Genetic Diversity and Molecular Analysis of Human Parainfluenza Virus Type 3 in Saint Petersburg (Russia) in 2017–2023: Emergence of a New Phylogenetic Cluster
Abstract
1. Introduction
2. Materials and Methods
2.1. Sample Collection
2.2. RNA Extraction
2.3. RT-PCR Detection of Respiratory Viruses
2.4. Viruses and Cells
2.5. Virus Isolation and Propagation
2.6. RT-PCR Amplification of HN Gene
2.7. DNA Sequencing and Phylogenetic Analysis
2.8. GenBank Accession Numbers
3. Results
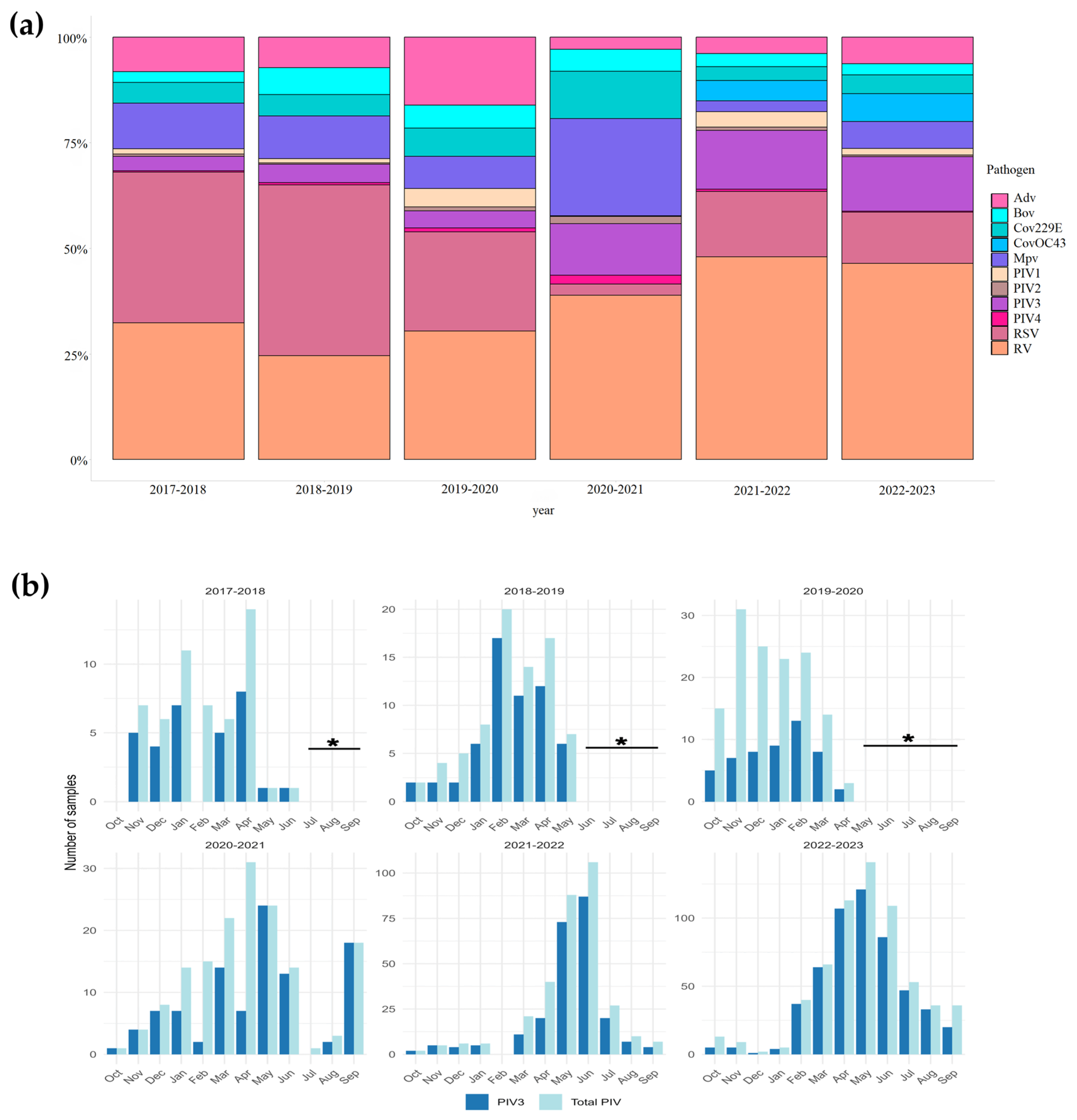
3.1. Detection of hPIV in Clinical Samples
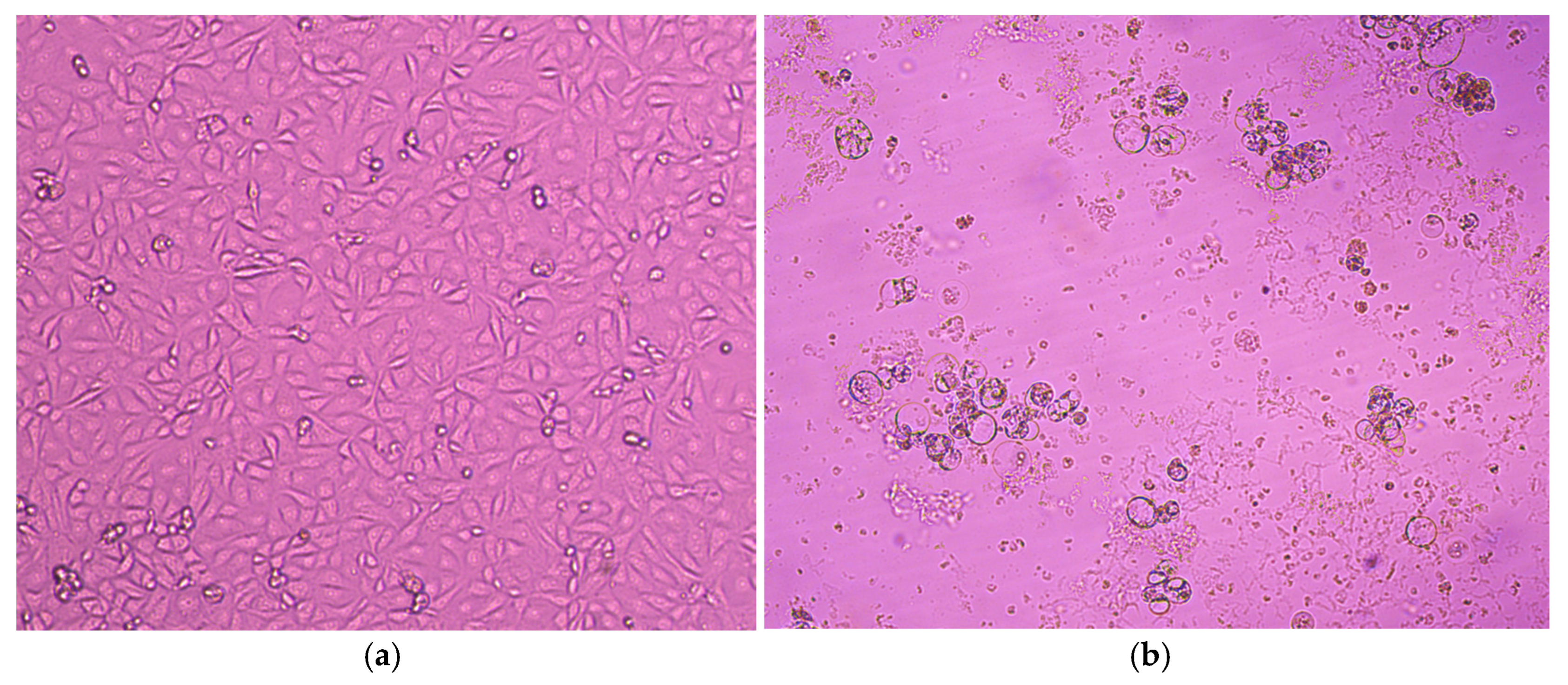
3.2. Virus Isolation on MA-104 and LLC-MK2 Cell Lines
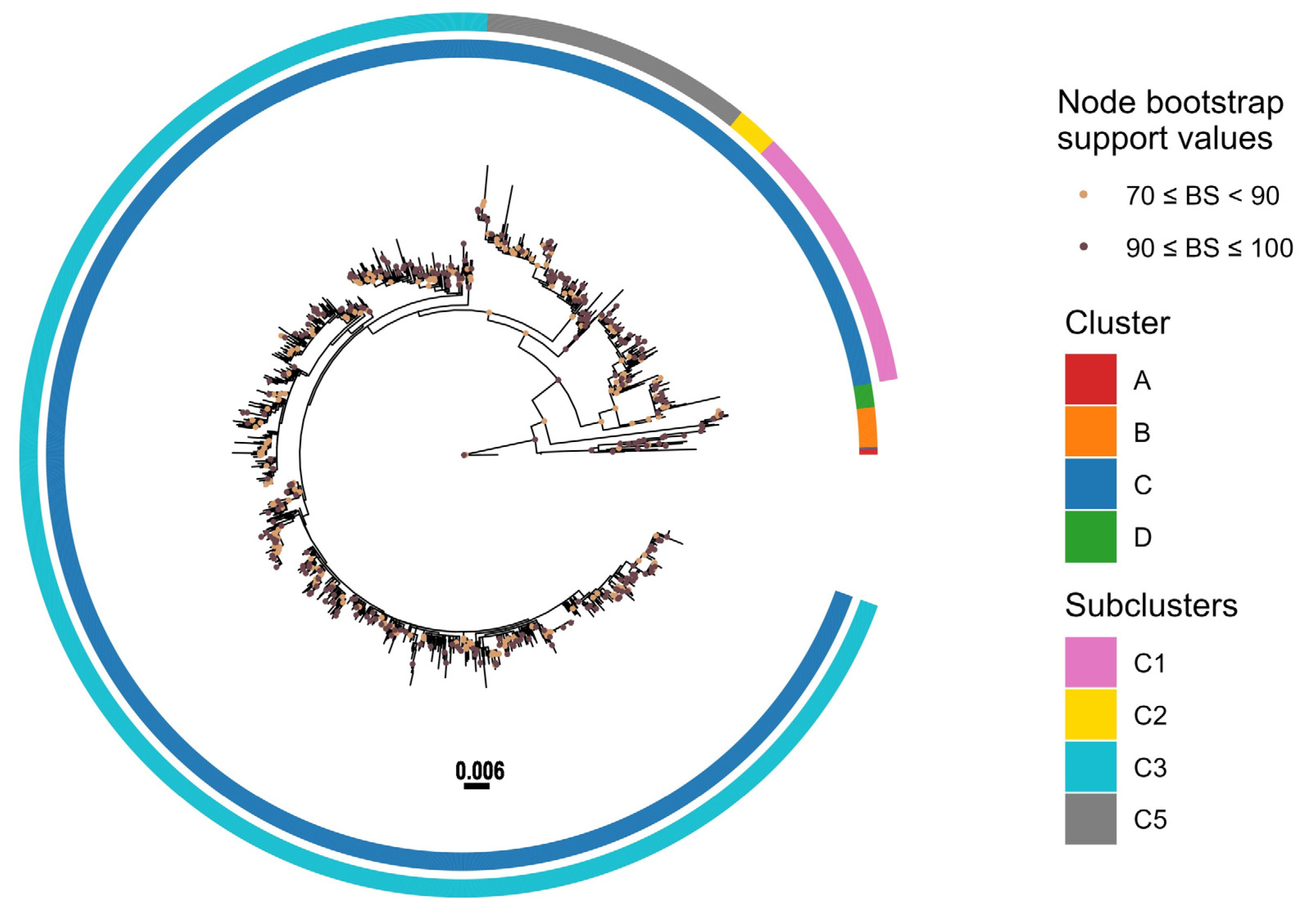
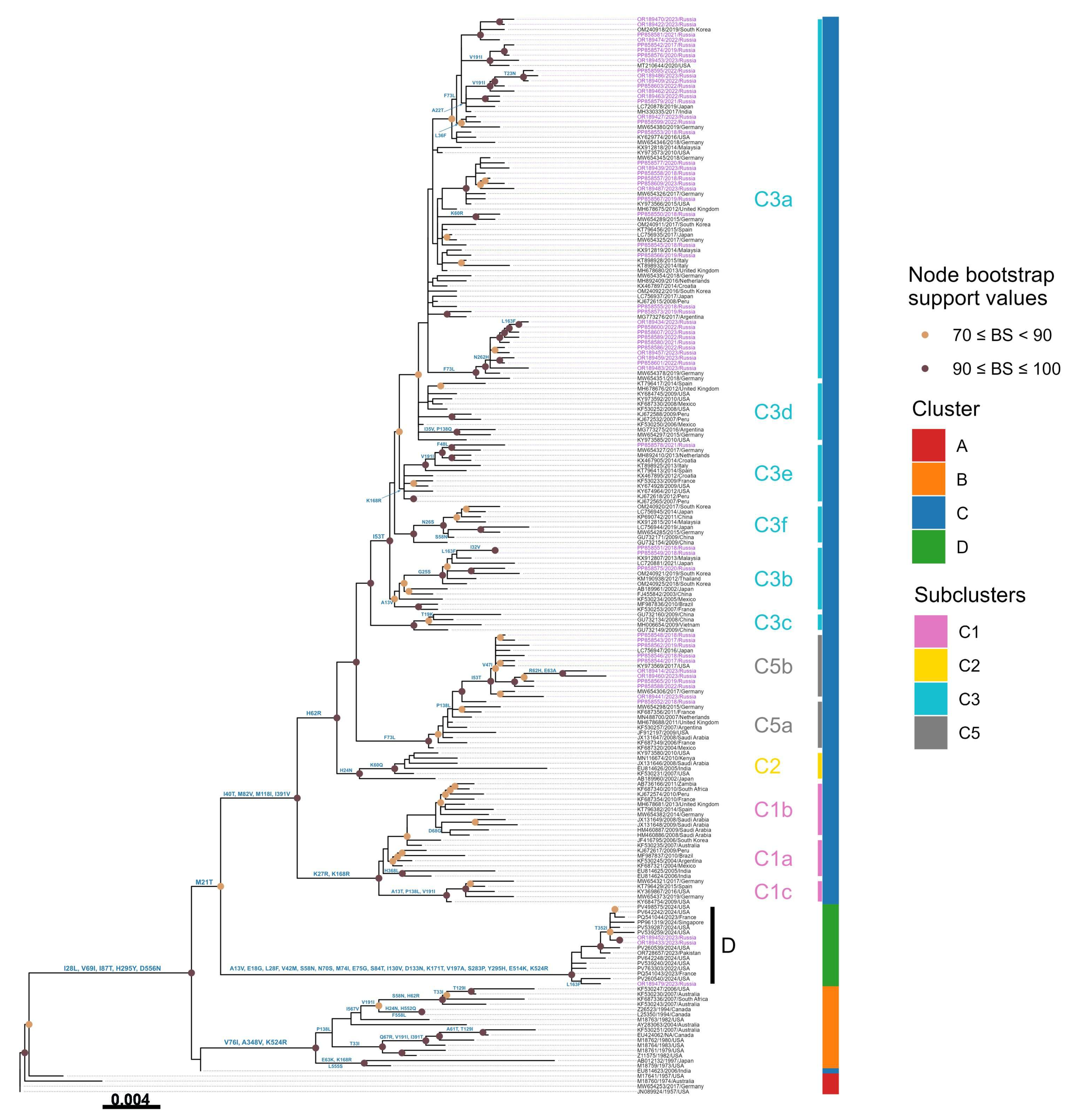
3.3. Phylogenetic Analysis
3.4. Amino Acid Variations
3.5. Potential Glycosylation Sites Analysis
4. Discussion
5. Conclusions
6. Limitations
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Karron, R.A.; Collins, P.L. Parainfluenza Viruses. In Fields Virology, 6th ed.; Knipe, D.M., Howley, P., Eds.; Lippincott Williams & Wilkins (LWW): Philadelphia, PA, USA, 2013; Volume 1, pp. 669–1023. [Google Scholar]
- Schomacker, H.; Schaap-Nutt, A.; Collins, P.L.; Schmidt, A.C. Pathogenesis of acute respiratory illness caused by human parainfluenza viruses. Curr. Opin. Virol. 2012, 2, 294–299. [Google Scholar] [CrossRef] [PubMed]
- Henrickson, K. Parainfluenza Viruses. Clin. Microbiol. Rev. 2003, 16, 242–264. [Google Scholar] [CrossRef]
- Glezen, W.P.; Frank, A.L.; Taber, L.H.; Kasel, J.A. Parainfluenza Virus Type 3: Seasonality and Risk of Infection and Reinfection in Young Children. J. Infect. Dis. 1984, 150, 851–857. [Google Scholar] [CrossRef]
