Phaeoviruses Present in Cultured and Natural Kelp Species, Saccharina latissima and Laminaria hyperborea (Phaeophyceae, Laminariales), in Norway
Abstract
1. Introduction
2. Materials and Methods
2.1. Sampling
2.2. DNA Extraction
2.3. DNA Amplification by PCR
2.4. Oxford Nanopore Technologies (ONT) Sequencing and Raw Sequence Processing
2.5. Downstream Analysis
3. Results
3.1. Viral Prevalence on Norwegian Kelp Species
3.2. Sequencing and Viral Variants
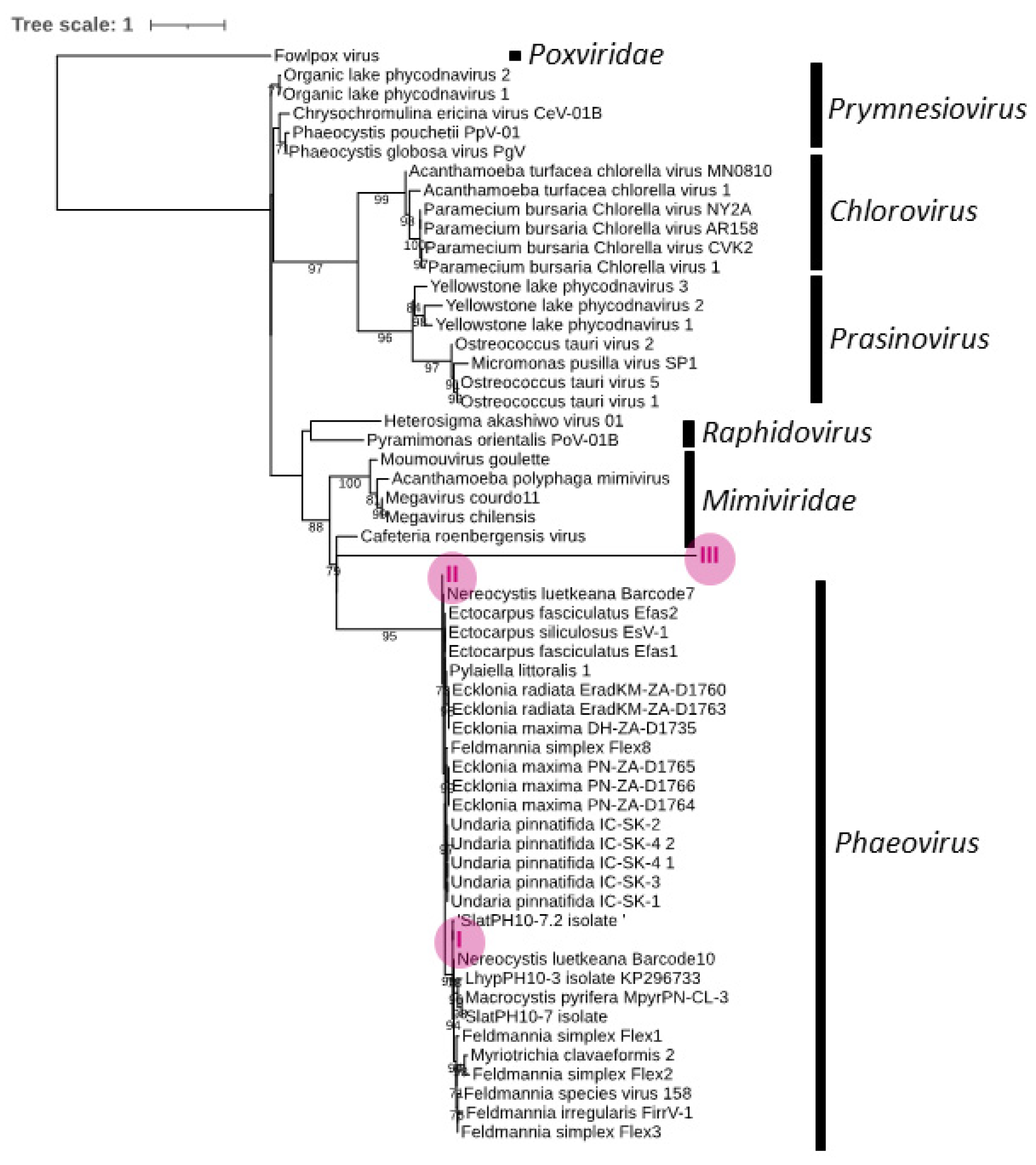
3.3. Phylogeny
4. Discussion
4.1. A Novel Virus Sequence Found in Kelp
4.2. Host Range and Blurred Spatio-Temporal Frontiers
4.3. Phaeoviral Prevalence on the Norwegian Coast and Climate Change
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Koonin, E.V.; Dolja, V.V.; Krupovic, M.; Varsani, A.; Wolf, Y.I.; Yutin, N.; Zerbini, F.M.; Kuhn, J.H. Global Organization and Proposed Megataxonomy of the Virus World. Microbiol. Mol. Biol. Rev. 2020, 84, e00061-19. [Google Scholar] [CrossRef] [PubMed]
- Wilson, W.H.; Allen, M.J. Viruses of Eukaryotic Algae. In Studies in Viral Ecology; Hurst, C.J., Ed.; John Wiley & Sons, Inc.: Hoboken, NJ, USA, 2011; pp. 189–203. [Google Scholar]
- Dunigan, D.D.; Fitzgerald, L.A.; Van Etten, J.L. Phycodnaviruses: A Peek at Genetic Diversity. Virus Res. 2006, 117, 119–132. [Google Scholar] [CrossRef] [PubMed]
- Müller, D.G.; Parodi, E. Transfer of a Marine DNA Virus from Ectocarpus to Feldmannia (Ectocarpales, Phaeophyceae): Aberrant Symptoms and Restitution of the Host. Protoplasma 1993, 175, 121–125. [Google Scholar] [CrossRef]
- Müller, D.G.; Brautigam, M.; Kntppers, R.; Muller, D.G. Virus Infection and Persistence of Foreign DNA in the Marine Brown Alga Feldmannia Simplex (Ectocarpales, Phaeophyceae). Phycologia 1996, 35, 61–63. [Google Scholar] [CrossRef]
- Müller, D.G.; Sengco, M.; Wolf, S.; Bräutigam, M.; Schmid, C.E.; Kapp, M.; Knippers, R. Comparison of Two DNA Viruses Infecting the Marine Brown Algae Ectocarpus Siliculosus and E. Fasciculatus. J. Gen. Virol. 1996, 77, 2329–2333. [Google Scholar] [CrossRef] [PubMed]
- McKeown, D.A.; Stevens, K.; Peters, A.F.; Bond, P.; Harper, G.M.; Brownlee, C.; Brown, M.T.; Schroeder, D.C. Phaeoviruses Discovered in Kelp (Laminariales). ISME J. 2017, 11, 2869–2873. [Google Scholar] [CrossRef] [PubMed]
- McKeown, D.; Schroeder, J.; Stevens, K.; Peters, A.; Sáez, C.; Park, J.; Rothman, M.; Bolton, J.; Brown, M.; Schroeder, D.; et al. Phaeoviral Infections Are Present in Macrocystis, Ecklonia and Undaria (Laminariales) and Are Influenced by Wave Exposure in Ectocarpales. Viruses 2018, 10, 410. [Google Scholar] [CrossRef] [PubMed]
- Maier, I.; Wolf, S.; Delaroque, N.; Müller, D.G.; Kawai, H. A DNA Virus Infecting the Marine Brown Alga Pilayella Littoralis (Ectocarpales, Phaeophyceae) in Culture. Eur. J. Phycol. 1998, 33, 213–220. [Google Scholar] [CrossRef]
- Maier, I.; Müller, D.G.; Katsaros, C. Entry of the DNA Virus, Ectocarpus Fasciculatus Virus Type 1 (Phycodnaviridae), into Host Cell Cytosol and Nucleus. Phycol. Res. 2002, 50, 227–231. [Google Scholar] [CrossRef]
- Meints, R.H.; Ivey, R.G.; Lee, A.M.; Choi, T.-J. Identification of Two Virus Integration Sites in the Brown Alga Feldmannia Chromosome. J. Virol. 2008, 82, 1407–1413. [Google Scholar] [CrossRef]
- Schroeder, D.C. Viruses of Seaweeds. In Studies in Viral Ecology; Hurst, C.J., Ed.; John Wiley & Sons, Inc.: Hoboken, NJ, USA, 2011; pp. 205–215. [Google Scholar]
- Müller, D.G.; Kawai, H.; Stache, B.; Lanka, S. A Virus Infection in the Marine Brown Alga Ectocarpus siliculosus (Phaeophyceae). Bot. Acta 1990, 103, 72–82. [Google Scholar] [CrossRef]
- Müller, D.G.; Stache, B. Worldwide Occurrence of Virus-Infections in Filamentous Marine Brown Algae. Helgol. Meeresunters. 1992, 46, 1–8. [Google Scholar] [CrossRef]
- Cock, J.M.; Sterck, L.; Rouzé, P.; Scornet, D.; Allen, A.E.; Amoutzias, G.; Anthouard, V.; Artiguenave, F.; Aury, J.-M.; Badger, J.H.; et al. The Ectocarpus Genome and the Independent Evolution of Multicellularity in Brown Algae. Nature 2010, 465, 617–621. [Google Scholar] [CrossRef] [PubMed]
- Schroeder, D.C. More to Phaeovirus Infections than First Meets the Eye. Perspect. Phycol. 2015, 2, 105–109. [Google Scholar] [CrossRef]
- Grimsley, N.H.; Thomas, R.; Kegel, J.U.; Jacquet, S.; Moreau, H.; Desdevises, Y. Chapter Nine—Genomics of Algal Host-Virus Interactions. In Advances in Botanical Research; Piganeau, G., Ed.; Elsevier: Amsterdam, The Netherlands, 2012; Volume 64, ISBN 00652296. [Google Scholar]
- Müller, D.G.; Kapp, M.; Knippers, R. Viruses in Marine Brown Algae. Adv. Virus Res. 1998, 50, 49–67. [Google Scholar] [CrossRef] [PubMed]
- Paul, K. Dayton Ecology of Kelp Communities. Annu. Rev. Ecol. Syst. 1985, 16, 215–245. [Google Scholar]
- Smale, D.A.; Moore, P.J. Variability in Kelp Forest Structure along a Latitudinal Gradient in Ocean Temperature. J. Exp. Mar. Biol. Ecol. 2017, 486, 255–264. [Google Scholar] [CrossRef]
- Smale, D.A.; Burrows, M.T.; Moore, P.; O’Connor, N.; Hawkins, S.J. Threats and Knowledge Gaps for Ecosystem Services Provided by Kelp Forests: A Northeast Atlantic Perspective. Ecol. Evol. 2013, 3, 4016–4038. [Google Scholar] [CrossRef]
- Mann, K.H. Seaweeds: Their Productivity and Strategy for Growth. Science 1973, 182, 975–981. [Google Scholar] [CrossRef]
