Occult HBV Infection: A Faceless Enemy in Liver Cancer Development
Abstract
:1. Introduction
2. The Hepatitis B Virus
3. Occult HBV Infection
| Author (Publication year) (Reference) | Population | n | Prevalence of anti–HBsAg | Prevalence of anti–HBc | Prevalence of occult HBV infection | Comments |
|---|---|---|---|---|---|---|
| Hemodialysis patients | ||||||
| Cabrerizo M, et al. (1997) [26] | Spain | 33 | 14 (42%) | 14 (42%) | 19 (58%) | |
| Besisik F, et al. (2003) [27] | Turkey | 33 | 12 (36.4%) | All patients had HCV infection. | ||
| Abu El Makarem MA, et al. (2012) [28] | Egypt | 145 | 15 (10.3%) | 29 (20%) | 6 (4.1%) | Patients with or without HCV infection were included. |
| Minuk GY, et al. (2004) [29] | Canada | 239 | 152 (63%) | 21 (8.7%) | 9 (3.8%) | |
| Albuquerque AC, et al. (2012) [30] | Brazil | 752 | 201 of 752 (26.7%) | 135 of 201 (67.2%) | 3 of 201 (1.5%)* | |
| Fabrizi F, et al. (2005) [31] | Italy | 213 | 120 of 316 (37.9%) † | 123 (57.7%) | 0 (0%) | Patients undergoing hemodialysis or peritoneal dialysis were included. |
| Goral V, et al. (2006) [32] | Turkey | 50 | 21 (42%) | 4 (8%) | 0 (0%) | |
| HIV+ patients | ||||||
| Hofer M, et al. (1998) [33] | Switzerland | 57 | 0 (0%) | 56 (98.2%) | 51 (89.5%) | Longitudinal observation |
| Marite B, et al. (2011) [34] | Cube | 325 | 45 of 99 (45.5) ‡ | 99 (30.5%) | 13 of 54 (24.1%) § | |
| Filippini, et al. (2006) [35] | Italy | 115 | 16 of 115 (13.9%) (baseline) | 58 of 115 (50.4%) (baseline) | 17 of 86 (19.8%) || | Longitudinal design |
| Panigrahi R, et al. (2012) [36] | India | 112 | 12 of 112 (10.7%) | |||
| Nunez M, et al. (2002) [37] | Spain | 85 | Not reported | Not reported | 0 (0%) |
4. Principal Molecular Mechanisms Associated with OBI