- Rima, B.; Balkema-Buschmann, A.; Dundon, W.G.; Duprex, P.; Easton, A.; Fouchier, R.; Kurath, G.; Lamb, R.; Lee, B.; Rota, P.; et al. ICTV Virus Taxonomy Profile: Paramyxoviridae. J. Gen. Virol. 2019, 100, 1593–1594. [Google Scholar] [CrossRef]
- Weinberg, G.A.; Hall, C.B.; Iwane, M.K.; Poehling, K.A.; Edwards, K.M.; Griffin, M.R.; Staat, M.A.; Curns, A.T.; Erdman, D.D.; Szilagyi, P.G. Parainfluenza Virus Infection of Young Children: Estimates of the Population-Based Burden of Hospitalization. J. Pediatr. 2009, 154, 694–699.e1. [Google Scholar] [CrossRef]
- Abedi, G.R.; Prill, M.M.; Langley, G.E.; Wikswo, M.E.; Weinberg, G.A.; Curns, A.T.; Schneider, E. Estimates of Parainfluenza Virus-Associated Hospitalizations and Cost Among Children Aged Less Than 5 Years in the United States, 1998–2010. J. Pediatr. Infect. Dis. Soc. 2014, 5, 7–13. [Google Scholar] [CrossRef] [PubMed]
- Maykowski, P.; Smithgall, M.; Zachariah, P.; Oberhardt, M.; Vargas, C.; Reed, C.; Demmer, R.T.; Stockwell, M.S.; Saiman, L. Seasonality and clinical impact of human parainfluenza viruses. Influ. Other Respir. Viruses 2018, 12, 706–716. [Google Scholar] [CrossRef] [PubMed]
- Reed, G.; Jewett, P.H.; Thompson, J.; Tollefson, S.; Wright, P.F. Epidemiology and clinical impact of parainfluenza virus infections in otherwise healthy infants and young children <5 years old. J. Infect. Dis. 1997, 175, 807–813. [Google Scholar] [CrossRef]
- Counihan, M.E.; Shay, D.K.; Holman, R.C.; Lowther, S.A.; Anderson, L.J. Human parainfluenza virus-associated hospitalizations among children less than five years of age in the United States. Pediatr. Infect. Dis. J. 2001, 20, 646–653. [Google Scholar] [CrossRef]
- Iwane, M.K.; Edwards, K.M.; Szilagyi, P.G.; Walker, F.J.; Griffin, M.R.; Weinberg, G.A.; Coulen, C.; Poehling, K.A.; Shone, L.P.; Balter, S.; et al. Population-Based Surveillance for Hospitalizations Associated With Respiratory Syncytial Virus, Influenza Virus, and Parainfluenza Viruses Among Young Children. Pediatrics 2004, 113, 1758–1764. [Google Scholar] [CrossRef]
- Morgan, O.W.; Chittaganpitch, M.; Clague, B.; Chantra, S.; Sanasuttipun, W.; Prapasiri, P.; Naorat, S.; Laosirithavorn, Y.; Peret, T.C.T.; Erdman, D.D.; et al. Hospitalization due to human parainfluenza virus-associated lower respiratory tract illness in rural Thailand. Influenza Other Respir. Viruses 2012, 7, 280–285. [Google Scholar] [CrossRef][Green Version]
- Goya, S.; Mistchenko, A.S.; Viegas, M. Phylogenetic and molecular analyses of human parainfluenza type 3 virus in Buenos Aires, Argentina, between 2009 and 2013: The emergence of new genetic lineages. Infect. Genet. Evol. 2016, 39, 85–91. [Google Scholar] [CrossRef]
- Lim, F.J.; Blyth, C.C.; Fathima, P.; de Klerk, N.; Moore, H.C. Record linkage study of the pathogen-specific burden of respiratory viruses in children. Influenza Other Respir. Viruses 2017, 11, 502–510. [Google Scholar] [CrossRef]
- Ramezannia, Z.; Sadeghi, J.; Oskouie, S.A.; Rezaee, M.A.; Baghi, H.B.; Azadi, A.; Oskouee, M.A.; Bautista, M.C. Evaluation of Human Respiratory Syncytial Virus and Human Parainfluenza Virus Type 3 among Hospitalized Children in Northwest of Iran. Can. J. Infect. Dis. Med. Microbiol. 2021, 2021, 2270307. [Google Scholar] [CrossRef] [PubMed]
