The Aqueous Leaf Extract of the Medicinal Herb Costus speciosus Suppresses Influenza A H1N1 Viral Activity under In Vitro and In Vivo Conditions
Abstract
1. Introduction
2. Materials and Methods
2.1. Preparation of C. speciosus TB100 Aqueous Extract
2.2. Removal of Endotoxin Contaminants
2.3. Cell Culture and In Vitro Infection of A/Puerto Rico/8/1934 (H1N1)
2.4. Determination of 50% Cytotoxic Concentration (CC50) of TB100
2.5. Determination of 50% Effective Concentration (EC50) of TB100
2.6. Preliminary Assessment of TB100 Time of Addition Assay
2.7. Preliminary Assessement of TB100 Virucidal, Attachment, and Entry Assay
2.8. Assessment of TB100 Antiviral Activity
2.9. TB100 Mediated Induction of IFN-β and IL6
2.10. Western Blot Detection of Transcriptional Regulator Phosphorylation
2.11. TB100 Antiviral Activity—In Vivo Dose Optimization
2.12. TB100 Antiviral Activity—In Vivo Challenge Study
2.13. HPLC Analysis of Chemical Compounds Present in TB100 Aqueous, and Methanol Extracts
2.14. Antiviral Activity of Active Compounds
2.15. Statistical Analysis
3. Results
3.1. Determination of the CC50, EC50 and Selectivity Index of TB100
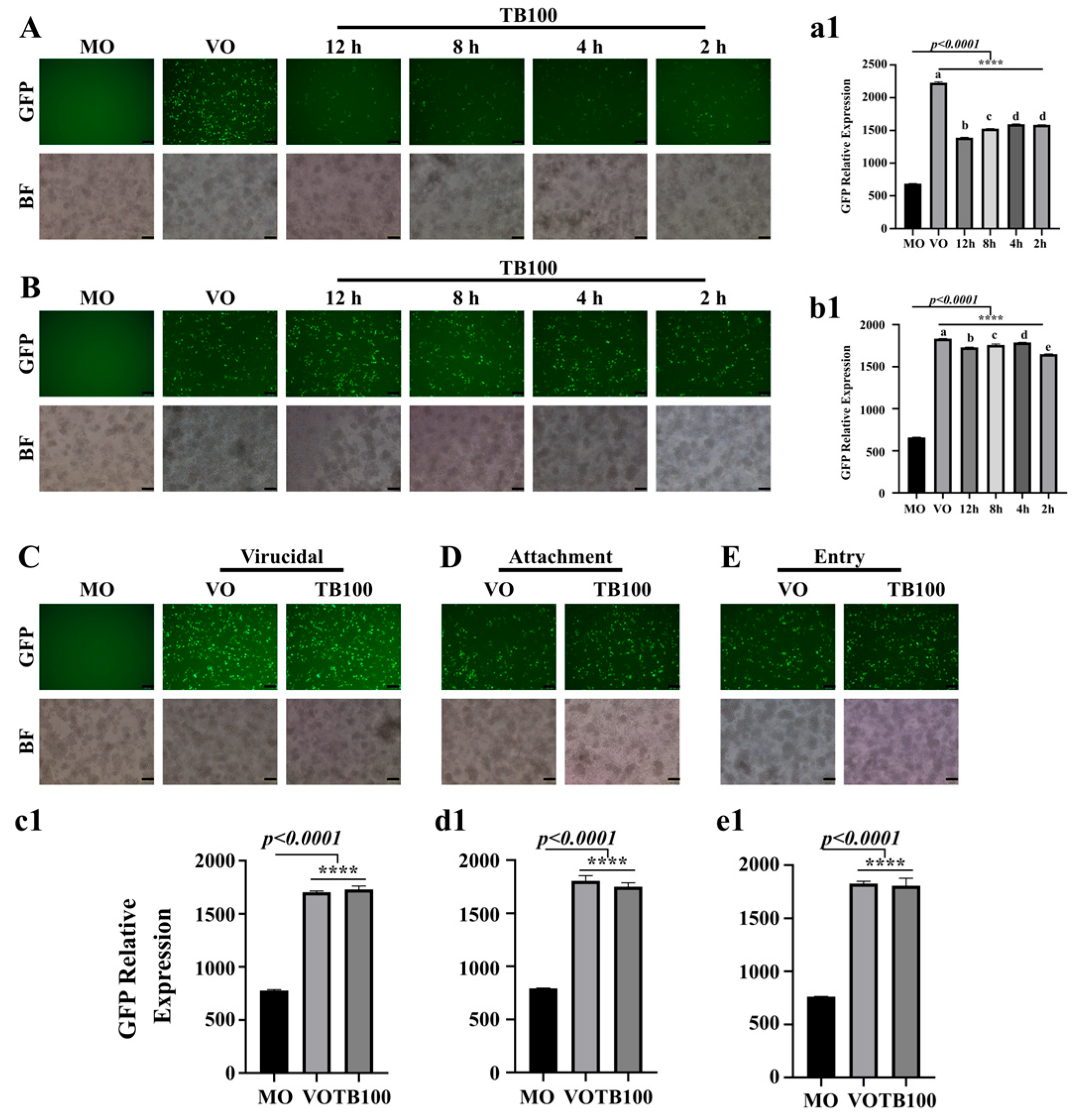
3.2. TB100 Effect on Time of Addition, Virucidal, Attachment, and Entry of A/Puerto Rico/8/1934(H1N1) Influenza
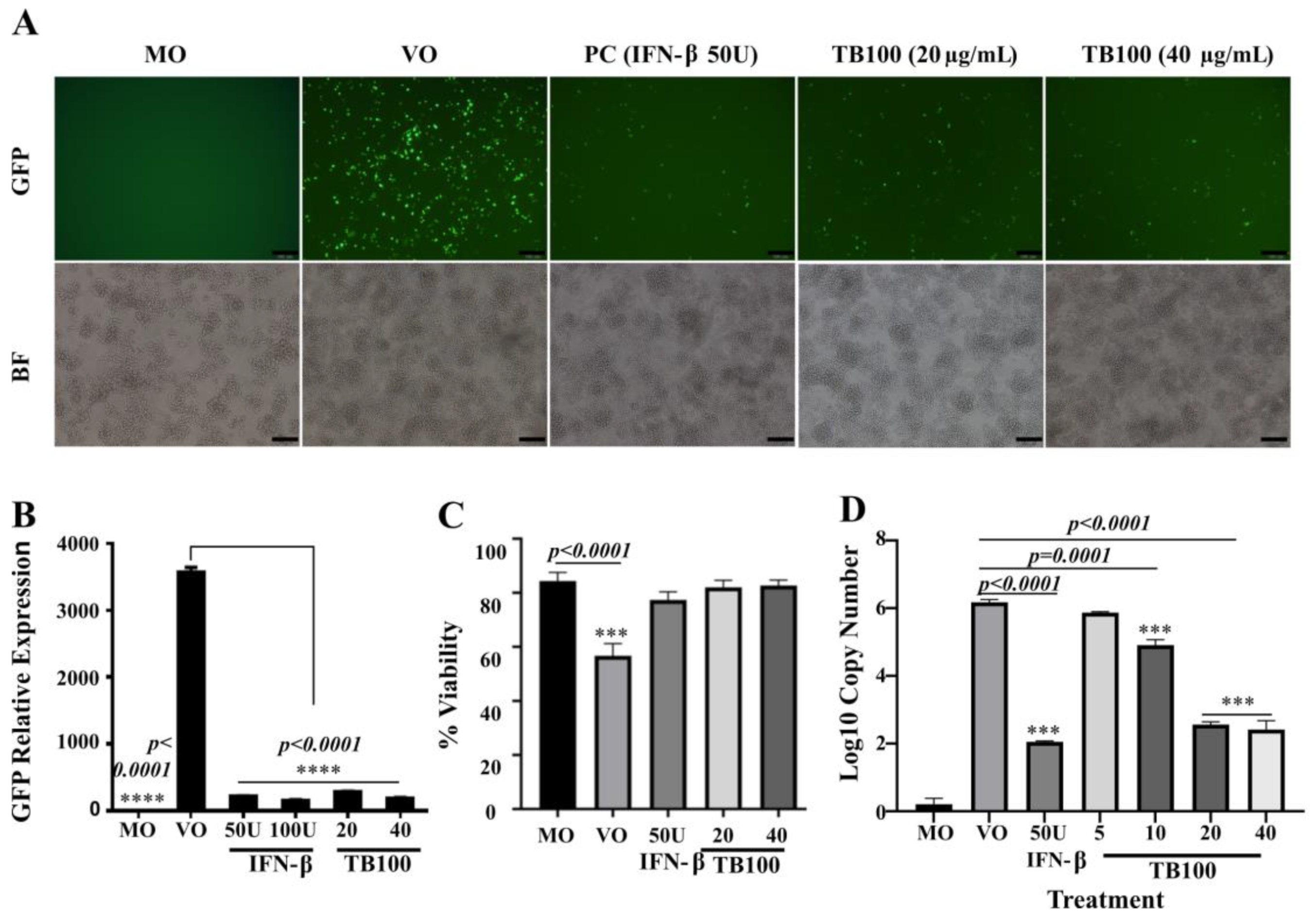
3.3. TB100 Possesses Pretreatment Antiviral Properties against A/Puerto Rico/8/1934 (H1N1) Influenza
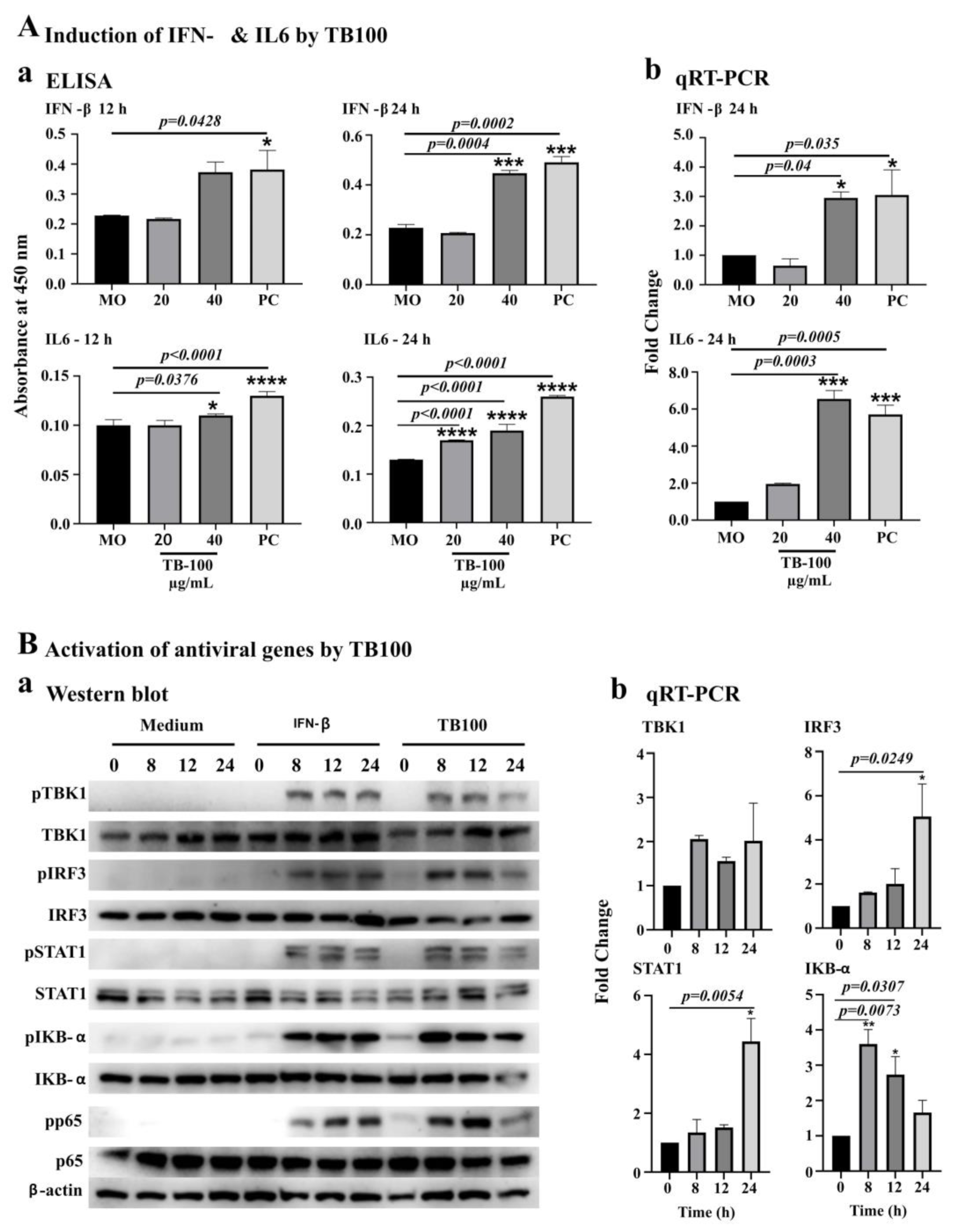
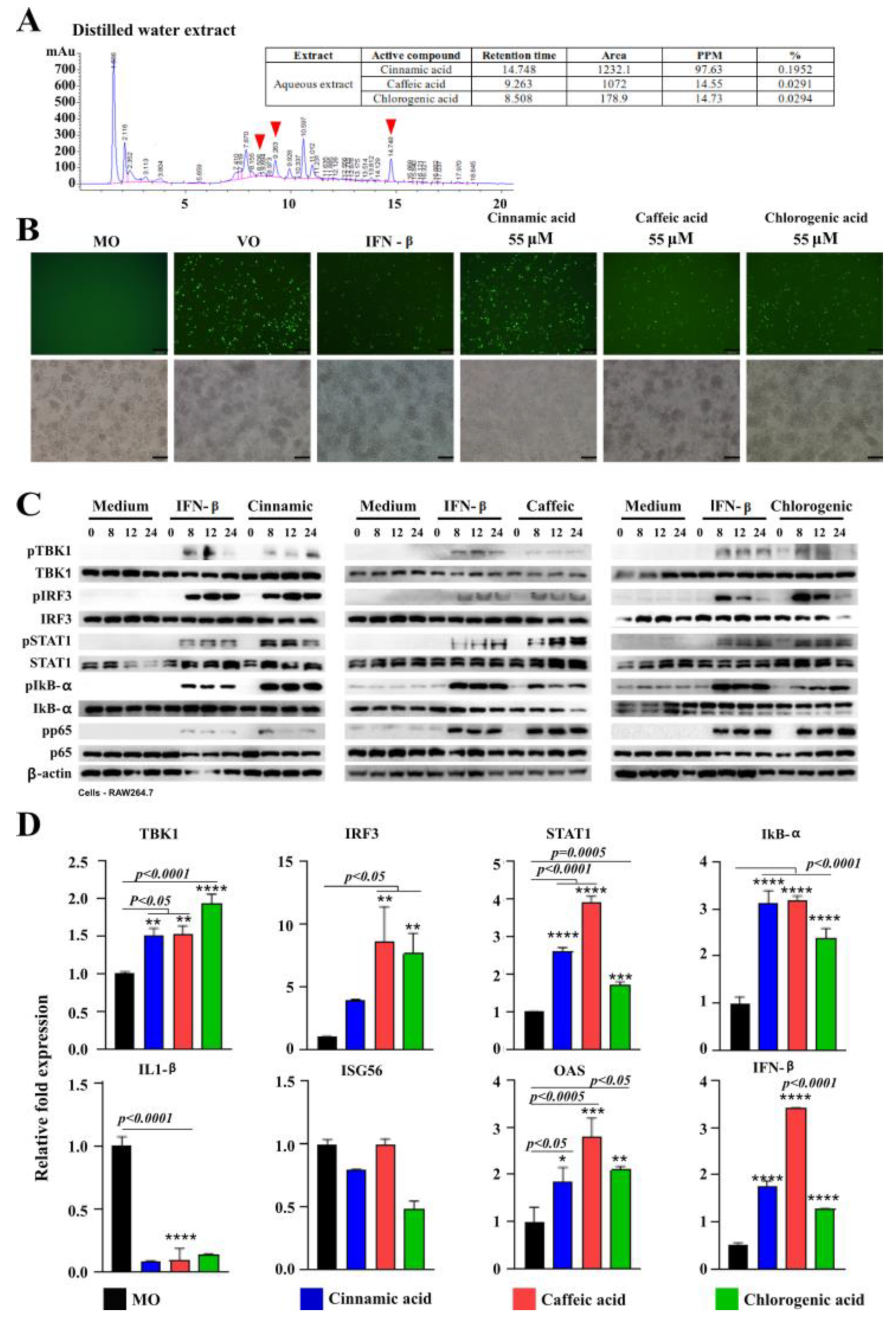
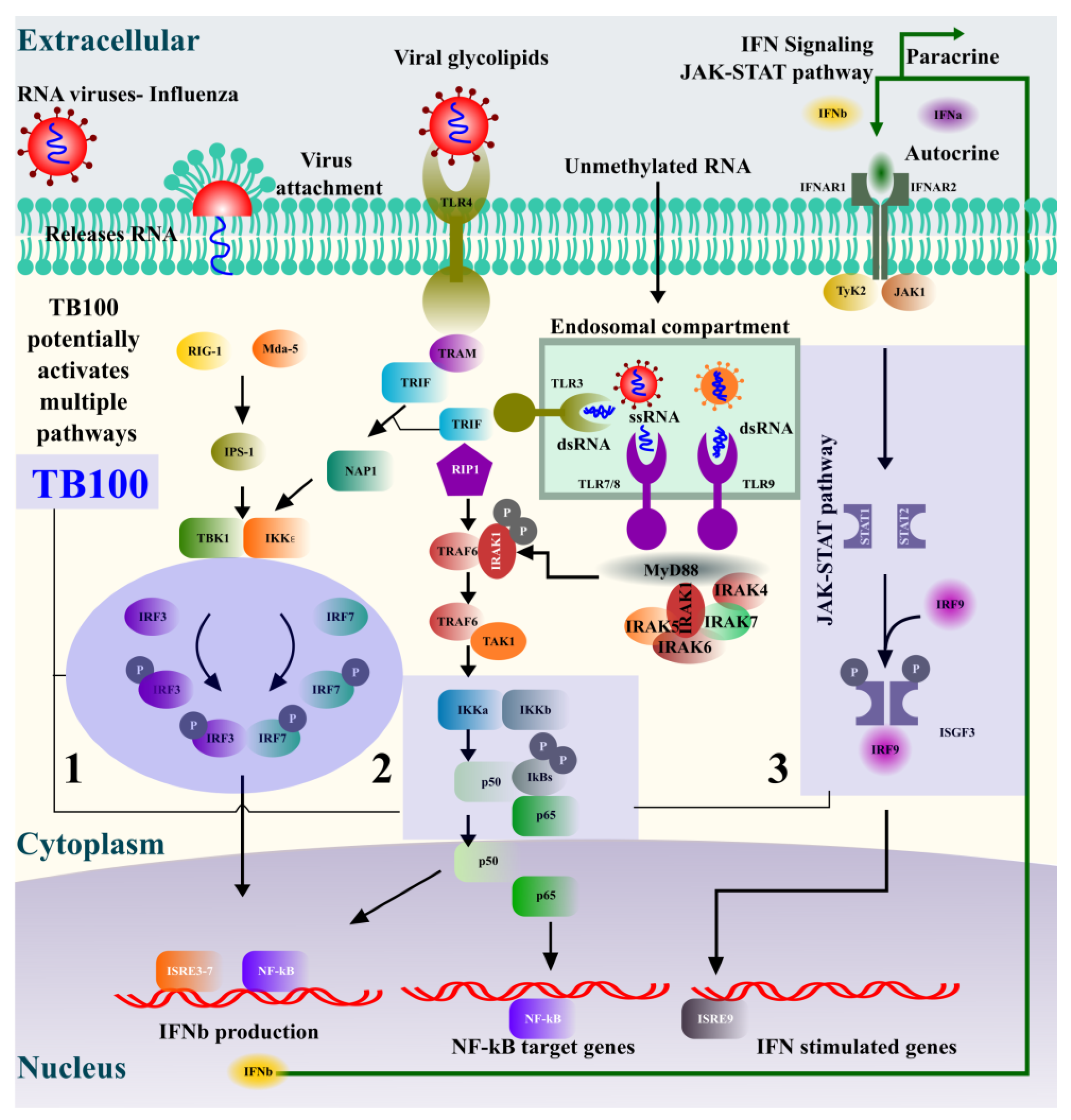
3.4. Assessment for Induction of IFN-β, IL6 and Transcriptional Activators by TB100
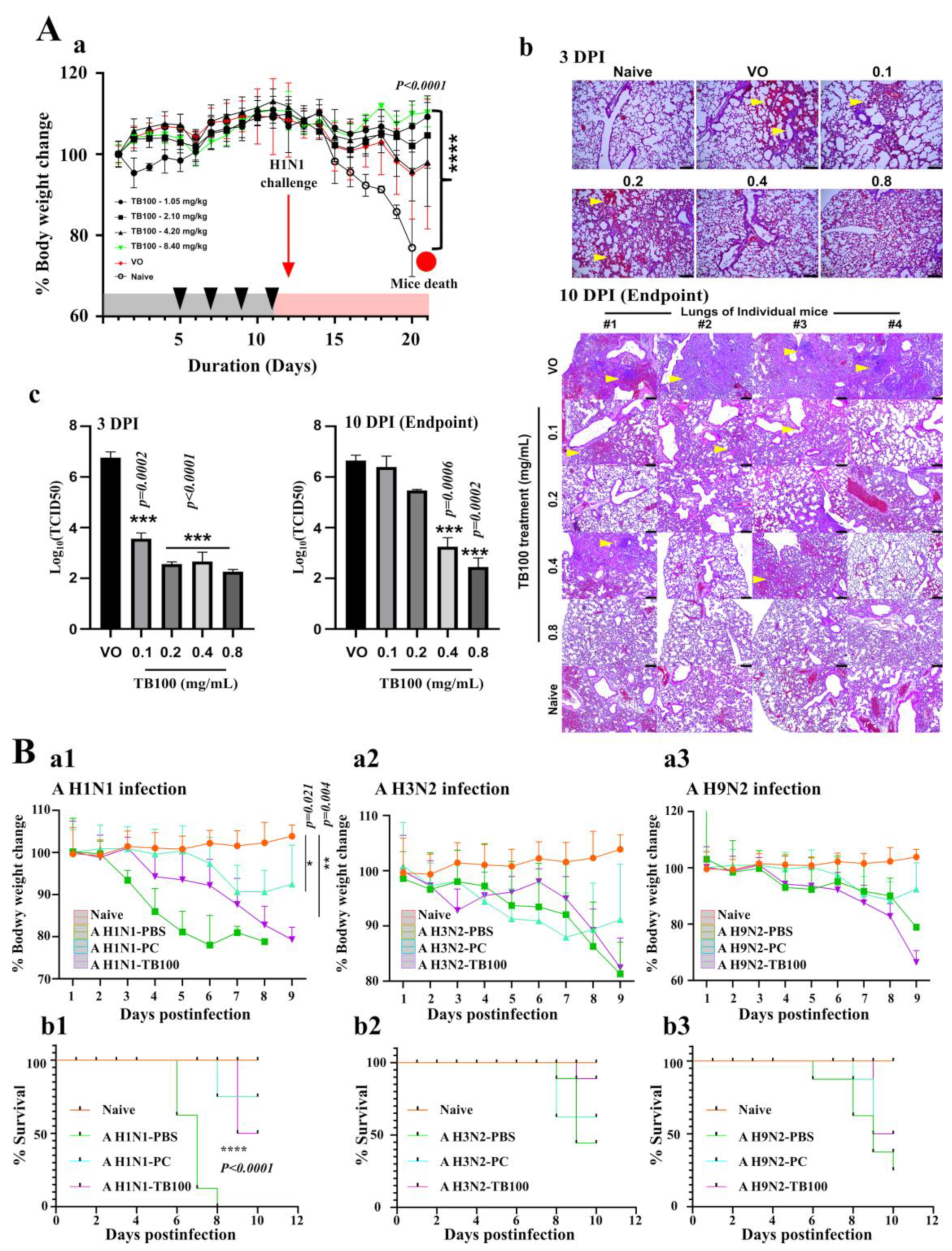
3.5. TB100 In Vivo Dose Optimization and Antiviral Activity
3.6. In Vivo Challenge Study
3.7. Identification ofTB100 Active Compounds and Their Antiviral Effects
3.8. Selectivity Index (SI) of Cinnamic, Caffeic, and Chlorogenic Acids
4. Discussion
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Conflicts of Interest
References
- Young-Xu, Y.; Van Aalst, R.; Russo, E.; Lee, J.K.H.; Chit, A. The Annual Burden of Seasonal Influenza in the Us Veterans Affairs Population. PLoS ONE 2017, 12, e0169344. [Google Scholar] [CrossRef] [PubMed]
- Carrat, F.; Flahault, A. Influenza Vaccine: The Challenge of Antigenic Drift. Vaccine 2007, 66, 45–53. [Google Scholar] [CrossRef] [PubMed]
- Maltezou, H.C.; Ioannidou, E.; De Schrijver, K.; François, G.; De Schryver, A. Influenza Vaccination Programs for Healthcare Personnel: Organizational Issues and Beyond. Int. J. Environ. Res. Public Health 2021, 18, 11122. [Google Scholar] [CrossRef] [PubMed]
- Karlsson Hedestam, G.B.; Fouchier, R.A.M.; Phogat, S.; Burton, D.R.; Sodroski, J.; Wyatt, R.T. The Challenges of Eliciting Neutralizing Antibodies to HIV-1 and to Influenza Virus. Nat. Rev. Microbiol. 2008, 6, 143–155. [Google Scholar] [CrossRef]
- Amarelle, L.; Lecuona, E.; Sznajder, J.I. Anti-Influenza Treatment: Drugs Currently Used and under Development. Arch. Bronconeumol 2017, 53, 19–26. [Google Scholar] [CrossRef]
- Świerczyńska, M.; Mirowska-Guzel, D.M.; Pindelska, E. Antiviral Drugs in Influenza. Int. J. Environ. Res. Public Health 2022, 19, 3018. [Google Scholar] [CrossRef]
- Laborda, P.; Wang, S.Y.; Voglmeir, J. Influenza Neuraminidase Inhibitors: Synthetic Approaches, Derivatives, and Biological Activity. Molecules 2016, 21, 1513. [Google Scholar] [CrossRef]
- Davies, B.E. Pharmacokinetics of Oseltamivir: An Oral Antiviral for the Treatment and Prophylaxis of Influenza in Diverse Populations. J. Antimicrob. Chemother. 2010, 65, ii5–ii10. [Google Scholar] [CrossRef]
- Rouse, B.T.; Sehrawat, S. Immunity and Immunopathology to Viruses: What Decides the Outcome? Nat. Rev. Immunol. 2010, 10, 514–526. [Google Scholar] [CrossRef]
- Lee, J.Y.; Abundo, M.E.C.; Lee, C.W. Herbal Medicines with Antiviral Activity against the Influenza Virus, a Systematic Review. Am. J. Chin. Med. 2018, 46, 1663–1700. [Google Scholar] [CrossRef]
- Lateef Mousa, H.A. Herbal Therapies for Prevention and Treatment of Influenza and Influenza-Like Illness. J. Infect. Dis. Ther. 2015. [Google Scholar] [CrossRef]
- Waisundara, V.Y.; Watawana, M.I.; Jayawardena, N. Costus Speciosus and Coccinia Grandis: Traditional Medicinal Remedies for Diabetes. South Afr. J. Bot. 2015, 98, 1–5. [Google Scholar] [CrossRef]