4.1. Genome Integration of HBV
4.2. Genetic Mutations in HBV
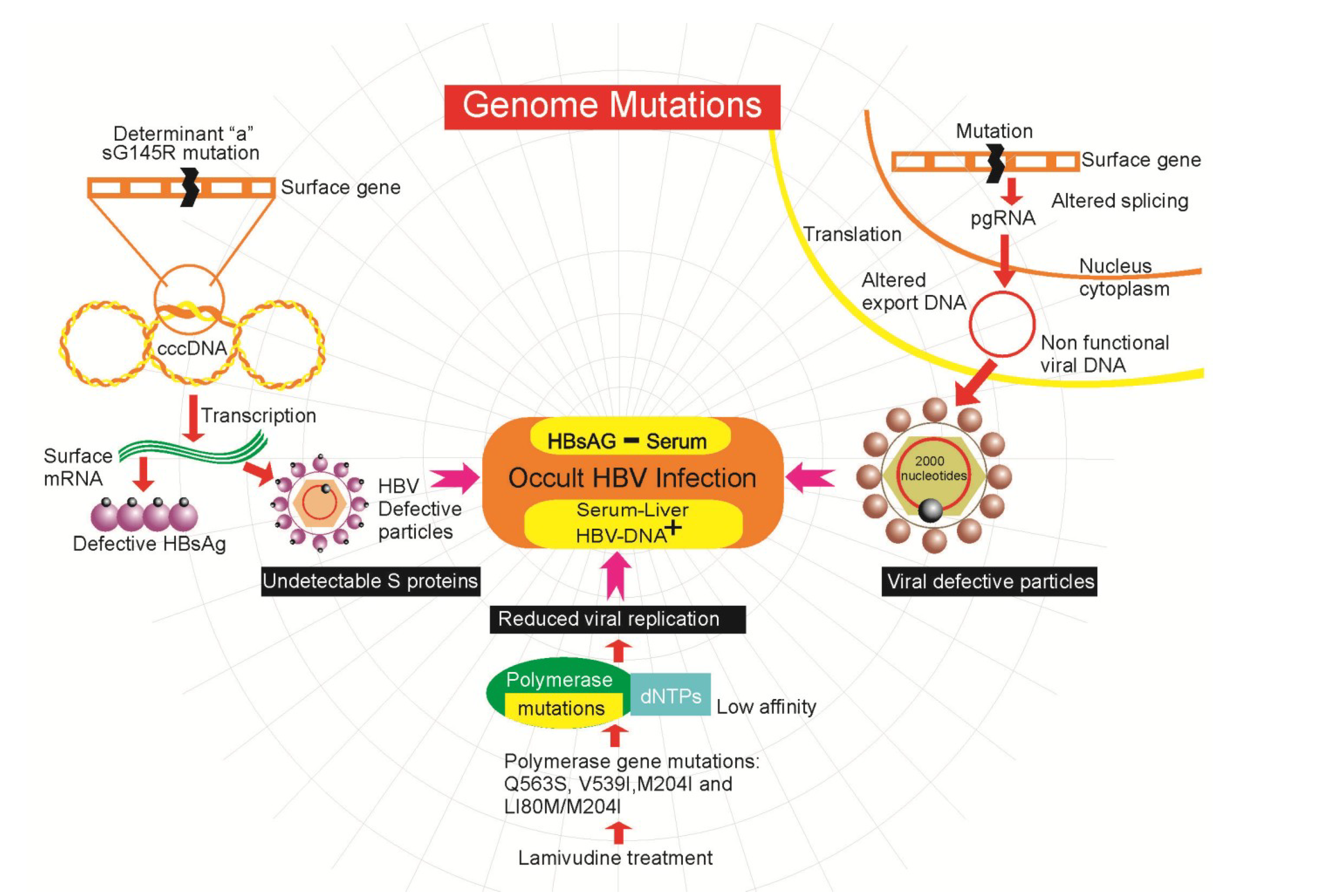
4.3. Epigenetics Mechanisms
4.4. Host Immune Response in OBI
4.5. Viral Co-Infection
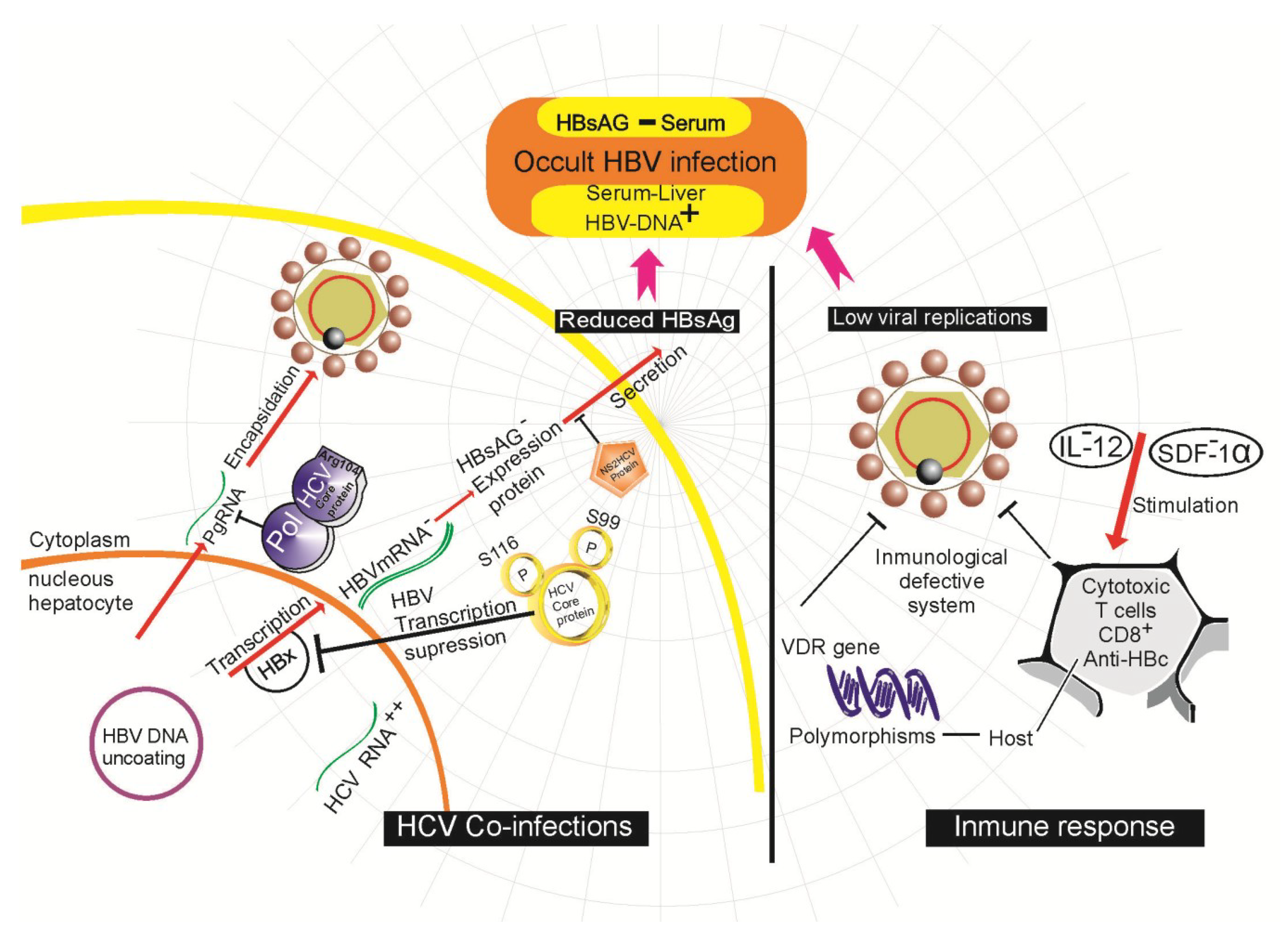
4.6. Other Mechanisms
5. The Influence of OBI on Development of HCC
6. Appropriate Diagnosis of OBI
7. Epidemiological Implications for Underdiagnosis of OBI in a Latin-American Region
Author Contributions
Conflicts of Interest
References and Notes
- Tan, Y.J. Hepatitis B virus infection and the risk of hepatocellular carcinoma. World J. Gastroenterol. (WJG) 2011, 17, 4853–4857. [Google Scholar] [CrossRef]
- Alves, R.C.; Alves, D.; Guz, B.; Matos, C.; Viana, M.; Harriz, M.; Terrabuio, D.; Kondo, M.; Gampel, O.; Polletti, P. Advanced hepatocellular carcinoma. Review of targeted molecular drugs. Ann. Hepatol. 2011, 10, 21–27. [Google Scholar]
- World Health Organization. World Hepatitis Day 2012. Available online: http://www.who.int/csr/disease/hepatitis/world_hepatitis_day/en/ (accessed on 10 March 2014).
- World Health Organization. Hepatitis B. (Fact Sheet No 204). Available online: http://www.who.int/csr/disease/hepatitis/world_hepatitis_day/en/ (accessed on 10 March 2014).
- Lavanchy, D.; Hepatitis, B. Virus epidemiology, disease burden, treatment, and current and emerging prevention and control measures. J. Viral Hepat 2004, 11, 97–107. [Google Scholar] [CrossRef]
- International Agency for Research on Cancer. Global battle against cancer won’t be won with treatment alone. Effective prevention measures urgently needed to prevent cancer crisis. (Press Release No. 224). 2014. Available online: http://www.who.int/csr/disease/hepatitis/world_hepatitis_day/en/ (accessed on 10 March 2014).
