A Novel Cogu-like Virus Identified in Wine Grapes
Abstract
1. Introduction
2. Materials and Methods
2.1. Grapevine Sampling and Sample Processing
2.2. RNA Extraction and HTS Analysis
2.3. Nucleic Acid Extraction, RT-PCR Testing, and Sanger Sequencing
2.4. Sequence and Phylogenetic Analysis
3. Results
3.1. Cogu-like Virus Sequences Revealed by HTS in Grapevine Leaf and Petiole Tissue
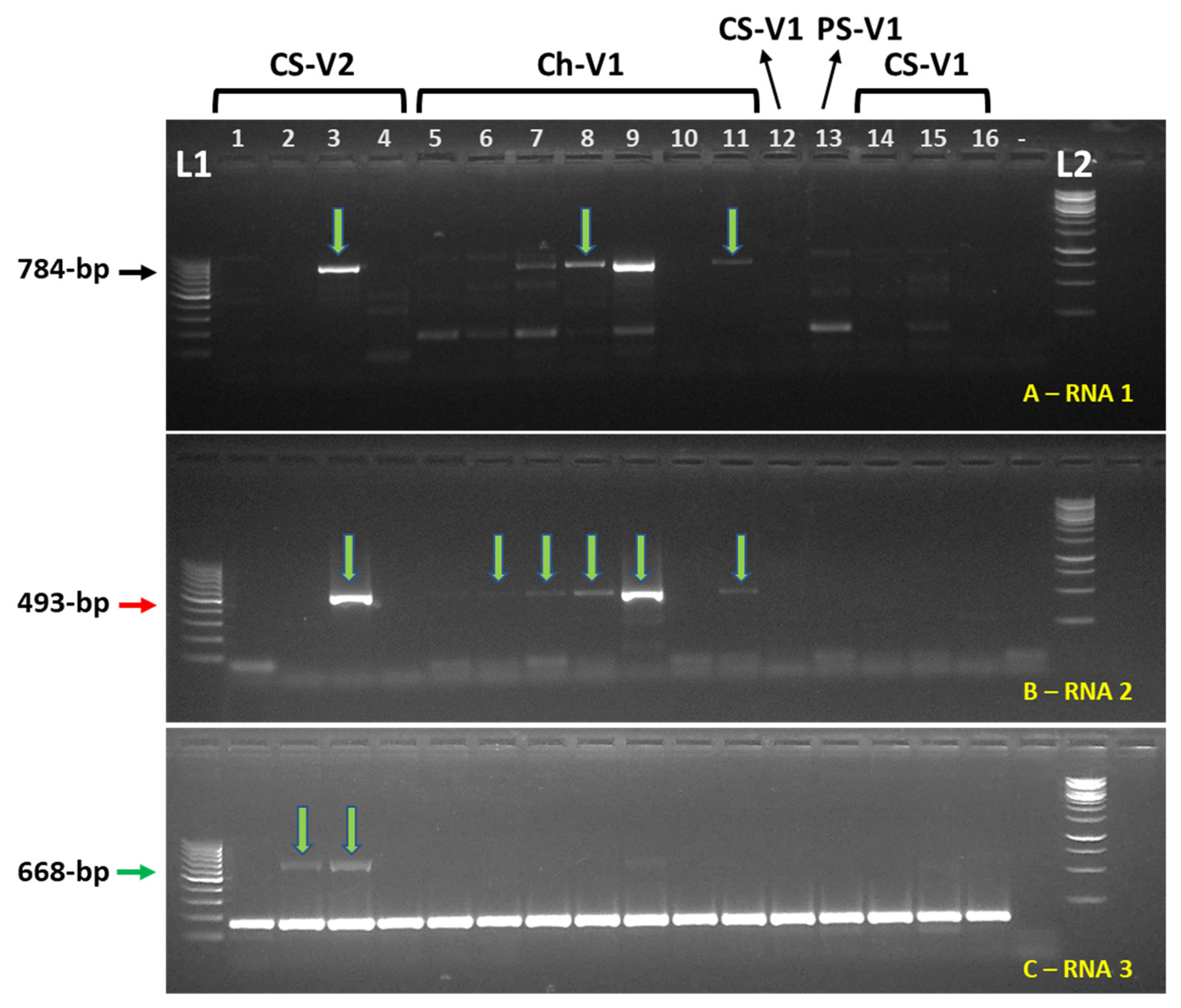
3.2. Prevalence of GaCLIdV in Wine Grapes in Southwestern Idaho and in Eastern Oregon
3.3. Phylogeny of the New Grapevine Cogu-like Virus
4. Discussion
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Martelli, G.P. An Overview on Grapevine Viruses, Viroids, and the Diseases They Cause. In Grapevine Viruses: Molecular Biology, Diagnostics and Management; Meng, B., Martelli, G.P., Golino, D.A., Fuchs, M., Eds.; Springer International Publishing: Cham, Switzerland, 2017; pp. 31–46. ISBN 978-3-319-57706-7. [Google Scholar]
- Fuchs, M. Grapevine Viruses: Did You Say More than a Hundred? J. Plant Pathol. 2025, 107, 217–227. [Google Scholar] [CrossRef]
- Diaz-Lara, A.; Navarro, B.; Di Serio, F.; Stevens, K.; Hwang, M.S.; Kohl, J.; Vu, S.T.; Falk, B.W.; Golino, D.; Al Rwahnih, M. Two Novel Negative-Sense RNA Viruses Infecting Grapevine Are Members of a Newly Proposed Genus within the Family Phenuiviridae. Viruses 2019, 11, 685. [Google Scholar] [CrossRef] [PubMed]
- Nabeshima, T.; Abe, J. High-Throughput Sequencing Indicates Novel Varicosavirus, Emaravirus, and Deltapartitivirus Infections in Vitis coignetiae. Viruses 2021, 13, 827. [Google Scholar] [CrossRef] [PubMed]
- Bertazzon, N.; Chitarra, W.; Angelini, E.; Nerva, L. Two New Putative Plant Viruses from Wood Metagenomics Analysis of an Esca Diseased Vineyard. Plants 2020, 9, 835. [Google Scholar] [CrossRef]
- Chiapello, M.; Rodríguez-Romero, J.; Nerva, L.; Forgia, M.; Chitarra, W.; Ayllón, M.A.; Turina, M. Putative New Plant Viruses Associated with Plasmopara viticola-Infected Grapevine Samples. Ann. Appl. Biol. 2020, 176, 180–191. [Google Scholar] [CrossRef]
- Sasaya, T.; Palacios, G.; Briese, T.; Di Serio, F.; Groschup, M.H.; Neriya, Y.; Song, J.-W.; Tomitaka, Y. ICTV Virus Taxonomy Profile: Phenuiviridae 2023. J. Gen. Virol. 2023, 104, 001893. [Google Scholar] [CrossRef]