- Sohrab, S.; Mishra, P.; Mishra, S.K. Phytochemical Competence and Pharmacological Perspectives of an Endangered Boon—Costus Speciosus (Koen.) Sm.: A Comprehensive Review. Bull. Natl. Res. Cent. 2021, 45, 209. [Google Scholar] [CrossRef]
- Wang, Y.J.; Pan, K.L.; Hsieh, T.C.; Chang, T.Y.; Lin, W.H.; Hsu, J.T.A. Diosgenin, a Plant-Derived Sapogenin, Exhibits Antiviral Activity In Vitro against Hepatitis C Virus. J. Nat. Prod. 2011, 74, 580–584. [Google Scholar] [CrossRef]
- Sokolova, A.S.; Putilova, V.P.; Yarovaya, O.I.; Zybkina, A.V.; Mordvinova, E.D.; Zaykovskaya, A.V.; Shcherbakov, D.N.; Orshanskaya, I.R.; Sinegubova, E.O.; Esaulkova, I.L.; et al. Synthesis and Antiviral Activity of Camphene Derivatives against Different Types of Viruses. Molecules 2021, 26, 2235. [Google Scholar] [CrossRef]
- Abubakar, A.R.; Haque, M. Preparation of Medicinal Plants: Basic Extraction and Fractionation ProceduRes. for Experimental Purposes. J. Pharm. Bioallied. Sci. 2020, 12, 1–10. [Google Scholar] [CrossRef]
- Karakus, U.; Crameri, M.; Lanz, C.; Yángüez, E. Propagation and Titration of Influenza Viruses. Methods Mol. Biol. 2018, 1836, 59–88. [Google Scholar]
- Baer, A.; Kehn-Hall, K. Viral Concentration Determination through Plaque Assays: Using Traditional and Novel Overlay Systems. J. Vis. Exp. 2014, 93, e52065. [Google Scholar]
- Verma, A.; Prasad, K.N.; Singh, A.K.; Nyati, K.K.; Gupta, R.K.; Paliwal, V.K. Evaluation of the MTT Lymphocyte Proliferation Assay for the Diagnosis of Neurocysticercosis. J. Microbiol. Methods 2010, 81, 175–178. [Google Scholar] [CrossRef]
- Kumar, P.; Nagarajan, A.; Uchil, P.D. Analysis of Cell Viability by the MTT Assay. Cold Spring Harb. Protoc. 2018, 6, pdb-prot095505. [Google Scholar] [CrossRef]
- Bortoluzzi, A.A.M.; Staffen, I.V.; Banhuk, F.W.; Griebler, A.; Matos, P.K.; Ayala, T.S.; da Silva, E.A.A.; Sarragiotto, M.H.; Schuquel, I.T.A.; Jorge, T.C.M.; et al. Determination of Chemical Structure and Anti-Trypanosoma Cruzi Activity of Extracts from the Roots of Lonchocarpus Cultratus (Vell.) A.M.G. Azevedo & H.C. Lima. Saudi J. Biol. Sci. 2021, 28, 99–108. [Google Scholar] [PubMed]
- Strober, W. Trypan Blue Exclusion Test of Cell Viability. Curr. Protoc. Immunol. 2015, 111, A3.B.1–A3.B.3. [Google Scholar] [CrossRef] [PubMed]
- Hockett, R.D.; Kilby, J.M.; Derdeyn, C.A.; Saag, M.S.; Sillers, M.; Squires, K.; Chiz, S.; Nowak, M.A.; Shaw, G.M.; Bucy, R.P. Constant Mean Viral Copy Number per Infected Cell in Tissues Regardless of High, Low, or Undetectable Plasma HIV RNA. J. Exp. Med. 1999, 189, 1545–1554. [Google Scholar] [CrossRef]
- Čikoš, Š.; Bukovská, A.; Koppel, J. Relative Quantification of MRNA: Comparison of Methods Currently Used for Real-Time PCR Data Analysis. BMC Mol. Biol. 2007, 8, 113. [Google Scholar] [CrossRef]
- OriGene IRF3 Mouse QPCR Primer Pair (NM_016849). Available online: https://www.origene.com/catalog/gene-expression/qpcr-primer-pairs/mp206702/irf3-mouse-qpcr-primer-pair-nm_016849 (accessed on 8 July 2022).
- OriGene TBK1 Mouse QPCR Primer Pair (NM_019786). Available online: https://www.origene.com/catalog/gene-expression/qpcr-primer-pairs/mp216588/tbk1-mouse-qpcr-primer-pair-nm_019786 (accessed on 8 July 2022).
- Chen, E.Q.; Bai, L.; Gong, D.Y.; Tang, H. Employment of Digital Gene Expression Profiling to Identify Potential Pathogenic and Therapeutic Targets of Fulminant Hepatic Failure. J. Transl. Med. 2015, 13, 22. [Google Scholar] [CrossRef]
- Kruger, N.J. The Bradford Method for Protein Quantitation. In The Protein Protocols Handbook; Springer: Berlin/Heidelberg, Germany, 2002; pp. 15–21. ISBN 0-89603-940-4. [Google Scholar]
- Scneider, C.A.; Rasband, W.S.; Eliceiri, K.W. NIH Image to ImageJ: 25 yeard of image analysis. Nat. Methods 2012, 9, 671–675. [Google Scholar] [CrossRef] [PubMed]
- Cardiff, R.D.; Miller, C.H.; Munn, R.J. Manual Hematoxylin and Eosin Staining of Mouse Tissue Sections. Cold Spring Harb. Protoc. 2014, 6, 655–658. [Google Scholar] [CrossRef]
- Wulff, N.H.; Tzatzaris, M.; Young, P.J. Monte Carlo Simulation of the Spearman-Kaerber TCID50. J. Clin. Bioinform. 2012, 2, 5. [Google Scholar] [CrossRef]
- Kim, J.H.; Weeratunga, P.; Kim, M.S.; Nikapitiya, C.; Lee, B.H.; Uddin, M.B.; Kim, T.H.; Yoon, J.E.; Park, C.; Ma, J.Y.; et al. Inhibitory Effects of an Aqueous Extract from Cortex Phellodendri on the Growth and Replication of Broad-Spectrum of Viruses In Vitro and in Vivo. BMC Complement. Altern. Med. 2016, 16, 265. [Google Scholar] [CrossRef]
