Nuclear Imprisonment: Viral Strategies to Arrest Host mRNA Nuclear Export
Abstract
:1. Introduction
2. Biogenesis and Nuclear Export of Cellular mRNA
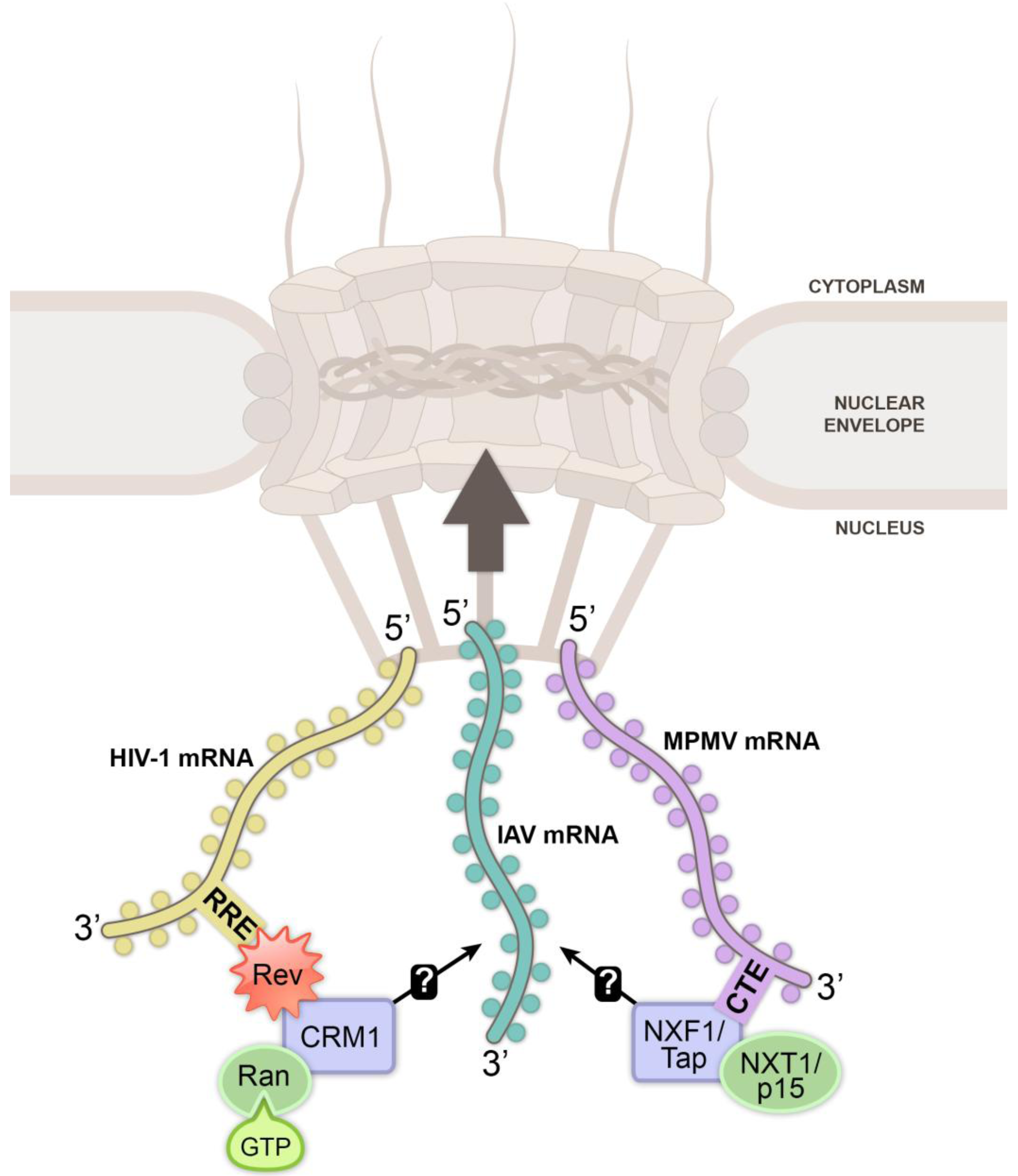
3. Viral Mechanisms to Exploit Nuclear Export Receptors
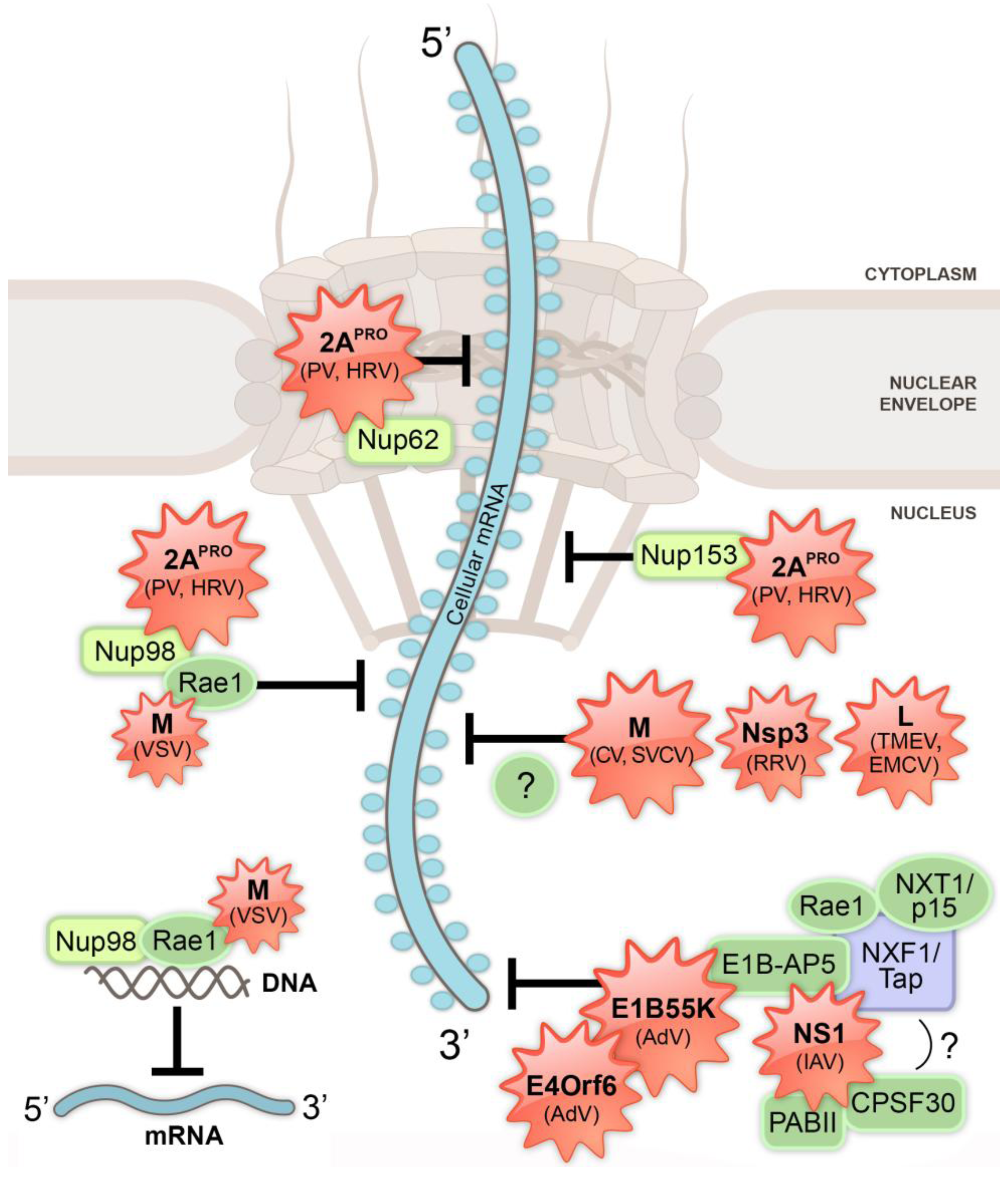
4. Inhibition of Nucleo-Cytoplasmic Trafficking by RNA Viruses
4.1. Influenza Virus Inhibits Host mRNA Processing and Nuclear Export
4.2. Vesiculoviruses and mRNA Nuclear Export Inhibition
4.4. Rotavirus Inhibits Host mRNA Nuclear Export
5. Inhibition of Nucleo-Cytoplasmic Trafficking by DNA Viruses
Adenovirus Inhibits Host mRNA Nuclear Export
6. Conclusions and Future Perspectives
Acknowledgments
Conflict of Interest
References and Notes
- Fensterl, V.; Sen, G.C. Interferons and viral infections. Biofactors 2009, 35, 14–20. [Google Scholar] [CrossRef]
- Ball, L.A. Virus replication strategies. In Fields Virology, 5th ed.; Knipe, D.M., Howley, P.M., Eds.; Lippincott Williams & Wilkins: Philadelphia, PA, USA, 2007; pp. 119–139. [Google Scholar]
- Fornerod, M.; Ohno, M.; Yoshida, M.; Mattaj, I.W. Crm1 is an export receptor for leucine-rich nuclear export signals. Cell 1997, 90, 1051–1060. [Google Scholar] [CrossRef]
- Stade, K.; Ford, C.S.; Guthrie, C.; Weis, K. Exportin 1 (crm1p) is an essential nuclear export factor. Cell 1997, 90, 1041–1050. [Google Scholar] [CrossRef]
- Gruter, P.; Tabernero, C.; von Kobbe, C.; Schmitt, C.; Saavedra, C.; Bachi, A.; Wilm, M.; Felber, B.K.; Izaurralde, E. Tap, the human homolog of mex67p, mediates cte-dependent rna export from the nucleus. Mol. Cell 1998, 1, 649–659. [Google Scholar] [CrossRef]
- Sandri-Goldin, R.M. Viral regulation of mrna export. J. Virol. 2004, 78, 4389–4396. [Google Scholar] [CrossRef]
- Fontoura, B.M.; Faria, P.A.; Nussenzveig, D.R. Viral interactions with the nuclear transport machinery: Discovering and disrupting pathways. IUBMB Life 2005, 57, 65–72. [Google Scholar] [CrossRef]
- Gustin, K.E. Inhibition of nucleo–cytoplasmic trafficking by rna viruses: Targeting the nuclear pore complex. Virus Res. 2003, 95, 35–44. [Google Scholar] [CrossRef]
- Katahira, J. Mrna export and the trex complex. Biochim. Biophys. Acta 2012, 1819, 507–513. [Google Scholar] [CrossRef]
- Rodriguez-Navarro, S.; Hurt, E. Linking gene regulation to mrna production and export. Curr. Opin. Cell Biol. 2011, 23, 302–309. [Google Scholar] [CrossRef]
- Millevoi, S.; Vagner, S. Molecular mechanisms of eukaryotic pre-mrna 3' end processing regulation. Nucleic Acids Res. 2010, 38, 2757–2774. [Google Scholar] [CrossRef]
- Mandel, C.R.; Bai, Y.; Tong, L. Protein factors in pre–mrna 3'-end processing. Cell Mol. Life Sci. 2008, 65, 1099–1122. [Google Scholar] [CrossRef]
- Shatkin, A.J.; Manley, J.L. The ends of the affair: Capping and polyadenylation. Nat. Struct. Biol. 2000, 7, 838–842. [Google Scholar] [CrossRef]
- Cheng, H.; Dufu, K.; Lee, C.S.; Hsu, J.L.; Dias, A.; Reed, R. Human mrna export machinery recruited to the 5' end of mrna. Cell 2006, 127, 1389–1400. [Google Scholar] [CrossRef]
- Izaurralde, E.; Lewis, J.; Gamberi, C.; Jarmolowski, A.; McGuigan, C.; Mattaj, I.W. A cap-binding protein complex mediating u snrna export. Nature 1995, 376, 709–712. [Google Scholar] [CrossRef]
- Zhou, Z.; Luo, M.J.; Straesser, K.; Katahira, J.; Hurt, E.; Reed, R. The protein aly links pre-messenger–rna splicing to nuclear export in metazoans. Nature 2000, 407, 401–405. [Google Scholar] [CrossRef]
- Masuda, S.; Das, R.; Cheng, H.; Hurt, E.; Dorman, N.; Reed, R. Recruitment of the human trex complex to mrna during splicing. Genes Dev. 2005, 19, 1512–1517. [Google Scholar] [CrossRef]
- Chavez, S.; Beilharz, T.; Rondon, A.G.; Erdjument–Bromage, H.; Tempst, P.; Svejstrup, J.Q.; Lithgow, T.; Aguilera, A. A protein complex containing tho2, hpr1, mft1 and a novel protein, thp2, connects transcription elongation with mitotic recombination in saccharomyces cerevisiae. EMBO J. 2000, 19, 5824–5834. [Google Scholar] [CrossRef]
- Rehwinkel, J.; Herold, A.; Gari, K.; Kocher, T.; Rode, M.; Ciccarelli, F.L.; Wilm, M.; Izaurralde, E. Genome-wide analysis of mrnas regulated by the tho complex in drosophila melanogaster. Nat. Struct. Mol. Biol. 2004, 11, 558–566. [Google Scholar] [CrossRef]
- Piruat, J.I.; Aguilera, A. A novel yeast gene, tho2, is involved in rna pol ii transcription and provides new evidence for transcriptional elongation-associated recombination. EMBO J. 1998, 17, 4859–4872. [Google Scholar] [CrossRef]
- Fleckner, J.; Zhang, M.; Valcarcel, J.; Green, M.R. U2af65 recruits a novel human dead box protein required for the u2 snrnp-branchpoint interaction. Genes Dev. 1997, 11, 1864–1872. [Google Scholar] [CrossRef]
- Libri, D.; Graziani, N.; Saguez, C.; Boulay, J. Multiple roles for the yeast sub2/yuap56 gene in splicing. Genes Dev. 2001, 15, 36–41. [Google Scholar] [CrossRef]
- Strasser, K.; Masuda, S.; Mason, P.; Pfannstiel, J.; Oppizzi, M.; Rodriguez-Navarro, S.; Rondon, A.G.; Aguilera, A.; Struhl, K.; Reed, R.; et al. Trex is a conserved complex coupling transcription with messenger rna export. Nature 2002, 417, 304–308. [Google Scholar] [CrossRef]
- Luo, M.L.; Zhou, Z.; Magni, K.; Christoforides, C.; Rappsilber, J.; Mann, M.; Reed, R. Pre-mrna splicing and mrna export linked by direct interactions between uap56 and aly. Nature 2001, 413, 644–647. [Google Scholar] [CrossRef]