- Torre, A. Hepatopatía crónica y hepatocarcinoma. Rev. Gastroenterol. Méx. 2008, 73, 58–66. [Google Scholar]
- Aguirre, I; Fernández, J.; Fustea, L.; Torrasb, R. Estado actual del hepatocarcinoma y perspectivas futuras. Actualización 2010, 0, 1762. [Google Scholar]
- Valerio, J.; Vasquez, F.; Perez, J.; Cortazar-Benitez, L.F.; Chávez-Tapia, N.C.; Ruvalcaba-Rojas, O.A.; Torres-Medina, V.; Ocejo-Rodríguez, A. Prevalence of VHB and VHC serological markers among blood donors in the capital state of Veracruz, Mexico. Gac. Med. Mex. 2009, 145, 183–187. [Google Scholar]
- Bottcher, B.; Wynne, S.A.; Crowther, R.A. Determination of the fold of the core protein of hepatitis B virus by electron cryomicroscopy. Nature 1997, 386, 88–91. [Google Scholar] [CrossRef]
- Quarleri, J. Core promoter: A critical region where the hepatitis B virus makes decisions. World J. Gastroenterol. (WJG) 2014, 20, 425–435. [Google Scholar] [CrossRef]
- Leung, N. Treatment of HBeAg-positive chronic hepatitis B with nucleos(t)ide analogues. Liver Int. 2011, 31, 85–89. [Google Scholar] [CrossRef]
- Bruss, V. A short linear sequence in the pre-S domain of the large hepatitis B virus envelope protein required for virion formation. J. Virol. 1997, 71, 9350–9357. [Google Scholar]
- Qin, S.; Tang, H.; Zhao, L.S.; He, F.; Lin, Y.; Liu, L.; He, X.M. Cloning of HBsAg-encoded genes in different vectors and their expression in eukaryotic cells. World J. Gastroenterol. (WJG) 2003, 9, 1111–1113. [Google Scholar]
- Persing, D.H.; Varmus, H.E.; Ganem, D. The preS1 protein of hepatitis B virus is acylated at its amino terminus with myristic acid. J. Virol. 1987, 61, 1672–1677. [Google Scholar]
- Krugman, S. The newly licensed hepatitis B vaccine. Characteristics and indications for use. J. Am. Med. Assoc. 1982, 247, 2012–2015. [Google Scholar] [CrossRef]
- Wu, J.Y.; Newton, S.; Judd, A.; Stocker, B.; Robinson, W.S. Expression of immunogenic epitopes of hepatitis B surface antigen with hybrid flagellin proteins by a vaccine strain of Salmonella. Proc. Natl. Acad. Sci. USA 1989, 86, 4726–4730. [Google Scholar] [CrossRef]
- Milich, D.R. Genetic and molecular basis for T- and B-cell recognition of hepatitis B viral antigens. Immunol. Rev. 1987, 99, 71–103. [Google Scholar] [CrossRef]
- Milich, D.R. T- and B-cell recognition of hepatitis B viral antigens. Immunol. Today 1988, 9, 380–386. [Google Scholar] [CrossRef]
- Hu, W.G.; Wei, J.; Xia, H.C.; Yang, X.X.; Li, F.; Li, G.D.; Wang, Y.; Zhang, Z.C. Identification of the immunogenic domains in HBsAg preS1 region using overlapping preS1 fragment fusion proteins. World J. Gastroenterol. (WJG) 2005, 11, 2088–2094. [Google Scholar]
- Madalinski, K.; Sylvan, S.P.; Hellstrom, U.; Mikolajewicz, J.; Dzierzanowska-Fangrat, K. Presence of anti-preS1, anti-preS2, and anti-HBs antibodies in newborns immunized with Bio-Hep-B vaccine. Med. Sci. Monit. 2004, 10, PI10–PI17. [Google Scholar]
- Ge, G.; Wang, S.; Han, Y.; Zhang, C.; Lu, S.; Huang, Z. Removing N-terminal sequences in pre-S1 domain enhanced antibody and B-cell responses by an HBV large surface antigen DNA vaccine. PLoS One 2012, 7, e41573. [Google Scholar]
- Song, le H.; Xuan, N.T.; Toan, N.L.; Binh, V.Q.; Boldt, A.B.; Kremsner, P.G.; Kun, J.F. Association of two variants of the interferon-alpha receptor-1 gene with the presentation of hepatitis B virus infection. Eur. Cytokine Netw. 2008, 19, 204–210. [Google Scholar]
- Raimondo, G.; Allain, J.P.; Brunetto, M.R.; Buendia, M.A.; Chen, D.S.; Colombo, M.; Craxì, A.; Donato, F.; Ferrari, C.; Gaeta, G.B.; et al. Statements from the Taormina expert meeting on occult hepatitis B virus infection. J. Hepatol. 2008, 49, 652–657. [Google Scholar] [CrossRef]
- Larrubia, J.R. Occult hepatitis B virus infection: A complex entity with relevant clinical implications. World J. Gastroenterol. (WJG) 2011, 17, 1529–1530. [Google Scholar] [CrossRef]
- Cabrerizo, M.; Bartolome, J.; De Sequera, P.; Caramelo, C.; Carreno, V. Hepatitis B virus DNA in serum and blood cells of hepatitis B surface antigen-negative hemodialysis patients and staff. J. Am. Soc. Nephrol. (JASN) 1997, 8, 1443–1447. [Google Scholar]
- Besisik, F.; Karaca, C.; Akyuz, F.; Horosanli, S.; Onel, D.; Badur, S.; Sever, M.S.; Danalioglu, A.; Demir, K.; Kaymakoglu, S.; et al. Occult HBV infection and YMDD variants in hemodialysis patients with chronic HCV infection. J. Hepatol. 2003, 38, 506–510. [Google Scholar]
