Abstract
The Coronaviridae family, an enveloped RNA virus family, and, more particularly, human coronaviruses (HCoV), were historically known to be responsible for a large portion of common colds and other upper respiratory tract infections. HCoV are now known to be involved in more serious respiratory diseases, i.e. bronchitis, bronchiolitis or pneumonia, especially in young children and neonates, elderly people and immunosuppressed patients. They have also been involved in nosocomial viral infections. In 2002–2003, the outbreak of severe acute respiratory syndrome (SARS), due to a newly discovered coronavirus, the SARS-associated coronavirus (SARS-CoV); led to a new awareness of the medical importance of the Coronaviridae family. This pathogen, responsible for an emerging disease in humans, with high risk of fatal outcome; underline the pressing need for new approaches to the management of the infection, and primarily to its prevention. Another interesting feature of coronaviruses is their potential environmental resistance, despite the accepted fragility of enveloped viruses. Indeed, several studies have described the ability of HCoVs (i.e. HCoV 229E, HCoV OC43 (also known as betacoronavirus 1), NL63, HKU1 or SARS-CoV) to survive in different environmental conditions (e.g. temperature and humidity), on different supports found in hospital settings such as aluminum, sterile sponges or latex surgical gloves or in biological fluids. Finally, taking into account the persisting lack of specific antiviral treatments (there is, in fact, no specific treatment available to fight coronaviruses infections), the Coronaviridae specificities (i.e. pathogenicity, potential environmental resistance) make them a challenging model for the development of efficient means of prevention, as an adapted antisepsis-disinfection, to prevent the environmental spread of such infective agents. This review will summarize current knowledge on the capacity of human coronaviruses to survive in the environment and the efficacy of well-known antiseptic-disinfectants against them, with particular focus on the development of new methodologies to evaluate the activity of new antiseptic-disinfectants on viruses.
1. Introduction
The worldwide epidemic of SARS (Severe Acute Respiratory Syndrome) in 2002–2003, due to a newly discovered coronavirus, the SARS-CoV (SARS-associated coronavirus), reinforced the interest into the Coronaviridae family. Human coronaviruses 229E and OC43 (HCoV 229E and OC43) were previously already known to be responsible for mild and upper respiratory tract diseases. Since then, two further members of this family have been identified (HCoV HUK1 and NL63) and HCoVs have been involved in more serious respiratory tract infections. Moreover, these viruses show an environmental resistance that increases their probability of transfer between contaminated hosts via surfaces, hands, etc. This resistance leads to the urgent need for development of efficient and targeted modes of prevention. As no treatment or vaccines are available to cure HCoVs infections, it is fundamental to dispose of adapted antiseptics-disinfectants, whose efficiency should be rigorously evaluated.
3. HCoVs: Enveloped, but not that Fragile
In this section, we highlight the potency of coronaviruses to survive in different conditions, despite their enveloped nature. This knowledge is essential for a better understanding of the possibility of virus transfer and cross-contamination, and for formulating appropriate infection-control measures. Indeed, despite the fact that transmission was believed to be mainly achieved by direct physical contact with infected patient or by respiratory droplets, several well-described clusters of infection were difficult to explain by these routes. Examples include transmission to 22 persons on an aircraft [102], to 13 guests sharing the same floor of a hotel, and more than 300 persons in an apartment complex [103]. These observations led to some speculations about a possible transmission by other means including surfaces, hands, etc., and to the study of SARS-CoV (and other HCoVs) survival in different conditions.
Despite the fact that this review is devoted to human coronaviruses, some data concerning the murine hepatitis virus (MHV) and the transmissible gastroenteritis virus (TGEV), now called alphacoronavirus 1 [1], are recorded here because they have been used as SARS-CoV surrogates.
3.1. Survival Under Different Conditions of Humidity and Temperature
Some decades ago, a study compared the survival rates of the HCoV 229E to the ones of a non-enveloped virus, the type 1-poliovirus, under different conditions of temperature and humidity. Results are reported in Table 1.

Table 1.
Survival rates of the HCoV 229E and the poliovirus, type 1, under different conditions of temperature and humidity [104].
| HCoV 229E | Type 1-Poliovirus, Sabin strain | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Relative humidity | 20 °C | 6 °C | 20 °C | 6 °C | ||||||
| 15 min | 24 h | 72 h | 6 days | 15 min | 24 h | 15 min | 24 h | 15 min | 24 h | |
| 30% | 87% | 65% | >50% | n.d. | 91% | 65% | 0% | 0% | n.d. | n.d. |
| 50% | 90.9% | 75% | >50% | 20% | 96.5% | 80% | 0% | 0% | n.d. | n.d. |
| 80% | 55% | 3% | 0% | n.d. | 104.8% | 86% | 90% | 30% | n.d. | n.d. |
(n.d.: not done)
Thus, at 20 °C, aerosolized HCoV 229E was found to better survive at 50% relative humidity than at 30%. Indeed, nearly 20% of the original infectious virus was still detectable after six days. High relative humidity seemed less favorable to the virus, unless the temperature came down to 6 °C. At this temperature, the survival of the HCoV 229E was significantly enhanced whatever the rate of relative humidity. This enhanced survival rate at high relative humidity and low temperature may explain the winter propagation of coronaviruses. Moreover, the HCoV 229E survival was significantly higher at 30% and 50% of relative humidity than those of the poliovirus in the same experimental conditions, which could be a striking result according to its non-enveloped nature [104].
Sensitivity of SARS-CoV to temperature has also been assayed. The exposure of the virus to a temperature of 56 °C over 30 min reduced virus titer under an undetectable level, except if SARS-CoV is associated with proteins, such as 20% fetal calf serum (FCS), which bring a protection for the virus. In this case, the temperature needs to reach 60 °C over 30 min to bring virus titer below the detection limit. This emphasizes the importance of organic material in which viruses could be embedded in the real conditions and could protect the virus, mostly from disinfection procedures. When the virus was placed at 4 °C, there was no loss of infectivity [105]. Another study confirms the viral stability at 4 °C, and also at 20 °C and 37 °C for at least 2 hrs, but SARS-CoV lost its infectivity after 90, 60 and 30 min exposure at 56 °C, 67 °C and 75 °C, respectively [106].
3.2. Suspension vs. Desiccation
Coronaviruses also well survive in suspension. At 37 °C, HCoV 229E and OC43 displayed survival rates of 80% and 100%, respectively, in phosphate buffered saline (PBS) over three days and of 30% and 55%, respectively, over six days. These survival rates came down to 50% for HCoV 229E and 30% for HCoV OC43 after three days in culture medium and after ten days, they were of 0% and 10% for each virus, respectively. The same study also showed that desiccation has a more severe effect on coronaviruses. Indeed, in standard environmental conditions (21 °C and 50% to 70% of relative humidity), HCoV 229E infectivity came down to 30% after three hrs of desiccation on various surfaces that can be found in hospital settings, such as aluminum, sterile sponges or surgical latex gloves. HCoV OC43 was more sensitive to desiccation, since its infectivity was below the detectable threshold after three hrs of drying [107].
Rabenau et al. made a comparative study on the stability of different viruses, i.e. SARS-CoV, HCoV 229E, type 1-herpes simplex virus (HSV-1) and the type 3-adenovirus, in suspension and after drying. In medium culture, with and without 10% FCS, the HCoV 229E progressively lost its infectivity over nine days, which is consistent with the previous study. The infectious titers of the three other viruses, including the SARS-CoV, were stable over nine days, with and without proteins. After drying on a plastic surface, the HCoV 229E and the HSV-1 lost their infectivity in 72 hrs, in the presence or absence of FCS. In contrast, the SARS-CoV retained its infectivity for as long as six days, with a further protecting effect of proteins. It took nine days in a dried state, for SARS-CoV to completely lose its infectivity. The adenovirus was the most stable virus assayed as it conserved its infectivity throughout the nine days of the experiment [105].
Some other studies confirm these results. SARS-CoV has been shown to survive after drying on different kinds of materials or diluted in water, revealing a decreased infectivity only after 72 to 96 hrs, depending on the conditions. However, its infectivity is reduced more rapidly if it is deposited on porous surfaces such as cotton or paper [106,108].
Thus, RNA of SARS-CoV was found on different environment samples, such as chair, elevator, computer mouse, etc., and this may have contributed to contamination of health-care workers who had not been in direct contact with SARS-patients [109,110].
A more recent study implicated water and sewage in the transmission of SARS-CoV, taking the MHV and the TGEV as surrogates for their experiments. At 25 °C, the time required for 99% reduction in water was 22 days for TGEV and 17 days for MHV, and, in sewage, it took nine days for TGEV and seven days for MHV. After four weeks in almost the same conditions but at 4 °C, there was less than <1 log10 infectivity decrease for both viruses. The authors concluded that in case of SARS-CoV re-emergence water contaminated with fecal waste should be considered as a potential vehicle of transmission [111].
These studies firmly illustrated the potency of coronaviruses and especially the SARS-CoV, to be transmitted via other routes than respiratory droplets and the likely risk of contamination via surfaces and fomites. It should also be noticed that the residual infectivity of those enveloped viruses in different conditions can almost reach the one of non-enveloped viruses. This reappraises the environmental stability of these two types of viruses.
3.4. Survival in Biological Fluids
As it has been noted earlier, HCoVs are excreted in respiratory secretions but also in other biological fluids such as feces. Knowing and understanding viral survival is then essential to estimate the risk of potential transmission through this route.
Studies have been conducted on SARS-CoV, which was shown to survive at least 96 hrs in sputum, serum and feces. Its infectivity level is nevertheless lower when it is suspended in urines [106]. It is noteworthy that SARS-CoV survival depends on the kind of feces whose pH may vary. Some studies have shown certain surprising results in regard of the previously quoted studies. Indeed, SARS-CoV did not survive beyond 24 hrs in normal feces of an adult or beyond three hrs in newborns’ feces, which is slightly acidic. In contrast, it could survive longer, up to four days, in diarrheic feces whose pH could reach pH 9. The same study revealed a SARS-CoV survival until four to five days in respiratory specimen [108,117].
According to these data, transfer of viruses and cross-contamination should be carefully considered. Indeed, under certain circumstances, for instance in health-care settings, contamination of inanimate materials or other people by infectious respiratory secretions or other body fluids (saliva, urine or feces) seems to play a role in SARS-CoV transmission, and it is likely the same for the other HCoVs. Thus, it is essential to dispose of adapted, targeted and efficient ways of disinfection whose efficiency has to be correctly evaluated.
4. Antisepsis-Disinfection: An Efficient Weapon, with Room for Improvement
4.1. How Prevention Measures Halted the Propagation of SARS-CoV
The absence of treatment, the high mortality rate and the transmission patterns of SARS-CoV involved the setting of powerful and coordinated means of prevention to stop the worldwide spread of this virus. Indeed, the SARS-CoV epidemic has been brought under control thanks to basic public health measures, including rapid case detection and isolation, contact tracing, quarantine and good precautionary control measures (hand washing, use of personal protective equipment) [54,59]. Additionally, the WHO expressed recommendations for travelers coming from areas affected by the SARS with screening of potential cases and in-flight care of suspected cases followed by aircraft disinfection [121].
Thus, besides these standard measures, our knowledge on HCoVs sensitivity to antiseptics-disinfectants should improve, in order to use these fundamental prevention tools in a targeted and coherent manner.
4.2 What is Antisepsis-Disinfection and how do we Evaluate its Efficiency?
Facing the lack in a specific antiviral treatment, it is necessary to develop new means of prevention and to ensure that the existing ones are efficient according to the field situation. Proper evaluation of the efficiency of antiseptics-disinfectants on viruses is thus crucial.
Essentially, antiseptic-disinfectant antiviral activity is evaluated by combining viruses and the product to be tested for an appropriately defined and precise contact time, according to the expected use of the product (surface or hands disinfection, for instance). Product activity and its eventual cytotoxicity are then neutralized and the loss of viral infectivity due to the product activity is estimated. Neutralization of the antiseptic-disinfectant activity plays a key role in the test procedure; it ensures a precise contact time, the elimination of the residual activity and cytotoxicity of the tested product, and the successful recovery of viruses that are not killed by the product. These tests require appropriate controls, especially to check the absence of interference on viral infectivity, due to the test itself. It is also important to test the efficiency of neutralization, removal of cytotoxicity under reproducible and well-defined test conditions (e.g., contact time and environmental temperature). A germicide can be considered to have an efficient antiseptic-disinfectant antiviral activity if it induces, in a well-defined contact time, a reduction in viral titers higher than 3 or 4 log10, depending on American and European regulatory agencies, respectively [122,123].
4.4. International Standardization Context
One of the challenges of antiviral antiseptics-disinfectants testing is the standardization to obtain valuable and comparable results. This is illustrated in the next section, where, even if the results concerning the activity of antiseptics-disinfectants on HCoVs are generally consistent with each other, they are still difficult to compare. It is then extremely important to set standards to test these antiseptic activities.
To date, only one European Standard (NF EN 14476+A1) on virucidal antiseptic-disinfectant activity testing in human medicine has been published [122]. This protocol, from January 2007, specifies the test method and the minimum requirements to establish virucidal activity according to the potential use of the products tested, e.g. disinfection of surfaces and instruments, hygienic hand wash or thermochemical disinfection. Virus strains, temperatures, contact times and interfering substances are specified for each potential use. According to this standard, a product is considered to have an antiseptic-disinfectant antiviral activity if it induces a loss of infectivity of at least 4 log10 in viral titers during an accurate contact time.
In the United States, the principal standard is relatively close to the European one but it specifies an efficacy criterion of 3 log10. Several standards have been then published to cover the different field situations such as two standards concerning the evaluation of hygienic hand wash, a standard concerning the evaluation of efficacy of virucidal agents intended for inanimate environmental surfaces and, finally, a specific standard concerning the neutralization step [123,124,125,126,127].
4.5. Sensitivity of HCoVs to Antiseptics-Disinfectants
4.5.1. Sensitivity of “Classic” HCoVs (other than SARS-CoV) to Antiseptics-Disinfectants
A study, by Sattar et al., evaluated the efficiency of 15 antiseptics-disinfectants of various chemical families on four different viruses: two non-enveloped viruses (type b-coxsackievirus and type 5-adenovirus) and two enveloped viruses (HCoV 229E and type 3-parainfluenzavirus). With this aim in view, viral inocula were suspended in feces or mucin to mimic organic matter and left to dry on stainless-steel disks. The contact time was 1 min and the efficacy criterion was a reduction in viral titers of 3 log10. Results are gathered in Table 2.

Table 2.
Comparison of non-enveloped and enveloped viruses (HCoV 229E, type 3-parainfluenzavirus, type b-coxsackievirus and type 5-adenovirus) sensitivity to different antiseptics-disinfectants formulations, thanks to carrier tests [128]. The efficiency is validated if the reduction in viral titers after a contact-time of 1 min is ≥ 3 log10.
| Tested antiseptics-disinfectants | Concentration (%) - (pH at used concentration) | HCoV 229E | Type 3-parainfluenzavirus | Type B-Coxsackievirus | Type 5-Adenovirus |
|---|---|---|---|---|---|
| Enveloped | Enveloped | Non-enveloped | Non-enveloped | ||
| Halogenous compounds | |||||
| Sodium hypochlorite | 0.01 (8.0) | No | No | No | No |
| 0.10 (9.4) | Yes | Yes | No | No | |
| 0.50 (11.0) | Yes | Yes | Yes | Yes | |
| Chloramine T | 0.01 (7.0) | No | Yes | No | No |
| 0.10 (8.0) | Yes | No | No | No | |
| 0.30 (8.0) | Yes | Yes | Yes | Yes | |
| Sodium hypochlorite and potassium bromide | 0.01 (10.0) | No | No | No | No |
| 0.05 (11.5) | Yes | Yes | No | No | |
| 0.10 (12.0) | Yes | Yes | No | No | |
| Povidone-iodine | 10.0 (3.0) (1% available iodine) | Yes | Yes | No | No |
| Ethanol | 70.0 (4.0) | Yes | Yes | No | Yes |
| Glutaraldehyde | 2.0 (7.0) | Yes | Yes | Yes | Yes |
| Quaternary ammonium compounds | |||||
| n-alkyl-dimethylbenzyl chloride | 0.04 (6.0) | No | No | No | No |
| n-alkyl-dimethylbenzyl chloride | 0.04 (1.0) | Yes | Yes | Yes | Yes |
| + HCl | 7.00 | ||||
| n-alkyl-dimethylbenzyl chloride | 0.04 (5.0) | Yes | Yes | No | Yes |
| + ethanol | 70.0 | ||||
| n-alkyl-dimethylbenzyl chloride | 0.04 (11.0) | Yes | Yes | No | Yes |
| + sodium metasilicate | 0.5 | ||||
| Chlorhexidine gluconate | 0.008 (5.0) | No | Yes | No | No |
| + cetrimide | 0.08 | ||||
| Chlorhexidine gluconate | 0.05 (4.5) | Yes | Yes | No | Yes |
| + cetrimide | 0.50 | ||||
| + ethanol | 70.0 | ||||
| Phenolic compounds | |||||
| o-phenylphenol | 0.02 (9.0) | No | No | No | No |
| + o-benzyl-chlorophenol | 0.03 | ||||
| + p-tert-amylphenol | 0.01 | ||||
| o-phenylphenol | 0.02 (9.0) | Yes | Yes | No | No |
| + o-benzyl-chlorophenol l | 0.03 | ||||
| + p-tert-amylpheno | 0.01 | ||||
| + SDS | 0.60 | ||||
| o-phenylphenol | 0.02 (9.0) | Yes | Yes | No | Yes |
| + o-benzyl-chlorophenol | 0.03 | ||||
| + p-tert-amylphenol | 0.01 | ||||
| + ethanol | 70.0 | ||||
| Sodium o-benzyl-p-chlorophenate | 0.50 (13.0) | Yes | Yes | Yes | Yes |
| +Sodium dodecyl sulfate | 0.60 |
This study highlighted the fact that enveloped viruses are more sensitive than non-enveloped viruses to the action of antiseptics-disinfectants, despite sensitivity discrepancies within each group. However, enveloped viruses are not that fragile and they are not inactivated by a number of antiseptics-disinfectants such as quaternary ammoniums compounds or phenolic compounds. The association chlorhexidine and cetrimide, widely used in human medicine, did not seem to be effective on HCoV 229E, except if ethanol is added [128].
A more recent study investigated the action of antiseptics-disinfectants on HCoVs 229E and OC43 with suspension tests and contact times of 5 min. The neutralization step was achieved by dilution in medium culture. The povidone-iodine (0.75% free iodine) caused a 50% reduction in infectivity of both of the viruses, which is not enough to claim a virucidal activity. Moreover, to obtain a 50% reduction in HCoV 229E titers, tenfold increase in concentration of povidone-iodine was required. Some other products (70% ethanol, soap or 5% bleach) were assayed but without success because they interfered with the biological viral titration assay [107].
This also highlights the importance of the neutralization step and the necessity of developing means to eliminate the toxicity of the tested products.
This result was also confirmed on the SARS-CoV by Kariwa et al. who tested different formulations of povidone-iodine with suspensions tests and contact times of 1 and 2 min. The neutralization step was achieved chemically by the addition of sodium thiosulfate. All formulations reduced the viral infectivity under the detectable level after 2 min of contact-time. The same result was obtained with 70% ethanol in 1 min [129].
Two other studies conducted in our laboratory concerned the HCoV 229E and its sensitivity to two widely used antiseptics, chlorhexidine and hexamidine, but also to new molecules belonging to the calixarene family [130,131]. In these studies, antiseptic antiviral activities were assayed thanks to suspension tests and the efficacy criterion was a reduction of 4 log10, as recommended by the European Standard [122]. A novel methodology of gel filtration for the neutralization step was developed in these studies, using homemade and reproducible Sephadex™ columns.
Chlorhexidine was shown to have a time and concentration-dependent anti-HCoV 229E activity allowing a 3 log10 reduction, but only after a 60 min contact time (Figure 1a). It was then not sufficient to claim an antiseptic anti-HCoV 229E activity. Hexamidine did not show any activity against HCoV 229E [130,131]. These results highlighted the necessity of (i) evaluating the activity of commonly used antiseptics-disinfectants against different viruses, to be sure of their efficiency and to develop a targeted antisepsis, and (ii) developing new active noncytotoxic molecules.
The second study concerned the antiseptic anti-HCoV 229E activity of two calixarenic compounds, i.e. the tetra-para-sulfonato-calix[4]arene (C[4]S) and the 1,3-bis(bithiazolyl)-tetra-para-sulfonato-calix[4]arene (C[4]S-BTZ) [130,131]. These molecules were attractive targets at first because they did not show any cytotoxicity. Then, the C[4]S-BTZ showed an equivalent, and even better, activity than that of chlorhexidine. Indeed, its activity reached almost 3 log10 reduction in viral titers from 5 min of contact time (Figure 1b). Some further studies are needed, but calixarenes appear as interesting candidates to be new antiseptics-disinfectants.
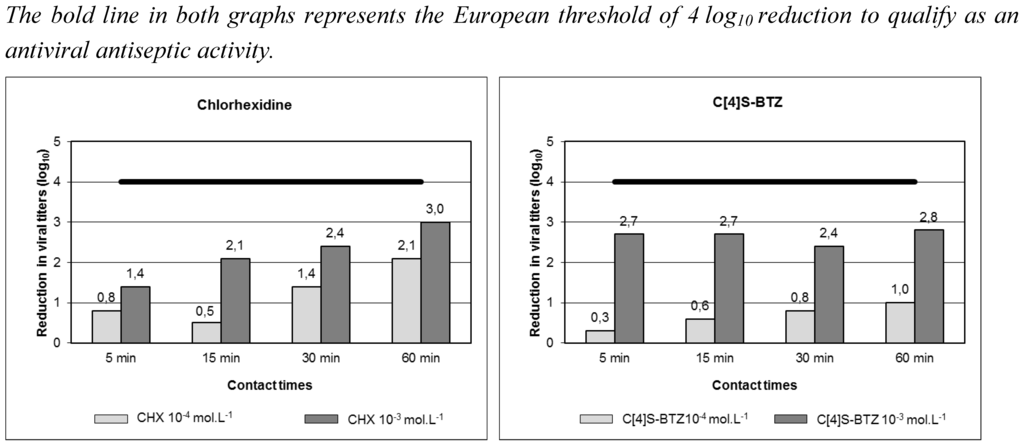
Figure 1.
Evaluation of antiseptic HCoV 229E activity of (a) chlorhexidine (CHX) and (b) the 1,3-bis(bithiazolyl)-tetra-para-sulfonato-calix[4]arene (C[4]S-BTZ) [130,131].
4.5.2. SARS-CoV Sensitivity to Antiseptics-Disinfectants
Rabenau et al. achieved a study using suspension tests with different organic loads (albumin, FCS or sheep erythrocytes) and following the recommendations of the European Standard [122]. Most of the tested alcoholic-based solutions (isopropanol or ethanol) has been shown to allow a reduction > 4 log10 in viral titers over 30 sec, whatever the added organic load. They also investigated the activity of three surface and instrument disinfectants (one based on benzalkonium chloride and laurylamine; one based on benzalkonium chloride, glutaraldehyde and didecyldimonium chloride; and one based on magnesium monoperphthalate). Contact times were then, still in accordance to the European Standard, 30 and 60 min. SARS-CoV was inactivated by all the disinfectants to below the limit of detection (the smaller reduction factor was 3.25 log10), regardless of the type of organic load [132]. The same team pursued its investigation evaluating the SARS-CoV virucidal activity of different disinfectants based on alcohols (propanol, ethanol used for hands disinfection), aldehydes (formaldehyde, glutardialdehyde), glucoprotamin and wine vinegar. The methodology was the same that previously described, except for the organic load, which was FCS. In case of cytotoxic effect after the dilution-neutralization step, the virus-disinfectant mixture was membrane filtered. This allowed the concentration of the viral particles, which could then be tittered, while retaining the disinfectant. The results are recorded in Table 3. The variation in reduction factors was due to the filtration used as neutralization step when disinfectant toxicity was too strong [105].

Table 3.
Virucidal activity on SARS-CoV of different hand-rub formulations and surfaces disinfectants thanks to suspension tests [105].
| Tested formulations | Contact times | Minimal reduction factor (log10) |
|---|---|---|
| 100% 2-propanol | 30 s | ≥ 3.31 |
| 70% 2-propanol | 30 s | ≥ 3.31 |
| 78% ethanol | 30 s | ≥ 5.01 |
| 45% 2-propanol, 30% 1-propanol | 30 s | ≥ 2.78 |
| Wine vinegar | 60 s | ≥ 3.0 |
| 0.7% formaldehyde | 2 min | ≥ 3.01 |
| 1.0% formaldehyde | 2 min | ≥ 3.01 |
| 0.5% glutardialdehyde | 2 min | ≥ 4.01 |
| 26% glucoprotamin | 2 min | ≥ 1.68 |
Recently, a study used MHV and TGEV as SARS-CoV surrogates. Thanks to carrier tests on stainless steel surfaces and a chemical neutralization step, the anti-SARS-CoV efficacy of six different formulations was evaluated. The efficacy criterion was a reduction of 3 log10 in viral titers after 1 min contact time. Results are reported in Table 4.

Table 4.
Virucidal activity on MHV and TGEV, used as SARS-CoV surrogates, of different hand-rub formulations and surface disinfectants using carrier test methodology [133] (MHV: Murine hepatitis virus, TGEV: Transmissible gastro-enteritis virus).
| Concentration of active ingredients of the tested commercial formulations | MHV | TGEV |
|---|---|---|
| Bleach (6% sodium hypochlorite – use dilution: 1:100, ≈ 600 mg/mL) | No | No |
| 9.09% o-phenylphenol, 7.66% p-tertiary amylphenol | No | No |
| 0.55% ortho-phthalaldehyde | No | No |
| 70% ethanol | Yes | Yes |
| 62% ethanol | No | Yes |
| 71% ethanol | No | Yes |
This study revealed first that there were some behavioral differences between the two viruses chosen as surrogates. This raises the question of the pertinence of surrogates use. However, SARS-CoV is a virus which requires a level 3 containment laboratory. Therefore, virus surrogates allow laboratories, which do not dispose of this type of equipment, to conduct studies and produce precious data without working on a virus, which had already caused a worldwide epidemic.
Another important point revealed by this study is the inefficiency of bleach, a widely used disinfectant, when applied at the 1:100 (0.06%) use-dilution prescribed by the manufacturer. Sattar et al., whose results are recorded in Table 2, have found higher reductions of HCoV 229E viral titers with concentrations of hypochlorite greater than the one tested here. These results are then consistent with a concentration-dependent effect [133].
Another recent study used MHV as the SARS-CoV surrogate, and carrier tests on Petri dishes. Antiseptic antiviral activity of common household disinfectants or antiseptics, containing either 0.05% of triclosan, 0.12% of chloroxylenol, 0.21% of sodium hypochlorite, 0.23% of pine oil, or 0.10% of a quaternary compound with 79.0% of ethanol, were investigated. All of them provided at least a 3 log10 reduction in viral titers within a 30 sec contact time, which is consistent with the previous results [134].
Despite the fact that these studies bring vital information, they also highlight the necessity of standardization of the antiseptics-disinfectants activity evaluation. We should also develop in-field tests in order to have a better appreciation of the true action of antiseptics-disinfectants.
5. Conclusions
The four HCoVs 229E, OC43, NL63 and HKU1 cause mild respiratory illnesses compared to SARS-CoV, but these infectious agents are involved in 10 to 20% of hospitalizations of young children and immunocompromised adults with respiratory tract illness and they are also involved in nosocomial infections. Moreover, although the SARS-epidemic has been contained, the possibility of re-emergence of SARS-CoV or emergence of another zoonotic strain remains.
Besides the absence of specific treatment and vaccine, HCoVs are now known to show a significant environmental resistance. Their survival in different biological fluids such as respiratory secretions or feces has been proved. Furthermore, some parameters seem of benefit for HCoVs such as the stabilizing effect of low temperature and high relative humidity or the protective action of organic materials. This protective effect should be carefully considered when developing antiseptic-disinfection strategies. Indeed, this often involves a higher quantity and/or concentration of the antiseptic-disinfectant product and so, a higher toxicity. Thus, an efficient disinfection process should include a precleaning step to get rid of these organic materials. The old well-known principle of antisepsis-disinfection that only clean things can be efficiently disinfected is still valuable.
Finally, in regard to the different studies on HCoVs’s sensitivity to antiseptics-disinfectants, only few formulations are efficient within an adapted contact time and without a too-strong toxicity. For instance, considering their lack of efficiency against HCoVs, and also their toxicity, products only based on quaternary ammoniums or phenolic compounds should be avoided. Some largely used antiseptics-disinfectants such as ethanol or bleach show a significant activity on the HCoVs. However, some critical parameters should be considered, especially in the case of chlorine-derived compounds, such as the presence of organic materials that could prevent their antiseptic activity, or their dose-dependent effect on the HCoVs. The povidone-iodine or the chlorhexidine, when associated to ethanol and/or cetrimide, could be recommended when there is a risk of HCoVs contamination, contrary to another widely used antiseptic, the hexamidine.
It is now essential to pursue investigations on (i) HCoVs’s environmental stability and the role of inanimate material in their spread, (ii) their sensitivity to antiseptics-disinfectants formulations in standardized and targeted conditions, and (iii) the development of new efficient and nontoxic antiseptic-disinfectant molecules such as the calixarenic compounds.
Acknowledgments
Financial support was provided by the French Ministry for Education and Research and the French National Scientific Research Center.
Conflict of Interest
The authors declare no conflict of interest.
References
- ICTV (International Committee on Taxonomy on Viruses). Virus Taxonomy: 2011 Release (current). Available online: http://ictvonline.org/virusTaxonomy.asp?version=2011 (Accessed on 24 September 2012).
- Almeida, J.D.; Tyrrell, D.A. The morphology of three previously uncharacterized human respiratory viruses that grow in organ culture. J. Gen. Virol. 1967, 1, 175–178. [Google Scholar] [CrossRef]
- Bradburne, A.F.; Bynoe, M.L.; Tyrrell, D.A. Effects of a "new" human respiratory virus in volunteers. Br. Med. J. 1967, 3, 767–769. [Google Scholar] [CrossRef]
- Hamre, D.; Procknow, J.J. A new virus isolated from the human respiratory tract. Proc. Soc. Exp. Biol. Med. 1966, 121, 190–193. [Google Scholar]
- McIntosh, K.; Dees, J.H.; Becker, W.B.; Kapikian, A.Z.; Chanock, R.M. Recovery in tracheal organ cultures of novel viruses from patients with respiratory disease. Proc. Natl. Acad. Sci. USA 1967, 57, 933–940. [Google Scholar] [CrossRef]
- Larson, H.E.; Reed, S.E.; Tyrrell, D.A. Isolation of rhinoviruses and coronaviruses from 38 colds in adults. J. Med. Virol. 1980, 5, 221–229. [Google Scholar] [CrossRef]
- Esper, F.; Weibel, C.; Ferguson, D.; Landry, M.L.; Kahn, J.S. Evidence of a novel human coronavirus that is associated with respiratory tract disease in infants and young children. J. Infect. Dis. 2005, 191, 492–498. [Google Scholar] [CrossRef]
- Fouchier, R.A.; Hartwig, N.G.; Bestebroer, T.M.; Niemeyer, B.; de Jong, J.C.; Simon, J.H.; Osterhaus, A.D. A previously undescribed coronavirus associated with respiratory disease in humans. Proc. Natl. Acad. Sci. USA 2004, 101, 6212–6216. [Google Scholar]
- Van der Hoek, L.; Pyrc, K.; Jebbink, M.F.; Vermeulen-Oost, W.; Berkhout, R.J.; Wolthers, K.C.; Wertheim-van Dillen, P.M.; Kaandorp, J.; Spaargaren, J.; Berkhout, B. Identification of a new human coronavirus. Nat. Med. 2004, 10, 368–373. [Google Scholar]
- Woo, P.C.; Lau, S.K.; Chu, C.M.; Chan, K.H.; Tsoi, H.W.; Huang, Y.; Wong, B.H.; Poon, R.W.; Cai, J.J.; Luk, W.K.; et al. Characterization and complete genome sequence of a novel coronavirus, coronavirus HKU1, from patients with pneumonia. J. Virol. 2005, 79, 884–895. [Google Scholar]
- Drosten, C.; Gunther, S.; Preiser, W.; van der Werf, S.; Brodt, H.R.; Becker, S.; Rabenau, H.; Panning, M.; Kolesnikova, L.; Fouchier, R.A.; et al. Identification of a novel coronavirus in patients with severe acute respiratory syndrome. N. Engl. J. Med. 2003, 348, 1967–1976. [Google Scholar] [CrossRef]
- Ksiazek, T.G.; Erdman, D.; Goldsmith, C.S.; Zaki, S.R.; Peret, T.; Emery, S.; Tong, S.; Urbani, C.; Comer, J.A.; Lim, W.; et al. A novel coronavirus associated with severe acute respiratory syndrome. N. Engl. J. Med. 2003, 348, 1953–1966. [Google Scholar] [CrossRef]
- Peiris, J.S.; Lai, S.T.; Poon, L.L.; Guan, Y.; Yam, L.Y.; Lim, W.; Nicholls, J.; Yee, W.K.; Yan, W.W.; Cheung, M.T.; et al. Coronavirus as a possible cause of severe acute respiratory syndrome. Lancet 2003, 361, 1319–1325. [Google Scholar]
- WHO multicentre collaborative network for Severe Acute Respiratory Syndrome diagnosis. A multicentre collaboration to investigate the cause of severe acute respiratory syndrome. Lancet 2003, 361, 1730–1733. [CrossRef]
- Arden, K.E.; Nissen, M.D.; Sloots, T.P.; Mackay, I.M. New human coronavirus, HCoV-NL63, associated with severe lower respiratory tract disease in Australia. J. Med. Virol. 2005, 75, 455–462. [Google Scholar] [CrossRef]
- Bastien, N.; Anderson, K.; Hart, L.; Van Caeseele, P.; Brandt, K.; Milley, D.; Hatchette, T.; Weiss, E.C.; Li, Y. Human coronavirus NL63 infection in Canada. J. Infect. Dis. 2005, 191, 503–506. [Google Scholar] [CrossRef]
- Vabret, A.; Mourez, T.; Dina, J.; van der Hoek, L.; Gouarin, S.; Petitjean, J.; Brouard, J.; Freymuth, F. Human coronavirus NL63, France. Emerg. Infect. Dis. 2005, 11, 1225–1229. [Google Scholar] [CrossRef]
- Van der Hoek, L.; Sure, K.; Ihorst, G.; Stang, A.; Pyrc, K.; Jebbink, M.F.; Petersen, G.; Forster, J.; Berkhout, B.; Uberla, K. Croup is associated with the novel coronavirus NL63. PLoS Med. 2005, 2, e240. [Google Scholar] [CrossRef]
- Gerna, G.; Percivalle, E.; Sarasini, A.; Campanini, G.; Piralla, A.; Rovida, F.; Genini, E.; Marchi, A.; Baldanti, F. Human respiratory coronavirus HKU1 versus other coronavirus infections in Italian hospitalised patients. J. Clin. Virol. 2007, 38, 244–250. [Google Scholar] [CrossRef]
- Sloots, T.P.; McErlean, P.; Speicher, D.J.; Arden, K.E.; Nissen, M.D.; Mackay, I.M. Evidence of human coronavirus HKU1 and human bocavirus in Australian children. J. Clin. Virol. 2006, 35, 99–102. [Google Scholar] [CrossRef]
- Chiu, S.S.; Chan, K.H.; Chu, K.W.; Kwan, S.W.; Guan, Y.; Poon, L.L.; Peiris, J.S. Human coronavirus NL63 infection and other coronavirus infections in children hospitalized with acute respiratory disease in Hong Kong, China. Clin. Infect. Dis. 2005, 40, 1721–1729. [Google Scholar] [CrossRef]
- Kon, M.; Watanabe, K.; Tazawa, T.; Tamura, T.; Tsukagoshi, H.; Noda, M.; Kimura, H.; Mizuta, K. Detection of human coronavirus NL63 and OC43 in children with acute respiratory infections in Niigata, Japan, between 2010 and 2011. Jpn. J. Infect. Dis. 2012, 65, 270–272. [Google Scholar] [CrossRef]
- Gerna, G.; Campanini, G.; Rovida, F.; Percivalle, E.; Sarasini, A.; Marchi, A.; Baldanti, F. Genetic variability of human coronavirus OC43-, 229E-, and NL63-like strains and their association with lower respiratory tract infections of hospitalized infants and immunocompromised patients. J. Med. Virol. 2006, 78, 938–949. [Google Scholar] [CrossRef]
- Esposito, S.; Bosis, S.; Niesters, H.G.; Tremolati, E.; Begliatti, E.; Rognoni, A.; Tagliabue, C.; Principi, N.; Osterhaus, A.D. Impact of human coronavirus infections in otherwise healthy children who attended an emergency department. J. Med. Virol. 2006, 78, 1609–1615. [Google Scholar] [CrossRef]
- Vabret, A.; Dina, J.; Gouarin, S.; Petitjean, J.; Tripey, V.; Brouard, J.; Freymuth, F. Human (non-severe acute respiratory syndrome) coronavirus infections in hospitalised children in France. J. Paediatr. Child. Health. 2008, 44, 176–181. [Google Scholar] [CrossRef]
- Talbot, H.K.; Shepherd, B.E.; Crowe, J.E., Jr.; Griffin, M.R.; Edwards, K.M.; Podsiad, A.B.; Tollefson, S.J.; Wright, P.F.; Williams, J.V. The pediatric burden of human coronaviruses evaluated for twenty years. Pediatr. Infect. Dis. J. 2009, 28, 682–687. [Google Scholar] [CrossRef]
- Vabret, A.; Mourez, T.; Gouarin, S.; Petitjean, J.; Freymuth, F. An outbreak of coronavirus OC43 respiratory infection in Normandy, France. Clin. Infect. Dis. 2003, 36, 985–989. [Google Scholar] [CrossRef]
- Van Elden, L.J.; van Loon, A.M.; van Alphen, F.; Hendriksen, K.A.; Hoepelman, A.I.; van Kraaij, M.G.; Oosterheert, J.J.; Schipper, P.; Schuurman, R.; Nijhuis, M. Frequent detection of human coronaviruses in clinical specimens from patients with respiratory tract infection by use of a novel real-time reverse-transcriptase polymerase chain reaction. J. Infect. Dis. 2004, 189, 652–657. [Google Scholar] [CrossRef]
- Riski, H.; Hovi, T. Coronavirus infections of man associated with diseases other than the common cold. J. Med. Virol. 1980, 6, 259–265. [Google Scholar] [CrossRef]
- Talbot, H.K.; Crowe, J.E., Jr.; Edwards, K.M.; Griffin, M.R.; Zhu, Y.; Weinberg, G.A.; Szilagyi, P.G.; Hall, C.B.; Podsiad, A.B.; Iwane, M.; et al. Coronavirus infection and hospitalizations for acute respiratory illness in young children. J. Med. Virol. 2009, 81, 853–856. [Google Scholar] [CrossRef]
- Woo, P.C.; Lau, S.K.; Tsoi, H.W.; Huang, Y.; Poon, R.W.; Chu, C.M.; Lee, R.A.; Luk, W.K.; Wong, G.K.; Wong, B.H.; et al. Clinical and molecular epidemiological features of coronavirus HKU1-associated community-acquired pneumonia. J. Infect. Dis. 2005, 192, 1898–1907. [Google Scholar] [CrossRef]
- Gagneur, A.; Sizun, J.; Vallet, S.; Legr, M.C.; Picard, B.; Talbot, P.J. Coronavirus-related nosocomial viral respiratory infections in a neonatal and paediatric intensive care unit: a prospective study. J. Hosp. Infect. 2002, 51, 59–64. [Google Scholar] [CrossRef]
- Sizun, J.; Soupre, D.; Legrand, M.C.; Giroux, J.D.; Rubio, S.; Cauvin, J.M.; Chastel, C.; Alix, D.; de Parscau, L. Neonatal nosocomial respiratory infection with coronavirus: a prospective study in a neonatal intensive care unit. Acta Paediatr. 1995, 84, 617–620. [Google Scholar] [CrossRef]
- Falsey, A.R.; Walsh, E.E.; Hayden, F.G. Rhinovirus and coronavirus infection-associated hospitalizations among older adults. J. Infect. Dis. 2002, 185, 1338–1341. [Google Scholar] [CrossRef]
- Nicholson, K.G.; Kent, J.; Hammersley, V.; Cancio, E. Acute viral infections of upper respiratory tract in elderly people living in the community: comparative, prospective, population based study of disease burden. BMJ 1997, 315, 1060–1064. [Google Scholar] [CrossRef]
- Pene, F.; Merlat, A.; Vabret, A.; Rozenberg, F.; Buzyn, A.; Dreyfus, F.; Cariou, A.; Freymuth, F.; Lebon, P. Coronavirus 229E-related pneumonia in immunocompromised patients. Clin. Infect. Dis. 2003, 37, 929–932. [Google Scholar] [CrossRef]
- Folz, R.J.; Elkordy, M.A. Coronavirus pneumonia following autologous bone marrow transplantation for breast cancer. Chest 1999, 115, 901–905. [Google Scholar] [CrossRef]
- Chany, C.; Moscovici, O.; Lebon, P.; Rousset, S. Association of coronavirus infection with neonatal necrotizing enterocolitis. Pediatrics 1982, 69, 209–214. [Google Scholar]
- Vabret, A.; Dina, J.; Gouarin, S.; Petitjean, J.; Corbet, S.; Freymuth, F. Detection of the new human coronavirus HKU1: a report of 6 cases. Clin. Infect. Dis. 2006, 42, 634–639. [Google Scholar] [CrossRef]
- Zhang, X.M.; Herbst, W.; Kousoulas, K.G.; Storz, J. Biological and genetic characterization of a hemagglutinating coronavirus isolated from a diarrhoeic child. J. Med. Virol. 1994, 44, 152–161. [Google Scholar] [CrossRef]
- Arbour, N.; Cote, G.; Lachance, C.; Tardieu, M.; Cashman, N.R.; Talbot, P.J. Acute and persistent infection of human neural cell lines by human coronavirus OC43. J. Virol. 1999, 73, 3338–3350. [Google Scholar]
- Arbour, N.; Ekande, S.; Cote, G.; Lachance, C.; Chagnon, F.; Tardieu, M.; Cashman, N.R.; Talbot, P.J. Persistent infection of human oligodendrocytic and neuroglial cell lines by human coronavirus 229E. J. Virol. 1999, 73, 3326–3337. [Google Scholar]
- Bonavia, A.; Arbour, N.; Yong, V.W.; Talbot, P.J. Infection of primary cultures of human neural cells by human coronaviruses 229E and OC43. J. Virol. 1997, 71, 800–806. [Google Scholar]
- Arbour, N.; Day, R.; Newcombe, J.; Talbot, P.J. Neuroinvasion by human respiratory coronaviruses. J. Virol. 2000, 74, 8913–8921. [Google Scholar] [CrossRef]
- Murray, R.S.; Brown, B.; Brian, D.; Cabirac, G.F. Detection of coronavirus RNA and antigen in multiple sclerosis brain. Ann. Neurol. 1992, 31, 525–533. [Google Scholar] [CrossRef]
- Stewart, J.N.; Mounir, S.; Talbot, P.J. Human coronavirus gene expression in the brains of multiple sclerosis patients. Virology 1992, 191, 502–505. [Google Scholar] [CrossRef]
- St-Jean, J.R.; Jacomy, H.; Desforges, M.; Vabret, A.; Freymuth, F.; Talbot, P.J. Human respiratory coronavirus OC43: genetic stability and neuroinvasion. J. Virol. 2004, 78, 8824–8834. [Google Scholar] [CrossRef]
- Riski, H.; Hovi, T.; Frick, M.H. Carditis associated with coronavirus infection. Lancet 1980, 2, 100–101. [Google Scholar]
- Hsu, L.Y.; Lee, C.C.; Green, J.A.; Ang, B.; Paton, N.I.; Lee, L.; Villacian, J.S.; Lim, P.L.; Earnest, A.; Leo, Y.S. Severe acute respiratory syndrome (SARS) in Singapore: clinical features of index patient and initial contacts. Emerg. Infect. Dis. 2003, 9, 713–717. [Google Scholar] [CrossRef]
- Poutanen, S.M.; Low, D.E.; Henry, B.; Finkelstein, S.; Rose, D.; Green, K.; Tellier, R.; Draker, R.; Adachi, D.; Ayers, M.; et al. Identification of severe acute respiratory syndrome in Canada. N. Engl. J. Med. 2003, 348, 1995–2005. [Google Scholar] [CrossRef]
- WHO (World Health Organization). Summary of probable SARS cases with onset of illness from 1 November 2002 to 31 July. Available online: http://www.who.int/csr/sars/country/table2004_04_21/en/index.html (Accessed on 21 February 2010).
- Lee, N.; Hui, D.; Wu, A.; Chan, P.; Cameron, P.; Joynt, G.M.; Ahuja, A.; Yung, M.Y.; Leung, C.B.; To, K.F.; et al. A major outbreak of severe acute respiratory syndrome in Hong Kong. N. Engl. J. Med. 2003, 348, 1986–1994. [Google Scholar] [CrossRef]
- Varia, M.; Wilson, S.; Sarwal, S.; McGeer, A.; Gournis, E.; Galanis, E.; Henry, B. Investigation of a nosocomial outbreak of severe acute respiratory syndrome (SARS) in Toronto, Canada. CMAJ 2003, 169, 285–292. [Google Scholar]
- Zhao, Z.; Zhang, F.; Xu, M.; Huang, K.; Zhong, W.; Cai, W.; Yin, Z.; Huang, S.; Deng, Z.; Wei, M.; et al. Description and clinical treatment of an early outbreak of severe acute respiratory syndrome (SARS) in Guangzhou, PR China. J. Med. Microbiol. 2003, 52, 715–720. [Google Scholar] [CrossRef]
- Booth, C.M.; Matukas, L.M.; Tomlinson, G.A.; Rachlis, A.R.; Rose, D.B.; Dwosh, H.A.; Walmsley, S.L.; Mazzulli, T.; Avendano, M.; Derkach, P.; et al. Clinical features and short-term outcomes of 144 patients with SARS in the greater Toronto area. JAMA 2003, 289, 2801–2809. [Google Scholar] [CrossRef]
- Donnelly, C.A.; Ghani, A.C.; Leung, G.M.; Hedley, A.J.; Fraser, C.; Riley, S.; Abu-Raddad, L.J.; Ho, L.M.; Thach, T.Q.; Chau, P.; et al. Epidemiological determinants of spread of causal agent of severe acute respiratory syndrome in Hong Kong. Lancet 2003, 361, 1761–1766. [Google Scholar]
- Peiris, J.S.; Chu, C.M.; Cheng, V.C.; Chan, K.S.; Hung, I.F.; Poon, L.L.; Law, K.I.; Tang, B.S.; Hon, T.Y.; Chan, C.S.; et al. Clinical progression and viral load in a community outbreak of coronavirus-associated SARS pneumonia: a prospective study. Lancet 2003, 361, 1767–1772. [Google Scholar]
- Peiris, J.S.; Yuen, K.Y.; Osterhaus, A.D.; Stohr, K. The severe acute respiratory syndrome. N. Engl. J. Med. 2003, 349, 2431–2441. [Google Scholar] [CrossRef]
- WHO (World Health Organization). Consensus document on the epidemiology of severe acute respiratory syndrome (SARS). Available online: http://www.who.int/csr/sars/en/WHOconsensus.pdf (Accessed on 25-09-12).
- Hamming, I.; Timens, W.; Bulthuis, M.L.; Lely, A.T.; Navis, G.J.; van Goor, H. Tissue distribution of ACE2 protein, the functional receptor for SARS coronavirus. A first step in understanding SARS pathogenesis. J. Pathol. 2004, 203, 631–637. [Google Scholar] [CrossRef]
- Li, W.; Moore, M.J.; Vasilieva, N.; Sui, J.; Wong, S.K.; Berne, M.A.; Somasundaran, M.; Sullivan, J.L.; Luzuriaga, K.; Greenough, T.C.; et al. Angiotensin-converting enzyme 2 is a functional receptor for the SARS coronavirus. Nature 2003, 426, 450–454. [Google Scholar] [CrossRef]
- Dwosh, H.A.; Hong, H.H.; Austgarden, D.; Herman, S.; Schabas, R. Identification and containment of an outbreak of SARS in a community hospital. CMAJ 2003, 168, 1415–1420. [Google Scholar]
- Ng, S.K. Possible role of an animal vector in the SARS outbreak at Amoy Gardens. Lancet 2003, 362, 570–572. [Google Scholar] [CrossRef]
- Vijgen, L.; Keyaerts, E.; Moes, E.; Thoelen, I.; Wollants, E.; Lemey, P.; Vandamme, A.M.; Van Ranst, M. Complete genomic sequence of human coronavirus OC43: molecular clock analysis suggests a relatively recent zoonotic coronavirus transmission event. J. Virol. 2005, 79, 1595–1604. [Google Scholar]
- Lau, S.K.; Woo, P.C.; Li, K.S.; Huang, Y.; Tsoi, H.W.; Wong, B.H.; Wong, S.S.; Leung, S.Y.; Chan, K.H.; Yuen, K.Y. Severe acute respiratory syndrome coronavirus-like virus in Chinese horseshoe bats. Proc. Natl. Acad. Sci. USA 2005, 102, 14040–14045. [Google Scholar]
- Li, W.; Shi, Z.; Yu, M.; Ren, W.; Smith, C.; Epstein, J.H.; Wang, H.; Crameri, G.; Hu, Z.; Zhang, H.; et al. Bats are natural reservoirs of SARS-like coronaviruses. Science 2005, 310, 676–679. [Google Scholar] [CrossRef]
- Poon, L.L.; Chu, D.K.; Chan, K.H.; Wong, O.K.; Ellis, T.M.; Leung, Y.H.; Lau, S.K.; Woo, P.C.; Suen, K.Y.; Yuen, K.Y.; et al. Identification of a novel coronavirus in bats. J. Virol. 2005, 79, 2001–2009. [Google Scholar]
- Woo, P.C.; Lau, S.K.; Li, K.S.; Poon, R.W.; Wong, B.H.; Tsoi, H.W.; Yip, B.C.; Huang, Y.; Chan, K.H.; Yuen, K.Y. Molecular diversity of coronaviruses in bats. Virology 2006, 351, 180–187. [Google Scholar] [CrossRef]
- Kan, B.; Wang, M.; Jing, H.; Xu, H.; Jiang, X.; Yan, M.; Liang, W.; Zheng, H.; Wan, K.; Liu, Q.; et al. Molecular evolution analysis and geographic investigation of severe acute respiratory syndrome coronavirus-like virus in palm civets at an animal market and on farms. J. Virol. 2005, 79, 11892–11900. [Google Scholar] [CrossRef]
- Cinatl, J., Jr.; Michaelis, M.; Morgenstern, B.; Doerr, H.W. High-dose hydrocortisone reduces expression of the pro-inflammatory chemokines CXCL8 and CXCL10 in SARS coronavirus-infected intestinal cells. Int. J. Mol. Med. 2005, 15, 323–327. [Google Scholar]
- Cinatl, J.; Morgenstern, B.; Bauer, G.; Chandra, P.; Rabenau, H.; Doerr, H.W. Glycyrrhizin, an active component of liquorice roots, and replication of SARS-associated coronavirus. Lancet 2003, 361, 2045–2046. [Google Scholar] [CrossRef]
- Morgenstern, B.; Michaelis, M.; Baer, P.C.; Doerr, H.W.; Cinatl, J., Jr. Ribavirin and interferon-beta synergistically inhibit SARS-associated coronavirus replication in animal and human cell lines. Biochem. Biophys. Res. Commun. 2005, 326, 905–908. [Google Scholar] [CrossRef]
- Mazzulli, T.; Farcas, G.A.; Poutanen, S.M.; Willey, B.M.; Low, D.E.; Butany, J.; Asa, S.L.; Kain, K.C. Severe acute respiratory syndrome-associated coronavirus in lung tissue. Emerg. Infect. Dis. 2004, 10, 20–24. [Google Scholar] [CrossRef]
- Yamamoto, N.; Yang, R.; Yoshinaka, Y.; Amari, S.; Nakano, T.; Cinatl, J.; Rabenau, H.; Doerr, H.W.; Hunsmann, G.; Otaka, A.; et al. HIV protease inhibitor nelfinavir inhibits replication of SARS-associated coronavirus. Biochem. Biophys. Res. Commun. 2004, 318, 719–725. [Google Scholar] [CrossRef]
- Cinatl, J., Jr.; Michaelis, M.; Hoever, G.; Preiser, W.; Doerr, H.W. Development of antiviral therapy for severe acute respiratory syndrome. Antiviral Res. 2005, 66, 81–97. [Google Scholar] [CrossRef]
- Haagmans, B.L.; Osterhaus, A.D. Coronaviruses and their therapy. Antiviral Res. 2006, 71, 397–403. [Google Scholar] [CrossRef]
- Bisht, H.; Roberts, A.; Vogel, L.; Bukreyev, A.; Collins, P.L.; Murphy, B.R.; Subbarao, K.; Moss, B. Severe acute respiratory syndrome coronavirus spike protein expressed by attenuated vaccinia virus protectively immunizes mice. Proc. Natl. Acad. Sci. USA. 2004, 101, 6641–6646. [Google Scholar]
- Kapadia, S.U.; Rose, J.K.; Lamirande, E.; Vogel, L.; Subbarao, K.; Roberts, A. Long-term protection from SARS coronavirus infection conferred by a single immunization with an attenuated VSV-based vaccine. Virology 2005, 340, 174–182. [Google Scholar] [CrossRef]
- Chen, Z.; Zhang, L.; Qin, C.; Ba, L.; Yi, C.E.; Zhang, F.; Wei, Q.; He, T.; Yu, W.; Yu, J.; et al. Recombinant modified vaccinia virus Ankara expressing the spike glycoprotein of severe acute respiratory syndrome coronavirus induces protective neutralizing antibodies primarily targeting the receptor binding region. J. Virol. 2005, 79, 2678–2688. [Google Scholar]
- Bukreyev, A.; Lamirande, E.W.; Buchholz, U.J.; Vogel, L.N.; Elkins, W.R.; St Claire, M.; Murphy, B.R.; Subbarao, K.; Collins, P.L. Mucosal immunisation of African green monkeys (Cercopithecus aethiops) with an attenuated parainfluenza virus expressing the SARS coronavirus spike protein for the prevention of SARS. Lancet 2004, 363, 2122–2127. [Google Scholar]
- Ishii, K.; Hasegawa, H.; Nagata, N.; Mizutani, T.; Morikawa, S.; Suzuki, T.; Taguchi, F.; Tashiro, M.; Takemori, T.; Miyamura, T.; et al. Induction of protective immunity against severe acute respiratory syndrome coronavirus (SARS-CoV) infection using highly attenuated recombinant vaccinia virus DIs. Virology 2006, 351, 368–380. [Google Scholar] [CrossRef]
- See, R.H.; Zakhartchouk, A.N.; Petric, M.; Lawrence, D.J.; Mok, C.P.; Hogan, R.J.; Rowe, T.; Zitzow, L.A.; Karunakaran, K.P.; Hitt, M.M.; et al. Comparative evaluation of two severe acute respiratory syndrome (SARS) vaccine candidates in mice challenged with SARS coronavirus. J. Gen. Virol. 2006, 87, 641–650. [Google Scholar] [CrossRef]
- Czub, M.; Weingartl, H.; Czub, S.; He, R.; Cao, J. Evaluation of modified vaccinia virus Ankara based recombinant SARS vaccine in ferrets. Vaccine 2005, 23, 2273–2279. [Google Scholar] [CrossRef]
- He, Y.; Li, J.; Heck, S.; Lustigman, S.; Jiang, S. Antigenic and immunogenic characterization of recombinant baculovirus-expressed severe acute respiratory syndrome coronavirus spike protein: implication for vaccine design. J. Virol. 2006, 80, 5757–5767. [Google Scholar] [CrossRef]
- Zhou, Z.; Post, P.; Chubet, R.; Holtz, K.; McPherson, C.; Petric, M.; Cox, M. A recombinant baculovirus-expressed S glycoprotein vaccine elicits high titers of SARS-associated coronavirus (SARS-CoV) neutralizing antibodies in mice. Vaccine 2006, 24, 3624–3631. [Google Scholar] [CrossRef]
- He, Y.; Zhou, Y.; Siddiqui, P.; Jiang, S. Inactivated SARS-CoV vaccine elicits high titers of spike protein-specific antibodies that block receptor binding and virus entry. Biochem. Biophys. Res. Commun. 2004, 325, 445–452. [Google Scholar] [CrossRef]
- Spruth, M.; Kistner, O.; Savidis-Dacho, H.; Hitter, E.; Crowe, B.; Gerencer, M.; Bruhl, P.; Grillberger, L.; Reiter, M.; Tauer, C.; et al. A double-inactivated whole virus candidate SARS coronavirus vaccine stimulates neutralising and protective antibody responses. Vaccine 2006, 24, 652–661. [Google Scholar]
- Tang, L.; Zhu, Q.; Qin, E.; Yu, M.; Ding, Z.; Shi, H.; Cheng, X.; Wang, C.; Chang, G.; Fang, F.; et al. Inactivated SARS-CoV vaccine prepared from whole virus induces a high level of neutralizing antibodies in BALB/c mice. DNA Cell Biol. 2004, 23, 391–394. [Google Scholar] [CrossRef]
- Xiong, S.; Wang, Y.F.; Zhang, M.Y.; Liu, X.J.; Zhang, C.H.; Liu, S.S.; Qian, C.W.; Li, J.X.; Lu, J.H.; Wan, Z.Y.; et al. Immunogenicity of SARS inactivated vaccine in BALB/c mice. Immunol. Lett. 2004, 95, 139–143. [Google Scholar] [CrossRef]
- Takasuka, N.; Fujii, H.; Takahashi, Y.; Kasai, M.; Morikawa, S.; Itamura, S.; Ishii, K.; Sakaguchi, M.; Ohnishi, K.; Ohshima, M.; et al. A subcutaneously injected UV-inactivated SARS coronavirus vaccine elicits systemic humoral immunity in mice. Int. Immunology 2004, 16, 1423–1430. [Google Scholar] [CrossRef]
- Stadler, K.; Roberts, A.; Becker, S.; Vogel, L.; Eickmann, M.; Kolesnikova, L.; Klenk, H.D.; Murphy, B.; Rappuoli, R.; Abrignani, S.; et al. SARS vaccine protective in mice. Emerg. Infect. Dis. 2005, 11, 1312–1314. [Google Scholar]
- Woo, P.C.; Lau, S.K.; Tsoi, H.W.; Chen, Z.W.; Wong, B.H.; Zhang, L.; Chan, J.K.; Wong, L.P.; He, W.; Ma, C.; et al. SARS coronavirus spike polypeptide DNA vaccine priming with recombinant spike polypeptide from Escherichia coli as booster induces high titer of neutralizing antibody against SARS coronavirus. Vaccine 2005, 23, 4959–4968. [Google Scholar] [CrossRef]
- Yang, Z.Y.; Kong, W.P.; Huang, Y.; Roberts, A.; Murphy, B.R.; Subbarao, K.; Nabel, G.J. A DNA vaccine induces SARS coronavirus neutralization and protective immunity in mice. Nature 2004, 428, 561–564. [Google Scholar]
- Zhao, P.; Ke, J.S.; Qin, Z.L.; Ren, H.; Zhao, L.J.; Yu, J.G.; Gao, J.; Zhu, S.Y.; Qi, Z.T. DNA vaccine of SARS-Cov S gene induces antibody response in mice. Acta Biochim. Biophys. Sin. 2004, 36, 37–41. [Google Scholar] [CrossRef]
- Wang, Z.; Yuan, Z.; Matsumoto, M.; Hengge, U.R.; Chang, Y.F. Immune responses with DNA vaccines encoded different gene fragments of severe acute respiratory syndrome coronavirus in BALB/c mice. Biochem. Biophys. Res. Commun. 2005, 327, 130–135. [Google Scholar] [CrossRef]
- Perlman, S.; Dandekar, A.A. Immunopathogenesis of coronavirus infections: implications for SARS. Nat. Rev. Immunol. 2005, 5, 917–927. [Google Scholar] [CrossRef]
- Bolles, M.; Deming, D.; Long, K.; Agnihothram, S.; Whitmore, A.; Ferris, M.; Funkhouser, W.; Gralinski, L.; Totura, A.; Heise, M.; et al. A double-inactivated severe acute respiratory syndrome coronavirus vaccine provides incomplete protection in mice and induces increased eosinophilic proinflammatory pulmonary response upon challenge. J. Virol. 2011, 85, 12201–12215. [Google Scholar] [CrossRef]
- Deming, D.; Sheahan, T.; Heise, M.; Yount, B.; Davis, N.; Sims, A.; Suthar, M.; Harkema, J.; Whitmore, A.; Pickles, R.; et al. Vaccine efficacy in senescent mice challenged with recombinant SARS-CoV bearing epidemic and zoonotic spike variants. PLoS Med. 2006, 3, e525. [Google Scholar] [CrossRef]
- Lokugamage, K.G.; Yoshikawa-Iwata, N.; Ito, N.; Watts, D.M.; Wyde, P.R.; Wang, N.; Newman, P.; Kent Tseng, C.T.; Peters, C.J.; Makino, S. Chimeric coronavirus-like particles carrying severe acute respiratory syndrome coronavirus (SCoV) S protein protect mice against challenge with SCoV. Vaccine 2008, 26, 797–808. [Google Scholar]
- Yasui, F.; Kai, C.; Kitabatake, M.; Inoue, S.; Yoneda, M.; Yokochi, S.; Kase, R.; Sekiguchi, S.; Morita, K.; Hishima, T.; et al. Prior immunization with severe acute respiratory syndrome (SARS)-associated coronavirus (SARS-CoV) nucleocapsid protein causes severe pneumonia in mice infected with SARS-CoV. J. Immunol. 2008, 181, 6337–6348. [Google Scholar]
- Tseng, C.T.; Sbrana, E.; Iwata-Yoshikawa, N.; Newman, P.C.; Garron, T.; Atmar, R.L.; Peters, C.J.; Couch, R.B. Immunization with SARS coronavirus vaccines leads to pulmonary immunopathology on challenge with the SARS virus. PLoS ONE. 2012, 7, e35421. [Google Scholar]
- Olsen, S.J.; Chang, H.L.; Cheung, T.Y.; Tang, A.F.; Fisk, T.L.; Ooi, S.P.; Kuo, H.W.; Jiang, D.D.; Chen, K.T.; Lando, J.; et al. Transmission of the severe acute respiratory syndrome on aircraft. N. Engl. J. Med. 2003, 349, 2416–2422. [Google Scholar] [CrossRef]
- Lee, S.H. The SARS epidemic in Hong Kong. J. Epidemiol. Community Health. 2003, 57, 652–654. [Google Scholar] [CrossRef]
- Ijaz, M.K.; Brunner, A.H.; Sattar, S.A.; Nair, R.C.; Johnson-Lussenburg, C.M. Survival characteristics of airborne human coronavirus 229E. J. Gen. Virol. 1985, 66 ( Pt 12). [Google Scholar]
- Rabenau, H.F.; Cinatl, J.; Morgenstern, B.; Bauer, G.; Preiser, W.; Doerr, H.W. Stability and inactivation of SARS coronavirus. Med. Microbiol. Immunol. 2005, 194, 1–6. [Google Scholar] [CrossRef]
- Duan, S.M.; Zhao, X.S.; Wen, R.F.; Huang, J.J.; Pi, G.H.; Zhang, S.X.; Han, J.; Bi, S.L.; Ruan, L.; Dong, X.P. Stability of SARS coronavirus in human specimens and environment and its sensitivity to heating and UV irradiation. Biomed. Environ. Sci. 2003, 16, 246–255. [Google Scholar]
- Sizun, J.; Yu, M.W.; Talbot, P.J. Survival of human coronaviruses 229E and OC43 in suspension and after drying on surfaces: a possible source of hospital-acquired infections. J. Hosp. Infect. 2000, 46, 55–60. [Google Scholar] [CrossRef]
- Lai, M.Y.; Cheng, P.K.; Lim, W.W. Survival of severe acute respiratory syndrome coronavirus. Clin. Infect. Dis. 2005, 41, e67–71. [Google Scholar] [CrossRef]
- Chen, Y.C.; Huang, L.M.; Chan, C.C.; Su, C.P.; Chang, S.C.; Chang, Y.Y.; Chen, M.L.; Hung, C.C.; Chen, W.J.; Lin, F.Y.; et al. SARS in hospital emergency room. Emerg. Infect. Dis. 2004, 10, 782–788. [Google Scholar]
- Dowell, S.F.; Simmerman, J.M.; Erdman, D.D.; Wu, J.S.; Chaovavanich, A.; Javadi, M.; Yang, J.Y.; Anderson, L.J.; Tong, S.; Ho, M.S. Severe acute respiratory syndrome coronavirus on hospital surfaces. Clin. Infect. Dis. 2004, 39, 652–657. [Google Scholar] [CrossRef]
- Casanova, L.; Rutala, W.A.; Weber, D.J.; Sobsey, M.D. Survival of surrogate coronaviruses in water. Water Res. 2009, 43, 1893–1898. [Google Scholar] [CrossRef]
- Lamarre, A.; Talbot, P.J. Effect of pH and temperature on the infectivity of human coronavirus 229E. Can. J. Microbiol. 1989, 35, 972–974. [Google Scholar]
- Sturman, L.S.; Ricard, C.S.; Holmes, K.V. Conformational change of the coronavirus peplomer glycoprotein at pH 8.0 and 37 degrees C correlates with virus aggregation and virus-induced cell fusion. J. Virol. 1990, 64, 3042–3050. [Google Scholar]
- Daniel, C.; Talbot, P.J. Physico-chemical properties of murine hepatitis virus, strain A 59. Brief report. Arch. Virol. 1987, 96, 241–248. [Google Scholar] [CrossRef]
- Pocock, D.H.; Garwes, D.J. The influence of pH on the growth and stability of transmissible gastroenteritis virus in vitro. Arch. Virol. 1975, 49, 239–247. [Google Scholar] [CrossRef]
- Pratelli, A. Canine coronavirus inactivation with physical and chemical agents. Vet. J. 2008, 177, 71–79. [Google Scholar] [CrossRef]
- WHO (World Health Organization). First data on stability and resistance of SARS coronavirus compiled by members of WHO laboratory network. Available online: http://www.who.int/csr/sars/survival_2003_05_04/en/index.html (Accessed on 16 February 2010).
- Lipsitch, M.; Cohen, T.; Cooper, B.; Robins, J.M.; Ma, S.; James, L.; Gopalakrishna, G.; Chew, S.K.; Tan, C.C.; Samore, M.H.; et al. Transmission dynamics and control of severe acute respiratory syndrome. Science 2003, 300, 1966–1970. [Google Scholar]
- Riley, S.; Fraser, C.; Donnelly, C.A.; Ghani, A.C.; Abu-Raddad, L.J.; Hedley, A.J.; Leung, G.M.; Ho, L.M.; Lam, T.H.; Thach, T.Q.; et al. Transmission dynamics of the etiological agent of SARS in Hong Kong: impact of public health interventions. Science 2003, 300, 1961–1966. [Google Scholar]
- Seto, W.H.; Tsang, D.; Yung, R.W.; Ching, T.Y.; Ng, T.K.; Ho, M.; Ho, L.M.; Peiris, J.S. Effectiveness of precautions against droplets and contact in prevention of nosocomial transmission of severe acute respiratory syndrome (SARS). Lancet 2003, 361, 1519–1520. [Google Scholar]
- WHO (World Health Organization). Global surveillance for Severe Acute Respiratory Syndrome (SARS). Wkly. Epidemiol. Rec. 2003, 78, 100–109.
- AFNOR, Chemical disinfectants and- Virucidal quantitative suspension test for chemical disinfectants and antiseptics used in human medicine - Test method and requirements (phase 2, step 1). NF EN 14476+A1. 2007.
- ASTM, Standard Test Method for Efficacy of Virucidal Agents Intended for Inanimate Environmental Surfaces. E1053-97 (last reapproval in 2002). 1997.
- ASTM, Standard Test Method for Efficacy of Antimicrobial Agents Against Viruses in Suspension. E1052-96 (last reapproval in 2002), 1996.
- ASTM, Standard Test Method for Neutralization of Virucidal Agents in Virucidal Efficacy Evaluations. E1482-04. 2004.
- ASTM (American Society for Testing and Materials), Standard Test Method for Determining the Virus-Eliminating Effectiveness of Liquid Hygienic Handwash and Handrub Agents Using the Fingerpads of Adult Volunteers. E1838-02. 2002.
- ASTM (American Society for Testing and Materials), Standard Test Method for Evaluation of Hygienic Handwash and Handrub Formulations for Virus-Eliminating Activity Using the Entire Hand. E2011-09. 2009.
- Sattar, S.A.; Springthorpe, V.S.; Karim, Y.; Loro, P. Chemical disinfection of non-porous inanimate surfaces experimentally contaminated with four human pathogenic viruses. Epidemiol. Infect. 1989, 102, 493–505. [Google Scholar] [CrossRef]
- Kariwa, H.; Fujii, N.; Takashima, I. Inactivation of SARS coronavirus by means of povidone-iodine, physical conditions and chemical reagents. Dermatology. 2006, 212 Suppl 1, 119–123. [Google Scholar]
- Geller, C.; Fontanay, S.; Finance, C.; Duval, R.E. A new Sephadex-based method for removing microbicidal and cytotoxic residues when testing antiseptics against viruses: Experiments with a human coronavirus as a model. J. Virol. Methods. 2009, 159, 217–226. [Google Scholar] [CrossRef]
- Geller, C.; Fontanay, S.; Mourer, M.; Dibama, H.M.; Regnouf-de-Vains, J.B.; Finance, C.; Duval, R.E. Antiseptic properties of two calix[4]arenes derivatives on the human coronavirus 229E. Antiviral Res. 2010, 88, 343–346. [Google Scholar] [CrossRef]
- Rabenau, H.F.; Kampf, G.; Cinatl, J.; Doerr, H.W. Efficacy of various disinfectants against SARS coronavirus. J. Hosp. Infect. 2005, 61, 107–111. [Google Scholar] [CrossRef]
- Hulkower, R.L.; Casanova, L.M.; Rutala, W.A.; Weber, D.J.; Sobsey, M.D. Inactivation of surrogate coronaviruses on hard surfaces by health care germicides. Am. J. Infect. Control. 2011, 39, 401–407. [Google Scholar] [CrossRef]
- Dellanno, C.; Vega, Q.; Boesenberg, D. The antiviral action of common household disinfectants and antiseptics against murine hepatitis virus, a potential surrogate for SARS coronavirus. Am. J. Infect. Control. 2009, 37, 649–652. [Google Scholar] [CrossRef]
© 2012 by the authors; licensee MDPI, Basel, Switzerland. This article is an open-access article distributed under the terms and conditions of the Creative Commons Attribution license (http://creativecommons.org/licenses/by/3.0/).