- Jensen, T.H.; Boulay, J.; Rosbash, M.; Libri, D. The decd box putative atpase sub2p is an early mrna export factor. Curr. Biol. 2001, 11, 1711–1715. [Google Scholar] [CrossRef]
- MacMorris, M.; Brocker, C.; Blumenthal, T. Uap56 levels affect viability and mrna export in caenorhabditis elegans. RNA 2003, 9, 847–857. [Google Scholar] [CrossRef]
- Strasser, K.; Hurt, E. Splicing factor sub2p is required for nuclear mrna export through its interaction with yra1p. Nature 2001, 413, 648–652. [Google Scholar] [CrossRef]
- Dufu, K.; Livingstone, M.J.; Seebacher, J.; Gygi, S.P.; Wilson, S.A.; Reed, R. Atp is required for interactions between uap56 and two conserved mrna export proteins, aly and cip29, to assemble the trex complex. Genes Dev. 2010, 24, 2043–2053. [Google Scholar] [CrossRef]
- Taniguchi, I.; Ohno, M. Atp-dependent recruitment of export factor aly/ref onto intronless mrnas by rna helicase uap56. Mol. Cell Biol. 2008, 28, 601–608. [Google Scholar] [CrossRef]
- Stutz, F.; Bachi, A.; Doerks, T.; Braun, I.C.; Seraphin, B.; Wilm, M.; Bork, P.; Izaurralde, E. Ref, an evolutionary conserved family of hnrnp-like proteins, interacts with tap/mex67p and participates in mrna nuclear export. RNA 2000, 6, 638–650. [Google Scholar] [CrossRef]
- Strasser, K.; Hurt, E. Yra1p, a conserved nuclear rna-binding protein, interacts directly with mex67p and is required for mrna export. EMBO J. 2000, 19, 410–420. [Google Scholar] [CrossRef]
- Hautbergue, G.M.; Hung, M.L.; Golovanov, A.P.; Lian, L.Y.; Wilson, S.A. Mutually exclusive interactions drive handover of mrna from export adaptors to tap. Proc. Natl. Acad. Sci. USA 2008, 105, 5154–5159. [Google Scholar] [CrossRef]
- Gabler, S.; Schutt, H.; Groitl, P.; Wolf, H.; Shenk, T.; Dobner, T. E1b 55–kilodalton–associated protein: A cellular protein with rna-binding activity implicated in nucleocytoplasmic transport of adenovirus and cellular mrnas. J. Virol. 1998, 72, 7960–7971. [Google Scholar]
- Bachi, A.; Braun, I.C.; Rodrigues, J.P.; Pante, N.; Ribbeck, K.; von Kobbe, C.; Kutay, U.; Wilm, M.; Gorlich, D.; Carmo-Fonseca, M.; et al. The c-terminal domain of tap interacts with the nuclear pore complex and promotes export of specific cte-bearing rna substrates. RNA 2000, 6, 136–158. [Google Scholar] [CrossRef]
- Segref, A.; Sharma, K.; Doye, V.; Hellwig, A.; Huber, J.; Luhrmann, R.; Hurt, E. Mex67p, a novel factor for nuclear mrna export, binds to both poly(a)+ rna and nuclear pores. EMBO J. 1997, 16, 3256–3271. [Google Scholar]
- Katahira, J.; Strasser, K.; Podtelejnikov, A.; Mann, M.; Jung, J.U.; Hurt, E. The mex67p-mediated nuclear mrna export pathway is conserved from yeast to human. EMBO J. 1999, 18, 2593–2609. [Google Scholar]
- Braun, I.C.; Herold, A.; Rode, M.; Izaurralde, E. Nuclear export of mrna by tap/nxf1 requires two nucleoporin–binding sites but not p15. Mol. Cell Biol. 2002, 22, 5405–5418. [Google Scholar] [CrossRef]
- Farny, N.G.; Hurt, J.A.; Silver, P.A. Definition of global and transcript-specific mrna export pathways in metazoans. Genes Dev. 2008, 22, 66–78. [Google Scholar] [CrossRef]
- Carmody, S.R.; Wente, S.R. Mrna nuclear export at a glance. J. Cell Sci. 2009, 122, 1933–1937. [Google Scholar] [CrossRef]
- Blevins, M.B.; Smith, A.M.; Phillips, E.M.; Powers, M.A. Complex formation among the rna export proteins nup98, rae1/gle2, and tap. J. Biol. Chem. 2003, 278, 20979–20988. [Google Scholar]
- Fontoura, B.M.; Blobel, G.; Yaseen, N.R. The nucleoporin nup98 is a site for gdp/gtp exchange on ran and termination of karyopherin beta 2-mediated nuclear import. J. Biol. Chem. 2000, 275, 31289–31296. [Google Scholar] [CrossRef]
- Fontoura, B.M.; Dales, S.; Blobel, G.; Zhong, H. The nucleoporin nup98 associates with the intranuclear filamentous protein network of tpr. Proc. Natl. Acad. Sci. USA 2001, 98, 3208–3213. [Google Scholar] [CrossRef]
- Powers, M.A.; Forbes, D.J.; Dahlberg, J.E.; Lund, E. The vertebrate glfg nucleoporin, nup98, is an essential component of multiple rna export pathways. J. Cell Biol. 1997, 136, 241–250. [Google Scholar] [CrossRef]
- Pritchard, C.E.; Fornerod, M.; Kasper, L.H.; van Deursen, J.M. Rae1 is a shuttling mrna export factor that binds to a glebs-like nup98 motif at the nuclear pore complex through multiple domains. J. Cell Biol. 1999, 145, 237–254. [Google Scholar] [CrossRef]
- Radu, A.; Moore, M.S.; Blobel, G. The peptide repeat domain of nucleoporin nup98 functions as a docking site in transport across the nuclear pore complex. Cell 1995, 81, 215–222. [Google Scholar] [CrossRef]
- Zolotukhin, A.S.; Felber, B.K. Nucleoporins nup98 and nup214 participate in nuclear export of human immunodeficiency virus type 1 rev. J. Virol. 1999, 73, 120–127. [Google Scholar]
- Oka, M.; Asally, M.; Yasuda, Y.; Ogawa, Y.; Tachibana, T.; Yoneda, Y. The mobile fg nucleoporin nup98 is a cofactor for crm1-dependent protein export. Mol. Biol. Cell 2010, 21, 1885–1896. [Google Scholar] [CrossRef]
- Kraemer, D.; Blobel, G. Mrna binding protein mrnp 41 localizes to both nucleus and cytoplasm. Proc. Natl. Acad. Sci. USA 1997, 94, 9119–9124. [Google Scholar] [CrossRef]
- Bharathi, A.; Ghosh, A.; Whalen, W.A.; Yoon, J.H.; Pu, R.; Dasso, M.; Dhar, R. The human rae1 gene is a functional homologue of schizosaccharomyces pombe rae1 gene involved in nuclear export of poly(a)+ rna. Gene 1997, 198, 251–258. [Google Scholar] [CrossRef]
- Bailer, S.M.; Siniossoglou, S.; Podtelejnikov, A.; Hellwig, A.; Mann, M.; Hurt, E. Nup116p and nup100p are interchangeable through a conserved motif which constitutes a docking site for the mrna transport factor gle2p. EMBO J. 1998, 17, 1107–1119. [Google Scholar] [CrossRef]
- Brown, J.A.; Bharathi, A.; Ghosh, A.; Whalen, W.; Fitzgerald, E.; Dhar, R. A mutation in the schizosaccharomyces pombe rae1 gene causes defects in poly(a)+ rna export and in the cytoskeleton. J. Biol. Chem. 1995, 270, 7411–7419. [Google Scholar]
- Whalen, W.A.; Bharathi, A.; Danielewicz, D.; Dhar, R. Advancement through mitosis requires rae1 gene function in fission yeast. Yeast 1997, 13, 1167–1179. [Google Scholar] [CrossRef]
- Faria, P.A.; Chakraborty, P.; Levay, A.; Barber, G.N.; Ezelle, H.J.; Enninga, J.; Arana, C.; van Deursen, J.; Fontoura, B.M. Vsv disrupts the rae1/mrnp41 mrna nuclear export pathway. Mol. Cell 2005, 17, 93–102. [Google Scholar] [CrossRef]
- Ren, Y.; Seo, H.S.; Blobel, G.; Hoelz, A. Structural and functional analysis of the interaction between the nucleoporin nup98 and the mrna export factor rae1. Proc. Natl. Acad. Sci. USA 2010, 107, 10406–10411. [Google Scholar] [CrossRef]
- Alcazar–Roman, A.R.; Tran, E.J.; Guo, S.; Wente, S.R. Inositol hexakisphosphate and gle1 activate the dead–box protein dbp5 for nuclear mrna export. Nat. Cell Biol. 2006, 8, 711–716. [Google Scholar] [CrossRef]
- Weirich, C.S.; Erzberger, J.P.; Flick, J.S.; Berger, J.M.; Thorner, J.; Weis, K. Activation of the dexd/h–box protein dbp5 by the nuclear-pore protein gle1 and its coactivator insp6 is required for mrna export. Nat. Cell Biol. 2006, 8, 668–676. [Google Scholar] [CrossRef]
- Murphy, R.; Wente, S.R. An rna–export mediator with an essential nuclear export signal. Nature 1996, 383, 357–360. [Google Scholar] [CrossRef]
- Snay-Hodge, C.A.; Colot, H.V.; Goldstein, A.L.; Cole, C.N. Dbp5p/rat8p is a yeast nuclear pore–associated dead–box protein essential for rna export. EMBO J. 1998, 17, 2663–2676. [Google Scholar] [CrossRef]
- Tseng, S.S.; Weaver, P.L.; Liu, Y.; Hitomi, M.; Tartakoff, A.M.; Chang, T.H. Dbp5p, a cytosolic rna helicase, is required for poly(a)+ rna export. EMBO J. 1998, 17, 2651–2662. [Google Scholar] [CrossRef]
- Schmitt, C.; von Kobbe, C.; Bachi, A.; Pante, N.; Rodrigues, J.P.; Boscheron, C.; Rigaut, G.; Wilm, M.; Seraphin, B.; Carmo-Fonseca, M.; et al. Dbp5, a dead-box protein required for mrna export, is recruited to the cytoplasmic fibrils of nuclear pore complex via a conserved interaction with can/nup159p. EMBO J. 1999, 18, 4332–4347. [Google Scholar] [CrossRef]
- Strahm, Y.; Fahrenkrog, B.; Zenklusen, D.; Rychner, E.; Kantor, J.; Rosbach, M.; Stutz, F. The rna export factor gle1p is located on the cytoplasmic fibrils of the npc and physically interacts with the fg–nucleoporin rip1p, the dead-box protein rat8p/dbp5p and a new protein ymr 255p. EMBO J. 1999, 18, 5761–5777. [Google Scholar] [CrossRef]
- Weirich, C.S.; Erzberger, J.P.; Berger, J.M.; Weis, K. The n-terminal domain of nup159 forms a beta-propeller that functions in mrna export by tethering the helicase dbp5 to the nuclear pore. Mol. Cell 2004, 16, 749–760. [Google Scholar] [CrossRef]
- Kendirgi, F.; Rexer, D.J.; Alcazar-Roman, A.R.; Onishko, H.M.; Wente, S.R. Interaction between the shuttling mrna export factor gle1 and the nucleoporin hcg1: A conserved mechanism in the export of hsp70 mrna. Mol. Biol. Cell 2005, 16, 4304–4315. [Google Scholar] [CrossRef]
- Tran, E.J.; Zhou, Y.; Corbett, A.H.; Wente, S.R. The dead-box protein dbp5 controls mrna export by triggering specific rna:Protein remodeling events. Mol. Cell 2007, 28, 850–859. [Google Scholar] [CrossRef]
- von Moeller, H.; Basquin, C.; Conti, E. The mrna export protein dbp5 binds rna and the cytoplasmic nucleoporin nup214 in a mutually exclusive manner. Nat. Struct. Mol. Biol. 2009, 16, 247–254. [Google Scholar] [CrossRef]
- Lund, M.K.; Guthrie, C. The dead-box protein dbp5p is required to dissociate mex67p from exported mrnps at the nuclear rim. Mol. Cell 2005, 20, 645–651. [Google Scholar] [CrossRef]
- Tran, E.J.; Wente, S.R. Dynamic nuclear pore complexes: Life on the edge. Cell 2006, 125, 1041–1053. [Google Scholar] [CrossRef]
- Jin, L.; Guzik, B.W.; Bor, Y.C.; Rekosh, D.; Hammarskjold, M.L. Tap and nxt promote translation of unspliced mrna. Genes Dev. 2003, 17, 3075–3086. [Google Scholar] [CrossRef]
- Bolger, T.A.; Folkmann, A.W.; Tran, E.J.; Wente, S.R. The mrna export factor gle1 and inositol hexakisphosphate regulate distinct stages of translation. Cell 2008, 134, 624–633. [Google Scholar] [CrossRef]
- Natalizio, B.J.; Wente, S.R. Postage for the messenger: Designating routes for nuclear mrna export. Trends Cell Biol. 2013. [Google Scholar] [CrossRef]
- Culjkovic-Kraljacic, B.; Borden, K.L. Aiding and abetting cancer: Mrna export and the nuclear pore. Trends Cell Biol. 2013, 23, 328–335. [Google Scholar] [CrossRef]
- Speese, S.D.; Ashley, J.; Jokhi, V.; Nunnari, J.; Barria, R.; Li, Y.; Ataman, B.; Koon, A.; Chang, Y.T.; Li, Q.; et al. Nuclear envelope budding enables large ribonucleoprotein particle export during synaptic wnt signaling. Cell 2012, 149, 832–846. [Google Scholar] [CrossRef]
- Buser, C.; Walther, P.; Mertens, T.; Michel, D. Cytomegalovirus primary envelopment occurs at large infoldings of the inner nuclear membrane. J. Virol. 2007, 81, 3042–3048. [Google Scholar] [CrossRef]
- Darlington, R.W.; Moss, L.H., 3rd. Herpesvirus envelopment. J. Virol. 1968, 2, 48–55. [Google Scholar]
- Lee, C.P.; Chen, M.R. Escape of herpesviruses from the nucleus. Rev. Med. Virol. 2010, 20, 214–230. [Google Scholar] [CrossRef]
- Burke, B.; Stewart, C.L. The nuclear lamins: Flexibility in function. Nat. Rev. Mol. Cell Biol. 2013, 14, 13–24. [Google Scholar] [CrossRef]
- Malim, M.H.; Hauber, J.; Le, S.Y.; Maizel, J.V.; Cullen, B.R. The hiv-1 rev trans-activator acts through a structured target sequence to activate nuclear export of unspliced viral mrna. Nature 1989, 338, 254–257. [Google Scholar] [CrossRef]
- Yoon, D.W.; Lee, H.; Seol, W.; DeMaria, M.; Rosenzweig, M.; Jung, J.U. Tap: A novel cellular protein that interacts with tip of herpesvirus saimiri and induces lymphocyte aggregation. Immunity 1997, 6, 571–582. [Google Scholar] [CrossRef]
- Pasquinelli, A.E.; Ernst, R.K.; Lund, E.; Grimm, C.; Zapp, M.L.; Rekosh, D.; Hammarskjold, M.L.; Dahlberg, J.E. The constitutive transport element (cte) of mason-pfizer monkey virus (mpmv) accesses a cellular mrna export pathway. EMBO J. 1997, 16, 7500–7510. [Google Scholar] [CrossRef]
- Cullen, B.R. Nuclear mrna export: Insights from virology. Trends Biochem. Sci. 2003, 28, 419–424. [Google Scholar] [CrossRef]
- Mears, W.E.; Rice, S.A. The herpes simplex virus immediate-early protein icp27 shuttles between nucleus and cytoplasm. Virology 1998, 242, 128–137. [Google Scholar] [CrossRef]
- Phelan, A.; Clements, J.B. Herpes simplex virus type 1 immediate early protein ie63 shuttles between nuclear compartments and the cytoplasm. J. Gen. Virol. 1997, 78, 3327–3331. [Google Scholar]
- Sandri-Goldin, R.M. Icp27 mediates hsv rna export by shuttling through a leucine–rich nuclear export signal and binding viral intronless rnas through an rgg motif. Genes Dev. 1998, 12, 868–879. [Google Scholar] [CrossRef]
- Soliman, T.M.; Sandri-Goldin, R.M.; Silverstein, S.J. Shuttling of the herpes simplex virus type 1 regulatory protein icp27 between the nucleus and cytoplasm mediates the expression of late proteins. J. Virol. 1997, 71, 9188–9197. [Google Scholar]
- Bryant, H.E.; Wadd, S.E.; Lamond, A.I.; Silverstein, S.J.; Clements, J.B. Herpes simplex virus ie63 (icp27) protein interacts with spliceosome-associated protein 145 and inhibits splicing prior to the first catalytic step. J. Virol. 2001, 75, 4376–4385. [Google Scholar] [CrossRef]
- Hardy, W.R.; Sandri-Goldin, R.M. Herpes simplex virus inhibits host cell splicing, and regulatory protein icp27 is required for this effect. J. Virol. 1994, 68, 7790–7799. [Google Scholar]
- Lindberg, A.; Kreivi, J.P. Splicing inhibition at the level of spliceosome assembly in the presence of herpes simplex virus protein icp27. Virology 2002, 294, 189–198. [Google Scholar] [CrossRef]
- Koffa, M.D.; Clements, J.B.; Izaurralde, E.; Wadd, S.; Wilson, S.A.; Mattaj, I.W.; Kuersten, S. Herpes simplex virus icp27 protein provides viral mrnas with access to the cellular mrna export pathway. EMBO J. 2001, 20, 5769–5778. [Google Scholar] [CrossRef]
- Chen, I.H.; Sciabica, K.S.; Sandri-Goldin, R.M. Icp27 interacts with the rna export factor aly/ref to direct herpes simplex virus type 1 intronless mrnas to the tap export pathway. J. Virol. 2002, 76, 12877–12889. [Google Scholar] [CrossRef]
- Chen, I.H.; Li, L.; Silva, L.; Sandri-Goldin, R.M. Icp27 recruits aly/ref but not tap/nxf1 to herpes simplex virus type 1 transcription sites although tap/nxf1 is required for icp27 export. J. Virol. 2005, 79, 3949–3961. [Google Scholar] [CrossRef]
- Johnson, L.A.; Li, L.; Sandri-Goldin, R.M. The cellular rna export receptor tap/nxf1 is required for icp27-mediated export of herpes simplex virus 1 rna, but the trex complex adaptor protein aly/ref appears to be dispensable. J. Virol. 2009, 83, 6335–6346. [Google Scholar] [CrossRef]
- Johnson, L.A.; Sandri-Goldin, R.M. Efficient nuclear export of herpes simplex virus 1 transcripts requires both rna binding by icp27 and icp27 interaction with tap/nxf1. J. Virol. 2009, 83, 1184–1192. [Google Scholar] [CrossRef]
- Soliman, T.M.; Silverstein, S.J. Identification of an export control sequence and a requirement for the kh domains in icp27 from herpes simplex virus type 1. J. Virol. 2000, 74, 7600–7609. [Google Scholar] [CrossRef]
- Soliman, T.M.; Silverstein, S.J. Herpesvirus mrnas are sorted for export via crm1-dependent and –independent pathways. J. Virol. 2000, 74, 2814–2825. [Google Scholar] [CrossRef]
- Boyne, J.R.; Colgan, K.J.; Whitehouse, A. Recruitment of the complete htrex complex is required for kaposi's sarcoma-associated herpesvirus intronless mrna nuclear export and virus replication. PLoS Pathog. 2008, 4, e1000194. [Google Scholar] [CrossRef]
- Jackson, B.R.; Boyne, J.R.; Noerenberg, M.; Taylor, A.; Hautbergue, G.M.; Walsh, M.J.; Wheat, R.; Blackbourn, D.J.; Wilson, S.A.; Whitehouse, A. An interaction between kshv orf57 and uif provides mrna-adaptor redundancy in herpesvirus intronless mrna export. PLoS Pathog. 2011, 7, e1002138. [Google Scholar] [CrossRef]
- Williams, B.J.; Boyne, J.R.; Goodwin, D.J.; Roaden, L.; Hautbergue, G.M.; Wilson, S.A.; Whitehouse, A. The prototype gamma-2 herpesvirus nucleocytoplasmic shuttling protein, orf 57, transports viral rna through the cellular mrna export pathway. Biochem. J. 2005, 387, 295–308. [Google Scholar] [CrossRef]
- Hiriart, E.; Bardouillet, L.; Manet, E.; Gruffat, H.; Penin, F.; Montserret, R.; Farjot, G.; Sergeant, A. A region of the epstein-barr virus (ebv) mrna export factor eb2 containing an arginine-rich motif mediates direct binding to rna. J. Biol. Chem. 2003, 278, 37790–37798. [Google Scholar] [CrossRef]
- Ote, I.; Lebrun, M.; Vandevenne, P.; Bontems, S.; Medina-Palazon, C.; Manet, E.; Piette, J.; Sadzot-Delvaux, C. Varicella-zoster virus ie4 protein interacts with sr proteins and exports mrnas through the tap/nxf1 pathway. PLoS One 2009, 4, e7882. [Google Scholar] [CrossRef]
- Lischka, P.; Toth, Z.; Thomas, M.; Mueller, R.; Stamminger, T. The ul69 transactivator protein of human cytomegalovirus interacts with dexd/h-box rna helicase uap56 to promote cytoplasmic accumulation of unspliced rna. Mol. Cell Biol. 2006, 26, 1631–1643. [Google Scholar]
- Gatfield, D.; Izaurralde, E. Ref1/aly and the additional exon junction complex proteins are dispensable for nuclear mrna export. J. Cell Biol. 2002, 159, 579–588. [Google Scholar]
- Boyle, S.M.; Ruvolo, V.; Gupta, A.K.; Swaminathan, S. Association with the cellular export receptor crm 1 mediates function and intracellular localization of epstein-barr virus sm protein, a regulator of gene expression. J. Virol. 1999, 73, 6872–6881. [Google Scholar]
- Farjot, G.; Buisson, M.; Duc Dodon, M.; Gazzolo, L.; Sergeant, A.; Mikaelian, I. Epstein-barr virus eb2 protein exports unspliced rna via a crm-1-independent pathway. J. Virol. 2000, 74, 6068–6076. [Google Scholar] [CrossRef]
- Elton, D.; Simpson-Holley, M.; Archer, K.; Medcalf, L.; Hallam, R.; McCauley, J.; Digard, P. Interaction of the influenza virus nucleoprotein with the cellular crm1-mediated nuclear export pathway. J. Virol. 2001, 75, 408–419. [Google Scholar]
- Watanabe, K.; Takizawa, N.; Katoh, M.; Hoshida, K.; Kobayashi, N.; Nagata, K. Inhibition of nuclear export of ribonucleoprotein complexes of influenza virus by leptomycin b. Virus Res. 2001, 77, 31–42. [Google Scholar]
- Hao, L.; Sakurai, A.; Watanabe, T.; Sorensen, E.; Nidom, C.A.; Newton, M.A.; Ahlquist, P.; Kawaoka, Y. Drosophila rnai screen identifies host genes important for influenza virus replication. Nature 2008, 454, 890–893. [Google Scholar] [CrossRef]
- Morita, M.; Kuba, K.; Ichikawa, A.; Nakayama, M.; Katahira, J.; Iwamoto, R.; Watanebe, T.; Sakabe, S.; Daidoji, T.; Nakamura, S.; et al. The lipid mediator protectin d1 inhibits influenza virus replication and improves severe influenza. Cell 2013, 153, 112–125. [Google Scholar] [CrossRef]
- Wang, W.; Cui, Z.Q.; Han, H.; Zhang, Z.P.; Wei, H.P.; Zhou, Y.F.; Chen, Z.; Zhang, X.E. Imaging and characterizing influenza a virus mrna transport in living cells. Nucleic Acids Res. 2008, 36, 4913–4928. [Google Scholar] [CrossRef]
- Read, E.K.; Digard, P. Individual influenza a virus mrnas show differential dependence on cellular nxf1/tap for their nuclear export. J. Gen. Virol. 2010, 91, 1290–1301. [Google Scholar]
- Plotch, S.J.; Bouloy, M.; Ulmanen, I.; Krug, R.M. A unique cap(m7gpppxm)-dependent influenza virion endonuclease cleaves capped rnas to generate the primers that initiate viral rna transcription. Cell 1981, 23, 847–858. [Google Scholar] [CrossRef]
- Hale, B.G.; Randall, R.E.; Ortin, J.; Jackson, D. The multifunctional ns1 protein of influenza a viruses. J. Gen. Virol. 2008, 89, 2359–2376. [Google Scholar] [CrossRef]
- Isaacs, A.; Lindenmann, J. Virus interference. I. The interferon. Proc. R. Soc. Lond. B. Biol. Sci. 1957, 147, 258–267. [Google Scholar]
- Garcia-Sastre, A. Induction and evasion of type i interferon responses by influenza viruses. Virus Res. 2011, 162, 12–18. [Google Scholar] [CrossRef]
- Qiu, Y.; Krug, R.M. The influenza virus ns1 protein is a poly(a)-binding protein that inhibits nuclear export of mrnas containing poly(a). J. Virol. 1994, 68, 2425–2432. [Google Scholar]
- Alonso–Caplen, F.V.; Nemeroff, M.E.; Qiu, Y.; Krug, R.M. Nucleocytoplasmic transport: The influenza virus ns1 protein regulates the transport of spliced ns2 mrna and its precursor ns1 mrna. Genes Dev. 1992, 6, 255–267. [Google Scholar] [CrossRef]
- Fortes, P.; Beloso, A.; Ortin, J. Influenza virus ns1 protein inhibits pre-mrna splicing and blocks mrna nucleocytoplasmic transport. EMBO J. 1994, 13, 704–712. [Google Scholar]
- Qiu, Y.; Nemeroff, M.; Krug, R.M. The influenza virus ns1 protein binds to a specific region in human u6 snrna and inhibits u6-u2 and u6-u4 snrna interactions during splicing. RNA 1995, 1, 304–316. [Google Scholar]
- Nemeroff, M.E.; Barabino, S.M.; Li, Y.; Keller, W.; Krug, R.M. Influenza virus ns1 protein interacts with the cellular 30 kda subunit of cpsf and inhibits 3'end formation of cellular pre-mrnas. Mol. Cell 1998, 1, 991–1000. [Google Scholar] [CrossRef]
- Chen, Z.; Li, Y.; Krug, R.M. Influenza a virus ns1 protein targets poly(a)-binding protein ii of the cellular 3'-end processing machinery. EMBO J. 1999, 18, 2273–2283. [Google Scholar] [CrossRef]
- Poon, L.L.; Pritlove, D.C.; Fodor, E.; Brownlee, G.G. Direct evidence that the poly(a) tail of influenza a virus mrna is synthesized by reiterative copying of a u track in the virion rna template. J. Virol. 1999, 73, 3473–3476. [Google Scholar]
- Robertson, J.S.; Schubert, M.; Lazzarini, R.A. Polyadenylation sites for influenza virus mrna. J. Virol. 1981, 38, 157–163. [Google Scholar]
- Robb, N.C.; Fodor, E. The accumulation of influenza a virus segment 7 spliced mrnas is regulated by the ns1 protein. J. Gen. Virol. 2012, 93, 113–118. [Google Scholar] [CrossRef]