- Zhao, H.; Harris, R.J.; Ellis, J.; Donati, M.; Pebody, R.G. Epidemiology of parainfluenza infection in England and Wales, 1998–2013: Any evidence of change? Epidemiol. Infect. 2017, 145, 1210–1220. [Google Scholar] [CrossRef]
- DeGroote, N.P.; Haynes, A.K.; Taylor, C.; Killerby, M.E.; Dahl, R.M.; Mustaquim, D.; Gerber, S.I.; Watson, J.T. Human parainfluenza virus circulation, United States, 2011–2019. J. Clin. Virol. 2020, 124, 104261. [Google Scholar] [CrossRef] [PubMed]
- Mao, N.; Ji, Y.; Xie, Z.; Wang, H.; Wang, H.; An, J.; Zhang, X.; Zhang, Y.; Zhu, Z.; Cui, A.; et al. Human Parainfluenza Virus-Associated Respiratory Tract Infection among Children and Genetic Analysis of HPIV-3 Strains in Beijing, China. PLoS ONE 2012, 7, e43893. [Google Scholar] [CrossRef]
- Durbin, A.P.; McAuliffe, J.M.; Collins, P.L.; Murphy, B.R. Mutations in the C, D, and V Open Reading Frames of Human Parainfluenza Virus Type 3 Attenuate Replication in Rodents and Primates. Virology 1999, 261, 319–330. [Google Scholar] [CrossRef]
- Moscona, A. Entry of parainfluenza virus into cells as a target for interrupting childhood respiratory disease. J. Clin. Investig. 2005, 115, 1688–1698. [Google Scholar] [CrossRef]
- Vainionpää, R.; Hyypiä, T. Biology of parainfluenza viruses. Clin. Microbiol. Rev. 1994, 7, 265–275. [Google Scholar] [CrossRef] [PubMed]
- Huberman, K.; Peluso, R.W.; Moscona, A. Hemagglutinin–Neuraminidase of Human Parainfluenza 3: Role of the Neuraminidase in the Viral Life Cycle. Virology 1995, 214, 294–300. [Google Scholar] [CrossRef] [PubMed]
- Suzuki, T.; Portner, A.; Scroggs, R.A.; Uchikawa, M.; Koyama, N.; Matsuo, K.; Suzuki, Y.; Takimoto, T. Receptor Specificities of Human Respiroviruses. J. Virol. 2001, 75, 4604–4613. [Google Scholar] [CrossRef]
- Lawrence, M.C.; Borg, N.A.; Streltsov, V.A.; Pilling, P.A.; Epa, V.; Varghese, J.N.; McKimm-Breschkin, J.L.; Colman, P.M. Structure of the Haemagglutinin-neuraminidase from Human Parainfluenza Virus Type III. J. Mol. Biol. 2004, 335, 1343–1357. [Google Scholar] [CrossRef]
- Porotto, M.; Murrell, M.; Greengard, O.; Doctor, L.; Moscona, A. Influence of the Human Parainfluenza Virus 3 Attachment Protein’s Neuraminidase Activity on Its Capacity To Activate the Fusion Protein. J. Virol. 2005, 79, 2383–2392. [Google Scholar] [CrossRef]
- Palermo, L.M.; Porotto, M.; Yokoyama, C.C.; Palmer, S.G.; Mungall, B.A.; Greengard, O.; Niewiesk, S.; Moscona, A. Human Parainfluenza Virus Infection of the Airway Epithelium: Viral Hemagglutinin-Neuraminidase Regulates Fusion Protein Activation and Modulates Infectivity. J. Virol. 2009, 83, 6900–6908. [Google Scholar] [CrossRef]
- Jiang, J.; Wen, H.; Chi, M.; Liu, Y.; Liu, J.; Cao, Z.; Zhao, L.; Song, Y.; Liu, N.; Chi, L.; et al. Functional analysis of amino acids at stalk/head interface of human parainfluenza virus type 3 hemagglutinin-neuraminidase protein in the membrane fusion process. Virus Genes 2018, 54, 333–342. [Google Scholar] [CrossRef]