- Duggins, D.O.; Eckman, J.E.; Sewell, A.T. Ecology of Understory Kelp Environments. II. Effects of Kelps on Recruitment of Benthic Invertebrates. J. Exp. Mar. Biol. Ecol. 1990, 143, 27–45. [Google Scholar] [CrossRef]
- Steneck, R.S.; Graham, M.H.; Bourque, B.J.; Corbett, D.; Erlandson, J.M.; Estes, J.A.; Tegner, M.J. Kelp Forest Ecosystems: Biodiversity, Stability, Resilience and Future. Environ. Conserv. 2002, 29, 436–459. [Google Scholar] [CrossRef]
- Duggins, D.O.; Simenstad, C.A.; Estes, J.A. Magnification of Secondary Production by Kelp Detritus in Coastal Marine Ecosystems. Science 1989, 245, 170–173. [Google Scholar] [CrossRef] [PubMed]
- Krumhansl, K.A.; Okamoto, D.K.; Rassweiler, A.; Novak, M.; Bolton, J.J.; Cavanaugh, K.C.; Connell, S.D.; Johnson, C.R.; Konar, B.; Ling, S.D.; et al. Global Patterns of Kelp Forest Change over the Past Half-Century. Proc. Natl. Acad. Sci. USA 2016, 113, 13785–13790. [Google Scholar] [CrossRef]
- Moy, F.E.; Christie, H. Large-Scale Shift from Sugar Kelp (Saccharina latissima) to Ephemeral Algae along the South and West Coast of Norway. Mar. Biol. Res. 2012, 8, 309–321. [Google Scholar] [CrossRef]
- Christie, H.; Andersen, G.S.; Bekkby, T.; Fagerli, C.W.; Gitmark, J.K.; Gundersen, H.; Rinde, E. Shifts between Sugar Kelp and Turf Algae in Norway: Regime Shifts or Fluctuations between Different Opportunistic Seaweed Species? Front. Mar. Sci. 2019, 6, 72. [Google Scholar] [CrossRef]
- Sengco, M.R.; Bräutigam, M.; Kapp, M.; Müller, D.G. Detection of Virus DNA in Ectocarpus siliculosus and E. fasciculatus (Phaeophyceae) from Various Geographic Areas. Eur. J. Phycol. 1996, 31, 73–78. [Google Scholar] [CrossRef]
- Müller, D.G.; Westermeier, R.; Morales, J.; Reina, G.G.; del Campo, E.; Correa, J.A.; Rometscha, E. Massive Prevalence of Viral DNA in Ectocarpus (Phaeophyceae, Ectocarpales) from Two Habitats in the North Atlantic and South Pacific. Bot. Mar. 2000, 43, 157–159. [Google Scholar] [CrossRef]
- Meland, M.; Rebours, C. The Norwegian Seaweed Industry. Bioforsk Fokus 2012, 7, 275–277. [Google Scholar]
- Skjermo, J.; Aasen, I.M.; Arff, J.; Broch, O.J.; Carvajal, A.K.; Christie, H.C.; Forbord, S.; Olsen, Y.; Reitan, K.I.; Rustad, T.; et al. A New Norwegian Bioeconomy Based on Cultivation and Processing of Seaweeds: Opportunities and R&D Needs SINTEF Fisheries and Aquaculture SINTEF Fisheries and Aquaculture; SINTEF Fisheries and Aquaculture: Trondheim, Norway, 2014. [Google Scholar]
- Lima, F.P.; Ribeiro, P.A.; Queiroz, N.; Hawkins, S.J.; Santos, A.M. Do Distributional Shifts of Northern and Southern Species of Algae Match the Warming Pattern? Glob. Chang. Biol. 2007, 13, 2592–2604. [Google Scholar] [CrossRef]
