Differentiating Cell Entry Potentials of SARS-CoV-2 Omicron Subvariants on Human Lung Epithelium Cells
Abstract
1. Introduction
2. Materials and Methods
2.1. Cell Lines Culture
2.2. Plasmids
2.3. Spike Pseudotyped Lentivirus Transduction through Gaussia Luciferase Reporter Assay
2.4. Single-Round Virus-to-Cell Entry
2.5. Spike Protein Processing in Cells
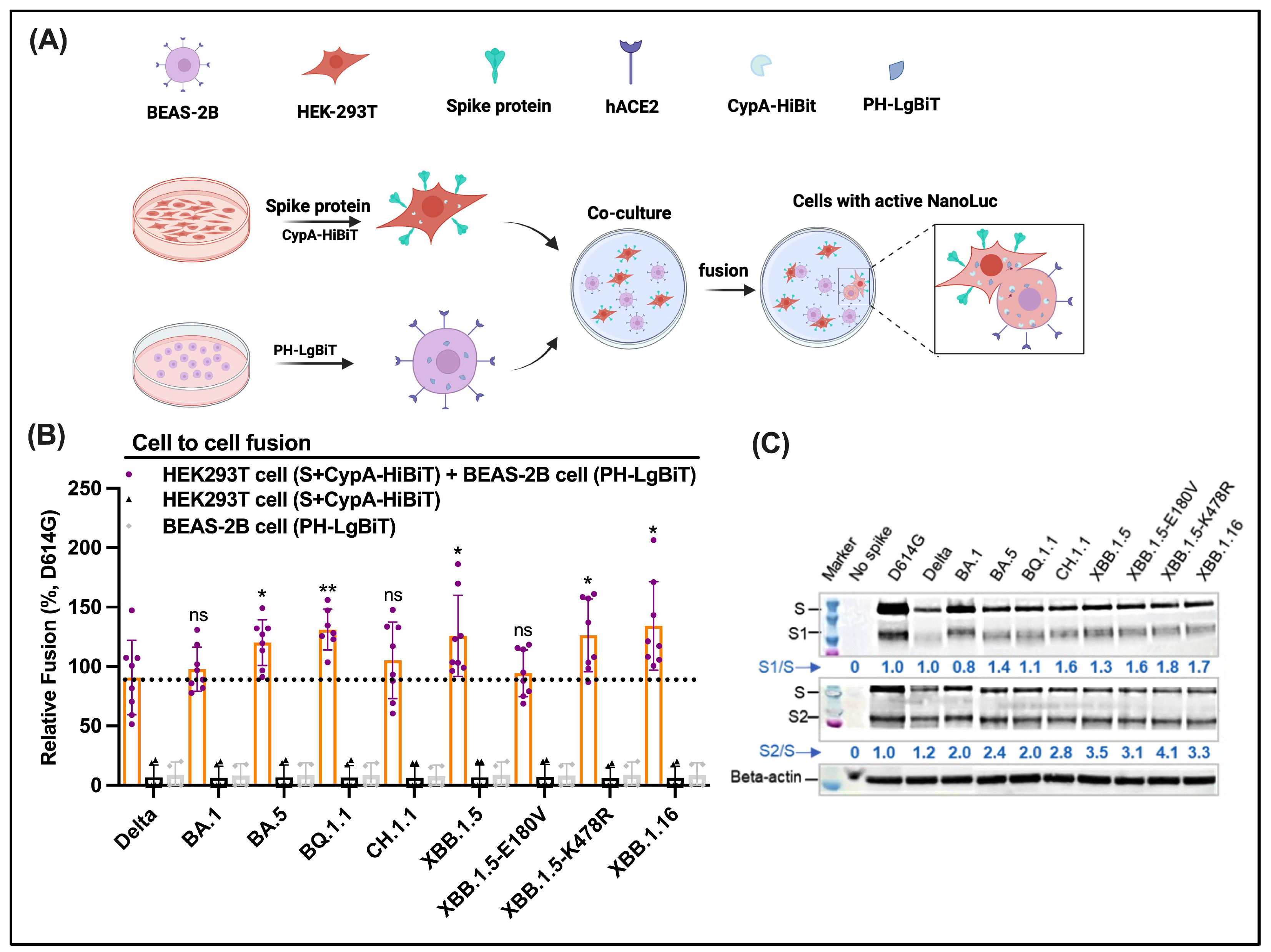
2.6. Cell-to-Cell Fusion
2.7. Statistics and Reproducibility
3. Results
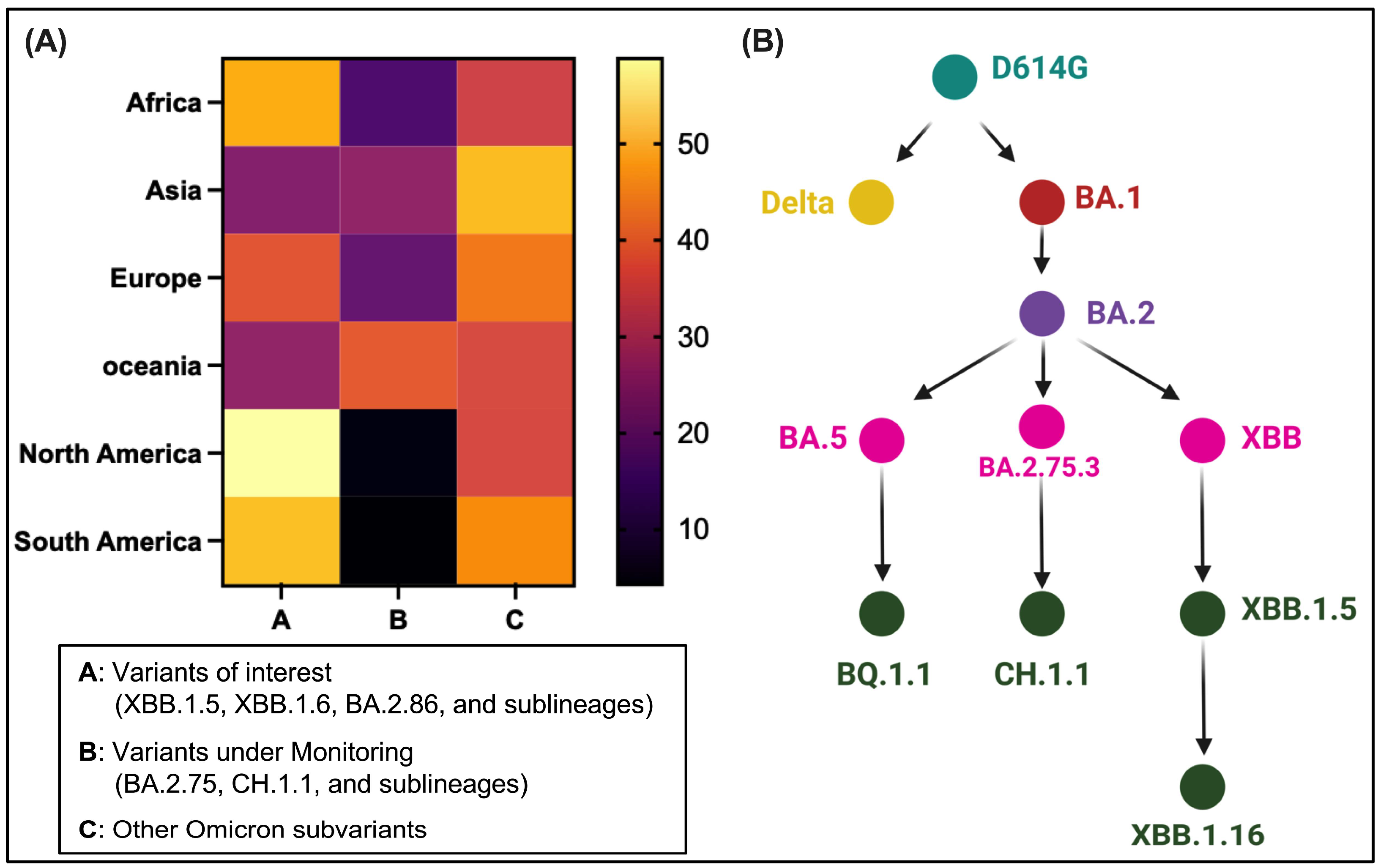
3.1. Dominance and Unique S Mutations of Omicron Descendants
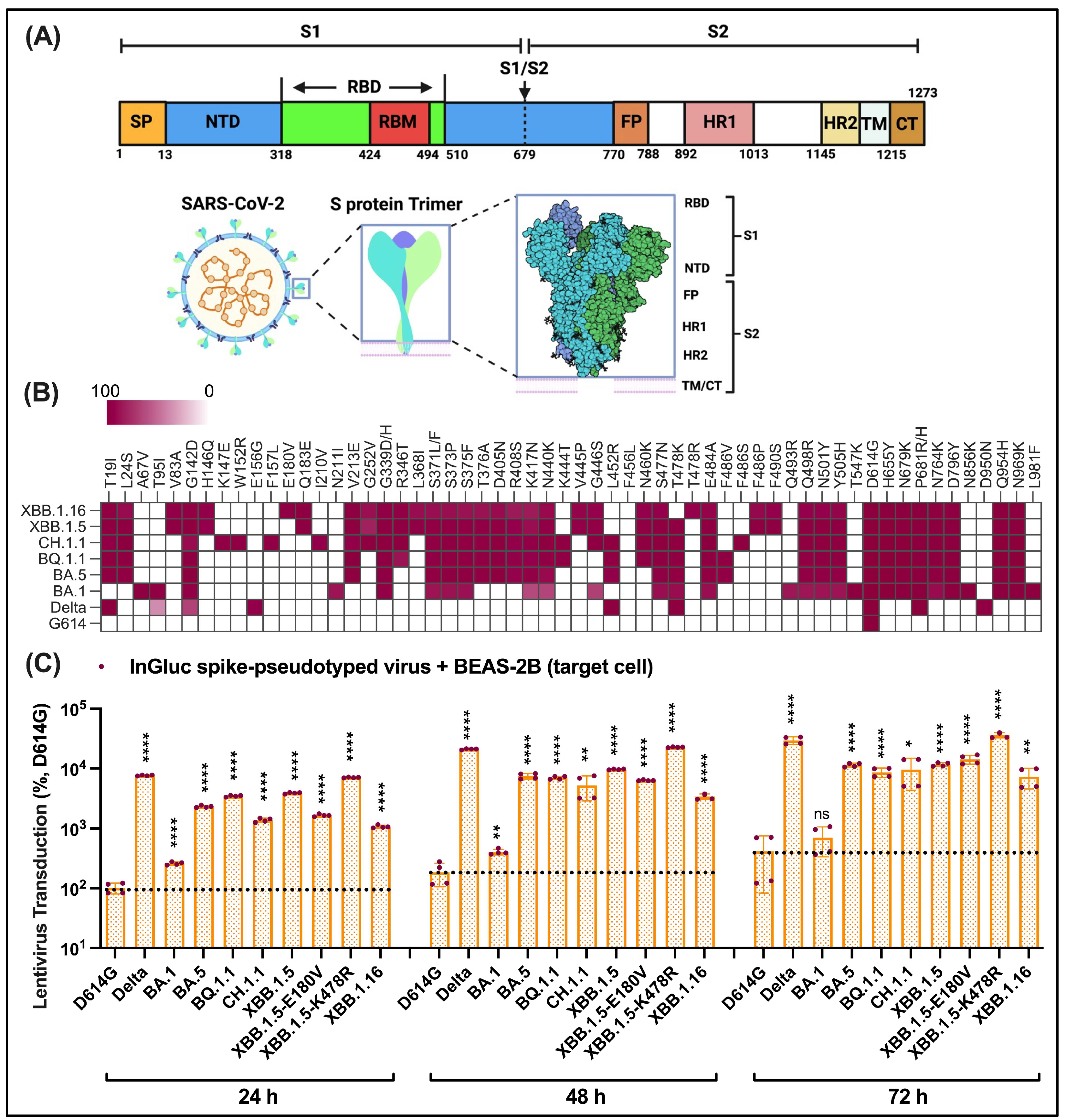
3.2. Elevated Pseudovirus Transduction of Omicron Subvariants Compared to Prototype D614G but Reduced Compared to Delta
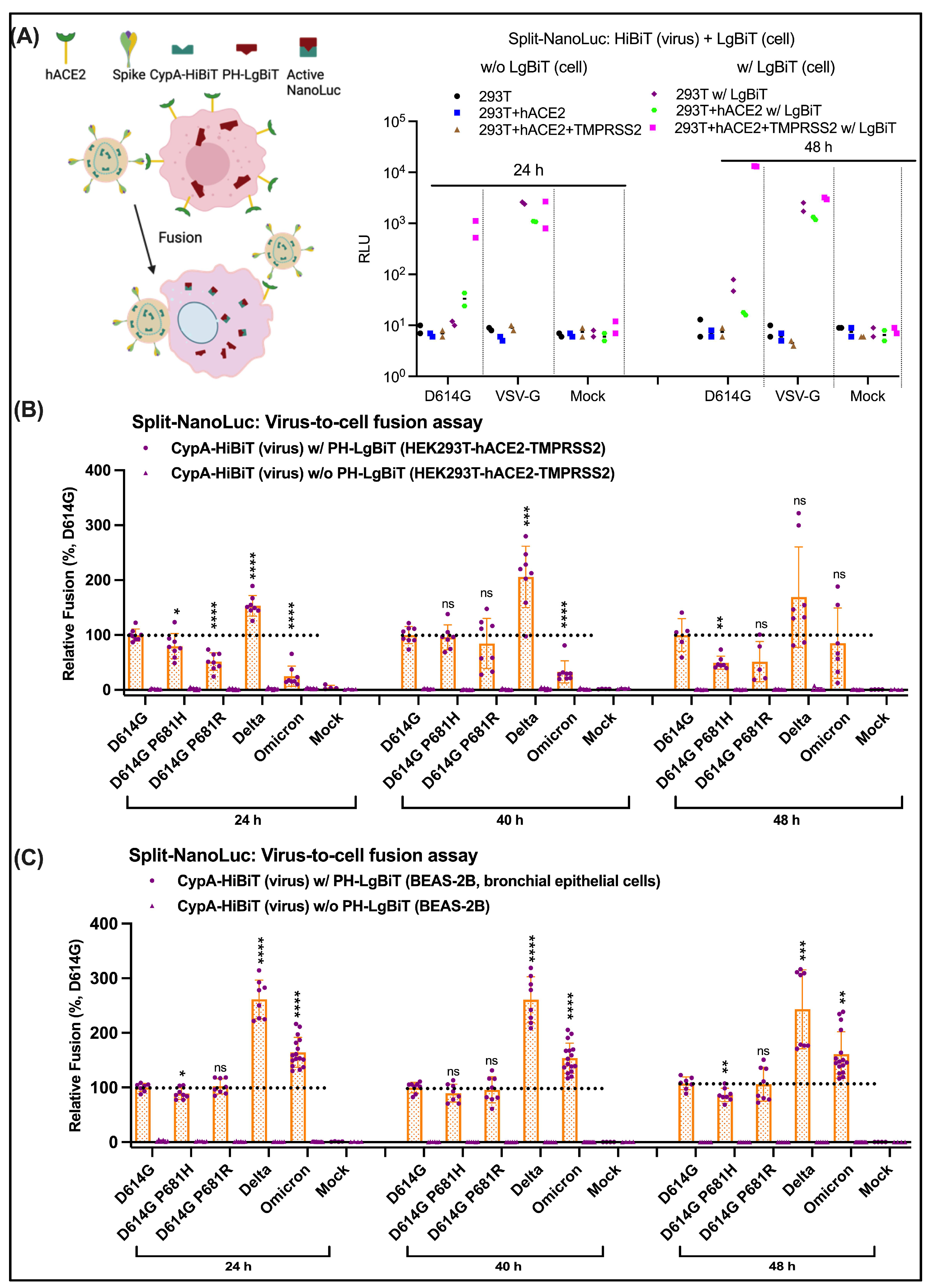
3.3. Suboptimal Use of HEK293T-hACE2 in Quantifying SARS-CoV-2 Cell Entry
3.4. Entry Affected by P681H/R at Cleavage Sites Is Cell-Type Dependent
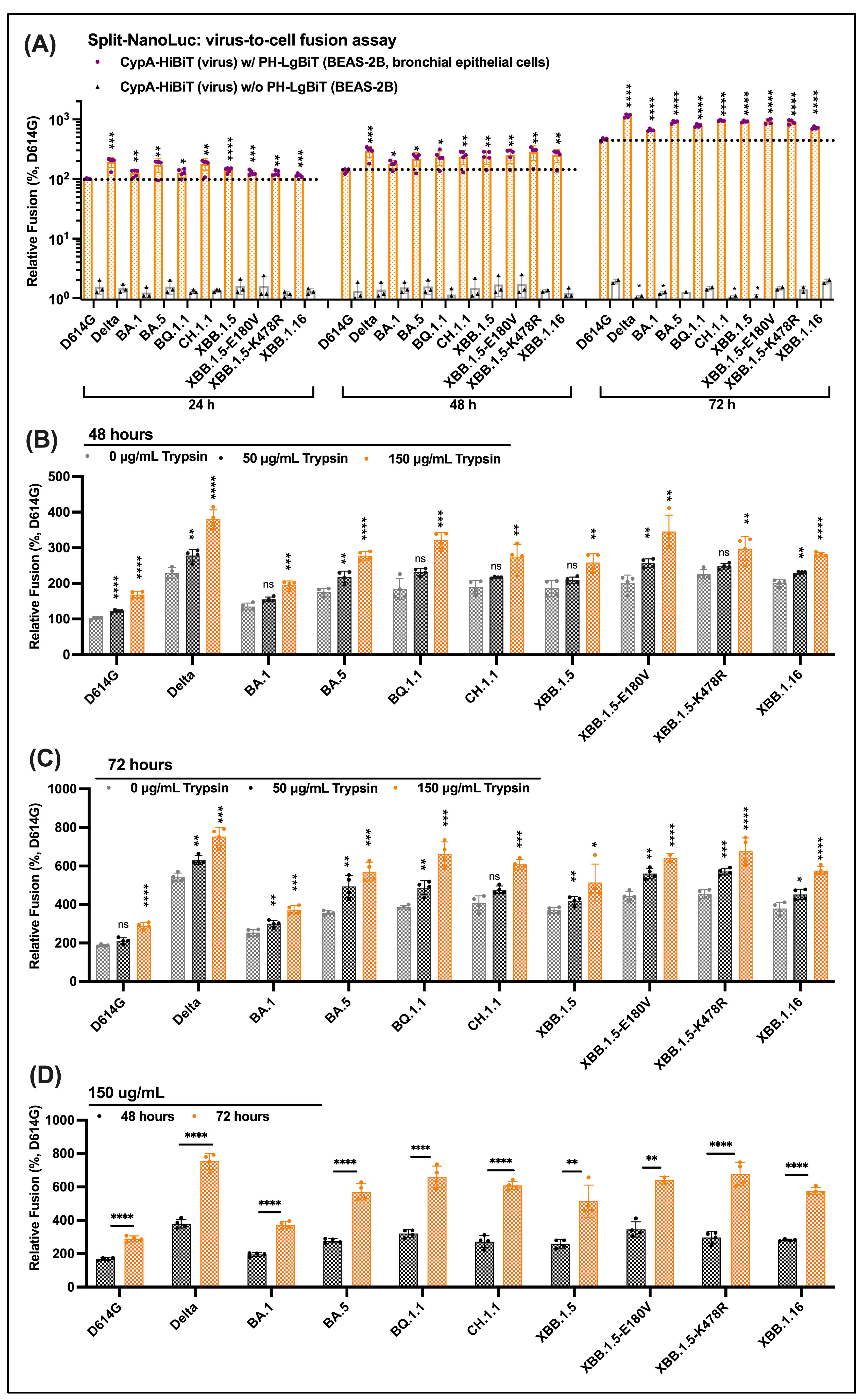
3.5. Efficient Virus-to-Cell Entry of Omicron Descendants on BEAS-2B Cells
3.6. Serine Protease Trypsin Promotes Virus-to-Cell Entry of Delta, Omicron and Its Descendants
3.7. No Obvious Difference in Cell-to-Cell Fusion Mediated by Omicron S Descendants
3.8. Omicron Subvariants Total S1/S2 Proteolytic Processing in Cells
4. Discussion
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Contreras, S.; Iftekhar, E.N.; Priesemann, V. From emergency response to long-term management: The many faces of the endemic state of COVID-19. Lancet Reg. Health-Eur. 2023, 30, 100664. [Google Scholar] [CrossRef]
- Altmann, D.M.; Whettlock, E.M.; Liu, S.Y.; Arachchillage, D.J.; Boyton, R.J. The immunology of long COVID. Nat. Rev. Immunol. 2023, 23, 618–634. [Google Scholar] [CrossRef]
- Wu, A.; Wang, L.; Zhou, H.Y.; Ji, C.Y.; Xia, S.Z.; Cao, Y.; Meng, J.; Ding, X.; Gold, S.; Jiang, T.; et al. One year of SARS-CoV-2 evolution. Cell Host Microbe 2021, 29, 503–507. [Google Scholar] [CrossRef]
- Markov, P.V.; Ghafari, M.; Beer, M.; Lythgoe, K.; Simmonds, P.; Stilianakis, N.I.; Katzourakis, A. The evolution of SARS-CoV-2. Nat. Rev. Microbiol. 2023, 21, 361–379. [Google Scholar] [CrossRef]
- Lipsitch, M.; Krammer, F.; Regev-Yochay, G.; Lustig, Y.; Balicer, R.D. SARS-CoV-2 breakthrough infections in vaccinated individuals: Measurement, causes and impact. Nat. Rev. Immunol. 2022, 22, 57–65. [Google Scholar] [CrossRef]
- Ke, Z.; Oton, J.; Qu, K.; Cortese, M.; Zila, V.; McKeane, L.; Nakane, T.; Zivanov, J.; Neufeldt, C.J.; Cerikan, B.; et al. Structures and distributions of SARS-CoV-2 spike proteins on intact virions. Nature 2020, 588, 498–502. [Google Scholar] [CrossRef] [PubMed]
- Walls, A.C.; Park, Y.J.; Tortorici, M.A.; Wall, A.; McGuire, A.T.; Veesler, D. Structure, function, and antigenicity of the SARS-CoV-2 spike glycoprotein. Cell 2020, 181, 281–292.e6. [Google Scholar] [CrossRef] [PubMed]
- Ghimire, D.; Han, Y.; Lu, M. Structural plasticity and immune evasion of SARS-CoV-2 spike variants. Viruses 2022, 14, 1255. [Google Scholar] [CrossRef] [PubMed]
- Lu, M.; Uchil, P.D.; Li, W.; Zheng, D.; Terry, D.S.; Gorman, J.; Shi, W.; Zhang, B.; Zhou, T.; Ding, S.; et al. Real-Time conformational dynamics of SARS-CoV-2 spikes on virus particles. Cell Host Microbe 2020, 28, 880–891.e8. [Google Scholar] [CrossRef] [PubMed]
- Yang, Z.; Han, Y.; Ding, S.; Shi, W.; Zhou, T.; Finzi, A.; Kwong, P.D.; Mothes, W.; Lu, M. SARS-CoV-2 variants increase kinetic stability of open spike conformations as an evolutionary strategy. mBio 2022, 13, e03227-21. [Google Scholar] [CrossRef]
- Parsons, R.J.; Acharya, P. Evolution of the SARS-CoV-2 Omicron spike. Cell Rep. 2023, 42, 113444. [Google Scholar] [CrossRef]
- Cai, Y.; Zhang, J.; Xiao, T.; Peng, H.; Sterling, S.M.; Walsh, R.M., Jr.; Rawson, S.; Rits-Volloch, S.; Chen, B. Distinct conformational states of SARS-CoV-2 spike protein. Science 2020, 369, 1586–1592. [Google Scholar] [CrossRef]
- Wrapp, D.; Wang, N.; Corbett, K.S.; Goldsmith, J.A.; Hsieh, C.L.; Abiona, O.; Graham, B.S.; McLellan, J.S. Cryo-EM structure of the 2019-nCoV spike in the prefusion conformation. Science 2020, 367, 1260–1263. [Google Scholar] [CrossRef]
- Baden, L.R.; El Sahly, H.M.; Essink, B.; Kotloff, K.; Frey, S.; Novak, R.; Diemert, D.; Spector, S.A.; Rouphael, N.; Creech, C.B.; et al. Efficacy and safety of the mRNA-1273 SARS-CoV-2 vaccine. N. Engl. J. Med. 2020, 384, 403–416. [Google Scholar] [CrossRef]
- Polack, F.P.; Thomas, S.J.; Kitchin, N.; Absalon, J.; Gurtman, A.; Lockhart, S.; Perez, J.L.; Perez Marc, G.; Moreira, E.D.; Zerbini, C.; et al. Safety and efficacy of the BNT162b2 mRNA COVID-19 vaccine. N. Engl. J. Med. 2020, 383, 2603–2615. [Google Scholar] [CrossRef]
- Voysey, M.; Clemens, S.A.C.; Madhi, S.A.; Weckx, L.Y.; Folegatti, P.M.; Aley, P.K.; Angus, B.; Baillie, V.L.; Barnabas, S.L.; Bhorat, Q.E.; et al. Safety and efficacy of the ChAdOx1 nCoV-19 vaccine (AZD1222) against SARS-CoV-2: An interim analysis of four random-ised controlled trials in Brazil, South Africa, and the UK. Lancet 2021, 397, 99–111. [Google Scholar] [CrossRef] [PubMed]
- Chary, M.; Barbuto, A.F.; Izadmehr, S.; Tarsillo, M.; Fleischer, E.; Burns, M.M. COVID-19 Therapeutics: Use, Mechanism of Action, and Toxicity (Vaccines, Monoclonal Antibodies, and Immunotherapeu-tics). J. Med. Toxicol. 2023, 19, 205–218. [Google Scholar] [CrossRef] [PubMed]
- Tauzin, A.; Gong, S.Y.; Beaudoin-Bussières, G.; Vézina, D.; Gasser, R.; Nault, L.; Marchitto, L.; Benlarbi, M.; Chatterjee, D.; Nayrac, M.; et al. Strong humoral immune responses against SARS-CoV-2 Spike after BNT162b2 mRNA vaccination with a 16-week interval between doses. Cell Host Microbe 2022, 30, 97–109.e5. [Google Scholar] [CrossRef] [PubMed]
- Chatterjee, D.; Tauzin, A.; Marchitto, L.; Gong, S.Y.; Boutin, M.; Bourassa, C.; Beaudoin-Bussières, G.; Bo, Y.X.; Ding, S.L.; Laumaea, A.; et al. SARS-CoV-2 Omicron Spike recognition by plasma from individuals receiving BNT162b2 mRNA vaccination with a 16-week interval between doses. Cell Rep. 2022, 38, 110429. [Google Scholar] [CrossRef] [PubMed]
- Guenthoer, J.; Lilly, M.; Starr, T.N.; Dadonaite, B.; Lovendahl, K.N.; Croft, J.T.; Stoddard, C.I.; Chohan, V.; Ding, S.L.; Ruiz, F.; et al. Identification of broad, potent antibodies to functionally constrained regions of SARS-CoV-2 spike following a breakthrough infection. Proc. Natl. Acad. Sci. USA 2023, 120, e2220948120. [Google Scholar] [CrossRef] [PubMed]
- Li, W.; Chen, Y.; Prévost, J.; Ullah, I.; Lu, M.; Gong, S.Y.; Tauzin, A.; Gasser, R.; Vézina, D.; Anand, S.P.; et al. Structural basis and mode of action for two broadly neutralizing antibodies against SARS-CoV-2 emerging variants of con-cern. Cell Rep. 2022, 38, 110210. [Google Scholar] [CrossRef]
- Yao, H.; Song, Y.; Chen, Y.; Wu, N.; Xu, J.; Sun, C.; Zhang, J.; Weng, T.; Zhang, Z.; Wu, Z.; et al. Molecular architecture of the SARS-CoV-2 virus. Cell 2020, 183, 730–738.e13. [Google Scholar] [CrossRef] [PubMed]
- Shang, J.; Wan, Y.; Luo, C.; Ye, G.; Geng, Q.; Auerbach, A.; Li, F. Cell entry mechanisms of SARS-CoV-2. Proc. Natl. Acad. Sci. USA 2020, 117, 11727–11734. [Google Scholar] [CrossRef]
- Hoffmann, M.; Kleine-Weber, H.; Pohlmann, S. A multibasic cleavage site in the spike protein of SARS-CoV-2 is essential for infection of human lung cells. Mol. Cell 2020, 78, 779–784.e5. [Google Scholar] [CrossRef] [PubMed]
- Hoffmann, M.; Kleine-Weber, H.; Schroeder, S.; Kruger, N.; Herrler, T.; Erichsen, S.; Schiergens, T.S.; Herrler, G.; Wu, N.H.; Nitsche, A.; et al. SARS-CoV-2 cell entry depends on ACE2 and TMPRSS2 and is blocked by a clinically proven protease inhibitor. Cell 2020, 181, 271–280.e8. [Google Scholar] [CrossRef] [PubMed]
- Jackson, C.B.; Farzan, M.; Chen, B.; Choe, H. Mechanisms of SARS-CoV-2 entry into cells. Nat. Rev. Mol. Cell Biol. 2022, 23, 3–20. [Google Scholar] [CrossRef]
- Bestle, D.; Heindl, M.R.; Limburg, H.; Van Lam van, T.; Pilgram, O.; Moulton, H.; Stein, D.A.; Hardes, K.; Eickmann, M.; Dolnik, O.; et al. TMPRSS2 and furin are both essential for proteolytic activation of SARS-CoV-2 in human airway cells. Life Sci. Alliance 2020, 3, e202000786. [Google Scholar] [CrossRef]
- Zhou, T.; Tsybovsky, Y.; Gorman, J.; Rapp, M.; Cerutti, G.; Chuang, G.-Y.; Katsamba, P.S.; Sampson, J.M.; Schön, A.; Bimela, J.; et al. Cryo-EM structures of SARS-CoV-2 spike without and with ACE2 reveal a pH-dependent switch to mediate endosomal posi-tioning of receptor-binding domains. Cell Host Microbe 2020, 28, 867–879.e5. [Google Scholar] [CrossRef]
- Wang, Q.; Zhang, Y.; Wu, L.; Niu, S.; Song, C.; Zhang, Z.; Lu, G.; Qiao, C.; Hu, Y.; Yuen, K.Y.; et al. Structural and functional basis of SARS-CoV-2 entry by using human ACE2. Cell 2020, 181, 894–904.e9. [Google Scholar] [CrossRef]
- Lan, J.; Ge, J.; Yu, J.; Shan, S.; Zhou, H.; Fan, S.; Zhang, Q.; Shi, X.; Wang, Q.; Zhang, L.; et al. Structure of the SARS-CoV-2 spike receptor-binding domain bound to the ACE2 receptor. Nature 2020, 581, 215–220. [Google Scholar] [CrossRef]
- Chatterjee, D.; Tauzin, A.; Laumaea, A.; Gong, S.Y.; Bo, Y.X.; Guilbault, A.; Goyette, G.; Bourassa, C.; Gendron-Lepage, G.; Medjahed, H.; et al. Antigenicity of the Mu (B.1.621) and A.2.5 SARS-CoV-2 Spikes. Viruses 2022, 14, 144. [Google Scholar] [CrossRef]
- Cai, Y.; Zhang, J.; Xiao, T.; Lavine, C.L.; Rawson, S.; Peng, H.; Zhu, H.; Anand, K.; Tong, P.; Gautam, A.; et al. Structural basis for enhanced infectivity and immune evasion of SARS-CoV-2 variants. Science 2021, 373, 642–648. [Google Scholar] [CrossRef]
- Carabelli, A.M.; Peacock, T.P.; Thorne, L.G.; Harvey, W.T.; Hughes, J.; Peacock, S.J.; Barclay, W.S.; de Silva, T.; Towers, G.J.; Robertson, D.L.; et al. SARS-CoV-2 variant biology: Immune escape, transmission and fitness. Nat. Rev. Microbiol. 2023, 21, 162–177. [Google Scholar] [CrossRef]
- Prevost, J.; Finzi, A. The great escape? SARS-CoV-2 variants evading neutralizing responses. Cell Host Microbe 2021, 29, 322–324. [Google Scholar] [CrossRef] [PubMed]
- Zhang, J.; Cai, Y.; Lavine, C.L.; Peng, H.; Zhu, H.; Anand, K.; Tong, P.; Gautam, A.; Mayer, M.L.; Rits-Volloch, S.; et al. Structural and functional impact by SARS-CoV-2 Omicron spike mutations. Cell Rep. 2022, 39, 110729. [Google Scholar] [CrossRef] [PubMed]
- McCallum, M.; Czudnochowski, N.; Rosen, L.E.; Zepeda, S.K.; Bowen, J.E.; Walls, A.C.; Hauser, K.; Joshi, A.; Stewart, C.; Dillen, J.R.; et al. Structural basis of SARS-CoV-2 Omicron immune evasion and receptor engagement. Science 2022, 375, 864–868. [Google Scholar] [CrossRef]
- Planas, D.; Saunders, N.; Maes, P.; Guivel-Benhassine, F.; Planchais, C.; Buchrieser, J.; Bolland, W.H.; Porrot, F.; Staropoli, I.; Lemoine, F.; et al. Considerable escape of SARS-CoV-2 Omicron to antibody neutralization. Nature 2021, 602, 671–675. [Google Scholar] [CrossRef]
- Cele, S.; Jackson, L.; Khoury, D.S.; Khan, K.; Moyo-Gwete, T.; Tegally, H.; San, J.E.; Cromer, D.; Scheepers, C.; Amoako, D.G.; et al. Omicron extensively but incompletely escapes Pfizer BNT162b2 neutralization. Nature 2021, 602, 654–656. [Google Scholar] [CrossRef]
- Cao, Y.; Wang, J.; Jian, F.; Xiao, T.; Song, W.; Yisimayi, A.; Huang, W.; Li, Q.; Wang, P.; An, R. Omicron escapes the majority of existing SARS-CoV-2 neutralizing antibodies. Nature 2022, 602, 657–663. [Google Scholar] [CrossRef] [PubMed]
- Harvey, W.T.; Carabelli, A.M.; Jackson, B.; Gupta, R.K.; Thomson, E.C.; Harrison, E.M.; Ludden, C.; Reeve, R.; Rambaut, A.; Consortium, C.-G.U.; et al. SARS-CoV-2 variants, spike mutations and immune escape. Nat. Rev. Microbiol. 2021, 19, 409–424. [Google Scholar] [CrossRef]
- Liu, L.; Iketani, S.; Guo, Y.; Chan, J.F.-W.; Wang, M.; Liu, L.; Luo, Y.; Chu, H.; Huang, Y.; Nair, M.S.; et al. Striking antibody evasion manifested by the Omicron variant of SARS-CoV-2. Nature 2022, 602, 676–681. [Google Scholar] [CrossRef]
- Faraone, J.N.; Qu, P.; Zheng, Y.M.; Carlin, C.; Jones, D.; Panchal, A.R.; Saif, L.J.; Oltz, E.M.; Gumina, R.J.; Liu, S.L. Continued evasion of neutralizing antibody response by Omicron XBB.1.16. Cell Rep. 2023, 42, 113193. [Google Scholar] [CrossRef]
- Wang, Q.; Iketani, S.; Li, Z.; Liu, L.; Guo, Y.; Huang, Y.; Bowen, A.D.; Liu, M.; Wang, M.; Yu, J. Alarming antibody evasion properties of rising SARS-CoV-2 BQ and XBB subvariants. Cell 2023, 186, 279–286.e8. [Google Scholar] [CrossRef]
- Yisimayi, A.; Song, W.; Wang, J.; Jian, F.; Yu, Y.; Chen, X.; Xu, Y.; Yang, S.; Niu, X.; Xiao, T.; et al. Repeated Omicron exposures override ancestral SARS-CoV-2 immune imprinting. Nature 2023, 625, 148–156. [Google Scholar] [CrossRef]
- Qu, P.; Evans, J.P.; Kurhade, C.; Zeng, C.; Zheng, Y.M.; Xu, K.; Shi, P.Y.; Xie, X.; Liu, S.L. Determinants and Mechanisms of the Low Fusogenicity and High Dependence on Endosomal Entry of Omicron Subvariants. mBio 2023, 14, e03176-22. [Google Scholar] [CrossRef] [PubMed]
- Zeng, C.; Evans, J.P.; King, T.; Zheng, Y.M.; Oltz, E.M.; Whelan, S.P.J.; Saif, L.; Peeples, M.E.; Liu, S.L. SARS-CoV-2 spreads through cell-to-cell transmission. Proc. Natl. Acad. Sci. USA 2021, 119, e2111400119. [Google Scholar] [CrossRef]
- Changrob, S.; Fu, Y.B.; Guthmiller, J.J.; Halfmann, P.J.; Li, L.; Stamper, C.T.; Dugan, H.L.; Accola, M.; Rehrauer, W.; Zheng, N.Y.; et al. Cross-Neutralization of Emerging SARS-CoV-2 Variants of Concern by Antibodies Targeting Distinct Epitopes on Spike. mBio 2021, 12, e02975-21. [Google Scholar] [CrossRef] [PubMed]
- Wang, P.; Nair, M.S.; Liu, L.; Iketani, S.; Luo, Y.; Guo, Y.; Wang, M.; Yu, J.; Zhang, B.; Kwong, P.D.; et al. Antibody resistance of SARS-CoV-2 variants B.1.351 and B.1.1.7. Nature 2021, 593, 130–135. [Google Scholar] [CrossRef]
- Wang, P.; Casner, R.G.; Nair, M.S.; Wang, M.; Yu, J.; Cerutti, G.; Liu, L.; Kwong, P.D.; Huang, Y.; Shapiro, L.; et al. Increased resistance of SARS-CoV-2 variant P.1 to antibody neutralization. Cell Host Microbe 2021, 29, 747–751.e4. [Google Scholar] [CrossRef]
- Hu, J.; Gao, Q.Z.; He, C.L.; Huang, A.L.; Tang, N.; Wang, K. Development of cell-based pseudovirus entry assay to identify potential viral entry inhibitors and neutralizing antibodies against SARS-CoV-2. Genes Dis. 2020, 7, 551–557. [Google Scholar] [CrossRef] [PubMed]
- Xue, S.Y.; Wang, X.L.; Wang, L.; Xu, W.; Xia, S.; Sun, L.J.; Wang, S.H.; Shen, N.; Yang, Z.Q.; Huang, B.; et al. A novel cyclic γ-AApeptide-based long-acting pan-coronavirus fusion inhibitor with potential oral bioavailability by tar-geting two sites in spike protein. Cell Discov. 2022, 8, 88. [Google Scholar] [CrossRef] [PubMed]
- Zhong, P.; Agosto, L.M.; Ilinskaya, A.; Dorjbal, B.; Truong, R.; Derse, D.; Uchil, P.D.; Heidecker, G.; Mothes, W. Cell-to-Cell transmission can overcome multiple donor and target cell barriers imposed on Cell-Free HIV. PLoS ONE 2013, 8, e53138. [Google Scholar] [CrossRef] [PubMed]
- Yamamoto, M.; Du, Q.L.; Song, J.P.; Wang, H.Y.; Watanabe, A.; Tanaka, Y.; Kawaguchi, Y.; Inoue, J.; Matsuda, Z. Cell–cell and virus–cell fusion assay–based analyses of alanine insertion mutants in the distal alpha9 portion of the JRFL gp41 subunit from HIV-1. J. Biol. Chem. 2019, 294, 5677–5687. [Google Scholar] [CrossRef] [PubMed]
- Mykytyn, A.Z.; Breugem, T.I.; Riesebosch, S.; Schipper, D.; van den Doel, P.B.; Rottier, R.J.; Lamers, M.M.; Haagmans, B.L. SARS-CoV-2 entry into human airway organoids is serine protease-mediated and facilitated by the multibasic cleavage site. eLife 2021, 10, 64508. [Google Scholar] [CrossRef] [PubMed]
- Chen, Y.Z.; Prévost, J.; Ullah, I.; Romero, H.; Lisi, V.; Tolbert, W.D.; Grover, J.R.; Ding, S.L.; Gong, S.Y.; Beaudoin-Bussières, G.; et al. Molecular basis for antiviral activity of two pediatric neutralizing antibodies targeting SARS-CoV-2 Spike RBD. iScience 2023, 26, 105783. [Google Scholar] [CrossRef]
- Tauzin, A.; Beaudoin-Bussières, G.; Benlarbi, M.; Nayrac, M.; Bo, Y.X.; Gendron-Lepage, G.; Medjahed, H.; Perreault, J.; Gokool, L.; Arlotto, P.; et al. Humoral Responses Elicited after a Fifth Dose of SARS-CoV-2 mRNA Bivalent Vaccine. Viruses 2023, 15, 1926. [Google Scholar] [CrossRef]
- Bridges, J.P.; Vladar, E.K.; Huang, H.; Mason, R.J. Respiratory epithelial cell responses to SARS-CoV-2 in COVID-19. Thorax 2021, 77, 203–209. [Google Scholar] [CrossRef]
- Whittaker, G.R. SARS-CoV-2 spike and its adaptable furin cleavage site. Lancet Microbe 2021, 2, e488–e489. [Google Scholar] [CrossRef]
- Liu, Y.; Liu, J.; Johnson, B.A.; Xia, H.; Ku, Z.; Schindewolf, C.; Widen, S.G.; An, Z.; Weaver, S.C.; Menachery, V.D.; et al. Delta spike P681R mutation enhances SARS-CoV-2 fitness over Alpha variant. bioRxiv 2021. [Google Scholar] [CrossRef]
- Saito, A.; Irie, T.; Suzuki, R.; Maemura, T.; Nasser, H.; Uriu, K.; Kosugi, Y.; Shirakawa, K.; Sadamasu, K.; Kimura, I.; et al. Enhanced fusogenicity and pathogenicity of SARS-CoV-2 Delta P681R mutation. Nature 2022, 602, 300–306. [Google Scholar] [CrossRef]
- Lubinski, B.; Fernandes, M.H.V.; Frazier, L.; Tang, T.; Daniel, S.; Diel, D.G.; Jaimes, J.A.; Whittaker, G.R. Functional evaluation of the P681H mutation on the proteolytic activation of the SARS-CoV-2 variant B.1.1.7 (Alpha) spike. iScience 2021, 25, 103589. [Google Scholar] [CrossRef] [PubMed]