- Coelingh, K.L.v.W.; Winter, C.C.; Jorgensen, E.D.; Murphy, B.R. Antigenic and structural properties of the hemagglutinin-neuraminidase glycoprotein of human parainfluenza virus type 3: Sequence analysis of variants selected with monoclonal antibodies which inhibit infectivity, hemagglutination, and neuraminidase activities. J. Virol. 1987, 61, 1473–1477. [Google Scholar] [CrossRef]
- Klippmark, E.; Rydbeck, R.; Shibuta, H.; Norrby, E. Antigenic variation of human and bovine parainfluenza virus type 3 strains. J. Gen. Virol. 1990, 71, 1577–1580. [Google Scholar] [CrossRef] [PubMed]
- Villaran, M.V.; García, J.; Gomez, J.; Arango, A.E.; Gonzales, M.; Chicaiza, W.; Alemán, W.; de Rivera, I.L.; Sanchez, F.; Aguayo, N.; et al. Human parainfluenza virus in patients with influenza-like illness from Central and South America during 2006–2010. Influenza Other Respir. Viruses 2013, 8, 217–227. [Google Scholar] [CrossRef] [PubMed]
- Almajhdi, F.N. Hemagglutinin-Neuraminidase Gene Sequence-Based Reclassification of Human Parainfluenza Virus 3 Variants. Intervirology 2015, 58, 35–40. [Google Scholar] [CrossRef]
- Puig-Barberà, J.; Tormos, A.; Trushakova, S.; Sominina, A.; Pisareva, M.; Ciblak, M.A.; Badur, S.; Yu, H.; Cowling, B.J.; Burtseva, E.; et al. The Global Influenza Hospital Surveillance Network (GIHSN): A new platform to describe the epidemiology of severe influenza. Influenza Other Respir. Viruses 2015, 9, 277–286. [Google Scholar] [CrossRef]
- World Health Organization. Global Epidemiological Surveillance Standards for Influenza; World Health Organization: Geneva, Switzerland, 2013; ISBN 9789241506601. Available online: https://www.who.int/publications/i/item/9789241506601 (accessed on 25 August 2025).