- Harvell, C.D.; Mitchell, C.E.; Ward, J.R.; Altizer, S.; Dobson, A.P.; Ostfeld, R.S.; Samuel, M.D. Climate Warming and Disease Risks for Terrestrial and Marine Biota. Science 2002, 296, 2158–2162. [Google Scholar] [CrossRef]
- Gachon, C.M.M.; Sime-Ngando, T.; Strittmatter, M.; Chambouvet, A.; Kim, G.H. Algal Diseases: Spotlight on a Black Box. Trends Plant Sci. 2010, 15, 633–640. [Google Scholar] [CrossRef] [PubMed]
- Ribeiro, P.A.; Næss, T.; Dahle, G.; Asplin, L.; Meland, K.; Fredriksen, S.; Sjøtun, K. Going with the Flow—Population Genetics of the Kelp Saccharina Latissima (Phaeophyceae, Laminariales). Front. Mar. Sci. 2022, 9, 876420. [Google Scholar] [CrossRef]
- Maeda, T.; Kawai, T.; Nakaoka, M.; Yotsukura, N. Effective DNA Extraction Method for Fragment Analysis Using Capillary Sequencer of the Kelp, Saccharina. J. Appl. Phycol. 2013, 25, 337–347. [Google Scholar] [CrossRef]
- Næss, T. Population Genetics of Saccharina Latissima (Sugar Kelp) in Norway; UiB: Bergen, Norway, 2019. [Google Scholar]
- Fort, A.; Guiry, M.D.; Sulpice, R. Magnetic Beads, a Particularly Effective Novel Method for Extraction of NGS-Ready DNA from Macroalgae. Algal Res. 2018, 32, 308–313. [Google Scholar] [CrossRef]
- Stevens, K.; Weynberg, K.; Bellas, C.; Brown, S.; Brownlee, C.; Brown, M.T.; Schroeder, D.C. A Novel Evolutionary Strategy Revealed in the Phaeoviruses. PLoS ONE 2014, 9, e86040. [Google Scholar] [CrossRef]
- Li, H. Minimap2: Pairwise Alignment for Nucleotide Sequences. Bioinformatics 2018, 34, 3094–3100. [Google Scholar] [CrossRef] [PubMed]
- Katoh, K.; Misawa, K.; Kuma, K.; Miyata, T. MAFFT: A Novel Method for Rapid Multiple Sequence Alignment Based on Fast Fourier Transform. Nucleic Acids Res. 2002, 30, 3059–3066. [Google Scholar] [CrossRef]
- Nguyen, L.-T.; Schmidt, H.A.; von Haeseler, A.; Minh, B.Q. IQ-TREE: A Fast and Effective Stochastic Algorithm for Estimating Maximum Likelihood Phylogenies. Mol. Biol. Evol. 2015, 32, 268–274. [Google Scholar] [CrossRef]
- Kalyaanamoorthy, S.; Minh, B.Q.; Wong, T.K.F.; von Haeseler, A.; Jermiin, L.S. ModelFinder: Fast Model Selection for Accurate Phylogenetic Estimates. Nat. Methods 2017, 14, 587–589. [Google Scholar] [CrossRef]
- Hoang, D.T.; Chernomor, O.; von Haeseler, A.; Minh, B.Q.; Vinh, L.S. UFBoot2: Improving the Ultrafast Bootstrap Approximation. Mol. Biol. Evol. 2017; In Press. [Google Scholar] [CrossRef]
- Letunic, I.; Bork, P. Interactive Tree Of Life (ITOL) v5: An Online Tool for Phylogenetic Tree Display and Annotation. Nucleic. Acids. Res. 2021, 49, W293–W296. [Google Scholar] [CrossRef]
- Wickham, H.; Averick, M.; Bryan, J.; Chang, W.; Mcgowan, L.D.A.; François, R.; Grolemund, G.; Hayes, A.; Henry, L.; Hester, J.; et al. Welcome to the Tidyverse. J. Open Source Softw. 2019, 4, 1686. [Google Scholar] [CrossRef]