- Abu El Makarem, M.A.; Abdel Hamid, M.; Abdel Aleem, A.; Ali, A.; Shatat, M.; Sayed, D.; Deaf, A.; Hamdy, L.; Tony, E.A. Prevalence of occult hepatitis B virus infection in hemodialysis patients from egypt with or without hepatitis C virus infection. Hepat. Mon. 2012, 12, 253–258. [Google Scholar]
- Minuk, G.Y.; Sun, D.F.; Greenberg, R.; Zhang, M.; Hawkins, K.; Uhanova, J.; Gutkin, A.; Bernstein, K.; Giulivi, A.; Osiowy, C. Occult hepatitis B virus infection in a North American adult hemodialysis patient population. J. Hepatol. 2004, 40, 1072–1077. [Google Scholar] [CrossRef]
- Albuquerque, A.C.; Coelho, M.R.; Lemos, M.F.; Moreira, R.C. Occult Hepatitis B Virus Infection in Hemodialysis Patients in Recife; Revista da Sociedade Brasileira de Medicina Tropical: State of Pernambuco, Brazil, 2012; Volume 45, pp. 558–562. [Google Scholar]
- Fabrizi, F.; Messa, P.G.; Lunghi, G.; Aucella, F.; Bisegna, S.; Mangano, S.; Villa, M.; Barbisoni, F.; Rusconi, E.; Martin, P. Occult hepatitis B virus infection in dialysis patients: A multicentre survey. Aliment. Pharmacol. Ther. 2005, 21, 1341–1347. [Google Scholar] [CrossRef]
- Goral, V.; Ozkul, H.; Tekes, S.; Sit, D.; Kadiroglu, A.K. Prevalence of occult HBV infection in haemodialysis patients with chronic HCV. World J. Gastroenterol. (WJG) 2006, 12, 3420–3424. [Google Scholar]
- Hofer, M.; Joller-Jemelka, H.I.; Grob, P.J.; Luthy, R.; Opravil, M. Frequent chronic hepatitis B virus infection in HIV-infected patients positive for antibody to hepatitis B core antigen only. Swiss HIV Cohort Study. Eur. J. Clin. Microbiol. Infect. Dis. 1998, 17, 6–13. [Google Scholar] [CrossRef]
- Marite, B.; Montalvo, M.C.; Rodriguez, L.de L.; Sariego, S.; Verdasquera, D.; Vincent, M.; Gutiérrez, A.; Sánchez, M. Occult hepatitis B in Cuban HIV patients. MEDICC Rev. 2011, 13, 32–37. [Google Scholar]
- Filippini, P.; Coppola, N.; Pisapia, R.; Scolastico, C.; Marrocco, C.; Zaccariello, A.; Nacca, C.; Sagnelli, C.; de Stefano, G.; Ferraro, T.; et al. Impact of occult hepatitis B virus infection in HIV patients naive for antiretroviral therapy. Aids 2006, 20, 1253–1260. [Google Scholar] [CrossRef]
- Panigrahi, R.; Majumder, S.; Gooptu, M.; Biswas, A.; Datta, S.; Chandra, P.K.; Banerjee, A.; Chakrabarti, S.; Bandopadhyay, D.; De, B.K.; et al. Occult HBV infection among anti-HBc positive HIV-infected patients in apex referral centre, Eastern India. Ann. Hepatol. 2012, 11, 870–875. [Google Scholar]
- Nunez, M.; Rios, P.; Perez-Olmeda, M.; Soriano, V. Lack of ‘occult’ hepatitis B virus infection in HIV-infected patients. Aids 2002, 16, 2099–2101. [Google Scholar] [CrossRef]
- Hollinger, F.B.; Hepatitis, B. Virus infection and transfusion medicine: Science and the occult. Transfusion 2008, 48, 1001–1026. [Google Scholar] [CrossRef]
- Fang, Y.; Shang, Q.L.; Liu, J.Y.; Li, D.; Xu, W.Z.; Teng, X.; Zhao, H.W.; Fu, L.J.; Zhang, F.M.; Gu, H.X. Prevalence of occult hepatitis B virus infection among hepatopathy patients and healthy people in China. J. Infect. 2009, 58, 383–388. [Google Scholar] [CrossRef]
- Raimondo, G.; Pollicino, T.; Romano, L.; Zanetti, A.R. A 2010 update on occult hepatitis B infection. Pathol.-Biol. 2010, 58, 254–257. [Google Scholar] [CrossRef]
- Cacciola, I.; Pollicino, T.; Squadrito, G.; Cerenzia, G.; Orlando, M.E.; Raimondo, G. Occult hepatitis B virus infection in patients with chronic hepatitis C liver disease. N. Eng. J. Med. 1999, 341, 22–26. [Google Scholar] [CrossRef]
- Akarsu, M.; Kantar, F.U.; Sayiner, A.A. Occult hepatitis B: Evolving challenges and new perspectives. Hepat. Mon. 2011, 11, 475–476. [Google Scholar]
- Raimondo, G.; Navarra, G.; Mondello, S.; Costantino, L.; Colloredo, G.; Cucinotta, E.; di Vita, G.; Scisca, C.; Squadrito, G.; Pollicino, T. Occult hepatitis B virus in liver tissue of individuals without hepatic disease. J. Hepatol. 2008, 48, 743–746. [Google Scholar] [CrossRef]
- Zerbini, A.; Pilli, M.; Boni, C.; Penna, A.; di Vincenzo, P.; Giuberti, T.; Orlandini, A.; Raffa, G.; Pollicino, T.; Raimondo, G.; et al. The characteristics of the cell-mediated immune response identify different profiles of occult hepatitis B virus infection. Gastroenterology 2008, 134, 1470–1481. [Google Scholar] [CrossRef]
- Fang, Y.; Teng, X.; Xu, W.Z.; Li, D.; Zhao, H.W.; Fu, L.J.; Zhang, F.M.; Gu, H.X. Molecular characterization and functional analysis of occult hepatitis B virus infection in Chinese patients infected with genotype C. J. Med. Virol. 2009, 81, 826–835. [Google Scholar] [CrossRef]
- Hollinger, F.B.; Sood, G. Occult hepatitis B virus infection: A covert operation. J. Viral Hepat. 2010, 17, 1–15. [Google Scholar] [CrossRef]
