li, J.; Guo, Y.; Liang, X.; Sun, M.; Wang, G.; De, W.; Wu, W. Microrna-223 functions as an oncogene in human gastric cancer by targeting FBXW7/HCDC4. J. Cancer Res. Clin. Oncol. 2012, 138, 763–774. [Google Scholar] [CrossRef] [PubMed]
- Li, X.; Zhang, Y.; Zhang, H.; Liu, X.; Gong, T.; Li, M.; Sun, L.; Ji, G.; Shi, Y.; Han, Z.; et al. MirRNA-223 promotes gastric cancer invasion and metastasis by targeting tumor suppressor EPB41l3. Mol. Cancer Res. 2011, 9, 824–833. [Google Scholar] [CrossRef] [PubMed]
- Sun, G.; Li, H.; Rossi, J.J. Sequence context outside the target region influences the effectiveness of mir-223 target sites in the rhob 3′utr. Nucleic Acids Res. 2010, 38, 239–252. [Google Scholar] [CrossRef] [PubMed]
- Wong, Q.W.; Lung, R.W.; Law, P.T.; Lai, P.B.; Chan, K.Y.; To, K.F.; Wong, N. Microrna-223 is commonly repressed in hepatocellular carcinoma and potentiates expression of stathmin1. Gastroenterology 2008, 135, 257–269. [Google Scholar] [CrossRef] [PubMed]
- Pulikkan, J.A.; Dengler, V.; Peramangalam, P.S.; Peer Zada, A.A.; Muller-Tidow, C.; Bohlander, S.K.; Tenen, D.G.; Behre, G. Cell-cycle regulator E2F1 and microRNA-223 comprise an autoregulatory negative feedback loop in acute myeloid leukemia. Blood 2010, 115, 1768–1778. [Google Scholar] [CrossRef] [PubMed]
- Haneklaus, M.; Gerlic, M.; O’Neill, L.A.; Masters, S.L. Mir-223: Infection, inflammation and cancer. J. Intern. Med. 2013, 274, 215–226. [Google Scholar] [CrossRef] [PubMed]
- McGirt, L.Y.; Adams, C.M.; Baerenwald, D.A.; Zwerner, J.P.; Zic, J.A.; Eischen, C.M. Mir-223 regulates cell growth and targets proto-oncogenes in mycosis fungoides/cutaneous t-cell lymphoma. J. Investig. Dermatol. 2014, 134, 1101–1107. [Google Scholar] [CrossRef] [PubMed]
- Decker, T.; Stockinger, S.; Karaghiosoff, M.; Muller, M.; Kovarik, P. Ifns and stats in innate immunity to microorganisms. J. Clin. Invest. 2002, 109, 1271–1277. [Google Scholar] [CrossRef] [PubMed]
- Kovacic, B.; Stoiber, D.; Moriggl, R.; Weisz, E.; Ott, R.G.; Kreibich, R.; Levy, D.E.; Beug, H.; Freissmuth, M.; Sexl, V. Stat1 acts as a tumor promoter for leukemia development. Cancer Cell 2006, 10, 77–87. [Google Scholar] [CrossRef] [PubMed]
- Sanda, T.; Tyner, J.W.; Gutierrez, A.; Ngo, V.N.; Glover, J.; Chang, B.H.; Yost, A.; Ma, W.; Fleischman, A.G.; Zhou, W.; et al. TYK2-STAT1-BCL2 pathway dependence in T-cell acute lymphoblastic leukemia. Cancer Discov. 2013, 3, 564–577. [Google Scholar] [CrossRef] [PubMed]
- Moles, R.; Bellon, M.; Nicot, C. Stat1: A novel target of mir-150 and mir-223 is involved in the proliferation of HTLV-I-transformed and ATL cells. Neoplasia 2015, 17, 449–462. [Google Scholar] [CrossRef] [PubMed]
- Valastyan, S.; Reinhardt, F.; Benaich, N.; Calogrias, D.; Szasz, A.M.; Wang, Z.C.; Brock, J.E.; Richardson, A.L.; Weinberg, R.A. A pleiotropically acting microrna, miR-31, inhibits breast cancer metastasis. Cell 2009, 137, 1032–1046. [Google Scholar] [CrossRef] [PubMed]
- Augoff, K.; Das, M.; Bialkowska, K.; McCue, B.; Plow, E.F.; Sossey-Alaoui, K. MiR-31 is a broad regulator of β 1-integrin expression and function in cancer cells. Mol. Cancer Res. 2011, 9, 1500–1508. [Google Scholar] [CrossRef] [PubMed]
- Fujikawa, D.; Yamagishi, M.; Kurokawa, N.; Soejima, A.; Ishida, T.; Tanaka, Y.; Nakano, K.; Watanabe, T. HTLV-1 Tax disrupts the host epigenome by interacting with a Polycomb group protein EZH2. Retrovirology 2014, 11, e144. [Google Scholar] [CrossRef]
- Sparmann, A.; van Lohuizen, M. Polycomb silencers control cell fate, development and cancer. Nat. Rev. Cancer 2006, 6, 846–856. [Google Scholar] [CrossRef] [PubMed]
- Richly, H.; Aloia, L.; di Croce, L. Roles of the polycomb group proteins in stem cells and cancer. Cell Death Dis. 2011, 2, e204. [Google Scholar] [CrossRef] [PubMed]
- Thu, Y.M.; Richmond, A. Nf-κb inducing kinase: A key regulator in the immune system and in cancer. Cytokine Growth Factor Rev. 2010, 21, 213–226. [Google Scholar] [CrossRef] [PubMed]
- Mori, N.; Fujii, M.; Ikeda, S.; Yamada, Y.; Tomonaga, M.; Ballard, D.W.; Yamamoto, N. Constitutive activation of NF-κb in primary adult T-cell leukemia cells. Blood 1999, 93, 2360–2368. [Google Scholar] [PubMed]
- Prasad, S.; Ravindran, J.; Aggarwal, B.B. NF-κb and cancer: How intimate is this relationship. Mol. Cell. Biochem. 2010, 336, 25–37. [Google Scholar] [CrossRef] [PubMed]
- Colangelo, T.; Fucci, A.; Votino, C.; Sabatino, L.; Pancione, M.; Laudanna, C.; Binaschi, M.; Bigioni, M.; Maggi, C.A.; Parente, D.; et al. MicroRNA-130b promotes tumor development and is associated with poor prognosis in colorectal cancer. Neoplasia 2013, 15, 1086–1099. [Google Scholar] [CrossRef] [PubMed]
- Lai, K.W.; Koh, K.X.; Loh, M.; Tada, K.; Subramaniam, M.M.; Lim, X.Y.; Vaithilingam, A.; Salto-Tellez, M.; Iacopetta, B.; Ito, Y.; et al. MicroRNA-130b regulates the tumour suppressor RUNX3 in gastric cancer. Eur. J. Cancer 2010, 46, 1456–1463. [Google Scholar] [CrossRef] [PubMed]
- Scheffer, A.R.; Holdenrieder, S.; Kristiansen, G.; von Ruecker, A.; Muller, S.C.; Ellinger, J. Circulating micrornas in serum: Novel biomarkers for patients with bladder cancer? World J. Urol. 2014, 32, 353–358. [Google Scholar] [CrossRef] [PubMed]
- Sand, M.; Skrygan, M.; Georgas, D.; Sand, D.; Gambichler, T.; Altmeyer, P.; Bechara, F.G. The miRNA machinery in primary cutaneous malignant melanoma, cutaneous malignant melanoma metastases and benign melanocytic nevi. Cell Tissue Res. 2012, 350, 119–126. [Google Scholar] [CrossRef] [PubMed]
- Chen, Z.; Jin, Y.; Yu, D.; Wang, A.; Mahjabeen, I.; Wang, C.; Liu, X.; Zhou, X. Down-regulation of the microrna-99 family members in head and neck squamous cell carcinoma. Oral Oncol. 2012, 48, 686–691. [Google Scholar] [CrossRef] [PubMed]
- Yip, L.; Kelly, L.; Shuai, Y.; Armstrong, M.J.; Nikiforov, Y.E.; Carty, S.E.; Nikiforova, M.N. Microrna signature distinguishes the degree of aggressiveness of papillary thyroid carcinoma. Ann. Surg. Oncol. 2011, 18, 2035–2041. [Google Scholar] [CrossRef] [PubMed]
- Yang, C.; Cai, J.; Wang, Q.; Tang, H.; Cao, J.; Wu, L.; Wang, Z. Epigenetic silencing of miR-130b in ovarian cancer promotes the development of multidrug resistance by targeting colony-stimulating factor 1. Gynecol. Oncol. 2012, 124, 325–334. [Google Scholar] [CrossRef] [PubMed]
- Dong, P.; Karaayvaz, M.; Jia, N.; Kaneuchi, M.; Hamada, J.; Watari, H.; Sudo, S.; Ju, J.; Sakuragi, N. Mutant p53 gain-of-function induces epithelial-mesenchymal transition through modulation of the miR-130b-ZEB1 axis. Oncogene 2013, 32, 3286–3295. [Google Scholar] [CrossRef] [PubMed]
- Zhao, G.; Zhang, J.G.; Shi, Y.; Qin, Q.; Liu, Y.; Wang, B.; Tian, K.; Deng, S.C.; Li, X.; Zhu, S.; et al. MiR-130b is a prognostic marker and inhibits cell proliferation and invasion in pancreatic cancer through targeting STAT3. PLoS ONE 2013, 8, e73803. [Google Scholar] [CrossRef] [PubMed]
- Yu, T.; Cao, R.; Li, S.; Fu, M.; Ren, L.; Chen, W.; Zhu, H.; Zhan, Q.; Shi, R. MiR-130b plays an oncogenic role by repressing pten expression in esophageal squamous cell carcinoma cells. BMC Cancer 2015, 15, e29. [Google Scholar] [CrossRef] [PubMed]
- Castro, N.E.; Kato, M.; Park, J.T.; Natarajan, R. Transforming growth factor beta1 (TGF-β1) enhances expression of profibrotic genes through a novel signaling cascade and micrornas in renal mesangial cells. J. Biol. Chem. 2014, 289, 29001–29013. [Google Scholar] [CrossRef] [PubMed]
- Petrocca, F.; Vecchione, A.; Croce, C.M. Emerging role of miR-106b-25/mir-17–92 clusters in the control of transforming growth factor beta signaling. Cancer Res. 2008, 68, 8191–8194. [Google Scholar] [CrossRef] [PubMed]
- Mendell, J.T. Miriad roles for the miR-17–92 cluster in development and disease. Cell 2008, 133, 217–222. [Google Scholar] [CrossRef] [PubMed]
- Hayashita, Y.; Osada, H.; Tatematsu, Y.; Yamada, H.; Yanagisawa, K.; Tomida, S.; Yatabe, Y.; Kawahara, K.; Sekido, Y.; Takahashi, T. A polycistronic microrna cluster, miR-17–92, is overexpressed in human lung cancers and enhances cell proliferation. Cancer Res. 2005, 65, 9628–9632. [Google Scholar] [CrossRef] [PubMed]
- Fu, X.; Tian, J.; Zhang, L.; Chen, Y.; Hao, Q. Involvement of microrna-93, a new regulator of PTEN/AKT signaling pathway, in regulation of chemotherapeutic drug cisplatin chemosensitivity in ovarian cancer cells. FEBS Lett. 2012, 586, 1279–1286. [Google Scholar] [CrossRef] [PubMed]
- Long, J.; Wang, Y.; Wang, W.; Chang, B.H.; Danesh, F.R. Identification of microrna-93 as a novel regulator of vascular endothelial growth factor in hyperglycemic conditions. J. Biol. Chem. 2010, 285, 23457–23465. [Google Scholar] [CrossRef] [PubMed]
- Fang, L.; Deng, Z.; Shatseva, T.; Yang, J.; Peng, C.; Du, W.W.; Yee, A.J.; Ang, L.C.; He, C.; Shan, S.W.; et al. Microrna miR-93 promotes tumor growth and angiogenesis by targeting integrin-beta8. Oncogene 2011, 30, 806–821. [Google Scholar] [CrossRef] [PubMed]
- Du, L.; Zhao, Z.; Ma, X.; Hsiao, T.H.; Chen, Y.; Young, E.; Suraokar, M.; Wistuba, I.; Minna, J.D.; Pertsemlidis, A. Mir-93-directed downregulation of dab2 defines a novel oncogenic pathway in lung cancer. Oncogene 2014, 33, 4307–4315. [Google Scholar] [CrossRef] [PubMed]
- Fang, L.; Du, W.W.; Yang, W.; Rutnam, Z.J.; Peng, C.; Li, H.; O’Malley, Y.Q.; Askeland, R.W.; Sugg, S.; Liu, M.; et al. MiR-93 enhances angiogenesis and metastasis by targeting LATS2. Cell Cycle 2012, 11, 4352–4365. [Google Scholar] [CrossRef] [PubMed]
- Okamura, S.; Arakawa, H.; Tanaka, T.; Nakanishi, H.; Ng, C.C.; Taya, Y.; Monden, M.; Nakamura, Y. p53DINP1, a p53-inducible gene, regulates p53-dependent apoptosis. Mol. Cell 2001, 8, 85–94. [Google Scholar] [CrossRef]
- Tomasini, R.; Seux, M.; Nowak, J.; Bontemps, C.; Carrier, A.; Dagorn, J.C.; Pebusque, M.J.; Iovanna, J.L.; Dusetti, N.J. TP53INP1 is a novel p73 target gene that induces cell cycle arrest and cell death by modulating p73 transcriptional activity. Oncogene 2005, 24, 8093–8104. [Google Scholar] [CrossRef] [PubMed]
- Gironella, M.; Seux, M.; Xie, M.J.; Cano, C.; Tomasini, R.; Gommeaux, J.; Garcia, S.; Nowak, J.; Yeung, M.L.; Jeang, K.T.; et al. Tumor protein 53-induced nuclear protein 1 expression is repressed by miR-155, and its restoration inhibits pancreatic tumor development. Proc. Natl. Acad. Sci. USA 2007, 104, 16170–16175. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Jiang, F.; Liu, T.; He, Y.; Yan, Q.; Chen, X.; Wang, H.; Wan, X. MiR-125b promotes proliferation and migration of type II endometrial carcinoma cells through targeting TP53INP1 tumor suppressor in vitro and in vivo. BMC Cancer 2011, 11, e425. [Google Scholar] [CrossRef] [PubMed]
- Hershko, T.; Chaussepied, M.; Oren, M.; Ginsberg, D. Novel link between e2f and p53: Proapoptotic cofactors of p53 are transcriptionally upregulated by e2f. Cell Death Differ. 2005, 12, 377–383. [Google Scholar] [CrossRef] [PubMed]
- Seux, M.; Peuget, S.; Montero, M.P.; Siret, C.; Rigot, V.; Clerc, P.; Gigoux, V.; Pellegrino, E.; Pouyet, L.; N’Guessan, P.; et al. TP53INP1 decreases pancreatic cancer cell migration by regulating sparc expression. Oncogene 2011, 30, 3049–3061. [Google Scholar] [CrossRef] [PubMed]
- Varrin-Doyer, M.; Nicolle, A.; Marignier, R.; Cavagna, S.; Benetollo, C.; Wattel, E.; Giraudon, P. Human T lymphotropic virus type 1 increases T lymphocyte migration by recruiting the cytoskeleton organizer CRMP2. J. Immunol. 2012, 188, 1222–1233. [Google Scholar] [CrossRef] [PubMed]
- Beimling, P.; Moelling, K. Direct interaction of CREB protein with 21 bp tax-response elements of HTLV-Iltr. Oncogene 1992, 7, 257–262. [Google Scholar] [PubMed]
- Zhao, L.J.; Giam, C.Z. Human T-cell lymphotropic virus type I (HTLV-I) transcriptional activator, tax, enhances CREB binding to HTLV-I 21-base-pair repeats by protein-protein interaction. Proc. Natl. Acad. Sci. USA 1992, 89, 7070–7074. [Google Scholar] [CrossRef] [PubMed]
- Adya, N.; Giam, C.Z. Distinct regions in human T-cell lymphotropic virus type I tax mediate interactions with activator protein CREB and basal transcription factors. J. Virol. 1995, 69, 1834–1841. [Google Scholar] [PubMed]
- Harrod, R.; Tang, Y.; Nicot, C.; Lu, H.S.; Vassilev, A.; Nakatani, Y.; Giam, C.Z. An exposed kid-like domain in human T-cell lymphotropic virus type 1 tax is responsible for the recruitment of coactivators CBP/P300. Mol. Cell. Biol. 1998, 18, 5052–5061. [Google Scholar] [PubMed]
- Harrod, R.; Kuo, Y.L.; Tang, Y.; Yao, Y.; Vassilev, A.; Nakatani, Y.; Giam, C.Z. P300 and P300/CAMP-responsive element-binding protein associated factor interact with human T-cell lymphotropic virus type-1 tax in a multi-histone acetyltransferase/activator-enhancer complex. J. Biol. Chem. 2000, 275, 11852–11857. [Google Scholar] [CrossRef] [PubMed]
- Bogenberger, J.M.; Laybourn, P.J. Human T lymphotropic virus type 1 protein tax reduces histone levels. Retrovirology 2008, 5, e9. [Google Scholar] [CrossRef] [PubMed]
- Rahman, S.; Quann, K.; Pandya, D.; Singh, S.; Khan, Z.K.; Jain, P. HTLV-1 tax mediated downregulation of mirnas associated with chromatin remodeling factors in T cells with stably integrated viral promoter. PLoS ONE 2012, 7, e34490. [Google Scholar] [CrossRef] [PubMed]
- Lin, R.J.; Lin, Y.C.; Yu, A.L. MiR-149* induces apoptosis by inhibiting AKT1 and E2F1 in human cancer cells. Mol. Carcinog. 2010, 49, 719–727. [Google Scholar] [CrossRef] [PubMed]
- Jin, L.; Hu, W.L.; Jiang, C.C.; Wang, J.X.; Han, C.C.; Chu, P.; Zhang, L.J.; Thorne, R.F.; Wilmott, J.; Scolyer, R.A.; et al. MicroRNA-149*, a p53-responsive microRNA, functions as an oncogenic regulator in human melanoma. Proc. Natl. Acad. Sci. USA 2011, 108, 15840–15845. [Google Scholar] [CrossRef] [PubMed]
- Schaefer, A.; Jung, M.; Mollenkopf, H.J.; Wagner, I.; Stephan, C.; Jentzmik, F.; Miller, K.; Lein, M.; Kristiansen, G.; Jung, K. Diagnostic and prognostic implications of microrna profiling in prostate carcinoma. Int. J. Cancer 2010, 126, 1166–1176. [Google Scholar] [CrossRef] [PubMed]
- Li, D.; Chen, P.; Li, X.Y.; Zhang, L.Y.; Xiong, W.; Zhou, M.; Xiao, L.; Zeng, F.; Li, X.L.; Wu, M.H.; et al. Grade-specific expression profiles of miRNAs/mRNAs and docking study in human grade I–III astrocytomas. OMICS 2011, 15, 673–682. [Google Scholar] [CrossRef] [PubMed]
- Liu, H.; Brannon, A.R.; Reddy, A.R.; Alexe, G.; Seiler, M.W.; Arreola, A.; Oza, J.H.; Yao, M.; Juan, D.; Liou, L.S.; et al. Identifying mRNA targets of microrna dysregulated in cancer: With application to clear cell renal cell carcinoma. BMC Syst. Biol. 2010, 4, e51. [Google Scholar] [CrossRef] [PubMed]
- Skalsky, R.L.; Cullen, B.R. Reduced expression of brain-enriched micrornas in glioblastomas permits targeted regulation of a cell death gene. PLoS ONE 2011, 6, e24248. [Google Scholar] [CrossRef] [PubMed]
- Zhang, L.; Volinia, S.; Bonome, T.; Calin, G.A.; Greshock, J.; Yang, N.; Liu, C.G.; Giannakakis, A.; Alexiou, P.; Hasegawa, K.; et al. Genomic and epigenetic alterations deregulate microrna expression in human epithelial ovarian cancer. Proc. Natl. Acad. Sci. USA 2008, 105, 7004–7009. [Google Scholar] [CrossRef] [PubMed]
- Cui, J.; Bi, M.; Overstreet, A.M.; Yang, Y.; Li, H.; Leng, Y.; Qian, K.; Huang, Q.; Zhang, C.; Lu, Z.; et al. MiR-873 regulates ERα transcriptional activity and tamoxifen resistance via targeting CDK3 in breast cancer cells. Oncogene 2014. [Google Scholar] [CrossRef] [PubMed]
- Vernin, C.; Thenoz, M.; Pinatel, C.; Gessain, A.; Gout, O.; Delfau-Larue, M.H.; Nazaret, N.; Legras-Lachuer, C.; Wattel, E.; Mortreux, F. HTLV-1 BZIP factor HBZ promotes cell proliferation and genetic instability by activating oncomirs. Cancer Res. 2014, 74, 6082–6093. [Google Scholar] [CrossRef] [PubMed]
- Sayed, D.; Rane, S.; Lypowy, J.; He, M.; Chen, I.Y.; Vashistha, H.; Yan, L.; Malhotra, A.; Vatner, D.; Abdellatif, M. MicroRNA-21 targets sprouty2 and promotes cellular outgrowths. Mol. Biol. Cell 2008, 19, 3272–3282. [Google Scholar] [CrossRef] [PubMed]
- Thum, T.; Gross, C.; Fiedler, J.; Fischer, T.; Kissler, S.; Bussen, M.; Galuppo, P.; Just, S.; Rottbauer, W.; Frantz, S.; et al. Microrna-21 contributes to myocardial disease by stimulating map kinase signalling in fibroblasts. Nature 2008, 456, 980–984. [Google Scholar] [CrossRef] [PubMed]
- Asangani, I.A.; Rasheed, S.A.; Nikolova, D.A.; Leupold, J.H.; Colburn, N.H.; Post, S.; Allgayer, H. MicroRNA-21 (miR-21) post-transcriptionally downregulates tumor suppressor PDCD4 and stimulates invasion, intravasation and metastasis in colorectal cancer. Oncogene 2008, 27, 2128–2136. [Google Scholar] [CrossRef] [PubMed]
- Meng, F.; Henson, R.; Wehbe-Janek, H.; Ghoshal, K.; Jacob, S.T.; Patel, T. Microrna-21 regulates expression of the pten tumor suppressor gene in human hepatocellular cancer. Gastroenterology 2007, 133, 647–658. [Google Scholar] [CrossRef] [PubMed]
- Zhu, S.; Si, M.L.; Wu, H.; Mo, Y.Y. Microrna-21 targets the tumor suppressor gene tropomyosin 1 (TPM1). J. Biol. Chem. 2007, 282, 14328–14336. [Google Scholar] [CrossRef] [PubMed]
- Volinia, S.; Calin, G.A.; Liu, C.G.; Ambs, S.; Cimmino, A.; Petrocca, F.; Visone, R.; Iorio, M.; Roldo, C.; Ferracin, M.; et al. A microrna expression signature of human solid tumors defines cancer gene targets. Proc. Natl. Acad. Sci. USA 2006, 103, 2257–2261. [Google Scholar] [CrossRef] [PubMed]
- O’Donnell, K.A.; Wentzel, E.A.; Zeller, K.I.; Dang, C.V.; Mendell, J.T. c-Myc -regulated microRNAs modulate E2F1 expression. Nature 2005, 435, 839–843. [Google Scholar] [CrossRef] [PubMed]
- Monzo, M.; Navarro, A.; Bandres, E.; Artells, R.; Moreno, I.; Gel, B.; Ibeas, R.; Moreno, J.; Martinez, F.; Diaz, T.; et al. Overlapping expression of micrornas in human embryonic colon and colorectal cancer. Cell Res. 2008, 18, 823–833. [Google Scholar] [CrossRef] [PubMed]
- Novotny, G.W.; Sonne, S.B.; Nielsen, J.E.; Jonstrup, S.P.; Hansen, M.A.; Skakkebaek, N.E.; Rajpert-de Meyts, E.; Kjems, J.; Leffers, H. Translational repression of e2f1 mrna in carcinoma in situ and normal testis correlates with expression of the mir-17–92 cluster. Cell Death Differ. 2007, 14, 879–882. [Google Scholar] [CrossRef] [PubMed]
- Xiao, C.; Srinivasan, L.; Calado, D.P.; Patterson, H.C.; Zhang, B.; Wang, J.; Henderson, J.M.; Kutok, J.L.; Rajewsky, K. Lymphoproliferative disease and autoimmunity in mice with increased miR-17–92 expression in lymphocytes. Nat. Immunol. 2008, 9, 405–414. [Google Scholar] [CrossRef] [PubMed]
- Gandhi, N.S.; Tekade, R.K.; Chougule, M.B. Nanocarrier mediated delivery of siRNA/miRNA in combination with chemotherapeutic agents for cancer therapy: Current progress and advances. J. Control. Release 2014, 194, 238–256. [Google Scholar] [CrossRef] [PubMed]
- Sethi, S.; Li, Y.; Sarkar, F.H. Regulating mirna by natural agents as a new strategy for cancer treatment. Curr. Drug Targets 2013, 14, 1167–1174. [Google Scholar] [CrossRef] [PubMed]
- Ling, H.; Fabbri, M.; Calin, G.A. Micrornas and other non-coding RNAs as targets for anticancer drug development. Nat. Rev. Drug Disc. 2013, 12, 847–865. [Google Scholar] [CrossRef] [PubMed]
© 2015 by the authors; licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Moles, R.; Nicot, C. The Emerging Role of miRNAs in HTLV-1 Infection and ATLL Pathogenesis. Viruses 2015, 7, 4047-4074. https://doi.org/10.3390/v7072805
Moles R, Nicot C. The Emerging Role of miRNAs in HTLV-1 Infection and ATLL Pathogenesis. Viruses. 2015; 7(7):4047-4074. https://doi.org/10.3390/v7072805
Chicago/Turabian StyleMoles, Ramona, and Christophe Nicot. 2015. "The Emerging Role of miRNAs in HTLV-1 Infection and ATLL Pathogenesis" Viruses 7, no. 7: 4047-4074. https://doi.org/10.3390/v7072805