- Paradis, E.; Schliep, K. Ape 5.0: An Environment for Modern Phylogenetics and Evolutionary Analyses in R. Bioinformatics 2019, 35, 526–528. [Google Scholar] [CrossRef] [PubMed]
- Oksanen, J.; Simpson, G.; Blanchet, F.; Kindt, R.; Legendre, P.M.P.; O’Hara, R.; Solymos, P.; Stevens, M.; Szoecs, E.; Wagner, H.B.; et al. Version 2022. Vegan: Community Ecology Package_. R Package 2.6-4. 2022. Available online: https://CRAN.R-Project.Org/Package=vegan (accessed on 16 October 2023).
- Wickham, H. Ggplot2: Elegant Graphics for Data Analysis; Springer: New York, NY, USA, 2016. [Google Scholar]
- R Core Team. A Language and Environment for Statistical Computing; R Foundation for Statistical Computing: Vienna, Austria, 2017. [Google Scholar]
- RStudio Team. RStudio: Integrated Development for R; RStudio, Inc.: Boston, MA, USA, 2020. [Google Scholar]
- Microsoft Corporation. Microsoft Excel. 2018. Available online: https://Office.Microsoft.Com/Excel (accessed on 10 October 2023).
- Asplin, L.; Albretsen, J.; Johnsen, I.A.; Sandvik, A.D. The Hydrodynamic Foundation for Salmon Lice Dispersion Modeling along the Norwegian Coast. Ocean Dyn. 2020, 70, 1151–1167. [Google Scholar] [CrossRef]
- Schroeder, D.C.; Schoenrock, K.M.; McHugh, T.A.; Ray, J.; Krueger-Hadfield, S.A. Phaeoviruses Found in Recovering Nereocystis luetkeana Kelp Forest Community. J. Phycol. 2023, 59, 818–821. [Google Scholar] [CrossRef] [PubMed]
- Kawai, H.; Hanyuda, T.; Draisma, S.G.A.; Wilce, R.T.; Andersen, R.A. Molecular Phylogeny of Two Unusual Brown Algae, Phaeostrophion irregulare and Platysiphon glacialis, Proposal of the Stschapoviales Ord. Nov. and Platysiphonaceae Fam. Nov., and a Re-Examination of Divergence Times for Brown Algal Orders. J. Phycol. 2015, 51, 918–928. [Google Scholar] [CrossRef]
- Bringloe, T.T.; Starko, S.; Wade, R.M.; Vieira, C.; Kawai, H.; De Clerck, O.; Cock, J.M.; Coelho, S.M.; Destombe, C.; Valero, M.; et al. Phylogeny and Evolution of the Brown Algae. CRC Crit. Rev. Plant Sci. 2020, 39, 281–321. [Google Scholar] [CrossRef]
- IPCC. Climate Change 2014 Synthesis Report; IPCC: Geneva, Switzerland, 2014. [Google Scholar]
- Miljødirektoratet. Marine Heatwaves in Northen Sea Areas: Occurrence, Effects, and Expected Frequencies; Miljødirektoratet: Trondheim, Norway, 2022. [Google Scholar]
- Raybaud, V.; Beaugrand, G.; Goberville, E.; Delebecq, G.; Destombe, C.; Valero, M.; Davoult, D.; Morin, P.; Gevaert, F. Decline in Kelp in West Europe and Climate. PLoS ONE 2013, 8, e66044. [Google Scholar] [CrossRef]
- Arafeh-Dalmau, N.; Montaño-Moctezuma, G.; Martinez, J.A.; Beas-Luna, R.; Schoeman, D.S.; Torres-Moye, G. Extreme Marine Heatwaves Alter Kelp Forest Community near Its Equatorward Distribution Limit. Front. Mar. Sci. 2019, 6, 499. [Google Scholar] [CrossRef]