- Mason, A.L.; Xu, L.; Guo, L.; Kuhns, M.; Perrillo, R.P. Molecular basis for persistent hepatitis B virus infection in the liver after clearance of serum hepatitis B surface antigen. Hepatology 1998, 27, 1736–1742. [Google Scholar] [CrossRef]
- Laras, A.; Koskinas, J.; Dimou, E.; Kostamena, A.; Hadziyannis, S.J. Intrahepatic levels and replicative activity of covalently closed circular hepatitis B virus DNA in chronically infected patients. Hepatology 2006, 44, 694–702. [Google Scholar]
- Bock, C.T.; Schwinn, S.; Locarnini, S.; Fyfe, J.; Manns, M.P.; Trautwein, C.; Zentgraf, H. Structural organization of the hepatitis B virus minichromosome. J. Mol. Biol. 2001, 307, 183–196. [Google Scholar] [CrossRef]
- Zoulim, F. New insight on hepatitis B virus persistence from the study of intrahepatic viral cccDNA. J. Hepatol. 2005, 42, 302–308. [Google Scholar] [CrossRef]
- Raimondo, G.; Burk, R.D.; Lieberman, H.M.; Muschel, J.; Hadziyannis, S.J.; Will, H.; Kew, M.C.; Dusheiko, G.M.; Shafritz, D.A. Interrupted replication of hepatitis B virus in liver tissue of HBsAg carriers with hepatocellular carcinoma. Virology 1988, 166, 103–112. [Google Scholar] [CrossRef]
- Urashima, T.; Saigo, K.; Kobayashi, S.; Imaseki, H.; Matsubara, H.; Koide, Y.; Asano, T.; Kondo, Y.; Koike, K.; Isono, K. Identification of hepatitis B virus integration in hepatitis C virus-infected hepatocellular carcinoma tissues. J. Hepat. 1997, 26, 771–778. [Google Scholar]
- Samal, J.; Kandpal, M.; Vivekanandan, P. Molecular mechanisms underlying occult hepatitis B virus infection. Clin. Microbiol. Rev. 2012, 25, 142–163. [Google Scholar] [CrossRef]
- De la Fuente, R.A.; Gutierrez, M.L.; Garcia-Samaniego, J.; Fernandez-Rodriguez, C.; Lledo, J.L.; Castellano, G. Pathogenesis of occult chronic hepatitis B virus infection. World J. Gastroenterol. (WJG) 2011, 17, 1543–1548. [Google Scholar] [CrossRef]
- Candotti, D.; Lin, C.K.; Belkhiri, D.; Sakuldamrongpanich, T.; Biswas, S.; Lin, S.; Teo, D.; Ayob, Y.; Allain, J.P. Occult hepatitis B infection in blood donors from South East Asia: Molecular characterisation and potential mechanisms of occurrence. Gut 2012, 61, 1744–1753. [Google Scholar] [CrossRef]
- Biswas, S.; Candotti, D.; Allain, J.P. Specific amino acid substitutions in the S protein prevent its excretion in vitro and may contribute to occult hepatitis B virus infection. J. Virol. 2013, 87, 7882–7892. [Google Scholar] [CrossRef]
- Muroyama, R.; Kato, N.; Yoshida, H.; Otsuka, M.; Moriyama, M.; Wang, Y.; Shao, R.X.; Dharel, N.; Tanaka, Y.; Ohta, M.; et al. Nucleotide change of codon 38 in the X gene of hepatitis B virus genotype C is associated with an increased risk of hepatocellular carcinoma. J. Hepatol. 2006, 45, 805–812. [Google Scholar] [CrossRef]
- Yuan, Q.; Ou, S.H.; Chen, C.R.; Ge, S.X.; Pei, B.; Chen, Q.R.; Yan, Q.; Lin, Y.C.; Ni, H.Y.; Huang, C.H.; et al. Molecular characteristics of occult hepatitis B virus from blood donors in southeast China. J. Clin. Microbiol. 2010, 48, 357–362. [Google Scholar] [CrossRef]
- Gaillard, R.K.; Barnard, J.; Lopez, V.; Hodges, P.; Bourne, E.; Johnson, L.; Allen, M.I.; Condreay, P.; Miller, W.H.; Condreay, L.D. Kinetic analysis of wild-type and YMDD mutant hepatitis B virus polymerases and effects of deoxyribonucleotide concentrations on polymerase activity. Antimicrob. Agents Chemother. 2002, 46, 1005–1013. [Google Scholar] [CrossRef]
- Bowden, S.; Bartholomeusz, A.; Locarnini, S. Lamivudine resistant occult HBV: Implications for public health? J. Hepatol. 2003, 38, 526–528. [Google Scholar]
- Torresi, J.; Earnest-Silveira, L.; Deliyannis, G.; Edgtton, K.; Zhuang, H.; Locarnini, S.A.; Fyfe, J.; Sozzi, T.; Jackson, D.C. Reduced antigenicity of the hepatitis B virus HBsAg protein arising as a consequence of sequence changes in the overlapping polymerase gene that are selected by lamivudine therapy. Virology 2002, 293, 305–313. [Google Scholar] [CrossRef]
- Hass, M.; Hannoun, C.; Kalinina, T.; Sommer, G.; Manegold, C.; Gunther, S. Functional analysis of hepatitis B virus reactivating in hepatitis B surface antigen-negative individuals. Hepatology 2005, 42, 93–103. [Google Scholar]
- Gunther, S.; Sommer, G.; Iwanska, A.; Will, H. Heterogeneity and common features of defective hepatitis B virus genomes derived from spliced pregenomic RNA. Virology 1997, 238, 363–371. [Google Scholar] [CrossRef]
- Van Hemert, F.J.; Zaaijer, H.L.; Berkhout, B.; Lukashov, V.V. Occult hepatitis B infection: An evolutionary scenario. Virol. J. 2008, 5, 146. [Google Scholar] [CrossRef]