- Li, R.; Chowdhury, M.Y.E.; Kim, J.H.; Kim, T.H.; Pathinayake, P.; Koo, W.S.; Park, M.E.; Yoon, J.E.; Roh, J.B.; Hong, S.P.; et al. Mucosally Administered Lactobacillus Surface-Displayed Influenza Antigens (SM2 and HA2) with Cholera Toxin Subunit A1 (CTA1) Induce Broadly Protective Immune Responses against Divergent Influenza Subtypes. Vet Microbiol. 2015, 179, 250–263. [Google Scholar] [CrossRef] [PubMed]
- Gheraibia, S.; Belattar, N.; Wahhab, M.A.A. HPLC analysis, anatioxidant and cytotoxic activity of different extracts of Costus speciousis against HepG-2 cell lines. S. Afr. J. Bot 2020, 131, 222–228. [Google Scholar] [CrossRef]
- Inkyu, C.; Park, Y.; Ryu, I.Y.; Jung, H.J.; Ullah, S.; Choi, H.; Park, C.; Kang, D.; Lee., S.; Chun, P.; et al. In silico and in vitro insight into tyrosinase inhibitors with a 2-thioxooxazoline-4-one template. Comput. Struct. Biotechnol. 2021, 19, 37–50. [Google Scholar]
- Amano, R.; Yamashita, A.; Kasai, H.; Hori, T.; Miyasato, S.; Saito, S.; Yokoe, H.; Takahashi, K.; Tanaka, T.; Otoguro, T.; et al. Cinnamic Acid Derivatives Inhibit Hepatitis C Virus Replication via the Induction of Oxidative Stress. Antiviral. Res. 2017, 145, 123–130. [Google Scholar] [CrossRef] [PubMed]
- Chen, Y.; Li, Z.; Pan, P.; Lao, Z.; Xu, J.; Li, Z.; Zhan, S.; Liu, X.; Wu, Y.; Wang, W.; et al. Cinnamic Acid Inhibits Zika Virus by Inhibiting RdRp Activity. Antiviral. Res. 2021, 192, 105117. [Google Scholar] [CrossRef]
- Utsunomiya, H.; Ichinose, M.; Ikeda, K.; Uozaki, M.; Morishita, J.; Kuwahara, T.; Koyama, A.H.; Yamasaki, H. Inhibition by Caffeic Acid of the Influenza a Virus Multiplication In Vitro. Int. J. Mol. Med. 2014, 34, 1020–1024. [Google Scholar] [CrossRef]
- Ding, Y.; Cao, Z.; Cao, L.; DIng, G.; Wang, Z.; Xiao, W. Antiviral Activity of Chlorogenic Acid against Influenza A (H1N1/H3N2) Virus and Its Inhibition of Neuraminidase. Sci. Rep. 2017, 7, 45723. [Google Scholar] [CrossRef]
- Li, X.; Liu, Y.; Hou, X.; Peng, H.; Zhang, L.; Jiang, Q.; Shi, M.; Ji, Y.; Wang, Y.; Shi, W. Chlorogenic Acid Inhibits the Replication and Viability of Enterovirus 71 In Vitro. PLoS ONE 2013, 8, e76007. [Google Scholar] [CrossRef]
- Maji, P.; Ghosh Dhar, D.; Misra, P.; Dhar, P. Costus Speciosus (Koen Ex. Retz.) Sm.: Current Status and Future Industrial Prospects. Ind. Crops Prod. 2020, 152, 112571. [Google Scholar] [CrossRef]
- Bolívar, S.; Anfossi, R.; Humeres, C.; Vivar, R.; Boza, P.; Muñoz, C.; Pardo-Jimenez, V.; Olivares-Silva, F.; Díaz-Araya, G. IFN-β Plays Both pro- and Anti-Inflammatory Roles in the Rat Cardiac Fibroblast through Differential STAT Protein Activation. Front. Pharmacol. 2018, 9, 1368. [Google Scholar] [CrossRef]
- Velazquez-Salinas, L.; Verdugo-Rodriguez, A.; Rodriguez, L.L.; Borca, M.V. The Role of Interleukin 6 during Viral Infections. Front. Microbiol. 2019, 10, 1057. [Google Scholar] [CrossRef] [PubMed]
- Magro, G. SARS-CoV-2 and COVID-19: Is Interleukin-6 (IL-6) the ‘Culprit Lesion’ of ARDS Onset? What Is There besides Tocilizumab? SGP130Fc. Cytokine X 2020, 2, 100029. [Google Scholar] [CrossRef]
- Yang, C.; Murti, A.; Pfeffer, S.; Basu, L.; Kim, J.; Pfeffer, L. IFNα/β promotes cell survival by activating NF-kB. Proc. Natl. Acad. Sci. USA 2000, 97, 13631–13636. [Google Scholar] [CrossRef] [PubMed]
- Lei, X.; Dong, X.; Ma, R.; Wang, W.; Xiao, X.; Tian, Z.; Wang, C.; Wang, Y.; Li, L.; Ren, L.; et al. Activation and Evasion of Type I Interferon Responses by SARS-CoV-2. Nat. Commun. 2020, 11, 32733001. [Google Scholar] [CrossRef] [PubMed]
- Teijaro, J.R. Type i Interferons in Viral Control and Immune Regulation. Curr. Opin. Virol. 2016, 16, 31–40. [Google Scholar] [CrossRef] [PubMed]
- Pahl, H.L. Activators and Target Genes of Rel/NF-ΚB Transcription Factors. Oncogene 1999, 18, 6853–6866. [Google Scholar] [CrossRef]
- Guo, B.; Cheng, G. Modulation of the Interferon Antiviral Response by the TBK1/IKKi Adaptor Protein TANK. J. Biol. Chem. 2007, 282, 11817–11826. [Google Scholar] [CrossRef]
- Tsuchida, T.; Kawai, T.; Akira, S. Inhibition of IRF3-Dependent Antiviral Responses by Cellular and Viral Proteins. Cell Res. 2009, 19, 3–4. [Google Scholar] [CrossRef]
- Shen, Y.; Darnell, J.E. Antiviral Response in Cells Containing Stat1 with Heterologous Transactivation Domains. J. Virol. 2001, 75, 2627–2633. [Google Scholar] [CrossRef]
- Pham, A.M.; TenOever, B.R. The IKK Kinases: Operators of Ativiral Signaling. Viruses 2010, 2, 55–72. [Google Scholar] [CrossRef]
- Bartlett, N.W.; Slater, L.; Glanville, N.; Haas, J.J.; Caramori, G.; Casolari, P.; Clarke, D.L.; Message, S.D.; Aniscenko, J.; Kebadze, T.; et al. Defining Critical Roles for NF-ΚB P65 and Type I Interferon in Innate Immunity to Rhinovirus. EMBO Mol. Med. 2012, 4, 1244–1260. [Google Scholar] [CrossRef] [PubMed]
- Hao, X.Y.; Li, F.-D.; Lv, Q.; Xu, Y.F.; Han, Y.L.; Gao, H. Establishment of BALB/C Mouse Models of Influenza A H1N1 Aerosol Inhalation. J. Med. Virol. 2019, 91, 1918–1929. [Google Scholar] [CrossRef]
- Fukushi, M.; Ito, T.; Oka, T.; Kitazawa, T.; Miyoshi-Akiyama, T.; Kirikae, T.; Yamashita, M.; Kudo, K. Serial Histopathological Examination of the Lungs of Mice Infected with Influenza A Virus PR8 Strain. PLoS ONE 2011, 6, e21207. [Google Scholar] [CrossRef] [PubMed]
- Xie, Y.; Huang, B.; Yu, K.; Shi, F.; Liu, T.; Xu, W. Caffeic Acid Derivatives: A New Type of Influenza Neuraminidase Inhibitors. Bioorg. Med. Chem. Lett. 2013, 23, 3556–3560. [Google Scholar] [CrossRef] [PubMed]
- Song, P.; Li, W.; Xie, J.; Hou, Y.; You, C. Cytokine Storm Induced by SARS-CoV-2. Clin. Chim. Acta 2020, 509, 280–287. [Google Scholar] [CrossRef]
| Target Gene | Primer Name | Sequence (5′-3′) | Ref./Gene Accession |
|---|---|---|---|
| TBK1 (Mouse) | TBK1 FP | CGGAAAGAAGTGTTGCGGTTAGC | [25] |
| TBK1 RP | CAGGCTGCTTTTGCCATTGGTG | ||
| IRF3 (Mouse) | IRF3 FP | CGGAAAGAAGTGTTGCGGTTAGC | [26] |
| IRF3 RP | CAGGCTGCTTTTGCCATTGGTG | ||
| STAT1 (Mouse) | STAT1 FP | GCCTCTCATTGTCACCGAAGAAC | NM_001357627 |
| STAT1 RP | TGGCTGACGTTGGAGATCACCA | ||
| IKB-α (Mouse) | IKB-α FP | GACATGCCTCTCTCCTGTAGTC | [27] |
| IKB-α RP | GGTGAAGCACATCACTGGTCTC | ||
| IL-1β (Mouse) | IL1-β FP | GGTGTGTGACGTTCCCATTA | XM_006498795 |
| IL1-β RP | ATTGAGGTGGAGAGCTTTCAG | ||
| ISG56 (Mouse) | ISG56 FP | AGAGAACAGCTACCACCTTT | AH004732 |
| ISG56 RP | TGGACCTGCTCTGAGATTCT | ||
| OAS (Mouse) | OAS FP | GAGGCGGTTGGCTGAAGAGG | NR_126529.1 |
| OAS RP | GAGGAAGGCTGGCTGTGATTGG | ||
| IFN-β (Mouse) | IFN-β FP | TCCAAGAAAGGACGAACATTCG | X14029 |
| IFN-β RP | TGAGGACATCTCCCACGTCAA | ||
| GFP | GFP FP | GGCAAGCTGACCCTGAAGTT | ON755320 |
| GFP RP | CTTGTAGTTGCCGTCGTCCT | ||
| GAPDH (Mouse) | GAPDH FP | AGGTCATCCCAGAGCTGAACG | NG_148718 |
| GAPDH RP | CACCCTGTTGCTGTAGCCGTAT |
| Cell line | CC50 (µg/mL) | EC50 (µg/mL) | Selectivity Index (SI) |
|---|---|---|---|
| RAW264.7 | 117.12 ± 18.31 | 15.19 ± 0.61 | 7.68 ± 0.89 |
| A549 | 71.86 ± 8.91 | 13.64 ± 0.55 | 5.25 ± 0.44 |
| HEp2 | >300 | 16.78 ± 0.42 | >17.87 |
| Active compound | CC50 (mM) | EC50 (µM) | Selectivity Index (SI) |
|---|---|---|---|
| Cinnamic acid | 6.076 | 28.64 | 212.150 |
| Caffeic acid | 8.925 | 28.18 | 316.713 |
| Chlorogenic acid | 4.875 | 15.62 | 312.099 |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2023 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Senevirathne, A.; Jayathilaka, E.H.T.T.; Haluwana, D.K.; Chathuranga, K.; Senevirathne, M.; Jeong, J.-S.; Kim, T.-W.; Lee, J.-S.; De Zoysa, M. The Aqueous Leaf Extract of the Medicinal Herb Costus speciosus Suppresses Influenza A H1N1 Viral Activity under In Vitro and In Vivo Conditions. Viruses 2023, 15, 1375. https://doi.org/10.3390/v15061375
Senevirathne A, Jayathilaka EHTT, Haluwana DK, Chathuranga K, Senevirathne M, Jeong J-S, Kim T-W, Lee J-S, De Zoysa M. The Aqueous Leaf Extract of the Medicinal Herb Costus speciosus Suppresses Influenza A H1N1 Viral Activity under In Vitro and In Vivo Conditions. Viruses. 2023; 15(6):1375. https://doi.org/10.3390/v15061375
Chicago/Turabian StyleSenevirathne, Amal, E. H. T. Thulshan Jayathilaka, D. K. Haluwana, Kiramage Chathuranga, Mahinda Senevirathne, Ji-Soo Jeong, Tae-Won Kim, Jong-Soo Lee, and Mahanama De Zoysa. 2023. "The Aqueous Leaf Extract of the Medicinal Herb Costus speciosus Suppresses Influenza A H1N1 Viral Activity under In Vitro and In Vivo Conditions" Viruses 15, no. 6: 1375. https://doi.org/10.3390/v15061375
APA StyleSenevirathne, A., Jayathilaka, E. H. T. T., Haluwana, D. K., Chathuranga, K., Senevirathne, M., Jeong, J.-S., Kim, T.-W., Lee, J.-S., & De Zoysa, M. (2023). The Aqueous Leaf Extract of the Medicinal Herb Costus speciosus Suppresses Influenza A H1N1 Viral Activity under In Vitro and In Vivo Conditions. Viruses, 15(6), 1375. https://doi.org/10.3390/v15061375