- Wolff, T.; O'Neill, R.E.; Palese, P. Ns1-binding protein (ns1-bp): A novel human protein that interacts with the influenza a virus nonstructural ns1 protein is relocalized in the nuclei of infected cells. J. Virol. 1998, 72, 7170–7180. [Google Scholar]
- Tsai, P.-L.; Chiou, N.-T.; Kuss, S.; Garcia-Sastre, A.; Lynch, K.W.; Fontoua, B.M.A. Cellular rna binding proteins ns1-bp and hnrnp k regulate influenza a virus rna splicing. PLoS Pathog. 2013, 9, e1003460. [Google Scholar] [CrossRef]
- Satterly, N.; Tsai, P.L.; van Deursen, J.; Nussenzveig, D.R.; Wang, Y.; Faria, P.A.; Levay, A.; Levy, D.E.; Fontoura, B.M. Influenza virus targets the mrna export machinery and the nuclear pore complex. Proc. Natl. Acad. Sci. USA 2007, 104, 1853–1858. [Google Scholar]
- Enninga, J.; Levy, D.E.; Blobel, G.; Fontoura, B.M. Role of nucleoporin induction in releasing an mrna nuclear export block. Science 2002, 295, 1523–1525. [Google Scholar] [CrossRef]
- Castello, A.; Izquierdo, J.M.; Welnowska, E.; Carrasco, L. Rna nuclear export is blocked by poliovirus 2a protease and is concomitant with nucleoporin cleavage. J. Cell. Sci. 2009, 122, 3799–3809. [Google Scholar]
- Qu, X.; Lykke-Andersen, S.; Nasser, T.; Saguez, C.; Bertrand, E.; Jensen, T.H.; Moore, C. Assembly of an export-competent mrnp is needed for efficient release of the 3'-end processing complex after polyadenylation. Mol. Cell Biol. 2009, 29, 5327–5338. [Google Scholar]
- Hammell, C.M.; Gross, S.; Zenklusen, D.; Heath, C.V.; Stutz, F.; Moore, C.; Cole, C.N. Coupling of termination, 3' processing, and mrna export. Mol. Cell Biol. 2002, 22, 6441–6457. [Google Scholar]
- Hilleren, P.; Parker, R. Defects in the mrna export factors rat7p, gle1p, mex67p, and rat8p cause hyperadenylation during 3'-end formation of nascent transcripts. RNA 2001, 7, 753–764. [Google Scholar]
- Jensen, T.H.; Patricio, K.; McCarthy, T.; Rosbash, M. A block to mrna nuclear export in s. Cerevisiae leads to hyperadenylation of transcripts that accumulate at the site of transcription. Mol. Cell 2001, 7, 887–898. [Google Scholar] [CrossRef]
- Lyles, D.S.; Rupprecht, C.E. Rhabdoviridae. In Fields Virology, 5th ed.; Knipe, D.M., Howley, P.M., Eds.; Lippincott Williams & Wilkins: Philadelphia, PA, USA, 2007; pp. 1363–1408. [Google Scholar]
- Glodowski, D.R.; Petersen, J.M.; Dahlberg, J.E. Complex nuclear localization signals in the matrix protein of vesicular stomatitis virus. J. Biol. Chem. 2002, 277, 46864–46870. [Google Scholar] [CrossRef]
- von Kobbe, C.; van Deursen, J.M.; Rodrigues, J.P.; Sitterlin, D.; Bachi, A.; Wu, X.; Wilm, M.; Carmo-Fonseca, M.; Izaurralde, E. Vesicular stomatitis virus matrix protein inhibits host cell gene expression by targeting the nucleoporin nup98. Mol. Cell 2000, 6, 1243–1252. [Google Scholar] [CrossRef]
- Her, L.S.; Lund, E.; Dahlberg, J.E. Inhibition of ran guanosine triphosphatase-dependent nuclear transport by the matrix protein of vesicular stomatitis virus. Science 1997, 276, 1845–1848. [Google Scholar] [CrossRef]
- Petersen, J.M.; Her, L.S.; Varvel, V.; Lund, E.; Dahlberg, J.E. The matrix protein of vesicular stomatitis virus inhibits nucleocytoplasmic transport when it is in the nucleus and associated with nuclear pore complexes. Mol. Cell Biol. 2000, 20, 8590–8601. [Google Scholar] [CrossRef]
- Zhang, L.; Das, P.; Schmolke, M.; Manicassamy, B.; Wang, Y.; Deng, X.; Cai, L.; Tu, B.P.; Forst, C.V.; Roth, M.G.; et al. Inhibition of pyrimidine synthesis reverses viral virulence factor-mediated block of mrna nuclear export. J. Cell Biol. 2012, 196, 315–326. [Google Scholar] [CrossRef]
- Petersen, J.M.; Her, L.S.; Dahlberg, J.E. Multiple vesiculoviral matrix proteins inhibit both nuclear export and import. Proc. Natl. Acad. Sci. USA 2001, 98, 8590–8595. [Google Scholar] [CrossRef]
- Coulon, P.; Deutsch, V.; Lafay, F.; Martinet-Edelist, C.; Wyers, F.; Herman, R.C.; Flamand, A. Genetic evidence for multiple functions of the matrix protein of vesicular stomatitis virus. J. Gen. Virol. 1990, 71, 991–996. [Google Scholar] [CrossRef]
- Rajani, K.R.; Pettit Kneller, E.L.; McKenzie, M.O.; Horita, D.A.; Chou, J.W.; Lyles, D.S. Complexes of vesicular stomatitis virus matrix protein with host rae1 and nup98 involved in inhibition of host transcription. PLoS Pathog. 2012, 8, e1002929. [Google Scholar] [CrossRef]
- Capelson, M.; Liang, Y.; Schulte, R.; Mair, W.; Wagner, U.; Hetzer, M.W. Chromatin-bound nuclear pore components regulate gene expression in higher eukaryotes. Cell 2010, 140, 372–383. [Google Scholar] [CrossRef]
- Babu, J.R.; Jeganathan, K.B.; Baker, D.J.; Wu, X.; Kang-Decker, N.; van Deursen, J.M. Rae1 is an essential mitotic checkpoint regulator that cooperates with bub3 to prevent chromosome missegregation. J. Cell Biol. 2003, 160, 341–353. [Google Scholar] [CrossRef]
- Griffis, E.R.; Altan, N.; Lippincott-Schwartz, J.; Powers, M.A. Nup98 is a mobile nucleoporin with transcription–dependent dynamics. Mol. Biol. Cell 2002, 13, 1282–1297. [Google Scholar] [CrossRef]
- Egecioglu, D.; Brickner, J.H. Gene positioning and expression. Curr. Opin. Cell Biol. 2011, 23, 338–345. [Google Scholar] [CrossRef]
- Singer, S.; Zhao, R.; Barsotti, A.M.; Ouwehand, A.; Fazollahi, M.; Coutavas, E.; Breuhahn, K.; Neumann, O.; Longerich, T.; Pusterla, T.; et al. Nuclear pore component nup98 is a potential tumor suppressor and regulates posttranscriptional expression of select p53 target genes. Mol. Cell 2012, 48, 799–810. [Google Scholar] [CrossRef]