- Wernberg, T.; Thomsen, M.S.; Tuya, F.; Kendrick, G.A.; Staehr, P.A.; Toohey, B.D. Decreasing Resilience of Kelp Beds along a Latitudinal Temperature Gradient: Potential Implications for a Warmer Future. Ecol. Lett. 2010, 13, 685–694. [Google Scholar] [CrossRef]
- Filbee-Dexter, K.; Wernberg, T.; Grace, S.P.; Thormar, J.; Fredriksen, S.; Narvaez, C.N.; Feehan, C.J.; Norderhaug, K.M. Marine Heatwaves and the Collapse of Marginal North Atlantic Kelp Forests. Sci. Rep. 2020, 10, 13388. [Google Scholar] [CrossRef]
- Deysher, L.E.; Dean, T.A. In Situ Recruitment of Sporophytes of the Giant Kelp, Macrocystis pyrifera (L.) C.A. Agardh: Effects of Physical Factors. J. Exp. Mar. Biol. Ecol. 1986, 103, 41–63. [Google Scholar] [CrossRef]
- Le, D.M.; Desmond, M.J.; Pritchard, D.W.; Hepburn, C.D. Effect of Temperature on Sporulation and Spore Development of Giant Kelp (Macrocystis pyrifera). PLoS ONE 2022, 17, e0278268. [Google Scholar] [CrossRef] [PubMed]
- Lind, A.C.; Konar, B. Effects of Abiotic Stressors on Kelp Early Life-History Stages. ALGAE 2017, 32, 223–233. [Google Scholar] [CrossRef]
- Jiang, S.C.; Paul, J.H. Significance of Lysogeny in the Marine Environment: Studies with Isolates and a Model of Lysogenic Phage Production. Microb. Ecol. 1998, 35, 235–243. [Google Scholar] [CrossRef] [PubMed]
- Weinbauerl, M.G.; Suttle, A. Lysogeny and Prophage Induction in Coastal and Offshore Bacterial Communities. Aquat. Microb. Ecol. 1999, 18, 217–225. [Google Scholar] [CrossRef]
- Wilson, W.H.; Francis, I.; Ryan, K. Temperature Induction of Viruses in Symbiotic Dinoflagellates. Aquat. Microb. Ecol. 2001, 25, 99–102. [Google Scholar] [CrossRef][Green Version]
- Laffy, P.W.; Botté, E.S.; Wood-Charlson, E.M.; Weynberg, K.D.; Rattei, T.; Webster, N.S. Thermal Stress Modifies the Marine Sponge Virome. Environ. Microbiol. Rep. 2019, 11, 690–698. [Google Scholar] [CrossRef]
- Danovaro, R.; Corinaldesi, C.; Dell’anno, A.; Fuhrman, J.A.; Middelburg, J.J.; Noble, R.T.; Suttle, C.A. Marine Viruses and Global Climate Change. FEMS Microbiol. Rev. 2011, 35, 993–1034. [Google Scholar] [CrossRef]
- Ottersen, G.; Knut, Y.; Børsheim, L.; Arneborg, M.; Maar Aau, V.; Schourup-Kristensen, A.; Rosell, E.A.; Hieronymus, M. Observed and Expected Future Impacts of Climate Change on Marine Environment and Ecosystems in the Nordic Region; Havforskningsinstituttet: Nordnes, Norway, 2023. [Google Scholar]
- Norwegian Ministry of Climate and Environment. Norway’s Eighth National Communication under the Framework Convention on Climate Change; Norwegian Ministry of Climate and Environment: Oslo, Norway, 2023.