- Falk, B.W.; Tsai, J.H. Biology and Molecular Biology of Viruses in the Genus Tenuivirus. Annu. Rev. Phytopathol. 1998, 36, 139–163. [Google Scholar] [CrossRef]
- Mekuria, T.A.; Karasev, A.V.; Martin, R.R.; Naidu, R.A. First Report of Grapevine Leafroll-Associated Virus-3 in Six Wine Grape Cultivars in Idaho. Plant Dis. 2009, 93, 1218. [Google Scholar] [CrossRef]
- Thompson, B.D.; Dahan, J.; Lee, J.; Martin, R.R.; Karasev, A.V. A Novel Genetic Variant of Grapevine Leafroll-Associated Virus-3 (GLRaV-3) from Idaho Grapevines. Plant Dis. 2019, 103, 509–518. [Google Scholar] [CrossRef]
- Kanuya, E.; Clayton, L.A.; Naidu, R.A.; Karasev, A.V. First Report of Grapevine Fleck Virus in Idaho Grapevines. Plant Dis. 2012, 96, 1705. [Google Scholar] [CrossRef]
- Thompson, B.D.; Eid, S.; Vander Pol, D.; Lee, J.; Karasev, A.V. First Report of Grapevine Red Blotch Virus in Idaho Grapevines. Plant Dis. 2019, 103, 2704. [Google Scholar] [CrossRef]
- Dahan, J.; Thompson, B.D.; Lee, J.; Karasev, A.V. First Report of Grapevine Rupestris Vein Feathering Virus in Wine Grapes in Idaho. Plant Dis. 2021, 105, 3309. [Google Scholar] [CrossRef]
- Dahan, J.; Orellana, G.E.; Lee, J.; Karasev, A.V. Occurrence of Grapevine-Associated Tymo-like Virus in Wine Grapes in the United States. Plant Dis. 2023, 107, 592. [Google Scholar] [CrossRef]
- Dahan, J.; Orellana, G.E.; Lee, J.; Karasev, A.V. Genome Sequences of Two Grapevine Rupestris Stem Pitting-Associated Virus Variants from Vitis vinifera Cv. Riesling in Idaho, USA. Microbiol. Resour. Announc. 2023, 12, e01366-22. [Google Scholar] [CrossRef]
- Dahan, J.; Orellana, G.E.; Lee, J.; Karasev, A.V. Grapevine Endophyte Endornavirus and Two New Endornaviruses Found Associated with Grapevines (Vitis vinifera L.) in Idaho, USA. Viruses 2023, 15, 1347. [Google Scholar] [CrossRef]
- Dahan, J.; Orellana, G.E.; Lee, J.; Karasev, A.V. Occurrence of Grapevine Yellow Speckle Viroid 2 and Australian Grapevine Viroid in Idaho Grapevines. Plant Dis. 2024, 108, 1121. [Google Scholar] [CrossRef]
- Lee, J.; Rennaker, C.D.; Thompson, B.D.; Karasev, A.V. Influence of Grapevine Red Blotch Virus (GRBV) on Idaho ‘Syrah’ Grape Composition. Sci. Hortic. 2021, 282, 110055. [Google Scholar] [CrossRef]
- Lee, J.; Rennaker, C.D.; Thompson, B.D.; Dahan, J.; Karasev, A.V. Idaho ‘Cabernet Sauvignon’ Grape Composition Altered by Grapevine Leafroll-Associated Virus 3. NFS J. 2023, 31, 1–6. [Google Scholar] [CrossRef]