- Yarbrough, M.L.; White, M.A.; Fontoura, B.M. Shaping the p53 response with nucleoporins. Mol. Cell 2012, 48, 665–666. [Google Scholar] [CrossRef]
- Blower, M.D.; Nachury, M.; Heald, R.; Weis, K. A rae1-containing ribonucleoprotein complex is required for mitotic spindle assembly. Cell 2005, 121, 223–234. [Google Scholar] [CrossRef]
- Cross, M.K.; Powers, M.A. Nup98 regulates bipolar spindle assembly through association with microtubules and opposition of mcak. Mol. Biol. Cell 2011, 22, 661–672. [Google Scholar] [CrossRef]
- Chakraborty, P.; Seemann, J.; Mishra, R.K.; Wei, J.H.; Weil, L.; Nussenzveig, D.R.; Heiber, J.; Barber, G.N.; Dasso, M.; Fontoura, B.M. Vesicular stomatitis virus inhibits mitotic progression and triggers cell death. EMBO Rep. 2009, 10, 1154–1160. [Google Scholar] [CrossRef]
- Hastie, E.; Grdzelishvili, V.Z. Vesicular stomatitis virus as a flexible platform for oncolytic virotherapy against cancer. J. Gen. Virol. 2012, 93, 2529–2545. [Google Scholar] [CrossRef]
- Renukaradhya, G.J.; Webb, T.J.; Khan, M.A.; Lin, Y.L.; Du, W.; Gervay-Hague, J.; Brutkiewicz, R.R. Virus-induced inhibition of cd1d1–mediated antigen presentation: Reciprocal regulation by p38 and erk. J. Immunol. 2005, 175, 4301–4308. [Google Scholar]
- Gustin, K.E.; Sarnow, P. Effects of poliovirus infection on nucleo-cytoplasmic trafficking and nuclear pore complex composition. EMBO J. 2001, 20, 240–249. [Google Scholar] [CrossRef]
- Gustin, K.E.; Sarnow, P. Inhibition of nuclear import and alteration of nuclear pore complex composition by rhinovirus. J. Virol. 2002, 76, 8787–8796. [Google Scholar] [CrossRef]
- Ricour, C.; Borghese, F.; Sorgeloos, F.; Hato, S.V.; van Kuppeveld, F.J.; Michiels, T. Random mutagenesis defines a domain of theiler's virus leader protein that is essential for antagonism of nucleocytoplasmic trafficking and cytokine gene expression. J. Virol. 2009, 83, 11223–11232. [Google Scholar] [CrossRef]
- Ricour, C.; Delhaye, S.; Hato, S.V.; Olenyik, T.D.; Michel, B.; van Kuppeveld, F.J.; Gustin, K.E.; Michiels, T. Inhibition of mrna export and dimerization of interferon regulatory factor 3 by theiler's virus leader protein. J. Gen. Virol. 2009, 90, 177–186. [Google Scholar] [CrossRef]
- Porter, F.W.; Bochkov, Y.A.; Albee, A.J.; Wiese, C.; Palmenberg, A.C. A picornavirus protein interacts with ran-gtpase and disrupts nucleocytoplasmic transport. Proc. Natl. Acad. Sci. USA 2006, 103, 12417–12422. [Google Scholar]
- Belov, G.A.; Lidsky, P.V.; Mikitas, O.V.; Egger, D.; Lukyanov, K.A.; Bienz, K.; Agol, V.I. Bidirectional increase in permeability of nuclear envelope upon poliovirus infection and accompanying alterations of nuclear pores. J. Virol. 2004, 78, 10166–10177. [Google Scholar]
- Lidsky, P.V.; Hato, S.; Bardina, M.V.; Aminev, A.G.; Palmenberg, A.C.; Sheval, E.V.; Polyakov, V.Y.; van Kuppeveld, F.J.; Agol, V.I. Nucleocytoplasmic traffic disorder induced by cardioviruses. J. Virol. 2006, 80, 2705–2717. [Google Scholar] [CrossRef]
- Park, N.; Katikaneni, P.; Skern, T.; Gustin, K.E. Differential targeting of nuclear pore complex proteins in poliovirus-infected cells. J. Virol. 2008, 82, 1647–1655. [Google Scholar] [CrossRef]
- Park, N.; Skern, T.; Gustin, K.E. Specific cleavage of the nuclear pore complex protein nup62 by a viral protease. J. Biol. Chem. 2010, 285, 28796–28805. [Google Scholar] [CrossRef]
- Watters, K.; Palmenberg, A.C. Differential processing of nuclear pore complex proteins by rhinovirus 2a proteases from different species and serotypes. J. Virol. 2011, 85, 10874–10883. [Google Scholar] [CrossRef]
- Ghildyal, R.; Jordan, B.; Li, D.; Dagher, H.; Bardin, P.G.; Gern, J.E.; Jans, D.A. Rhinovirus 3c protease can localize in the nucleus and alter active and passive nucleocytoplasmic transport. J. Virol. 2009, 83, 7349–7352. [Google Scholar] [CrossRef]
- Racaniello, V. Picornaviridae: The viruses and their replication. In Fields Virology, 5th ed.; Knipe, D.M., Howley, P.M., Eds.; Lippincott Williams & Wilkins: Philadelphia, PA, USA, 2007; pp. 795–838. [Google Scholar]
- Porter, F.W.; Palmenberg, A.C. Leader-induced phosphorylation of nucleoporins correlates with nuclear trafficking inhibition by cardioviruses. J. Virol. 2009, 83, 1941–1951. [Google Scholar] [CrossRef]
- Bardina, M.V.; Lidsky, P.V.; Sheval, E.V.; Fominykh, K.V.; van Kuppeveld, F.J.; Polyakov, V.Y.; Agol, V.I. Mengovirus-induced rearrangement of the nuclear pore complex: Hijacking cellular phosphorylation machinery. J. Virol. 2009, 83, 3150–3161. [Google Scholar] [CrossRef]
- Delhaye, S.; van Pesch, V.; Michiels, T. The leader protein of theiler's virus interferes with nucleocytoplasmic trafficking of cellular proteins. J. Virol. 2004, 78, 4357–4362. [Google Scholar] [CrossRef]
- Rubio, R.M.; Mora, S.I.; Romero, P.; Arias, C.F.; Lopez, S. Rotavirus prevents the expression of host responses by blocking the nucleocytoplasmic transport of polyadenylated mrnas. J. Virol. 2013, 87, 6336–6345. [Google Scholar] [CrossRef]
- Beltz, G.A.; Flint, S.J. Inhibition of hela cell protein synthesis during adenovirus infection. Restriction of cellular messenger rna sequences to the nucleus. J. Mol. Biol. 1979, 131, 353–373. [Google Scholar] [CrossRef]
- Babiss, L.E.; Ginsberg, H.S. Adenovirus type 5 early region 1b gene product is required for efficient shutoff of host protein synthesis. J. Virol. 1984, 50, 202–212. [Google Scholar]