- Müller, D.G. Mendelian Segregation of a Virus Genome during Host Meiosis in the Marine Brown Alga Ectocarpus Siliculosus. J. Plant Physiol. 1991, 137, 739–743. [Google Scholar] [CrossRef]
- Bartsch, I.; Wiencke, C.; Bischof, K.; Buchholz, C.M.; Buck, B.H.; Eggert, A.; Feuerpfeil, P.; Hanelt, D.; Jacobsen, S.; Karez, R.; et al. The Genus Laminaria Sensu Lato: Recent Insights and Developments. Eur. J. Phycol. 2008, 43, 1–86. [Google Scholar] [CrossRef]
- Roossinck, M.J.; García-Arenal, F. Ecosystem Simplification, Biodiversity Loss and Plant Virus Emergence. Curr. Opin. Virol. 2015, 10, 56–62. [Google Scholar] [CrossRef] [PubMed]
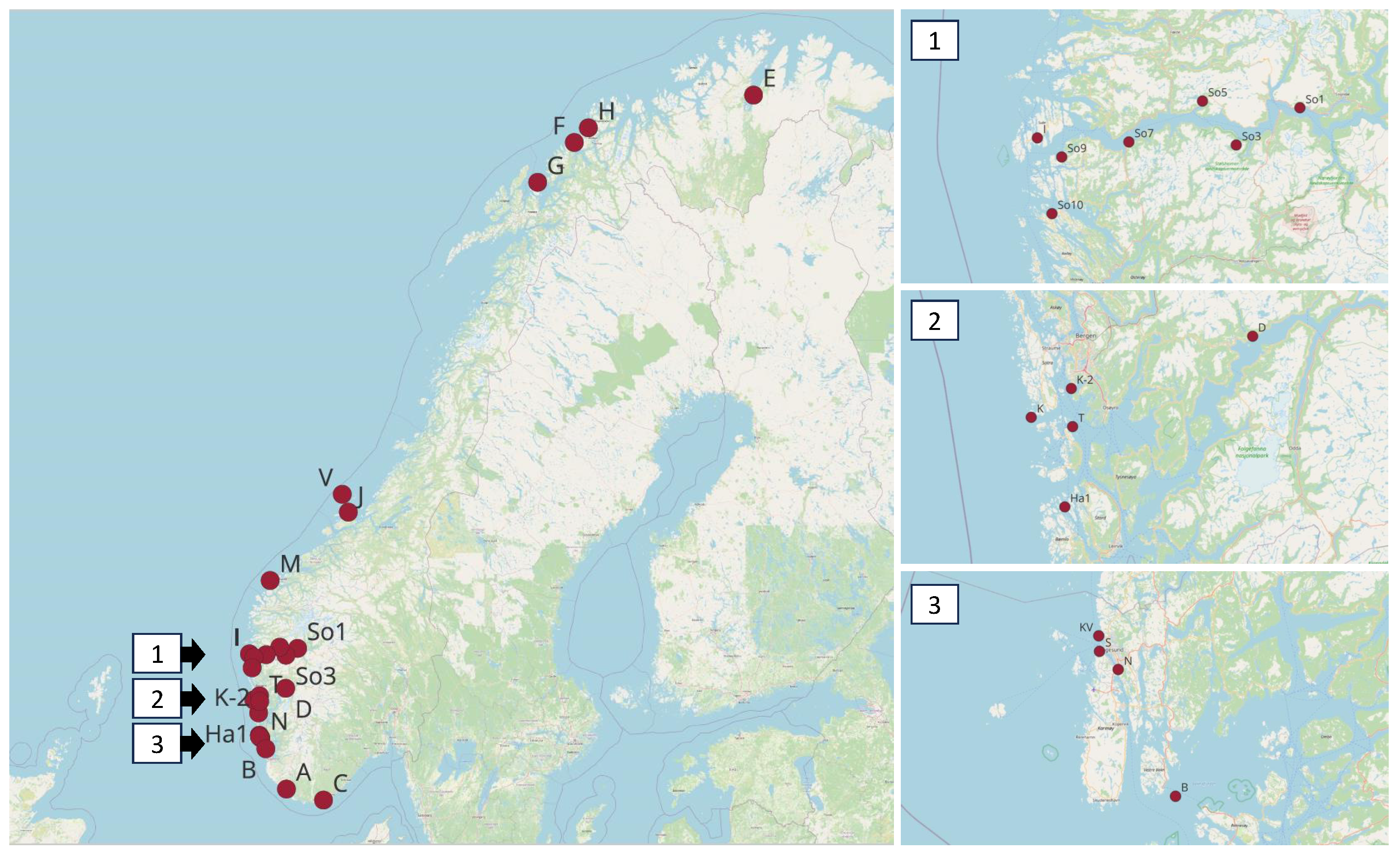


| Location | Sampling Station | Latitude | Longitude | Area |
|---|---|---|---|---|
| Bud, Jøssingfjorden, Vest-Agder | A | 58.308861 | 6.332444 | South |
| Søgne, Vest-Agder | C | 58.073583 | 7.837028 | South |
| Kvam, Hardangerfjorden | D | 60.392111 | 6.300194 | Fjords |
| Porsanger, Finmark | E | 70.297278 | 25.294500 | North |
| Sommarøy, Troms | F | 69.639222 | 18.017917 | North |
| Krøttøy, Troms | G | 69.069472 | 16.531444 | North |
| Vengsøya, Troms | H | 69.844972 | 18.588639 | North |
| Solund, Sognefjorden | I | 61.073931 | 4.831590 | South |
| Frøya, Sør-Trøndelag | J | 63.735389 | 8.845333 | South |
| Korsfjorden, Hordaland | K | 60.237907 | 5.238756 | South |
| Leikanger, Sogn og Fjordane | So1 | 61.182194 | 6.784817 | Fjords |
| Finnefjorden, Sogn og Fjordane | So3 | 61.048350 | 6.311100 | Fjords |
| Høyanger, Sogn og Fjordane | So5 | 61.206100 | 6.060062 | Fjords |
| Oppedalsvika, Sogn og Fjordane | So7 | 61.059367 | 5.512367 | Fjords |
| Nyhamnarsundet, Sogn og Fjordane | So9 | 61.005300 | 5.012750 | South |
| Kilstraumen, Hordaland | So10 | 60.800100 | 4.940267 | South |
| Bårdholmen vest av Fitjar, Hordaland (Hardangerfjorden) | Ha1 | 59.896000 | 5.202667 | South |
| Korsfjorden, Hordaland (Laminaria hyperborea) | K | 60.157397 | 5.006340 | South |
| Korsfjorden, Hordaland (Saccharina latissima) | K-2 | 60.240977 | 5.240332 | South |
| Norheimsvågen, Karmøy, Rogaland. | N | 59.378957 | 5.298491 | South |
| Kvalsvikvegen, Haugesund, Rogaland | KV | 59.437829 | 5.231808 | South |
| Storøy, Karmøy, Rogaland | S | 59.410729 | 5.234212 | South |
| Vikna, Nord-Trondelag | V | 64.054167 | 8.599167 | South |
| Bona Sea Rogaland | B | 59.156389 | 5.495556 | South |
| Møre og Romsdal | M | 62.481667 | 5.670833 | South |
| Trollsøy, Austevoll | T | 60.130350 | 5.248183 | South |
| Ectocarpus siliculosus | Kelp | |
|---|---|---|
| Phaeoviruses present in reproductive structures (gametangia and sporangia). | yes | yes |
| Phaeoviruses present in vegetative cells. | no | yes |
| Phaeoviral symptoms are temperature-sensitive. | yes | ? |
| Virus genome is duplicated during mitosis and segregates in a Mendelian fashion during meiosis. | yes | yes |
| Vertical and lateral (gametes or spores) viral transmission. | yes | yes |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2023 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Ruiz Martínez, E.; Mckeown, D.A.; Schroeder, D.C.; Thuestad, G.; Sjøtun, K.; Sandaa, R.-A.; Larsen, A.; Hoell, I.A. Phaeoviruses Present in Cultured and Natural Kelp Species, Saccharina latissima and Laminaria hyperborea (Phaeophyceae, Laminariales), in Norway. Viruses 2023, 15, 2331. https://doi.org/10.3390/v15122331
Ruiz Martínez E, Mckeown DA, Schroeder DC, Thuestad G, Sjøtun K, Sandaa R-A, Larsen A, Hoell IA. Phaeoviruses Present in Cultured and Natural Kelp Species, Saccharina latissima and Laminaria hyperborea (Phaeophyceae, Laminariales), in Norway. Viruses. 2023; 15(12):2331. https://doi.org/10.3390/v15122331
Chicago/Turabian StyleRuiz Martínez, Eliana, Dean A. Mckeown, Declan C. Schroeder, Gunnar Thuestad, Kjersti Sjøtun, Ruth-Anne Sandaa, Aud Larsen, and Ingunn Alne Hoell. 2023. "Phaeoviruses Present in Cultured and Natural Kelp Species, Saccharina latissima and Laminaria hyperborea (Phaeophyceae, Laminariales), in Norway" Viruses 15, no. 12: 2331. https://doi.org/10.3390/v15122331
APA StyleRuiz Martínez, E., Mckeown, D. A., Schroeder, D. C., Thuestad, G., Sjøtun, K., Sandaa, R.-A., Larsen, A., & Hoell, I. A. (2023). Phaeoviruses Present in Cultured and Natural Kelp Species, Saccharina latissima and Laminaria hyperborea (Phaeophyceae, Laminariales), in Norway. Viruses, 15(12), 2331. https://doi.org/10.3390/v15122331