- Portela, A.; Esteller, M. Epigenetic modifications and human disease. Nature Biotechnol. 2010, 28, 1057–1068. [Google Scholar] [CrossRef]
- Vivekanandan, P.; Thomas, D.; Torbenson, M. Methylation regulates hepatitis B viral protein expression. J. Infect. Dis. 2009, 199, 1286–1291. [Google Scholar] [CrossRef]
- Vivekanandan, P.; Thomas, D.; Torbenson, M. Hepatitis B viral DNA is methylated in liver tissues. J. Viral Hepat. 2008, 15, 103–107. [Google Scholar]
- Kaur, P.; Paliwal, A.; Durantel, D.; Hainaut, P.; Scoazec, J.Y.; Zoulim, F.; Chemin, I.; Herceg, Z. DNA methylation of hepatitis B virus (HBV) genome associated with the development of hepatocellular carcinoma and occult HBV infection. J. Infect. Dis. 2010, 202, 700–704. [Google Scholar] [CrossRef]
- Belloni, L.; Pollicino, T.; De Nicola, F.; Guerrieri, F.; Raffa, G.; Fanciulli, M.; Raimondo, G.; Levrero, M. Nuclear HBx binds the HBV minichromosome and modifies the epigenetic regulation of cccDNA function. Proc. Natl. Acad. Sci. USA 2009, 106, 19975–19979. [Google Scholar] [CrossRef]
- Said, Z.N. An overview of occult hepatitis B virus infection. World J. Gastroenterol. (WJG) 2011, 17, 1927–1938. [Google Scholar] [CrossRef]
- Ceneli, O.; Ozkurt, Z.N.; Acar, K.; Rota, S.; Aki, S.Z.; Yegin, Z.A.; Yagci, M.; Ozenirler, S.; Sucak, G.T. Hepatitis B-related events in autologous hematopoietic stem cell transplantation recipients. World J. Gastroenterol. (WJG) 2010, 16, 1765–1771. [Google Scholar] [CrossRef]
- Fierro, N.A.; Roman, S.; Realpe, M.; Hernandez-Nazara, Z.; Zepeda-Carrillo, E.A.; Panduro, A. Multiple cytokine expression profiles reveal immune-based differences in occult hepatitis B genotype H-infected Mexican Nahua patients. Mem. Do Inst. Oswaldo Cruz 2011, 106, 1007–1013. [Google Scholar]
- Arababadi, M.K.; Pourfathollah, A.A.; Jafarzadeh, A.; Hassanshahi, G. Serum Levels of IL-10 and IL-17A in Occult HBV-Infected South-East Iranian Patients. Hepat. Mon. 2010, 10, 31–35. [Google Scholar]
- Arababadi, M.K.; Pourfathollah, A.A.; Jafarzadeh, A.; Hassanshahi, G.; Daneshmandi, S.; Shamsizadeh, A.; Kennedy, D. Non-association of IL-12 +1188 and IFN-gamma +874 polymorphisms with cytokines serum level in occult HBV infected patients. Saudi J. Gastroenterol. 2011, 17, 30–35. [Google Scholar] [CrossRef]
- Hassanshahi, G.; Arababadi, M.K.; Khoramdelazad, H.; Yaghini, N.; Zarandi, E.R. Assessment of CXCL12 (SDF-1alpha) polymorphisms and its serum level in posttransfusion occult HBV-infected patients in Southeastern Iran. Arch. Med. Res. 2010, 41, 338–342. [Google Scholar] [CrossRef]
- Arababadi, M.K.; Nasiri Ahmadabadi, B.; Kennedy, D. Current information on the immunologic status of occult hepatitis B infection. Transfusion 2012, 52, 1819–1826. [Google Scholar] [CrossRef]
- Sheen, I.S.; Liaw, Y.F.; Lin, D.Y.; Chu, C.M. Role of hepatitis C and delta viruses in the termination of chronic hepatitis B surface antigen carrier state: A multivariate analysis in a longitudinal follow-up study. J. Infect. Dis. 1994, 170, 358–361. [Google Scholar] [CrossRef]
- Mimms, L.T.; Mosley, J.W.; Hollinger, F.B.; Aach, R.D.; Stevens, C.E.; Cunningham, M.; Vallari, D.V.; Barbosa, L.H.; Nemo, G.J. Effect of concurrent acute infection with hepatitis C virus on acute hepatitis B virus infection. BMJ 1993, 307, 1095–1097. [Google Scholar] [CrossRef]
- Rodriguez-Inigo, E.; Bartolome, J.; Ortiz-Movilla, N.; Platero, C.; Lopez-Alcorocho, J.M.; Pardo, M.; Castillo, I.; Carreno, V. Hepatitis C virus (HCV) and hepatitis B virus (HBV) can coinfect the same hepatocyte in the liver of patients with chronic HCV and occult HBV infection. J. Virol. 2005, 79, 15578–15581. [Google Scholar] [CrossRef]
- Chen, S.Y.; Kao, C.F.; Chen, C.M.; Shih, C.M.; Hsu, M.J.; Chao, C.H.; Wang, S.H.; You, L.R.; Lee, Y.H. Mechanisms for inhibition of hepatitis B virus gene expression and replication by hepatitis C virus core protein. J. Biol. Chem. 2003, 278, 591–607. [Google Scholar]
- Dumoulin, F.L.; von dem Bussche, A.; Li, J.; Khamzina, L.; Wands, J.R.; Sauerbruch, T.; Spengler, U. Hepatitis C virus NS2 protein inhibits gene expression from different cellular and viral promoters in hepatic and nonhepatic cell lines. Virology 2003, 305, 260–266. [Google Scholar] [CrossRef]
- Huang, Y.W.; Liao, Y.T.; Chen, W.; Chen, C.L.; Hu, J.T.; Liu, C.J.; Lai, M.Y.; Chen, P.J.; Chen, D.S.; Yang, S.S.; et al. Vitamin D receptor gene polymorphisms and distinct clinical phenotypes of hepatitis B carriers in Taiwan. Genes Immun. 2010, 11, 87–93. [Google Scholar] [CrossRef]