- Park, K.S.; Yang, M.H.; Lee, C.K.; Song, K. Genetic analysis of human parainfluenza viruses circulating in Korea, 2006. J. Med. Virol. 2014, 86, 1041–1047. [Google Scholar] [CrossRef]
- Katoh, K.; Standley, D.M. MAFFT Multiple Sequence Alignment Software Version 7: Improvements in Performance and Usability. Mol. Biol. Evol. 2013, 30, 772–780. [Google Scholar] [CrossRef] [PubMed]
- Tamura, K.; Stecher, G.; Kumar, S.; Battistuzzi, F.U. MEGA11: Molecular Evolutionary Genetics Analysis Version 11. Mol. Biol. Evol. 2021, 38, 3022–3027. [Google Scholar] [CrossRef] [PubMed]
- Stamatakis, A. RAxML version 8: A tool for phylogenetic analysis and post-analysis of large phylogenies. Bioinformatics 2014, 30, 1312–1313. [Google Scholar] [CrossRef] [PubMed]
- Menardo, F.; Loiseau, C.; Brites, D.; Coscolla, M.; Gygli, S.M.; Rutaihwa, L.K.; Trauner, A.; Beisel, C.; Borrell, S.; Gagneux, S. Treemmer: A tool to reduce large phylogenetic datasets with minimal loss of diversity. BMC Bioinform. 2018, 19, 164. [Google Scholar] [CrossRef]
- Godoy, C.; Peremiquel-Trillas, P.; Andrés, C.; Gimferrer, L.; Uriona, S.M.; Codina, M.G.; Armadans, L.; Martí, M.d.C.; Fuentes, F.; Esperalba, J.; et al. A molecular epidemiological study of human parainfluenza virus type 3 at a tertiary university hospital during 2013–2015 in Catalonia, Spain. Diagn. Microbiol. Infect. Dis. 2016, 86, 153–159. [Google Scholar] [CrossRef]
- Aso, J.; Kimura, H.; Ishii, H.; Saraya, T.; Kurai, D.; Nagasawa, K.; Matsushima, Y.; Ryo, A.; Takizawa, H. Molecular evolution of the hemagglutinin-neuraminidase (HN) gene in human respirovirus 3. Virus Res. 2020, 277, 197824. [Google Scholar] [CrossRef]
- Kim, H.N.; Yoon, S.-Y.; Lim, C.S.; Lee, C.K.; Yoon, J. Phylogenetic analysis of human parainfluenza type 3 virus strains responsible for the outbreak during the COVID-19 pandemic in Seoul, South Korea. J. Clin. Virol. 2022, 153, 105213. [Google Scholar] [CrossRef]
- Pan, Y.; Zhang, Y.; Shi, W.; Peng, X.; Cui, S.; Zhang, D.; Lu, G.; Liu, Y.; Wu, S.; Yang, P.; et al. Human parainfluenza virus infection in severe acute respiratory infection cases in Beijing, 2014–2016: A molecular epidemiological study. Influenza Other Respir. Viruses 2017, 11, 564–568. [Google Scholar] [CrossRef]
- Košutić-Gulija, T.; Slovic, A.; Ljubin-Sternak, S.; Mlinarić-Galinović, G.; Forčić, D. Genetic analysis of human parainfluenza virus type 3 obtained in Croatia, 2011–2015. J. Med. Microbiol. 2017, 66, 502–510. [Google Scholar] [CrossRef]
- Takahashi, M.; Nagasawa, K.; Saito, K.; Maisawa, S.-I.; Fujita, K.; Murakami, K.; Kuroda, M.; Ryo, A.; Kimura, H. Detailed genetic analyses of the HN gene in human respirovirus 3 detected in children with acute respiratory illness in the Iwate Prefecture, Japan. Infect. Genet. Evol. 2018, 59, 155–162. [Google Scholar] [CrossRef] [PubMed]
- Elusah, J.; Bulimo, W.D.; Opanda, S.M.; Symekher, S.L.; Wamunyokoli, F.; Kumar, B. Genetic diversity and evolutionary analysis of human respirovirus type 3 strains isolated in Kenya using complete hemagglutinin-neuraminidase (HN) gene. PLoS ONE 2020, 15, e0229355. [Google Scholar] [CrossRef] [PubMed]
- Porotto, M.; Fornabaio, M.; Kellogg, G.E.; Moscona, A. A Second Receptor Binding Site on Human Parainfluenza Virus Type 3 Hemagglutinin-Neuraminidase Contributes to Activation of the FusionMechanism. J. Virol. 2007, 81, 3216–3228. [Google Scholar] [CrossRef]
- Palermo, L.M.; Porotto, M.; Greengard, O.; Moscona, A. Fusion Promotion by a Paramyxovirus Hemagglutinin-Neuraminidase Protein: pH Modulation of Receptor Avidity of Binding Sites I and II. J. Virol. 2007, 81, 9152–9161. [Google Scholar] [CrossRef] [PubMed]
