Abstract
The clinical course and outcome of COVID-19 are highly variable, ranging from asymptomatic infections to severe disease and death. Understanding the risk factors of severe COVID-19 is relevant both in the clinical setting and at the epidemiological level. Here, we provide an overview of host, viral and environmental factors that have been shown or (in some cases) hypothesized to be associated with severe clinical outcomes. The factors considered in detail include the age and frailty, genetic polymorphisms, biological sex (and pregnancy), co- and superinfections, non-communicable comorbidities, immunological history, microbiota, and lifestyle of the patient; viral genetic variation and infecting dose; socioeconomic factors; and air pollution. For each category, we compile (sometimes conflicting) evidence for the association of the factor with COVID-19 outcomes (including the strength of the effect) and outline possible action mechanisms. We also discuss the complex interactions between the various risk factors.
1. Introduction
Coronavirus disease 2019 (COVID-19) has affected all human populations worldwide, but its toll in mortality and morbidity has been distributed very unevenly across geographical regions, across age groups, and along the spectra of other host, viral and environmental factors. Characterizing the factors that are associated with the outcome of infections with severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2) has relevance at multiple levels. First, an understanding of these factors is required for the assessment of the risk of severe disease in individual patients, which may guide therapeutic decisions in patient care. Antiviral treatments against SARS-CoV-2 are most effective when administered early, before the onset of severe symptoms [1,2]; however, the cost, availability and side-effects of the treatments preclude broad prophylactic application. Under these conditions, effective treatment protocols require reliable prognosis in the early stages of infection, which can be aided by considering known risk factors.
Second, well-characterized risk factors, combined with the prevalence and distribution of these factors in a population can be used to forecast potential mortality and morbidity at the population level, which can inform policy decisions, and guide an optimized public health response.
Third, while a statistical association (between a potential risk factor and COVID-19 outcome) does not necessarily imply causality, identified risk factors may provide clues for causative mechanisms of pathogenesis. Elucidating these mechanisms can guide the development of new therapeutic options, as well as effective non-pharmaceutical interventions against COVID-19.
We provide a structured overview of host, viral and environmental factors that have been shown to be associated with severe clinical outcomes. These associations are typically quantified in terms of risk, odds, or hazard ratios—we provide a brief explanation of these terms in Appendix A.
2. Host Factors
2.1. Age
Age is among the strongest risk factors of COVID-19 mortality. This effect was first reported in early 2020 [3,4], and has since been confirmed by several meta-analyses [5,6]. The risk of death in particular is best reflected by the infection fatality ratio (IFR, probability of death upon infection) of SARS-CoV-2, which has been reliably estimated in relation to age for the first waves of the pandemic by combining information on age-specific mortality and seroprevalence data [7,8,9]. These results indicate that in adults, the IFR increases exponentially with age, doubling the risk of death with approximately every 6–7 years of age, and (for the first large wave of the pandemic) exceeding 1% between 65 and 75 years [9]. In addition to increased mortality, older patients typically experience more severe symptoms [5,10,11] and require hospitalization [12,13], intensive care [5,11,12] and mechanical ventilation (MV) [11,12] more often.
Age-related changes in the human body can affect COVID-19 pathogenesis in a multitude of ways. Aging of the lungs involves increased cellular senescence, epigenetic dysregulation, oxidative stress, mitochondrial dysfunction, inflamm-aging and immunosenescence associated with NK cell cytotoxicity and immune surveillance [14]. Several immunological changes connected to aging may also exacerbate COVID-19 pathogenesis such as altered IFN-γ signaling; neutrophilic infiltration; decreased CD4+ or CD8+ T cell, and naïve B cell levels; alveolar macrophage activation; and elevated release of pro-inflammatory cytokines [14]. Shorter telomeres have also been linked to COVID-19 severity [15]. Moreover, an analysis using single-cell transcriptomics data from multiple cell types identified a handful of genes that are progressively upregulated with age, are dysregulated by SARS-CoV-2 infection, and have important roles both in the aging of the lung and in the pathogenesis of COVID-19 [16]. The genes identified are involved in altered immune cell recruiting, impaired mitochondrial functions, and increased neutrophil attraction (neutrophil extracellular trap (NET) formation).
Age-related changes in the expression of the SARS-CoV-2 cellular entry receptor ACE2 have also been suggested to contribute to the severity of COVID-19. Although pre-COVID-19 analyses reported no significant difference in ACE2 levels according to age [17,18], ACE2 expression in the lungs of COVID-19-infected patients has been demonstrated to correlate with age [16,19], and disease severity also showed an association with ACE2 levels in the respiratory system [20,21]. Furthermore, chronic comorbidities also positively correlated with ACE2 levels in multiple analyses [22,23], which might explain the higher severity of disease in these risk groups. In turn, a higher expression of ACE2 may also counteract oxidative stress and inflammation through the role of the receptor in the renin–angiotensin system (RAS) [24,25], although the significance of this anti-inflammatory mechanism on COVID-19 severity is not well supported by clinical data. The increased expression of TMPRSS2 with age (the serin protease responsible for priming the SARS-CoV-2 spike protein of pre-omicron variants) [26,27] has also been linked to the greater susceptibility of adults to severe COVID-19 [28,29].
Decreased apoptotic sensitivity of aged lung tissue after viral infection has also been implicated in the age dependence of COVID-19 [19]. Early apoptosis mitigates SARS-CoV-2 production [30,31] and, in the case of other infections, it has been shown to decrease both disease severity and mortality [32].
Inflamm-aging is a progressive immunophysiological process associated with increased levels of basal inflammatory mediators (such as IL-6, IL-1β, TNF-α and CRP [33]) driven by the stimulation of the NF-κB signaling pathway and mediated by immune and senescent cells [34]. Severe COVID-19 and advanced age are both correlated with biomarkers of systemic inflammation [35], such as the neutrophil/leukocyte ratio (NLR) [36], weaker type-I IFN responses [37], NLRP3 inflammasome activation [38], and IL-6, IL-12 and IL-1β secretion [39]. Inflamm-aging is a part of a diverse set of mechanisms responsible for the progressive development of hyporesponsiveness and dysregulation of immunity during aging [34,40], called immunosenescence, which is believed to be a major driving force of COVID-19 pathogenesis [41]. Dysregulated immune processes correlated with severe COVID-19 include deficient early type-I IFN production, dysregulated inflammation by neutrophils and monocytes and diminished T cell responses [33,42].
Chronological age is strongly correlated with the molecular and physiological mechanisms of aging but does not reflect individual variation in the rate of these processes. To overcome this problem, several markers of ‘biological age’ have been proposed (telomere length, transcriptomic and metabolic signals, composite markers, etc.), second-generation epigenetic clocks (e.g., PhenoAge and GrimAge) being the most reliable estimators of all-cause mortality and aging-related physiological changes [43,44]. According to recent evidence, telomere length shortening [45,46] and the acceleration of epigenetic age (independent of chronological age) [45,47] are both associated with adverse outcomes during COVID-19. Epigenetic markers of disease severity are concentrated near promoter regions (including the promoters of certain aging-related genes [48]) and include the hypermethylation of IFN-related and the hypomethylation of inflammatory genes [49,50]. In turn, COVID-19 might influence the markers included in biological age estimators and possibly, the process of aging as well [51,52].
Finally, although other respiratory viral infections (e.g., respiratory syncytial virus (RSV) and influenza) affect children more severely compared to adults [53,54], most children are protected against severe COVID-19. Potential contributing factors include the low prevalence of chronic comorbidities [55], strong and rapid innate immune responses [56,57] and a more naïve character of immunity (higher ratio of naïve T cells, lower levels of cytotoxic T cells and NK cells and less T cell exhaustion [58]) compared to adults and especially to the elderly [59]. The observed robust innate immunity in children might be the consequence of differences in IFN responses resulting from the dominance of Orf1ab-specific CD4+ T cells [60], which act against non-structural proteins diminishing IFN signalization [61] or higher trained immunity resulting from frequent viral infections during childhood [62]. We note that all these factors may also affect the course of other respiratory viral infections, and the lower severity typically observed in adults with endemic respiratory viruses may be a consequence of pathogen-specific adaptive immunity acquired in childhood, which was also evidently absent in adults against SARS-CoV-2 during the first waves of the pandemic.
We conclude that the strong impact of age on the outcome of COVID-19 in adults may be a combined effect of multiple factors associated with aging.
2.2. Human Genetic Variation
Hundreds of thousands of SARS-CoV-2-infected cases have been screened for genetic information using whole-genome or whole-exome sequencing, or genotyping microarrays. These analyses have identified dozens of genes/loci that appear to be correlated with COVID-19 severity; in Table 1, we list the most plausible candidates that have either been confirmed by independent analyses and/or that have been regularly listed in reviews based on both bioinformatic and empirical studies.

Table 1.
Genes with rare or common variants that have highly supported association with COVID-19 severity.
The variants identified by high-throughput genetic screening might affect COVID-19 by interfering with viral entry, by modulating antiviral immunity, or by modulating the renin–angiotensin system responsible for the regulation of blood pressure that SARS-CoV-2 interferes with through its binding to ACE2; for some other variants, the mechanistic link is less clear. The effect sizes (odds ratios, OR) for individual variants ranged between OR = 0.819 (0.781–0.858) and OR = 1.885 (1.748–2.032) for critical illness, and between OR = 0.861 (0.834–0.889) and OR = 1.649 (1.562–1.741) for hospitalization in the largest GWAS study to date [83]. In another study, the overall effect of human genetic variation was assessed by estimating a polygenic risk score, and a high risk score was associated with a higher risk of severe COVID-19 compared with individuals with low genetic risk score (OR = 1.50 (1.18–1.92)) [84].
While genome-wide screening is able to identify genetic effects in an unbiased, systematic way, it has limited power to detect the effect of rare genetic variants [85]. Several rare genetic disorders have been implicated in severe COVID-19 by targeted analyses of a heterogeneous group of genetic variants called inborn errors of immunity (also called primary immunodeficiencies) [86]. The strongest candidates are loss-of-function variants of the X-chromosomal TLR7 gene (see also Table 1), which are associated with impaired IFN I and III responses, and are enriched in males under 60 years with critical COVID-19 outcomes [76,87,88,89]. A large study found that antiviral interferon responses might be compromised in at least 3.5% of patients with severe COVID-19 pneumonia via other components of the signaling pathway, such as TLR3, IRF7 or IFNAR1 [90], but independent analyses failed to reproduce the association of these rare alleles with severe disease [91,92]. While the impact of individual variants is hard to estimate due to the low sample sizes, a systematic review of primary immunodeficiencies estimated an overall hospitalization rate of 49%, and a case fatality rate of 9% for all cases combined [93].
Finally, although the role of host genetics is unknown in this case, the role of interferon signaling and the renin–angiotensin system in COVID-19 is further supported by the observation that IFN-I-neutralizing [94] and AngII autoantibodies [95] can be detected in a significant portion of deceased and hospitalized COVID-19 patients, respectively.
Most of the identified rare variants are associated with an increased probability of adverse clinical outcomes in COVID-19. While there is no known genetic variant that would offer complete resistance against SARS-CoV-2 infection [96,97], a small number of candidate variants might decrease the severity of disease (Table 1). An exonic mutation in TMPRSS2 [64,98], an indel in ACE1 [64], a mucus-oversecreting MUC5B variant [67,69,99], some HLA class 1 alleles [100,101,102] and blood group O (possibly by lowering the risk of cardiovascular complications) [103] may have a protective effect against severe COVID-19.
Several studies have addressed the impact of ancestry, which defines the large-scale patterns of human genetic variation, on the risk of severe COVID-19 [104,105,106]. However, while non-European ancestry has been repeatedly linked to a higher risk of severe outcomes [107], it seems unlikely that genetic differences account for the observed effect [67,68]. Instead, ancestry is often correlated with socioeconomic factors that affect COVID-19 outcomes (see Section 4.1. Socioeconomic factors), and the statistical effect of ancestry appears to be explained by correlated socioeconomic variables [106,108,109]. A rare known example related to ancestry is presented by some genetic components that have entered the human genome through Neanderthal introgression, such as the 3p21.31 locus (LTZFL1/SLC6A20) that increases [110], and the OAS cluster that decreases [111] the risk of severe COVID-19. Accordingly, both variants are more common in European and South Asian compared to African and East Asian populations.
We conclude that a few human genetic polymorphisms might have a relatively strong negative impact on the outcome of COVID-19, but most individual variants, and ancestry/ethnicity, appear to have no or only small effect.
2.3. Sex and Pregnancy
Certain respiratory viral infections disproportionately affect the two sexes. Males typically experience more frequent lower respiratory tract infections [112] and, in the case of influenza A and B viruses, RSV, SARS-CoV, and other sources of community-acquired pneumonia, more severe disease compared to females [113,114]. Similarly, with SARS-CoV-2 infection, males are hospitalized [8,115], are admitted to intensive care unit (ICU) [8,116] and die [117,118] more often, although infection rates appear to be similar to those of women [119]. Sex-disaggregated estimates of the infection fatality ratio show consistently higher fatality rates among males in every age category [8,9]. We summarize some of the largest studies and meta-analyses demonstrating the association of COVID-19 outcomes with biological sex in Table 2.

Table 2.
Large clinical studies and systematic reviews demonstrating the association of male sex with increased COVID-19 severity.
The underlying causes (which might apply for respiratory infections in general) might include differences in the prevalence of comorbidities [125], lifestyle choices [126], immunogenetic [127] and immunoendocrine [128] factors.
Sex hormones alter the expression of certain genes (through interaction with hormone response elements) related to physiological and immunological functions [129]. Hormonal differences between males and females may thus be partly responsible for sex-specific differences in the responses to respiratory viral infections and, particularly, COVID-19 [130].
Androgens modulate immune and inflammatory processes, typically in an immunosuppressive and anti-inflammatory manner [131]. In COVID-19 patients, this immune-modulating role may even be beneficially connected to the pathogenesis of disease, indicated by the observation that low levels of the major androgen, testosterone (which is a natural consequence of aging in males [132]) are associated with pro-inflammatory states and unfavorable disease outcomes, such as severe symptoms, acute respiratory distress syndrome (ARDS) and ICU admission, even when controlling for age [133,134]. Lastly, a genetic polymorphism, the length of the N-terminal polyQ tract in the androgen receptor is correlated with COVID-19 severity, with shorter alleles implying higher vulnerability against adverse outcomes [135]. Possible mechanisms explaining the negative impact of low testosterone are enhanced replication as a result of the SARS-CoV-2-induced impairment of intracellular Ca2+-regulation [136] and increased pro-inflammatory activity of innate immune cells [137]. In turn, androgens might also facilitate viral entry by the induction of TMPRSS2 expression [138].
Postmenopausal women also showed increased severity of COVID-19 in a small study [139], which may be partly attributed to low estrogen production [140], which is associated with higher risk of COVID-19 hospitalization and increased systemic inflammation [141]. Although age-related changes in sexual functions are difficult to disentangle from other effects of aging, several hypotheses have been proposed for the possible role of sex-linked factors. Estrogen hormones influence the expression of SARS-CoV-2 entry receptors, increasing ACE2 [142] and decreasing TMPRSS2 [143] levels; they induce innate [144] and adaptive [145] immune cells, promote anti-inflammatory functions [144], influence insulin secretion [146] and protection against cardiovascular diseases (CVDs) [147].
In addition to the influence of sex hormones, some X-linked genes may also contribute to sex-specific differences in COVID-19 immunity and outcome. The random inactivation of one of two X chromosomes in females creates variation in the expression of both alleles (‘X mosaicism’). Furthermore, this inactivation might be incomplete in some cases, which increases gene dosage in females. The gene of ACE2 is located on the X chromosome and displays X mosaicism. Although ACE2 expression is similar in respiratory tissues between the two sexes [148,149], age-related declines of its expression might be more pronounced in men [150]. Remarkably, while the respiratory expression of TMPRSS2 is also similar between the sexes [148], cells co-expressing both ACE2 and TMPRSS2 are 3-fold more abundant in males [151]. However, the role of these sex-specific differences in SARS-CoV-2 entry receptor expression on the observed clinical outcomes is unclear. The immune receptor TLR7 shows incomplete X-inactivation with higher expression and downstream signaling in females and reduced functions in males [152]. The Y chromosome is also involved in immune functions, altering gene expression in CD4+ T cells and macrophages [153]. Furthermore, loss of Y chromosome in immune cells is associated with impaired lymphoid functions [154,155].
Adaptive immune responses in women are also stronger against SARS-CoV-2 [156], indicated by greater IFN signalization [157], more robust T cell activation [158,159,160], and effective humoral immunity [161,162]. Moreover, autoantibody production against IFNs [163,164], and inborn errors of immunity such as impaired IFN signalization [76] (see Section 2.2. Human Genetic Variation) are also more often found in male patients. These factors might explain the observed lower concentration of pro-inflammatory (NLR, CRP, IL-6, and TNF-α) [161,165] and cellular damage markers (ALT and AST) [158] in women compared with men.
Although female patients are somewhat more protected against unfavorable outcomes of COVID-19, they are more severely affected by its long-term complications [166] and by lifestyle changes in the COVID-19 era [167].
Finally, pregnancy is a risk factor of severe COVID-19 that only affects women. Pregnant women show increased susceptibility to several viruses such as influenza A [168], SARS-CoV and MERS-CoV [169]. Similarly, even though the overall risk of unfavorable outcomes is rare [170,171,172], the risk of severe outcomes such as pneumonia [173], ICU admission [173,174], MV [174] and mortality [175,176] seems to be higher in pregnant, compared with non-pregnant women. In addition to the effect of pregnancy on the severity of COVID-19, SARS-CoV-2 infection also seems to influence the risk of complications in pregnancy. Compared with pregnant SARS-CoV-2-negative women, pregnant infected women more often develop hypertensive disorders (preeclampsia, eclampsia) [177,178], are admitted to ICU [174,179,180], receive MV [181] or die [174]. Adverse outcomes also affect the development of the fetus. Before or during delivery, fetuses of SARS-CoV-2-positive women are more likely to experience hyperbilirubinemia [182], intrapartum fetal distress [183], cesarean delivery [180,184,185], fetal growth restriction (resulting in low birthweight) [184], preterm birth [180,184,186,187] and stillbirth [177,186,187]. SARS-CoV-2 infection early in gestation seems to produce more severe consequences; however, most SARS-CoV-2-positive women are in the second or third trimester at the time of diagnosis [187]. Newborns of SARS-CoV-2-infected mothers have an increased risk of ARDS [182], ICU admission [174,180,182] and neonatal death [176].
Pregnancy is accompanied by complex endocrine, physiological, and immunological changes which might affect COVID-19 pathogenesis. While increased severity of COVID-19 in pregnant women is not properly understood, SARS-CoV-2 infection may damage the fetus in direct and indirect ways. The co-expression of ACE2 and TMPRSS2 on placental cells is rare [188], but several alternative SARS-CoV-2 entry receptors (CTSL, CTSB, and BSG/CD147) are abundant on multiple placental cell types [189]. Direct infection of the placenta is possible ex vivo [190], but seems to be rare in vivo [191]. However, the rare cases of viral invasion can involve extensive areas of the placenta [192]. Similarly, intrauterine vertical transmission of SARS-CoV-2 has been documented [193], but seems to be rare [194,195], possibly due to the relatively low viraemia in COVID-19 patients [196]. Inflammatory markers have been found in umbilical cord blood [197], indicating that indirect damage to the placenta and to the fetus by pro-inflammatory cytokines might also contribute to neonatal complications [198]. Whether directly or indirectly, damage to the placenta might induce a severe hypoxic state responsible for the adverse effects of COVID-19 [199]. Inflammation due to SARS-CoV-2 infection might also contribute to cytokine-driven neonatal respiratory distress in the fetus [200].
To conclude, compelling evidence supports that both male biological sex and pregnancy are strong risk factors of severe COVID-19, with many possible action mechanisms, but no compelling evidence on the relative importance of each.
2.4. Comorbidities
2.4.1. Non-Communicable Diseases (NCDs)
Chronic non-communicable diseases often involve the deterioration of one or multiple physiological functions, which might modulate the course of COVID-19. Indeed, many common NCDs, including cardiovascular, chronic respiratory or metabolic diseases, and various types of cancer, have been consistently associated with an increased risk of poor clinical outcomes in SARS-CoV-2-infected individuals (Table 3), while for some other comorbid conditions the evidence is less clear (Table 4).

Table 3.
Prevalent comorbidities with highly supported association with the risk of in-hospital COVID-19 mortality. The sources shown are either meta-analyses that found a significant summary effect, or large clinical studies.

Table 4.
Comorbidities with weakly supported or controversial association with COVID-19 severity.
Chronic obstructive pulmonary disease (COPD) was the third leading cause of death worldwide in 2019 [210], and this common condition has been associated with an increased risk of hospitalization, ICU admission and in-hospital death in COVID-19 [203,211]. The mechanisms potentially involved in the increased severity of COVID-19 in COPD patients are multifold. COPD is characterized by abnormal lung structure, impaired tissue repair, and severe loss of respiratory function [212,213]. Dysfunctional innate and adaptive immune responses [214,215,216], increased chronic inflammation [212,216] and upregulated ACE2 expression [213,216,217,218] have also been hypothesized to contribute to the risk of severe COVID-19. Exacerbation of lung pathology by other respiratory infections is common in COPD patients [219,220,221,222,223,224], suggesting shared mechanisms. It is also important to note that COPD frequently coexists with other NCDs [225,226,227] and its prevalence increases with age [228]. However, the studies controlling for age [3,7] or comorbidities [3] still found a significant independent effect of COPD on COVID-19 outcomes.
Interstitial lung disease (ILD) is a heterogeneous group of chronic conditions involving endothelial (alveolar) injury and fibrosis resulting in impaired gas exchange and limited pulmonary reserve [229,230], and has been associated with an increased risk of severe disease [231], ICU admission [232] or death [232,233,234] in COVID-19. In addition to the direct effect of impaired respiratory functions and resilience, the use of immunosuppressive medications in ILD has also been hypothesized to contribute to the observed increased likelihood of COVID-19 mortality [232].
Contrary to expectations, asthma has failed to show a clear effect on COVID-19 outcomes [235]. Both COPD and asthma are chronic lung diseases involving shortness of breath, chest tightness, wheezing and cough, but the causes and mechanisms of the two conditions are largely different. As opposed to COPD, in allergic (atopic) asthma ACE2 levels are decreased both in upper and lower airways [216,236,237], inflammation is characterized by a type 2 [238] instead of a type 1 response, and T cell levels are not decreased [26,52]. A component of type 2 inflammation, eosinophilia has been shown to be independently associated with decreased severity of COVID-19 [239]. In turn, other respiratory viral infections (influenza, rhinovirus, respiratory syncytial virus) have been associated with the exacerbation of asthma [240,241,242], and impaired IFN signalization has been observed in cells obtained from asthmatic patients upon viral infection [243,244,245]. Asthma is also a risk factor for hospitalization after influenza infection [246]. Fortunately, an aggravating effect has not been detected in SARS-CoV-2-infected patients with atopic asthma. However, non-atopic asthma has been associated with exacerbated symptoms [247,248]. It is characterized by type 1 inflammation (without eosinophilia) and is associated with age and other NCDs [247].
The associated conditions of obesity, diabetes, hypertension and cardiovascular disease (CVD) have all been consistently shown to considerably increase the risk of severe COVID-19 outcomes [201,202]. Given the high worldwide prevalence of these conditions (e.g., 13% of all adults globally have been estimated to be obese in 2020 [249]), they may have had the greatest contribution to COVID-19 mortality among all comorbidities, and may have only been exceeded by the effect of old age overall. In the most likely scenario, obesity, diabetes, hypertension and CVD are all part of the same interconnected pathophysiological pathway [250,251]. The interrelated nature of these chronic diseases is supported by their similarly high prevalence and co-occurrence in MERS and SARS patients [251,252,253]. An individual’s genetic background, existing insulin resistance, dyslipidemia and obesity are main risk factors of diabetes and hypertension, which result in hyperglycemia (which is associated with COVID-19 severity independently from diabetes [254,255,256]), dysregulation of the RAS, heightened immune activation, oxidative stress, and chronic inflammation (IL-1β, IL-6, and TNF-α) [250]. These complex changes give rise to chronic CVD. During COVID-19 pulmonary distress puts an increasing burden on the previously weakened cardiovascular system with a damaged pulmonary endothelial barrier, fluid extravasation, hypoxia, heightened inflammation (possibly through the decreased airway ACE2 levels in CVD patients [216,236,237]) and hypercoagulability, which culminate in possible acute consequences such as myocardial injury, infarction, heart failure, thrombosis or arrythmias [251,257,258,259]. Obesity is further associated with an increased risk of obstructive sleep apnea, asthma, and COPD [260,261,262,263] and adipose tissue seems to be a direct target of SARS-CoV-2 replication further exacerbating hyperglycemia and hyperinsulinemia [264].
Chronic kidney disease (CKD) has been associated with the severity of pneumonia [265], and it is among the strongest risk factors for hospitalization [266] and in-hospital death [201,202,206] with COVID-19. Similarly to cells of the heart and vascular endothelium, it has been proposed that kidney cells might be susceptible to direct SARS-CoV-2 infection [26,27], and infectious virus has been successfully isolated from urine to support this claim [267]. The observed renal damage in COVID-19 can probably be attributed to indirect mechanisms: increased inflammation (which is amplified by CKD), hemodynamic instability, rhabdomyolysis, microthrombi and hypoxia are plausible causes of acute kidney injury (AKI) [268].
The possible contribution of liver diseases to the risk of severe COVID-19 [208,280,281] might be related to the M1 polarization of macrophages causing an increased level of systemic inflammation [386]. However, it has been pointed out that liver diseases often co-occur with other important NCDs, and may not have an independent effect on COVID-19 [387]. Liver injury during COVID-19 might be a result of direct infection [26,388], immune-mediated inflammation [389] and/or antiviral medication use [390,391].
Cancer patients tend to be old, comorbid and immunocompromised in a variety of ways [209,392,393], which may influence COVID-19 outcomes, but certain malignancies are likely to have also a direct impact [394]. The risk of hospitalization and death with COVID-19 have both been associated with cancer [201,208,209,395]. Hematologic [280,281,395,396] and lung cancers [397], and cancers with advanced/metastatic stages [397,398] may have the strongest effect. In addition to the direct effects of cancer, anticancer therapy (chemotherapy, immunotherapy, radiotherapy) might also influence COVID-19 outcomes [394]. However, current limited evidence does not support a strong effect on disease severity [397,399,400].
Both immune-mediated inflammatory diseases [311] (e.g., rheumatoid arthritis [201]) which are characterized by dysfunctional cytokine responses, and the use of immunosuppressant medications [13,280], have been associated with poor COVID-19 outcomes. Special attention has been given to glucocorticoids [319,321,401,402]. While dexamethasone has been demonstrated to reduce mortality with severe/critical COVID-19 [403], it has also been shown that chronic (and especially high-dose) intake of glucocorticoids, and use in mild cases are connected to increased hospitalization and mortality [401]. This effect is likely mediated by the suppression of IFN responses and antimicrobial peptide secretion (causing respiratory dysbiosis) [404,405]. Patients who receive immunosuppressive drugs after organ transplantation are also at a higher risk of severe outcomes during COVID-19 [201,269,280]. Primarily immunocompromised individuals may similarly face poor COVID-19 outcomes (see Section 2.2. Human Genetic Variation); in this group, chronic lung diseases, insufficient vaccine responses and, in the most prevalent subgroup (common variable immunodeficiency), T- and B cell dysfunctions are commonly observed [406].
Some neurological conditions have also been linked to COVID-19 severity [201,280,407]. Possible mechanisms include immunosenescence, heightened IFN responses or genetic predisposition to severe COVID-19 (OAS1, APOE ε4 allele) in Alzheimer’s disease [350,408,409,410], respiratory muscle rigidity and insufficient cough reflex in Parkinson’s disease [350,411], systemic inflammation in epilepsy (with much uncertainty) [362,412] and susceptibility to acute stress in cerebrovascular diseases [413,414]. Increased levels of chronic inflammatory mediators have been observed in several mental disorders (major depressive disorder, bipolar disorder, schizophrenia, and sleeping disorders) [415].
We conclude that some common chronic comorbidities appear to have a strong impact on the risk of severe COVID-19 outcomes, while other conditions have weaker or less clear effect on the course of infection.
2.4.2. Coinfections/Superinfections
Coinfection refers to the simultaneous infection of a host by two or more pathogen species or strains, while superinfection is the acquisition of a second infection after, and in addition to, the first. In both cases, the simultaneous presence of two pathogens can modulate—exacerbate or ameliorate—the effects of either or both. Of note, exacerbating interactions have had a substantial impact on past influenza pandemics, where most deaths were often caused by secondary bacterial infections [416,417,418].
Co- and superinfections are often hard to distinguish in clinical settings (due to limitations in sample collection and pathogen identification, and the lack of clear definitions), and most studies on SARS-CoV-2 did not differentiate between the two scenarios [419]. According to a meta-analysis, in the studies that did distinguish between the two forms of dual infection, the overall rate of superinfections (19–30%) was slightly higher than the prevalence of coinfections (14–25%) [420]; most studies reported coinfections and superinfections among hospitalized cases.
In Table 5, we compiled the results of studies that tested the association between coinfection with specific pathogens and the severity of COVID-19, hypothetical causative mechanisms for the associations, and potential mechanisms by which a pre-existing infection might facilitate the acquisition of SARS-CoV-2 (or vice versa). Data about coinfections were not available beyond case reports or case series for dengue virus [421], certain human herpesviruses [422] (cytomegalovirus [423], Epstein–Barr virus, human herpesvirus 6), fungi causing mucormycosis [424] and Pneumocystis jirovecii [425], and we therefore did not include detailed information on these pathogens in the table. We also note that the effect of some pathogens on COVID-19 severity is highly debated and may depend on the severity and the degree of clinical control of the coinfections.

Table 5.
Co- and superinfections tested for associations with COVID-19 severity.
Infection by SARS-CoV-2 might also facilitate co- or superinfections with some pathogens. Enhanced adherence to infected cell lines [530,531,532], reduced ciliary function and clearance [533,534], altered mucus secretion (goblet cell hyperplasia) [535,536], reduced oxygen exchange [537,538,539], virus-induced [533] and immune-mediated (e.g., by NETs) [540] cytotoxic airway damage, disruption of innate immunity followed by hyperinflammation [541], immunosuppressive effects of platelet activation [542], decreased levels of adaptive immune cells [35,543] and induced microbiota dysbiosis (both respiratory and gastrointestinal through the gut–lung axis) [426,544] are all possible mechanisms. In turn, some other acute infections might promote superinfection with SARS-CoV-2 through similar mechanisms; however, we are not aware of any studies designed to test this effect. Finally, in the context of long-term chronic infections, superinfection with SARS-CoV-2 (over a pre-existing condition) may be more likely than acquiring the other pathogen during the brief course of a COVID-19 episode. Uncontrolled HIV infection, in particular, appears to increase susceptibility to SARS-CoV-2 [468], and to promote persistent COVID-19 in some patients [545], by suppressing efficient immunity. Certain chronic coinfections predispose to NCDs as well (see Section 5.1. Interactions Between Risk Factors of Severe COVID-19). Furthermore, SARS-CoV-2 infection has been reportedly connected to the reactivation of latent hepatitis virus [546] and Mycobacterium tuberculosis infections [442,547].
Recent meta-analyses estimate the prevalence of bacterial co- or superinfections at approximately 15–20% among hospitalized cases [548,549] with an even higher prevalence in severe cases [430,549,550,551]. While (community-acquired) bacterial coinfections seem to be relatively rare (3–8%) [420,430,550] even compared to RSV or influenza virus patients [434], (hospital-acquired) secondary infections are quite common (14–24%) [419,420,550]. At hospital admission, Klebsiella pneumoniae, Streptococcus pneumoniae, Staphylococcus aureus and Haemophilus influenzae were the most frequently detected coinfecting species, while superinfections in the hospital were typically caused by Acinetobacter spp., Pseudomonas aeruginosa, Escherichia coli, Staphylococcus aureus and Klebsiella spp. [420,430]. Unfortunately, most studies of bacterial superinfections had low sample sizes, which resulted in very wide margins for the effect sizes and indicates a low certainty of demonstrated effects.
The overall prevalence of viral infections detected concurrently or following COVID-19 is approximately 7–12% [548,549,552] with a higher rate of coinfections (5–10%) [420,553] compared to superinfections (~4%) [420]. The probability of acquiring a viral superinfection during COVID-19 may be lower than observed in the general population [554], and there were no differences found in prevalence between cohorts of severe and mild cases [549]. Common coinfecting viruses are influenza viruses, RSV and enteroviruses (particularly, rhinoviruses) [420,553]. Epstein–Barr virus (EBV), human herpesviruses (HHV), seasonal human coronaviruses (HCoV), adenoviruses and human metapneumovirus (HMPV) were also reported in several studies [490,549,552,555,556]. The overall effect of viral co/superinfections on the severity of COVID-19 is unclear [550,553,556,557], but the common phenomenon of viral interference [558], and differences between the relative proportion and effect of specific pathogens in the studies may be responsible for these mixed results.
Fungal superinfections are more common (4–13%) [420] than coinfections (2–7%) [420] and both are connected to increased mortality [431,559]. Invasive pulmonary aspergillosis (Aspergillus spp.) is dominantly present [420,549,551], especially among coinfections [420], while invasive candidiasis (Candida spp.) is the most frequent complication among superinfected cases [420]. Mucormycosis (Rhizopus spp., Mucor spp.etc.) [424] and pneumocystis pneumonia (Pneumocystis jirovecii) [425,560] have also been reported repeatedly. Individuals with DM or patients receiving corticosteroid treatment have a higher risk of severe SARS-CoV-2–fungal coinfections (aspergillosis and mucormycosis) [509,561,562].
The prevalence of parasitic coinfections is strongly heterogeneous geographically and has not been assessed systematically in COVID-19 patients. In an Ethiopian cohort study, Entamoeba spp. (~20%) and Giardia spp. (~4%) were common parasitic protozoa, while Hymenolepis nana (~17%), Schistosoma mansoni (~5%) and Ascaris lumbricoides (~4%) were commonly identified helminth species [519]. In Egypt, Toxoplasma gondii (~22%), Cryptosporidium spp. (~20%), Blastocystis spp. (~17.6%) and Giardia spp. (~9%) were reported [563]. In Sub-Saharan Africa and South Asia, a meta-analysis estimated the overall prevalence of Plasmodium spp. approximately 11% among COVID-19 patients [564]. In Brazil, COVID-19 patients coinfected with Chagas disease (Trypanosoma spp.) were rare (~0.4%) [528].
Finally, we note that in the study of associations between co/superinfections and COVID-19 severity, causality is often hard to establish. In particular, individuals with coinfections might be more likely to have weakened immunity or general health, which would imply a hidden shared common cause for the coinfection and subsequent severe COVID-19, rather than direct causality.
2.5. Frailty
Frailty is a medical condition affecting multiple organ systems, characterized by reduced strength and endurance, impaired physiologic and immunological functions, and a reduced ability to combat acute stressors, leading to increased dependency and/or death [565,566,567]. The prevalence of frailty is estimated to be approximately 5–9% worldwide over the age of 50, but it varies considerably by demographic variables (e.g., age and sex) and also geographically (higher in low- or middle-income countries in spite of younger populations) [568], with notably higher rates among nursing home residents (~50% over the age of 60) [569] compared to community-dwelling individuals (~10% among people older than 65) [570]. Importantly, frailty is strongly linked to, but is not equivalent to, aging, and it is typically quantified by the Clinical Frailty Scale (CFS) from 1 (very fit) to 9 (terminally ill), which is a composite marker based on a clinical assessment of physical ability, comorbidity, cognitive impairment, and disability [571].
The effect of frailty on in-hospital COVID-19 mortality has been demonstrated in large clinical studies [572,573,574,575], and confirmed by meta-analyses [576,577]. Mildly frail patients (CFS 4–5) had increased risk of severe outcomes compared to fit individuals (CFS 1–3) in several (but not all) clinical studies [573,576]; severe frailty (CFS 6–9) showed a correlation with severe disease consistently and with a greater effect size [572,573,578]. COVID-19 mortality is gradually increasing with the CFS even when controlling for age and sex [579,580]. A similar relationship has been observed with all-cause mortality and different frailty measures as well [581,582,583].
The development of frailty has been linked to chronic inflammation, which is a major factor in the pathogenesis of severe COVID-19 [584,585]. Both conditions share inflammatory and immunological biomarkers (IL-6, CRP, LDH, PCT, and cortisol) [584] and lead to heightened coagulation, development of sarcopenia and decline in multi-system function [586].
In summary, although frailty syndrome is a biologically overlapping condition with aging and multimorbidity, it appears to also have an independent effect on the outcome of COVID-19, and it is an important predictor of disease severity upon hospital admission.
2.6. Microbiota
SARS-CoV-2 infection is often accompanied by respiratory and intestinal dysbiosis with characteristic patterns that are distinguishable from those induced by influenza infection [587] or other forms of community-acquired pneumonia [426,588]. While most of these changes can attributed to the presence of SARS-CoV-2 and host–virus interactions, some evidence supports a bidirectional relationship between the composition of the microbiota and COVID-19 disease severity, implying that pre-infection variation in the microbiota might influence clinical outcomes. Gastrointestinal microbiota composition has a complex relationship with several COVID-19 risk factors, such as age, NCDs, lifestyle and the frailty syndrome (for details see Section 5.1. Interactions Between Risk Factors of Severe COVID-19), indicating at least a mediator role in the determination of severe COVID-19.
Probiotics inhibit the growth of pathogenic microorganisms [589], enhance immune responses [590], and have been effectively used in the treatment of metabolic diseases (obesity and DM) [591]. As a therapeutic option in the management of COVID-19 probiotic treatments have yielded limited and mixed results [592,593]. Antibiotics constrain the growth of opportunistic pathogens and can prevent secondary infections, but they also perturb the healthy microbiota. This disruption of the microbial communities might have an impact on COVID-19 severity. Individuals with repeated recent exposure to antibiotic treatment have been shown to have increased severity of disease [594,595], although this might have a causal relationship with the original cause that necessitated treatment, rather than with the treatment itself. Those who received antibiotics during early (non-severe) stages of COVID-19 subsequently had increased severity and stayed longer in hospital [596], but did not have increased mortality [596]. Altogether, several lines of indirect evidence indicate that the composition and diversity of the human microbiota might play a role in the determination of COVID-19 severity, possibly independently from other COVID-19 risk factors.
The effect of the microbiota on clinical outcomes is likely to be mediated by its response to SARS-CoV-2 infection. In addition to other important functions, the microbiota are responsible for promoting and maintaining a stable and immunologically stimulating environment both in the respiratory and gastrointestinal tracts [597,598]. The disruption of the microbiota involves several processes that might contribute to COVID-19 pathogenesis; conversely, microbiota that is resilient to the effects of COVID-19 may have a protective effect.
SARS-CoV-2 replication initiates in the URT and continues in the alveoli of the lungs influencing the local immunological environment. This causes characteristic alterations in the diversity and composition of the microbiota in the oral cavity [599,600] and the upper [426,601] and lower respiratory tract (LRT) [602,603]. The diversity of the microbiota typically increases in the URT [604,605,606] due to the emergence of opportunistic pathogens (Klebsiella, Streptococcus, Veillonella, Prevotella, Enterococcus, Rothia, etc.). Consequently, local inflammation increases in the oral cavity (IL-6, IL-17) [605], the URT (IL-6, IL-8, IL-1β) [607] and the lungs (white blood cell (WBC) and lymphocyte (LYM) counts) [544] together with changes in the host metabolic profile in the RT (reduced protein, lipid and glycan metabolism, induced nucleotide and amino acid biosynthesis and carbohydrate metabolism) [588,606].
Pro-inflammatory cytokines from the lungs are transferred to the circulatory system [608], which then induces changes in the gastrointestinal microbiota (gut–lung axis) as has been shown in influenza infections [609,610,611]. This connection is confirmed by the fact that COVID-19 is often accompanied by GI symptoms [612]. These affect approximately 5–10% of patients (most often anorexia, nausea, vomiting, and diarrhea) [613,614], and might persist long after viral clearance [615]. As in the RT, the proportion of anti-inflammatory probiotic and beneficial commensal bacteria (Lactobacillus, Bifidobacterium, Eubacterium, Faecalibacterium, Roseburia, Lachnospiraceae, etc.) typically decreases, while opportunistic pathogens and pro-inflammatory species (Streptococcus, Veillonella, Actinomyces, Clostridium, Bacteroides, etc.) expand in the GIT during COVID-19 [616]. Contrary to the RT, microbial diversity declines in the intestines [587,617] due to the depletion of rich commensal communities. Unique alterations occur to the fungal (higher levels of Candida, Aspergillus, Auris, etc.) [600,617], viral [602,606] and archaeal [606] communities as well both in the RT and the GIT. The microbial diversity in most cases quickly returns to normal values after recovery [618] and the magnitude of dysbiosis is proportional to certain immunological and metabolic signatures of COVID-19 and severity of disease [544,619,620].
In line with compositional changes in the microbiota, altered biosynthetic and metabolic pathways, including more intense vitamin B12 and urea production along with impaired short chain fatty acid (SCFA), L-isoleucine, tryptophan and polyamine biosynthesis and sulfur oxidation were typical in SARS-CoV-2-infected patients [618,621,622,623,624]. Some of these metabolic signatures seem to show sexual dimorphism in COVID-19 patients compared to uninfected controls [625]. Rise in inflammatory molecules in the GIT has also been shown during COVID-19 (i.e., CRP, PCT, D-dimer, LDH, AAT, and GGT) [587,626]. In particular, fecal butyrate levels were found to be negatively associated with some of these biomarkers (IL-10, CXCL-10, and CRP), similarly to L-isoleucine (CXCL-10) [622], SCFAs [627] and derivatives of bile acids [628], which have broad effects on the immune system. Butyrate in particular has important roles in the induction of Treg cells [629,630], it reduces several pro-inflammatory pathways [631], participates in the secretion of mucins and defensins [629] and helps to maintain the intestinal barrier [632]. Causal connections between beneficial and harmful microbes, inflammation and metabolic responses have been established by multi-omics analyses [620,633].
Local inflammation and dysbiosis damage the integrity and increase the permeability of the intestinal barrier [623,634], which might facilitate the infection of intestinal epithelial cells by SARS-CoV-2 [635]. In turn, direct infection and viral replication might exacerbate dysbiosis in different ways. First, decline in intestinal ACE2 levels by infection and/or the loss of beneficial bacteria [636] might aggravate inflammation through the RAS [637]. As an alternative mechanism, reduction in ACE2 might also downregulate the amino acid transporter B0AT1 (heterodimer formation with ACE2 [638]), which is responsible for tryptophan absorption [639]. With reduced levels of tryptophan the secretion of antimicrobial peptides decreases [639,640], which in turn aggravates dysbiosis. Severe dysbiosis and increased intestinal permeability might lead to the translocation of pathogens, toxins and cytokines to the circulatory system leading to severe complications and multi-organ failure [641]. The causal role of COVID-19-induced gastrointestinal dysbiosis in the development of symptoms and disease severity is further supported by results obtained with a gnotobiotic mouse model [642]. Fecal microbiota transplantation from COVID-19 patients to germ-free mice resulted in lung histopathology, an inflammatory cytokine profile, cognitive impairment, and increased susceptibility towards bacterial infection in the animal model indicating that pre-infection differences in microbiota composition might influence COVID-19 susceptibility and severity as well. Further studies on mice suggest that gut dysbiosis also damages the blood–brain barrier [643,644], induces neuroinflammation [644,645] and facilitates direct neuroinvasion by SARS-CoV-2 [646].
To conclude, microbial dysbiosis is a characteristic trait of SARS-CoV-2 infection that may be both cause and consequence in the pathogenesis of COVID-19. Multiple factors associated with an altered microbiota have been connected to COVID-19 severity. However, further studies are needed to explore the causal relationships between the microbiota and COVID-19 pathogenesis, controlling for the interrelated effects of age, lifestyle, and comorbidities.
2.7. Immunological History
2.7.1. Previous SARS-CoV-2 Infection
SARS-CoV-2 infection elicits both cellular and humoral immunity, which strongly reduces the risk of severe clinical outcome in subsequent re-infections, and provides partial protection against re-infection (reviewed in [647,648]).
Following successful immunization, B cells (and antibodies) are thought to be responsible for SARS-CoV-2 inoculum neutralization, early control and inhibition of viral replication, while T cells are mainly the agents of cellular control of infection in addition to their role in the coordination of immune responses [648]. For this reason, humoral immunity might be effective against both reinfection and severity of disease, while cellular immunity mainly reduces severity of COVID-19 [649]. Regarding the molecular targets, cellular responses target mainly structural proteins (S, M, N) of SARS-CoV-2, but some CD4+ and CD8+ T cells recognize accessory and non-structural proteins as well [650]. Similarly, antibodies mainly target epitopes on the S and N proteins as potential targets of neutralization [651].
Following SARS-CoV-2 infection the diversity and affinity of antibodies keeps increasing for several months [652] along with the level of memory B cells [648], but the levels of most immune components (IgG, IgA, and T cells) decline exponentially even in the first month post infection [648]. In the absence of a new variant with substantial immune evasion capabilities (such as Omicron variants), natural immunity might retain its protective effect for 8–12 months against reinfection [647,652,653,654,655] and probably longer against severe manifestations [648]. This is consistent with the observation that relatively low antibody titers show 50% protective effect against symptomatic and severe disease (14.4–28.4% and 0.71–13% of the initial magnitude, respectively) [656]. In addition to immunoglobulin levels, the persistence of peripheral SARS-CoV-2-specific CD4+ and CD8+ T cells is also a determinant of effective immune protection against reinfection and disease control [657,658].
While a detailed discussion of the impact of vaccinations (and other medical interventions) on COVID-19 outcomes goes beyond the scope of this review, we note that there are some differences between vaccination and natural infection in the presentation of antigens and the qualities of the developing immune memory. While vaccination induces systemic immunity, and the most widely used vaccines elicit only spike-specific humoral and cellular immunity, natural infection generates immunity against all viral proteins, and induces tissue-specific (e.g., mucosal) responses as well [648]. However, some vaccines evoke higher IgG levels compared to natural infection (regardless of severity) [659,660]. Short-term immunity after infection and vaccination might be similarly effective [661], but the duration of protection seem to be significantly shorter in the latter case (~6 months for reinfections [647] compared to the previously mentioned 8–12 months following SARS-CoV-2 infection), possibly due to the more rapid decay of antibody titers [659]. This might also explain why in real-world settings reinfection results in less severe disease compared to COVID-19 following vaccination [662]. Nonetheless, this distinction is probably losing its importance as the pandemic progresses and increasing numbers of individuals accumulate exposure to both vaccination and infection, developing ‘hybrid immunity’. It has been shown repeatedly that individuals with hybrid immunity can acquire stronger neutralizing antibody levels compared to individuals with vaccination induced or natural immunity alone [659,663].
In the first two years of the pandemic, reinfections were rare (<2% of followed cases) [664,665,666] confirming the protective effect of specific immunity. Additionally, the severity of reinfections were significantly lower compared to primary infections (aOR = 0.10 (0.03–0.25) [667], aOR = 0.39 (0.35–0.44) [668]). At the end of 2021, with the rise of the Omicron variant, which showed similar transmissibility but more effective immune evasion compared to previous variants of concern (VOCs) [669], the rate of reinfections rose significantly [670,671,672]. However, those individuals who had had prior immunity were less infectious compared to immunologically naïve individuals [673] indicating lower viral burden and better disease control. Then, in 2022, repeated waves of Omicron subvariants were consistently characterized by the lack of evidence for (re-)increased severity of disease compared with the preceding wave [674,675], which indicates long-lasting protection against severe disease, while immune escape mutations appear to be able to erode protection against re-infection rapidly.
COVID-19 disease severity is modulated by additional factors even in the presence of immunological protection resulting from previous SARS-CoV-2 infection(s). Advanced age [668,676,677], the presence of comorbidities [676,677,678,679] and male sex [668] have an exacerbating effect on re-infections similar to first infections. Severe primary infection predicts higher risk of severe symptoms in re-infections [676,677], even though more severe primary infection appears to elicit higher levels of memory B cells [680] and antibodies [680,681,682,683], and stronger T cell responses [650]. A higher severity of reinfections is associated also with markers of dampened immune protection (low avidity IgG [684], and longer time between infections [678]).
In summary, previous SARS-CoV-2 infection provides substantial protection against severe COVID-19 in subsequent re-infections, possibly modulated by, but largely independent of other risk factors, and this protection appears to last longer and be more robust to viral evolution than the protection against re-infection.
2.7.2. Cross-Reactive Immunity from Other Infections
Some evidence indicates that immune responses elicited by previous, non-SARS-CoV-2 infections might also influence the outcome of COVID-19, if pre-existing immune responses can cross-react to SARS-CoV-2 epitopes. Based on sequence similarity, cross-reactive responses to SARS-CoV-2 are most likely to involve pre-existing immunity to other human coronaviruses (HCoV), of which two betacoronaviruses (HCoV-229E, HCoV-NL63) and two alphacoronaviruses (HCoV-OC43, HCoV-HKU1) are responsible for an estimated 10–15% of common cold episodes [685]. However, pre-existing cross-reactive immunity to SARS-CoV-2 may not be entirely explained by previous exposure to HCoVs [686,687], and may include responses to unrelated infections such as influenza or cytomegalovirus infections as well [688].
Of the distinct arms of adaptive immunity, CD4+ T cell responses against epitopes conserved across SARS-CoV-2 and other coronaviruses are present in COVID-19 convalescent patients [650,689,690,691] and more importantly, often in unexposed healthy individuals [689,692,693,694,695] as well, with proportions up to 50–80% in some of the unexposed populations analyzed [650,690,696]. Antigenic targets consist of structural (S, N, M) [650,689,690,695], non-structural [650,690] and accessory proteins [650,690]. These pre-existing T cells seem to cross-react with both SARS-CoV [650] and all four common cold coronaviruses [692]. Preexisting CD8+ T cells showed similar cross-reactivity [688,697,698,699,700]. While a correlation between the level of pre-existing cross-reactive T cells and clinical severity during COVID-19 has (to our knowledge) not been tested directly yet, potential effects on the course of the disease include the priming of protective immunity [701] and contributing to an early control of viral replication [702]. However, these T cells often have low avidity against SARS-CoV-2 antigens, and de novo immune responses are probably required for effective control of SARS-CoV-2 replication [703].
Humoral immune responses from previous infections might also display cross-reactivity against SARS-CoV-2. SARS-CoV-2 cross-reacting antibodies have been detected in several [704,705,706,707], but not all [708,709,710] studies that analyzed sera obtained or intravenous immunoglobulin manufactured before the pandemic. Some studies found cross-reactive antibodies in only a small percentage of individuals [27,28], implying that genuine variability might have existed within or between populations, possibly related to differences in infection history. The most likely source of SARS-CoV-2 cross-reactive antibodies are HCoV-specific memory B cells [711]. The molecular targets are mainly the S [704,705,711,712,713] and N proteins [694,704,705,713,714], but cross-recognition of conserved non-structural proteins [704] has also been reported. These antibodies recognize SARS and MERS coronaviruses [712] along with seasonal HCoV antigens [694,711,712,714,715,716], indicating broad cross-reactivity. Due to closer relatedness, the probability to cross-react to SARS-CoV-2 might be higher for immune responses that had been elicited against HCoVs belonging to the betacoronaviruses compared to those that targeted earlier alphacoronavirus infections. However, this prediction has not been tested by a systematic comparison, and while some studies suggest a primary role of betacoronavirus cross-immunity [711,712], others do not appear to support the hypothesis [694,707,713,716]. In addition, in COVID-19 convalescent individuals the levels of seasonal HCoV cross-reacting antibodies are typically also boosted, which provides further support for cross-reactivity. However, while some studies demonstrated this effect for antibody levels against all seasonal HCoVs [717,718,719], in others the effect was limited to titers against betacoronaviruses [720,721,722,723] (especially HCoV-OC43 [707,724,725,726,727]), or surprisingly, to alphacoronaviruses only [728,729]. Independent of the level of antibodies, SARS-CoV-2-specific IgG and IgM were shown to robustly recognize betacoronavirus antigens [712,715,730].
Several studies have investigated the potential impact of pre-existing HCoV-specific antibodies on COVID-19 severity; unfortunately, the conflicting results do not allow a firm conclusion. Some analyses demonstrated a correlation between HCoV-specific antibody titers and milder COVID-19 outcomes [718,728,729,731,732,733,734,735], other studies found no significant effect [709,736,737], and some studies reported a correlation between cross-reactive antibody titers and more severe COVID-19 outcomes [707,720,722,723,726,738]. A beneficial effect might be explained by effective cross-neutralization or the priming of effective humoral immune responses, while a negative effect might arise from low-avidity cross-reactive immunoglobulins hindering the production of high-avidity SARS-CoV-2-specific antibodies. Antibody-dependent enhancement of SARS-CoV-2 has also been proposed [739], but not confirmed.
It must be noted that cross-reactivity does not necessarily, or even typically, imply cross-neutralization against SARS-CoV-2. Neutralizing antibodies have been described in some studies [740,741,742], but most analyses have failed to demonstrate neutralization activity [707,709,722,743,744,745].
We conclude that while the existence of cross-reactive immunity to SARS-CoV-2 has been demonstrated by several studies, the impact of this immunity on COVID-19 outcomes remains largely hypothetical. Finally, we note that any impact of cross-reactive immunity to other infections is likely to have been restricted to the first waves of the pandemic, and immunity to SARS-CoV-2 in subsequent waves has probably been dominated by specific immunity from previous episodes of COVID-19.
2.8. Lifestyle
2.8.1. Physical Activity
Regular physical activity appears to have a beneficial effect on the outcome of COVID-19. Individuals living a sedentary lifestyle are exposed to a higher risk of COVID-19 hospitalization (OR = 2.26 (1.81–2.83)) [746], ICU admission (OR = 1.73 (1.18–2.55)) [746], severe disease [747] and mortality (OR = 2.49 (1.33–4.67)) [746,747,748,749] compared to individuals who exercise regularly. In detailed analyses, higher metabolic equivalent of task per week was associated with a lower risk of COVID-19 hospitalization [750], severe disease [747] and mortality [747,749]. Similarly, cardiorespiratory fitness was also correlated with the severity of disease [751] and the risk of death [752]. The beneficial effects of physical training seem to be long-lasting. Among male military conscripts, high cardiorespiratory fitness and muscle strength in late adolescence and early adulthood proved to be protective against the adverse effects of COVID-19 decades later [753].
Regular physical activity has broad effects on human metabolism and the immune system that might be protective against severe COVID-19. An active lifestyle has been linked to lower incidence and severity of URT viral infections (e.g., influenza) [754,755] in humans, and to attenuated inflammation following bacterial infection in mice [756]. Both aerobic and muscle strength training stimulate the release of myokines (e.g., myostatin, IL-6, IL-15, and LIF) [757], which in the long term counteract low-grade chronic inflammation [758]. Exercising can also boost innate [759] and adaptive immune responses [760,761,762], and helps to maintain local tissue immunity [763] (e.g., in the lungs [756]) and to delay immunosenescence [764]. In addition to immunological functions, regular exercising helps to slow down the deterioration of frailty by preserving muscle [765] and respiratory function [766], and prevents body fat accumulation [767] and the development of CVD [768]. Loss of adipose tissue lowers the leptin/adiponectin ratio and hence, chronic inflammation [769].
The impact of physical activity on COVID-19 may thus be mediated by its effects on immunity, comorbidities, and frailty. For further details on the effects of physical activity on COVID-19 pathogenesis, we recommend reading the review by Filgueira et al. [770].
2.8.2. Alcohol Consumption
High consumption of alcohol has been linked to adverse health effects, including an exacerbation of Mycobacterium tuberculosis infection and other sources of ARDS [771,772]. While some early reports failed to find a significant effect of alcohol consumption on COVID-19 severity [773,774,775], excess alcohol intake has since been repeatedly associated with worse clinical outcomes, such as severe disease [776], ARDS [777] and death [778]. A latent causal variable analysis that considered also genetic correlations using GWAS data also confirmed the link between alcohol consumption and severe COVID-19 [779]. Furthermore, as with many other risk factors, alcohol consumption has been shown to correlate with the level of proinflammatory biomarkers (e.g., CRP and NLR) [778] and proinflammatory cytokines (IL-1β, IL-6, and TNF-α) [780,781]. It causes oxidative stress [782], impacts the activity of alveolar macrophages [781,782], T lymphocyte proliferation and turnover [781] and the number and function of NK cells [783]. Alcohol use in the long term also has an impact on the endothelial cilia and respiratory clearance [784]. Alcohol-related liver disease has also been shown to increase the risk of COVID-19 mortality [310].
2.8.3. Smoking
The consumption of combusted tobacco products has long been known to have a detrimental effect on lung function and health. In accordance, smoking might induce more severe lung inflammation and respiratory distress during COVID-19, similarly to influenza virus infections [785]. However, the results of association studies in the case of COVID-19 have been somewhat controversial. Some analyses that considered smoking habits [786] and a genetically predicted tendency to smoke [787,788] have been linked to more adverse outcomes in COVID-19 patients. However, the frequency and duration of smoking, or time since quitting tobacco use also influence the increased risk posed by this habit. The long-term damaging effects of smoking have been studied by comparing never-smokers to former [204,789,790,791,792] or ever-smokers (former and current smokers combined) [124,204,793,794,795], which consistently showed more severe outcomes (e.g., hospitalization, ICU admission, MV and death) in the latter groups. However, although current smokers compared to (current) non-smokers seem to be more prone to experience severe symptoms and death [123,794,796,797], when comparing current smokers to never-smokers, some publications reported increased severity [204,788,796,798], while others no effect [791,792,799], and surprisingly, recent cohort studies found lower rates of severe outcomes [789,790] in the smoker group of the study. The short-term effects of tobacco use are harder to measure in clinical settings, but ambiguous results on the effect of current smoking behavior on COVID-19 might imply further physiological and immunological effects that differ from the mechanisms responsible for long-lasting damage in the lungs.
Several components of combusted tobacco products show immunomodulatory (e.g., polycyclic aromatic hydrocarbons (PAHs), acrolein, and CO) and/or harmful effect (e.g., volatile organic compounds, metals, oxidants, and nicotine) on human health [800]. These facilitate the development of chronic lung disease [801], CVD [802] and DM [803], which might confound the estimation of the direct effect of smoking if not controlled for properly. Possible direct effects of smoking on COVID-19 severity include impairment of mucociliary clearance [804], increased epithelial permeability [805], immune suppression [806] (IFN responses in particular [807]), elevated oxidative stress, inflammation and vascular injury [808,809]. Potential mechanisms proposed to explain a beneficial effect of current smoking demonstrated in some studies include the modulation of the RAS by increased ACE2 levels in the lungs of smokers [810,811], increased NO inhibiting viral replication [812], or inhibition of pro-inflammatory cytokine secretion [813,814] by nicotine.
We conclude that the effect of smoking on COVID-19 appears to be complicated and requires further study. The review by Benowitz et al. [800] provides a more detailed discussion of the topic.
2.8.4. Diet and Nutrition
Dietary habits influence metabolism and the risk of developing chronic diseases. Malnutrition can result in a wide range of unfavorable health effects [815], including impaired immunity [816], and is independently associated with mortality among older adults [817]. COVID-19 patients who experienced malnutrition either years before SARS-CoV-2 infection [818,819] or during COVID-19 [820,821] were more likely to experience adverse outcomes, such as prolonged hospitalization, MV or mortality. Among those considered well nourished, food choice may still have some influence on the severity of COVID-19. Some studies have reported that high-quality (defined by multiple measures) [822], vegetarian [823] or plant-based [824,825] diets were associated with more favorable disease outcomes. Adherence to the Mediterranean diet (high in fruits, vegetables, legumes, olive oil, and whole grains; low intake of processed foods and red meat), which involves a high intake of antioxidants [826], boosts immunity and reduces inflammation [827], was also associated with less severe COVID-19 in small cohorts [828,829]. Preliminary evidence supports that other anti-inflammatory dietary patterns such as intermittent fasting [830] and ketogenic diet [831,832] might be beneficial.
No single food item has been unambiguously connected to COVID-19 severity [833]. However, certain metabolic biomarkers, including essential nutrients, are useful biomarkers of COVID-19 outcome [834]. Current research focuses mostly on two nutrients, omega-3 polyunsaturated fatty acids (PUFAs) and vitamin D.
Although sample sizes were small, some studies indicate that regular consumption of omega-3 PUFAs might be favorable against severe COVID-19. Low omega-3 index [835] and omega-3 PUFA deficiency [836] have both been associated with adverse outcomes, such as MV and mortality. Similarly, omega-3 PUFA supplementation has been connected to milder symptoms [837,838], and omega-3 supplementation had previously been shown to improve symptoms in ARDS patients [839]. These effects might be mediated by the role of omega-3 PUFAs in the enzymatic conversion of specialized pro-resolving mediators (SPMs) [840]. SPMs play an important role in the termination of inflammatory reactions by preventing the infiltration of phagocytes, enhancing the removal of apoptotic cells and debris, inhibiting cytokine production and removing inflammatory mediators [841]. Omega-3 supplementation increases SPM levels [842], which also correlate with mild COVID-19 [843]. Lastly, certain PUFAs (such as the omega-3 eicosapentaenoic acid) have been shown to interfere with the binding of the SARS-CoV-2 RBD to hACE2, and to TMPRSS2 and CTSL in vitro [844].
The active form of vitamin D, 1,25(OH)2D has important functions in innate antiviral immunity against several respiratory viruses, such as rhinoviruses [845], RSV [846] and influenza viruses [847]. 1,25(OH)2D induces the production of antiviral effector molecules, enhances the activity of innate immune cells and Tregs, and also lowers TNFα, Th1 and Th17 cell levels, and the ACE2:ACE ratio [848,849]. In COVID-19, some studies have found an association between lower levels of the major circulating form of vitamin D, 25(OH)D, and an increased severity of COVID-19 [850,851,852]. However, as has been pointed out elsewhere [848], these results may indicate either worse COVID-19 outcomes due to low initial 25(OH)D levels [848], or, conversely, they might reflect relevant metabolic changes as a result of more severe disease [853]. A recent review by Martineau and Cantorna [848] reported that out of 11 randomized controlled trials that had been published at the time of writing, only 4 reported significant associations. Two recent large studies and a systematic review on vitamin D supplementation reported lower severity and mortality in the treated groups compared to controls [854,855,856], a study using Mendelian randomization found no association between genetically predicted 25(OH)D levels and COVID-19 severity [857], and another genetic analysis demonstrated an association between a risk score constructed from several genetic variants that influence vitamin D pathways and with 25(OH)D levels, and between 25(OH)D levels with disease outcome [858]. Recently, a mechanistic link has been proposed between vitamin D supplementation and reduced COVID-19 severity through the increased expression of interferon stimulating genes and higher protein levels both in vitro and in vivo [859].
We conclude that more studies are needed to clarify the importance and magnitude of the effect of Omega-3 PUFAs and vitamin D on COVID-19. For further details, we recommend reading the reviews by Mazidimoradi et al. [836], and by Martineau and Cantorna [848], respectively.
3. Viral Factors
3.1. Viral Genetic Variation
Viral genetic factors have long been known to influence the outcome of infection in other well-studied viral epidemics [860], and it has quickly become clear that the emerging variants of SARS-CoV-2 responsible for successive waves of the epidemic (designated Variants of Concern, VOCs) can influence not just the transmissibility of the virus, but the severity of COVID-19 as well (Table 6). Because the risk of severe outcome is influenced by multiple factors that can differ between countries and even between successive waves, risk ratios of severe disease could be reliably estimated when two variants were simultaneously present (typically: one replacing the other) in the same country or region.

Table 6.
Clinical studies investigating the impact of SARS-CoV-2 VOCs on COVID-19 disease severity compared to non-VOC or previous VOC.
The first marked effect was observed when the Alpha VOC (Pango designation: B.1.1.7) replaced basal non-VOC variants in the UK in late 2020, then spread to become the dominant lineage worldwide in the first half of 2021. Two large studies using S gene target failure as a proxy for Alpha infection estimated that the risk of hospitalization and death was approximately 64% and 55% higher in Alpha, compared with non-VOC infections [861,862].
Limited data are available on the severity of infections caused by the Beta (Pango: B.1.351) and Gamma (Pango: P.1) VOCs that had a geographically more limited spread. One study found that both Beta and Gamma infections were associated with increased number of hospitalizations and ICU admissions but not with deaths [864]. Another analysis estimated that the odds of death were approximately 1.5-fold higher in infections with the Beta variant compared with the Alpha variant [865]. The Lambda VOC (Pango: C37) fueled a particularly deadly epidemic wave in several countries of South America [873]; however, it is unclear whether or to what extent an increased risk of severe outcome with this VOC might have contributed to the high population-level mortality.
The Delta VOC (Pango: B.1.617.2; dominant lineage worldwide between July and December 2021) seems to have had further increased virulence compared to the first group of successful VOCs (Alpha, Beta and Gamma) that all shared the N501Y mutation in the spike protein. In a cohort of 212,326 patients, individuals infected with the Delta variant (identified by high-probability inference) had an increased chance of hospitalization, admission to ICU, or death, compared with N501Y-positive patients [863]. The probability of hospitalization after a positive test was higher for Delta compared to Alpha infections in two other cohorts [866,867].
After the Delta wave, several lineages of the Omicron VOC (BA.1, BA.2, BA.4, BA.5) were responsible for the next global waves of the pandemic. Analyses of the first Omicron wave (BA.1) reported significantly reduced risk of severe disease compared with the previously circulating Delta variant. Initial results suggested that after Omicron infection the risk of hospital admission decreased by 50–75% and mortality by 80–90% [868,869]. However, it has been pointed out that part of the observed reduction in the severity of COVID-19 can be attributed to the increased ability of the Omicron variants to infect individuals with preexisting immunity, which provides partial protection against severe COVID-19 and death [874]. Analyses taking into account documented antigen exposure and under-ascertainment of prior infections reported a reduction of approximately 30% in the probability of hospitalization, compared with the preceding Delta variant [675,870]. Omicron’s reduced ability to replicate effectively in TMPRSS2 expressing cells such as alveolar tissue (which is the main anatomical site of COVID-19 pathogenesis) and its effective entry into cells of the upper respiratory tract [875,876] (mainly through the cathepsin-L pathway) might explain this reduction in disease severity in immunologically naïve individuals. However, a chimeric recombinant SARS-CoV-2 encoding the S gene of Omicron in the backbone of an ancestral SARS-CoV-2 isolate proved to be highly lethal (similarly to the basal lineage of the virus) in K18-hACE2 mice that experience only mild symptoms when infected with Omicron [877]. This observation suggests that the reduced pathogenicity of Omicron might depend also on changes outside of the spike protein.
Importantly, the population-level impact of Omicron has been shaped by both its reduced per capita risk of mortality and severe disease, and by its increased transmissibility [878], particularly among those with pre-existing immunity, which resulted in higher total case counts. Most studies so far indicate that both BA.2 [872,879,880], BA.4 and BA.5 infected patients [674,881,882,883] show similar severity of COVID-19 compared to BA.1 infections.
The evolution of virulence is hard to predict for the future variants of the virus. Selection does not act directly on the virulence of SARS-CoV-2 (hospitalization and death usually occur after the main transmission period), and its direction most likely depends on how the virus can increase its transmissibility and escape host immune responses [884].
Finally, some attempts have also been made to correlate the severity of infection with individual mutations in the viral genome [885,886,887]. However, the effect of individual mutations is hard to estimate when most comparisons can only be based on observing competing VOCs that differ in multiple mutations that may also involve epistatic effects. We conclude that viral genetic variation can have a strong effect on the risk of severe COVID-19 outcomes, but this cannot easily be traced to individual allelic variants.
3.2. Infecting Dose (Inoculum Size)
Accumulating indirect evidence supports the hypothesis that the size of the viral inoculum might influence the outcome of SARS-CoV-2 infection. First, several dose-titration studies using animal models of SARS-CoV-2 (ferrets, mice or Syrian hamsters) have demonstrated an effect of the infecting viral dose on the severity of subsequent disease symptoms [888,889,890].
Second, several observations are compatible with a link between impaired transmission (lower infecting dose) and lower frequency of severe disease in humans. In particular, widespread use of masking (which is likely to reduce the infecting dose) appears to be associated not only with reduced transmission, but also with a reduced severity or frequency of symptoms among the remaining cases [891,892]. There are also several case studies in closely monitored settings that found an increase in the rate of asymptomatic infections following the introduction of masking, for example at a seafood processing facility and at a chicken plant [891]. Finally, in addition to masking, there are further documented cases where highly similar groups developed divergent clinical forms of COVID-19 potentially due to differences in the setting of exposure. In a study conducted in Spain, among three different clusters of infection the outcome of the disease was the mildest where individuals lived in a large house, less benign where they stayed in an apartment flat, and most severe in the case of attendees of a pre-lockdown meeting in a small conference hall [893]. A Swiss study compared outbreaks in a military setting before and after the introduction of social distancing and stringent hygiene measures [894]. In two of the three groups of young, predominantly male soldiers, the first outbreak occurred before the introduction of preventive measures, and in these groups, 30% of cases resulted in a symptomatic infection. In contrast, the third group experienced their first outbreak after the measures had been implemented, and all infections were asymptomatic.
One hypothesis for the possible causal link between higher infecting dose and more severe disease posits that a larger viral inoculum might overwhelm and evade the primary innate immune responses, resulting in the release of high levels of inflammatory mediators [895]. A larger initial dose might also allow the virus to replicate to higher levels before adaptive immune responses are launched. Note that these mechanisms do not depend on the specifics of COVID-19, and indeed, similar dose dependence of symptoms has been documented for human influenza viruses [896,897], SARS-CoV [898], respiratory syncytial virus [899] and for several non-human pathogens (see reviewed in [892,900]), which lends further indirect support to the “SARS-CoV-2 inoculum hypothesis” [895].
The route of transmission might also influence the clinical outcome of COVID-19. Multiple studies using animal models of COVID-19 indicate that airborne and especially aerosol transmission might result in a disease with an earlier onset and higher severity compared to infections acquired by fomite, oral or gastrointestinal exposures [888,901,902,903].
We conclude that current support for the dose response of COVID-19 severity arises from indirect evidence, and the magnitude of the effect is unclear. An ongoing human dose finding infection study [904] might soon yield the first direct estimation of the effect size.
4. Environmental Factors
4.1. Socioeconomic Factors
Socioeconomic status (SES) has a strong impact on general health and life expectancy [905,906], and could therefore be expected to affect the risk of severe COVID-19 as well. There are an estimated 435 million people in low-income countries (mostly in Sub-Saharan Africa, East and South Asia) who are at high risk from COVID-19 due to their lack of access to health care and safe drinking water, exposure to household air pollution, undernutrition, and other factors associated with low SES [907]. In addition, regional and individual-level studies reported a disproportionately high share of ethnic minorities among both COVID-19 cases and deaths in high-income countries [105,908,909,910,911], which also hints at the importance of multidimensional poverty in COVID-19 morbidity and mortality. Such effects may arise from differences in both the risk of acquiring the infection, and the risk of severe outcomes when infected.
A large body of accumulating evidence supports the idea that low SES is an independent risk factor of severe COVID-19. Multiple indicators related to housing, poverty, nutrition, health care, education and belonging to an ethnic minority have been associated with the outcome of the disease [104,911], and although not all studies confirmed the effects [912,913,914,915] (Table 7), the associations were typically stronger in the larger studies (death in the lowest income quantile compared to the highest OR = 1.95 (1.56–2.43) [202], OR = 1.79 (1.68–1.80) [281]). Although low SES is connected to several other COVID-19 severity risk factors (see Section 5.1. Interactions Between Risk Factors of Severe COVID-19), its independent effect on COVID-19 outcomes indicates that not all aspects of SES relevant for the course of the infection are captured in the known risk factors associated with it.

Table 7.
Socioeconomic variables implicated in the outcome of COVID-19: studies with or without evidence of association.
We note that socioeconomic factors are likely to contribute to the effect of ethnicity/ancestry on COVID-19, due to statistical associations between the two factors. Most meta-analyses (relying mostly on data from the UK and the US) have failed to show an independent effect of ethnicity/ancestry on mortality among hospitalized cases [108,926,927] (except [104]), implying that the higher COVID-19 fatality rates observed among some ethnic minorities might be explained mainly by the higher prevalence of certain comorbidities [928,929,930,931,932] and low SES [933] in these groups. For this reason, and in the absence of a strong case for genetic differences influencing COVID-19 severity that could be linked to ancestry (see Section 2.2. Human Genetic Variation), we do not discuss ethnicity/ancestry as an independent host factor of severe COVID-19.
4.2. Air Pollution
Air pollution has been a public health issue since the industrial revolution [934]. One and a half centuries of chemical manufacturing has resulted in an increased concentration of pollutants in the air, mainly in the form of SO2, NO2, NH3, CO, O3, volatile organic compounds and particulate matter (PM).
The association of the environmental concentration of these pollutants with COVID-19 severity has been extensively investigated (summarized in Table 8), with some, but not all, studies demonstrating statistically significant associations.

Table 8.
Individual-level studies investigating the association between air pollutant exposure and risk of severe clinical outcomes with COVID-19.
While the results of the direct association studies have been mixed, several lines of indirect evidence support that air pollution may have some detrimental effect on COVID-19 pathogenesis. Several components of particulate air pollution facilitate the formation of reactive oxygen signals, which causes inflammation in the lungs [950]. Inflammation caused by PM, and COVID-19 pathogenesis share multiple signaling pathways (TLR, NLR, Nrf2, NF-κB, TNF, IL-1, IL-17, and JAK-STAT) [951,952], which further reinforces this connection. The enhancement of pulmonary epithelial permeability, suppression of mucociliary clearance, interference with antimicrobial proteins, induction of antibacterial instead of antiviral innate immune responses, induced mitochondrial damage and apoptosis, inhibition of IFN production and overexpression of inflammatory metabolites have all been proposed as potential mechanisms [891,953,954]. PM2.5-mediated upregulation of ACE2 in the lungs has also been suggested and supported by an animal study [955] and a human cell culture experiment [956], but a bioinformatic analysis of transcriptomic data related to COVID-19 lung biopsy, SARS-CoV-2 infection in epithelial cells and PM exposure failed to find significant ACE2 upregulation in human infections [952].
We conclude that the link between air pollution and COVID-19 clinical outcome has relatively weak direct evidence, but may affect some subpopulations (e.g., with certain comorbidities) more strongly.
5. Interactions between Effects
5.1. Interactions between Risk Factors of Severe COVID-19
Risk factors of severe COVID-19 do not act in isolation but may influence both the impact and the occurrence of each other in multiple ways (Figure 1). The effect of low socioeconomic status is likely to be mediated, at least in part, through known risk factors that tend to be related to poverty. Low SES is associated with increased chronic and respiratory infection burden both among adults [957,958] and children [959,960], predisposing these disadvantaged individuals to NCDs. Furthermore, multidimensional poverty has a deteriorating effect on lifestyle (nutrition [961,962,963], smoking [964,965,966]), microbiota diversity and composition [967,968,969], and on the development of several comorbidities (cardiovascular [970], pulmonary [971], renal [972], metabolic [973,974] diseases and cancer [975]). Poor housing conditions can be associated with increased indoor air pollution [976,977], which increases the prevalence and severity of both chronic respiratory conditions [978,979] and respiratory infections [978,980,981]. Overcrowding, low-SES neighborhood characteristics and financial instability also escalate chronic stress levels [957,982,983,984], which is considered to be a main contributor to premature mortality in socioeconomically disadvantaged individuals [985,986,987], and may also affect COVID-19 outcomes.
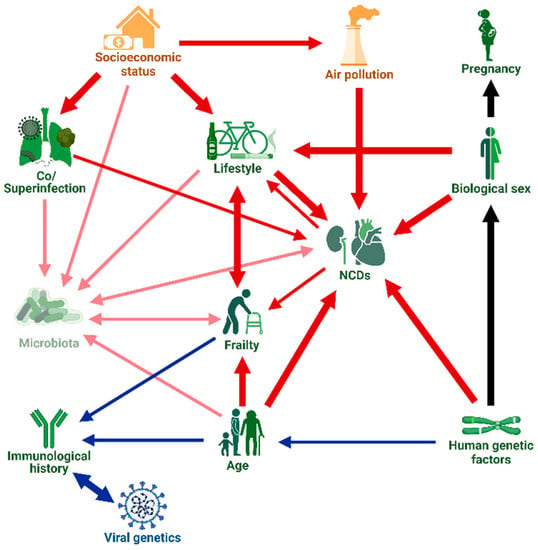
Figure 1.
The network of interactions between identified and proposed risk factors of severe COVID-19. Host (green), viral (blue) and environmental (yellow) factors are connected by arrows indicating three types of interactions between the factors: some factors might modulate the effect of other factors (blue arrows), e.g., advanced age shortens the duration of effective adaptive immune responses; some interactions may affect the occurrence of factors (red arrows), e.g., lifestyle choices influence the likelihood of certain comorbidities; finally, some factors act as the major determinant of another (black arrows), e.g., biological sex determines the possibility of pregnancy, and sex itself is genetically determined in humans. The width of the arrows indicates the perceived magnitude of these effects. The pictogram of microbiota alterations is faded out to illustrate the uncertainty of its causal role in the development of severe COVID-19. The figure was created with BioRender.com.
Lifestyle choices influence the development of non-communicable diseases (CVD [988,989], obesity [990], DM [991], CKD [992] and CLD [993]) independently from socioeconomic status, and they also affect the deterioration of physical functions (frailty) [994,995] and the composition and diversity of the gastrointestinal microbiota [996]. A few studies also examined the modulating effect of physical activity and dietary choices on COVID-19-related microbial alteration [594,997].
Air pollution, and long-term PM exposure in particular, have been linked to decreased lung function [998,999] and a number of comorbidities, such as asthma [1000], COPD [1001], cardiovascular disease [1002], lung cancer [1003], and type 2 diabetes mellitus [1004]. Subpopulations with respiratory diseases show a stronger association between air pollution and COVID-19 severity [940,941,943], which further supports the interaction of these two factors. In turn, chronic lung diseases can limit the capacity of patients for physical work, and affect lifestyle [1005,1006].
Certain chronic coinfections predispose to NCDs (e.g., HIV can facilitate the development of CVD, CKD, CLD, cancer and secondary immunodeficiency [1007], while hepatitis viruses promote liver cirrhosis and cancer [1008]). Furthermore, some parasite infections cause alterations in microbiota diversity [1009,1010], which is tightly connected with the regulation of inflammatory responses [1011,1012].
The gastrointestinal microbiota play an important role in several chronic diseases that are associated with the outcome of COVID-19, such as type 2 DM [1013,1014], obesity [1015,1016], CHD [1017,1018], hypertension [1019,1020], lung diseases (allergies, asthma, COPD) [1021,1022], CLD (liver cirrhosis [1023] and NAFLD [1024]), CKD [1025], neurodegenerative diseases [1026] and depression [1027]. Similarly to the effect of age, NCD- and frailty-related dysbiosis in the GIT [1028,1029,1030] correlates with increased sterile inflammation and inflamm-aging [1031,1032], low butyrate production [1033,1034], impaired barrier function [1035] and increased lipopolysaccharide (LPS) levels in the blood [1036].
Frailty syndrome is characterized by an increased level of sterile systemic inflammation and diminished immune responses, and it is closely related both to age and to age-related diseases [1037]. The deterioration in the physical condition of frail individuals makes it harder to maintain physical activity, possibly accelerating the decline in physical health [1038]. A large proportion of frail individuals have one or more comorbidities, which indicates an overlap between these two risk factors [1039,1040]. As an important example, metabolic inflammation [1039], due to the accumulation of adipose tissue, contributes to the observed inflammatory phenotype in frail individuals [1031]. Although frail individuals have functional adaptive immune responses after SARS-CoV-2 infection [1041] and vaccination [1042,1043], faster waning of IgG levels [1044,1045] and immune-senescent memory T cell functions [1046] indicates that immune memory might be less durable compared to in non-frail individuals.
Age and inflamm-aging is associated with comorbidities, such as obesity, atherosclerosis, rheumatoid arthritis, diabetes, and neurodegeneration [1047,1048]. Reduced ability to recognize cellular damage and the build-up of senescent cells during aging also contribute to the observed inflammatory phenotype in frail individuals [1031]. Aging is also likely to modulate the protective effect of both specific immunity from previous episodes of COVID-19 [677], and cross-reactive immunity from other infections [1049], and is associated with shifts in the upper respiratory tract (URT) [604] and GI microbiota [1050]. Compositional changes and the low diversity of the microbiota in advanced age are associated with a weakened intestinal barrier [1051], elevated levels of bacterial products (such as LPS) in the blood [1052] and heightened inflammation (inflamm-aging) [1053,1054]. In turn, preservation of the microbiota in elderly individuals is correlated with slower immunosenescence [1055]. One study found that age modifies the correlation between microbiota changes and COVID-19 symptom severity concluding that dysbiosis might be an important mediating mechanism between age and COVID-19 severity [604].
Women tend to attribute higher priority to physical health [1056,1057], be more prone to adapt a healthy diet [1058], adhere to hygiene habits [1059], seek professional care [1060] and comply to its recommendations [1061] compared with men [1062,1063,1064]. These behavioral and lifestyle factors contribute to the higher prevalence of chronic diseases, such as COPD [1065], obesity [1066], DM [1067], hypertension [1068] and CVD [1069] among men in high-income countries. However, sex differences in the severity of COVID-19 cannot be explained fully with differences in the prevalence of lifestyle-associated NCDs and health behaviors [1070], indicating further, independent effects of this factor.
Human genetic polymorphisms influence healthy aging and longevity [1071,1072], and predisposition for certain comorbidities, such as type I [1073,1074] and II [1075,1076] diabetes mellitus, obesity [1077,1078] and cardiovascular disease [1079,1080]. Several polymorphisms involved in the SARS-CoV-2-human protein contactome have been associated also with non-communicable diseases (cardiovascular diseases, obesity, schizophrenia) [1081].
Lastly, there is an interaction between the effects of specific immunity from past SARS-CoV-2 infections and viral variation, since the antigenic match between the strain(s) involved in the previous and the current exposures can modulate the protective effect of immunity [1082].
5.2. Direct and Indirect Effects of COVID-19 on the Risk Factors of Severe Disease
While the severity of COVID-19 is influenced by the factors discussed in this review paper, this interaction can be bidirectional, as some of the risk factors can themselves be affected either directly by COVID-19, or indirectly by the human interventions aimed at mitigating the impact of the epidemic. COVID-19 can lead to lasting damage to physical health, which can involve some of the risk factors as well. Acute organ injury caused by COVID-19 can cause long-term cardiovascular [1083,1084], pulmonary [1085], metabolic [1086], renal [1086] and neurological [1087] damage, potentially contributing to chronic conditions that can themselves be risk factors for severe COVID-19 in subsequent infections. COVID-19 can lead to the exacerbation of asthma [1088] and neurodegenerative disorders (both Parkinson’s and Alzheimer’s disease) [1088], and increase the risk of developing mental disorders [1088,1089], diabetes [1090,1091], CKD [1092], hypertension [1093] and CVD [1094]. COVID-19 also disturbs metabolic homeostasis, and can result in a low intake of calories and greatly reduced physical activity during hospitalization, which can exacerbate frailty and biological aging [586]. Indeed, in small cohorts, the incidence of frailty and the number of disabilities increased after COVID-19 hospitalization and critical care [1095,1096,1097].
In addition to the direct pathology of COVID-19, long-term stringent lockdown measures might also contribute to the increased prevalence of some COVID-19 risk factors. During lockdown periods, physical activity levels tend to decrease substantially [1098,1099], while the sales [1100] and consumption [1101] of alcoholic beverages tend to increase in several populations (e.g., older people, more depressed individuals, and essential workers) [1102], indicating that drinking might be a coping mechanism for some during social isolation [1103]. Finally, the economic consequences of the pandemic have negatively impacted the financial situation of hundreds of millions of people worldwide [1104,1105]. Those who had already been at the edge of poverty [1106,1107,1108,1109] or were not able to work from home [1110,1111] had been affected most strongly. Severely affected households experienced increasing food insecurity [1105,1108,1112] and deteriorating mental [1113,1114,1115,1116] and cardiovascular health [1117]. The effects of the pandemic on SES have thus created a positive feedback loop by putting millions of people into a socioeconomic position that makes them more susceptible to both SARS-CoV-2 infection and severe COVID-19.
6. Discussion and Conclusions
In this review, we aimed to provide a comprehensive overview of all ‘inherent’ risk factors of severe COVID-19, omitting only the effect of medical interventions (vaccination and therapy). This broad scope of the review entails some inevitable limitations: while we attempted to identify the most relevant studies for each factor, it was not feasible to conduct a systematic review of all factors. Furthermore, the definitions of ‘mild’ and ‘severe’ COVID-19, the markers used to quantify disease severity, and the potential confounding factors included in the analyses differed widely between the studies, which makes a systematic comparison and summary of this immensely broad field practically unfeasible. The study populations might also have differed in hidden background variables, and the sheer number of the potential contributing factors makes the unbiased estimation of the effect of individual factors very hard. To point the reader to the (in our subjective assessment) most reliable sources of more detailed information, we provide a selection of the largest and (in terms of the cofactors considered) most comprehensive cohort studies and meta-analyses that investigated multiple risk factors of severe COVID-19 in Supplementary Table S1.
Possibly due to the difficulties outlined above, the importance of several factors in COVID-19 remains controversial, with conflicting results in the published literature. Moreover, it is often hard to establish causality and to elucidate the causative mechanisms behind the associations identified. For each factor, we outlined the proposed (in some cases, largely hypothetical) mechanisms of action, and we discussed potential interactions (involving indirectly mediated effects) between the factors.
Based on our broad survey of host, viral and environmental factors implicated in the risk of severe COVID-19, we conclude that both in terms of the strength of the evidence and in the number of infections affected, the age and sex of the patient, the common comorbidities grouped as ‘metabolic syndrome’ (diabetes, hypertension, obesity), and the inherent propensity of the viral variant (VOC) to cause severe disease stand out as the most important factors for the first encounter with the virus, while subsequent re-infections are also strongly affected by SARS-CoV-2-specific adaptive immunity. Influenced by variation in these factors, the risk of individuals varies across a very broad spectrum (age alone spanned approximately four orders of magnitude in the risk of mortality in the first wave of the epidemic [9]). Awareness of these factors can help decisions in the clinic (whom to treat, how to allocate resources), and also public health decisions at the population level. While prevalent immunity from previous infections and vaccinations (and the emergence of the Omicron variants) has substantially reduced the overall risk of severe outcomes compared with the first waves of the pandemic, the risk factors identified in previous analyses are likely to continue to shape relative risks in the future.
Supplementary Materials
The following supporting information can be downloaded at: https://www.mdpi.com/article/10.3390/v15010175/s1, Table S1: List of, and compiled results from, selected large cohort studies and meta-analyses investigating multiple risk factors of severe COVID-19.
Author Contributions
Conceptualization, L.Z. and V.M.; literature study, L.Z. and V.M.; writing—original draft preparation, L.Z.; writing—review and editing, V.M. and L.Z.; visualization, L.Z.; supervision, V.M.; project administration, V.M.; funding acquisition, V.M. All authors have read and agreed to the published version of the manuscript.
Funding
This study was funded by the National Research, Development and Innovation Office in Hungary (RRF-2.3.1-21-2022-00006).
Institutional Review Board Statement
Not applicable.
Informed Consent Statement
Not applicable.
Data Availability Statement
Not applicable.
Acknowledgments
We thank participants of a symposium of the National Laboratory for Health Security (Hungary) for a constructive discussion of a presentation of our work.
Conflicts of Interest
The authors declare no conflict of interest. The funders had no role in the design of the study; in the collection, analyses, or interpretation of data; in the writing of the manuscript, or in the decision to publish the results.
Appendix A. Relative Risk Measures Used to Identify COVID-19 Risk Factors
The risk ratio (RR), the odds ratio (OR) and the hazard ratio (HR) are three frequently used statistical measures in medical research that compare the occurrence of a clinical outcome between two groups. These two groups may consist of a control group, and one in which some treatment was administered, or in which a clinically relevant characteristic can be observed (the “intervention group”). However, the exact goal and therefore the use and limitations differ between these statistics.
Risk is the probability of a measured outcome occurring in one of the groups. Hence, the risk ratio (RR) is the quotient of the risks in the intervention and in the control group. A RR = 1 means that there is no difference in the likelihood of the outcome occurring (“null hypothesis”), while a RR > 1 or RR < 1 shows increased or decreased risk in the intervention group compared to the control (the “alternative hypothesis”). The reliability and uncertainty of the result is typically indicated by the p-value and the confidence interval. The RR is informative and easy to understand, but its calculation requires a representative sample of the population, and it is therefore not applicable in some study designs (e.g., case–control studies).
Odds are the quotient of the number of events and non-events in one of the studied groups, where the event is the outcome of interest. The odds thus express how many times it is more likely that the outcome happens than it does not, and the odds ratio (OR) measures how much larger the odds are in one of the groups compared to the other. This measure can also be calculated in case–control studies, where a representative sample of the studies population is not available, and the overall risk and odds of the event in either group cannot be directly estimated. On the other hand, the OR is harder to intuitively understand. Importantly, the OR approximates the RR accurately when risks are very small; however, with larger risks, the OR overestimates the RR.
Finally, hazard estimates the instantaneous probability of the observed event during the observation period, as opposed to risk and odds that refer to the occurrence of an event during the entire observation period. The hazard ratio (HR) is therefore the ratio of the hazards in the intervention group and in the control group, and it shows how the intervention changes the rate of experiencing the outcome. HRs are meaningful if the rates are relatively constant in the two groups over the observation period.
For a more detailed explanation and helpful examples, we recommend reading the review by George et al. [1118].
References
- Vegivinti, C.T.R.; Evanson, K.W.; Lyons, H.; Akosman, I.; Barrett, A.; Hardy, N.; Kane, B.; Keesari, P.R.; Pulakurthi, Y.S.; Sheffels, E.; et al. Efficacy of Antiviral Therapies for COVID-19: A Systematic Review of Randomized Controlled Trials. BMC Infect. Dis. 2022, 22, 107. [Google Scholar] [CrossRef] [PubMed]
- Coronavirus Disease 2019 (COVID-19) Treatment Guidelines. National Institutes of Health COVID-19 Treatment Guidelines Panel. Available online: https://www.covid19treatmentguidelines.nih.gov/ (accessed on 9 December 2022).
- Wu, C.; Chen, X.; Cai, Y.; Xia, J.; Zhou, X.; Xu, S.; Huang, H.; Zhang, L.; Zhou, X.; Du, C.; et al. Risk Factors Associated with Acute Respiratory Distress Syndrome and Death in Patients with Coronavirus Disease 2019 Pneumonia in Wuhan, China. JAMA Intern. Med. 2020, 180, 934–943. [Google Scholar] [CrossRef] [PubMed]
- Yang, X.; Yu, Y.; Xu, J.; Shu, H.; Liu, H.; Wu, Y.; Zhang, L.; Yu, Z.; Fang, M.; Yu, T.; et al. Clinical Course and Outcomes of Critically Ill Patients with SARS-CoV-2 Pneumonia in Wuhan, China: A Single-Centered, Retrospective, Observational Study. Lancet Respir. Med. 2020, 8, 475–481. [Google Scholar] [CrossRef] [PubMed]
- Pijls, B.G.; Jolani, S.; Atherley, A.; Derckx, R.T.; Dijkstra, J.I.R.; Franssen, G.H.L.; Hendriks, S.; Richters, A.; Venemans-Jellema, A.; Zalpuri, S.; et al. Demographic Risk Factors for COVID-19 Infection, Severity, ICU Admission and Death: A Meta-Analysis of 59 Studies. BMJ Open 2021, 11, e044640. [Google Scholar] [CrossRef] [PubMed]
- Fang, X.; Li, S.; Yu, H.; Wang, P.; Zhang, Y.; Chen, Z.; Li, Y.; Cheng, L.; Li, W.; Jia, H.; et al. Epidemiological, Comorbidity Factors with Severity and Prognosis of COVID-19: A Systematic Review and Meta-Analysis. Aging 2020, 12, 12493–12503. [Google Scholar] [CrossRef] [PubMed]
- Verity, R.; Okell, L.C.; Dorigatti, I.; Winskill, P.; Whittaker, C.; Imai, N.; Cuomo-Dannenburg, G.; Thompson, H.; Walker, P.G.T.; Fu, H.; et al. Estimates of the Severity of Coronavirus Disease 2019: A Model-Based Analysis. Lancet Infect. Dis. 2020, 20, 669–677. [Google Scholar] [CrossRef] [PubMed]
- Salje, H.; Tran Kiem, C.; Lefrancq, N.; Courtejoie, N.; Bosetti, P.; Paireau, J.; Andronico, A.; Hozé, N.; Richet, J.; Dubost, C.-L.; et al. Estimating the Burden of SARS-CoV-2 in France. Science 2020, 369, 208–211. [Google Scholar] [CrossRef]
- O’Driscoll, M.; Ribeiro Dos Santos, G.; Wang, L.; Cummings, D.A.T.; Azman, A.S.; Paireau, J.; Fontanet, A.; Cauchemez, S.; Salje, H. Age-Specific Mortality and Immunity Patterns of SARS-CoV-2. Nature 2021, 590, 140–145. [Google Scholar] [CrossRef]
- Du, P.; Li, D.; Wang, A.; Shen, S.; Ma, Z.; Li, X. A Systematic Review and Meta-Analysis of Risk Factors Associated with Severity and Death in COVID-19 Patients. Can. J. Infect. Dis. Med. Microbiol. 2021, 2021, e6660930. [Google Scholar] [CrossRef]
- Booth, A.; Reed, A.B.; Ponzo, S.; Yassaee, A.; Aral, M.; Plans, D.; Labrique, A.; Mohan, D. Population Risk Factors for Severe Disease and Mortality in COVID-19: A Global Systematic Review and Meta-Analysis. PLoS ONE 2021, 16, e0247461. [Google Scholar] [CrossRef]
- Ebinger, J.E.; Achamallah, N.; Ji, H.; Claggett, B.L.; Sun, N.; Botting, P.; Nguyen, T.-T.; Luong, E.; Kim, E.H.; Park, E.; et al. Pre-Existing Traits Associated with COVID-19 Illness Severity. PLoS ONE 2020, 15, e0236240. [Google Scholar] [CrossRef] [PubMed]
- Carrillo-Vega, M.F.; Salinas-Escudero, G.; García-Peña, C.; Gutiérrez-Robledo, L.M.; Parra-Rodríguez, L. Early Estimation of the Risk Factors for Hospitalization and Mortality by COVID-19 in Mexico. PLoS ONE 2020, 15, e0238905. [Google Scholar] [CrossRef] [PubMed]
- Schneider, J.L.; Rowe, J.H.; Garcia-de-Alba, C.; Kim, C.F.; Sharpe, A.H.; Haigis, M.C. The Aging Lung: Physiology, Disease, and Immunity. Cell 2021, 184, 1990–2019. [Google Scholar] [CrossRef]
- Sanchez-Vazquez, R.; Guío-Carrión, A.; Zapatero-Gaviria, A.; Martínez, P.; Blasco, M.A. Shorter Telomere Lengths in Patients with Severe COVID-19 Disease. Aging 2021, 13, 1–15. [Google Scholar] [CrossRef] [PubMed]
- Chow, R.D.; Majety, M.; Chen, S. The Aging Transcriptome and Cellular Landscape of the Human Lung in Relation to SARS-CoV-2. Nat. Commun. 2021, 12, 4. [Google Scholar] [CrossRef]
- Chen, J.; Jiang, Q.; Xia, X.; Liu, K.; Yu, Z.; Tao, W.; Gong, W.; Han, J.-D.J. Individual Variation of the SARS-CoV-2 Receptor ACE2 Gene Expression and Regulation. Aging Cell 2020, 19, e13168. [Google Scholar] [CrossRef]
- Schouten, L.R.; van Kaam, A.H.; Kohse, F.; Veltkamp, F.; Bos, L.D.; de Beer, F.M.; van Hooijdonk, R.T.; Horn, J.; Straat, M.; Witteveen, E.; et al. Age-Dependent Differences in Pulmonary Host Responses in ARDS: A Prospective Observational Cohort Study. Ann. Intensive Care 2019, 9, 55. [Google Scholar] [CrossRef]
- Inde, Z.; Croker, B.A.; Yapp, C.; Joshi, G.N.; Spetz, J.; Fraser, C.; Qin, X.; Xu, L.; Deskin, B.; Ghelfi, E.; et al. Age-Dependent Regulation of SARS-CoV-2 Cell Entry Genes and Cell Death Programs Correlates with COVID-19 Severity. Sci. Adv. 2021, 7, eabf8609. [Google Scholar] [CrossRef]
- Gheware, A.; Ray, A.; Rana, D.; Bajpai, P.; Nambirajan, A.; Arulselvi, S.; Mathur, P.; Trikha, A.; Arava, S.; Das, P.; et al. ACE2 Protein Expression in Lung Tissues of Severe COVID-19 Infection. Sci. Rep. 2022, 12, 4058. [Google Scholar] [CrossRef]
- Zheng, M. ACE2 and COVID-19 Susceptibility and Severity. Aging Dis. 2022, 13, 360–372. [Google Scholar] [CrossRef]
- Pinto, B.G.G.; Oliveira, A.E.R.; Singh, Y.; Jimenez, L.; Gonçalves, A.N.A.; Ogava, R.L.T.; Creighton, R.; Schatzmann Peron, J.P.; Nakaya, H.I. ACE2 Expression Is Increased in the Lungs of Patients with Comorbidities Associated with Severe COVID-19. J. Infect. Dis. 2020, 222, 556–563. [Google Scholar] [CrossRef] [PubMed]
- Rodrigues, R.; Costa de Oliveira, S. The Impact of Angiotensin-Converting Enzyme 2 (ACE2) Expression Levels in Patients with Comorbidities on COVID-19 Severity: A Comprehensive Review. Microorganisms 2021, 9, 1692. [Google Scholar] [CrossRef] [PubMed]
- Chamsi-Pasha, M.A.R.; Shao, Z.; Tang, W.H.W. Angiotensin-Converting Enzyme 2 as a Therapeutic Target for Heart Failure. Curr. Heart Fail. Rep. 2014, 11, 58–63. [Google Scholar] [CrossRef] [PubMed]
- Arendse, L.B.; Danser, A.H.J.; Poglitsch, M.; Touyz, R.M.; Burnett, J.C.; Llorens-Cortes, C.; Ehlers, M.R.; Sturrock, E.D. Novel Therapeutic Approaches Targeting the Renin-Angiotensin System and Associated Peptides in Hypertension and Heart Failure. Pharmacol. Rev. 2019, 71, 539–570. [Google Scholar] [CrossRef] [PubMed]
- Hoffmann, M.; Kleine-Weber, H.; Schroeder, S.; Krüger, N.; Herrler, T.; Erichsen, S.; Schiergens, T.S.; Herrler, G.; Wu, N.-H.; Nitsche, A.; et al. SARS-CoV-2 Cell Entry Depends on ACE2 and TMPRSS2 and Is Blocked by a Clinically Proven Protease Inhibitor. Cell 2020, 181, 271–280. [Google Scholar] [CrossRef]
- Letko, M.; Marzi, A.; Munster, V. Functional Assessment of Cell Entry and Receptor Usage for SARS-CoV-2 and Other Lineage B Betacoronaviruses. Nat. Microbiol. 2020, 5, 562–569. [Google Scholar] [CrossRef]
- Schuler, B.A.; Habermann, A.C.; Plosa, E.J.; Taylor, C.J.; Jetter, C.; Negretti, N.M.; Kapp, M.E.; Benjamin, J.T.; Gulleman, P.; Nichols, D.S.; et al. Age-Determined Expression of Priming Protease TMPRSS2 and Localization of SARS-CoV-2 in Lung Epithelium. J. Clin. Investig. 2021, 131, e140766. [Google Scholar] [CrossRef]
- Wang, A.; Chiou, J.; Poirion, O.B.; Buchanan, J.; Valdez, M.J.; Verheyden, J.M.; Hou, X.; Kudtarkar, P.; Narendra, S.; Newsome, J.M.; et al. Single-Cell Multiomic Profiling of Human Lungs Reveals Cell-Type-Specific and Age-Dynamic Control of SARS-CoV2 Host Genes. eLife 2020, 9, e62522. [Google Scholar] [CrossRef]
- Deng, X.; Hackbart, M.; Mettelman, R.C.; O’Brien, A.; Mielech, A.M.; Yi, G.; Kao, C.C.; Baker, S.C. Coronavirus Nonstructural Protein 15 Mediates Evasion of DsRNA Sensors and Limits Apoptosis in Macrophages. Proc. Natl. Acad. Sci. USA 2017, 114, E4251–E4260. [Google Scholar] [CrossRef]
- DeDiego, M.L.; Nieto-Torres, J.L.; Jiménez-Guardeño, J.M.; Regla-Nava, J.A.; Alvarez, E.; Oliveros, J.C.; Zhao, J.; Fett, C.; Perlman, S.; Enjuanes, L. Severe Acute Respiratory Syndrome Coronavirus Envelope Protein Regulates Cell Stress Response and Apoptosis. PLoS Pathog. 2011, 7, e1002315. [Google Scholar] [CrossRef]
- Danthi, P. Viruses and the Diversity of Cell Death. Annu. Rev. Virol. 2016, 3, 533–553. [Google Scholar] [CrossRef] [PubMed]
- Rea, I.M.; Alexander, H.D. Triple Jeopardy in Ageing: COVID-19, Co-Morbidities and Inflamm-Ageing. Ageing Res. Rev. 2022, 73, 101494. [Google Scholar] [CrossRef] [PubMed]
- Salminen, A.; Huuskonen, J.; Ojala, J.; Kauppinen, A.; Kaarniranta, K.; Suuronen, T. Activation of Innate Immunity System during Aging: NF-KB Signaling Is the Molecular Culprit of Inflamm-Aging. Ageing Res. Rev. 2008, 7, 83–105. [Google Scholar] [CrossRef] [PubMed]
- Brodin, P. Immune Determinants of COVID-19 Disease Presentation and Severity. Nat. Med. 2021, 27, 28–33. [Google Scholar] [CrossRef] [PubMed]
- Lagunas-Rangel, F.A. Neutrophil-to-Lymphocyte Ratio and Lymphocyte-to-C-Reactive Protein Ratio in Patients with Severe Coronavirus Disease 2019 (COVID-19): A Meta-Analysis. J. Med. Virol. 2020, 92, 1733–1734. [Google Scholar] [CrossRef]
- Molony, R.D.; Nguyen, J.T.; Kong, Y.; Montgomery, R.R.; Shaw, A.C.; Iwasaki, A. Aging Impairs Both Primary and Secondary RIG-I Signaling for Interferon Induction in Human Monocytes. Sci. Signal. 2017, 10, eaan2392. [Google Scholar] [CrossRef]
- Rodrigues, T.S.; de Sá, K.S.G.; Ishimoto, A.Y.; Becerra, A.; Oliveira, S.; Almeida, L.; Gonçalves, A.V.; Perucello, D.B.; Andrade, W.A.; Castro, R.; et al. Inflammasomes Are Activated in Response to SARS-CoV-2 Infection and Are Associated with COVID-19 Severity in Patients. J. Exp. Med. 2021, 218, e20201707. [Google Scholar] [CrossRef]
- McGonagle, D.; Sharif, K.; O’Regan, A.; Bridgewood, C. The Role of Cytokines Including Interleukin-6 in COVID-19 Induced Pneumonia and Macrophage Activation Syndrome-Like Disease. Autoimmun. Rev. 2020, 19, 102537. [Google Scholar] [CrossRef]
- Sayed, N.; Huang, Y.; Nguyen, K.; Krejciova-Rajaniemi, Z.; Grawe, A.P.; Gao, T.; Tibshirani, R.; Hastie, T.; Alpert, A.; Cui, L.; et al. An Inflammatory Aging Clock (IAge) Based on Deep Learning Tracks Multimorbidity, Immunosenescence, Frailty and Cardiovascular Aging. Nat. Aging 2021, 1, 598–615. [Google Scholar] [CrossRef]
- Schmitt, C.A.; Tchkonia, T.; Niedernhofer, L.J.; Robbins, P.D.; Kirkland, J.L.; Lee, S. COVID-19 and Cellular Senescence. Nat. Rev. Immunol. 2022, 1–13. [Google Scholar] [CrossRef]
- Bartleson, J.M.; Radenkovic, D.; Covarrubias, A.J.; Furman, D.; Winer, D.A.; Verdin, E. SARS-CoV-2, COVID-19 and the Aging Immune System. Nat. Aging 2021, 1, 769–782. [Google Scholar] [CrossRef] [PubMed]
- Bergsma, T.; Rogaeva, E. DNA Methylation Clocks and Their Predictive Capacity for Aging Phenotypes and Healthspan. Neurosci. Insights 2020, 15, 263310552094222. [Google Scholar] [CrossRef] [PubMed]
- McCrory, C.; Fiorito, G.; Hernandez, B.; Polidoro, S.; O’Halloran, A.M.; Hever, A.; Ni Cheallaigh, C.; Lu, A.T.; Horvath, S.; Vineis, P.; et al. GrimAge Outperforms Other Epigenetic Clocks in the Prediction of Age-Related Clinical Phenotypes and All-Cause Mortality. J. Gerontol. Ser. A 2021, 76, 741–749. [Google Scholar] [CrossRef] [PubMed]
- Mongelli, A.; Barbi, V.; Gottardi Zamperla, M.; Atlante, S.; Forleo, L.; Nesta, M.; Massetti, M.; Pontecorvi, A.; Nanni, S.; Farsetti, A.; et al. Evidence for Biological Age Acceleration and Telomere Shortening in COVID-19 Survivors. Int. J. Mol. Sci. 2021, 22, 6151. [Google Scholar] [CrossRef] [PubMed]
- Wang, Q.; Codd, V.; Raisi-Estabragh, Z.; Musicha, C.; Bountziouka, V.; Kaptoge, S.; Allara, E.; Di Angelantonio, E.; Butterworth, A.S.; Wood, A.M.; et al. Shorter Leukocyte Telomere Length Is Associated with Adverse COVID-19 Outcomes: A Cohort Study in UK Biobank. EBioMedicine 2021, 70, 103485. [Google Scholar] [CrossRef] [PubMed]
- Cao, X.; Li, W.; Wang, T.; Ran, D.; Davalos, V.; Planas-Serra, L.; Pujol, A.; Esteller, M.; Wang, X.; Yu, H. Accelerated Biological Aging in COVID-19 Patients. Nat. Commun. 2022, 13, 2135. [Google Scholar] [CrossRef]
- Bohlin, J.; Page, C.M.; Lee, Y.; Pettersson, J.H.-O.; Jugessur, A.; Magnus, P.; Håberg, S.E. Age and Sex Effects on DNA Methylation Sites Linked to Genes Implicated in Severe COVID-19 and SARS-CoV-2 Host Cell Entry. PLoS ONE 2022, 17, e0269105. [Google Scholar] [CrossRef]
- Corley, M.J.; Pang, A.P.S.; Dody, K.; Mudd, P.A.; Patterson, B.K.; Seethamraju, H.; Bram, Y.; Peluso, M.J.; Torres, L.; Iyer, N.S.; et al. Genome-wide DNA Methylation Profiling of Peripheral Blood Reveals an Epigenetic Signature Associated with Severe COVID-19. J. Leukoc. Biol. 2021, 110, 21–26. [Google Scholar] [CrossRef]
- Balnis, J.; Madrid, A.; Hogan, K.J.; Drake, L.A.; Chieng, H.C.; Tiwari, A.; Vincent, C.E.; Chopra, A.; Vincent, P.A.; Robek, M.D.; et al. Blood DNA Methylation and COVID-19 Outcomes. Clin. Epigenetics 2021, 13, 118. [Google Scholar] [CrossRef]
- Pang, A.P.S.; Higgins-Chen, A.T.; Comite, F.; Raica, I.; Arboleda, C.; Went, H.; Mendez, T.; Schotsaert, M.; Dwaraka, V.; Smith, R.; et al. Longitudinal Study of DNA Methylation and Epigenetic Clocks Prior to and Following Test-Confirmed COVID-19 and MRNA Vaccination. Front. Genet. 2022, 13, 819749. [Google Scholar] [CrossRef]
- Attia, M.H. A Cautionary Note on Altered Pace of Aging in the COVID-19 Era. Forensic Sci. Int. Genet. 2022, 59, 102724. [Google Scholar] [CrossRef] [PubMed]
- Kenmoe, S.; Kengne-Nde, C.; Ebogo-Belobo, J.T.; Mbaga, D.S.; Modiyinji, A.F.; Njouom, R. Systematic Review and Meta-Analysis of the Prevalence of Common Respiratory Viruses in Children < 2 Years with Bronchiolitis in the Pre-COVID-19 Pandemic Era. PLoS ONE 2020, 15, e0242302. [Google Scholar] [CrossRef]
- Wang, X.; Li, Y.; O’Brien, K.L.; Madhi, S.A.; Widdowson, M.-A.; Byass, P.; Omer, S.B.; Abbas, Q.; Ali, A.; Amu, A.; et al. Global Burden of Respiratory Infections Associated with Seasonal Influenza in Children under 5 Years in 2018: A Systematic Review and Modelling Study. Lancet Glob. Health 2020, 8, e497–e510. [Google Scholar] [CrossRef] [PubMed]
- Sharma, V.; Coleman, S.; Nixon, J.; Sharples, L.; Hamilton-Shield, J.; Rutter, H.; Bryant, M. A Systematic Review and Meta-Analysis Estimating the Population Prevalence of Comorbidities in Children and Adolescents Aged 5 to 18 Years. Obes. Rev. 2019, 20, 1341–1349. [Google Scholar] [CrossRef] [PubMed]
- Pierce, C.A.; Preston-Hurlburt, P.; Dai, Y.; Aschner, C.B.; Cheshenko, N.; Galen, B.; Garforth, S.J.; Herrera, N.G.; Jangra, R.K.; Morano, N.C.; et al. Immune Responses to SARS-CoV-2 Infection in Hospitalized Pediatric and Adult Patients. Sci. Transl. Med. 2020, 12, eabd5487. [Google Scholar] [CrossRef]
- Pierce, C.A.; Sy, S.; Galen, B.; Goldstein, D.Y.; Orner, E.; Keller, M.J.; Herold, K.C.; Herold, B.C. Natural Mucosal Barriers and COVID-19 in Children. JCI Insight 2021, 6, e148694. [Google Scholar] [CrossRef]
- Yoshida, M.; Worlock, K.B.; Huang, N.; Lindeboom, R.G.H.; Butler, C.R.; Kumasaka, N.; Dominguez Conde, C.; Mamanova, L.; Bolt, L.; Richardson, L.; et al. Local and Systemic Responses to SARS-CoV-2 Infection in Children and Adults. Nature 2022, 602, 321–327. [Google Scholar] [CrossRef]
- Chou, J.; Thomas, P.G.; Randolph, A.G. Immunology of SARS-CoV-2 Infection in Children. Nat. Immunol. 2022, 23, 177–185. [Google Scholar] [CrossRef]
- Cohen, C.A.; Li, A.P.Y.; Hachim, A.; Hui, D.S.C.; Kwan, M.Y.W.; Tsang, O.T.Y.; Chiu, S.S.; Chan, W.H.; Yau, Y.S.; Kavian, N.; et al. SARS-CoV-2 Specific T Cell Responses Are Lower in Children and Increase with Age and Time after Infection. Nat. Commun. 2021, 12, 4678. [Google Scholar] [CrossRef]
- Vazquez, C.; Swanson, S.E.; Negatu, S.G.; Dittmar, M.; Miller, J.; Ramage, H.R.; Cherry, S.; Jurado, K.A. SARS-CoV-2 Viral Proteins NSP1 and NSP13 Inhibit Interferon Activation through Distinct Mechanisms. PLoS ONE 2021, 16, e0253089. [Google Scholar] [CrossRef]
- Loske, J.; Röhmel, J.; Lukassen, S.; Stricker, S.; Magalhães, V.G.; Liebig, J.; Chua, R.L.; Thürmann, L.; Messingschlager, M.; Seegebarth, A.; et al. Pre-Activated Antiviral Innate Immunity in the Upper Airways Controls Early SARS-CoV-2 Infection in Children. Nat. Biotechnol. 2022, 40, 319–324. [Google Scholar] [CrossRef] [PubMed]
- Hu, P.; Bauer, V.L.; Sawyer, S.L.; Diaz-Griffero, F. Human ACE2 Polymorphisms from Different Human Populations Modulate SARS-CoV-2 Infection. Viruses 2022, 14, 1451. [Google Scholar] [CrossRef]
- Dieter, C.; Brondani, L.d.A.; Leitão, C.B.; Gerchman, F.; Lemos, N.E.; Crispim, D. Genetic Polymorphisms Associated with Susceptibility to COVID-19 Disease and Severity: A Systematic Review and Meta-Analysis. PLoS ONE 2022, 17, e0270627. [Google Scholar] [CrossRef] [PubMed]
- Saengsiwaritt, W.; Jittikoon, J.; Chaikledkaew, U.; Udomsinprasert, W. Genetic Polymorphisms of ACE1, ACE2, and TMPRSS2 Associated with COVID-19 Severity: A Systematic Review with Meta-Analysis. Rev. Med. Virol. 2022, 32, e2323. [Google Scholar] [CrossRef]
- Russo, R.; Andolfo, I.; Lasorsa, V.A.; Iolascon, A.; Capasso, M. Genetic Analysis of the Coronavirus SARS-CoV-2 Host Protease TMPRSS2 in Different Populations. Front. Genet. 2020, 11, 872. [Google Scholar] [CrossRef] [PubMed]
- Niemi, M.E.K.; Daly, M.J.; Ganna, A. The Human Genetic Epidemiology of COVID-19. Nat. Rev. Genet. 2022, 23, 533–546. [Google Scholar] [CrossRef] [PubMed]
- Pathak, G.A.; Karjalainen, J.; Stevens, C.; Neale, B.M.; Daly, M.; Ganna, A.; Andrews, S.J.; Kanai, M.; Cordioli, M.; Polimanti, R.; et al. A First Update on Mapping the Human Genetic Architecture of COVID-19. Nature 2022, 608, E1–E10. [Google Scholar] [CrossRef]
- van Moorsel, C.H.M.; van der Vis, J.J.; Duckworth, A.; Scotton, C.J.; Benschop, C.; Ellinghaus, D.; Ruven, H.J.T.; Quanjel, M.J.R.; Grutters, J.C. The MUC5B Promoter Polymorphism Associates with Severe COVID-19 in the European Population. Front. Med. 2021, 8, 668024. [Google Scholar] [CrossRef]
- Roberts, G.H.L.; Partha, R.; Rhead, B.; Knight, S.C.; Park, D.S.; Coignet, M.V.; Zhang, M.; Berkowitz, N.; Turrisini, D.A.; Gaddis, M.; et al. Expanded COVID-19 Phenotype Definitions Reveal Distinct Patterns of Genetic Association and Protective Effects. Nat. Genet. 2022, 54, 374–381. [Google Scholar] [CrossRef]
- Karlsen, T.H. Understanding COVID-19 through Genome-Wide Association Studies. Nat. Genet. 2022, 54, 368–369. [Google Scholar] [CrossRef]
- Downes, D.J.; Cross, A.R.; Hua, P.; Roberts, N.; Schwessinger, R.; Cutler, A.J.; Munis, A.M.; Brown, J.; Mielczarek, O.; de Andrea, C.E.; et al. Identification of LZTFL1 as a Candidate Effector Gene at a COVID-19 Risk Locus. Nat. Genet. 2021, 53, 1606–1615. [Google Scholar] [CrossRef] [PubMed]
- Fischer, J.C.; Schmidt, A.G.; Bölke, E.; Uhrberg, M.; Keitel, V.; Feldt, T.; Jensen, B.; Häussinger, D.; Adams, O.; Schneider, E.M.; et al. Association of HLA Genotypes, AB0 Blood Type and Chemokine Receptor 5 Mutant CD195 with the Clinical Course of COVID-19. Eur. J. Med. Res. 2021, 26, 107. [Google Scholar] [CrossRef] [PubMed]
- Wickenhagen, A.; Sugrue, E.; Lytras, S.; Kuchi, S.; Noerenberg, M.; Turnbull, M.L.; Loney, C.; Herder, V.; Allan, J.; Jarmson, I.; et al. A Prenylated DsRNA Sensor Protects against Severe COVID-19. Science 2021, 374, eabj3624. [Google Scholar] [CrossRef] [PubMed]
- Zhou, S.; Butler-Laporte, G.; Nakanishi, T.; Morrison, D.R.; Afilalo, J.; Afilalo, M.; Laurent, L.; Pietzner, M.; Kerrison, N.; Zhao, K.; et al. A Neanderthal OAS1 Isoform Protects Individuals of European Ancestry against COVID-19 Susceptibility and Severity. Nat. Med. 2021, 27, 659–667. [Google Scholar] [CrossRef] [PubMed]
- Asano, T.; Boisson, B.; Onodi, F.; Matuozzo, D.; Moncada-Velez, M.; Maglorius Renkilaraj, M.R.L.; Zhang, P.; Meertens, L.; Bolze, A.; Materna, M.; et al. X-Linked Recessive TLR7 Deficiency in ~1% of Men under 60 Years Old with Life-Threatening COVID-19. Sci. Immunol. 2021, 6, eabl4348. [Google Scholar] [CrossRef]
- Namkoong, H.; Edahiro, R.; Takano, T.; Nishihara, H.; Shirai, Y.; Sonehara, K.; Tanaka, H.; Azekawa, S.; Mikami, Y.; Lee, H.; et al. DOCK2 Is Involved in the Host Genetics and Biology of Severe COVID-19. Nature 2022, 609, 754–760. [Google Scholar] [CrossRef] [PubMed]
- Verma, S.; Abbas, M.; Verma, S.; Khan, F.H.; Raza, S.T.; Siddiqi, Z.; Ahmad, I.; Mahdi, F. Impact of I/D Polymorphism of Angiotensin-Converting Enzyme 1 (ACE1) Gene on the Severity of COVID-19 Patients. Infect. Genet. Evol. 2021, 91, 104801. [Google Scholar] [CrossRef]
- Pietzner, M.; Chua, R.L.; Wheeler, E.; Jechow, K.; Willett, J.D.S.; Radbruch, H.; Trump, S.; Heidecker, B.; Zeberg, H.; Heppner, F.L.; et al. ELF5 Is a Potential Respiratory Epithelial Cell-Specific Risk Gene for Severe COVID-19. Nat. Commun. 2022, 13, 4484. [Google Scholar] [CrossRef]
- Hernández Cordero, A.I.; Li, X.; Milne, S.; Yang, C.X.; Bossé, Y.; Joubert, P.; Timens, W.; van den Berge, M.; Nickle, D.; Hao, K.; et al. Multi-Omics Highlights ABO Plasma Protein as a Causal Risk Factor for COVID-19. Hum. Genet. 2021, 140, 969–979. [Google Scholar] [CrossRef]
- Kurki, S.N.; Kantonen, J.; Kaivola, K.; Hokkanen, L.; Mäyränpää, M.I.; Puttonen, H.; Martola, J.; Pöyhönen, M.; Kero, M.; Tuimala, J.; et al. APOE Ε4 Associates with Increased Risk of Severe COVID-19, Cerebral Microhaemorrhages and Post-COVID Mental Fatigue: A Finnish Biobank, Autopsy and Clinical Study. Acta Neuropathol. Commun. 2021, 9, 199. [Google Scholar] [CrossRef]
- National Center for Biotechnology Information (NCBI). Available online: https://www.ncbi.nlm.nih.gov/ (accessed on 13 July 2022).
- COVID-19 Host Genetics Initiative; Gazon, H.; Juszczak, D.; Fadeur, M.; Camby, S.; Meuris, C.; Thys, M.; Jacques, J.; Henket, M.; Beguin, Y.; et al. Mapping the Human Genetic Architecture of COVID-19. Nature 2021, 600, 472–477. [Google Scholar] [CrossRef]
- Huang, Q.-M.; Zhang, P.-D.; Li, Z.-H.; Zhou, J.-M.; Liu, D.; Zhang, X.-R.; Zhong, W.-F.; Zhang, Y.-J.; Shen, D.; Liang, F.; et al. Genetic Risk and Chronic Obstructive Pulmonary Disease Independently Predict the Risk of Incident Severe COVID-19. Ann. Am. Thorac. Soc. 2022, 19, 58–65. [Google Scholar] [CrossRef] [PubMed]
- Fadista, J.; Manning, A.K.; Florez, J.C.; Groop, L. The (in)Famous GWAS P-Value Threshold Revisited and Updated for Low-Frequency Variants. Eur. J. Hum. Genet. 2016, 24, 1202–1205. [Google Scholar] [CrossRef] [PubMed]
- Tangye, S.G.; Al-Herz, W.; Bousfiha, A.; Chatila, T.; Cunningham-Rundles, C.; Etzioni, A.; Franco, J.L.; Holland, S.M.; Klein, C.; Morio, T.; et al. Human Inborn Errors of Immunity: 2019 Update on the Classification from the International Union of Immunological Societies Expert Committee. J. Clin. Immunol. 2020, 40, 24–64. [Google Scholar] [CrossRef]
- van der Made, C.I.; Simons, A.; Schuurs-Hoeijmakers, J.; van den Heuvel, G.; Mantere, T.; Kersten, S.; van Deuren, R.C.; Steehouwer, M.; van Reijmersdal, S.V.; Jaeger, M.; et al. Presence of Genetic Variants among Young Men with Severe COVID-19. JAMA 2020, 324, 663–673. [Google Scholar] [CrossRef]
- Fallerini, C.; Daga, S.; Mantovani, S.; Benetti, E.; Picchiotti, N.; Francisci, D.; Paciosi, F.; Schiaroli, E.; Baldassarri, M.; Fava, F.; et al. Association of Toll-like Receptor 7 Variants with Life-Threatening COVID-19 Disease in Males: Findings from a Nested Case-Control Study. eLife 2021, 10, e67569. [Google Scholar] [CrossRef]
- Mantovani, S.; Daga, S.; Fallerini, C.; Baldassarri, M.; Benetti, E.; Picchiotti, N.; Fava, F.; Gallì, A.; Zibellini, S.; Bruttini, M.; et al. Rare Variants in Toll-like Receptor 7 Results in Functional Impairment and Downregulation of Cytokine-Mediated Signaling in COVID-19 Patients. Genes Immun. 2022, 23, 51–56. [Google Scholar] [CrossRef]
- Zhang, Q.; Bastard, P.; Liu, Z.; Le Pen, J.; Moncada-Velez, M.; Chen, J.; Ogishi, M.; Sabli, I.K.D.; Hodeib, S.; Korol, C.; et al. Inborn Errors of Type I IFN Immunity in Patients with Life-Threatening COVID-19. Science 2020, 370, eabd4570. [Google Scholar] [CrossRef]
- Kosmicki, J.A.; Horowitz, J.E.; Banerjee, N.; Lanche, R.; Marcketta, A.; Maxwell, E.; Bai, X.; Sun, D.; Backman, J.D.; Sharma, D.; et al. Pan-Ancestry Exome-Wide Association Analyses of COVID-19 Outcomes in 586,157 Individuals. Am. J. Hum. Genet. 2021, 108, 1350–1355. [Google Scholar] [CrossRef]
- Povysil, G.; Butler-Laporte, G.; Shang, N.; Wang, C.; Khan, A.; Alaamery, M.; Nakanishi, T.; Zhou, S.; Forgetta, V.; Eveleigh, R.J.M.; et al. Rare Loss-of-Function Variants in Type I IFN Immunity Genes Are Not Associated with Severe COVID-19. J. Clin. Investig. 2021, 131, e147834. [Google Scholar] [CrossRef]
- Drzymalla, E.; Green, R.F.; Knuth, M.; Khoury, M.J.; Dotson, W.D.; Gundlapalli, A. COVID-19-Related Health Outcomes in People with Primary Immunodeficiency: A Systematic Review. Clin. Immunol. 2022, 243, 109097. [Google Scholar] [CrossRef] [PubMed]
- Bastard, P.; Gervais, A.; Le Voyer, T.; Rosain, J.; Philippot, Q.; Manry, J.; Michailidis, E.; Hoffmann, H.-H.; Eto, S.; Garcia-Prat, M.; et al. Autoantibodies Neutralizing Type I IFNs Are Present in ~4% of Uninfected Individuals over 70 Years Old and Account for ~20% of COVID-19 Deaths. Sci. Immunol. 2021, 6, eabl4340. [Google Scholar] [CrossRef] [PubMed]
- Briquez, P.S.; Rouhani, S.J.; Yu, J.; Pyzer, A.R.; Trujillo, J.; Dugan, H.L.; Stamper, C.T.; Changrob, S.; Sperling, A.I.; Wilson, P.C.; et al. Severe COVID-19 Induces Autoantibodies against Angiotensin II That Correlate with Blood Pressure Dysregulation and Disease Severity. Sci. Adv. 2022, 8, eabn3777. [Google Scholar] [CrossRef] [PubMed]
- Netea, M.G.; Domínguez-Andrés, J.; van de Veerdonk, F.L.; van Crevel, R.; Pulendran, B.; van der Meer, J.W.M. Natural Resistance against Infections: Focus on COVID-19. Trends Immunol. 2022, 43, 106–116. [Google Scholar] [CrossRef] [PubMed]
- Andreakos, E.; Abel, L.; Vinh, D.C.; Kaja, E.; Drolet, B.A.; Zhang, Q.; O’Farrelly, C.; Novelli, G.; Rodríguez-Gallego, C.; Haerynck, F.; et al. A Global Effort to Dissect the Human Genetic Basis of Resistance to SARS-CoV-2 Infection. Nat. Immunol. 2022, 23, 159–164. [Google Scholar] [CrossRef]
- David, A.; Parkinson, N.; Peacock, T.P.; Pairo-Castineira, E.; Khanna, T.; Cobat, A.; Tenesa, A.; Sancho-Shimizu, V.; Casanova, J.-L.; Abel, L.; et al. A Common TMPRSS2 Variant Has a Protective Effect against Severe COVID-19. Curr. Res. Transl. Med. 2022, 70, 103333. [Google Scholar] [CrossRef]
- Verma, A.; Minnier, J.; Wan, E.S.; Huffman, J.E.; Gao, L.; Joseph, J.; Ho, Y.-L.; Wu, W.-C.; Cho, K.; Gorman, B.R.; et al. A MUC5B Gene Polymorphism, Rs35705950-T Confers Protective Effects Against COVID-19 Hospitalization but Not Severe Disease or Mortality. Am. J. Respir. Crit. Care Med. 2022, 206, 1220–1229. [Google Scholar] [CrossRef]
- Shkurnikov, M.; Nersisyan, S.; Jankevic, T.; Galatenko, A.; Gordeev, I.; Vechorko, V.; Tonevitsky, A. Association of HLA Class I Genotypes with Severity of Coronavirus Disease-19. Front. Immunol. 2021, 12, 641900. [Google Scholar] [CrossRef]
- Hernández-Doño, S.; Sánchez-González, R.A.; Trujillo-Vizuet, M.G.; Zamudio-Castellanos, F.Y.; García-Silva, R.; Bulos-Rodríguez, P.; Vazquez-Guzmán, C.A.; Cárdenas-Ramos, X.; de León Rodríguez, D.; Elías, F.; et al. Protective HLA Alleles against Severe COVID-19: HLA-A*68 as an Ancestral Protection Allele in Tapachula-Chiapas, Mexico. Clin. Immunol. 2022, 238, 108990. [Google Scholar] [CrossRef]
- Abdelhafiz, A.S.; Ali, A.; Fouda, M.A.; Sayed, D.M.; Kamel, M.M.; Kamal, L.M.; Khalil, M.A.; Bakry, R.M. HLA-B*15 Predicts Survival in Egyptian Patients with COVID-19. Hum. Immunol. 2022, 83, 10–16. [Google Scholar] [CrossRef] [PubMed]
- Pereira, E.; Felipe, S.; de Freitas, R.; Araújo, V.; Soares, P.; Ribeiro, J.; Henrique dos Santos, L.; Alves, J.O.; Canabrava, N.; van Tilburg, M.; et al. ABO Blood Group and Link to COVID-19: A Comprehensive Review of the Reported Associations and Their Possible Underlying Mechanisms. Microb. Pathog. 2022, 169, 105658. [Google Scholar] [CrossRef] [PubMed]
- Magesh, S.; John, D.; Li, W.T.; Li, Y.; Mattingly-app, A.; Jain, S.; Chang, E.Y.; Ongkeko, W.M. Disparities in COVID-19 Outcomes by Race, Ethnicity, and Socioeconomic Status: A Systematic Review and Meta-Analysis. JAMA Netw. Open 2021, 4, e2134147. [Google Scholar] [CrossRef] [PubMed]
- Parcha, V.; Malla, G.; Suri, S.S.; Kalra, R.; Heindl, B.; Berra, L.; Fouad, M.N.; Arora, G.; Arora, P. Geographic Variation in Racial Disparities in Health and Coronavirus Disease-2019 (COVID-19) Mortality. Mayo Clin. Proc. Innov. Qual. Outcomes 2020, 4, 703–716. [Google Scholar] [CrossRef] [PubMed]
- Muñoz-Price, L.S.; Nattinger, A.B.; Rivera, F.; Hanson, R.; Gmehlin, C.G.; Perez, A.; Singh, S.; Buchan, B.W.; Ledeboer, N.A.; Pezzin, L.E. Racial Disparities in Incidence and Outcomes among Patients with COVID-19. JAMA Netw. Open 2020, 3, e2021892. [Google Scholar] [CrossRef] [PubMed]
- Shelton, J.F.; Shastri, A.J.; Ye, C.; Weldon, C.H.; Filshtein-Sonmez, T.; Coker, D.; Symons, A.; Esparza-Gordillo, J.; Aslibekyan, S.; Auton, A. Trans-Ancestry Analysis Reveals Genetic and Nongenetic Associations with COVID-19 Susceptibility and Severity. Nat. Genet. 2021, 53, 801–808. [Google Scholar] [CrossRef] [PubMed]
- Raharja, A.; Tamara, A.; Kok, L.T. Association Between Ethnicity and Severe COVID-19 Disease: A Systematic Review and Meta-Analysis. J. Racial Ethn. Health Disparities 2021, 8, 1563–1572. [Google Scholar] [CrossRef]
- Li, S.L.; Pereira, R.H.; Prete, C.A., Jr.; Zarebski, A.E.; Emanuel, L.; Alves, P.J.; Peixoto, P.S.; Braga, C.K.; de Souza Santos, A.A.; de Souza, W.M.; et al. Higher Risk of Death from COVID-19 in Low-Income and Non-White Populations of São Paulo, Brazil. BMJ Glob. Health 2021, 6, e004959. [Google Scholar] [CrossRef]
- Zeberg, H.; Pääbo, S. The Major Genetic Risk Factor for Severe COVID-19 Is Inherited from Neanderthals. Nature 2020, 587, 610–612. [Google Scholar] [CrossRef]
- Zeberg, H.; Pääbo, S. A Genomic Region Associated with Protection against Severe COVID-19 Is Inherited from Neandertals. Proc. Natl. Acad. Sci. USA 2021, 118, e2026309118. [Google Scholar] [CrossRef]
- Falagas, M.E.; Mourtzoukou, E.G.; Vardakas, K.Z. Sex Differences in the Incidence and Severity of Respiratory Tract Infections. Respir. Med. 2007, 101, 1845–1863. [Google Scholar] [CrossRef]
- Ursin, R.L.; Klein, S.L. Sex Differences in Respiratory Viral Pathogenesis and Treatments. Annu. Rev. Virol. 2021, 8, 393–414. [Google Scholar] [CrossRef] [PubMed]
- Corica, B.; Tartaglia, F.; D’Amico, T.; Romiti, G.F.; Cangemi, R. Sex and Gender Differences in Community-Acquired Pneumonia. Intern. Emerg. Med. 2022, 17, 1575–1588. [Google Scholar] [CrossRef]
- Sieurin, J.; Brandén, G.; Magnusson, C.; Hergens, M.-P.; Kosidou, K. A Population-Based Cohort Study of Sex and Risk of Severe Outcomes in COVID-19. Eur. J. Epidemiol. 2022, 37, 1159–1169. [Google Scholar] [CrossRef] [PubMed]
- Meijs, D.A.M.; van Bussel, B.C.T.; Stessel, B.; Mehagnoul-Schipper, J.; Hana, A.; Scheeren, C.I.E.; Peters, S.A.E.; van Mook, W.N.K.A.; van der Horst, I.C.C.; Marx, G.; et al. Better COVID-19 Intensive Care Unit Survival in Females, Independent of Age, Disease Severity, Comorbidities, and Treatment. Sci. Rep. 2022, 12, 734. [Google Scholar] [CrossRef]
- Her, A.-Y.; Bhak, Y.; Jun, E.J.; Yuan, S.L.; Garg, S.; Lee, S.; Bhak, J.; Shin, E.-S. Sex-Specific Difference of in-Hospital Mortality from COVID-19 in South Korea. PLoS ONE 2022, 17, e0262861. [Google Scholar] [CrossRef] [PubMed]
- Vahidy, F.S.; Pan, A.P.; Ahnstedt, H.; Munshi, Y.; Choi, H.A.; Tiruneh, Y.; Nasir, K.; Kash, B.A.; Andrieni, J.D.; McCullough, L.D. Sex Differences in Susceptibility, Severity, and Outcomes of Coronavirus Disease 2019: Cross-Sectional Analysis from a Diverse US Metropolitan Area. PLoS ONE 2021, 16, e0245556. [Google Scholar] [CrossRef]
- Peckham, H.; de Gruijter, N.M.; Raine, C.; Radziszewska, A.; Ciurtin, C.; Wedderburn, L.R.; Rosser, E.C.; Webb, K.; Deakin, C.T. Male Sex Identified by Global COVID-19 Meta-Analysis as a Risk Factor for Death and ITU Admission. Nat. Commun. 2020, 11, 6317. [Google Scholar] [CrossRef]
- Bennett, T.D.; Moffitt, R.A.; Hajagos, J.G.; Amor, B.; Anand, A.; Bissell, M.M.; Bradwell, K.R.; Bremer, C.; Byrd, J.B.; Denham, A.; et al. Clinical Characterization and Prediction of Clinical Severity of SARS-CoV-2 Infection among US Adults Using Data from the US National COVID Cohort Collaborative. JAMA Netw. Open 2021, 4, e2116901. [Google Scholar] [CrossRef]
- Gujski, M.; Jankowski, M.; Rabczenko, D.; Goryński, P.; Juszczyk, G. Characteristics and Clinical Outcomes of 116,539 Patients Hospitalized with COVID-19—Poland, March–December 2020. Viruses 2021, 13, 1458. [Google Scholar] [CrossRef]
- Kim, H.-J.; Hwang, H.; Hong, H.; Yim, J.-J.; Lee, J. A Systematic Review and Meta-Analysis of Regional Risk Factors for Critical Outcomes of COVID-19 during Early Phase of the Pandemic. Sci. Rep. 2021, 11, 9784. [Google Scholar] [CrossRef]
- Dessie, Z.G.; Zewotir, T. Mortality-Related Risk Factors of COVID-19: A Systematic Review and Meta-Analysis of 42 Studies and 423,117 Patients. BMC Infect. Dis. 2021, 21, 855. [Google Scholar] [CrossRef] [PubMed]
- Li, X.; Zhong, X.; Wang, Y.; Zeng, X.; Luo, T.; Liu, Q. Clinical Determinants of the Severity of COVID-19: A Systematic Review and Meta-Analysis. PLoS ONE 2021, 16, e0250602. [Google Scholar] [CrossRef]
- Rivero-Calle, I.; Pardo-Seco, J.; Aldaz, P.; Vargas, D.A.; Mascarós, E.; Redondo, E.; Díaz-Maroto, J.L.; Linares-Rufo, M.; Fierro-Alacio, M.J.; Gil, A.; et al. Incidence and Risk Factor Prevalence of Community-Acquired Pneumonia in Adults in Primary Care in Spain (NEUMO-ES-RISK Project). BMC Infect. Dis. 2016, 16, 645. [Google Scholar] [CrossRef] [PubMed]
- Higgins, S.T.; Kurti, A.N.; Redner, R.; White, T.J.; Gaalema, D.E.; Roberts, M.E.; Doogan, N.J.; Tidey, J.W.; Miller, M.E.; Stanton, C.A.; et al. A Literature Review on Prevalence of Gender Differences and Intersections with Other Vulnerabilities to Tobacco Use in the United States, 2004–2014. Prev. Med. 2015, 80, 89–100. [Google Scholar] [CrossRef] [PubMed]
- Chamekh, M.; Deny, M.; Romano, M.; Lefèvre, N.; Corazza, F.; Duchateau, J.; Casimir, G. Differential Susceptibility to Infectious Respiratory Diseases between Males and Females Linked to Sex-Specific Innate Immune Inflammatory Response. Front. Immunol. 2017, 8, 1806. [Google Scholar] [CrossRef] [PubMed]
- Kadel, S.; Kovats, S. Sex Hormones Regulate Innate Immune Cells and Promote Sex Differences in Respiratory Virus Infection. Front. Immunol. 2018, 9, 1653. [Google Scholar] [CrossRef]
- Shepherd, R.; Cheung, A.S.; Pang, K.; Saffery, R.; Novakovic, B. Sexual Dimorphism in Innate Immunity: The Role of Sex Hormones and Epigenetics. Front. Immunol. 2021, 11, 604000. [Google Scholar] [CrossRef]
- Brandi, M.L. Are Sex Hormones Promising Candidates to Explain Sex Disparities in the COVID-19 Pandemic? Rev. Endocr. Metab. Disord. 2022, 23, 171–183. [Google Scholar] [CrossRef]
- Gubbels Bupp, M.R.; Jorgensen, T.N. Androgen-Induced Immunosuppression. Front. Immunol. 2018, 9, 794. [Google Scholar] [CrossRef]
- Decaroli, M.C.; Rochira, V. Aging and Sex Hormones in Males. Virulence 2017, 8, 545–570. [Google Scholar] [CrossRef]
- Rastrelli, G.; Di Stasi, V.; Inglese, F.; Beccaria, M.; Garuti, M.; Di Costanzo, D.; Spreafico, F.; Greco, G.F.; Cervi, G.; Pecoriello, A.; et al. Low Testosterone Levels Predict Clinical Adverse Outcomes in SARS-CoV-2 Pneumonia Patients. Andrology 2021, 9, 88–98. [Google Scholar] [CrossRef] [PubMed]
- Pagano, M.T.; Peruzzu, D.; Busani, L.; Pierdominici, M.; Ruggieri, A.; Antinori, A.; D’Offizi, G.; Petrosillo, N.; Palmieri, F.; Piselli, P.; et al. Predicting Respiratory Failure in Patients Infected by SARS-CoV-2 by Admission Sex-Specific Biomarkers. Biol. Sex Differ. 2021, 12, 63. [Google Scholar] [CrossRef] [PubMed]
- Baldassarri, M.; Picchiotti, N.; Fava, F.; Fallerini, C.; Benetti, E.; Daga, S.; Valentino, F.; Doddato, G.; Furini, S.; Giliberti, A.; et al. Shorter Androgen Receptor PolyQ Alleles Protect against Life-Threatening COVID-19 Disease in European Males. EBioMedicine 2021, 65, 103246. [Google Scholar] [CrossRef]
- Montaño, L.M.; Sommer, B.; Solís-Chagoyán, H.; Romero-Martínez, B.S.; Aquino-Gálvez, A.; Gomez-Verjan, J.C.; Calixto, E.; González-Avila, G.; Flores-Soto, E. Could Lower Testosterone in Older Men Explain Higher COVID-19 Morbidity and Mortalities? Int. J. Mol. Sci. 2022, 23, 935. [Google Scholar] [CrossRef] [PubMed]
- Traish, A.; Bolanos, J.; Nair, S.; Saad, F.; Morgentaler, A. Do Androgens Modulate the Pathophysiological Pathways of Inflammation? Appraising the Contemporary Evidence. J. Clin. Med. 2018, 7, 549. [Google Scholar] [CrossRef] [PubMed]
- Lin, B.; Ferguson, C.; White, J.T.; Wang, S.; Vessella, R.; True, L.D.; Hood, L.; Nelson, P.S. Prostate-Localized and Androgen-Regulated Expression of the Membrane-Bound Serine Protease TMPRSS2. Cancer Res. 1999, 59, 4180–4184. [Google Scholar]
- Wang, X.-W.; Hu, H.; Xu, Z.-Y.; Zhang, G.-K.; Yu, Q.-H.; Yang, H.-L.; Zheng, J.-H. Association of Menopausal Status with COVID-19 Outcomes: A Propensity Score Matching Analysis. Biol. Sex Differ. 2021, 12, 16. [Google Scholar] [CrossRef]
- Sund, M.; Fonseca-Rodríguez, O.; Josefsson, A.; Welen, K.; Connolly, A.-M.F. Association between Pharmaceutical Modulation of Oestrogen in Postmenopausal Women in Sweden and Death Due to COVID-19: A Cohort Study. BMJ Open 2022, 12, e053032. [Google Scholar] [CrossRef]
- Costeira, R.; Lee, K.A.; Murray, B.; Christiansen, C.; Castillo-Fernandez, J.; Lochlainn, M.N.; Pujol, J.C.; Macfarlane, H.; Kenny, L.C.; Buchan, I.; et al. Estrogen and COVID-19 Symptoms: Associations in Women from the COVID Symptom Study. PLoS ONE 2021, 16, e0257051. [Google Scholar] [CrossRef]
- Mompeón, A.; Lázaro-Franco, M.; Bueno-Betí, C.; Pérez-Cremades, D.; Vidal-Gómez, X.; Monsalve, E.; Gironacci, M.M.; Hermenegildo, C.; Novella, S. Estradiol, Acting through ERα, Induces Endothelial Non-Classic Renin-Angiotensin System Increasing Angiotensin 1-7 Production. Mol. Cell. Endocrinol. 2016, 422, 1–8. [Google Scholar] [CrossRef]
- Baristaite, G.; Gurwitz, D. Estradiol Reduces ACE2 and TMPRSS2 MRNA Levels in A549 Human Lung Epithelial Cells. Drug Dev. Res. 2022, 83, 961–966. [Google Scholar] [CrossRef] [PubMed]
- Kovats, S. Estrogen Receptors Regulate Innate Immune Cells and Signaling Pathways. Cell. Immunol. 2015, 294, 63–69. [Google Scholar] [CrossRef] [PubMed]
- Pernis, A.B. Estrogen and CD4+ T Cells. Curr. Opin. Rheumatol. 2007, 19, 414–420. [Google Scholar] [CrossRef] [PubMed]
- Sharma, G.; Prossnitz, E.R. Mechanisms of Estradiol-Induced Insulin Secretion by the G Protein-Coupled Estrogen Receptor GPR30/GPER in Pancreatic β-Cells. Endocrinology 2011, 152, 3030–3039. [Google Scholar] [CrossRef] [PubMed]
- Iorga, A.; Cunningham, C.M.; Moazeni, S.; Ruffenach, G.; Umar, S.; Eghbali, M. The Protective Role of Estrogen and Estrogen Receptors in Cardiovascular Disease and the Controversial Use of Estrogen Therapy. Biol. Sex Differ. 2017, 8, 33. [Google Scholar] [CrossRef] [PubMed]
- Asselta, R.; Paraboschi, E.M.; Mantovani, A.; Duga, S. ACE2 and TMPRSS2 Variants and Expression as Candidates to Sex and Country Differences in COVID-19 Severity in Italy. Aging 2020, 12, 10087–10098. [Google Scholar] [CrossRef]
- Baughn, L.B.; Sharma, N.; Elhaik, E.; Sekulic, A.; Bryce, A.H.; Fonseca, R. Targeting TMPRSS2 in SARS-CoV-2 Infection. Mayo Clin. Proc. 2020, 95, 1989–1999. [Google Scholar] [CrossRef]
- Xie, X.; Xudong, X.; Chen, J.; Junzhu, C.; Wang, X.; Xingxiang, W.; Zhang, F.; Furong, Z.; Liu, Y.; Yanrong, L. Age- and Gender-Related Difference of ACE2 Expression in Rat Lung. Life Sci. 2006, 78, 2166–2171. [Google Scholar] [CrossRef]
- Okwan-Duodu, D.; Lim, E.-C.; You, S.; Engman, D.M. TMPRSS2 Activity May Mediate Sex Differences in COVID-19 Severity. Signal Transduct. Target. Ther. 2021, 6, 100. [Google Scholar] [CrossRef]
- Spiering, A.E.; de Vries, T.J. Why Females Do Better: The X Chromosomal TLR7 Gene-Dose Effect in COVID-19. Front. Immunol. 2021, 12, 756262. [Google Scholar] [CrossRef]
- Maan, A.A.; Eales, J.; Akbarov, A.; Rowland, J.; Xu, X.; Jobling, M.A.; Charchar, F.J.; Tomaszewski, M. The Y Chromosome: A Blueprint for Men’s Health? Eur. J. Hum. Genet. 2017, 25, 1181–1188. [Google Scholar] [CrossRef] [PubMed]
- Dumanski, J.P.; Halvardson, J.; Davies, H.; Rychlicka-Buniowska, E.; Mattisson, J.; Moghadam, B.T.; Nagy, N.; Węglarczyk, K.; Bukowska-Strakova, K.; Danielsson, M.; et al. Immune Cells Lacking Y Chromosome Show Dysregulation of Autosomal Gene Expression. Cell. Mol. Life Sci. 2021, 78, 4019–4033. [Google Scholar] [CrossRef]
- Thompson, D.J.; Genovese, G.; Halvardson, J.; Ulirsch, J.C.; Wright, D.J.; Terao, C.; Davidsson, O.B.; Day, F.R.; Sulem, P.; Jiang, Y.; et al. Genetic Predisposition to Mosaic Y Chromosome Loss in Blood. Nature 2019, 575, 652–657. [Google Scholar] [CrossRef] [PubMed]
- Klein, S.L.; Flanagan, K.L. Sex Differences in Immune Responses. Nat. Rev. Immunol. 2016, 16, 626–638. [Google Scholar] [CrossRef] [PubMed]
- Francis, M.E.; Richardson, B.; Goncin, U.; McNeil, M.; Rioux, M.; Foley, M.K.; Ge, A.; Pechous, R.D.; Kindrachuk, J.; Cameron, C.M.; et al. Sex and Age Bias Viral Burden and Interferon Responses during SARS-CoV-2 Infection in Ferrets. Sci. Rep. 2021, 11, 14536. [Google Scholar] [CrossRef] [PubMed]
- Jin, S.; An, H.; Zhou, T.; Li, T.; Xie, M.; Chen, S.; Chen, C.; Ying, B.; Xu, Z.; Li, X.; et al. Sex- and Age-Specific Clinical and Immunological Features of Coronavirus Disease 2019. PLOS Pathog. 2021, 17, e1009420. [Google Scholar] [CrossRef]
- Takahashi, T.; Ellingson, M.K.; Wong, P.; Israelow, B.; Lucas, C.; Klein, J.; Silva, J.; Mao, T.; Oh, J.E.; Tokuyama, M.; et al. Sex Differences in Immune Responses That Underlie COVID-19 Disease Outcomes. Nature 2020, 588, 315–320. [Google Scholar] [CrossRef]
- Shattuck-Heidorn, H.; Danielsen, A.C.; Gompers, A.; Bruch, J.D.; Zhao, H.; Boulicault, M.; Marsella, J.; Richardson, S.S. A Finding of Sex Similarities Rather than Differences in COVID-19 Outcomes. Nature 2021, 597, E7–E9. [Google Scholar] [CrossRef]
- Lieberman, N.A.P.; Peddu, V.; Xie, H.; Shrestha, L.; Huang, M.-L.; Mears, M.C.; Cajimat, M.N.; Bente, D.A.; Shi, P.-Y.; Bovier, F.; et al. In Vivo Antiviral Host Transcriptional Response to SARS-CoV-2 by Viral Load, Sex, and Age. PLoS Biol. 2020, 18, e3000849. [Google Scholar] [CrossRef]
- Vassilaki, N.; Gargalionis, A.N.; Bletsa, A.; Papamichalopoulos, N.; Kontou, E.; Gkika, M.; Patas, K.; Theodoridis, D.; Manolis, I.; Ioannidis, A.; et al. Impact of Age and Sex on Antibody Response Following the Second Dose of COVID-19 BNT162b2 MRNA Vaccine in Greek Healthcare Workers. Microorganisms 2021, 9, 1725. [Google Scholar] [CrossRef]
- Bastard, P.; Rosen, L.B.; Zhang, Q.; Michailidis, E.; Hoffmann, H.-H.; Zhang, Y.; Dorgham, K.; Philippot, Q.; Rosain, J.; Béziat, V.; et al. Autoantibodies against Type I IFNs in Patients with Life-Threatening COVID-19. Science 2020, 370, eabd4585. [Google Scholar] [CrossRef] [PubMed]
- Liu, Y.; Ebinger, J.E.; Mostafa, R.; Budde, P.; Gajewski, J.; Walker, B.; Joung, S.; Wu, M.; Bräutigam, M.; Hesping, F.; et al. Paradoxical Sex-Specific Patterns of Autoantibody Response to SARS-CoV-2 Infection. J. Transl. Med. 2021, 19, 524. [Google Scholar] [CrossRef] [PubMed]
- Scully, E.P.; Schumock, G.; Fu, M.; Massaccesi, G.; Muschelli, J.; Betz, J.; Klein, E.Y.; West, N.E.; Robinson, M.; Garibaldi, B.T.; et al. Sex and Gender Differences in Testing, Hospital Admission, Clinical Presentation, and Drivers of Severe Outcomes from COVID-19. Open Forum Infect. Dis. 2021, 8, ofab448. [Google Scholar] [CrossRef] [PubMed]
- Sylvester, S.V.; Rusu, R.; Chan, B.; Bellows, M.; O’Keefe, C.; Nicholson, S. Sex Differences in Sequelae from COVID-19 Infection and in Long COVID Syndrome: A Review. Curr. Med. Res. Opin. 2022, 38, 1391–1399. [Google Scholar] [CrossRef]
- Bucciarelli, V.; Nasi, M.; Bianco, F.; Seferovic, J.; Ivkovic, V.; Gallina, S.; Mattioli, A.V. Depression Pandemic and Cardiovascular Risk in the COVID-19 Era and Long COVID Syndrome: Gender Makes a Difference. Trends Cardiovasc. Med. 2022, 32, 12–17. [Google Scholar] [CrossRef]
- Robinson, D.P.; Lorenzo, M.E.; Jian, W.; Klein, S.L. Elevated 17β-Estradiol Protects Females from Influenza A Virus Pathogenesis by Suppressing Inflammatory Responses. PLoS Pathog. 2011, 7, e1002149. [Google Scholar] [CrossRef]
- Di Mascio, D.; Khalil, A.; Saccone, G.; Rizzo, G.; Buca, D.; Liberati, M.; Vecchiet, J.; Nappi, L.; Scambia, G.; Berghella, V.; et al. Outcome of Coronavirus Spectrum Infections (SARS, MERS, COVID-19) during Pregnancy: A Systematic Review and Meta-Analysis. Am. J. Obstet. Gynecol. MFM 2020, 2, 100107. [Google Scholar] [CrossRef]
- Huntley, B.J.F.; Mulder, I.A.; Di Mascio, D.; Vintzileos, W.S.; Vintzileos, A.M.; Berghella, V.; Chauhan, S.P. Adverse Pregnancy Outcomes among Individuals with and without Severe Acute Respiratory Syndrome Coronavirus 2 (SARS-CoV-2): A Systematic Review and Meta-Analysis. Obstet. Gynecol. 2021, 137, 585–596. [Google Scholar] [CrossRef]
- Gale, C.; Quigley, M.A.; Placzek, A.; Knight, M.; Ladhani, S.; Draper, E.S.; Sharkey, D.; Doherty, C.; Mactier, H.; Kurinczuk, J.J. Characteristics and Outcomes of Neonatal SARS-CoV-2 Infection in the UK: A Prospective National Cohort Study Using Active Surveillance. Lancet Child Adolesc. Health 2021, 5, 113–121. [Google Scholar] [CrossRef]
- Di Toro, F.; Gjoka, M.; Di Lorenzo, G.; De Santo, D.; De Seta, F.; Maso, G.; Risso, F.M.; Romano, F.; Wiesenfeld, U.; Levi-D’Ancona, R.; et al. Impact of COVID-19 on Maternal and Neonatal Outcomes: A Systematic Review and Meta-Analysis. Clin. Microbiol. Infect. 2021, 27, 36–46. [Google Scholar] [CrossRef]
- Martinez-Portilla, R.J.; Sotiriadis, A.; Chatzakis, C.; Torres-Torres, J.; Espino y Sosa, S.; Sandoval-Mandujano, K.; Castro-Bernabe, D.A.; Medina-Jimenez, V.; Monarrez-Martin, J.C.; Figueras, F.; et al. Pregnant Women with SARS-CoV-2 Infection Are at Higher Risk of Death and Pneumonia: Propensity Score Matched Analysis of a Nationwide Prospective Cohort (COV19Mx). Ultrasound Obstet. Gynecol. 2021, 57, 224–231. [Google Scholar] [CrossRef] [PubMed]
- Allotey, J.; Stallings, E.; Bonet, M.; Yap, M.; Chatterjee, S.; Kew, T.; Debenham, L.; Llavall, A.C.; Dixit, A.; Zhou, D.; et al. Clinical Manifestations, Risk Factors, and Maternal and Perinatal Outcomes of Coronavirus Disease 2019 in Pregnancy: Living Systematic Review and Meta-Analysis. BMJ 2020, 370, m3320. [Google Scholar] [CrossRef] [PubMed]
- Rozo, N.; Valencia, D.; Newton, S.M.; Avila, G.; Gonzalez, M.A.; Sancken, C.L.; Burkel, V.K.; Ellington, S.R.; Gilboa, S.M.; Rao, C.Y.; et al. Severity of Illness by Pregnancy Status among Laboratory-Confirmed SARS-CoV-2 Infections Occurring in Reproductive-Aged Women in Colombia. Paediatr. Perinat. Epidemiol. 2022, 36, 456–465. [Google Scholar] [CrossRef]
- Marchand, G.; Patil, A.S.; Masoud, A.T.; Ware, K.; King, A.; Ruther, S.; Brazil, G.; Calteux, N.; Ulibarri, H.; Parise, J.; et al. Systematic Review and Meta-Analysis of COVID-19 Maternal and Neonatal Clinical Features and Pregnancy Outcomes up to June 3, 2021. AJOG Glob. Rep. 2022, 2, 100049. [Google Scholar] [CrossRef] [PubMed]
- Wei, S.Q.; Bilodeau-Bertrand, M.; Liu, S.; Auger, N. The Impact of COVID-19 on Pregnancy Outcomes: A Systematic Review and Meta-Analysis. Can. Med. Assoc. J. 2021, 193, E540–E548. [Google Scholar] [CrossRef]
- Cruz Melguizo, S.; de la Cruz Conty, M.L.; Carmona Payán, P.; Abascal-Saiz, A.; Pintando Recarte, P.; González Rodríguez, L.; Cuenca Marín, C.; Martínez Varea, A.; Oreja Cuesta, A.B.; Rodríguez, P.P.; et al. Pregnancy Outcomes and SARS-CoV-2 Infection: The Spanish Obstetric Emergency Group Study. Viruses 2021, 13, 853. [Google Scholar] [CrossRef]
- Engjom, H.; Ramakrishnan, R.; Vousden, N.; Bunch, K.; Morris, E.; Simpson, N.A.B.; Gale, C.; O’Brien, P.; Quigley, M.; Brocklehurst, P.; et al. Severity of Maternal Infection and Perinatal Outcomes during Periods in Which Wildtype, Alpha and Delta SARS-CoV-2 Variants Were Dominant: Data from the UK Obstetric Surveillance System National Cohort Study. BMJ Med. 2022, 1, e000190. [Google Scholar] [CrossRef]
- Doyle, T.J.; Kiros, G.; Schmitt-Matzen, E.N.; Propper, R.; Thompson, A.; Phillips-Bell, G.S. Maternal and Perinatal Outcomes Associated with SARS-CoV-2 Infection during Pregnancy, Florida, 2020–2021: A Retrospective Cohort Study. Clin. Infect. Dis. 2022, 75, ciac441. [Google Scholar] [CrossRef]
- Ciapponi, A.; Bardach, A.; Comandé, D.; Berrueta, M.; Argento, F.J.; Rodriguez Cairoli, F.; Zamora, N.; Santa María, V.; Xiong, X.; Zaraa, S.; et al. COVID-19 and Pregnancy: An Umbrella Review of Clinical Presentation, Vertical Transmission, and Maternal and Perinatal Outcomes. PLoS ONE 2021, 16, e0253974. [Google Scholar] [CrossRef]
- Norman, M.; Navér, L.; Söderling, J.; Ahlberg, M.; Hervius Askling, H.; Aronsson, B.; Byström, E.; Jonsson, J.; Sengpiel, V.; Ludvigsson, J.F.; et al. Association of Maternal SARS-CoV-2 Infection in Pregnancy with Neonatal Outcomes. JAMA 2021, 325, 2076–2086. [Google Scholar] [CrossRef]
- Crovetto, F.; Crispi, F.; Llurba, E.; Pascal, R.; Larroya, M.; Trilla, C.; Camacho, M.; Medina, C.; Dobaño, C.; Gomez-Roig, M.D.; et al. Impact of Severe Acute Respiratory Syndrome Coronavirus 2 Infection on Pregnancy Outcomes: A Population-Based Study. Clin. Infect. Dis. 2021, 73, 1768–1775. [Google Scholar] [CrossRef] [PubMed]
- Regan, A.K.; Arah, O.A.; Fell, D.B.; Sullivan, S.G. SARS-CoV-2 Infection During Pregnancy and Associated Perinatal Health Outcomes: A National US Cohort Study. J. Infect. Dis. 2022, 225, 759–767. [Google Scholar] [CrossRef] [PubMed]
- Metz, T.D.; Clifton, R.G.; Hughes, B.L.; Sandoval, G.J.; Grobman, W.A.; Saade, G.R.; Manuck, T.A.; Longo, M.; Sowles, A.; Clark, K.; et al. Association of SARS-CoV-2 Infection with Serious Maternal Morbidity and Mortality from Obstetric Complications. JAMA 2022, 327, 748–759. [Google Scholar] [CrossRef] [PubMed]
- Gurol-Urganci, I.; Jardine, J.E.; Carroll, F.; Draycott, T.; Dunn, G.; Fremeaux, A.; Harris, T.; Hawdon, J.; Morris, E.; Muller, P.; et al. Maternal and Perinatal Outcomes of Pregnant Women with SARS-CoV-2 Infection at the Time of Birth in England: National Cohort Study. Am. J. Obstet. Gynecol. 2021, 225, 522.e1–522.e11. [Google Scholar] [CrossRef] [PubMed]
- Piekos, S.N.; Roper, R.T.; Hwang, Y.M.; Sorensen, T.; Price, N.D.; Hood, L.; Hadlock, J.J. The Effect of Maternal SARS-CoV-2 Infection Timing on Birth Outcomes: A Retrospective Multicentre Cohort Study. Lancet Digit. Health 2022, 4, e95–e104. [Google Scholar] [CrossRef] [PubMed]
- Pique-Regi, R.; Romero, R.; Tarca, A.L.; Luca, F.; Xu, Y.; Alazizi, A.; Leng, Y.; Hsu, C.-D.; Gomez-Lopez, N. Does the Human Placenta Express the Canonical Cell Entry Mediators for SARS-CoV-2? eLife 2020, 9, e58716. [Google Scholar] [CrossRef]
- Ashary, N.; Bhide, A.; Chakraborty, P.; Colaco, S.; Mishra, A.; Chhabria, K.; Jolly, M.K.; Modi, D. Single-Cell RNA-Seq Identifies Cell Subsets in Human Placenta That Highly Expresses Factors Driving Pathogenesis of SARS-CoV-2. Front. Cell Dev. Biol. 2020, 8, 783. [Google Scholar] [CrossRef]
- Fahmi, A.; Brügger, M.; Démoulins, T.; Zumkehr, B.; Oliveira Esteves, B.I.; Bracher, L.; Wotzkow, C.; Blank, F.; Thiel, V.; Baud, D.; et al. SARS-CoV-2 Can Infect and Propagate in Human Placenta Explants. Cell Rep. Med. 2021, 2, 100456. [Google Scholar] [CrossRef]
- Lu-Culligan, A.; Chavan, A.R.; Vijayakumar, P.; Irshaid, L.; Courchaine, E.M.; Milano, K.M.; Tang, Z.; Pope, S.D.; Song, E.; Vogels, C.B.F.; et al. Maternal Respiratory SARS-CoV-2 Infection in Pregnancy Is Associated with a Robust Inflammatory Response at the Maternal-Fetal Interface. Med 2021, 2, 591–610. [Google Scholar] [CrossRef]
- Verma, S.; Joshi, C.S.; Silverstein, R.B.; He, M.; Carter, E.B.; Mysorekar, I.U. SARS-CoV-2 Colonization of Maternal and Fetal Cells of the Human Placenta Promotes Alteration of Local Renin-Angiotensin System. Med 2021, 2, 575–590. [Google Scholar] [CrossRef]
- Allotey, J.; Chatterjee, S.; Kew, T.; Gaetano, A.; Stallings, E.; Fernández-García, S.; Yap, M.; Sheikh, J.; Lawson, H.; Coomar, D.; et al. SARS-CoV-2 Positivity in Offspring and Timing of Mother-to-Child Transmission: Living Systematic Review and Meta-Analysis. BMJ 2022, 376, e067696. [Google Scholar] [CrossRef] [PubMed]
- World Health Organization. Definition and Categorization of the Timing of Mother-to-Child Transmission of SARS-CoV-2: Scientific Brief, 8 February 2021; World Health Organization: Geneva, Switzerland, 2021. [Google Scholar]
- Babaei, R.; Bokharaei-Salim, F.; Khanaliha, K.; Kiani, S.J.; Marjani, A.; Garshasbi, S.; Dehghani-Dehej, F.; Chavoshpour, S. Prevalence of SARS-CoV-2 Infection in Neonates Born to Mothers or Relatives with COVID-19. BMC Infect. Dis. 2022, 22, 730. [Google Scholar] [CrossRef] [PubMed]
- Rodríguez-Serrano, D.A.; Roy-Vallejo, E.; Zurita Cruz, N.D.; Martín Ramírez, A.; Rodríguez-García, S.C.; Arevalillo-Fernández, N.; Galván-Román, J.M.; Fontán García-Rodrigo, L.; Vega-Piris, L.; Chicot Llano, M.; et al. Detection of SARS-CoV-2 RNA in Serum Is Associated with Increased Mortality Risk in Hospitalized COVID-19 Patients. Sci. Rep. 2021, 11, 13134. [Google Scholar] [CrossRef] [PubMed]
- Garcia-Flores, V.; Romero, R.; Xu, Y.; Theis, K.R.; Arenas-Hernandez, M.; Miller, D.; Peyvandipour, A.; Bhatti, G.; Galaz, J.; Gershater, M.; et al. Maternal-Fetal Immune Responses in Pregnant Women Infected with SARS-CoV-2. Nat. Commun. 2022, 13, 320. [Google Scholar] [CrossRef] [PubMed]
- Suhren, J.-T.; Meinardus, A.; Hussein, K.; Schaumann, N. Meta-Analysis on COVID-19-Pregnancy-Related Placental Pathologies Shows No Specific Pattern. Placenta 2022, 117, 72–77. [Google Scholar] [CrossRef] [PubMed]
- Dang, D.; Wang, L.; Zhang, C.; Li, Z.; Wu, H. Potential Effects of SARS-CoV-2 Infection during Pregnancy on Fetuses and Newborns Are Worthy of Attention. J. Obstet. Gynaecol. Res. 2020, 46, 1951–1957. [Google Scholar] [CrossRef]
- Foo, S.-S.; Cambou, M.C.; Mok, T.; Fajardo, V.M.; Jung, K.L.; Fuller, T.; Chen, W.; Kerin, T.; Mei, J.; Bhattacharya, D.; et al. The Systemic Inflammatory Landscape of COVID-19 in Pregnancy: Extensive Serum Proteomic Profiling of Mother-Infant Dyads with in Utero SARS-CoV-2. Cell Rep. Med. 2021, 2, 100453. [Google Scholar] [CrossRef]
- Ge, E.; Li, Y.; Wu, S.; Candido, E.; Wei, X. Association of Pre-Existing Comorbidities with Mortality and Disease Severity among 167,500 Individuals with COVID-19 in Canada: A Population-Based Cohort Study. PLoS ONE 2021, 16, e0258154. [Google Scholar] [CrossRef]
- Gutierrez, J.P.; Bertozzi, S.M. Non-Communicable Diseases and Inequalities Increase Risk of Death among COVID-19 Patients in Mexico. PLoS ONE 2020, 15, e0240394. [Google Scholar] [CrossRef]
- Pardhan, S.; Wood, S.; Vaughan, M.; Trott, M. The Risk of COVID-19 Related Hospitalsation, Intensive Care Unit Admission and Mortality in People with Underlying Asthma or COPD: A Systematic Review and Meta-Analysis. Front. Med. 2021, 8, 668808. [Google Scholar] [CrossRef]
- Mahamat-Saleh, Y.; Fiolet, T.; Rebeaud, M.E.; Mulot, M.; Guihur, A.; Fatouhi, D.E.; Laouali, N.; Peiffer-Smadja, N.; Aune, D.; Severi, G. Diabetes, Hypertension, Body Mass Index, Smoking and COVID-19-Related Mortality: A Systematic Review and Meta-Analysis of Observational Studies. BMJ Open 2021, 11, e052777. [Google Scholar] [CrossRef] [PubMed]
- Bae, S.; Kim, S.R.; Kim, M.-N.; Shim, W.J.; Park, S.-M. Impact of Cardiovascular Disease and Risk Factors on Fatal Outcomes in Patients with COVID-19 According to Age: A Systematic Review and Meta-Analysis. Heart 2021, 107, 373–380. [Google Scholar] [CrossRef] [PubMed]
- Cai, R.; Zhang, J.; Zhu, Y.; Liu, L.; Liu, Y.; He, Q. Mortality in Chronic Kidney Disease Patients with COVID-19: A Systematic Review and Meta-Analysis. Int. Urol. Nephrol. 2021, 53, 1623–1629. [Google Scholar] [CrossRef] [PubMed]
- Cai, Z.; Yang, Y.; Zhang, J. Obesity Is Associated with Severe Disease and Mortality in Patients with Coronavirus Disease 2019 (COVID-19): A Meta-Analysis. BMC Public Health 2021, 21, 1505. [Google Scholar] [CrossRef] [PubMed]
- Sohrabi, M.-R.; Amin, R.; Maher, A.; Bahadorimonfared, A.; Janbazi, S.; Hannani, K.; Kolahi, A.-A.; Zali, A.-R. Sociodemographic Determinants and Clinical Risk Factors Associated with COVID-19 Severity: A Cross-Sectional Analysis of over 200,000 Patients in Tehran, Iran. BMC Infect. Dis. 2021, 21, 474. [Google Scholar] [CrossRef] [PubMed]
- Han, S.; Zhuang, Q.; Chiang, J.; Tan, S.H.; Chua, G.W.Y.; Xie, C.; Chua, M.L.K.; Soon, Y.Y.; Yang, V.S. Impact of Cancer Diagnoses on the Outcomes of Patients with COVID-19: A Systematic Review and Meta-Analysis. BMJ Open 2022, 12, e044661. [Google Scholar] [CrossRef]
- The Top 10 Causes of Death. Available online: https://www.who.int/news-room/fact-sheets/detail/the-top-10-causes-of-death (accessed on 14 June 2022).
- Gerayeli, F.V.; Milne, S.; Cheung, C.; Li, X.; Yang, C.W.T.; Tam, A.; Choi, L.H.; Bae, A.; Sin, D.D. COPD and the Risk of Poor Outcomes in COVID-19: A Systematic Review and Meta-Analysis. EClinicalMedicine 2021, 33, 100789. [Google Scholar] [CrossRef]
- Agustí, A.; Hogg, J.C. Update on the Pathogenesis of Chronic Obstructive Pulmonary Disease. N. Engl. J. Med. 2019, 381, 1248–1256. [Google Scholar] [CrossRef]
- Higham, A.; Mathioudakis, A.; Vestbo, J.; Singh, D. COVID-19 and COPD: A Narrative Review of the Basic Science and Clinical Outcomes. Eur. Respir. Rev. 2020, 29, 200199. [Google Scholar] [CrossRef]
- Mallia, P.; Message, S.D.; Gielen, V.; Contoli, M.; Gray, K.; Kebadze, T.; Aniscenko, J.; Laza-Stanca, V.; Edwards, M.R.; Slater, L.; et al. Experimental Rhinovirus Infection as a Human Model of Chronic Obstructive Pulmonary Disease Exacerbation. Am. J. Respir. Crit. Care Med. 2011, 183, 734–742. [Google Scholar] [CrossRef]
- Barnes, P.J. Inflammatory Mechanisms in Patients with Chronic Obstructive Pulmonary Disease. J. Allergy Clin. Immunol. 2016, 138, 16–27. [Google Scholar] [CrossRef] [PubMed]
- Song, J.; Zeng, M.; Wang, H.; Qin, C.; Hou, H.; Sun, Z.; Xu, S.; Wang, G.; Guo, C.; Deng, Y.; et al. Distinct Effects of Asthma and COPD Comorbidity on Disease Expression and Outcome in Patients with COVID-19. Allergy 2021, 76, 483–496. [Google Scholar] [CrossRef] [PubMed]
- Leung, J.M.; Yang, C.X.; Tam, A.; Shaipanich, T.; Hackett, T.-L.; Singhera, G.K.; Dorscheid, D.R.; Sin, D.D. ACE-2 Expression in the Small Airway Epithelia of Smokers and COPD Patients: Implications for COVID-19. Eur. Respir. J. 2020, 55, 2000688. [Google Scholar] [CrossRef] [PubMed]
- Milne, S.; Yang, C.X.; Timens, W.; Bossé, Y.; Sin, D.D. SARS-CoV-2 Receptor ACE2 Gene Expression and RAAS Inhibitors. Lancet Respir. Med. 2020, 8, e50–e51. [Google Scholar] [CrossRef] [PubMed]
- Mulpuru, S.; Li, L.; Ye, L.; Hatchette, T.; Andrew, M.K.; Ambrose, A.; Boivin, G.; Bowie, W.; Chit, A.; Santos, G.D.; et al. Effectiveness of Influenza Vaccination on Hospitalizations and Risk Factors for Severe Outcomes in Hospitalized Patients with COPD. CHEST 2019, 155, 69–78. [Google Scholar] [CrossRef]
- Restrepo, M.I.; Mortensen, E.M.; Pugh, J.A.; Anzueto, A. COPD Is Associated with Increased Mortality in Patients with Community-Acquired Pneumonia. Eur. Respir. J. 2006, 28, 346–351. [Google Scholar] [CrossRef]
- Bafadhel, M.; McKenna, S.; Terry, S.; Mistry, V.; Reid, C.; Haldar, P.; McCormick, M.; Haldar, K.; Kebadze, T.; Duvoix, A.; et al. Acute Exacerbations of Chronic Obstructive Pulmonary Disease: Identification of Biologic Clusters and Their Biomarkers. Am. J. Respir. Crit. Care Med. 2011, 184, 662–671. [Google Scholar] [CrossRef]
- George, S.N.; Garcha, D.S.; Mackay, A.J.; Patel, A.R.C.; Singh, R.; Sapsford, R.J.; Donaldson, G.C.; Wedzicha, J.A. Human Rhinovirus Infection during Naturally Occurring COPD Exacerbations. Eur. Respir. J. 2014, 44, 87–96. [Google Scholar] [CrossRef]
- Wilkinson, T.M.A.; Hurst, J.R.; Perera, W.R.; Wilks, M.; Donaldson, G.C.; Wedzicha, J.A. Effect of Interactions between Lower Airway Bacterial and Rhinoviral Infection in Exacerbations of COPD. Chest 2006, 129, 317–324. [Google Scholar] [CrossRef]
- Mathioudakis, A.G.; Janssens, W.; Sivapalan, P.; Singanayagam, A.; Dransfield, M.T.; Jensen, J.-U.S.; Vestbo, J. Acute Exacerbations of Chronic Obstructive Pulmonary Disease: In Search of Diagnostic Biomarkers and Treatable Traits. Thorax 2020, 75, 520–527. [Google Scholar] [CrossRef]
- Vanfleteren, L.E.G.W.; Spruit, M.A.; Wouters, E.F.M.; Franssen, F.M.E. Management of Chronic Obstructive Pulmonary Disease beyond the Lungs. Lancet Respir. Med. 2016, 4, 911–924. [Google Scholar] [CrossRef]
- Terzano, C.; Colamesta, V.; Unim, B.; Romani, S.; Meneghini, A.; Volpe, G.; La Torre, G. Chronic Obstructive Pulmonary Disease (COPD) Exacerbation: Impact of Comorbidities on Length and Costs during Hospitalization. Eur. Rev. Med. Pharmacol. Sci. 2017, 21, 3680–3689. [Google Scholar] [CrossRef] [PubMed]
- Murphy, T.E.; McAvay, G.J.; Allore, H.G.; Stamm, J.A.; Simonelli, P.F. Contributions of COPD, Asthma, and Ten Comorbid Conditions to Health Care Utilization and Patient-Centered Outcomes among US Adults with Obstructive Airway Disease. Int. J. Chron. Obstruct. Pulmon. Dis. 2017, 12, 2515–2522. [Google Scholar] [CrossRef] [PubMed]
- Afonso, A.S.M.; Verhamme, K.M.C.; Sturkenboom, M.C.J.M.; Brusselle, G.G.O. COPD in the General Population: Prevalence, Incidence and Survival. Respir. Med. 2011, 105, 1872–1884. [Google Scholar] [CrossRef] [PubMed]
- Podolanczuk, A.J.; Richeldi, L. COVID-19 and Interstitial Lung Disease: Keep Them Separate. Am. J. Respir. Crit. Care Med. 2020, 202, 1614–1616. [Google Scholar] [CrossRef]
- Castelino, F.V.; Varga, J. Interstitial Lung Disease in Connective Tissue Diseases: Evolving Concepts of Pathogenesis and Management. Arthritis Res. Ther. 2010, 12, 213. [Google Scholar] [CrossRef]
- Lee, H.; Choi, H.; Yang, B.; Lee, S.-K.; Park, T.S.; Park, D.W.; Moon, J.-Y.; Kim, T.-H.; Sohn, J.W.; Yoon, H.J.; et al. Interstitial Lung Disease Increases Susceptibility to and Severity of COVID-19. Eur. Respir. J. 2021, 58, 2004125. [Google Scholar] [CrossRef]
- Esposito, A.J.; Menon, A.A.; Ghosh, A.J.; Putman, R.K.; Fredenburgh, L.E.; El-Chemaly, S.Y.; Goldberg, H.J.; Baron, R.M.; Hunninghake, G.M.; Doyle, T.J. Increased Odds of Death for Patients with Interstitial Lung Disease and COVID-19: A Case–Control Study. Am. J. Respir. Crit. Care Med. 2020, 202, 1710–1713. [Google Scholar] [CrossRef]
- Ouyang, L.; Gong, J.; Yu, M. Pre-Existing Interstitial Lung Disease in Patients with Coronavirus Disease 2019: A Meta-Analysis. Int. Immunopharmacol. 2021, 100, 108145. [Google Scholar] [CrossRef]
- Drake, T.M.; Docherty, A.B.; Harrison, E.M.; Quint, J.K.; Adamali, H.; Agnew, S.; Babu, S.; Barber, C.M.; Barratt, S.; Bendstrup, E.; et al. Outcome of Hospitalization for COVID-19 in Patients with Interstitial Lung Disease. An International Multicenter Study. Am. J. Respir. Crit. Care Med. 2020, 202, 1656–1665. [Google Scholar] [CrossRef]
- Lee, S.C.; Son, K.J.; Han, C.H.; Jung, J.Y.; Park, S.C. Impact of Comorbid Asthma on Severity of Coronavirus Disease (COVID-19). Sci. Rep. 2020, 10, 21805. [Google Scholar] [CrossRef] [PubMed]
- Wark, P.A.B.; Pathinayake, P.S.; Kaiko, G.; Nichol, K.; Ali, A.; Chen, L.; Sutanto, E.N.; Garratt, L.W.; Sohal, S.S.; Lu, W.; et al. ACE2 Expression Is Elevated in Airway Epithelial Cells from Older and Male Healthy Individuals but Reduced in Asthma. Respirol. Carlton Vic 2021, 26, 442–451. [Google Scholar] [CrossRef] [PubMed]
- Jackson, D.J.; Busse, W.W.; Bacharier, L.B.; Kattan, M.; O’Connor, G.T.; Wood, R.A.; Visness, C.M.; Durham, S.R.; Larson, D.; Esnault, S.; et al. Association of Respiratory Allergy, Asthma, and Expression of the SARS-CoV-2 Receptor ACE2. J. Allergy Clin. Immunol. 2020, 146, 203–206. [Google Scholar] [CrossRef] [PubMed]
- Skevaki, C.; Karsonova, A.; Karaulov, A.; Fomina, D.; Xie, M.; Chinthrajah, S.; Nadeau, K.C.; Renz, H. SARS-CoV-2 Infection and COVID-19 in Asthmatics: A Complex Relationship. Nat. Rev. Immunol. 2021, 21, 202–203. [Google Scholar] [CrossRef]
- Ho, K.S.; Howell, D.; Rogers, L.; Narasimhan, B.; Verma, H.; Steiger, D. The Relationship between Asthma, Eosinophilia, and Outcomes in Coronavirus Disease 2019 Infection. Ann. Allergy Asthma Immunol. Off. Publ. Am. Coll. Allergy Asthma Immunol. 2021, 127, 42–48. [Google Scholar] [CrossRef]
- Peltola, V.; Jartti, T.; Putto-Laurila, A.; Mertsola, J.; Vainionpää, R.; Waris, M.; Hyypiä, T.; Ruuskanen, O. Rhinovirus Infections in Children: A Retrospective and Prospective Hospital-Based Study. J. Med. Virol. 2009, 81, 1831–1838. [Google Scholar] [CrossRef]
- Nicholson, K.G.; Kent, J.; Ireland, D.C. Respiratory Viruses and Exacerbations of Asthma in Adults. Br. Med. J. 1993, 307, 982–986. [Google Scholar] [CrossRef]
- Johnston, S.L.; Pattemore, P.K.; Sanderson, G.; Smith, S.; Lampe, F.; Josephs, L.; Symington, P.; Toole, S.O.; Myint, S.H.; Tyrrell, D.A.J.; et al. Community Study of Role of Viral Infections in Exacerbations of Asthma in 9-11 Year Old Children. BMJ 1995, 310, 1225–1229. [Google Scholar] [CrossRef]
- Contoli, M.; Message, S.D.; Laza-Stanca, V.; Edwards, M.R.; Wark, P.A.B.; Bartlett, N.W.; Kebadze, T.; Mallia, P.; Stanciu, L.A.; Parker, H.L.; et al. Role of Deficient Type III Interferon-λ Production in Asthma Exacerbations. Nat. Med. 2006, 12, 1023–1026. [Google Scholar] [CrossRef]
- Gill, M.A.; Bajwa, G.; George, T.A.; Dong, C.C.; Dougherty, I.I.; Jiang, N.; Gan, V.N.; Gruchalla, R.S. Counterregulation between the FcεRI Pathway and Antiviral Responses in Human Plasmacytoid Dendritic Cells. J. Immunol. 2010, 184, 5999–6006. [Google Scholar] [CrossRef]
- Gonzales-van Horn, S.R.; David Farrar, J. Interferon at the Crossroads of Allergy and Viral Infections. J. Leukoc. Biol. 2015, 98, 185–194. [Google Scholar] [CrossRef] [PubMed]
- Coleman, B.L.; Fadel, S.A.; Fitzpatrick, T.; Thomas, S.-M. Risk Factors for Serious Outcomes Associated with Influenza Illness in High- versus Low- and Middle-Income Countries: Systematic Literature Review and Meta-Analysis. Influenza Other Respir. Viruses 2018, 12, 22–29. [Google Scholar] [CrossRef]
- Yang, J.M.; Koh, H.Y.; Moon, S.Y.; Yoo, I.K.; Ha, E.K.; You, S.; Kim, S.Y.; Yon, D.K.; Lee, S.W. Allergic Disorders and Susceptibility to and Severity of COVID-19: A Nationwide Cohort Study. J. Allergy Clin. Immunol. 2020, 146, 790–798. [Google Scholar] [CrossRef] [PubMed]
- Zhu, Z.; Hasegawa, K.; Ma, B.; Fujiogi, M.; Camargo, C.A.; Liang, L. Association of Asthma and Its Genetic Predisposition with the Risk of Severe COVID-19. J. Allergy Clin. Immunol. 2020, 146, 327–329.e4. [Google Scholar] [CrossRef] [PubMed]
- World Obesity Atlas 2022. Available online: https://www.worldobesity.org/resources/resource-library/world-obesity-atlas-2022 (accessed on 22 October 2022).
- Petrie, J.R.; Guzik, T.J.; Touyz, R.M. Diabetes, Hypertension, and Cardiovascular Disease: Clinical Insights and Vascular Mechanisms. Can. J. Cardiol. 2018, 34, 575–584. [Google Scholar] [CrossRef] [PubMed]
- Madjid, M.; Safavi-Naeini, P.; Solomon, S.D.; Vardeny, O. Potential Effects of Coronaviruses on the Cardiovascular System: A Review. JAMA Cardiol. 2020, 5, 831–840. [Google Scholar] [CrossRef]
- Badawi, A.; Ryoo, S.G. Prevalence of Comorbidities in the Middle East Respiratory Syndrome Coronavirus (MERS-CoV): A Systematic Review and Meta-Analysis. Int. J. Infect. Dis. 2016, 49, 129–133. [Google Scholar] [CrossRef]
- Yang, J.K.; Feng, Y.; Yuan, M.Y.; Yuan, S.Y.; Fu, H.J.; Wu, B.Y.; Sun, G.Z.; Yang, G.R.; Zhang, X.L.; Wang, L.; et al. Plasma Glucose Levels and Diabetes Are Independent Predictors for Mortality and Morbidity in Patients with SARS. Diabet. Med. J. Br. Diabet. Assoc. 2006, 23, 623–628. [Google Scholar] [CrossRef]
- Kesavadev, J.; Misra, A.; Saboo, B.; Aravind, S.R.; Hussain, A.; Czupryniak, L.; Raz, I. Blood Glucose Levels Should Be Considered as a New Vital Sign Indicative of Prognosis during Hospitalization. Diabetes Metab. Syndr. 2021, 15, 221–227. [Google Scholar] [CrossRef]
- Zhang, B.; Liu, S.; Zhang, L.; Dong, Y.; Zhang, S. Admission Fasting Blood Glucose Predicts 30-Day Poor Outcome in Patients Hospitalized for COVID-19 Pneumonia. Diabetes Obes. Metab. 2020, 22, 1955–1957. [Google Scholar] [CrossRef]
- Wang, S.; Ma, P.; Zhang, S.; Song, S.; Wang, Z.; Ma, Y.; Xu, J.; Wu, F.; Duan, L.; Yin, Z.; et al. Fasting Blood Glucose at Admission Is an Independent Predictor for 28-Day Mortality in Patients with COVID-19 without Previous Diagnosis of Diabetes: A Multi-Centre Retrospective Study. Diabetologia 2020, 63, 2102–2111. [Google Scholar] [CrossRef] [PubMed]
- Böhm, M.; Frey, N.; Giannitsis, E.; Sliwa, K.; Zeiher, A.M. Coronavirus Disease 2019 (COVID-19) and Its Implications for Cardiovascular Care: Expert Document from the German Cardiac Society and the World Heart Federation. Clin. Res. Cardiol. 2020, 109, 1446–1459. [Google Scholar] [CrossRef] [PubMed]
- Kai, H.; Kai, M. Interactions of Coronaviruses with ACE2, Angiotensin II, and RAS Inhibitors—Lessons from Available Evidence and Insights into COVID-19. Hypertens. Res. 2020, 43, 648–654. [Google Scholar] [CrossRef] [PubMed]
- Meng, M.; Zhao, Q.; Kumar, R.; Bai, C.; Deng, Y.; Wan, B. Impact of Cardiovascular and Metabolic Diseases on the Severity of COVID-19: A Systematic Review and Meta-Analysis. Aging 2020, 12, 23409–23421. [Google Scholar] [CrossRef]
- Falagas, M.E.; Kompoti, M. Obesity and Infection. Lancet Infect. Dis. 2006, 6, 438–446. [Google Scholar] [CrossRef]
- Poulain, M.; Doucet, M.; Major, G.C.; Drapeau, V.; Sériès, F.; Boulet, L.-P.; Tremblay, A.; Maltais, F. The Effect of Obesity on Chronic Respiratory Diseases: Pathophysiology and Therapeutic Strategies. CMAJ 2006, 174, 1293–1299. [Google Scholar] [CrossRef]
- Parameswaran, K.; Todd, D.C.; Soth, M. Altered Respiratory Physiology in Obesity. Can. Respir. J. 2006, 13, 203–210. [Google Scholar] [CrossRef]
- He, H.; Wang, B.; Zhou, M.; Cao, L.; Qiu, W.; Mu, G.; Chen, A.; Yang, S.; Chen, W. Systemic Inflammation Mediates the Associations Between Abdominal Obesity Indices and Lung Function Decline in a Chinese General Population. Diabetes Metab. Syndr. Obes. Targets Ther. 2020, 13, 141–150. [Google Scholar] [CrossRef]
- Stefan, N. SARS-CoV-2 Fires up Inflammation in Adipose Tissue. Nat. Rev. Endocrinol. 2022, 19, 8–9. [Google Scholar] [CrossRef]
- Sibbel, S.; Sato, R.; Hunt, A.; Turenne, W.; Brunelli, S.M. The Clinical and Economic Burden of Pneumonia in Patients Enrolled in Medicare Receiving Dialysis: A Retrospective, Observational Cohort Study. BMC Nephrol. 2016, 17, 199. [Google Scholar] [CrossRef]
- Hernández-Galdamez, D.R.; González-Block, M.Á.; Romo-Dueñas, D.K.; Lima-Morales, R.; Hernández-Vicente, I.A.; Lumbreras-Guzmán, M.; Méndez-Hernández, P. Increased Risk of Hospitalization and Death in Patients with COVID-19 and Pre-Existing Noncommunicable Diseases and Modifiable Risk Factors in Mexico. Arch. Med. Res. 2020, 51, 683–689. [Google Scholar] [CrossRef]
- Sun, J.; Zhu, A.; Li, H.; Zheng, K.; Zhuang, Z.; Chen, Z.; Shi, Y.; Zhang, Z.; Chen, S.-B.; Liu, X.; et al. Isolation of Infectious SARS-CoV-2 from Urine of a COVID-19 Patient. Emerg. Microbes Infect. 2020, 9, 991–993. [Google Scholar] [CrossRef] [PubMed]
- Legrand, M.; Bell, S.; Forni, L.; Joannidis, M.; Koyner, J.L.; Liu, K.; Cantaluppi, V. Pathophysiology of COVID-19-Associated Acute Kidney Injury. Nat. Rev. Nephrol. 2021, 17, 751–764. [Google Scholar] [CrossRef]
- Sandoval, M.; Nguyen, D.T.; Vahidy, F.S.; Graviss, E.A. Risk Factors for Severity of COVID-19 in Hospital Patients Age 18–29 Years. PLoS ONE 2021, 16, e0255544. [Google Scholar] [CrossRef] [PubMed]
- Sunjaya, A.P.; Allida, S.M.; Di Tanna, G.L.; Jenkins, C. Asthma and Risk of Infection, Hospitalization, ICU Admission and Mortality from COVID-19: Systematic Review and Meta-Analysis. J. Asthma 2022, 59, 866–879. [Google Scholar] [CrossRef] [PubMed]
- Han, X.; Xu, J.; Hou, H.; Yang, H.; Wang, Y. Impact of Asthma on COVID-19 Mortality in the United States: Evidence Based on a Meta-Analysis. Int. Immunopharmacol. 2022, 102, 108390. [Google Scholar] [CrossRef] [PubMed]
- Hou, H.; Xu, J.; Li, Y.; Wang, Y.; Yang, H. The Association of Asthma with COVID-19 Mortality: An Updated Meta-Analysis Based on Adjusted Effect Estimates. J. Allergy Clin. Immunol. Pract. 2021, 9, 3944–3968. [Google Scholar] [CrossRef]
- Hussein, M.H.; Elshazli, R.M.; Attia, A.S.; Nguyen, T.P.; Aboueisha, M.; Munshi, R.; Toraih, E.A.; Fawzy, M.S.; Kandil, E. Asthma and COVID-19; Different Entities, Same Outcome: A Meta-Analysis of 107,983 Patients. J. Asthma 2022, 59, 851–858. [Google Scholar] [CrossRef]
- Liu, S.; Cao, Y.; Du, T.; Zhi, Y. Prevalence of Comorbid Asthma and Related Outcomes in COVID-19: A Systematic Review and Meta-Analysis. J. Allergy Clin. Immunol. Pract. 2021, 9, 693–701. [Google Scholar] [CrossRef]
- Shi, L.; Xu, J.; Xiao, W.; Wang, Y.; Jin, Y.; Chen, S.; Duan, G.; Yang, H.; Wang, Y. Asthma in Patients with Coronavirus Disease 2019: A Systematic Review and Meta-Analysis. Ann. Allergy. Asthma. Immunol. 2021, 126, 524–534. [Google Scholar] [CrossRef]
- Wu, T.; Yu, P.; Li, Y.; Wang, J.; Li, Z.; Qiu, J.; Cui, L.; Mou, Y.; Sun, Y. Asthma Does Not Influence the Severity of COVID-19: A Meta-Analysis. J. Asthma 2022, 59, 1188–1194. [Google Scholar] [CrossRef] [PubMed]
- Otunla, A.; Rees, K.; Dennison, P.; Hobbs, R.; Suklan, J.; Schofield, E.; Gunnell, J.; Mighiu, A.; Hartmann-Boyce, J. Risks of Infection, Hospital and ICU Admission, and Death from COVID-19 in People with Asthma: Systematic Review and Meta-Analyses. BMJ Evid.-Based Med. 2022, 27, 263–273. [Google Scholar] [CrossRef]
- Sunjaya, A.P.; Allida, S.M.; Tanna, G.L.D.; Jenkins, C.R. Asthma and COVID-19 Risk: A Systematic Review and Meta-Analysis. Eur. Respir. J. 2022, 59, 2101209. [Google Scholar] [CrossRef] [PubMed]
- Reyes, F.M.; Hache-Marliere, M.; Karamanis, D.; Berto, C.G.; Estrada, R.; Langston, M.; Ntaios, G.; Gulani, P.; Shah, C.D.; Palaiodimos, L. Assessment of the Association of COPD and Asthma with In-Hospital Mortality in Patients with COVID-19. A Systematic Review, Meta-Analysis, and Meta-Regression Analysis. J. Clin. Med. 2021, 10, 2087. [Google Scholar] [CrossRef]
- Bhaskaran, K.; Bacon, S.; Evans, S.J.; Bates, C.J.; Rentsch, C.T.; MacKenna, B.; Tomlinson, L.; Walker, A.J.; Schultze, A.; Morton, C.E.; et al. Factors Associated with Deaths Due to COVID-19 versus Other Causes: Population-Based Cohort Analysis of UK Primary Care Data and Linked National Death Registrations within the OpenSAFELY Platform. Lancet Reg. Health—Eur. 2021, 6, 100109. [Google Scholar] [CrossRef] [PubMed]
- Williamson, E.J.; Walker, A.J.; Bhaskaran, K.; Bacon, S.; Bates, C.; Morton, C.E.; Curtis, H.J.; Mehrkar, A.; Evans, D.; Inglesby, P.; et al. Factors Associated with COVID-19-Related Death Using OpenSAFELY. Nature 2020, 584, 430–436. [Google Scholar] [CrossRef] [PubMed]
- Szarpak, L.; Mierzejewska, M.; Jurek, J.; Kochanowska, A.; Gasecka, A.; Truszewski, Z.; Pruc, M.; Blek, N.; Rafique, Z.; Filipiak, K.J.; et al. Effect of Coronary Artery Disease on COVID-19—Prognosis and Risk Assessment: A Systematic Review and Meta-Analysis. Biology 2022, 11, 221. [Google Scholar] [CrossRef] [PubMed]
- Hessami, A.; Shamshirian, A.; Heydari, K.; Pourali, F.; Alizadeh-Navaei, R.; Moosazadeh, M.; Abrotan, S.; Shojaie, L.; Sedighi, S.; Shamshirian, D.; et al. Cardiovascular Diseases Burden in COVID-19: Systematic Review and Meta-Analysis. Am. J. Emerg. Med. 2021, 46, 382–391. [Google Scholar] [CrossRef] [PubMed]
- Zhang, L.; Hou, J.; Ma, F.-Z.; Li, J.; Xue, S.; Xu, Z.-G. The Common Risk Factors for Progression and Mortality in COVID-19 Patients: A Meta-Analysis. Arch. Virol. 2021, 166, 2071–2087. [Google Scholar] [CrossRef]
- Loffi, M.; Piccolo, R.; Regazzoni, V.; Tano, G.D.; Moschini, L.; Robba, D.; Quinzani, F.; Esposito, G.; Franzone, A.; Danzi, G.B. Coronary Artery Disease in Patients Hospitalised with Coronavirus Disease 2019 (COVID-19) Infection. Open Heart 2020, 7, e001428. [Google Scholar] [CrossRef]
- Atkins, J.L.; Masoli, J.A.H.; Delgado, J.; Pilling, L.C.; Kuo, C.-L.; Kuchel, G.A.; Melzer, D. Preexisting Comorbidities Predicting COVID-19 and Mortality in the UK Biobank Community Cohort. J. Gerontol. Ser. A 2020, 75, 2224–2230. [Google Scholar] [CrossRef]
- Singh, S.; Khan, A. Clinical Characteristics and Outcomes of Coronavirus Disease 2019 among Patients with Preexisting Liver Disease in the United States: A Multicenter Research Network Study. Gastroenterology 2020, 159, 768–771. [Google Scholar] [CrossRef] [PubMed]
- Hassan, L.; Peek, N.; Lovell, K.; Carvalho, A.F.; Solmi, M.; Stubbs, B.; Firth, J. Disparities in COVID-19 Infection, Hospitalisation and Death in People with Schizophrenia, Bipolar Disorder, and Major Depressive Disorder: A Cohort Study of the UK Biobank. Mol. Psychiatry 2022, 27, 1248–1255. [Google Scholar] [CrossRef] [PubMed]
- Galiero, R.; Pafundi, P.C.; Simeon, V.; Rinaldi, L.; Perrella, A.; Vetrano, E.; Caturano, A.; Alfano, M.; Beccia, D.; Nevola, R.; et al. Impact of Chronic Liver Disease upon Admission on COVID-19 in-Hospital Mortality: Findings from COVOCA Study. PLoS ONE 2020, 15, e0243700. [Google Scholar] [CrossRef] [PubMed]
- Gimeno-Miguel, A.; Bliek-Bueno, K.; Poblador-Plou, B.; Carmona-Pírez, J.; Poncel-Falcó, A.; González-Rubio, F.; Ioakeim-Skoufa, I.; Pico-Soler, V.; Aza-Pascual-Salcedo, M.; Prados-Torres, A.; et al. Chronic Diseases Associated with Increased Likelihood of Hospitalization and Mortality in 68,913 COVID-19 Confirmed Cases in Spain: A Population-Based Cohort Study. PLoS ONE 2021, 16, e0259822. [Google Scholar] [CrossRef]
- Hashemi, N.; Viveiros, K.; Redd, W.D.; Zhou, J.C.; McCarty, T.R.; Bazarbashi, A.N.; Hathorn, K.E.; Wong, D.; Njie, C.; Shen, L.; et al. Impact of Chronic Liver Disease on Outcomes of Hospitalized Patients with COVID-19: A Multicentre United States Experience. Liver Int. 2020, 40, 2515–2521. [Google Scholar] [CrossRef]
- Khan, M.M.A.; Khan, M.N.; Mustagir, M.G.; Rana, J.; Islam, M.S.; Kabir, M.I. Effects of Underlying Morbidities on the Occurrence of Deaths in COVID-19 Patients: A Systematic Review and Meta-Analysis. J. Glob. Health 2020, 10, 020503. [Google Scholar] [CrossRef]
- Yin, T.; Li, Y.; Ying, Y.; Luo, Z. Prevalence of Comorbidity in Chinese Patients with COVID-19: Systematic Review and Meta-Analysis of Risk Factors. BMC Infect. Dis. 2021, 21, 200. [Google Scholar] [CrossRef]
- Wang, X.; Fang, X.; Cai, Z.; Wu, X.; Gao, X.; Min, J.; Wang, F. Comorbid Chronic Diseases and Acute Organ Injuries Are Strongly Correlated with Disease Severity and Mortality among COVID-19 Patients: A Systemic Review and Meta-Analysis. Research 2020, 2020, 1–17. [Google Scholar] [CrossRef]
- Kulkarni, A.V.; Kumar, P.; Tevethia, H.V.; Premkumar, M.; Arab, J.P.; Candia, R.; Talukdar, R.; Sharma, M.; Qi, X.; Rao, P.N.; et al. Systematic Review with Meta-Analysis: Liver Manifestations and Outcomes in COVID-19. Aliment. Pharmacol. Ther. 2020, 52, 584–599. [Google Scholar] [CrossRef]
- Lee, Y.R.; Kang, M.K.; Song, J.E.; Kim, H.J.; Kweon, Y.O.; Tak, W.Y.; Jang, S.Y.; Park, J.G.; Lee, C.; Hwang, J.S.; et al. Clinical Outcomes of Coronavirus Disease 2019 in Patients with Pre-Existing Liver Diseases: A Multicenter Study in South Korea. Clin. Mol. Hepatol. 2020, 26, 562–576. [Google Scholar] [CrossRef]
- Treskova-Schwarzbach, M.; Haas, L.; Reda, S.; Pilic, A.; Borodova, A.; Karimi, K.; Koch, J.; Nygren, T.; Scholz, S.; Schönfeld, V.; et al. Pre-Existing Health Conditions and Severe COVID-19 Outcomes: An Umbrella Review Approach and Meta-Analysis of Global Evidence. BMC Med. 2021, 19, 212. [Google Scholar] [CrossRef]
- Frager, S.Z.; Szymanski, J.; Schwartz, J.M.; Massoumi, H.S.; Kinkhabwala, M.; Wolkoff, A.W. Hepatic Predictors of Mortality in Severe Acute Respiratory Syndrome Coronavirus 2: Role of Initial Aspartate Aminotransferase/Alanine Aminotransferase and Preexisting Cirrhosis. Hepatol. Commun. 2021, 5, 424–433. [Google Scholar] [CrossRef] [PubMed]
- Kim, D.; Adeniji, N.; Latt, N.; Kumar, S.; Bloom, P.P.; Aby, E.S.; Perumalswami, P.; Roytman, M.; Li, M.; Vogel, A.S.; et al. Predictors of Outcomes of COVID-19 in Patients with Chronic Liver Disease: US Multi-Center Study. Clin. Gastroenterol. Hepatol. 2021, 19, 1469–1479. [Google Scholar] [CrossRef] [PubMed]
- Jeon, D.; Son, M.; Choi, J. Impact of Liver Cirrhosis on the Clinical Outcomes of Patients with COVID-19: A Nationwide Cohort Study of Korea. Korean J. Intern. Med. 2021, 36, 1092–1101. [Google Scholar] [CrossRef] [PubMed]
- Mahamid, M.; Nseir, W.; Khoury, T.; Mahamid, B.; Nubania, A.; Sub-Laban, K.; Schifter, J.; Mari, A.; Sbeit, W.; Goldin, E. Nonalcoholic Fatty Liver Disease Is Associated with COVID-19 Severity Independently of Metabolic Syndrome: A Retrospective Case-Control Study. Eur. J. Gastroenterol. Hepatol. 2021, 33, 1578–1581. [Google Scholar] [CrossRef]
- Gao, F.; Zheng, K.I.; Wang, X.; Yan, H.; Sun, Q.; Pan, K.; Wang, T.; Chen, Y.; George, J.; Zheng, M. Metabolic Associated Fatty Liver Disease Increases Coronavirus Disease 2019 Disease Severity in Nondiabetic Patients. J. Gastroenterol. Hepatol. 2021, 36, 204–207. [Google Scholar] [CrossRef]
- Ji, D.; Qin, E.; Xu, J.; Zhang, D.; Cheng, G.; Wang, Y.; Lau, G. Non-Alcoholic Fatty Liver Diseases in Patients with COVID-19: A Retrospective Study. J. Hepatol. 2020, 73, 451–453. [Google Scholar] [CrossRef]
- Pan, L.; Huang, P.; Xie, X.; Xu, J.; Guo, D.; Jiang, Y. Metabolic Associated Fatty Liver Disease Increases the Severity of COVID-19: A Meta-Analysis. Dig. Liver Dis. 2021, 53, 153–157. [Google Scholar] [CrossRef]
- Zhou, Y.-J.; Zheng, K.I.; Wang, X.-B.; Sun, Q.-F.; Pan, K.-H.; Wang, T.-Y.; Ma, H.-L.; Chen, Y.-P.; George, J.; Zheng, M.-H. Metabolic-Associated Fatty Liver Disease Is Associated with Severity of COVID-19. Liver Int. 2020, 40, 2160–2163. [Google Scholar] [CrossRef]
- Tao, Z.; Li, Y.; Cheng, B.; Zhou, T.; Gao, Y. Risk of Severe COVID-19 Increased by Metabolic Dysfunction-Associated Fatty Liver Disease. J. Clin. Gastroenterol. 2021, 55, 830–835. [Google Scholar] [CrossRef]
- Hegyi, P.J.; Váncsa, S.; Ocskay, K.; Dembrovszky, F.; Kiss, S.; Farkas, N.; Erőss, B.; Szakács, Z.; Hegyi, P.; Pár, G. Metabolic Associated Fatty Liver Disease Is Associated with an Increased Risk of Severe COVID-19: A Systematic Review with Meta-Analysis. Front. Med. 2021, 8, 626425. [Google Scholar] [CrossRef]
- Singh, A.; Hussain, S.; Antony, B. Non-Alcoholic Fatty Liver Disease and Clinical Outcomes in Patients with COVID-19: A Comprehensive Systematic Review and Meta-Analysis. Diabetes Metab. Syndr. Clin. Res. Rev. 2021, 15, 813–822. [Google Scholar] [CrossRef] [PubMed]
- Zhou, Y.-J.; Zheng, K.I.; Wang, X.-B.; Yan, H.-D.; Sun, Q.-F.; Pan, K.-H.; Wang, T.-Y.; Ma, H.-L.; Chen, Y.-P.; George, J.; et al. Younger Patients with MAFLD Are at Increased Risk of Severe COVID-19 Illness: A Multicenter Preliminary Analysis. J. Hepatol. 2020, 73, 719–721. [Google Scholar] [CrossRef] [PubMed]
- Marjot, T.; Moon, A.M.; Cook, J.A.; Abd-Elsalam, S.; Aloman, C.; Armstrong, M.J.; Pose, E.; Brenner, E.J.; Cargill, T.; Catana, M.-A.; et al. Outcomes Following SARS-CoV-2 Infection in Patients with Chronic Liver Disease: An International Registry Study. J. Hepatol. 2021, 74, 567–577. [Google Scholar] [CrossRef] [PubMed]
- Eder, L.; Croxford, R.; Drucker, A.M.; Mendel, A.; Kuriya, B.; Touma, Z.; Johnson, S.R.; Cook, R.; Bernatsky, S.; Haroon, N.; et al. COVID-19 Hospitalizations, Intensive Care Unit Stays, Ventilation, and Death among Patients with Immune-Mediated Inflammatory Diseases Compared to Controls. J. Rheumatol. 2022, 49, 523–530. [Google Scholar] [CrossRef]
- Yang, H.; Xu, J.; Liang, X.; Shi, L.; Wang, Y. Autoimmune Diseases Are Independently Associated with COVID-19 Severity: Evidence Based on Adjusted Effect Estimates. J. Infect. 2021, 82, e23–e26. [Google Scholar] [CrossRef] [PubMed]
- Faye, A.S.; Lee, K.E.; Laszkowska, M.; Kim, J.; Blackett, J.W.; McKenney, A.S.; Krigel, A.; Giles, J.T.; Wang, R.; Bernstein, E.J.; et al. Risk of Adverse Outcomes in Hospitalized Patients with Autoimmune Disease and COVID-19: A Matched Cohort Study from New York City. J. Rheumatol. 2021, 48, 454–462. [Google Scholar] [CrossRef] [PubMed]
- Liu, M.; Gao, Y.; Zhang, Y.; Shi, S.; Chen, Y.; Tian, J. The Association between Severe or Dead COVID-19 and Autoimmune Diseases: A Systematic Review and Meta-Analysis. J. Infect. 2020, 81, e93–e95. [Google Scholar] [CrossRef]
- Monreal, E.; Sainz de la Maza, S.; Fernández-Velasco, J.I.; Natera-Villalba, E.; Rita, C.G.; Rodríguez-Jorge, F.; Beltrán-Corbellini, Á.; Iturrieta-Zuazo, I.; Rodríguez de Santiago, E.; Espiño, M.; et al. The Impact of Immunosuppression and Autoimmune Disease on Severe Outcomes in Patients Hospitalized with COVID-19. J. Clin. Immunol. 2021, 41, 315–323. [Google Scholar] [CrossRef] [PubMed]
- Kjeldsen, J.; Nielsen, J.; Ellingsen, T.; Knudsen, T.; Nielsen, R.; Larsen, M.; Lund, K.; Nørgård, B. Outcome of COVID-19 in Hospitalized Patients with Chronic Inflammatory Diseases. A Population Based National Register Study in Denmark. J. Autoimmun. 2021, 120, 102632. [Google Scholar] [CrossRef]
- Moreno-Torres, V.; de Mendoza, C.; Mellor-Pita, S.; Martínez-Urbistondo, M.; Durán-del Campo, P.; Tutor-Ureta, P.; Vázquez-Comendador, J.-M.; Calderón-Parra, J.; Múñez-Rubio, E.; Ramos-Martínez, A.; et al. Systemic Autoimmune Diseases in Patients Hospitalized with COVID-19 in Spain: A Nation-Wide Registry Study. Viruses 2022, 14, 1631. [Google Scholar] [CrossRef]
- Jung, Y.; Kwon, M.; Choi, H.G. Association between Previous Rheumatoid Arthritis and COVID-19 and Its Severity: A Nationwide Cohort Study in South Korea. BMJ Open 2021, 11, e054753. [Google Scholar] [CrossRef] [PubMed]
- Raiker, R.; DeYoung, C.; Pakhchanian, H.; Ahmed, S.; Kavadichanda, C.; Gupta, L.; Kardeş, S. Outcomes of COVID-19 in Patients with Rheumatoid Arthritis: A Multicenter Research Network Study in the United States. Semin. Arthritis Rheum. 2021, 51, 1057–1066. [Google Scholar] [CrossRef] [PubMed]
- Nørgård, B.M.; Nielsen, J.; Knudsen, T.; Nielsen, R.G.; Larsen, M.D.; Jølving, L.R.; Kjeldsen, J. Hospitalization for COVID-19 in Patients Treated with Selected Immunosuppressant and Immunomodulating Agents, Compared to the General Population: A Danish Cohort Study. Br. J. Clin. Pharmacol. 2021, 87, 2111–2120. [Google Scholar] [CrossRef] [PubMed]
- Brodin, R.; Desirée van der Werff, S.; Hedberg, P.; Färnert, A.; Nauclér, P.; Bergman, P.; Requena-Méndez, A. The Association between Pre-Exposure to Glucocorticoids and Other Immunosuppressant Drugs with Severe COVID-19 Outcomes. Clin. Microbiol. Infect. 2022, 28, 1477–1485. [Google Scholar] [CrossRef]
- Rutherford, M.A.; Scott, J.; Karabayas, M.; Antonelou, M.; Gopaluni, S.; Gray, D.; Barrett, J.; Brix, S.R.; Dhaun, N.; McAdoo, S.P.; et al. Risk Factors for Severe Outcomes in Patients with Systemic Vasculitis and COVID-19: A Binational, Registry-Based Cohort Study. Arthritis Rheumatol. 2021, 73, 1713–1719. [Google Scholar] [CrossRef]
- Ahlström, B.; Frithiof, R.; Hultström, M.; Larsson, I.-M.; Strandberg, G.; Lipcsey, M. The Swedish COVID-19 Intensive Care Cohort: Risk Factors of ICU Admission and ICU Mortality. Acta Anaesthesiol. Scand. 2021, 65, 525–533. [Google Scholar] [CrossRef] [PubMed]
- Suárez-García, I.; Perales-Fraile, I.; González-García, A.; Muñoz-Blanco, A.; Manzano, L.; Fabregate, M.; Díez-Manglano, J.; Aizpuru, E.F.; Fernández, F.A.; García, A.G.; et al. In-Hospital Mortality among Immunosuppressed Patients with COVID-19: Analysis from a National Cohort in Spain. PLoS ONE 2021, 16, e0255524. [Google Scholar] [CrossRef] [PubMed]
- Vaid, N.; Ardissino, M.; Reed, T.a.N.; Goodall, J.; Utting, P.; Miscampbell, M.; Condurache, D.G.; Cohen, D.L. Clinical Characteristics and Outcomes of Immunosuppressed Patients Hospitalized with COVID-19: Experience from London. J. Intern. Med. 2021, 289, 385–394. [Google Scholar] [CrossRef]
- Calderón-Parra, J.; Cuervas-Mons, V.; Moreno-Torres, V.; Rubio-Rivas, M.; Blas, P.A.; Pinilla-Llorente, B.; Helguera-Amezua, C.; Jiménez-García, N.; Pesqueira-Fontan, P.-M.; Méndez-Bailón, M.; et al. Influence of Chronic Use of Corticosteroids and Calcineurin Inhibitors on COVID-19 Clinical Outcomes: Analysis of a Nationwide Registry. Int. J. Infect. Dis. 2022, 116, 51–58. [Google Scholar] [CrossRef]
- Pablos, J.L.; Galindo, M.; Carmona, L.; Lledó, A.; Retuerto, M.; Blanco, R.; Gonzalez-Gay, M.A.; Martinez-Lopez, D.; Castrejón, I.; Alvaro-Gracia, J.M.; et al. Clinical Outcomes of Hospitalised Patients with COVID-19 and Chronic Inflammatory and Autoimmune Rheumatic Diseases: A Multicentric Matched Cohort Study. Ann. Rheum. Dis. 2020, 79, 1544–1549. [Google Scholar] [CrossRef]
- El Fakih, R.; Haroon, A.; Alfraih, F.; Al-Khabori, M.K.; Alzahrani, M.; Alhuraiji, A.; Hamadah, A.; AlJohani, N.I.; Alahmari, B.; Essa, M.F.; et al. Clinical Course and Outcomes of COVID-19 in Hematopoietic Cell Transplant Patients, a Regional Report from the Middle East. Bone Marrow Transplant. 2021, 56, 2144–2151. [Google Scholar] [CrossRef]
- Sun, J.; Patel, R.C.; Zheng, Q.; Madhira, V.; Olex, A.L.; Islam, J.Y.; French, E.; Chiang, T.P.-Y.; Akselrod, H.; Moffitt, R.; et al. COVID-19 Disease Severity among People with HIV Infection or Solid Organ Transplant in the United States: A Nationally-Representative, Multicenter, Observational Cohort Study. medRxiv 2021. [Google Scholar] [CrossRef]
- Gatti, M.; Rinaldi, M.; Bussini, L.; Bonazzetti, C.; Pascale, R.; Pasquini, Z.; Faní, F.; Pinho Guedes, M.N.; Azzini, A.M.; Carrara, E.; et al. Clinical Outcome in Solid Organ Transplant Recipients Affected by COVID-19 Compared to General Population: A Systematic Review and Meta-Analysis. Clin. Microbiol. Infect. 2022, 28, 1057–1065. [Google Scholar] [CrossRef] [PubMed]
- Oltean, M.; Søfteland, J.M.; Bagge, J.; Ekelund, J.; Felldin, M.; Schult, A.; Magnusson, J.; Friman, V.; Karason, K. COVID-19 in Kidney Transplant Recipients: A Systematic Review of the Case Series Available Three Months into the Pandemic. Infect. Dis. 2020, 52, 830–837. [Google Scholar] [CrossRef] [PubMed]
- Genuardi, M.V.; Moss, N.; Najjar, S.S.; Houston, B.A.; Shore, S.; Vorovich, E.; Atluri, P.; Molina, M.; Chambers, S.; Sharkoski, T.; et al. Coronavirus Disease 2019 in Heart Transplant Recipients: Risk Factors, Immunosuppression, and Outcomes. J. Heart Lung Transplant. 2021, 40, 926–935. [Google Scholar] [CrossRef] [PubMed]
- Chaudhry, Z.S.; Williams, J.D.; Vahia, A.; Fadel, R.; Parraga Acosta, T.; Prashar, R.; Shrivastava, P.; Khoury, N.; Pinto Corrales, J.; Williams, C.; et al. Clinical Characteristics and Outcomes of COVID-19 in Solid Organ Transplant Recipients: A Cohort Study. Am. J. Transplant. 2020, 20, 3051–3060. [Google Scholar] [CrossRef] [PubMed]
- Chavarot, N.; Gueguen, J.; Bonnet, G.; Jdidou, M.; Trimaille, A.; Burger, C.; Amrouche, L.; Weizman, O.; Pommier, T.; Aubert, O.; et al. COVID-19 Severity in Kidney Transplant Recipients Is Similar to Nontransplant Patients with Similar Comorbidities. Am. J. Transplant. 2021, 21, 1285–1294. [Google Scholar] [CrossRef]
- Kamp, J.C.; Hinrichs, J.B.; Fuge, J.; Ewen, R.; Gottlieb, J. COVID-19 in Lung Transplant Recipients—Risk Prediction and Outcomes. PLoS ONE 2021, 16, e0257807. [Google Scholar] [CrossRef] [PubMed]
- Kulkarni, A.V.; Tevethia, H.V.; Premkumar, M.; Arab, J.P.; Candia, R.; Kumar, K.; Kumar, P.; Sharma, M.; Rao, P.N.; Reddy, D.N. Impact of COVID-19 on Liver Transplant Recipients–A Systematic Review and Meta-Analysis. EClinicalMedicine 2021, 38, 101025. [Google Scholar] [CrossRef]
- Sharma, P.; Chen, V.; Fung, C.M.; Troost, J.P.; Patel, V.N.; Combs, M.; Norman, S.; Garg, P.; Colvin, M.; Aaronson, K.; et al. COVID-19 Outcomes among Solid Organ Transplant Recipients: A Case-Control Study. Transplantation 2021, 105, 128–137. [Google Scholar] [CrossRef]
- Bojesen, A.B.; Lund, A.; Mortensen, F.V.; Kirkegård, J. Splenectomy and Risk of COVID-19 Infection, Hospitalisation, and Death. Infect. Dis. 2021, 53, 678–683. [Google Scholar] [CrossRef] [PubMed]
- Liu, L.; Ni, S.-Y.; Yan, W.; Lu, Q.-D.; Zhao, Y.-M.; Xu, Y.-Y.; Mei, H.; Shi, L.; Yuan, K.; Han, Y.; et al. Mental and Neurological Disorders and Risk of COVID-19 Susceptibility, Illness Severity and Mortality: A Systematic Review, Meta-Analysis and Call for Action. EClinicalMedicine 2021, 40, 101111. [Google Scholar] [CrossRef] [PubMed]
- Masoli, J.; Kuo, C.L.; Atkins, J.; Pilling, L.; Delgado, J.; Kuchel, G.; Melzer, D. 38 Dementia, Apoe and COVID-19 Severity. Age Ageing 2021, 50, i7–i11. [Google Scholar] [CrossRef]
- Hariyanto, T.I.; Putri, C.; Arisa, J.; Situmeang, R.F.V.; Kurniawan, A. Dementia and Outcomes from Coronavirus Disease 2019 (COVID-19) Pneumonia: A Systematic Review and Meta-Analysis. Arch. Gerontol. Geriatr. 2021, 93, 104299. [Google Scholar] [CrossRef] [PubMed]
- Yu, Y.; Travaglio, M.; Popovic, R.; Leal, N.S.; Martins, L.M. Alzheimer’s and Parkinson’s Diseases Predict Different COVID-19 Outcomes: A UK Biobank Study. Geriatrics 2021, 6, 10. [Google Scholar] [CrossRef] [PubMed]
- Zhou, J.; Liu, C.; Sun, Y.; Huang, W.; Ye, K. Cognitive Disorders Associated with Hospitalization of COVID-19: Results from an Observational Cohort Study. Brain. Behav. Immun. 2021, 91, 383–392. [Google Scholar] [CrossRef]
- Wang, Y.; Yang, Y.; Ren, L.; Shao, Y.; Tao, W.; Dai, X. Preexisting Mental Disorders Increase the Risk of COVID-19 Infection and Associated Mortality. Front. Public Health 2021, 9, 684112. [Google Scholar] [CrossRef]
- Zuin, M.; Guasti, P.; Roncon, L.; Cervellati, C.; Zuliani, G. Dementia and the Risk of Death in Elderly Patients with COVID-19 Infection: Systematic Review and Meta-Analysis. Int. J. Geriatr. Psychiatry 2021, 36, 697–703. [Google Scholar] [CrossRef]
- Bellou, V.; Tzoulaki, I.; Smeden, M.V.; Moons, K.G.M.; Evangelou, E.; Belbasis, L. Prognostic Factors for Adverse Outcomes in Patients with COVID-19: A Field-Wide Systematic Review and Meta-Analysis. Eur. Respir. J. 2022, 59, 2002964. [Google Scholar] [CrossRef]
- Liu, N.; Sun, J.; Wang, X.; Zhao, M.; Huang, Q.; Li, H. The Impact of Dementia on the Clinical Outcome of COVID-19: A Systematic Review and Meta-Analysis. J. Alzheimers Dis. 2020, 78, 1775–1782. [Google Scholar] [CrossRef]
- Filardo, T.D.; Khan, M.R.; Krawczyk, N.; Galitzer, H.; Karmen-Tuohy, S.; Coffee, M.; Schaye, V.E.; Eckhardt, B.J.; Cohen, G.M. Comorbidity and Clinical Factors Associated with COVID-19 Critical Illness and Mortality at a Large Public Hospital in New York City in the Early Phase of the Pandemic (March–April 2020). PLoS ONE 2020, 15, e0242760. [Google Scholar] [CrossRef] [PubMed]
- Samuels, S.; Niu, J.; Sareli, C.; Eckardt, P. The Epidemiology and Predictors of Outcomes among Confirmed COVID-19 Cases in a Large Community Healthcare System in South Florida. J. Community Health 2021, 46, 822–831. [Google Scholar] [CrossRef] [PubMed]
- Kim, J.H.; Chang, I.B.; Kim, Y.H.; Min, C.Y.; Yoo, D.M.; Choi, H.G. The Association of Pre-Existing Diagnoses of Alzheimer’s Disease and Parkinson’s Disease and Coronavirus Disease 2019 Infection, Severity and Mortality: Results from the Korean National Health Insurance Database. Front. Aging Neurosci. 2022, 14, 821235. [Google Scholar] [CrossRef] [PubMed]
- Chung, S.J.; Chang, Y.; Jeon, J.; Shin, J.I.; Song, T.-J.; Kim, J. Association of Alzheimer’s Disease with COVID-19 Susceptibility and Severe Complications: A Nationwide Cohort Study. J. Alzheimers Dis. 2022, 87, 701–710. [Google Scholar] [CrossRef]
- Wang, Y.; Li, M.; Kazis, L.E.; Xia, W. Clinical Outcomes of COVID-19 Infection among Patients with Alzheimer’s Disease or Mild Cognitive Impairment. Alzheimers Dement. 2022, 18, 911–923. [Google Scholar] [CrossRef]
- Honardoost, M.; Janani, L.; Aghili, R.; Emami, Z.; Khamseh, M.E. The Association between Presence of Comorbidities and COVID-19 Severity: A Systematic Review and Meta-Analysis. Cerebrovasc. Dis. 2021, 50, 132–140. [Google Scholar] [CrossRef]
- Yu, J.-N.; Wu, B.-B.; Yang, J.; Lei, X.-L.; Shen, W.-Q. Cardio-Cerebrovascular Disease Is Associated with Severity and Mortality of COVID-19: A Systematic Review and Meta-Analysis. Biol. Res. Nurs. 2021, 23, 258–269. [Google Scholar] [CrossRef]
- Siepmann, T.; Sedghi, A.; Barlinn, J.; de With, K.; Mirow, L.; Wolz, M.; Gruenewald, T.; Helbig, S.; Schroettner, P.; Winzer, S.; et al. Association of History of Cerebrovascular Disease with Severity of COVID-19. J. Neurol. 2021, 268, 773–784. [Google Scholar] [CrossRef]
- Pranata, R.; Huang, I.; Lim, M.A.; Wahjoepramono, E.J.; July, J. Impact of Cerebrovascular and Cardiovascular Diseases on Mortality and Severity of COVID-19–Systematic Review, Meta-Analysis, and Meta-Regression. J. Stroke Cerebrovasc. Dis. 2020, 29, 104949. [Google Scholar] [CrossRef]
- Ramphul, K.; Lohana, P.; Ramphul, Y.; Park, Y.; Mejias, S.; Dhillon, B.K.; Sombans, S.; Verma, R. Hypertension, Diabetes Mellitus, and Cerebrovascular Disease Predispose to a More Severe Outcome of COVID-19. Arch. Med. Sci. Atheroscler. Dis. 2021, 6, e30–e39. [Google Scholar] [CrossRef]
- Patel, U.; Malik, P.; Shah, D.; Patel, A.; Dhamoon, M.; Jani, V. Pre-Existing Cerebrovascular Disease and Poor Outcomes of COVID-19 Hospitalized Patients: A Meta-Analysis. J. Neurol. 2021, 268, 240–247. [Google Scholar] [CrossRef]
- Zhang, L.; Sun, W.; Wang, Y.; Wang, X.; Liu, Y.; Zhao, S.; Long, D.; Chen, L.; Yu, L. Clinical Course and Mortality of Stroke Patients with Coronavirus Disease 2019 in Wuhan, China. Stroke 2020, 51, 2674–2682. [Google Scholar] [CrossRef]
- Kummer, B.R.; Klang, E.; Stein, L.K.; Dhamoon, M.S.; Jetté, N. History of Stroke Is Independently Associated with In-Hospital Death in Patients with COVID-19. Stroke 2020, 51, 3112–3114. [Google Scholar] [CrossRef]
- Eskandar, E.N.; Altschul, D.J.; de la Garza Ramos, R.; Cezayirli, P.; Unda, S.R.; Benton, J.; Dardick, J.; Toma, A.; Patel, N.; Malaviya, A.; et al. Neurologic Syndromes Predict Higher In-Hospital Mortality in COVID-19. Neurology 2021, 96, e1527–e1538. [Google Scholar] [CrossRef]
- Yoo, J.; Kim, J.H.; Jeon, J.; Kim, J.; Song, T.-J. Risk of COVID-19 Infection and of Severe Complications among People with Epilepsy: A Nationwide Cohort Study. Neurology 2022, 98, e1886–e1892. [Google Scholar] [CrossRef]
- Hariyanto, T.I.; Kurniawan, A. Obstructive Sleep Apnea (OSA) and Outcomes from Coronavirus Disease 2019 (COVID-19) Pneumonia: A Systematic Review and Meta-Analysis. Sleep Med. 2021, 82, 47–53. [Google Scholar] [CrossRef]
- Rögnvaldsson, K.G.; Eyþórsson, E.S.; Emilsson, Ö.I.; Eysteinsdóttir, B.; Pálsson, R.; Gottfreðsson, M.; Guðmundsson, G.; Steingrímsson, V. Obstructive Sleep Apnea Is an Independent Risk Factor for Severe COVID-19: A Population-Based Study. Sleep 2022, 45, zsab272. [Google Scholar] [CrossRef]
- Chung, F.; Waseem, R.; Pham, C.; Penzel, T.; Han, F.; Bjorvatn, B.; Morin, C.M.; Holzinger, B.; Espie, C.A.; Benedict, C.; et al. The Association between High Risk of Sleep Apnea, Comorbidities, and Risk of COVID-19: A Population-Based International Harmonized Study. Sleep Breath. 2021, 25, 849–860. [Google Scholar] [CrossRef]
- Hu, M.; Han, X.; Ren, J.; Wang, Y.; Yang, H. Significant Association of Obstructive Sleep Apnoea with Increased Risk for Fatal COVID-19: A Quantitative Meta-Analysis Based on Adjusted Effect Estimates. Sleep Med. Rev. 2022, 63, 101624. [Google Scholar] [CrossRef]
- Goldstein, C.A.; Rizvydeen, M.; Conroy, D.A.; O’Brien Louise, M.; Gupta, G.; Somers, E.C.; Sharma, P.; Golob, J.L.; Troost, J.P.; Burgess, H.J. The Prevalence and Impact of Pre-Existing Sleep Disorder Diagnoses and Objective Sleep Parameters in Patients Hospitalized for COVID-19. J. Clin. Sleep Med. 2021, 17, 1039–1050. [Google Scholar] [CrossRef]
- Vignatelli, L.; Zenesini, C.; Belotti, L.M.B.; Baldin, E.; Bonavina, G.; Calandra-Buonaura, G.; Cortelli, P.; Descovich, C.; Fabbri, G.; Giannini, G.; et al. Risk of Hospitalization and Death for COVID-19 in People with Parkinson’s Disease or Parkinsonism. Mov. Disord. 2021, 36, 1–10. [Google Scholar] [CrossRef]
- El-Qushayri, A.E.; Ghozy, S.; Reda, A.; Kamel, A.M.A.; Abbas, A.S.; Dmytriw, A.A. The Impact of Parkinson’s Disease on Manifestations and Outcomes of COVID-19 Patients: A Systematic Review and Meta-Analysis. Rev. Med. Virol. 2022, 32, e2278. [Google Scholar] [CrossRef] [PubMed]
- Ceban, F.; Nogo, D.; Carvalho, I.P.; Lee, Y.; Nasri, F.; Xiong, J.; Lui, L.M.W.; Subramaniapillai, M.; Gill, H.; Liu, R.N.; et al. Association Between Mood Disorders and Risk of COVID-19 Infection, Hospitalization, and Death: A Systematic Review and Meta-Analysis. JAMA Psychiatry 2021, 78, 1079–1091. [Google Scholar] [CrossRef] [PubMed]
- Vai, B.; Mazza, M.G.; Delli Colli, C.; Foiselle, M.; Allen, B.; Benedetti, F.; Borsini, A.; Casanova Dias, M.; Tamouza, R.; Leboyer, M.; et al. Mental Disorders and Risk of COVID-19-Related Mortality, Hospitalisation, and Intensive Care Unit Admission: A Systematic Review and Meta-Analysis. Lancet Psychiatry 2021, 8, 797–812. [Google Scholar] [CrossRef] [PubMed]
- Castro, V.M.; Gunning, F.M.; McCoy, T.H.; Perlis, R.H. Mood Disorders and Outcomes of COVID-19 Hospitalizations. Am. J. Psychiatry 2021, 178, 541–547. [Google Scholar] [CrossRef]
- Egede, C.; Dawson, A.Z.; Walker, R.J.; Garacci, E.; Campbell, J.A.; Egede, L.E. Relationship between Mental Health Diagnoses and COVID-19 Test Positivity, Hospitalization, and Mortality in Southeast Wisconsin. Psychol. Med. 2021, 1–9. [Google Scholar] [CrossRef]
- Barcella, C.A.; Polcwiartek, C.; Mohr, G.H.; Hodges, G.; Søndergaard, K.; Niels Bang, C.; Andersen, M.P.; Fosbøl, E.; Køber, L.; Schou, M.; et al. Severe Mental Illness Is Associated with Increased Mortality and Severe Course of COVID-19. Acta Psychiatr. Scand. 2021, 144, 82–91. [Google Scholar] [CrossRef]
- Fond, G.; Pauly, V.; Leone, M.; Orleans, V.; Garosi, A.; Lancon, C.; Auquier, P.; Baumstarck, K.; Llorca, P.-M.; Boyer, L. Mortality among Inpatients with Bipolar Disorders and COVID-19: A Propensity Score Matching Analysis in a National French Cohort Study. Psychol. Med. 2021, 1–10. [Google Scholar] [CrossRef]
- Yang, H.; Chen, W.; Hu, Y.; Chen, Y.; Zeng, Y.; Sun, Y.; Ying, Z.; He, J.; Qu, Y.; Lu, D.; et al. Pre-Pandemic Psychiatric Disorders and Risk of COVID-19: A UK Biobank Cohort Analysis. Lancet Healthy Longev. 2020, 1, e69–e79. [Google Scholar] [CrossRef]
- Wang, S.; Quan, L.; Ding, M.; Kang, J.H.; Koenen, K.C.; Kubzansky, L.D.; Branch-Elliman, W.; Chavarro, J.E.; Roberts, A.L. Depression, Worry, and Loneliness Are Associated with Subsequent Risk of Hospitalization for COVID-19: A Prospective Study. Psychol. Med. 2022, 1–10. [Google Scholar] [CrossRef]
- Merzon, E.; Weiss, M.D.; Cortese, S.; Rotem, A.; Schneider, T.; Craig, S.G.; Vinker, S.; Golan Cohen, A.; Green, I.; Ashkenazi, S.; et al. The Association between ADHD and the Severity of COVID-19 Infection. J. Atten. Disord. 2022, 26, 491–501. [Google Scholar] [CrossRef]
- Velásquez García, H.A.; Wilton, J.; Smolina, K.; Chong, M.; Rasali, D.; Otterstatter, M.; Rose, C.; Prystajecky, N.; David, S.; Galanis, E.; et al. Mental Health and Substance Use Associated with Hospitalization among People with COVID-19: A Population-Based Cohort Study. Viruses 2021, 13, 2196. [Google Scholar] [CrossRef]
- Nishimi, K.; Neylan, T.C.; Bertenthal, D.; Dolsen, E.A.; Seal, K.H.; O’Donovan, A. Post-Traumatic Stress Disorder and Risk for Hospitalization and Death Following COVID-19 Infection. Transl. Psychiatry 2022, 12, 482. [Google Scholar] [CrossRef]
- Fond, G.; Pauly, V.; Leone, M.; Llorca, P.-M.; Orleans, V.; Loundou, A.; Lancon, C.; Auquier, P.; Baumstarck, K.; Boyer, L. Disparities in Intensive Care Unit Admission and Mortality among Patients with Schizophrenia and COVID-19: A National Cohort Study. Schizophr. Bull. 2021, 47, 624–634. [Google Scholar] [CrossRef]
- Nemani, K.; Li, C.; Olfson, M.; Blessing, E.M.; Razavian, N.; Chen, J.; Petkova, E.; Goff, D.C. Association of Psychiatric Disorders with Mortality among Patients with COVID-19. JAMA Psychiatry 2021, 78, 380–386. [Google Scholar] [CrossRef]
- Pardamean, E.; Roan, W.; Iskandar, K.T.A.; Prayangga, R.; Hariyanto, T.I. Mortality from Coronavirus Disease 2019 (COVID-19) in Patients with Schizophrenia: A Systematic Review, Meta-Analysis and Meta-Regression. Gen. Hosp. Psychiatry 2022, 75, 61–67. [Google Scholar] [CrossRef]
- Baillargeon, J.; Polychronopoulou, E.; Kuo, Y.-F.; Raji, M.A. The Impact of Substance Use Disorder on COVID-19 Outcomes. Psychiatr. Serv. 2021, 72, 578–581. [Google Scholar] [CrossRef]
- Hasin, D.S.; Fink, D.S.; Olfson, M.; Saxon, A.J.; Malte, C.; Keyes, K.M.; Gradus, J.L.; Cerdá, M.; Maynard, C.C.; Keyhani, S.; et al. Substance Use Disorders and COVID-19: An Analysis of Nation-Wide Veterans Health Administration Electronic Health Records. Drug Alcohol Depend. 2022, 234, 109383. [Google Scholar] [CrossRef]
- Lefere, S.; Tacke, F. Macrophages in Obesity and Non-Alcoholic Fatty Liver Disease: Crosstalk with Metabolism. JHEP Rep. 2019, 1, 30–43. [Google Scholar] [CrossRef]
- Da, B.L.; Im, G.Y.; Schiano, T.D. Coronavirus Disease 2019 Hangover: A Rising Tide of Alcohol Use Disorder and Alcohol-Associated Liver Disease. Hepatology 2020, 72, 1102–1108. [Google Scholar] [CrossRef] [PubMed]
- Zhao, B.; Ni, C.; Gao, R.; Wang, Y.; Yang, L.; Wei, J.; Lv, T.; Liang, J.; Zhang, Q.; Xu, W.; et al. Recapitulation of SARS-CoV-2 Infection and Cholangiocyte Damage with Human Liver Ductal Organoids. Protein Cell 2020, 11, 771–775. [Google Scholar] [CrossRef]
- Parveen, R.; Sehar, N.; Bajpai, R.; Agarwal, N.B. Association of Diabetes and Hypertension with Disease Severity in COVID-19 Patients: A Systematic Literature Review and Exploratory Meta-Analysis. Diabetes Res. Clin. Pract. 2020, 166, 108295. [Google Scholar] [CrossRef] [PubMed]
- Xu, D.; Zhou, F.; Sun, W.; Chen, L.; Lan, L.; Li, H.; Xiao, F.; Li, Y.; Kolachalama, V.B.; Li, Y.; et al. Relationship Between Serum Severe Acute Respiratory Syndrome Coronavirus 2 Nucleic Acid and Organ Damage in Coronavirus 2019 Patients: A Cohort Study. Clin. Infect. Dis. 2021, 73, 68–75. [Google Scholar] [CrossRef] [PubMed]
- Cai, Q.; Huang, D.; Yu, H.; Zhu, Z.; Xia, Z.; Su, Y.; Li, Z.; Zhou, G.; Gou, J.; Qu, J.; et al. COVID-19: Abnormal Liver Function Tests. J. Hepatol. 2020, 73, 566–574. [Google Scholar] [CrossRef] [PubMed]
- Finn, O.J. Immuno-Oncology: Understanding the Function and Dysfunction of the Immune System in Cancer. Ann. Oncol. 2012, 23, viii6–viii9. [Google Scholar] [CrossRef] [PubMed]
- Ménétrier-Caux, C.; Ray-Coquard, I.; Blay, J.-Y.; Caux, C. Lymphopenia in Cancer Patients and Its Effects on Response to Immunotherapy: An Opportunity for Combination with Cytokines? J. Immunother. Cancer 2019, 7, 85. [Google Scholar] [CrossRef] [PubMed]
- Chamilos, G.; Lionakis, M.S.; Kontoyiannis, D.P. Are All Patients with Cancer at Heightened Risk for Severe Coronavirus Disease 2019 (COVID-19)? Clin. Infect. Dis. 2021, 72, 351–356. [Google Scholar] [CrossRef]
- Di Felice, G.; Visci, G.; Teglia, F.; Angelini, M.; Boffetta, P. Effect of Cancer on Outcome of COVID-19 Patients: A Systematic Review and Meta-Analysis of Studies of Unvaccinated Patients. eLife 2022, 11, e74634. [Google Scholar] [CrossRef]
- Desai, A.; Gupta, R.; Advani, S.; Ouellette, L.; Kuderer, N.M.; Lyman, G.H.; Li, A. Mortality in Hospitalized Patients with Cancer and Coronavirus Disease 2019: A Systematic Review and Meta-Analysis of Cohort Studies. Cancer 2021, 127, 1459–1468. [Google Scholar] [CrossRef]
- Dai, M.; Liu, D.; Liu, M.; Zhou, F.; Li, G.; Chen, Z.; Zhang, Z.; You, H.; Wu, M.; Zheng, Q.; et al. Patients with Cancer Appear More Vulnerable to SARS-CoV-2: A Multicenter Study during the COVID-19 Outbreak. Cancer Discov. 2020, 10, 783–791. [Google Scholar] [CrossRef]
- Basse, C.; Diakite, S.; Servois, V.; Frelaut, M.; Noret, A.; Bellesoeur, A.; Moreau, P.; Massiani, M.-A.; Bouyer, A.-S.; Vuagnat, P.; et al. Characteristics and Outcome of SARS-CoV-2 Infection in Cancer Patients. JNCI Cancer Spectr. 2021, 5, pkaa090. [Google Scholar] [CrossRef]
- Yekedüz, E.; Utkan, G.; Ürün, Y. A Systematic Review and Meta-Analysis: The Effect of Active Cancer Treatment on Severity of COVID-19. Eur. J. Cancer 2020, 141, 92–104. [Google Scholar] [CrossRef]
- Robilotti, E.V.; Babady, N.E.; Mead, P.A.; Rolling, T.; Perez-Johnston, R.; Bernardes, M.; Bogler, Y.; Caldararo, M.; Figueroa, C.J.; Glickman, M.S.; et al. Determinants of COVID-19 Disease Severity in Patients with Cancer. Nat. Med. 2020, 26, 1218–1223. [Google Scholar] [CrossRef]
- Fagni, F.; Simon, D.; Tascilar, K.; Schoenau, V.; Sticherling, M.; Neurath, M.F.; Schett, G. COVID-19 and Immune-Mediated Inflammatory Diseases: Effect of Disease and Treatment on COVID-19 Outcomes and Vaccine Responses. Lancet Rheumatol. 2021, 3, e724–e736. [Google Scholar] [CrossRef]
- Chertok Shacham, E.; Ishay, A. New Insights on Effects of Glucocorticoids in SARS-CoV-2 Infection. Endocr. Pract. 2022, 28, 1100–1106. [Google Scholar] [CrossRef]
- Dexamethasone in Hospitalized Patients with COVID-19. N. Engl. J. Med. 2021, 384, 693–704. [CrossRef]
- Singanayagam, A.; Johnston, S.L. Long-Term Impact of Inhaled Corticosteroid Use in Asthma and Chronic Obstructive Pulmonary Disease (COPD): Review of Mechanisms That Underlie Risks. J. Allergy Clin. Immunol. 2020, 146, 1292–1294. [Google Scholar] [CrossRef]
- Singanayagam, A.; Glanville, N.; Cuthbertson, L.; Bartlett, N.W.; Finney, L.J.; Turek, E.; Bakhsoliani, E.; Calderazzo, M.A.; Trujillo-Torralbo, M.-B.; Footitt, J.; et al. Inhaled Corticosteroid Suppression of Cathelicidin Drives Dysbiosis and Bacterial Infection in Chronic Obstructive Pulmonary Disease. Sci. Transl. Med. 2019, 11, eaav3879. [Google Scholar] [CrossRef]
- Ameratunga, R.; Longhurst, H.; Steele, R.; Lehnert, K.; Leung, E.; Brooks, A.E.S.; Woon, S.-T. Common Variable Immunodeficiency Disorders, T-Cell Responses to SARS-CoV-2 Vaccines, and the Risk of Chronic COVID-19. J. Allergy Clin. Immunol. Pract. 2021, 9, 3575–3583. [Google Scholar] [CrossRef]
- Onisiforou, A.; Spyrou, G.M. Systems Bioinformatics Reveals Possible Relationship between COVID-19 and the Development of Neurological Diseases and Neuropsychiatric Disorders. Viruses 2022, 14, 2270. [Google Scholar] [CrossRef]
- Roy, E.R.; Wang, B.; Wan, Y.; Chiu, G.; Cole, A.; Yin, Z.; Propson, N.E.; Xu, Y.; Jankowsky, J.L.; Liu, Z.; et al. Type I Interferon Response Drives Neuroinflammation and Synapse Loss in Alzheimer Disease. J. Clin. Investig. 2020, 130, 1912–1930. [Google Scholar] [CrossRef]
- Finch, C.E.; Kulminski, A.M. The ApoE Locus and COVID-19: Are We Going Where We Have Been? J. Gerontol. A. Biol. Sci. Med. Sci. 2020, 76, e1–e3. [Google Scholar] [CrossRef] [PubMed]
- Chen, F.; Chen, Y.; Wang, Y.; Ke, Q.; Cui, L. The COVID-19 Pandemic and Alzheimer’s Disease: Mutual Risks and Mechanisms. Transl. Neurodegener. 2022, 11, 40. [Google Scholar] [CrossRef] [PubMed]
- Ebihara, S.; Saito, H.; Kanda, A.; Nakajoh, M.; Takahashi, H.; Arai, H.; Sasaki, H. Impaired Efficacy of Cough in Patients with Parkinson Disease. Chest 2003, 124, 1009–1015. [Google Scholar] [CrossRef] [PubMed]
- Dono, F.; Nucera, B.; Lanzone, J.; Evangelista, G.; Rinaldi, F.; Speranza, R.; Troisi, S.; Tinti, L.; Russo, M.; Di Pietro, M.; et al. Status Epilepticus and COVID-19: A Systematic Review. Epilepsy Behav. EB 2021, 118, 107887. [Google Scholar] [CrossRef]
- Romagnolo, A.; Imbalzano, G.; Artusi, C.A.; Balestrino, R.; Ledda, C.; De Rosa, F.G.; Riccardini, F.; Montanaro, E.; Bozzali, M.; Rizzone, M.G.; et al. Neurological Comorbidities and COVID-19-Related Case Fatality: A Cohort Study. J. Neurol. Sci. 2021, 428, 117610. [Google Scholar] [CrossRef]
- Galea, J.; Brough, D. The Role of Inflammation and Interleukin-1 in Acute Cerebrovascular Disease. J. Inflamm. Res. 2013, 6, 121–128. [Google Scholar] [CrossRef]
- Yuan, N.; Chen, Y.; Xia, Y.; Dai, J.; Liu, C. Inflammation-Related Biomarkers in Major Psychiatric Disorders: A Cross-Disorder Assessment of Reproducibility and Specificity in 43 Meta-Analyses. Transl. Psychiatry 2019, 9, 233. [Google Scholar] [CrossRef]
- Cauley, L.S.; Vella, A.T. Why Is Coinfection with Influenza Virus and Bacteria so Difficult to Control? Discov. Med. 2015, 19, 33–40. [Google Scholar]
- Gill, J.R.; Sheng, Z.-M.; Ely, S.F.; Guinee, D.G., Jr.; Beasley, M.B.; Suh, J.; Deshpande, C.; Mollura, D.J.; Morens, D.M.; Bray, M.; et al. Pulmonary Pathologic Findings of Fatal 2009 Pandemic Influenza A/H1N1 Viral Infections. Arch. Pathol. Lab. Med. 2010, 134, 235–243. [Google Scholar] [CrossRef]
- Bisno, A.L.; Griffin, J.P.; Van Epps, K.A.; Niell, H.B.; Rytel, M.W. Pneumonia and Hong Kong Influenza: A Prospective Study of the 1968-1969 Epidemic. Am. J. Med. Sci. 1971, 261, 251–263. [Google Scholar] [CrossRef]
- Omoush, S.A.; Alzyoud, J.A.M. The Prevalence and Impact of Coinfection and Superinfection on the Severity and Outcome of COVID-19 Infection: An Updated Literature Review. Pathogens 2022, 11, 445. [Google Scholar] [CrossRef] [PubMed]
- Musuuza, J.S.; Watson, L.; Parmasad, V.; Putman-Buehler, N.; Christensen, L.; Safdar, N. Prevalence and Outcomes of Co-Infection and Superinfection with SARS-CoV-2 and Other Pathogens: A Systematic Review and Meta-Analysis. PLoS ONE 2021, 16, e0251170. [Google Scholar] [CrossRef]
- Tsheten, T.; Clements, A.C.A.; Gray, D.J.; Adhikary, R.K.; Wangdi, K. Clinical Features and Outcomes of COVID-19 and Dengue Co-Infection: A Systematic Review. BMC Infect. Dis. 2021, 21, 729. [Google Scholar] [CrossRef]
- Simonnet, A.; Engelmann, I.; Moreau, A.-S.; Garcia, B.; Six, S.; El Kalioubie, A.; Robriquet, L.; Hober, D.; Jourdain, M. High Incidence of Epstein-Barr Virus, Cytomegalovirus, and Human-Herpes Virus-6 Reactivations in Critically Ill Patients with COVID-19. Infect. Dis. Now 2021, 51, 296–299. [Google Scholar] [CrossRef]
- Taherifard, E.; Movahed, H.; Kiani Salmi, S.; Taherifard, A.; Abdollahifard, S.; Taherifard, E. Cytomegalovirus Coinfection in Patients with Severe Acute Respiratory Syndrome Coronavirus 2 Infection: A Systematic Review of Reported Cases. Infect. Dis. 2022, 54, 543–557. [Google Scholar] [CrossRef]
- Singh, A.K.; Singh, R.; Joshi, S.R.; Misra, A. Mucormycosis in COVID-19: A Systematic Review of Cases Reported Worldwide and in India. Diabetes Metab. Syndr. Clin. Res. Rev. 2021, 15, 102146. [Google Scholar] [CrossRef]
- Szydłowicz, M.; Matos, O. Pneumocystis Pneumonia in the COVID-19 Pandemic Era: Similarities and Challenges. Trends Parasitol. 2021, 37, 859–862. [Google Scholar] [CrossRef]
- Hernández-Terán, A.; Mejía-Nepomuceno, F.; Herrera, M.T.; Barreto, O.; García, E.; Castillejos, M.; Boukadida, C.; Matias-Florentino, M.; Rincón-Rubio, A.; Avila-Rios, S.; et al. Dysbiosis and Structural Disruption of the Respiratory Microbiota in COVID-19 Patients with Severe and Fatal Outcomes. Sci. Rep. 2021, 11, 21297. [Google Scholar] [CrossRef]
- Sender, V.; Hentrich, K.; Henriques-Normark, B. Virus-Induced Changes of the Respiratory Tract Environment Promote Secondary Infections with Streptococcus Pneumoniae. Front. Cell. Infect. Microbiol. 2021, 11, 643326. [Google Scholar] [CrossRef]
- de Buhr, N.; von Köckritz-Blickwede, M. The Balance of Neutrophil Extracellular Trap Formation and Nuclease Degradation: An Unknown Role of Bacterial Coinfections in COVID-19 Patients? mBio 2021, 12, e03304-20. [Google Scholar] [CrossRef]
- Rangel, K.; Chagas, T.P.G.; De-Simone, S.G. Acinetobacter Baumannii Infections in Times of COVID-19 Pandemic. Pathogens 2021, 10, 1006. [Google Scholar] [CrossRef]
- Westblade, L.F.; Simon, M.S.; Satlin, M.J. Bacterial Coinfections in Coronavirus Disease 2019. Trends Microbiol. 2021, 29, 930–941. [Google Scholar] [CrossRef]
- Silva, D.L.; Lima, C.M.; Magalhães, V.C.R.; Baltazar, L.M.; Peres, N.T.A.; Caligiorne, R.B.; Moura, A.S.; Fereguetti, T.; Martins, J.C.; Rabelo, L.F.; et al. Fungal and Bacterial Coinfections Increase Mortality of Severely Ill COVID-19 Patients. J. Hosp. Infect. 2021, 113, 145–154. [Google Scholar] [CrossRef]
- Amin, D.; McKitish, K.; Shah, P.S. Association of Mortality and Recent Mycoplasma Pneumoniae Infection in COVID-19 Patients. J. Med. Virol. 2021, 93, 1180–1183. [Google Scholar] [CrossRef]
- Garcia-Vidal, C.; Sanjuan, G.; Moreno-García, E.; Puerta-Alcalde, P.; Garcia-Pouton, N.; Chumbita, M.; Fernandez-Pittol, M.; Pitart, C.; Inciarte, A.; Bodro, M.; et al. Incidence of Co-Infections and Superinfections in Hospitalized Patients with COVID-19: A Retrospective Cohort Study. Clin. Microbiol. Infect. 2021, 27, 83. [Google Scholar] [CrossRef]
- Hedberg, P.; Johansson, N.; Ternhag, A.; Abdel-Halim, L.; Hedlund, J.; Nauclér, P. Bacterial Co-Infections in Community-Acquired Pneumonia Caused by SARS-CoV-2, Influenza Virus and Respiratory Syncytial Virus. BMC Infect. Dis. 2022, 22, 108. [Google Scholar] [CrossRef]
- Falcone, M.; Tiseo, G.; Giordano, C.; Leonildi, A.; Menichini, M.; Vecchione, A.; Pistello, M.; Guarracino, F.; Ghiadoni, L.; Forfori, F.; et al. Predictors of Hospital-Acquired Bacterial and Fungal Superinfections in COVID-19: A Prospective Observational Study. J. Antimicrob. Chemother. 2021, 76, 1078–1084. [Google Scholar] [CrossRef]
- Wang, L.; Amin, A.K.; Khanna, P.; Aali, A.; McGregor, A.; Bassett, P.; Gopal Rao, G. An Observational Cohort Study of Bacterial Co-Infection and Implications for Empirical Antibiotic Therapy in Patients Presenting with COVID-19 to Hospitals in North West London. J. Antimicrob. Chemother. 2021, 76, 796–803. [Google Scholar] [CrossRef]
- Mitsi, E.; Reiné, J.; Urban, B.C.; Solórzano, C.; Nikolaou, E.; Hyder-Wright, A.D.; Pojar, S.; Howard, A.; Hitchins, L.; Glynn, S.; et al. Streptococcus Pneumoniae Colonization Associates with Impaired Adaptive Immune Responses against SARS-CoV-2. J. Clin. Investig. 2022, 132, e157124. [Google Scholar] [CrossRef]
- SARS-CoV-2, Bacterial Co-Infections, and AMR: The Deadly Trio in COVID-19? EMBO Mol. Med. 2020, 12, e12560. [CrossRef]
- Duployez, C.; Guern, R.L.; Tinez, C.; Lejeune, A.-L.; Robriquet, L.; Six, S.; Loïez, C.; Wallet, F. Panton-Valentine Leukocidin–Secreting Staphylococcus Aureus Pneumonia Complicating COVID-19. Emerg. Infect. Dis. 2020, 26, 1939. [Google Scholar] [CrossRef]
- Nieto-Moro, M.; Ecclesia, F.G.; Tomé-Masa, I.; De Lama Caro-Patón, G.; Leoz-Gordillo, I.; Cabrero-Hernández, M.; García-Salido, A. SARS-CoV-2 and Streptococcus Pneumoniae Coinfection as a Cause of Severe Pneumonia in an Infant. Pediatr. Pulmonol. 2020, 55, 2198–2200. [Google Scholar] [CrossRef]
- Riccò, M.; Ferraro, P.; Peruzzi, S.; Zaniboni, A.; Ranzieri, S. SARS-CoV-2–Legionella Co-Infections: A Systematic Review and Meta-Analysis (2020–2021). Microorganisms 2022, 10, 499. [Google Scholar] [CrossRef]
- Shah, T.; Shah, Z.; Yasmeen, N.; Baloch, Z.; Xia, X. Pathogenesis of SARS-CoV-2 and Mycobacterium Tuberculosis Coinfection. Front. Immunol. 2022, 13, 909011. [Google Scholar] [CrossRef]
- Sarkar, S.; Khanna, P.; Singh, A.K. Impact of COVID-19 in Patients with Concurrent Co-Infections: A Systematic Review and Meta-Analyses. J. Med. Virol. 2021, 93, 2385–2395. [Google Scholar] [CrossRef]
- Sy, K.T.L.; Haw, N.J.L.; Uy, J. Previous and Active Tuberculosis Increases Risk of Death and Prolongs Recovery in Patients with COVID-19. Infect. Dis. 2020, 52, 902–907. [Google Scholar] [CrossRef]
- Western Cape Department of Health in collaboration with the National Institute for Communicable Diseases, South Africa; Boulle, A.; Davies, M.-A.; Hussey, H.; Ismail, M.; Morden, E.; Vundle, Z.; Zweigenthal, V.; Mahomed, H.; Paleker, M.; et al. Risk Factors for Coronavirus Disease 2019 (COVID-19) Death in a Population Cohort Study from the Western Cape Province, South Africa. Clin. Infect. Dis. 2021, 73, e2005–e2015. [Google Scholar] [CrossRef]
- Mejia, O.R.; Gloag, E.S.; Li, J.; Ruane-Foster, M.; Claeys, T.A.; Farkas, D.; Wang, S.-H.; Farkas, L.; Xin, G.; Robinson, R.T. Mice Infected with Mycobacterium Tuberculosis Are Resistant to Acute Disease Caused by Secondary Infection with SARS-CoV-2. PLOS Pathog. 2022, 18, e1010093. [Google Scholar] [CrossRef]
- TB/COVID-19 Global Study Group. Tuberculosis and COVID-19 Co-Infection: Description of the Global Cohort. Eur. Respir. J. 2022, 59, 2102538. [Google Scholar] [CrossRef]
- Starshinova, A.A.; Kudryavtsev, I.; Malkova, A.; Zinchenko, U.; Karev, V.; Kudlay, D.; Glushkova, A.; Starshinova, A.Y.; Dominguez, J.; Villar-Hernández, R.; et al. Molecular and Cellular Mechanisms of M. Tuberculosis and SARS-CoV-2 Infections—Unexpected Similarities of Pathogenesis and What to Expect from Co-Infection. Int. J. Mol. Sci. 2022, 23, 2235. [Google Scholar] [CrossRef]
- Acharya, D.; Liu, G.; Gack, M.U. Dysregulation of Type I Interferon Responses in COVID-19. Nat. Rev. Immunol. 2020, 20, 397–398. [Google Scholar] [CrossRef]
- Cliff, J.M.; Kaufmann, S.H.E.; McShane, H.; van Helden, P.; O’Garra, A. The Human Immune Response to Tuberculosis and Its Treatment: A View from the Blood. Immunol. Rev. 2015, 264, 88–102. [Google Scholar] [CrossRef]
- Riou, C.; du Bruyn, E.; Stek, C.; Daroowala, R.; Goliath, R.T.; Abrahams, F.; Said-Hartley, Q.; Allwood, B.W.; Hsiao, N.-Y.; Wilkinson, K.A.; et al. Relationship of SARS-CoV-2–Specific CD4 Response to COVID-19 Severity and Impact of HIV-1 and Tuberculosis Coinfection. J. Clin. Investig. 2021, 131, e149125. [Google Scholar] [CrossRef]
- Petrone, L.; Petruccioli, E.; Vanini, V.; Cuzzi, G.; Gualano, G.; Vittozzi, P.; Nicastri, E.; Maffongelli, G.; Grifoni, A.; Sette, A.; et al. Coinfection of Tuberculosis and COVID-19 Limits the Ability to in Vitro Respond to SARS-CoV-2. Int. J. Infect. Dis. 2021, 113, S82–S87. [Google Scholar] [CrossRef]
- Wong, G.L.-H.; Wong, V.W.-S.; Yuen, B.W.-Y.; Tse, Y.-K.; Yip, T.C.-F.; Luk, H.W.-S.; Lui, G.C.-Y.; Chan, H.L.-Y. Risk of Hepatitis B Surface Antigen Seroreversion after Corticosteroid Treatment in Patients with Previous Hepatitis B Virus Exposure. J. Hepatol. 2020, 72, 57–66. [Google Scholar] [CrossRef]
- Yu, R.; Tan, S.; Dan, Y.; Lu, Y.; Zhang, J.; Tan, Z.; He, X.; Xiang, X.; Zhou, Y.; Guo, Y.; et al. Effect of SARS-CoV-2 Coinfection Was Not Apparent on the Dynamics of Chronic Hepatitis B Infection. Virology 2021, 553, 131–134. [Google Scholar] [CrossRef]
- Yang, S.; Wang, S.; Du, M.; Liu, M.; Liu, Y.; He, Y. Patients with COVID-19 and HBV Coinfection Are at Risk of Poor Prognosis. Infect. Dis. Ther. 2022, 11, 1229–1242. [Google Scholar] [CrossRef]
- Kang, S.H.; Cho, D.-H.; Choi, J.; Baik, S.K.; Gwon, J.G.; Kim, M.Y. Association between Chronic Hepatitis B Infection and COVID-19 Outcomes: A Korean Nationwide Cohort Study. PLoS ONE 2021, 16, e0258229. [Google Scholar] [CrossRef]
- Zou, X.; Fang, M.; Li, S.; Wu, L.; Gao, B.; Gao, H.; Ran, X.; Bian, Y.; Li, R.; Yu, S.; et al. Characteristics of Liver Function in Patients with SARS-CoV-2 and Chronic HBV Coinfection. Clin. Gastroenterol. Hepatol. 2021, 19, 597–603. [Google Scholar] [CrossRef]
- Lin, Y.; Yuan, J.; Long, Q.; Hu, J.; Deng, H.; Zhao, Z.; Chen, J.; Lu, M.; Huang, A. Patients with SARS-CoV-2 and HBV Co-Infection Are at Risk of Greater Liver Injury. Genes Dis. 2021, 8, 484–492. [Google Scholar] [CrossRef]
- Ali, N. Relationship Between COVID-19 Infection and Liver Injury: A Review of Recent Data. Front. Med. 2020, 7, 458. [Google Scholar] [CrossRef] [PubMed]
- Xiang, T.-D.; Zheng, X. Interaction between Hepatitis B Virus and SARS-CoV-2 Infections. World J. Gastroenterol. 2021, 27, 782–793. [Google Scholar] [CrossRef] [PubMed]
- Alothaid, H.; Aldughaim, M.S.K.; El Bakkouri, K.; AlMashhadi, S.; Al-Qahtani, A.A. Similarities between the Effect of SARS-CoV-2 and HCV on the Cellular Level, and the Possible Role of Ion Channels in COVID19 Progression: A Review of Potential Targets for Diagnosis and Treatment. Channels 2020, 14, 403–412. [Google Scholar] [CrossRef] [PubMed]
- Butt, A.A.; Yan, P.; Chotani, R.A.; Shaikh, O.S. Mortality Is Not Increased in SARS-CoV-2 Infected Persons with Hepatitis C Virus Infection. Liver Int. 2021, 41, 1824–1831. [Google Scholar] [CrossRef]
- Rehermann, B. Hepatitis C Virus versus Innate and Adaptive Immune Responses: A Tale of Coevolution and Coexistence. J. Clin. Investig. 2009, 119, 1745–1754. [Google Scholar] [CrossRef]
- Loftis, J.M.; Huckans, M.; Ruimy, S.; Hinrichs, D.J.; Hauser, P. Depressive Symptoms in Patients with Chronic Hepatitis C Are Correlated with Elevated Plasma Levels of Interleukin-1β and Tumor Necrosis Factor-α. Neurosci. Lett. 2008, 430, 264–268. [Google Scholar] [CrossRef]
- Shirley, K.; Loftis, J.M. A Spotlight on HCV and SARS-CoV-2 Co-Infection and Brain Function. Pharmacol. Biochem. Behav. 2022, 217, 173403. [Google Scholar] [CrossRef]
- Gill, K.; Ghazinian, H.; Manch, R.; Gish, R. Hepatitis C Virus as a Systemic Disease: Reaching beyond the Liver. Hepatol. Int. 2016, 10, 415–423. [Google Scholar] [CrossRef]
- Afify, S.; Eysa, B.; Hamid, F.A.; Abo-Elazm, O.M.; Edris, M.A.; Maher, R.; Abdelhalim, A.; Abdel Ghaffar, M.M.; Omran, D.A.; Shousha, H.I. Survival and Outcomes for Co-Infection of Chronic Hepatitis C with and without Cirrhosis and COVID-19: A Multicenter Retrospective Study. World J. Gastroenterol. 2021, 27, 7362–7375. [Google Scholar] [CrossRef]
- Lambarey, H.; Blumenthal, M.J.; Chetram, A.; Joyimbana, W.; Jennings, L.; Tincho, M.B.; Burgers, W.A.; Orrell, C.; Schäfer, G. SARS-CoV-2 Infection Is Associated with Uncontrolled HIV Viral Load in Non-Hospitalized HIV-Infected Patients from Gugulethu, South Africa. Viruses 2022, 14, 1222. [Google Scholar] [CrossRef]
- Moradi, Y.; Soheili, M.; Dehghanbanadaki, H.; Moradi, G.; Moradpour, F.; Mortazavi, S.M.M.; Kohan, H.G.; Zareie, M. The Effect of HIV/AIDS Infection on the Clinical Outcomes of COVID-19: A Meta-Analysis. J. Pharm. Pharm. Sci. 2022, 25, 183–192. [Google Scholar] [CrossRef]
- Danwang, C.; Noubiap, J.J.; Robert, A.; Yombi, J.C. Outcomes of Patients with HIV and COVID-19 Co-Infection: A Systematic Review and Meta-Analysis. AIDS Res. Ther. 2022, 19, 3. [Google Scholar] [CrossRef]
- Yang, X.; Sun, J.; Patel, R.C.; Zhang, J.; Guo, S.; Zheng, Q.; Olex, A.L.; Olatosi, B.; Weissman, S.B.; Islam, J.Y.; et al. Associations between HIV Infection and Clinical Spectrum of COVID-19: A Population Level Analysis Based on US National COVID Cohort Collaborative (N3C) Data. Lancet HIV 2021, 8, e690–e700. [Google Scholar] [CrossRef]
- Bhaskaran, K.; Rentsch, C.T.; MacKenna, B.; Schultze, A.; Mehrkar, A.; Bates, C.J.; Eggo, R.M.; Morton, C.E.; Bacon, S.C.J.; Inglesby, P.; et al. HIV Infection and COVID-19 Death: A Population-Based Cohort Analysis of UK Primary Care Data and Linked National Death Registrations within the OpenSAFELY Platform. Lancet HIV 2021, 8, e24–e32. [Google Scholar] [CrossRef]
- Hoffmann, C.; Casado, J.L.; Härter, G.; Vizcarra, P.; Moreno, A.; Cattaneo, D.; Meraviglia, P.; Spinner, C.D.; Schabaz, F.; Grunwald, S.; et al. Immune Deficiency Is a Risk Factor for Severe COVID-19 in People Living with HIV. HIV Med. 2021, 22, 372–378. [Google Scholar] [CrossRef]
- Ssentongo, P.; Heilbrunn, E.S.; Ssentongo, A.E.; Advani, S.; Chinchilli, V.M.; Nunez, J.J.; Du, P. Epidemiology and Outcomes of COVID-19 in HIV-Infected Individuals: A Systematic Review and Meta-Analysis. Sci. Rep. 2021, 11, 6283. [Google Scholar] [CrossRef]
- Tesoriero, J.M.; Swain, C.-A.E.; Pierce, J.L.; Zamboni, L.; Wu, M.; Holtgrave, D.R.; Gonzalez, C.J.; Udo, T.; Morne, J.E.; Hart-Malloy, R.; et al. COVID-19 Outcomes among Persons Living with or without Diagnosed HIV Infection in New York State. JAMA Netw. Open 2021, 4, e2037069. [Google Scholar] [CrossRef]
- Nomah, D.K.; Reyes-Urueña, J.; Díaz, Y.; Moreno, S.; Aceiton, J.; Bruguera, A.; Vivanco-Hidalgo, R.M.; Llibre, J.M.; Domingo, P.; Falcó, V.; et al. Sociodemographic, Clinical, and Immunological Factors Associated with SARS-CoV-2 Diagnosis and Severe COVID-19 Outcomes in People Living with HIV: A Retrospective Cohort Study. Lancet HIV 2021, 8, e701–e710. [Google Scholar] [CrossRef]
- Mirzaei, H.; McFarland, W.; Karamouzian, M.; Sharifi, H. COVID-19 among People Living with HIV: A Systematic Review. AIDS Behav. 2021, 25, 85–92. [Google Scholar] [CrossRef]
- Sharov, K.S. HIV/SARS-CoV-2 Co-Infection: T Cell Profile, Cytokine Dynamics and Role of Exhausted Lymphocytes. Int. J. Infect. Dis. 2021, 102, 163–169. [Google Scholar] [CrossRef]
- Krause, R.; Snyman, J.; Shi-Hsia, H.; Muema, D.; Karim, F.; Ganga, Y.; Ngoepe, A.; Zungu, Y.; Gazy, I.; Bernstein, M.; et al. HIV Skews the SARS-CoV-2 B Cell Response towards an Extrafollicular Maturation Pathway. eLife 2022, 11, e79924. [Google Scholar] [CrossRef]
- Corma-Gómez, A.; Fernández-Fuertes, M.; García, E.; Fuentes-López, A.; Gómez-Ayerbe, C.; Rivero-Juárez, A.; Domínguez, C.; Santos, M.; Viñuela, L.; Palacios, R.; et al. Severe Immunosuppression Is Related to Poorer Immunogenicity to SARS-CoV-2 Vaccines among People Living with HIV. Clin. Microbiol. Infect. 2022, 28, 1492–1498. [Google Scholar] [CrossRef]
- Hassold, N.; Brichler, S.; Ouedraogo, E.; Leclerc, D.; Carroue, S.; Gater, Y.; Alloui, C.; Carbonnelle, E.; Bouchaud, O.; Mechai, F.; et al. Impaired Antibody Response to COVID-19 Vaccination in Advanced HIV Infection. AIDS Lond. Engl. 2022, 36, F1–F5. [Google Scholar] [CrossRef]
- Woodruff, M.C.; Ramonell, R.P.; Nguyen, D.C.; Cashman, K.S.; Saini, A.S.; Haddad, N.S.; Ley, A.M.; Kyu, S.; Howell, J.C.; Ozturk, T.; et al. Extrafollicular B Cell Responses Correlate with Neutralizing Antibodies and Morbidity in COVID-19. Nat. Immunol. 2020, 21, 1506–1516. [Google Scholar] [CrossRef]
- Chen, Y.; Zuiani, A.; Fischinger, S.; Mullur, J.; Atyeo, C.; Travers, M.; Lelis, F.J.N.; Pullen, K.M.; Martin, H.; Tong, P.; et al. Quick COVID-19 Healers Sustain Anti-SARS-CoV-2 Antibody Production. Cell 2020, 183, 1496–1507. [Google Scholar] [CrossRef] [PubMed]
- Karim, F.; Gazy, I.; Cele, S.; Zungu, Y.; Krause, R.; Bernstein, M.; Khan, K.; Ganga, Y.; Rodel, H.; Mthabela, N.; et al. HIV Status Alters Disease Severity and Immune Cell Responses in Beta Variant SARS-CoV-2 Infection Wave. eLife 2021, 10, e67397. [Google Scholar] [CrossRef]
- Lagathu, C.; Cossarizza, A.; Béréziat, V.; Nasi, M.; Capeau, J.; Pinti, M. Basic Science and Pathogenesis of Ageing with HIV: Potential Mechanisms and Biomarkers. AIDS Lond. Engl. 2017, 31 (Suppl. 2), S105–S119. [Google Scholar] [CrossRef]
- Ho, H.; Peluso, M.J.; Margus, C.; Matias Lopes, J.P.; He, C.; Gaisa, M.M.; Osorio, G.; Aberg, J.A.; Mullen, M.P. Clinical Outcomes and Immunologic Characteristics of Coronavirus Disease 2019 in People with Human Immunodeficiency Virus. J. Infect. Dis. 2021, 223, 403–408. [Google Scholar] [CrossRef]
- Rosenthal, E.M.; Rosenberg, E.S.; Patterson, W.; Ferguson, W.P.; Gonzalez, C.; DeHovitz, J.; Udo, T.; Rajulu, D.T.; Hart-Malloy, R.; