Initiation of HIV Reverse Transcription
Abstract
:1. Introduction
2. Selective packaging of the primer tRNA
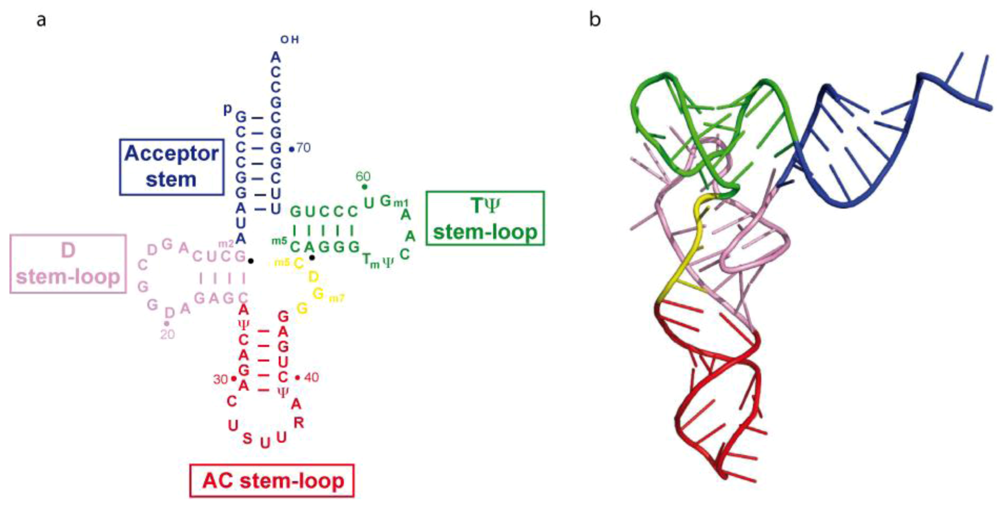
2.1. Structure of tRNA3Lys
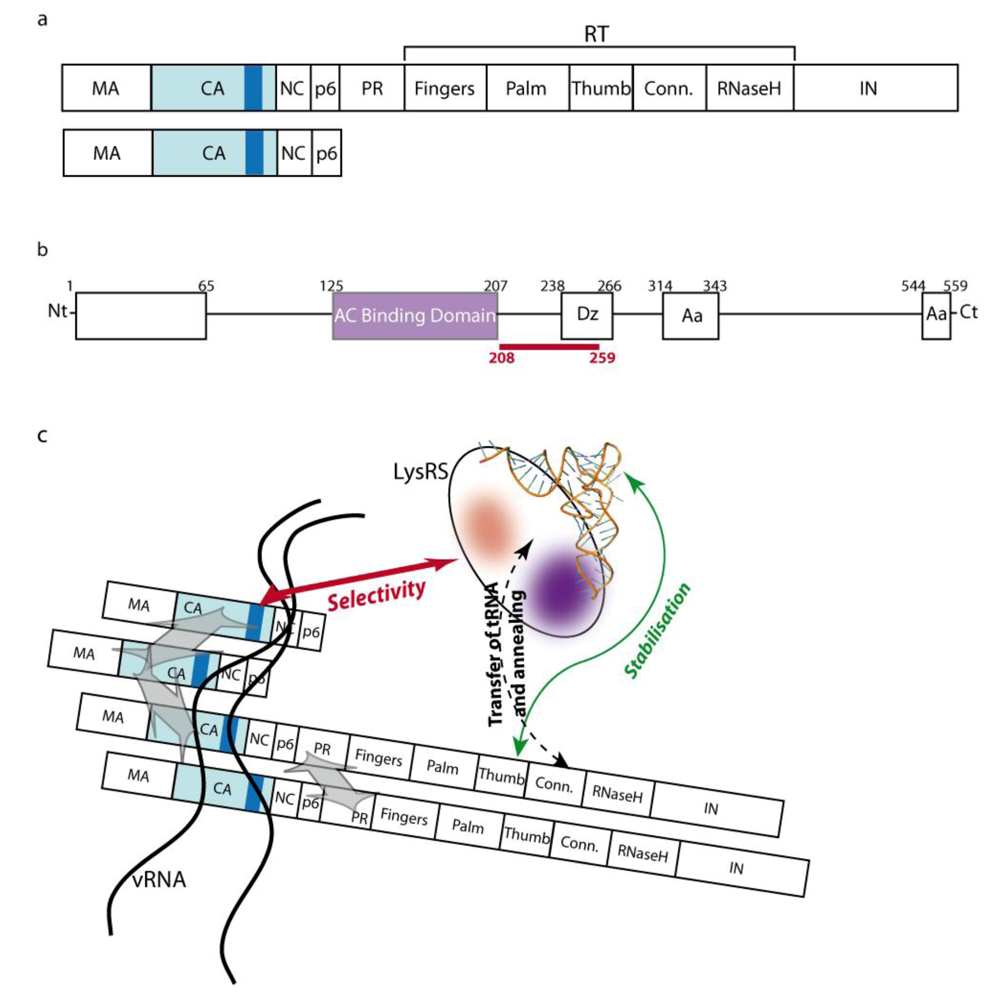
2.2. The role of Gag-Pol
2.3. The role of LysRS
2.4. The packaging complex
2.5. tRNA3Lys amino-acylation
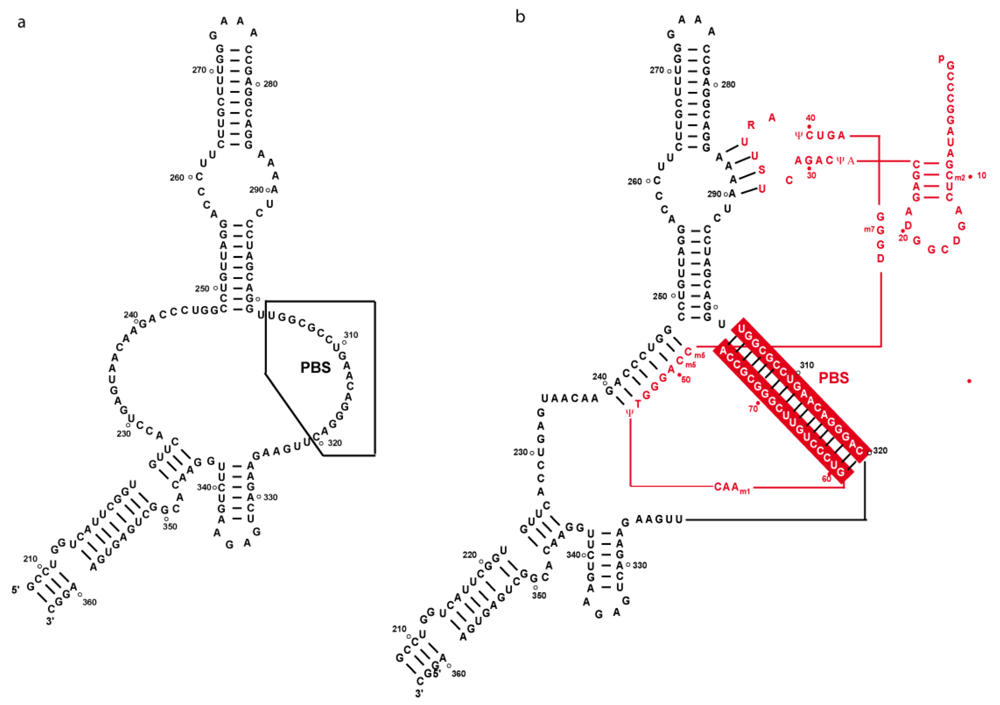
3. Secondary structure of the PBS domain
4. The Primer/Template Complex
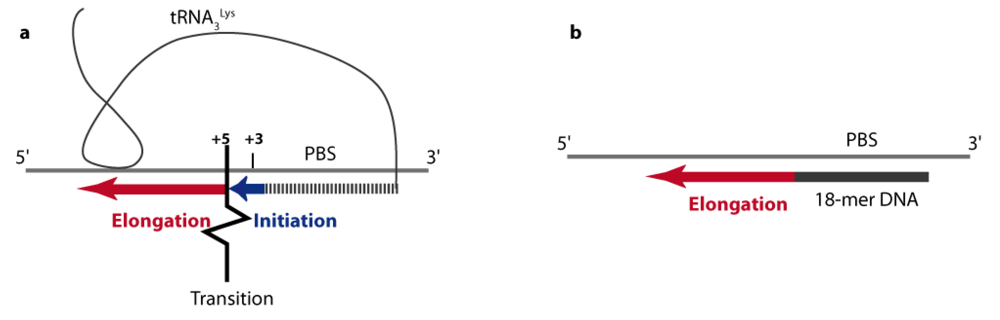
4.1. From initiation to elongation
4.2. Extended HIV-1 RNA/tRNA3Lys interactions and their implication in initiation of reverse transcription in vitro
4.2.1 Two and three-dimensional structures of a representative subtype A initiation complex
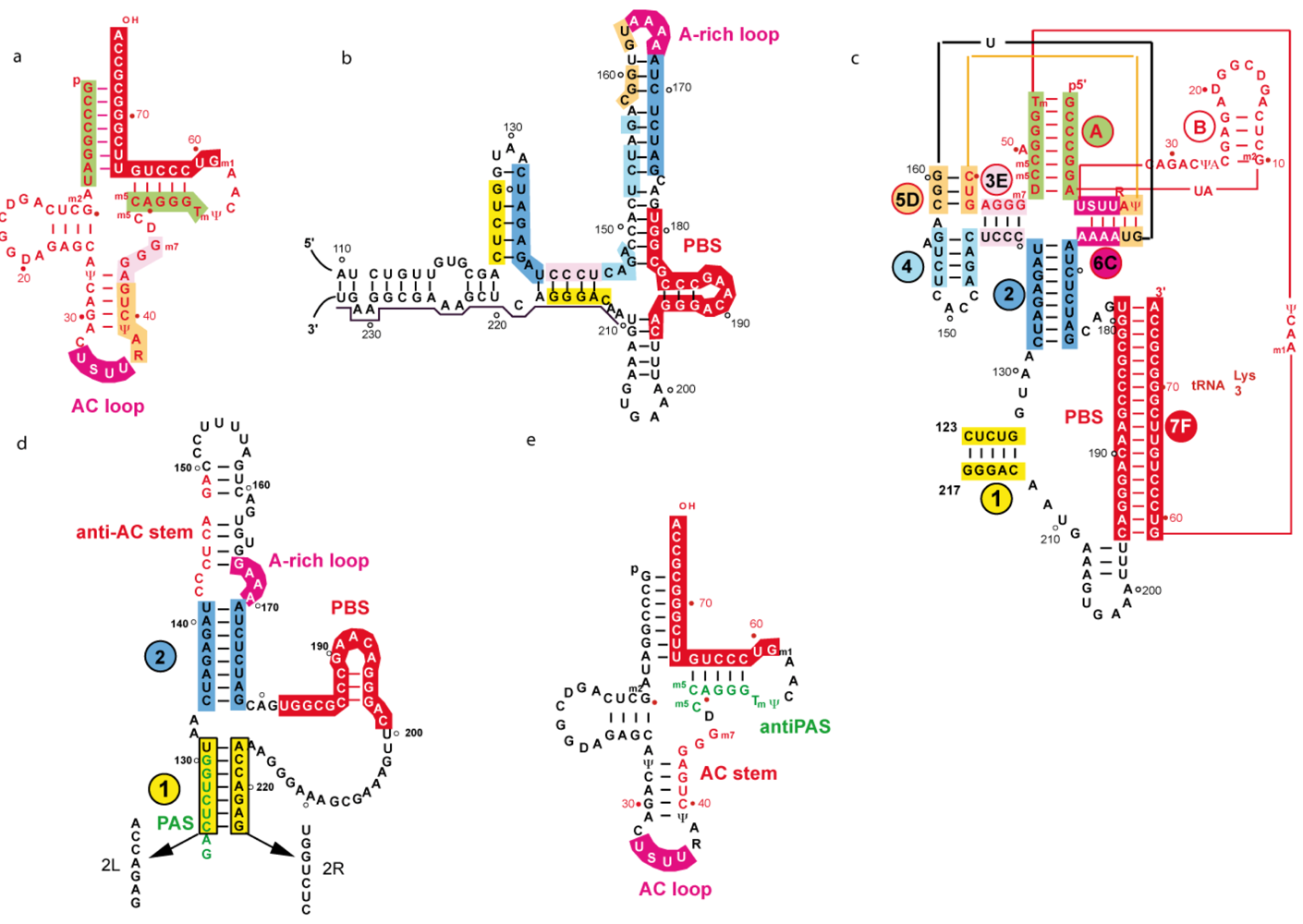
4.2.2. Secondary structure of tRNA/vRNA complexes representative of subtype B isolates
4.2.3. The PAS/anti-PAS interaction
4.2.4. The interaction involving the tRNA anticodon stem and the variable loop
4.3. Biological significance of extended HIV-1 RNA / tRNA3Lys interactions for virus replication
4.3.1. HIV-1 viruses carrying sequence deletions upstream and downstream of the PBS
4.3.2. HIV-1 variant viruses that use a non-self tRNA primer for reverse transcription
4.3.3. Reaching a consensus?
4.4. The case of HIV-2
7. Spatio-temporal regulation of reverse transcription
7.1. In producer cells
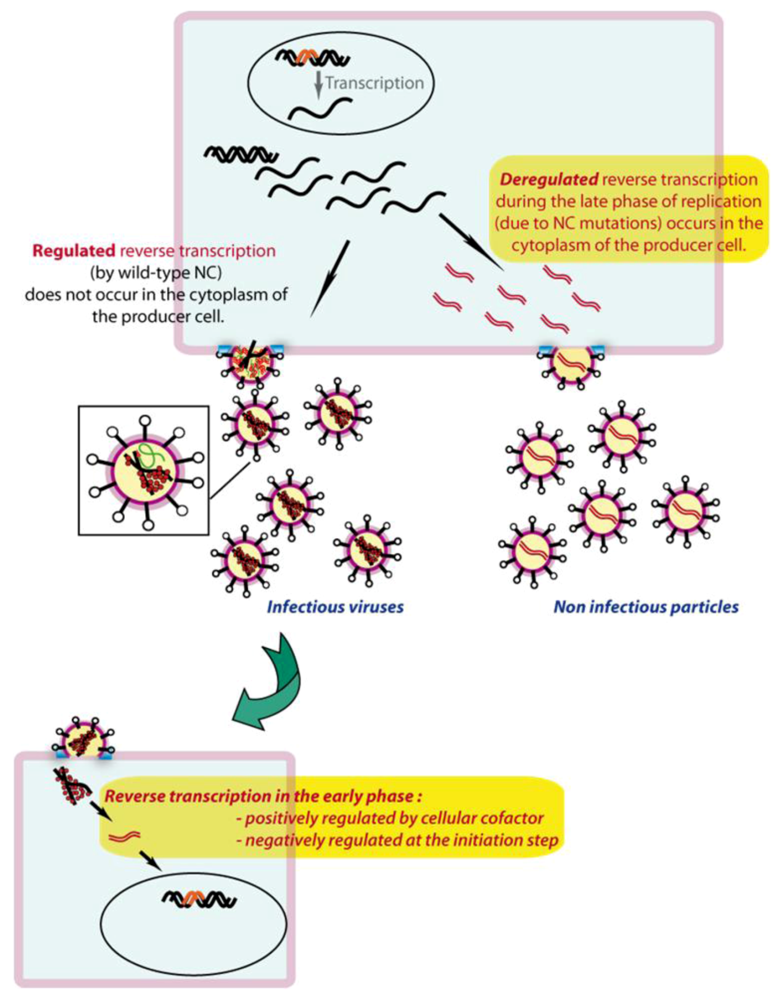
7.2. In target cells
8. Initiation of reverse transcription as a drug target
9. Conclusions
Acknowledgments
References
- Gilboa, E.; Mitra, S.W.; Goff, S.; Baltimore, D. A detailed model of reverse transcription and tests of crucial aspects. Cell 1979, 18, 93–100. [Google Scholar] [CrossRef] [PubMed]
- Mölling, K.; Bolognesi, D.P.; Bauer, H.; Büsen, W.; Plassmann, H.W.; Hausen, P. Association of viral reverse transcriptase with an enzyme degrading the RNA moiety of RNA-DNA hybrids. Nature New Biol. 1971, 234, 240–243. [Google Scholar] [CrossRef] [PubMed]
- Marquet, R.; Isel, C.; Ehresmann, C.; Ehresmann, B. tRNAs as primer of reverse transcriptase. Biochimie 1995, 77, 113–124. [Google Scholar] [CrossRef] [PubMed]
- Kleiman, L.; Jones, C.P.; Musier-Forsyth, K. Formation of the tRNALys packaging complex in HIV-1. FEBS Lett. 2010, 584, 359–365. [Google Scholar] [CrossRef] [PubMed]
- Kleiman, L.; Halwani, R.; Javanbakht, H. The selective packaging and annealing of primer tRNALys3 in HIV-1. Curr. HIV Res. 2004, 2, 163–175. [Google Scholar] [CrossRef]
- Jiang, M.; Mak, J.; Ladha, A.; Cohen, E.; Klein, M.; Rovinski, B.; Kleiman, L. Identification of tRNAs incorporated into wild-type and mutant human immunodeficiency virus type-1. J. Virol. 1993, 67, 3246–3253. [Google Scholar] [PubMed]
- Jiang, M.; Mak, J.; Wainberg, M.A.; Parniak, M.A.; Cohen, E.; Kleiman, L. Variable transfer RNA content in HIV-1(IIIB). Biochem. Biophys. Res. Commun. 1992, 185, 1005–1015. [Google Scholar] [CrossRef] [PubMed]
- Mak, J.; Jiang, M.; Wainberg, M.A.; Hammarskjöld, M.-L.; Rekosh, D.; Kleiman, L. Role of Pr160gag-pol in mediating the selective incorporation of tRNALys into human immunodeficiency virus type 1 particles. J. Virol. 1994, 68, 2065–2072. [Google Scholar] [PubMed]
- Huang, Y.; Mak, J.; Cao, Q.; Li, Z.; Wainberg, M.A.; Kleiman, L. Incorporation of excess wild-type and mutant tRNA3Lys into human immunodeficiency virus type 1. J. Virol. 1994, 68, 7676–7683. [Google Scholar] [PubMed]
- Bénas, P.; Bec, G.; Keith, G.; Marquet, R.; Ehresmann, C.; Ehresmann, B.; Dumas, P. The crystal structure of HIV reverse transcription primer tRNALys,3 shows a canonical anticodon loop. RNA 2000, 6, 1347–1355. [Google Scholar] [CrossRef]
- Agris, P.F.; Guenther, R.; Ingram, P.C.; Basti, M.M.; Stuart, J.W.; Sochacka, E.; Malkiewicz, A. Unconventional structure of tRNA(Lys)SUU anticodon explains tRNA's role in bacterial and mammalian ribosomal frameshifting and primer selection by HIV-1. RNA 1997, 3, 420–428. [Google Scholar] [PubMed]
- Durant, P.C.; Davis, D.R. Stabilization of the anticodon stem-loop of tRNALys,3 by an A+-C base-pair and by pseudouridine. J. Mol. Biol. 1999, 285, 115–131. [Google Scholar] [CrossRef] [PubMed]
- Puglisi, E.V.; Puglisi, J.D. Probing the conformation of human tRNA(3)(Lys) in solution by NMR. FEBS Lett. 2007, 581, 5307–5314. [Google Scholar] [CrossRef] [PubMed]
- Jiang, M.; Mak, J.; Huang, Y.; Kleiman, L. Reverse transcriptase is an important factor for the primer tRNA selection in HIV-1. Leukemia 1994, 8, S149–S151. [Google Scholar] [PubMed]
- Mak, J.; Khorchid, A.; Cao, Q.; Huang, Y.; Lowy, I.; Parniak, M.A.; Prasad, V.R.; Wainberg, M.A.; Kleiman, L. Effects of mutations in Pr160(gag-pol) upon tRNA3Lys and Pr160(gag-pol) incorporation into HIV-1. J. Mol. Biol. 1997, 265, 419–431. [Google Scholar] [CrossRef] [PubMed]
- Arts, E.J.; Miller, J.T.; Ehresmann, B.; Le Grice, S. F. J. Mutating a region of HIV-1 reverse transcriptase implicated in tRNALys-3 binding and the consequence for (-)-strand DNA synthesis. J. Biol. Chem. 1998, 273, 14523–14532. [Google Scholar] [CrossRef] [PubMed]
- Dufour, E.; El Dirani-Diab, R.; Boulme, F.; Fournier, M.; Nevinsky, G.; Tarrago-Litvak, L.; Litvak, S.; Andreola, M. L. p66/p51 and p51/p51 recombinant forms of reverse transcriptase from human immunodeficiency virus type 1--interactions with primer tRNA(Lys3), initiation of cDNA synthesis, and effect of inhibitors. Eur. J. Biochem. 1998, 251, 487–495. [Google Scholar] [PubMed]
- Mishima, Y.; Steitz, J.A. Site-specific crosslinking of 4-thiouridine-modified human tRNA3Lys to reverse transcriptase from human immunodeficiency virus type 1. EMBO J. 1995, 14, 2679–2687. [Google Scholar] [PubMed]
- Khorchid, A.; Javanbakht, H.; Wise, S.; Halwani, R.; Parniak, M.A.; Wainberg, M.A.; Kleiman, L. Sequences Within Pr160(gag-pol) Affecting the Selective Packaging of Primer tRNA(Lys3) into HIV-1. J. Mol. Biol. 2000, 299, 17–26. [Google Scholar] [CrossRef] [PubMed]
- Saadatmand, J.; Guo, F.; Cen, S.; Niu, M.; Kleiman, L. Interactions of reverse transcriptase sequences in Pol with Gag and LysRS in the HIV-1 tRNALys3 packaging/annealing complex. Virology 2008, 380, 109–117. [Google Scholar] [CrossRef] [PubMed]
- Halwani, R.; Cen, S.; Javanbakht, H.; Saadatmand, J.; Kim, S.; Shiba, K.; Kleiman, L. Cellular distribution of Lysyl-tRNA synthetase and its interaction with Gag during human immunodeficiency virus type 1 assembly. J. Virol. 2004, 78, 7553–7564. [Google Scholar] [CrossRef] [PubMed]
- Cen, S.; Khorchid, A.; Javanbakht, H.; Gabor, J.; Stello, T.; Shiba, K.; Musier-Forsyth, K.; Kleiman, L. Incorporation of lysyl-tRNA synthetase into human immunodeficiency virus type 1. J. Virol. 2001, 75, 5043–5048. [Google Scholar] [CrossRef] [PubMed]
- Cen, S.; Javanbakht, H.; Kim, S.; Shiba, K.; Craven, R.; Rein, A.; Ewalt, K.; Schimmel, P.; Musier-Forsyth, K.; Kleiman, L. Retrovirus-specific packaging of aminoacyl-tRNA synthetases with cognate primer tRNAs. J. Virol. 2002, 76, 13111–13115. [Google Scholar] [CrossRef] [PubMed]
- Gabor, J.; Cen, S.; Javanbakht, H.; Niu, M.; Kleiman, L. Effect of altering the tRNA(Lys)(3) concentration in human immunodeficiency virus type 1 upon its annealing to viral RNA, GagPol incorporation, and viral infectivity. J. Virol. 2002, 76, 9096–9102. [Google Scholar] [CrossRef] [PubMed]
- Guo, F.; Cen, S.; Niu, M.; Javanbakht, H.; Kleiman, L. Specific inhibition of the synthesis of human lysyl-tRNA synthetase results in decreases in tRNA(Lys) incorporation, tRNA(3)(Lys) annealing to viral RNA, and viral infectivity in human immunodeficiency virus type 1. J. Virol. 2003, 77, 9817–9822. [Google Scholar] [CrossRef] [PubMed]
- Cen, S.; Javanbakht, H.; Niu, M.; Kleiman, L. Ability of wild-type and mutant lysyl-tRNA synthetase to facilitate tRNA(Lys) incorporation into human immunodeficiency virus type 1. J. Virol. 2004, 78, 1595–1601. [Google Scholar] [CrossRef] [PubMed]
- Francin, M.; Kaminska, M.; Kerjan, P.; Mirande, M. The N-terminal domain of mammalian Lysyl-tRNA synthetase is a functional tRNA-binding domain. J. Biol. Chem. 2002, 277, 1762–1769. [Google Scholar] [CrossRef] [PubMed]
- Francin, M.; Mirande, M. Functional dissection of the eukaryotic-specific tRNA-interacting factor of lysyl-tRNA synthetase. J. Biol. Chem. 2003, 278, 1472–1479. [Google Scholar] [CrossRef] [PubMed]
- Javanbakht, H.; Cen, S.; Musier-Forsyth, K.; Kleiman, L. Correlation between tRNALys3 aminoacylation and its incorporation into HIV-1. J. Biol. Chem. 2002, 277, 17389–17396. [Google Scholar] [CrossRef] [PubMed]
- Kaminska, M.; Shalak, V.; Francin, M.; Mirande, M. Viral Hijacking of Mitochondrial Lysyl-tRNA Synthetase. J. Virol. 2006, 81, 68–73. [Google Scholar] [CrossRef] [PubMed]
- Javanbakht, H. The Interaction between HIV-1 Gag and Human Lysyl-tRNA Synthetase during Viral Assembly. J. Biol. Chem. 2003, 278, 27644–27651. [Google Scholar] [CrossRef] [PubMed]
- Kovaleski, B.J.; Kennedy, R.; Hong, M.K.; Datta, S.A.; Kleiman, L.; Rein, A.; Musier-Forsyth, K. In vitro characterization of the interaction between HIV-1 Gag and human lysyl-tRNA synthetase. J. Biol. Chem. 2006, 281, 19449–19456. [Google Scholar] [CrossRef] [PubMed]
- Kovaleski, B.J.; Kennedy, R.; Khorchid, A.; Kleiman, L.; Matsuo, H.; Musier-Forsyth, K. Critical role of helix 4 of HIV-1 capsid C-terminal domain in interactions with human lysyl-tRNA synthetase. J. Biol. Chem. 2007, 282, 32274–32279. [Google Scholar] [CrossRef] [PubMed]
- Ganser-Pornillos, B.K.; Yeager, M.; Sundquist, W.I. The structural biology of HIV assembly. Curr. Opin. Struct. Biol. 2008, 18, 203–217. [Google Scholar] [CrossRef] [PubMed]
- Pettit, S.C.; Gulnik, S.; Everitt, L.; Kaplan, A.H. The dimer interfaces of protease and extra-protease domains influence the activation of protease and the specificity of GagPol cleavage. J. Virol. 2003, 77, 366–374. [Google Scholar] [CrossRef] [PubMed]
- Berkowitz, R.D.; Luban, J.; Goff, S.P. Specific binding of human immunodeficiency virus type 1 gag polyprotein and nucleocapsid protein to viral RNAs detected by RNA mobility shift assays. J. Virol. 1993, 67, 7190–7200. [Google Scholar] [PubMed]
- Berkowitz, R.D.; Ohagen, A.; Hoglund, S.; Goff, S.P. Retroviral nucleocapsid domains mediate the specific recognition of genomic viral RNAs by chimeric Gag polyproteins during RNA packaging in vivo. J. Virol. 1995, 69, 6445–6456. [Google Scholar] [PubMed]
- Clever, J.; Sassetti, C.; Parslow, T.G. RNA secondary structure and binding sites for gag gene products in the 5' packaging signal of human immunodeficiency virus type 1. J. Virol. 1995, 69, 2101–2109. [Google Scholar] [PubMed]
- Damgaard, C.K.; Dyhr-Mikkelsen, H.; Kjems, J. Mapping the RNA binding sites for human immunodeficiency virus type-1 gag and NC proteins within the complete HIV-1 and -2 untranslated leader regions. Nucleic Acids Res. 1998, 26, 3667–3676. [Google Scholar] [CrossRef] [PubMed]
- Nikolaitchik, O.; Rhodes, T.D.; Ott, D.; Hu, W.S. Effects of mutations in the human immunodeficiency virus type 1 Gag gene on RNA packaging and recombination. J. Virol. 2006, 80, 4691–4697. [Google Scholar] [CrossRef] [PubMed]
- Cen, S.; Niu, M.; Kleiman, L. The connection domain in reverse transcriptase facilitates the in vivo annealing of tRNALys3 to HIV-1 genomic RNA. Retrovirology 2004, 1, 33. [Google Scholar] [CrossRef] [PubMed]
- Rigourd, M.; Bec, G.; Benas, P.; Le Grice, S.F.; Ehresmann, B.; Ehresmann, C.; Marquet, R. Effects of tRNA(3)(Lys) aminoacylation on the initiation of HIV-1 reverse transcription. Biochimie 2003, 85, 521–525. [Google Scholar] [CrossRef] [PubMed]
- Beerens, N.; Berkhout, B. The tRNA primer activation signal in the human immunodeficiency virus type 1 genome is important for initiation and processive elongation of reverse transcription. J. Virol. 2002, 76, 2329–2339. [Google Scholar] [CrossRef] [PubMed]
- Goldschmidt, V.; Rigourd, M.; Ehresmann, C.; Le Grice, S.F.; Ehresmann, B.; Marquet, R. Direct and indirect contributions of RNA secondary structure elements to the initiation of HIV-1 reverse transcription. J. Biol. Chem. 2002, 277, 43233–43242. [Google Scholar] [CrossRef] [PubMed]
- Paillart, J.C.; Dettenhofer, M.; Yu, X.F.; Ehresmann, C.; Ehresmann, B.; Marquet, R. First snapshots of the HIV-1 RNA structure in infected cells and in virions. J. Biol. Chem. 2004, 279, 48397–48403. [Google Scholar] [CrossRef] [PubMed]
- Watts, J.M.; Dang, K.K.; Gorelick, R.J.; Leonard, C.W.; Bess, J.W.; Swanstrom, R.; Burch, C.L.; Weeks, K. Architecture and secondary structure of an entire HIV-1 RNA genome. Nature 2009, 460, 711–716. [Google Scholar] [CrossRef] [PubMed]
- Wilkinson, K.; Gorelick, R.J.; Vasa, S.; Guex, N.; Rein, A.; Mathews, D.; Giddings, M.; Weeks, K. High-throughput SHAPE analysis reveals structures in HIV-1 genomic RNA strongly conserved across distinct biological states. PLoS Biol. 2008, 6, e96. [Google Scholar] [CrossRef] [PubMed]
- Goldschmidt, V.; Paillart, J.C.; Rigourd, M.; Ehresmann, B.; Aubertin, A.M.; Ehresmann, C.; Marquet, R. Structural variability of the initiation complex of HIV-1 reverse transcription. J. Biol. Chem. 2004, 279, 35923–35931. [Google Scholar] [CrossRef] [PubMed]
- Baudin, F.; Marquet, R.; Isel, C.; Darlix, J.L.; Ehresmann, B.; Ehresmann, C. Functional sites in the 5' region of human immunodeficiency virus type-1 RNA form defined structural domains. J. Mol. Biol. 1993, 229, 382–397. [Google Scholar] [CrossRef] [PubMed]
- Beerens, N.; Groot, F.; Berkhout, B. Initiation of HIV-1 reverse transcription is regulated by a primer activation signal. J. Biol. Chem. 2001, 276, 31247–31256. [Google Scholar] [CrossRef] [PubMed]
- Berkhout, B. Structure and function of the human immunodeficiency virus leader RNA. Prog. Nucleic Acids Res. Mol. Biol. 1996, 54, 1–34. [Google Scholar]
- Rizvi, T.A.; Panganiban, A.T. Simian immunodeficiency virus RNA is efficiently encapsidated by human immunodeficiency virus type 1 particles. J. Virol. 1993, 67, 2681–2688. [Google Scholar] [PubMed]
- Isel, C.; Lanchy, J.M.; Le Grice, S.F.J.; Ehresmann, C.; Ehresmann, B.; Marquet, R. Specific initiation and switch to elongation of human immunodeficiency virus type 1 reverse transcription require the post-transcriptional modifications of primer tRNA3Lys. EMBO J. 1996, 15, 917–924. [Google Scholar] [PubMed]
- Lanchy, J.M.; Ehresmann, C.; Le Grice, S.F.J.; Ehresmann, B.; Marquet, R. Binding and kinetic properties of HIV-1 reverse transcriptase markedly differ during initiation and elongation of reverse transcription. EMBO J. 1996, 15, 7178–7187. [Google Scholar] [PubMed]
- Lanchy, J.M.; Keith, G.; Le Grice, S.F.J.; Ehresmann, B.; Ehresmann, C.; Marquet, R. Contacts between reverse transcriptase and the primer strand govern the transition from initiation to elongation of HIV-1 reverse transcription. J. Biol. Chem. 1998, 273, 24425–24432. [Google Scholar] [CrossRef] [PubMed]
- Arts, E.J.; Stetor, S.R.; Li, X.G.; Rausch, J.W.; Howard, K.J.; Ehresmann, B.; North, T.W.; Wohrl, B.M.; Goody, R.S.; Wainberg, M.A.; Le Grice, S.F.J. Initiation of (-) strand DNA synthesis from tRNA3Lys on lentiviral RNAs: Implications of specific HIV-1 RNA-tRNA3Lys interactions inhibiting primer utilization by retroviral reverse transcriptases. Proc. Natl. Acad. Sci. U.S.A. 1996, 93, 10063–10068. [Google Scholar] [CrossRef] [PubMed]
- Tisne, C.; Rigourd, M.; Marquet, R.; Ehresmann, C.; Dardel, F. NMR and biochemical characterization of recombinant human tRNALys3 expressed in Escherichia coli: identification of posttranscriptional nucleotide modifications required for efficient initiation of HIV-1 reverse transcription. RNA 2000, 6, 1403–1412. [Google Scholar] [CrossRef]
- Boulme, F.; Freund, F.; Litvak, S. Initiation of in vitro reverse transcription from tRNA(Lys3) on HIV-1 or HIV-2 RNAs by both type 1 and 2 reverse transcriptases. FEBS Lett. 1998, 430, 165–170. [Google Scholar] [CrossRef] [PubMed]
- Isel, C.; Ehresmann, C.; Keith, G.; Ehresmann, B.; Marquet, R. Two step synthesis of (-) strong-stop DNA by avian and murine reverse transcriptases in vitro. Nucleic Acids Res. 1997, 25, 545–552. [Google Scholar] [CrossRef] [PubMed]
- Das, A.T.; Klaver, B.; Berkhout, B. Reduced replication of human immunodeficiency virus type 1 mutants that use reverse transcription primers other than the natural tRNA3Lys. J. Virol. 1995, 69, 3090–3097. [Google Scholar] [PubMed]
- Li, X.G.; Mak, J.; Arts, E. J.; Gu, Z.X.; Kleiman, L.; Wainberg, M.A.; Parniak, M.A. Effects of alterations of primer-binding site sequences on human immunodeficiency virus type 1 replication. J. Virol. 1994, 68, 6198–6206. [Google Scholar] [PubMed]
- Wakefield, J.K.; Wolf, A.G.; Morrow, C.D. Human immunodeficiency virus type 1 can use different tRNAs as primers for reverse transcription but selectively maintains a primer binding site complementary to tRNA3Lys. J Virol. 1995, 69, 6021–6029. [Google Scholar] [PubMed]
- Whitcomb, J.M.; Ortiz-Conde, B.A.; Hughes, S.H. Replication of avian leukosis viruses with mutations at the primer binding site: use of alternative tRNAs as primers. J. Virol. 1995, 69, 6228–6238. [Google Scholar] [PubMed]
- Colicelli, J.; Goff, S.P. Isolation of a recombinant murine leukemia virus utilizing a new primer tRNA. J. Virol. 1986, 57, 37–45. [Google Scholar] [PubMed]
- Lund, A.H.; Duch, M.; Lovmand, J.; Jørgensen, P.; Pedersen, F.S. Mutated primer binding sites interacting with different tRNAs allow efficient murine leukemia virus replication. J. Virol. 1993, 67, 7125–7130. [Google Scholar] [PubMed]
- Isel, C.; Ehresmann, C.; Keith, G.; Ehresmann, B.; Marquet, R. Initiation of reverse transcription of HIV-1: secondary structure of the HIV-1 RNA/tRNA3Lys (template/primer) complex. J. Mol. Biol. 1995, 247, 236–250. [Google Scholar] [CrossRef] [PubMed]
- Isel, C.; Marquet, R.; Keith, G.; Ehresmann, C.; Ehresmann, B. Modified nucleotides of transfer-RNA3Lys modulate primer / template loop-loop interaction in the initiation complex of HIV-1 reverse transcription. J. Biol. Chem. 1993, 268, 25269–25272. [Google Scholar] [PubMed]
- Brulé, F.; Marquet, R.; Rong, L.; Wainberg, M.A.; Roques, B.P.; Le Grice, S.F.J.; Ehresmann, B.; Ehresmann, C. Structural and functional properties of the HIV-1 RNA-tRNA3Lys primer annealed by the nucleocapsid protein: Comparison with the heat annealed complex. RNA 2002, 8, 8–15. [Google Scholar] [CrossRef] [PubMed]
- Skripkin, E.; Isel, C.; Marquet, R.; Ehresmann, B.; Ehresmann, C. Psoralen crosslinking between human immunodeficiency virys type 1 RNA and primer tRNA3Lys. Nucleic Acids Res. 1996, 24, 509–514. [Google Scholar] [CrossRef] [PubMed]
- Isel, C.; Keith, G.; Ehresmann, B.; Ehresmann, C.; Marquet, R. Mutational analysis of the tRNA3Lys/HIV-1 RNA (primer/template) complex. Nucleic Acids Res. 1998, 26, 1198–1204. [Google Scholar] [CrossRef] [PubMed]
- Puglisi, E.V.; Puglisi, J.D. HIV-1 A-rich RNA loop mimics the tRNA anticodon structure. Nat. Struct. Biol. 1998, 5, 1033–1036. [Google Scholar] [CrossRef] [PubMed]
- Bilbille, Y.; Vendeix, F.A.; Guenther, R.; Malkiewicz, A.; Ariza, X.; Vilarrasa, J.; Agris, P. The structure of the human tRNALys3 anticodon bound to the HIV genome is stabilized by modified nucleosides and adjacent mismatch base pairs. Nucleic Acids Res. 2009, 37, 3342–3353. [Google Scholar] [CrossRef] [PubMed]
- Isel, C.; Westhof, E.; Massire, C.; Le Grice, S.F.J.; Ehresmann, C.; Ehresmann, B.; Marquet, R. Structural basis for the specificity of the initiation of HIV-1 reverse transcription. EMBO J. 1999, 18, 1038–1048. [Google Scholar] [CrossRef] [PubMed]
- Iwatani, Y.; Rosen, A.E.; Guo, J.; Musier-Forsyth, K.; Levin, J.G. Efficient initiation of HIV-1 reverse transcription in vitro: Requirement for RNA sequences downstream of the primer binding site abrogated by nucleocapsid protein-dependent primer-template interactions. J. Biol. Chem. 2003, 278, 14185–14195. [Google Scholar] [CrossRef] [PubMed]
- Liang, C.; Li, X.; Rong, L.; Inouye, P.; Quan, Y.; Kleiman, L.; Wainberg, M.A. The importance of the A-rich loop in human immunodeficiency virus type 1 reverse transcription and infectivity. J. Virol. 1997, 71, 5750–5757. [Google Scholar] [PubMed]
- Liang, C.; Rong, L.; Gotte, M.; Li, X.; Quan, Y.; Kleiman, L.; Wainberg, M.A. Mechanistic studies of early pausing events during initiation of HIV-1 reverse transcription. J. Biol. Chem. 1998, 273, 21309–21315. [Google Scholar] [CrossRef] [PubMed]
- Li, X.; Quan, Y.; Arts, E.J.; Li, Z.; Preston, B.D.; de Rocquigny, H.; Roques, B.P.; Darlix, J.L.; Kleiman, L.; Parniak, M.A.; Wainberg, M.A. Human immunodeficiency virus Type 1 nucleocapsid protein (NCp7) directs specific initiation of minus-strand DNA synthesis primed by human tRNA(Lys3) in vitro: studies of viral RNA molecules mutated in regions that flank the primer binding site. J. Virol. 1996, 70, 4996–5004. [Google Scholar] [PubMed]
- Beerens, N.; Berkhout, B. Switching the in vitro tRNA usage of HIV-1 by simultaneous adaptation of the PBS and PAS. RNA 2002, 8, 357–369. [Google Scholar] [CrossRef]
- Ooms, M.; Cupac, D.; Abbink, T.E.; Huthoff, H.; Berkhout, B. The availability of the primer activation signal (PAS) affects the efficiency of HIV-1 reverse transcription initiation. Nucleic Acids Res. 2007, 35, 1649–1659. [Google Scholar] [CrossRef] [PubMed]
- Aiyar, A.; Cobrinik, D.; Ge, Z.; Kung, H.J.; Leis, J. Interaction between retroviral U5 RNA and the TψC loop of the tRNATrp primer is required for efficient initiation of reverse transcription. J. Virol. 1992, 66, 2464–2472. [Google Scholar] [PubMed]
- Goldschmidt, V.; Ehresmann, C.; Ehresmann, B.; Marquet, R. Does the HIV-1 primer activation signal interact with tRNA3Lys during the initiation of reverse transcription? Nucleic Acids Res. 2003, 31, 850–859. [Google Scholar] [CrossRef] [PubMed]
- Thrall, S.H.; Krebs, R.; Wohrl, B.M.; Cellai, L.; Goody, R.S.; Restle, T. Pre-steady-state kinetic characterization of RNA-primed initiation of transcription by HIV-1 reverse transcriptase and analysis of the transition to a processive DNA-primed polymerization mode. Biochemistry 1998, 37, 13349–13358. [Google Scholar] [CrossRef] [PubMed]
- Vicenzi, E.; Dimitrov, D.S.; Engelman, A.; Migone, T.S.; Purcell, D.F.; Leonard, J.; Englund, G.; Martin, M.A. An integration-defective U5 deletion mutant of human immunodeficiency virus type 1 reverts by eliminating additional long terminal repeat sequences. J. Virol. 1994, 68, 7879–7890. [Google Scholar] [PubMed]
- Li, Y.; Zhang, Z.; Wakefield, J.K.; Kang, S.M.; Morrow, C.D. Nucleotide substitutions within U5 are critical for efficient reverse transcription of human immunodeficiency virus type 1 with a primer binding site complementary to tRNAHis. J. Virol. 1997, 71, 6315–6322. [Google Scholar] [PubMed]
- Wakefield, J.K.; Kang, S.-M.; Morrow, C.D. Construction of a type 1 human immunodeficiency virus that maintains a primer binding site complementary to tRNAHis. J. Virol. 1996, 70, 966–975. [Google Scholar] [PubMed]
- Zhang, Z.; Kang, S.M.; LeBlanc, A.; Hajduk, S.L.; Morrow, C.D. Nucleotide sequences within the U5 region of the viral RNA genome are the major determinants for an human immunodeficiency virus type 1 to maintain a primer binding site complementary to tRNAHis. Virology 1996, 226, 306–317. [Google Scholar] [CrossRef] [PubMed]
- Kang, S.M.; Morrow, C.D. Genetic analysis of a unique human immunodeficiency virus type 1 (HIV- 1) with a primer binding site complementary to tRNAMet supports a role for U5-PBS stem-loop RNA structures in initiation of HIV-1 reverse transcription. J. Virol. 1999, 73, 1818–1827. [Google Scholar] [PubMed]
- Kang, S.M.; Zhang, Z.J.; Morrow, C.D. Identification of a sequence within U5 required for human immunodeficiency virus type 1 to stably maintain a primer binding site complementary to tRNAMet. J. Virol. 1997, 71, 207–217. [Google Scholar] [PubMed]
- Kang, S.M.; Zhang, Z.; Morrow, C.D. Identification of a human immunodeficiency virus type 1 that stably uses tRNALys1,2 rather than tRNALys,3 for initiation of reverse transcription. Virology 1999, 257, 95–105. [Google Scholar] [CrossRef] [PubMed]
- Brown, H.E.; Chen, H.; Engelman, A. Structure-based mutagenesis of the human immunodeficiency virus type 1 DNA attachment site: effects on integration and cDNA synthesis. J. Virol. 1999, 73, 9011–9020. [Google Scholar] [PubMed]
- Esposito, D.; Craigie, R. Sequence specificity of viral end DNA binding by HIV-1 integrase reveals critical regions for protein-DNA interaction. EMBO J. 1998, 17, 5832–5843. [Google Scholar] [CrossRef] [PubMed]
- Huang, Y.; Shalom, A.; Li, Z.; Wang, J.; Mak, J.; Wainberg, M.A.; Kleiman, L. Effects of modifying the tRNA3Lys anticodon on the initiation of human immunodeficiency virus type 1 reverse transcription. J. Virol. 1996, 70, 4700–4706. [Google Scholar] [PubMed]
- Zhang, Z.J.; Kang, S.M.; Li, Y.; Morrow, C.D. Genetic analysis of the U5-PBS of a novel HIV-1 reveals multiple interactions between the tRNA and RNA genome required for initiation of reverse transcription. RNA 1998, 4, 394–406. [Google Scholar] [CrossRef]
- Wei, M.; Yang, Y.; Niu, M.; Desfosse, L.; Kennedy, R.; Musier-Forsyth, K.; Kleiman, L. Inability of human immunodeficiency virus type 1 produced in murine cells to selectively incorporate primer formula. J. Virol. 2008, 82, 12049–12059. [Google Scholar] [CrossRef] [PubMed]
- Rigourd, M.; Goldschmidt, V.; Brule, F.; Morrow, C.D.; Ehresmann, B.; Ehresmann, C.; Marquet, R. Structure-function relationships of the initiation complex of HIV-1 reverse transcription: the case of mutant viruses using tRNA(His) as primer. Nucleic Acids Res. 2003, 31, 5764–5775. [Google Scholar] [CrossRef] [PubMed]
- Abbink, T.E.; Beerens, N.; Berkhout, B. Forced selection of a human immunodeficiency virus type 1 variant that uses a non-self tRNA primer for reverse transcription: involvement of viral RNA sequences and the reverse transcriptase enzyme. J. Virol. 2004, 78, 10706–10714. [Google Scholar] [CrossRef] [PubMed]
- Fossé, P.; Mougel, M.; Keith, G.; Westhof, E.; Ehresmann, B.; Ehresmann, C. Modified nucleotides of tRNAPro restrict interactions in the binary primer/template complex of M-MLV. J. Mol. Biol. 1998, 275, 731–746. [Google Scholar] [CrossRef] [PubMed]
- Morris, S.; Johnson, M.; Stavnezer, E.; Leis, J. Replication of Avian Sarcoma Virus In Vivo Requires an Interaction between the Viral RNA and the TψC Loop of the tRNATrp Primer. J. Virol. 2002, 76, 7571–7577. [Google Scholar] [CrossRef] [PubMed]
- Das, A.T.; Vink, M.; Berkhout, B. Alternative tRNA priming of human immunodeficiency virus type 1 reverse transcription explains sequence variation in the primer-binding site that has been attributed to APOBEC3G activity. J. Virol. 2005, 79, 3179–3181. [Google Scholar] [CrossRef] [PubMed]
- Dupuy, L.C.; Kelly, N.J.; Elgavish, T.E.; Harvey, S.C.; Morrow, C.D. Probing the importance of tRNA anticodon: human immunodeficiency virus type 1 (HIV-1) RNA genome complementarity with an HIV-1 that selects tRNA(Glu) for replication. J. Virol. 2003, 77, 8756–8764. [Google Scholar] [CrossRef] [PubMed]
- Berkhout, B.; Schoneveld, I. Secondary structure of the HIV-2 leader RNA comprising the tRNA-primer binding site. Nucleic Acids Res. 1993, 21, 1171–1178. [Google Scholar] [CrossRef] [PubMed]
- Freund, F.; Boulme, F.; Litvak, S.; Tarrago-Litvak, L. Initiation of HIV-2 reverse transcription: a secondary structure model of the RNA-tRNALys3 duplex. Nucleic Acids Res. 2001, 29, 2757–2765. [Google Scholar] [CrossRef] [PubMed]
- Chan, B.; Weidemaier, K.; Yip, W.T.; Barbara, P.F.; Musier-Forsyth, K. Intra-tRNA distance measurements for nucleocapsid proteindependent tRNA unwinding during priming of HIV reverse transcription. Proc. Natl. Acad. Sci. USA 1999, 96, 459–464. [Google Scholar] [CrossRef]
- de Rocquigny, H.; Gabus, C.; Vincent, A.; Fournié-Zaluski, M.C.; Roques, B.; Darlix, J.L. Viral RNA annealing activities of HIV-1 nucleocapsid protein require only peptide domains outside the zinc fingers. Proc. Natl. Acad. Sci. USA 1992, 89, 6472–6476. [Google Scholar] [CrossRef]
- Lapadat-Tapolsky, M.; de Rocquigny, H.; Van Gent, D.; Roques, B.; Plasterk, R.; Darlix, J.L. Interactions between HIV-1 nucleocapsid protein and viral DNA may have important functions in the viral life cycle [published erratum appears in Nucleic Acids Res. 1993 Apr 25;21(8):2024]. Nucleic Acids Res. 1993, 21, 831–839. [Google Scholar] [CrossRef] [PubMed]
- Prats, A.C.; Sarih, L.; Gabus, C.; Litvak, S.; Keith, G.; Darlix, J.L. Small finger protein of avian and murine retroviruses has nucleic acid annealing activity and positions the replication primer tRNA onto genomic RNA. EMBO J. 1988, 7, 1777–1783. [Google Scholar] [PubMed]
- Levin, J.G.; Guo, J.; Rouzina, I.; Musier-Forsyth, K. Nucleic acid chaperone activity of HIV-1 nucleocapsid protein: critical role in reverse transcription and molecular mechanism. Prog. Nucleic Acid Res. Mol. Biol. 2005, 80, 217–286. [Google Scholar] [PubMed]
- Cen, S.; Huang, Y.; Khorchid, A.; Darlix, J.L.; Wainberg, M.A.; Kleiman, L. The role of Pr55(gag) in the annealing of tRNA3Lys to human immunodeficiency virus type 1 genomic RNA. J. Virol. 1999, 73, 4485–4488. [Google Scholar] [PubMed]
- Feng, Y.X.; Campbell, S.; Harvin, D.; Ehresmann, B.; Ehresmann, C.; Rein, A. The human immunodeficiency virus type 1 Gag polyprotein has nucleic acid chaperone activity: possible role in dimerization of genomic RNA and placement of tRNA on the primer binding site. J. Virol. 1999, 73, 4251–4256. [Google Scholar] [PubMed]
- Guo, F.; Saadatmand, J.; Niu, M.; Kleiman, L. Roles of Gag and NCp7 in facilitating tRNA(Lys)(3) Annealing to viral RNA in human immunodeficiency virus type 1. J. Virol. 2009, 83, 8099–8107. [Google Scholar] [CrossRef] [PubMed]
- Huang, Y.; Khorchid, A.; Wang, J.; Parniak, M.A.; Darlix, J.L.; Wainberg, M.A.; Kleiman, L. Effect of mutations in the nucleocapsid protein (NCp7) upon Pr160(gag- pol) and tRNA(Lys) incorporation into human immunodeficiency virus type 1. J. Virol. 1997, 71, 4378–4384. [Google Scholar] [PubMed]
- Saadatmand, J.; Niu, M.; Kleiman, L.; Guo, F. The contribution of the primer activation signal to differences between Gag- and NCp7-facilitated tRNA(Lys3) annealing in HIV-1. Virology 2009, 391, 334–341. [Google Scholar] [CrossRef] [PubMed]
- Hargittai, M.R.; Mangla, A.T.; Gorelick, R.J.; Musier-Forsyth, K. HIV-1 nucleocapsid protein zinc finger structures induce tRNA(Lys,3) structural changes but are not critical for primer/template annealing. J. Mol. Biol. 2001, 312, 985–997. [Google Scholar] [CrossRef] [PubMed]
- Tisné, C.; Roques, B.P.; Dardel, F. The annealing mechanism of HIV-1 reverse transcription primer onto the viral genome. J. Biol. Chem. 2004, 279, 3588–3595. [Google Scholar] [CrossRef] [PubMed]
- Williams, M.C.; Gorelick, R.J.; Musier-Forsyth, K. Specific zinc-finger architecture required for HIV-1 nucleocapsid protein's nucleic acid chaperone function. Proc. Natl. Acad. Sci. USA 2002, 99, 8614–8619. [Google Scholar] [CrossRef]
- Lapadat-Tapolsky, M.; Pernelle, C.; Borie, C.; Darlix, J.L. Analysis of the nucleic acid annealing activities of nucleocapsid protein from HIV-1. Nucleic Acids Res. 1995, 23, 2434–2441. [Google Scholar] [CrossRef] [PubMed]
- Hargittai, M.R.; Gorelick, R.J.; Rouzina, I.; Musier-Forsyth, K. Mechanistic insights into the kinetics of HIV-1 nucleocapsid protein-facilitated tRNA annealing to the primer binding site. J. Mol. Biol. 2004, 337, 951–968. [Google Scholar] [CrossRef] [PubMed]
- Barraud, P.; Gaudin, C.; Dardel, F.; Tisné, C. New insights into the formation of HIV-1 reverse transcription initiation complex. Biochimie 2007, 89, 1204–1210. [Google Scholar] [CrossRef] [PubMed]
- Tisné, C.; Roques, B.P.; Dardel, F. Specific recognition of primer tRNA Lys 3 by HIV-1 nucleocapsid protein: involvement of the zinc fingers and the N-terminal basic extension. Biochimie 2003, 85, 557–561. [Google Scholar] [CrossRef] [PubMed]
- Tisné, C.; Roques, B.P.; Dardel, F. Heteronuclear NMR studies of the interaction of tRNA(Lys)3 with HIV-1 nucleocapsid protein. J. Mol. Biol. 2001, 306, 443–454. [Google Scholar] [CrossRef] [PubMed]
- Henriet, S.; Mercenne, G.; Bernacchi, S.; Paillart, J.C.; Marquet, R. Tumultuous relationship between the human immunodeficiency virus type 1 viral infectivity factor (Vif) and the human APOBEC-3G and APOBEC-3F restriction factors. Microbiol. Mol. Biol. Rev. 2009, 73, 211–232. [Google Scholar] [CrossRef] [PubMed]
- Conticello, S.G. The AID/APOBEC family of nucleic acid mutators. Genome Biol. 2008, 9, 229. [Google Scholar] [CrossRef]
- Bishop, K.N.; Holmes, R.K.; Sheehy, A.M.; Davidson, N.O.; Cho, S.J.; Malim, M.H. Cytidine deamination of retroviral DNA by diverse APOBEC proteins. Curr. Biol. 2004, 14, 1392–1396. [Google Scholar] [CrossRef] [PubMed]
- Holmes, R.K.; Malim, M.H.; Bishop, K.N. APOBEC-mediated viral restriction: not simply editing? Trends Biochem. Sci. 2007, 32, 118–128. [Google Scholar] [CrossRef] [PubMed]
- Dettenhofer, M.; Cen, S.; Carlson, B.A.; Kleiman, L.; Yu, X.F. Association of human immunodeficiency virus type 1 Vif with RNA and its role in reverse transcription. J. Virol. 2000, 74, 8938–8945. [Google Scholar] [CrossRef] [PubMed]
- Bishop, K.N.; Holmes, R.K.; Malim, M.H. Antiviral potency of APOBEC proteins does not correlate with cytidine deamination. J. Virol. 2006, 80, 8450–8458. [Google Scholar] [CrossRef] [PubMed]
- Cen, S.; Guo, F.; Niu, M.; Saadatmand, J.; Deflassieux, J.; Kleiman, L. The interaction between HIV-1 Gag and APOBEC3G. J. Biol. Chem. 2004, 279, 33177–33184. [Google Scholar] [CrossRef] [PubMed]
- Guo, F.; Cen, S.; Niu, M.; Saadatmand, J.; Kleiman, L. Inhibition of formula-primed reverse transcription by human APOBEC3G during human immunodeficiency virus type 1 replication. J. Virol. 2006, 80, 11710–11722. [Google Scholar] [CrossRef] [PubMed]
- Guo, F.; Cen, S.; Niu, M.; Yang, Y.; Gorelick, R.J.; Kleiman, L. The interaction of APOBEC3G with human immunodeficiency virus type 1 nucleocapsid inhibits tRNA3Lys annealing to viral RNA. J. Virol. 2007, 81, 11322–11331. [Google Scholar] [CrossRef] [PubMed]
- Yang, Y.; Guo, F.; Cen, S.; Kleiman, L. Inhibition of initiation of reverse transcription in HIV-1 by human APOBEC3F. Virology 2007, 365, 92–100. [Google Scholar] [CrossRef] [PubMed]
- Henriet, S.; Sinck, L.; Bec, G.; Gorelick, R. J.; Marquet, R.; Paillart, J.C. Vif is a RNA chaperone that could temporally regulate RNA dimerization and the early steps of HIV-1 reverse transcription. Nucleic Acids Res. 2007, 35, 5141–5153. [Google Scholar] [CrossRef] [PubMed]
- Bishop, K.N.; Verma, M.; Kim, E.Y.; Wolinsky, S.M.; Malim, M.H. APOBEC3G inhibits elongation of HIV-1 reverse transcripts. PLoS Pathog. 2008, 4, e1000231. [Google Scholar] [CrossRef] [PubMed]
- Iwatani, Y.; Chan, D.S.; Wang, F.; Maynard, K.S.; Sugiura, W.; Gronenborn, A.M.; Rouzina, I.; Williams, M.C.; Musier-Forsyth, K.; Levin, J.G. Deaminase-independent inhibition of HIV-1 reverse transcription by APOBEC3G. Nucleic Acids Res. 2007, 35, 7096–7108. [Google Scholar] [CrossRef] [PubMed]
- Huang, Y.; Wang, J.; Shalom, A.; Li, Z.; Khorchid, A.; Wainberg, M.A.; Kleiman, L. Primer tRNA3Lys on the viral genome exists in unextended and two-base extended forms within mature human immunodeficiency virus type 1. J. Virol. 1997, 71, 726–728. [Google Scholar] [PubMed]
- Lori, F.; di Marzo Veronese, F.; de Vico, A.L.; Lusso, P.; Reitz Jr., M.S.; Gallo, R.C. Viral DNA carried by human immunodeficiency virus type 1 virions. J. Virol. 1992, 66, 5067–5074. [Google Scholar] [PubMed]
- Trono, D. Partial reverse transcripts in virions from human immunodeficiency and murine leukemia viruses. J. Virol. 1992, 66, 4893–4900. [Google Scholar] [PubMed]
- Houzet, L.; Morichaud, Z.; Mougel, M. Fully-spliced HIV-1 RNAs are reverse transcribed with similar efficiencies as the genomic RNA in virions and cells, but more efficiently in AZT-treated cells. Retrovirology 2007, 4, 30. [Google Scholar] [CrossRef]
- Houzet, L.; Morichaud, Z.; Didierlaurent, L.; Muriaux, D.; Darlix, J.L.; Mougel, M. Nucleocapsid mutations turn HIV-1 into a DNA-containing virus. Nucleic Acids Res. 2008, 36, 2311–2319. [Google Scholar] [CrossRef] [PubMed]
- Didierlaurent, L.; Houzet, L.; Morichaud, Z.; Darlix, J.L.; Mougel, M. The conserved N-terminal basic residues and zinc-finger motifs of HIV-1 nucleocapsid restrict the viral cDNA synthesis during virus formation and maturation. Nucleic Acids Res. 2008, 36, 4745–4753. [Google Scholar] [CrossRef] [PubMed]
- Thomas, J.A.; Bosche, W.J.; Shatzer, T.L.; Johnson, D.G.; Gorelick, R.J. Mutations in human immunodeficiency virus type 1 nucleocapsid protein zinc fingers cause premature reverse transcription. J. Virol. 2008, 82, 9318–9328. [Google Scholar] [CrossRef] [PubMed]
- Arhel, N.J.; Souquere-Besse, S.; Munier, S.; Souque, P.; Guadagnini, S.; Rutherford, S.; Prevost, M.C.; Allen, T.D.; Charneau, P. HIV-1 DNA Flap formation promotes uncoating of the pre-integration complex at the nuclear pore. EMBO J. 2007, 26, 3025–3037. [Google Scholar] [CrossRef] [PubMed]
- Zennou, V.; Petit, C.; Guetard, D.; Nerhbass, U.; Montagnier, L.; Charneau, P. HIV-1 genome nuclear import is mediated by a central DNA flap. Cell 2000, 101, 173–185. [Google Scholar] [CrossRef] [PubMed]
- Warrilow, D.; Meredith, L.; Davis, A.; Burrell, C.; Li, P.; Harrich, D. Cell factors stimulate human immunodeficiency virus type 1 reverse transcription in vitro. J. Virol. 2008, 82, 1425–1437. [Google Scholar] [CrossRef] [PubMed]
- Meredith, L.; Ducloux, C.; Isel, C.; Lin, M.H.; Marquet, R.; Harrich, D. A U5 represssor of reverse transcription is required for optimal HIV-1 infectivity and replication. 2010, submitted. [Google Scholar]
- El Safadi, Y.; Vivet-Boudou, V.; Marquet, R. HIV-1 reverse transcriptase inhibitors. Appl. Microbiol. Biotechnol. 2007, 75, 723–737. [Google Scholar] [CrossRef] [PubMed]
- Goldschmidt, V.; Marquet, R. Primer unblocking by HIV-1 reverse transcriptase and resistance to nucleoside RT inhibitors (NRTIs). Int. J. Biochem. Cell Biol. 2004, 36, 1687–1705. [Google Scholar] [CrossRef] [PubMed]
- Isel, C.; Ehresmann, C.; Walter, P.; Ehresmann, B.; Marquet, R. The emergence of different resistance mechanisms towards nucleoside inhibitors is explained by the properties of the wild type HIV-1 reverse transcriptase. J. Biol. Chem. 2001, 176, 48725–48732. [Google Scholar] [CrossRef]
- Vivet-Boudou, V.; Didierjean, J.; Isel, C.; Marquet, R. Nucleoside and nucleotide inhibitors of HIV-1 replication. Cell Mol. Life Sci. 2006, 63, 163–186. [Google Scholar] [CrossRef] [PubMed]
- Rigourd, M.; Lanchy, J.M.; Le Grice, S.F.; Ehresmann, B.; Ehresmann, C.; Marquet, R. Inhibition of the initiation of HIV-1 reverse transcription by 3'-azido- 3'-deoxythymidine. Comparison with elongation. J. Biol. Chem. 2000, 275, 26944–26951. [Google Scholar] [PubMed]
- Rigourd, M.; Ehresmann, C.; Parniak, M.A.; Ehresmann, B.; Marquet, R. Primer unblocking and rescue of DNA synthesis by azidothymidine (AZT)-resistant HIV-1 reverse transcriptase: comparison between initiation and elongation of reverse transcription and between (-) and (+) strand DNA synthesis. J. Biol. Chem. 2002, 277, 18611–18618. [Google Scholar] [CrossRef] [PubMed]
- Thomas, J.; Hergenrother, P. Targeting RNA with small molecules. Chem. Rev. 2008, 108, 1171–1224. [Google Scholar] [CrossRef] [PubMed]
- Tisne, C.; Guilliere, F.; Dardel, F. NMR-based identification of peptides that specifically recognize the d-arm of tRNA. Biochimie 2005, 87, 885–888. [Google Scholar] [CrossRef] [PubMed]
- Chung, F.; Tisne, C.; Lecourt, T.; Dardel, F.; Micouin, L. NMR-guided fragment-based approach for the design of tRNA(Lys3) ligands. Angew. Chem. Int. Ed. Engl. 2007, 46, 4489–4491. [Google Scholar] [CrossRef] [PubMed]
- Chung, F.; Tisne, C.; Lecourt, T.; Seijo, B.; Dardel, F.; Micouin, L. Design of tRNA(Lys)3 ligands: fragment evolution and linker selection guided by NMR spectroscopy. Chemistry 2009, 15, 7109–7116. [Google Scholar] [CrossRef] [PubMed]
- Lee, R.; Kaushik, N.; Modak, M.J.; Vinayak, R.; Pandey, V.N. Polyamide nucleic acid targeted to the primer binding site of the HIV-1 RNA genome blocks in vitro HIV-1 reverse transcription. Biochemistry 1998, 37, 900–910. [Google Scholar] [CrossRef] [PubMed]
- Tripathi, S.; Chaubey, B.; Barton, B.E.; Pandey, V.N. Anti HIV-1 virucidal activity of polyamide nucleic acid-membrane transducing peptide conjugates targeted to primer binding site of HIV-1 genome. Virology 2007, 363, 91–103. [Google Scholar] [CrossRef] [PubMed]
- Freund, F.; Boulme, F.; Michel, J.; Ventura, M.; Moreau, S.; Litvak, S. Inhibition of HIV-1 replication in vitro and in human infected cells by modified antisense oligonucleotides targeting the tRNALys3/RNA initiation complex. Antisense Nucleic Acid Drug Dev. 2001, 11, 301–315. [Google Scholar] [CrossRef] [PubMed]
- Kaushik, N.; Pandey, V.N. PNA targeting the PBS and A-loop sequences of HIV-1 genome destabilizes packaged tRNA3(Lys) in the virions and inhibits HIV-1 replication. Virology 2002, 303, 297–308. [Google Scholar] [CrossRef] [PubMed]
- Kaushik, N.; Talele, T.T.; Monel, R.; Palumbo, P.; Pandey, V.N. Destabilization of tRNA3Lys from the primer-binding site of HIV-1 genome by anti-A loop polyamide nucleotide analog. Nucleic Acids Res. 2001, 29, 5099–5106. [Google Scholar] [CrossRef] [PubMed]
- Coburn, G.A.; Cullen, B.R. Potent and specific inhibition of human immunodeficiency virus type 1 replication by RNA interference. J. Virol. 2002, 76, 9225–9231. [Google Scholar] [CrossRef] [PubMed]
- Jacque, J.M.; Triques, K.; Stevenson, M. Modulation of HIV-1 replication by RNA interference. Nature 2002, 418, 435–438. [Google Scholar] [CrossRef] [PubMed]
- Surabhi, R.M.; Gaynor, R.B. RNA interference directed against viral and cellular targets inhibits human immunodeficiency Virus Type 1 replication. J. Virol. 2002, 76, 12963–12973. [Google Scholar] [CrossRef] [PubMed]
- von Eije, K.J.; ter Brake, O.; Berkhout, B. Stringent testing identifies highly potent and escape-proof anti-HIV short hairpin RNAs. J. Gene Med. 2009, 11, 459–467. [Google Scholar] [CrossRef] [PubMed]
- von Eije, K.J.; ter Brake, O.; Berkhout, B. Human immunodeficiency virus type 1 escape is restricted when conserved genome sequences are targeted by RNA interference. J. Virol. 2008, 82, 2895–2903. [Google Scholar] [CrossRef] [PubMed]
- Das, A.T.; Brummelkamp, T.R.; Westerhout, E.M.; Vink, M.; Madiredjo, M.; Bernards, R.; Berkhout, B. Human immunodeficiency virus type 1 escapes from RNA interference-mediated inhibition. J. Virol. 2004, 78, 2601–2605. [Google Scholar] [CrossRef] [PubMed]
© 2010 by the authors; licensee Molecular Diversity Preservation International, Basel, Switzerland This is an open-access article distributed under the terms of the Creative Commons Attribution License, which permits unrestricted use, distribution, and reproduction in any medium, provided the original work is properly cited.
Share and Cite
Isel, C.; Ehresmann, C.; Marquet, R. Initiation of HIV Reverse Transcription. Viruses 2010, 2, 213-243. https://doi.org/10.3390/v2010213
Isel C, Ehresmann C, Marquet R. Initiation of HIV Reverse Transcription. Viruses. 2010; 2(1):213-243. https://doi.org/10.3390/v2010213
Chicago/Turabian StyleIsel, Catherine, Chantal Ehresmann, and Roland Marquet. 2010. "Initiation of HIV Reverse Transcription" Viruses 2, no. 1: 213-243. https://doi.org/10.3390/v2010213
APA StyleIsel, C., Ehresmann, C., & Marquet, R. (2010). Initiation of HIV Reverse Transcription. Viruses, 2(1), 213-243. https://doi.org/10.3390/v2010213