- Mizuta, K.; Tsukagoshi, H.; Ikeda, T.; Aoki, Y.; Abiko, C.; Itagaki, T.; Nagano, M.; Noda, M.; Kimura, H. Molecular evolution of the haemagglutinin–neuraminidase gene in human parainfluenza virus type 3 isolates from children with acute respiratory illness in Yamagata prefecture, Japan. J. Med. Microbiol. 2014, 63, 570–577. [Google Scholar] [CrossRef]
- Tappert, M.M.; Smith, D.F.; Air, G.M. Fixation of Oligosaccharides to a Surface May Increase the Susceptibility to Human Parainfluenza Virus 1, 2, or 3 Hemagglutinin-Neuraminidase. J. Virol. 2011, 85, 12146–12159. [Google Scholar] [CrossRef]
- Xu, R.; Palmer, S.G.; Porotto, M.; Palermo, L.M.; Niewiesk, S.; Wilson, I.A.; Moscona, A.; Griffin, D.E. Interaction between the Hemagglutinin-Neuraminidase and Fusion Glycoproteins of Human Parainfluenza Virus Type III Regulates Viral Growth In Vivo. mBio 2013, 4, e00803-13. [Google Scholar] [CrossRef]
- Moscona, A.; Peluso, R.W. Relative affinity of the human parainfluenza virus type 3 hemagglutinin-neuraminidase for sialic acid correlates with virus-induced fusion activity. J. Virol. 1993, 67, 6463–6468. [Google Scholar] [CrossRef]
- Chu, F.-L.; Wen, H.-L.; Hou, G.-H.; Lin, B.; Zhang, W.-Q.; Song, Y.-Y.; Ren, G.-J.; Sun, C.-X.; Li, Z.-M.; Wang, Z. Role of N-linked glycosylation of the human parainfluenza virus type 3 hemagglutinin-neuraminidase protein. Virus Res. 2013, 174, 137–147. [Google Scholar] [CrossRef]
- Chen, J.; Deng, S.; Xu, X.; Chen, S.; Abo, Y.-N.; Bassat, Q.; Deng, J.; Komissarov, A.B.; Liu, E.; Muñoz-Almagro, C.; et al. Regional and type-specific variations in the global seasonality of human parainfluenza viruses and the influence of climatic factors: A systematic review and meta-analysis. Lancet Glob. Health 2025, 13, e1425–e1435. [Google Scholar] [CrossRef] [PubMed]

| Cluster/Subcluster | 2017–2018 | 2018–2019 | 2019–2020 | 2020–2021 | 2021–2022 | 2022–2023 | Total * |
|---|---|---|---|---|---|---|---|
| D | 3 | 3 | |||||
| C5a | 1 | 1 | |||||
| C5b | 4 | 2 | 1 | 3 | 10 | ||
| C3b | 2 | 1 | 3 | ||||
| C3e | 1 | 1 | |||||
| C3a | 6 | 3 | 4 | 3 | 7 | 17 | 40 |
| Total | 13 | 5 | 5 | 4 | 8 | 23 | 58 |
| A | B | C | D | |
|---|---|---|---|---|
| A | ||||
| B | 0.050 | |||
| C | 0.048 | 0.051 | ||
| D | 0.062 | 0.062 | 0.064 |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2025 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Mansour, O.; Fadeev, A.V.; Perederiy, A.A.; Zadirienko, M.I.; Danilenko, D.M.; Lioznov, D.A.; Komissarov, A.B. Genetic Diversity and Molecular Analysis of Human Parainfluenza Virus Type 3 in Saint Petersburg (Russia) in 2017–2023: Emergence of a New Phylogenetic Cluster. Viruses 2025, 17, 1197. https://doi.org/10.3390/v17091197
Mansour O, Fadeev AV, Perederiy AA, Zadirienko MI, Danilenko DM, Lioznov DA, Komissarov AB. Genetic Diversity and Molecular Analysis of Human Parainfluenza Virus Type 3 in Saint Petersburg (Russia) in 2017–2023: Emergence of a New Phylogenetic Cluster. Viruses. 2025; 17(9):1197. https://doi.org/10.3390/v17091197
Chicago/Turabian StyleMansour, Oula, Artem V. Fadeev, Alexander A. Perederiy, Marina I. Zadirienko, Daria M. Danilenko, Dmitry A. Lioznov, and Andrey B. Komissarov. 2025. "Genetic Diversity and Molecular Analysis of Human Parainfluenza Virus Type 3 in Saint Petersburg (Russia) in 2017–2023: Emergence of a New Phylogenetic Cluster" Viruses 17, no. 9: 1197. https://doi.org/10.3390/v17091197
APA StyleMansour, O., Fadeev, A. V., Perederiy, A. A., Zadirienko, M. I., Danilenko, D. M., Lioznov, D. A., & Komissarov, A. B. (2025). Genetic Diversity and Molecular Analysis of Human Parainfluenza Virus Type 3 in Saint Petersburg (Russia) in 2017–2023: Emergence of a New Phylogenetic Cluster. Viruses, 17(9), 1197. https://doi.org/10.3390/v17091197