- Bolger, A.M.; Lohse, M.; Usadel, B. Trimmomatic: A Flexible Trimmer for Illumina Sequence Data. Bioinformatics 2014, 30, 2114–2120. [Google Scholar] [CrossRef]
- Buchfink, B.; Reuter, K.; Drost, H.-G. Sensitive Protein Alignments at Tree-of-Life Scale Using DIAMOND. Nat. Methods 2021, 18, 366–368. [Google Scholar] [CrossRef] [PubMed]
- Marchler-Bauer, A.; Bo, Y.; Han, L.; He, J.; Lanczycki, C.J.; Lu, S.; Chitsaz, F.; Derbyshire, M.K.; Geer, R.C.; Gonzales, N.R.; et al. CDD/SPARCLE: Functional Classification of Proteins via Subfamily Domain Architectures. Nucleic Acids Res. 2017, 45, D200–D203. [Google Scholar] [CrossRef]
- Lu, S.; Wang, J.; Chitsaz, F.; Derbyshire, M.K.; Geer, R.C.; Gonzales, N.R.; Gwadz, M.; Hurwitz, D.I.; Marchler, G.H.; Song, J.S.; et al. CDD/SPARCLE: The Conserved Domain Database in 2020. Nucleic Acids Res. 2020, 48, D265–D268. [Google Scholar] [CrossRef]
- Green, K.J.; Brown, C.J.; Gray, S.M.; Karasev, A.V. Phylogenetic Study of Recombinant Strains of Potato Virus Y. Virology 2017, 507, 40–52. [Google Scholar] [CrossRef]
- Minh, B.Q.; Schmidt, H.A.; Chernomor, O.; Schrempf, D.; Woodhams, M.D.; von Haeseler, A.; Lanfear, R. IQ-TREE 2: New Models and Efficient Methods for Phylogenetic Inference in the Genomic Era. Mol. Biol. Evol. 2020, 37, 1530–1534. [Google Scholar] [CrossRef] [PubMed]
- Kalyaanamoorthy, S.; Minh, B.Q.; Wong, T.K.F.; von Haeseler, A.; Jermiin, L.S. ModelFinder: Fast Model Selection for Accurate Phylogenetic Estimates. Nat. Methods 2017, 14, 587–589. [Google Scholar] [CrossRef] [PubMed]
- Anisimova, M.; Gil, M.; Dufayard, J.-F.; Dessimoz, C.; Gascuel, O. Survey of Branch Support Methods Demonstrates Accuracy, Power, and Robustness of Fast Likelihood-Based Approximation Schemes. Syst. Biol. 2011, 60, 685–699. [Google Scholar] [CrossRef] [PubMed]
- Yu, G.; Smith, D.K.; Zhu, H.; Guan, Y.; Lam, T.T.-Y. Ggtree: An R Package for Visualization and Annotation of Phylogenetic Trees with Their Covariates and Other Associated Data. Methods Ecol. Evol. 2017, 8, 28–36. [Google Scholar] [CrossRef]
- Tokarz, R.; Sameroff, S.; Tagliafierro, T.; Jain, K.; Williams, S.H.; Cucura, D.M.; Rochlin, I.; Monzon, J.; Carpi, G.; Tufts, D.; et al. Identification of Novel Viruses in Amblyomma americanum, Dermacentor variabilis, and Ixodes scapularis Ticks. mSphere 2018, 3, e00614-17. [Google Scholar] [CrossRef]
- Fuchs, M. Grapevine Viruses: A Multitude of Diverse Species with Simple but Overall Poorly Adopted Management Solutions in the Vineyard. J. Plant Pathol. 2020, 102, 643–653. [Google Scholar] [CrossRef]
- Saldarelli, P.; Giampetruzzi, A.; Maree, H.J.; Al Rwahnih, M. High-Throughput Sequencing: Advantages Beyond Virus Identification. In Grapevine Viruses: Molecular Biology, Diagnostics and Management; Meng, B., Martelli, G.P., Golino, D.A., Fuchs, M., Eds.; Springer International Publishing: Cham, Switzerland, 2017; pp. 625–642. ISBN 978-3-319-57706-7. [Google Scholar]
- Gingery, R.E.; Nault, L.R.; Bradfute, O.E. Maize Stripe Virus: Characteristics of a Member of a New Virus Class. Virology 1981, 112, 99–108. [Google Scholar] [CrossRef] [PubMed]
- Sõmera, M.; Kvarnheden, A.; Desbiez, C.; Blystad, D.-R.; Sooväli, P.; Kundu, J.K.; Gantsovski, M.; Nygren, J.; Lecoq, H.; Verdin, E.; et al. Sixty Years after the First Description: Genome Sequence and Biological Characterization of European Wheat Striate Mosaic Virus Infecting Cereal Crops. Phytopathology 2020, 110, 68–79. [Google Scholar] [CrossRef] [PubMed]



| Sample ID | Read Pairs | Cleaned Read Pairs | Unmapped Read Pairs | Contigs > 1000 nt |
|---|---|---|---|---|
| 18RB02 | 20,840,871 | 19,368,940 | 6,340,098 | 22,692 |
| RB09 | 15,095,042 | 14,203,522 | 1,925,857 | 2295 |
| RB12 | 20,279,532 | 19,116,250 | 1,763,090 | 2327 |
| SgPhLV-1 | LLV | |||
|---|---|---|---|---|
| nt | aa | nt | aa | |
| RNA1/L(RdRP) | NS a | 40.6% | NS | 36.3% |
| RNA2/MP | NS | 49.7% | NS | 27.7% |
| RNA3/CP | NS | 40.1% | NS | 34.9% |
| Cultivar | Vineyard | RNA1_RdRP | RNA2_MP | RNA3_NP | ||||
|---|---|---|---|---|---|---|---|---|
| nt identity to RNA1, % | GenBank Acc. Number | nt identity to RNA2, % | GenBank Acc. Number | nt identity to RNA3, % | GenBank Acc. Number | |||
| 2020 | Chardonnay | V1 | 99.6% | PX114093 | 100.0% | PX124612 | 99.7% | PX123833 |
| Chardonnay | V1 | 99.8% | PX114094 | 88.7% | PX124613 | 100.0% | PX123834 | |
| 2021 | Chardonnay | V1 | NB a | - | NB | - | 98.0% | PX123835 |
| Chardonnay | V1 | 100.0% | PX114095 | bad seq | - | NB | - | |
| Chardonnay | V1 | 99.9% | PX114096 | 91.2% | PX124614 | bad seq | - | |
| 2023 | Cab. Franc | V3 | 99.4% | PX114097 | NB | - | NB | - |
| Cab. Sauv. | V2 | NB | - | NB | - | 99.1% | PX123836 | |
| Cab. Sauv. | V2 | 97.4% | PX114098 | 99.8% | PX124615 | 99.8% | PX123837 | |
| Chardonnay | V1 | bad seq | - | 80.7% | PX124616 | bad seq | - | |
| Chardonnay | V1 | bad seq | - | 92.5% | PX124617 | NB | - | |
| Chardonnay | V1 | 98.0% | PX114099 | 91.2% | PX124618 | NB | - | |
| Chardonnay | V1 | bad seq | - | 90.8% | PX124619 | 98.7% | PX123838 | |
| Chardonnay | V1 | 100.0% | PX114100 | 99.5% | PX124620 | NB | - | |
| Merlot | V4 | 96.6% | PX114101 | NB | - | NB | - | |
| Merlot | V4 | NB | - | 99.8% | PX124621 | NB | - | |
| Merlot | V4 | 99.9% | PX114102 | 99.8% | PX124622 | NB | - | |
| Merlot | V4 | 89.9% | PX114103 | 99.6% | PX123845 | NB | - | |
| Merlot | V4 | 100.0% | PX114104 | 91.7% | PX124623 | NB | - | |
| 2024 | Merlot | V3 | 97.5% | PX114105 | 99.8% | PX124624 | 99.8% | PX123839 |
| Riesling | V3 | NB | - | 99.8% | PX124625 | NB | - | |
| Cab. Franc | V5 | NB | - | 100.0% | PX124626 | 100.0% | PX123840 | |
| Tempranillo | V5 | 99.8% | PX114106 | 88.8% | PX124627 | 99.8% | PX123841 | |
| Tempranillo | V5 | NB | - | 100.0% | PX124628 | 100.0% | PX123842 | |
| Tempranillo | V5 | bad seq | - | 94.5% | PX124629 | bad seq | - | |
| Chardonnay | V1 | bad seq | - | 91.1% | PX124630 | 97.2% | PX123843 | |
| Chardonnay | V1 | NB | - | 91.3% | PX124631 | 100.0% | PX123844 | |
| Tempranillo | V6 | NB | - | 98.4% | PX123846 | NB | - | |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2025 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Dahan, J.; Orellana, G.E.; Reyes-Proaño, E.; Lee, J.; Karasev, A.V. A Novel Cogu-like Virus Identified in Wine Grapes. Viruses 2025, 17, 1175. https://doi.org/10.3390/v17091175
Dahan J, Orellana GE, Reyes-Proaño E, Lee J, Karasev AV. A Novel Cogu-like Virus Identified in Wine Grapes. Viruses. 2025; 17(9):1175. https://doi.org/10.3390/v17091175
Chicago/Turabian StyleDahan, Jennifer, Gardenia E. Orellana, Edison Reyes-Proaño, Jungmin Lee, and Alexander V. Karasev. 2025. "A Novel Cogu-like Virus Identified in Wine Grapes" Viruses 17, no. 9: 1175. https://doi.org/10.3390/v17091175
APA StyleDahan, J., Orellana, G. E., Reyes-Proaño, E., Lee, J., & Karasev, A. V. (2025). A Novel Cogu-like Virus Identified in Wine Grapes. Viruses, 17(9), 1175. https://doi.org/10.3390/v17091175