- Pilder, S.; Moore, M.; Logan, J.; Shenk, T. The adenovirus e1b–55k transforming polypeptide modulates transport or cytoplasmic stabilization of viral and host cell mrnas. Mol. Cell. Biol. 1986, 6, 470–476. [Google Scholar]
- Flint, S.J.; Gonzalez, R.A. Regulation of mrna production by the adenoviral e1b 55-kda and e4 orf6 proteins. Curr. Top. Microbiol. Immunol. 2003, 272, 287–330. [Google Scholar] [CrossRef]
- Halbert, D.N.; Cutt, J.R.; Shenk, T. Adenovirus early region 4 encodes functions required for efficient DNA replication, late gene expression, and host cell shutoff. J. Virol. 1985, 56, 250–257. [Google Scholar]
- Blanchette, P.; Kindsmuller, K.; Groitl, P.; Dallaire, F.; Speiseder, T.; Branton, P.E.; Dobner, T. Control of mrna export by adenovirus e4orf6 and e1b55k proteins during productive infection requires e4orf6 ubiquitin ligase activity. J. Virol. 2008, 82, 2642–2651. [Google Scholar] [CrossRef]
- Liang, S.; Hitomi, M.; Tartakoff, A.M. Adenoviral e1b-55kda protein inhibits yeast mrna export and perturbs nuclear structure. Proc. Natl. Acad. Sci. USA 1995, 92, 7372–7375. [Google Scholar] [CrossRef]
- Yang, U.C.; Huang, W.; Flint, S.J. Mrna export correlates with activation of transcription in human subgroup c adenovirus-infected cells. J. Virol. 1996, 70, 4071–4080. [Google Scholar]
- Yatherajam, G.; Huang, W.; Flint, S.J. Export of adenoviral late mrna from the nucleus requires the nxf1/tap export receptor. J. Virol. 2011, 85, 1429–1438. [Google Scholar] [CrossRef]
- Dosch, T.; Horn, F.; Schneider, G.; Kratzer, F.; Dobner, T.; Hauber, J.; Stauber, R.H. The adenovirus type 5 e1b–55k oncoprotein actively shuttles in virus-infected cells, whereas transport of e4orf6 is mediated by a crm1–independent mechanism. J. Virol. 2001, 75, 5677–5683. [Google Scholar] [CrossRef]
- Carter, C.C.; Izadpanah, R.; Bridge, E. Evaluating the role of crm1-mediated export for adenovirus gene expression. Virology 2003, 315, 224–233. [Google Scholar] [CrossRef]
- Flint, S.J.; Huang, W.; Goodhouse, J.; Kyin, S. A peptide inhibitor of exportin1 blocks shuttling of the adenoviral e1b 55 kda protein but not export of viral late mrnas. Virology 2005, 337, 7–17. [Google Scholar] [CrossRef]
- Rabino, C.; Aspegren, A.; Corbin-Lickfett, K.; Bridge, E. Adenovirus late gene expression does not require a rev-like nuclear rna export pathway. J. Virol. 2000, 74, 6684–6688. [Google Scholar] [CrossRef]
- Johnson, D.C.; Baines, J.D. Herpesviruses remodel host membranes for virus egress. Nat. Rev. Microbiol. 2011, 9, 382–394. [Google Scholar] [CrossRef]
- Mata, M.A.; Satterly, N.; Versteeg, G.A.; Frantz, D.; Wei, S.; Williams, N.; Schmolke, M.; Pena-Llopis, S.; Brugarolas, J.; Forst, C.V.; et al. Chemical inhibition of rna viruses reveals redd1 as a host defense factor. Nat. Chem. Biol. 2011, 7, 712–719. [Google Scholar] [CrossRef]
- Basu, D.; Walkiewicz, M.P.; Frieman, M.; Baric, R.S.; Auble, D.T.; Engel, D.A. Novel influenza virus ns1 antagonists block replication and restore innate immune function. J. Virol. 2009, 83, 1881–1891. [Google Scholar] [CrossRef]
- Konig, R.; Stertz, S.; Zhou, Y.; Inoue, A.; Hoffmann, H.H.; Bhattacharyya, S.; Alamares, J.G.; Tscherne, D.M.; Ortigoza, M.B.; Liang, Y.; et al. Human host factors required for influenza virus replication. Nature 2010, 463, 813–817. [Google Scholar] [CrossRef]
- Waldman, W.J.; Knight, D.A.; Blinder, L.; Shen, J.; Lurain, N.S.; Miller, D.M.; Sedmak, D.D.; Williams, J.W.; Chong, A.S. Inhibition of cytomegalovirus in vitro and in vivo by the experimental immunosuppressive agent leflunomide. Intervirology 1999, 42, 412–418. [Google Scholar] [CrossRef]
- Lin, Y.H.; Yadav, P.; Ravatn, R.; Stollar, V. A mutant of sindbis virus that is resistant to pyrazofurin encodes an altered rna polymerase. Virology 2000, 272, 61–71. [Google Scholar] [CrossRef]
- Bernhoff, E.; Tylden, G.D.; Kjerpeseth, L.J.; Gutteberg, T.J.; Hirsch, H.H.; Rinaldo, C.H. Leflunomide inhibition of bk virus replication in renal tubular epithelial cells. J. Virol. 2010, 84, 2150–2156. [Google Scholar] [CrossRef]
- Qing, M.; Zou, G.; Wang, Q.Y.; Xu, H.Y.; Dong, H.; Yuan, Z.; Shi, P.Y. Characterization of dengue virus resistance to brequinar in cell culture. Antimicrob. Agents Chemother. 2010, 54, 3686–3695. [Google Scholar] [CrossRef]
- Hoffmann, H.H.; Kunz, A.; Simon, V.A.; Palese, P.; Shaw, M.L. Broad–spectrum antiviral that interferes with de novo pyrimidine biosynthesis. Proc. Natl. Acad. Sci. USA 2011, 108, 5777–5782. [Google Scholar]
© 2013 by the authors; licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution license (http://creativecommons.org/licenses/by/3.0/).
Share and Cite
Kuss, S.K.; Mata, M.A.; Zhang, L.; Fontoura, B.M.A. Nuclear Imprisonment: Viral Strategies to Arrest Host mRNA Nuclear Export. Viruses 2013, 5, 1824-1849. https://doi.org/10.3390/v5071824
Kuss SK, Mata MA, Zhang L, Fontoura BMA. Nuclear Imprisonment: Viral Strategies to Arrest Host mRNA Nuclear Export. Viruses. 2013; 5(7):1824-1849. https://doi.org/10.3390/v5071824
Chicago/Turabian StyleKuss, Sharon K., Miguel A. Mata, Liang Zhang, and Beatriz M. A. Fontoura. 2013. "Nuclear Imprisonment: Viral Strategies to Arrest Host mRNA Nuclear Export" Viruses 5, no. 7: 1824-1849. https://doi.org/10.3390/v5071824
APA StyleKuss, S. K., Mata, M. A., Zhang, L., & Fontoura, B. M. A. (2013). Nuclear Imprisonment: Viral Strategies to Arrest Host mRNA Nuclear Export. Viruses, 5(7), 1824-1849. https://doi.org/10.3390/v5071824