- Meng, B.; Abdullahi, A.; Ferreira, I.; Goonawardane, N.; Saito, A.; Kimura, I.; Yamasoba, D.; Gerber, P.P.; Fatihi, S.; Rathore, S.; et al. Altered TMPRSS2 usage by SARS-CoV-2 Omicron impacts infectivity and fusogenicity. Nature 2022, 603, 706–714. [Google Scholar] [CrossRef] [PubMed]
- Willett, B.J.; Grove, J.; MacLean, O.A.; Wilkie, C.; De Lorenzo, G.; Furnon, W.; Cantoni, D.; Scott, S.; Logan, N.; Ashraf, S.; et al. SARS-CoV-2 Omicron is an immune escape variant with an altered cell entry pathway. Nat. Microbiol. 2022, 7, 1709. [Google Scholar] [CrossRef]
- Suzuki, R.; Yamasoba, D.; Kimura, I.; Wang, L.; Kishimoto, M.; Ito, J.; Morioka, Y.; Nao, N.; Nasser, H.; Uriu, K.; et al. Attenuated fusogenicity and pathogenicity of SARS-CoV-2 Omicron variant. Nature 2022, 603, 700–705. [Google Scholar] [CrossRef] [PubMed]
- Petersen, M.S.; Kongsstovu, S.I.; Eliasen, E.H.; Larsen, S.; Hansen, J.L.; Vest, N.; Dahl, M.M.; Christiansen, D.H.; Moller, L.F.; Kristiansen, M.F. Clinical characteristics of the Omicron variant—Results from a Nationwide Symptoms Survey in the Faroe Islands. Int. J. Infect. Dis. 2022, 122, 636–643. [Google Scholar] [CrossRef]
- Bálint, G.; Vörös-Horváth, B.; Széchenyi, A. Omicron: Increased transmissibility and decreased pathogenicity. Signal Transduct. Target. Ther. 2022, 7, 151. [Google Scholar] [CrossRef]
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2024 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Katte, R.H.; Ao, Y.; Xu, W.; Han, Y.; Zhong, G.; Ghimire, D.; Florence, J.; Tucker, T.A.; Lu, M. Differentiating Cell Entry Potentials of SARS-CoV-2 Omicron Subvariants on Human Lung Epithelium Cells. Viruses 2024, 16, 391. https://doi.org/10.3390/v16030391
Katte RH, Ao Y, Xu W, Han Y, Zhong G, Ghimire D, Florence J, Tucker TA, Lu M. Differentiating Cell Entry Potentials of SARS-CoV-2 Omicron Subvariants on Human Lung Epithelium Cells. Viruses. 2024; 16(3):391. https://doi.org/10.3390/v16030391
Chicago/Turabian StyleKatte, Revansiddha H., Yuanyun Ao, Wang Xu, Yang Han, Guohua Zhong, Dibya Ghimire, Jon Florence, Torry A. Tucker, and Maolin Lu. 2024. "Differentiating Cell Entry Potentials of SARS-CoV-2 Omicron Subvariants on Human Lung Epithelium Cells" Viruses 16, no. 3: 391. https://doi.org/10.3390/v16030391
APA StyleKatte, R. H., Ao, Y., Xu, W., Han, Y., Zhong, G., Ghimire, D., Florence, J., Tucker, T. A., & Lu, M. (2024). Differentiating Cell Entry Potentials of SARS-CoV-2 Omicron Subvariants on Human Lung Epithelium Cells. Viruses, 16(3), 391. https://doi.org/10.3390/v16030391