- Arababadi, M.K.; Pourfathollah, A.A.; Jafarzadeh, A.; Hassanshahi, G.; Rezvani, M.E. Association of exon 9 but not intron 8 VDR polymorphisms with occult HBV infection in south-eastern Iranian patients. J. Gastroenterol. Hepatol. 2010, 25, 90–93. [Google Scholar] [CrossRef]
- Shi, Y.; Wu, Y.H.; Wu, W.; Zhang, W.J.; Yang, J.; Chen, Z. Association between occult hepatitis B infection and the risk of hepatocellular carcinoma: A meta-analysis. Liver Int. 2012, 32, 231–240. [Google Scholar] [CrossRef]
- Shire, A.M.; Roberts, L.R. Occult hepatitis B virus infection: Bit player or role player? Hepatology 2011, 54, 760–763. [Google Scholar] [CrossRef]
- Ikeda, K.; Kobayashi, M.; Someya, T.; Saitoh, S.; Hosaka, T.; Akuta, N.; Suzuki, F.; Suzuki, Y.; Arase, Y.; Kumada, H. Occult hepatitis B virus infection increases hepatocellular carcinogenesis by eight times in patients with non-B, non-C liver cirrhosis: A cohort study. J. Viral Hepat. 2009, 16, 437–443. [Google Scholar] [CrossRef]
- Paterlini, P.; Driss, F.; Nalpas, B.; Pisi, E.; Franco, D.; Berthelot, P.; Brechot, C. Persistence of hepatitis B and hepatitis C viral genomes in primary liver cancers from HBsAg-negative patients: A study of a low-endemic area. Hepatology 1993, 17, 20–29. [Google Scholar]
- Matsuoka, S.; Nirei, K.; Tamura, A.; Nakamura, H.; Matsumura, H.; Oshiro, S.; Arakawa, Y.; Yamagami, H.; Tanaka, N.; Moriyama, M. Influence of occult hepatitis B virus coinfection on the incidence of fibrosis and hepatocellular carcinoma in chronic hepatitis C. Intervirology 2008, 51, 352–361. [Google Scholar] [CrossRef]
- Stroffolini, T.; Almasio, P.L.; Persico, M.; Bollani, S.; Benvegnu, L.; Di Costanzo, G.; Pastore, G.; Aghemo, A.; Stornaiuolo, G.; Mangia, A.; et al. Lack of correlation between serum anti-HBcore detectability and hepatocellular carcinoma in patients with HCV-related cirrhosis. Am. J. Gastroenterol. 2008, 103, 1966–1972. [Google Scholar] [CrossRef]
- Chu, C.J.; Lee, S.D. Hepatitis B virus/hepatitis C virus coinfection: Epidemiology, clinical features, viral interactions and treatment. J. Gastroenterol. Hepatol. 2008, 23, 512–520. [Google Scholar] [CrossRef]
- Wong, D.K.; Huang, F.Y.; Lai, C.L.; Poon, R.T.; Seto, W.K.; Fung, J.; Hung, I.F.; Yuen, M.F. Occult hepatitis B infection and HBV replicative activity in patients with cryptogenic cause of hepatocellular carcinoma. Hepatology 2011, 54, 829–836. [Google Scholar] [CrossRef]
- Chen, C.H.; Changchien, C.S.; Lee, C.M.; Tung, W.C.; Hung, C.H.; Hu, T.H.; Wang, J.H.; Wang, J.C.; Lu, S.N. A study on sequence variations in pre-S/surface, X and enhancer II/core promoter/precore regions of occult hepatitis B virus in non-B, non-C hepatocellular carcinoma patients in Taiwan. Int. J. Cancer 2009, 125, 621–629. [Google Scholar] [CrossRef]
- Ding, X.; Park, Y.N.; Taltavull, T.C.; Thung, S.N.; Jin, X.; Jin, Y.; Trung, N.S.; Edamoto, Y.; Sata, T.; Abe, K. Geographic characterization of hepatitis virus infections, genotyping of hepatitis B virus, and p53 mutation in hepatocellular carcinoma analyzed by in situ detection of viral genomes from carcinoma tissues: Comparison among six different countries. Jpn. J. Infect. Dis. 2003, 56, 12–18. [Google Scholar]
- Sorrell, M.F.; Belongia, E.A.; Costa, J.; Gareen, I.F.; Grem, J.L.; Inadomi, J.M.; Kern, E.R.; McHugh, J.A.; Petersen, G.M.; Rein, M.F.; et al. National Institutes of Health consensus development conference statement: Management of hepatitis B. Hepatology 2009, 49, S4–S12. [Google Scholar] [CrossRef]
- Madejon, A.; Manzano, M.L.; Arocena, C.; Castillo, I.; Carreno, V. Effects of delayed freezing of liver biopsies on the detection of hepatitis C virus RNA strands. J. Hepatol. 2000, 32, 1019–1025. [Google Scholar] [CrossRef]
- Baylis, S.A.; Heath, A.B.; Chudy, M.; Pisani, G.; Klotz, A.; Kerby, S.; Gerlich, W. An international collaborative study to establish the 2nd World Health Organization International Standard for hepatitis B virus DNA nucleic acid amplification technology-based assays. Vox Sang. 2008, 94, 358–362. [Google Scholar] [CrossRef]
- Raimondo, G.; Pollicino, T.; Levrero, M.; Craxi, A. Occult hepatitis B virus infection and hepatocellular carcinoma development in patients with chronic hepatitis C. Hepatology 2011, 54, 373–374, author reply 374. [Google Scholar]
- Arababadi, M.K.; Hassanshahi, G.; Pourfathollah, A.A.; Zarandi, E.R.; Kennedy, D. Post-transfusion occult hepatitis B (OBI): A global challenge for blood recipients and health authorities. Hepatit. Mon. 2011, 11, 714–718. [Google Scholar] [CrossRef]
- Roman, S.; Panduro, A.; Aguilar-Gutierrez, Y.; Maldonado, M.; Vazquez-Vandyck, M.; Martinez-Lopez, E.; Ruiz-Madrigal, B.; Hernandez-Nazara, Z. A low steady HBsAg seroprevalence is associated with a low incidence of HBV-related liver cirrhosis and hepatocellular carcinoma in Mexico: A systematic review. Hepatol. Int. 2009, 3, 343–355. [Google Scholar] [CrossRef]
- Silveira, T.R.; da Fonseca, J.C.; Rivera, L.; Fay, O.H.; Tapia, R.; Santos, J.I.; Urdeneta, E.; Clemens, S.A. Hepatitis B seroprevalence in Latin America. Pan Am. J. Public Health 1999, 6, 378–383. [Google Scholar]
- Valdespino, J.L.; Conde-González, C.J.; Olaiz-Fernández, G.; Palma, O.; Sepulveda, J. Prevalence of hepatitis B infection and carrier status among adults in Mexico. Salud publica de Mexico. 2007, 49, S404–S411. [Google Scholar] [CrossRef]
- Valerio-Urena, J.; Vasquez-Fernandez, F.; Perez-Sosa, J.A.; Cortazar-Benitez, L.F.; Chavez-Tapia, N.C.; Ruvalcaba-Rojas, O.A.; Torres-Medina, V.; Ocejo-Rodriguez, A. Prevalence of VHB and VHC serological markers among blood donors in the capital state of Veracruz, Mexico. Gac. Med. de Mex. 2009, 145, 183–187. [Google Scholar]
- Panduro, A.; Escobedo Melendez, G.; Fierro, N.A.; Ruiz Madrigal, B.; Zepeda-Carrillo, E.A.; Roman, S. Epidemiology of viral hepatitis in Mexico. Salud publica de Mexico. 2011, 53, S37–S45. [Google Scholar]
- Cusumano, A.M.; Gonzalez Bedat, M.C.; Garcia-Garcia, G.; Maury Fernandez, S.; Lugon, J.R.; Poblete Badal, H.; Elgueta Miranda, S.; Gomez, R.; Cerdas Calderon, M.; Almaguer Lopez, M.; et al. Latin American Dialysis and Renal Transplant Registry: 2008 report (data 2006). Clin. Nephrol. 2010, 74, S3–S8. [Google Scholar]
- Valdespino, J.L.; García-García, M.d.L.; Conde-González, C.J.; Olaiz-Fernández, G.; Palma, O.; Sepúlveda, J. Prevalencia de infección por VIH en la población adulta en México: Una epidemia en ascenso y expansión. Salud publica de Mexico. 2007, 49, S386–S394. [Google Scholar] [CrossRef]
- Tramuto, F.; Maida, C.M.; Colomba, G.M.; Di Carlo, P.; Vitale, F. Prevalence of occult hepatitis B virus infection in a cohort of HIV-positive patients resident in Sicily, Italy. Biomed. Res. Int. 2013, 859583. [Google Scholar]
- Costantini, A.; Marinelli, K.; Biagioni, G.; Monachetti, A.; Ferreri, M.L.; Butini, L.; Montroni, M.; Manzin, A.; Bagnarelli, P. Molecular analysis of hepatitis B virus (HBV) in an HIV co-infected patient with reactivation of occult HBV infection following discontinuation of lamivudine-including antiretroviral therapy. BMC Infect. Dis. 2011, 11, 310. [Google Scholar] [CrossRef]
- Roman, S.; Tanaka, Y.; Khan, A.; Kurbanov, F.; Kato, H.; Mizokami, M.; Panduro, A. Occult hepatitis B in the genotype H-infected Nahuas and Huichol native Mexican population. J. Med. Virol. 2010, 82, 1527–1536. [Google Scholar]
- Garcia, B.M.; Farfan, J.A.; Acosta, K.Y.; Puerto, F.I. Hepatitis B virus DNA in blood donors with anti-HBc as a possible indicator of active hepatitis B virus infection in Yucatan, Mexico. Transfus. Med. 2005, 15, 371–378. [Google Scholar] [CrossRef]
- Garcia-Montalvo, B.M.; Ventura-Zapata, L.P. Molecular and serological characterization of occult hepatitis B infection in blood donors from Mexico. Ann. Hepatol. 2011, 10, 133–141. [Google Scholar]
- Yuan, Q.; Ge, S.; Xiong, J.; Yan, Q.; Li, Z.; Hao, X.; Tian, D.; Niu, J.; Su, Z.; Chen, C.; Shih, J.W.; Zhang, J.; Xia, N. A novel immunoassay for PreS1 and/or core-related antigens for detection of HBsAg variants. J. Virol. Methods 2010, 168, 108–113. [Google Scholar] [CrossRef]
© 2014 by the authors; licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution license (http://creativecommons.org/licenses/by/3.0/).
Share and Cite
Morales-Romero, J.; Vargas, G.; García-Román, R. Occult HBV Infection: A Faceless Enemy in Liver Cancer Development. Viruses 2014, 6, 1590-1611. https://doi.org/10.3390/v6041590
Morales-Romero J, Vargas G, García-Román R. Occult HBV Infection: A Faceless Enemy in Liver Cancer Development. Viruses. 2014; 6(4):1590-1611. https://doi.org/10.3390/v6041590
Chicago/Turabian StyleMorales-Romero, Jaime, Gustavo Vargas, and Rebeca García-Román. 2014. "Occult HBV Infection: A Faceless Enemy in Liver Cancer Development" Viruses 6, no. 4: 1590-1611. https://doi.org/10.3390/v6041590
APA StyleMorales-Romero, J., Vargas, G., & García-Román, R. (2014). Occult HBV Infection: A Faceless Enemy in Liver Cancer Development. Viruses, 6(4), 1590-1611. https://doi.org/10.3390/v6041590




