Whole-Exome Sequencing Reveals Rare Genetic Variants in Saudi COVID-19 Patients with Extreme Phenotypes
Abstract
1. Introduction
2. Methodology
2.1. Study Subjects and Selection Criteria
2.2. Genomic DNA and Library Preparation
2.3. Next-Generation Sequencing
2.4. Quality Control and Preprocessing of Sequencing Reads
2.5. Read Mapping
2.6. Variant Discovery and Annotation
2.7. Variant Filtering and Classification
2.8. Protein–Protein Interaction Network Analysis
2.9. Functional Enrichment Analysis
- (1)
- Count in-network: The number of proteins in the user-provided network that are annotated with the term.
- (2)
- Strength: The log10 of the observed-to-expected ratio, quantifies the magnitude of the enrichment effect.
- (3)
- Signal: A weighted harmonic mean of the observed/expected ratio and the negative log10 of the False Discovery Rate (FDR), representing the overall enrichment significance.
- (4)
- False Discovery Rate (FDR): A measure of the statistical significance of the enrichment, accounting for multiple testing.
3. Results
3.1. Clinical Characteristics of COVID-19 Patients
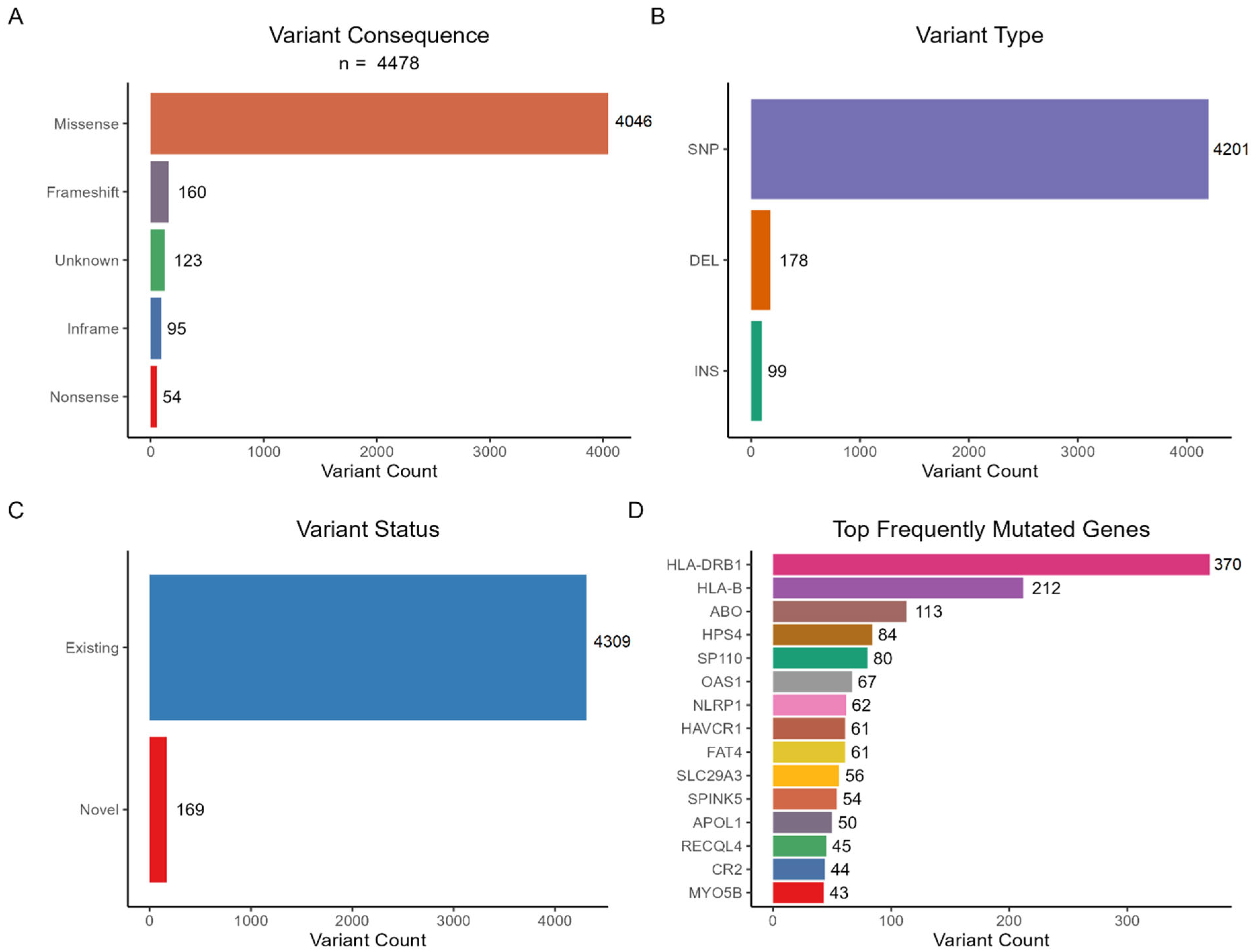
3.2. Identification of COVID-19-Specific Variants in Cohort
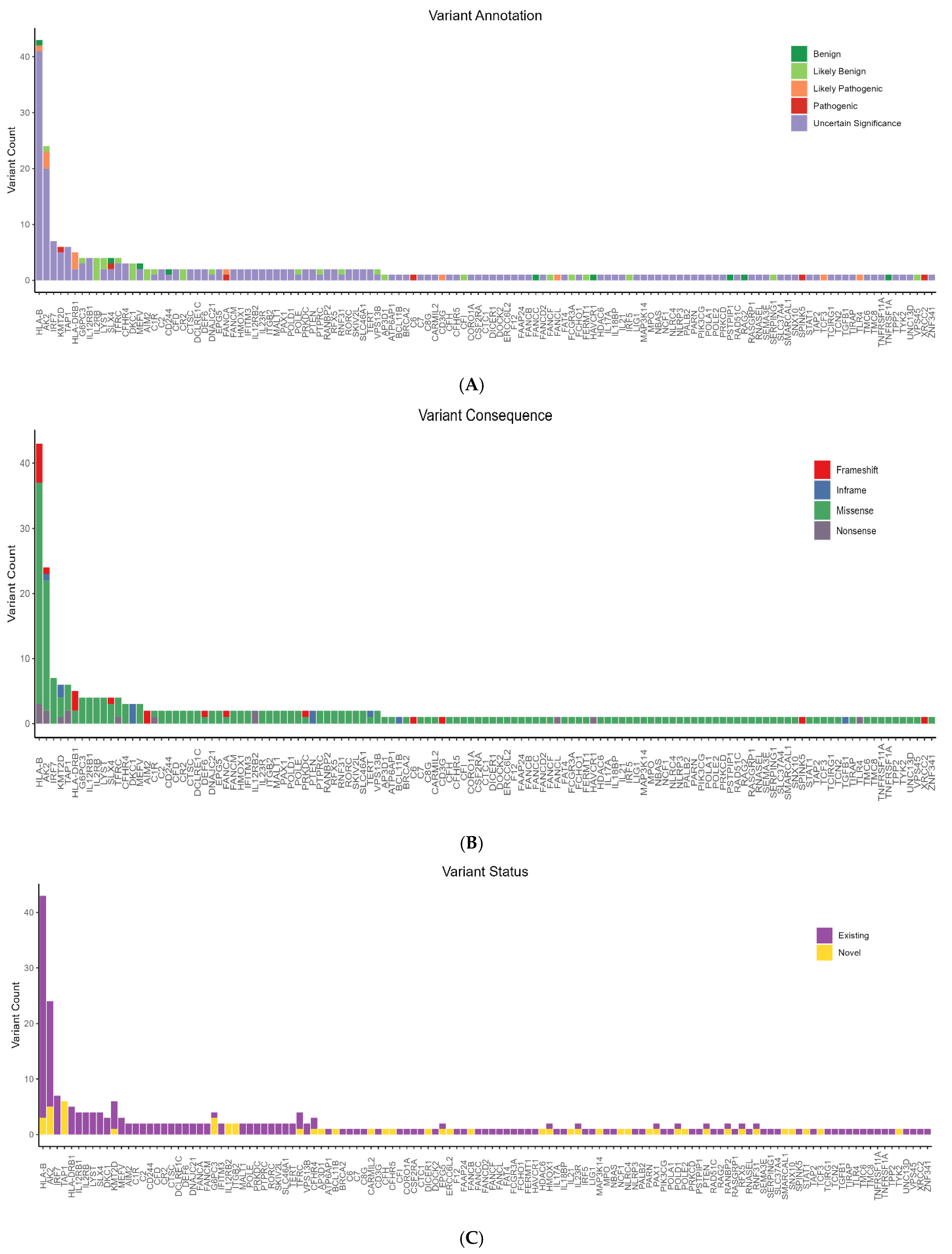
3.3. Identification of Rare Candidate Variants in the Cohort
3.4. Rare Candidate Pathogenic Variants in the Cohort
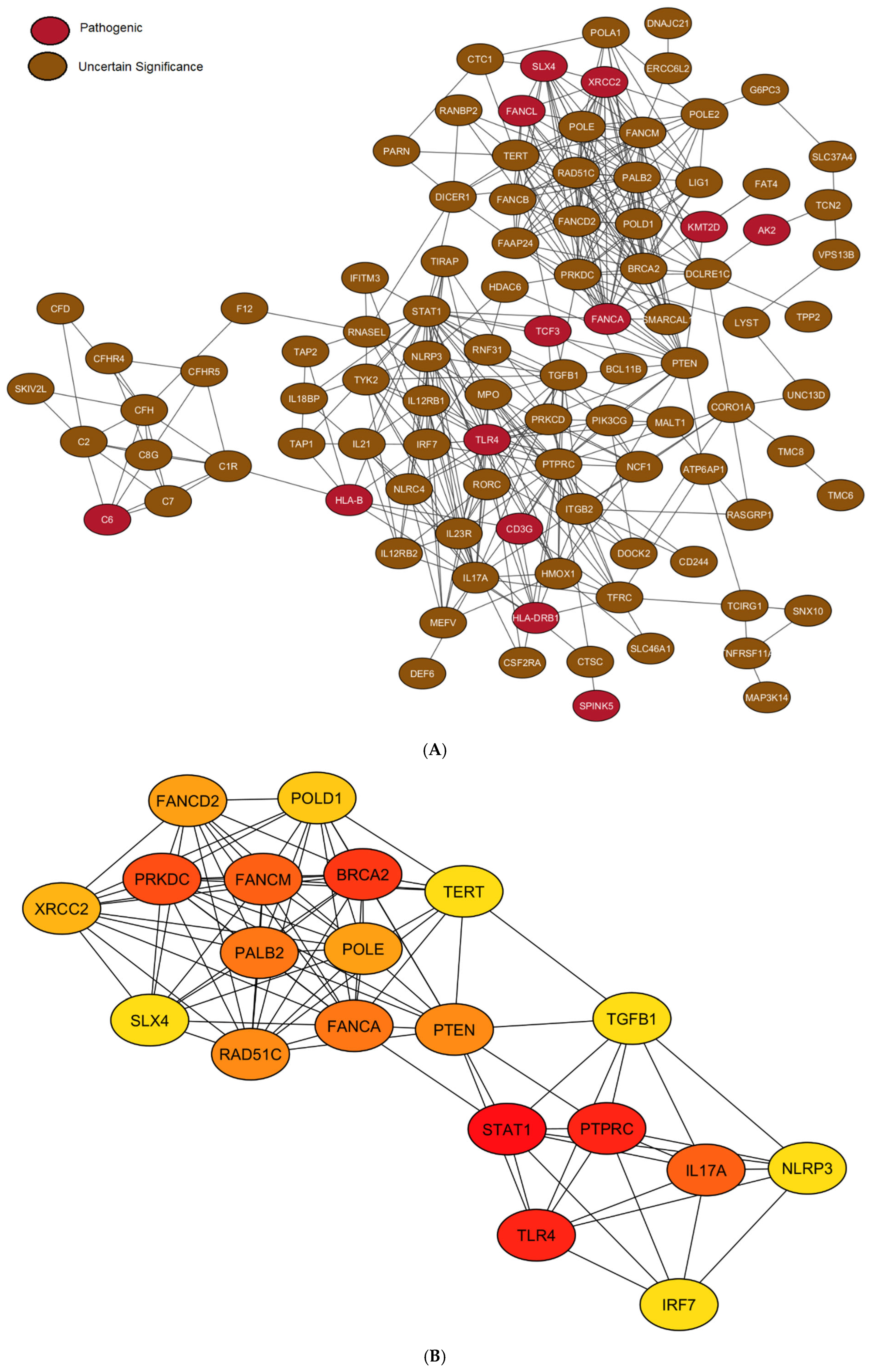
3.5. Protein–Protein Interaction Network Analysis
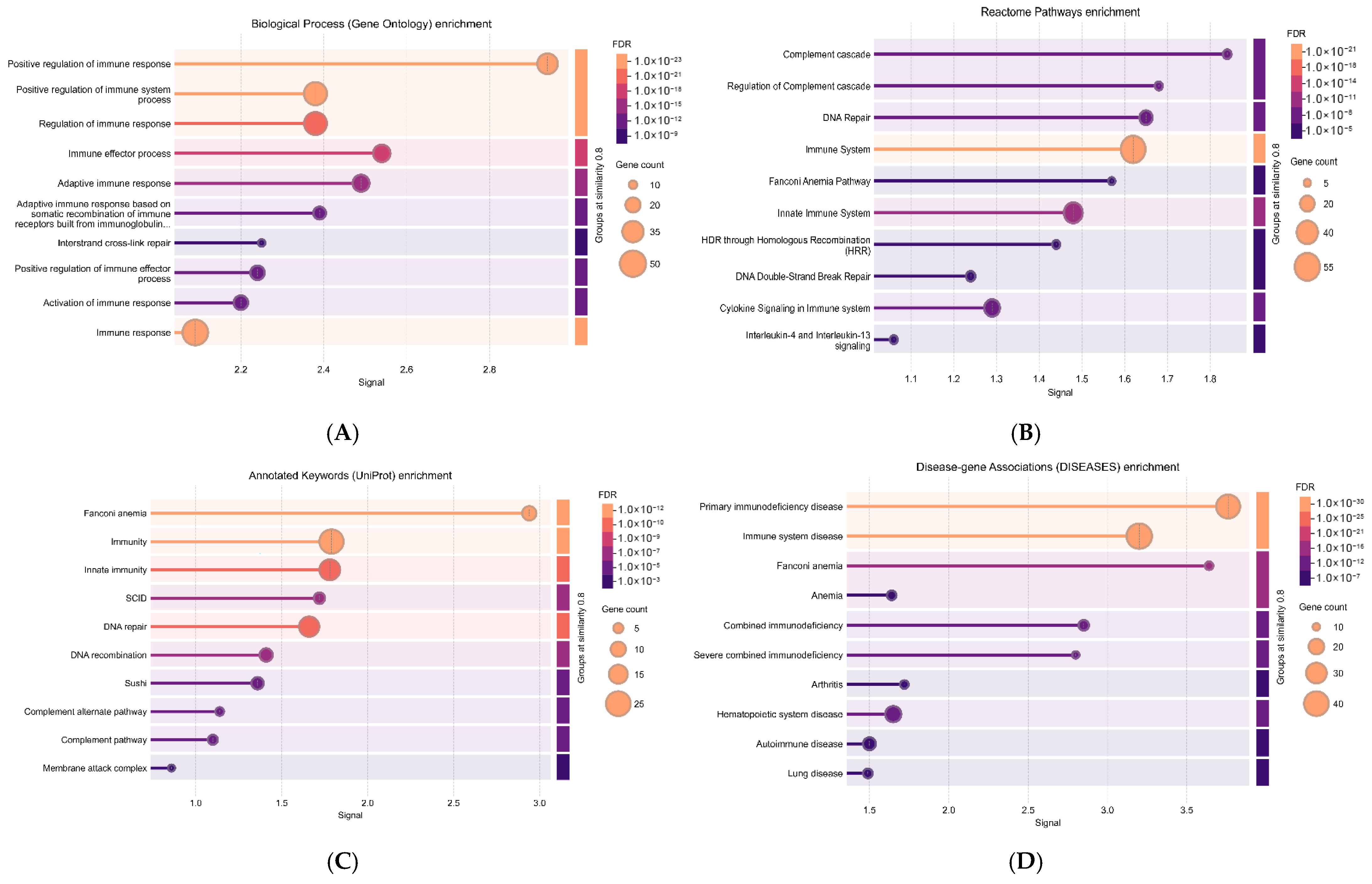
3.6. Functional Enrichment Analysis
4. Discussion
- The AK2 variants, which were observed in multiple patients, have a frequency of 0 in the control population data.
- The HLA-DRB1 variant, a key finding, also has a frequency of 0.
- Other variants like those in C6, CD3G, and SPINK5 have very low frequencies (e.g., 0.0032, 0.0006, and 0.00022, respectively) and are completely absent in the Middle Eastern population data shown.
5. Conclusions
6. Limitations of the Study
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Muralidar, S.; Ambi, S.V.; Sekaran, S.; Krishnan, U.M. The emergence of COVID-19 as a global pandemic: Understanding the epidemiology, immune response and potential therapeutic targets of SARS-CoV-2. Biochimie 2020, 179, 85–100. [Google Scholar] [CrossRef] [PubMed]
- World Health Organization. WHO COVID-19 Dashboard. Available online: https://data.who.int/dashboards/covid19/cases (accessed on 1 January 2025).
- Crits-Christoph, A.; Levy, J.I.; Pekar, J.E.; Goldstein, S.A.; Singh, R.; Hensel, Z.; Gangavarapu, K.; Rogers, M.B.; Moshiri, N.; Garry, R.F.; et al. Genetic tracing of market wildlife and viruses at the epicenter of the COVID-19 pandemic. Cell 2024, 187, 5468–5482.e11. [Google Scholar] [CrossRef]
- Petrosillo, N.; Viceconte, G.; Ergonul, O.; Ippolito, G.; Petersen, E. COVID-19, SARS and MERS: Are they closely related? Clin. Microbiol. Infect. 2020, 26, 729–734. [Google Scholar] [CrossRef] [PubMed]
- Liu, Y.; A Gayle, A.; Wilder-Smith, A.; Rocklöv, J. The reproductive number of COVID-19 is higher compared to SARS coronavirus. J. Travel Med. 2020, 27, taaa021. [Google Scholar] [CrossRef]
- Bigdelou, B.; Sepand, M.R.; Najafikhoshnoo, S.; Negrete, J.A.T.; Sharaf, M.; Ho, J.Q.; Sullivan, I.; Chauhan, P.; Etter, M.; Shekarian, T.; et al. COVID-19 and Preexisting Comorbidities: Risks, Synergies, and Clinical Outcomes. Front. Immunol. 2022, 13, 890517. [Google Scholar] [CrossRef]
- Beyerstedt, S.; Casaro, E.B.; Rangel, É.B. COVID-19: Angiotensin-converting enzyme 2 (ACE2) expression and tissue susceptibility to SARS-CoV-2 infection. Eur. J. Clin. Microbiol. Infect. Dis. 2021, 40, 905–919. [Google Scholar] [CrossRef]
- Pfortmueller, C.A.; Spinetti, T.; Urman, R.D.; Luedi, M.M.; Schefold, J.C. COVID-19-associated acute respiratory distress syndrome (CARDS): Current knowledge on pathophysiology and ICU treatment—A narrative review. Best Pract. Res. Clin. Anaesthesiol. 2021, 35, 351–368. [Google Scholar] [CrossRef]
- Ovsyannikova, I.G.; Haralambieva, I.H.; Crooke, S.N.; Poland, G.A.; Kennedy, R.B. The role of host genetics in the immune response to SARS-CoV-2 and COVID-19 susceptibility and severity. Immunol. Rev. 2020, 296, 205–219. [Google Scholar] [CrossRef]
- Rabaan, A.A.; Al Mutair, A.; Aljeldah, M.; Al Shammari, B.R.; Sulaiman, T.; Alshukairi, A.N.; Alfaresi, M.; Al-Jishi, J.M.; Al Bati, N.A.; Al-Mozaini, M.A.; et al. Genetic Variants and Protective Immunity Against SARS-CoV-2. Genes 2022, 13, 2355. [Google Scholar] [CrossRef]
- Clement, M.; Forbester, J.L.; Marsden, M.; Sabberwal, P.; Sommerville, M.S.; Wellington, D.; Dimonte, S.; Clare, S.; Harcourt, K.; Yin, Z.; et al. IFITM3 restricts virus-induced inflammatory cytokine production by limiting Nogo-B mediated TLR responses. Nat. Commun. 2022, 13, 5294. [Google Scholar] [CrossRef] [PubMed]
- Pei, X.M.; Yeung, M.H.Y.; Wong, A.N.N.; Tsang, H.F.; Yu, A.C.S.; Yim, A.K.Y.; Wong, S.C.C. Targeted Sequencing Approach and Its Clinical Applications for the Molecular Diagnosis of Human Diseases. Cells 2023, 12, 493. [Google Scholar] [CrossRef]
- Andrews, S. FastQC: A Quality Control Tool for High Throughput Sequence Data. 2010. Available online: http://www.bioinformatics.babraham.ac.uk/projects/fastqc/ (accessed on 25 August 2025).
- Krueger, F. Trim Galore: A Wrapper Around Cutadapt and FastQC to Consistently Apply Adapter and Quality Trimming to FastQ Files, with Extra Functionality for RRBS Data, v0.6.10; Zenodo: Meyrin, Switzerland, 2015. [Google Scholar] [CrossRef]
- Martin, M. Cutadapt removes adapter sequences from high-throughput sequencing reads. EMBnet. J. 2011, 17, 10–12. [Google Scholar] [CrossRef]
- Li, H. Aligning sequence reads, clone sequences and assembly contigs with BWA-MEM. arXiv 2013, arXiv:1303.3997. [Google Scholar] [CrossRef]
- Li, H.; Handsaker, B.; Wysoker, A.; Fennell, T.; Ruan, J.; Homer, N.; Marth, G.; Abecasis, G.; Durbin, R.; 1000 Genome Project Data Processing Subgroup. The Sequence Alignment/Map format and SAMtools. Bioinformatics 2009, 25, 2078–2079. [Google Scholar] [CrossRef]
- Van der Auwera, G.A.; Carneiro, M.O.; Hartl, C.; Poplin, R.; Del Angel, G.; Levy-Moonshine, A.; Jordan, T.; Shakir, K.; Roazen, D.; Thibault, J.; et al. From FastQ data to high confidence variant calls: The Genome Analysis Toolkit best practices pipeline. Curr. Protoc. Bioinform. 2013, 43, 11.10.1–11.10.33. [Google Scholar] [CrossRef]
- Pruitt, K.D.; Tatusova, T.; Brown, G.R.; Maglott, D.R. NCBI Reference Sequences (RefSeq): Current status, new features and genome annotation policy. Nucleic Acids Res. 2012, 40, D130–D135. [Google Scholar] [CrossRef] [PubMed]
- Amberger, J.S.; Bocchini, C.A.; Schiettecatte, F.; Scott, A.F.; Hamosh, A. OMIM.org: Online Mendelian Inheritance in Man (OMIM®), an online catalog of human genes and genetic disorders. Nucleic Acids Res. 2015, 43, D789–D798. [Google Scholar] [CrossRef] [PubMed]
- Landrum, M.J.; Lee, J.M.; Riley, G.R.; Jang, W.; Rubinstein, W.S.; Church, D.M.; Maglott, D.R. ClinVar: Public archive of relationships among sequence variation and human phenotype. Nucleic Acids Res. 2014, 42, D980–D985. [Google Scholar] [CrossRef] [PubMed]
- 1000 Genomes Project Consortium; Auton, A.; Brooks, L.D.; Durbin, R.M.; Garrison, E.P.; Kang, H.M.; Korbel, J.O.; Marchini, J.L.; McCarthy, S.; McVean, G.A.; et al. A global reference for human genetic variation. Nature 2015, 526, 68–74. [Google Scholar] [CrossRef]
- Lek, M.; Karczewski, K.J.; Minikel, E.V.; Samocha, K.E.; Banks, E.; Fennell, T.; O’Donnell-Luria, A.H.; Ware, J.S.; Hill, A.J.; Cummings, B.B.; et al. Analysis of protein-coding genetic variation in 60,706 humans. Nature 2016, 536, 285–291. [Google Scholar] [CrossRef]
- NHLBI GO Exome Sequencing Project. Sequencing Project. ESP 6500 exomes. EVS Release Version: V.0.0.25. (7 February 2014). Available online: https://genome.ucsc.edu/cgi-bin/hgTables?db=hg19&hgta_group=varRep&hgta_track=evsEsp6500&hgta_table=evsEsp6500&hgta_doSchema=describe+table+schema (accessed on 25 August 2025).
- Kumar, P.; Henikoff, S.; Ng, P.C. Predicting the effects of coding non-synonymous variants on protein function using the SIFT algorithm. Nat. Protoc. 2009, 4, 1073–1081. [Google Scholar] [CrossRef]
- Adzhubei, I.; Jordan, D.M.; Sunyaev, S.R. Predicting functional effect of human missense mutations using PolyPhen-2. Curr. Protoc. Hum. Genet. 2013, 7, Unit7.20. [Google Scholar] [CrossRef]
- Shihab, H.A.; Gough, J.; Mort, M.; Cooper, D.N.; Day, I.N.; Gaunt, T.R. Ranking non-synonymous single nucleotide polymorphisms based on disease concepts. Hum. Genom. 2014, 8, 11. [Google Scholar] [CrossRef] [PubMed]
- Schwarz, J.M.; Rödelsperger, C.; Schuelke, M.; Seelow, D. MutationTaster evaluates disease-causing potential of sequence alterations. Nat. Methods 2010, 7, 575–576. [Google Scholar] [CrossRef]
- Reva, B.; Antipin, Y.; Sander, C. Predicting the functional impact of protein mutations: Application to cancer genomics. Nucleic Acids Res. 2011, 39, e118. [Google Scholar] [CrossRef] [PubMed]
- Choi, Y.; Chan, A.P. PROVEAN web server: A tool to predict the functional effect of amino acid substitutions and indels. Bioinformatics 2015, 31, 2745–2747. [Google Scholar] [CrossRef]
- Kircher, M.; Witten, D.M.; Jain, P.; O’Roak, B.J.; Cooper, G.M.; Shendure, J. A general framework for estimating the relative pathogenicity of human genetic variants. Nat. Genet. 2014, 46, 310–315. [Google Scholar] [CrossRef]
- Hu, J.; Ng, P.C. SIFT Indel: Predictions for the functional effects of amino acid insertions/deletions in proteins. PLoS ONE 2013, 8, e77940. [Google Scholar] [CrossRef] [PubMed]
- Martin, A.R.; Williams, E.; Foulger, R.E.; Leigh, S.; Daugherty, L.C.; Niblock, O.; Leong, I.U.S.; Smith, K.R.; Gerasimenko, O.; Haraldsdottir, E.; et al. PanelApp crowdsources expert knowledge to establish consensus diagnostic gene panels. Nat. Genet. 2019, 51, 1560–1565. [Google Scholar] [CrossRef]
- Karczewski, K.J.; Francioli, L.C.; Tiao, G.; Cummings, B.B.; Alföldi, J.; Wang, Q.; Collins, R.L.; Laricchia, K.M.; Ganna, A.; Birnbaum, D.P.; et al. The mutational constraint spectrum quantified from variation in 141,456 humans. Nature 2020, 581, 434–443. [Google Scholar] [CrossRef]
- Richards, S.; Aziz, N.; Bale, S.; Bick, D.; Das, S.; Gastier-Foster, J.; Grody, W.W.; Hegde, M.; Lyon, E.; Spector, E.; et al. Standards and guidelines for the interpretation of sequence variants: A joint consensus recommendation of the American College of Medical Genetics and Genomics and the Association for Molecular Pathology. Genet. Med. Off. J. Am. Coll. Med. Genet. 2015, 17, 405–424. [Google Scholar] [CrossRef] [PubMed]
- Li, Q.; Wang, K. InterVar: Clinical Interpretation of Genetic Variants by the 2015 ACMG-AMP Guidelines. Am. J. Hum. Genet. 2017, 100, 267–280. [Google Scholar] [CrossRef] [PubMed]
- Kopanos, C.; Tsiolkas, V.; Kouris, A.; Chapple, C.E.; Albarca Aguilera, M.; Meyer, R.; Massouras, A. VarSome: The human genomic variant search engine. Bioinformatics 2019, 35, 1978–1980. [Google Scholar] [CrossRef] [PubMed]
- Szklarczyk, D.; Gable, A.L.; Nastou, K.C.; Lyon, D.; Kirsch, R.; Pyysalo, S.; Doncheva, N.T.; Legeay, M.; Fang, T.; Bork, P.; et al. The STRING database in 2021: Customizable protein-protein networks, and functional characterization of user-uploaded gene/measurement sets. Nucleic Acids Res. 2021, 49, D605–D612, Erratum in Nucleic Acids Res. 2021, 49, 10800. [Google Scholar] [CrossRef]
- Shannon, P.; Markiel, A.; Ozier, O.; Baliga, N.S.; Wang, J.T.; Ramage, D.; Amin, N.; Schwikowski, B.; Ideker, T. Cytoscape: A software environment for integrated models of biomolecular interaction networks. Genome Res. 2003, 13, 2498–2504. [Google Scholar] [CrossRef]
- Chin, C.H.; Chen, S.H.; Wu, H.H.; Ho, C.W.; Ko, M.T.; Lin, C.Y. Cytohubba: Identifying hub objects and sub-networks from complex interactome. BMC Syst. Biol. 2014, 8 (Suppl. 4), S11. [Google Scholar] [CrossRef]
- Fabião, J.; Sassi, B.; Pedrollo, E.F.; Gerchman, F.; Kramer, C.K.; Leitão, C.B.; Pinto, L.C. Why do men have worse COVID-19-related outcomes? A systematic review and meta-analysis with sex adjusted for age. Braz. J. Med. Biol. Res. 2022, 55, e11711. [Google Scholar] [CrossRef]
- Zhang, X.; Li, S.; Niu, S. ACE2 and COVID-19 and the resulting ARDS. Postgrad. Med. J. 2020, 96, 403–407. [Google Scholar] [CrossRef]
- Taborda, M.D.R.; Magana, N.V.; Palomera, C.D.; Pineda, J.C.; Rubi, M.G.; Gonzalez, R.A.; Arellano, A.R.; Suarez, A.L. A comparative analysis of estrogen receptors, ACE2 and cytokines in pre-and postmenopausal women and men with COVID-19. Front. Public Health 2025, 13, 1554024. [Google Scholar]
- Griffith, D.M.; Sharma, G.; Holliday, C.S.; Enyia, O.K.; Valliere, M.; Semlow, A.R.; Stewart, E.C.; Blumenthal, R.S. Men and COVID-19: A biopsychosocial approach to understanding sex differences in mortality and recommendations for practice and policy interventions. Prev. Chronic. Dis. 2020, 17, E63. [Google Scholar] [CrossRef]
- Spartano, S.; Faggiano, M.V.; Guidi, G.; D’Ambrosio, P.; Vaisfeld, A.; Novelli, A.; Falqui, S.; Cingolani, A.; Lambertenghi, L.; Visentin, A.; et al. Sex-Specific HLA Alleles Contribute to the Modulation of COVID-19 Severity. Int. J. Mol. Sci. 2024, 25, 13198. [Google Scholar] [CrossRef]
- Ejaz, H.; Alsrhani, A.; Zafar, A.; Javed, H.; Junaid, K.; Abdalla, A.E.; Abosalif, K.O.A.; Ahmed, Z.; Younas, S. COVID-19 and comorbidities: Deleterious impact on infected patients. J. Infect. Public Health 2020, 13, 1833–1839. [Google Scholar] [CrossRef]
- Basu, A.; Agwu, J.C.; Barlow, N.; Lee, B. Hypertension is the major predictor of poor outcomes among inpatients with COVID-19 infection in the UK: A retrospective cohort study. BMJ Open 2021, 11, e047561. [Google Scholar] [CrossRef] [PubMed]
- Szarpak, L.; Mierzejewska, M.; Jurek, J.; Kochanowska, A.; Gasecka, A.; Truszewski, Z.; Pruc, M.; Blek, N.; Rafique, Z.; Filipiak, K.J.; et al. Effect of Coronary Artery Disease on COVID-19-Prognosis and Risk Assessment: A Systematic Review and Meta-Analysis. Biology 2022, 11, 221. [Google Scholar] [CrossRef] [PubMed]
- Bo, K.; Li, W.; Zhang, H.; Wang, Y.; Zhou, Z.; Gao, Y.; Sun, Z.; Lian, J.; Wang, H.; Xu, L. Association of stress hyperglycemia ratio with left ventricular function and microvascular obstruction in patients with ST-segment elevation myocardial infarction: A 3.0 T cardiac magnetic resonance study. Cardiovasc. Diabetol. 2024, 23, 179. [Google Scholar] [CrossRef]
- Ran, E.; Zou, Y.; Zhao, C.; Liu, K.; Yuan, J.; Yang, W.; Zhao, L.; Yang, Q.; Yang, J.; Ju, X.; et al. COVID-19 in discharged patients with diabetes and chronic kidney disease: One-year follow-up and evaluation. Front. Endocrinol. 2025, 16, 1519993. [Google Scholar] [CrossRef] [PubMed]
- Migliorini, F.; Torsiello, E.; Spiezia, F.; Oliva, F.; Tingart, M.; Maffulli, N. Association between HLA genotypes and COVID-19 susceptibility, severity and progression: A comprehensive review of the literature. Eur. J. Med. Res. 2021, 26, 84. [Google Scholar] [CrossRef]
- Alduaij, W.; Al-Youha, S.; Al-Serri, A.; Almazeedi, S.; Al-Haddad, M.; Jamal, M.H.; Al-Sabah, S. The Impact of ABO Blood Group on Clinical Outcomes and Susceptibility in COVID-19: A Retrospective Population Study of 3305 Patients. Blood 2020, 136, 32–33. [Google Scholar] [CrossRef]
- Bubnova, L.; Pavlova, I.; Terentieva, M.; Glazanova, T.; Belyaeva, E.; Sidorkevich, S.; Bashketova, N.; Chkhingeria, I.; Kozhemyakina, M.; Azarov, D.; et al. HLA Genotypes in Patients with Infection Caused by Different Strains of SARS-CoV-2. Int. J. Env. Res. Public Health 2022, 19, 14024. [Google Scholar] [CrossRef]
- Hoseinnezhad, T.; Soltani, N.; Ziarati, S.; Behboudi, E.; Mousavi, M.J. The role of HLA genetic variants in COVID-19 susceptibility, severity, and mortality: A global review. J. Clin. Lab. Anal. 2024, 38, e25005. [Google Scholar] [CrossRef]
- Fischer, J.C.; Schmidt, A.G.; Bölke, E.; Uhrberg, M.; Keitel, V.; Feldt, T.; Jensen, B.; Häussinger, D.; Adams, O.; Schneider, E.M.; et al. Association of HLA genotypes, AB0 blood type and chemokine receptor 5 mutant CD195 with the clinical course of COVID-19. Eur. J. Med. Res. 2021, 26, 107. [Google Scholar] [CrossRef] [PubMed]
- Fakhkhari, M.; Caidi, H.; Sadki, K. HLA alleles associated with COVID-19 susceptibility and severity in different populations: A systematic review. Egypt. J. Med. Hum. Genet. 2023, 24, 10. [Google Scholar] [CrossRef]
- Goel, R.; Bloch, E.M.; Pirenne, F.; Al-Riyami, A.Z.; Crowe, E.; Dau, L.; Land, K.; Townsend, M.; Jecko, T.; Rahimi-Levene, N.; et al. ISBT COVID-19 Working Group. ABO blood group and COVID-19: A review on behalf of the ISBT COVID-19 Working Group. Vox Sang. 2021, 116, 849–861. [Google Scholar] [CrossRef]
- O’Sullivan, J.M.; Ward, S.; Fogarty, H.; O’Donnell, J.S. More on ‘Association between ABO blood groups and risk of SARS-CoV-2 pneumonia’. Br. J. Haematol. 2020, 190, 27–28. [Google Scholar] [CrossRef]
- Avila, J.; Long, B.; Holladay, D.; Gottlieb, M. Thrombotic complications of COVID-19. Am. J. Emerg. Med. 2021, 39, 213–218. [Google Scholar] [CrossRef] [PubMed]
- Thomason, P.A.; Corbyn, R.; Lilla, S.; Sumpton, D.; Gilbey, T.; Insall, R.H. Biogenesis of lysosome-related organelles complex-2 is an evolutionarily ancient proto-coatomer complex. Curr. Biol. 2024, 34, 3564–3581.e6. [Google Scholar] [CrossRef]
- Merideth, M.A.; Introne, W.J.; Wang, J.A.; O’Brien, K.J.; Huizing, M.; Gochuico, B.R. Genetic variants associated with Hermansky-Pudlak syndrome. Platelets 2020, 31, 544–547. [Google Scholar] [CrossRef]
- Fraschilla, I.; Jeffrey, K.L. The Speckled Protein (SP) Family: Immunity’s Chromatin Readers. Trends Immunol. 2020, 41, 572–585. [Google Scholar] [CrossRef]
- Ma, W.; Huang, G.; Wang, Z.; Wang, L.; Gao, Q. IRF7: Role and regulation in immunity and autoimmunity. Front. Immunol. 2023, 14, 1236923. [Google Scholar] [CrossRef]
- Wang, L.; Zhu, Y.; Zhang, N.; Xian, Y.; Tang, Y.; Ye, J.; Reza, F.; He, G.; Wen, X.; Jiang, X. The multiple roles of interferon regulatory factor family in health and disease. Signal Transduct. Target. Ther. 2024, 9, 282. [Google Scholar] [CrossRef] [PubMed]
- Martínez-Pomar, N.; Cunill, V.; Segura-Guerrero, M.; Pol-Pol, E.; Oblitas, D.E.; Pons, J.; Ayestarán, I.; Pruneda, P.C.; Losada, I.; Toledo-Pons, N.; et al. Hyperinflammatory Immune Response in COVID-19: Host Genetic Factors in Pyrin Inflammasome and Immunity to Virus in a Spanish Population from Majorca Island. Biomedicines 2023, 11, 2548. [Google Scholar] [CrossRef] [PubMed]
- Ritz, U.; Seliger, B. The transporter associated with antigen processing (TAP): Structural integrity, expression, function, and its clinical relevance. Mol. Med. 2001, 7, 149–158. [Google Scholar] [CrossRef]
- Fujisawa, K. Regulation of Adenine Nucleotide Metabolism by Adenylate Kinase Isozymes: Physiological Roles and Diseases. Int. J. Mol. Sci. 2023, 24, 5561. [Google Scholar] [CrossRef]
- Pannicke, U.; Hönig, M.; Hess, I.; Friesen, C.; Holzmann, K.; Rump, E.-M.; Barth, T.F.; Rojewski, M.T.; Schulz, A.; Boehm, T.; et al. Reticular dysgenesis (aleukocytosis) is caused by mutations in the gene encoding mitochondrial adenylate kinase 2. Nat. Genet. 2009, 41, 101–105. [Google Scholar] [CrossRef]
- Froimchuk, E.; Jang, Y.; Ge, K. Histone H3 lysine 4 methyltransferase KMT2D. Gene 2017, 627, 337–342. [Google Scholar] [CrossRef]
- Wu, J.; Chun, C.; Lagunas, A.M.; Crowe, D.L. Lysine Methyltransferase 2D Regulates Immune Response and Metastasis in Head and Neck Cancer. Anticancer Res. 2024, 44, 3231–3242. [Google Scholar] [CrossRef]
- Potter, S.J.; Zhang, L.; Kotliar, M.; Wu, Y.; Schafer, C.; Stefan, K.; Boukas, L.; Qu’d, D.; Bodamer, O.; Simpson, B.N.; et al. KMT2D regulates activation, localization, and integrin expression by T-cells. Front. Immunol. 2024, 15, 1341745. [Google Scholar] [CrossRef]
- McKinney, C.; Ellison, M.; Briones, N.J.; Baroffio, A.; Murphy, J.; Tran, A.D.; Reisz, J.A.; D’Alessandro, A.; Ambruso, D.R. Metabolic abnormalities in G6PC3-deficient human neutrophils result in severe functional defects. Blood Adv. 2020, 4, 5888–5901. [Google Scholar] [CrossRef] [PubMed]
- Ackermann, M.; Anders, H.-J.; Bilyy, R.; Bowlin, G.L.; Daniel, C.; De Lorenzo, R.; Egeblad, M.; Henneck, T.; Hidalgo, A.; Hoffmann, M.; et al. Patients with COVID-19: In the dark-NETs of neutrophils. Cell Death Differ. 2021, 28, 3125–3139. [Google Scholar] [CrossRef] [PubMed]
- Ullrich, K.A.; Schulze, L.L.; Paap, E.M.; Müller, T.M.; Neurath, M.F.; Zundler, S. Immunology of IL-12: An update on functional activities and implications for disease. EXCLI J. 2020, 19, 1563–1589. [Google Scholar]
- Henry, C.J.; Ornelles, D.A.; Mitchell, L.M.; Brzoza-Lewis, K.L.; Hiltbold, E.M. IL-12 produced by dendritic cells augments CD8+ T cell activation through the production of the chemokines CCL1 and CCL17. J. Immunol. 2008, 181, 8576–8584. [Google Scholar] [CrossRef] [PubMed]
- Fayyaz, H.; Zaman, A.; Haider, N.; Waris, R.; Hussain, M.; Raza, S.I.; Ahmad, W.; Ullah, I. Sequence variants underlying severe combined immunodeficiency and leukocyte adhesion deficiency type 1 in six consanguineous families. Immunogenetics 2024, 76, 351–360. [Google Scholar] [CrossRef] [PubMed]
- Siekacz, K.; Kumor-Kisielewska, A.; Miłkowska-Dymanowska, J.; Pietrusińska, M.; Bartczak, K.; Majewski, S.; Stańczyk, A.; Piotrowski, W.J.; Białas, A.J. Soluble ITGaM and ITGb2 Integrin Subunits Are Involved in Long-Term Pulmonary Complications after COVID-19 Infection. J. Clin. Med. 2023, 12, 342. [Google Scholar] [CrossRef] [PubMed]

| COVID-19 Patients | |
|---|---|
| Characteristic | n = 16 |
| Gender | |
| Female | 9 (56%) 1 |
| Male | 7 (44%) |
| Age | 59 (50–69) |
| Hypertension | 12 (75%) |
| Diabetes | |
| No | 2 (13%) |
| Unknown | 9 (56%) |
| Yes | 5 (31%) |
| Smoking | |
| No | 6 (38%) |
| Unknown | 9 (56%) |
| Yes | 1 (6.3%) |
| Fever | |
| No | 3 (19%) |
| Unknown | 3 (19%) |
| Yes | 10 (63%) |
| Cough | |
| Unknown | 2 (13%) |
| Yes | 14 (88%) |
| Chronic kidney disease | |
| No | 7 (44%) |
| Unknown | 8 (50%) |
| Yes | 1 (6.3%) |
| Ischemic heart disease | 8 (50%) |
| Immunosuppressive medications | |
| No | 2 (13%) |
| Unknown | 3 (19%) |
| Yes | 11 (69%) |
| Hospital duration (Days) | 12 (9–16) |
| ICU duration (Days) | 8 (5–12) |
| Outcome | |
| Death | 8 (50%) |
| Improved | 8 (50%) |
| Gene | Variation | Protein Change | Variant ID | Clinvar Classification | ACMG Classification 1 | Interpreted Classification |
|---|---|---|---|---|---|---|
| AK2 | NM_001319139.2:c.52_553insACATC5>C | p.*185delinsTS* | rs2035782928 | - | LP | LP |
| AK2 | NM_001319141.2:c.500_501insGACAT | p.I167Mfs*8 | rs1398317449 | LP | - | LP |
| C6 | NM_000065.5:c.1879delG | p.D627Tfs*4 | rs61469168 | P | - | P |
| CD3G | NM_000073.3:c.205delA | p.K71Nfs*40 | rs570768621 | P/LP | LP | LP |
| FANCA | NM_000135.4:c.2762A>T | p.K921I | rs879255255 | LP | - | LP |
| FANCA | NM_000135.4:c.3761_3762del | p.E1254Gfs*23 | rs868273545 | P | - | P |
| FANCL | NM_001114636.1:c.223C>T | p.Q75X | rs1693048371 | - | LP | LP |
| HLA-B | NM_005514.8:c.604_605del | p.T202Afs*18 | rs761596463 | - | LP | LP |
| HLA-DRB1 | NM_002124.3:c.593_612del | p.E198Gfs*18 | rs140357311 | - | LP | LP |
| KMT2D | NM_003482.4:c.11203C>T | p.Q3735X | rs1943059195 | P | LP | P |
| SLX4 | NM_032444.4:c.1093delC | p.Q365Sfs*32 | rs1218169126 | P | - | P |
| SPINK5 | NM_001127698.2:c.2459dupA | p.K824Efs*4 | rs587777750 | P | - | P |
| TCF3 | NM_001136139.4:c.1783G>A | p.V595I | Novel | - | LP | LP |
| TLR4 | NM_138554.5:c.2191C>T | p.R731X | rs201670644 | - | LP | LP |
| XRCC2 | NM_005431.2:c.350dupT | p.L117Ffs*6 | rs764640893 | LP | P | P |
| Gene | Variation | Protein Change | Variant ID | Control Frequency (1000 Genome Combined Populations) | Control Frequency (GenomeAD Exome Combined Populations) | Control Frequency (GenomeAD Exome Middle East Populations) | Control Frequency (GenomeAD Genome Middle East Populations) |
|---|---|---|---|---|---|---|---|
| AK2 | NM_001319139.2:c.52_553insACATC5>C | p.*185delinsTS* | rs2035782928 | - | - | - | - |
| AK2 | NM_001319141.2:c.500_501insGACAT | p.I167Mfs*8 | rs1398317449 | - | 3.57 × 10−5 | 0 | 0 |
| C6 | NM_000065.5:c.1879delG | p.D627Tfs*4 | rs61469168 | 0.0032 | 0.000314 | 0.000349 | 0 |
| CD3G | NM_000073.3:c.205delA | p.K71Nfs*40 | rs570768621 | 0.0006 | 5.87 × 10−5 | 0.000175 | 0 |
| FANCA | NM_000135.4:c.2762A>T | p.K921I | rs879255255 | - | - | - | - |
| FANCA | NM_000135.4:c.3761_3762del | p.E1254Gfs*23 | rs868273545 | - | 6.84 × 10−7 | 0 | 0 |
| FANCL | NM_001114636.1:c.223C>T | p.Q75X | rs1693048371 | - | 1.37 × 10−6 | 0 | - |
| HLA-B | NM_005514.8:c.604_605del | p.T202Afs*18 | rs761596463 | - | - | - | 0 |
| HLA-DRB1 | NM_002124.3:c.593_612del | p.E198Gfs*18 | rs140357311 | - | - | - | - |
| KMT2D | NM_003482.4:c.11203C>T | p.Q3735X | rs1943059195 | - | - | - | - |
| SLX4 | NM_032444.4:c.1093delC | p.Q365Sfs*32 | rs1218169126 | - | 5.47 × 10−6 | 0 | 0 |
| SPINK5 | NM_001127698.2:c.2459dupA | p.K824Efs*4 | rs587777750 | - | 0.00022 | 0 | 0 |
| TCF3 | NM_001136139.4:c.1783G>A | p.V595I | Novel | - | - | - | - |
| TLR4 | NM_138554.5:c.2191C>T | p.R731X | rs201670644 | - | 5.82 × 10−5 | 0.001057 | 0 |
| XRCC2 | NM_005431.2:c.350dupT | p.L117Ffs*6 | rs764640893 | - | - | - | - |
| Term Category | Enriched Term | Strength | Signal | padj | Enriched Genes |
|---|---|---|---|---|---|
| Diseases | Primary immunodeficiency disease | 1.17 | 3.76 | 8.2 × 10−30 | UNC13D, CORO1A, MEFV, MPO, CFHR5, IL12RB2, NCF1, RFX5, C2, RASGRP1, PRKDC, IL23R, TMC8, PRKCD, CARMIL2, NLRP3, TAP1, BCL11B, HLA-DRB1, STAT1, CFH, ATP6AP1, TAP2, ZNF341, DCLRE1C, IRF7, ITGB2, HLA-B, PTPRC, DOCK2, CD3G, TYK2, IL18BP, TMC6, IL12RB1, IL21, IL17A, AK2 |
| Fanconi anemia | 1.90 | 3.64 | 2.1 × 10−14 | PALB2, FANCM, FANCD2, SLX4, RAD51C, XRCC2, BRCA2, FANCA, FANCL, TNFRSF11A, FANCB | |
| Immune system disease | 1.05 | 3.20 | 2.5 × 10−29 | UNC13D, CORO1A, MEFV, MPO, CFHR5, IL12RB2, NCF1, RFX5, C2, RASGRP1, PRKDC, RNF31, IL23R, TMC8, PRKCD, CARMIL2, NLRP3, TAP1, BCL11B, HLA-DRB1, STAT1, CFH, ATP6AP1, TAP2, ZNF341, DCLRE1C, LYST, FAT4, IRF7, ITGB2, HLA-B, PTPRC, DOCK2, CD3G, TYK2, IL18BP, TMC6, IL12RB1, IL21, IL17A, MALT1, AK2 | |
| Combined immunodeficiency | 1.49 | 2.85 | 8.2 × 10−13 | CORO1A, RFX5, PRKDC, CARMIL2, TAP1, BCL11B, TAP2, DCLRE1C, ITGB2, PTPRC, DOCK2, CD3G, AK2 | |
| Severe combined immunodeficiency | 1.72 | 2.80 | 1.7 × 10−11 | CORO1A, RFX5, PRKDC, TAP1, BCL11B, TAP2, DCLRE1C, PTPRC, CD3G, AK2 | |
| GO Process | Positive regulation of immune response | 1.08 | 2.94 | 2.0 × 10−22 | TGFB1, CFHR5, C6, C2, RASGRP1, PRKDC, RNF31, IL23R, C7, PRKCD, NLRP3, HLA-DRB1, CFHR4, CFH, C8G, TLR4, TAP2, TFRC, TIRAP, IRF7, ITGB2, NLRC4, HLA-B, PTPRC, CD3G, TYK2, C1R, CFD, IL12RB1, FCHO1, IL21, IL17A, MALT1 |
| Immune effector process | 1.08 | 2.54 | 1.1 × 10−16 | UNC13D, CORO1A, MPO, CTSC, CFHR5, C6, TCIRG1, C2, RASGRP1, C7, RORC, PRKCD, CFHR4, CFH, CD244, C8G, TLR4, TAP2, LYST, IRF7, PIK3CG, DOCK2, C1R, CFD, IL21 | |
| Adaptive immune response | 1.09 | 2.49 | 4.9 × 10−16 | UNC13D, TGFB1, CTSC, C6, TCIRG1, C2, C7, RORC, PRKCD, TAP1, HLA-DRB1, CD244, C8G, TLR4, TAP2, DCLRE1C, IRF7, HLA-B, PIK3CG, CD3G, IL18BP, C1R, TNFRSF11A, IL17A | |
| Adaptive immune response based on somatic recombination of immune receptors built from immunoglobulin superfamily domains | 1.24 | 2.39 | 2.4 × 10−12 | UNC13D, TGFB1, CTSC, C6, TCIRG1, C2, C7, RORC, PRKCD, HLA-DRB1, C8G, TLR4, TAP2, IRF7, IL18BP, C1R | |
| Positive regulation of immune system process | 0.92 | 2.38 | 2.0 × 10−22 | UNC13D, HMOX1, CORO1A, TGFB1, CTSC, CFHR5, IL12RB2, TCF3, C6, C2, RASGRP1, PRKDC, RNF31, IL23R, C7, PRKCD, NLRP3, HLA-DRB1, CFHR4, CFH, CD244, C8G, TLR4, TAP2, TFRC, TIRAP, IRF7, ITGB2, NLRC4, HLA-B, PTPRC, CD3G, TYK2, C1R, CFD, IL12RB1, FCHO1, IL21, IL17A, MALT1 | |
| Reactome | Complement cascade | 1.44 | 1.84 | 5.2 × 10−8 | CFHR5, C6, C2, C7, CFHR4, CFH, C8G, C1R, CFD |
| Regulation of complement cascade | 1.47 | 1.68 | 3.0 × 10−7 | CFHR5, C6, C2, C7, CFHR4, CFH, C8G, C1R | |
| DNA repair | 1.00 | 1.65 | 1.8 × 10−9 | POLE2, PALB2, LIG1, FANCM, FANCD2, SLX4, PRKDC, POLE, RAD51C, XRCC2, DCLRE1C, BRCA2, FANCA, FANCL, FAAP24, POLD1, FANCB | |
| Immune system | 0.69 | 1.62 | 3.9 × 10−21 | UNC13D, HMOX1, MEFV, TGFB1, MPO, CTSC, CFHR5, IL12RB2, C6, TCIRG1, RANBP2, NCF1, C2, RASGRP1, PRKDC, IL23R, C7, RORC, PRKCD, NLRP3, TAP1, HLA-DRB1, STAT1, CFHR4, CFH, RNASEL, C8G, PTEN, TLR4, TAP2, TPP2, TIRAP, IRF7, ITGB2, IFITM3, NLRC4, CSF2RA, HLA-B, PTPRC, PIK3CG, DOCK2, CD3G, TYK2, IL18BP, C1R, TMC6, TNFRSF11A, CFD, IL12RB1, MAP3K14, IL21, IL17A, MALT1 | |
| Fanconi anemia pathway | 1.54 | 1.57 | 1.2 × 10−6 | FANCM, FANCD2, SLX4, FANCA, FANCL, FAAP24, FANCB | |
| UniProt Keywords | Fanconi anemia | 1.89 | 2.94 | 1.3 × 10−11 | PALB2, FANCD2, SLX4, RAD51C, XRCC2, BRCA2, FANCA, FANCL, FANCB |
| Immunity | 0.91 | 1.79 | 2.1 × 10−12 | MEFV, C6, C2, PRKDC, IL23R, C7, NLRP3, TAP1, HLA-DRB1, CFH, CD244, C8G, TLR4, TAP2, DCLRE1C, TIRAP, IRF7, IFITM3, NLRC4, HLA-B, PIK3CG, CD3G, C1R, CFD | |
| Innate immunity | 1.00 | 1.78 | 1.2 × 10−10 | MEFV, C6, C2, PRKDC, IL23R, C7, NLRP3, CFH, CD244, C8G, TLR4, TIRAP, IRF7, IFITM3, NLRC4, HLA-B, C1R, CFD | |
| SCID | 1.74 | 1.72 | 5.4 × 10−7 | RFX5, PRKDC, BCL11B, DCLRE1C, PTPRC, AK2 | |
| DNA repair | 0.98 | 1.66 | 8.1 × 10−10 | PALB2, LIG1, FANCM, FANCD2, SLX4, PRKDC, POLE, RAD51C, XRCC2, DCLRE1C, BRCA2, FANCA, FANCL, FAAP24, POLD1, FANCB, ERCC6L2 |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2025 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Mir, R.; Ullah, M.F.; Elfaki, I.; Alanazi, M.A.; Algehainy, N.A.; Altemani, F.H.; Moawadh, M.S.; Tayeb, F.J.; Alsayed, B.A.; Mir, M.M.; et al. Whole-Exome Sequencing Reveals Rare Genetic Variants in Saudi COVID-19 Patients with Extreme Phenotypes. Viruses 2025, 17, 1198. https://doi.org/10.3390/v17091198
Mir R, Ullah MF, Elfaki I, Alanazi MA, Algehainy NA, Altemani FH, Moawadh MS, Tayeb FJ, Alsayed BA, Mir MM, et al. Whole-Exome Sequencing Reveals Rare Genetic Variants in Saudi COVID-19 Patients with Extreme Phenotypes. Viruses. 2025; 17(9):1198. https://doi.org/10.3390/v17091198
Chicago/Turabian StyleMir, Rashid, Mohammad Fahad Ullah, Imadeldin Elfaki, Mohammad A. Alanazi, Naseh A. Algehainy, Faisal H. Altemani, Mamdoh S. Moawadh, Faris J. Tayeb, Badr A. Alsayed, Mohammad Muzaffar Mir, and et al. 2025. "Whole-Exome Sequencing Reveals Rare Genetic Variants in Saudi COVID-19 Patients with Extreme Phenotypes" Viruses 17, no. 9: 1198. https://doi.org/10.3390/v17091198
APA StyleMir, R., Ullah, M. F., Elfaki, I., Alanazi, M. A., Algehainy, N. A., Altemani, F. H., Moawadh, M. S., Tayeb, F. J., Alsayed, B. A., Mir, M. M., Alfaifi, J., Mustafa, S. K., Barnawi, J., & Alrdahe, S. S. (2025). Whole-Exome Sequencing Reveals Rare Genetic Variants in Saudi COVID-19 Patients with Extreme Phenotypes. Viruses, 17(9), 1198. https://doi.org/10.3390/v17091198







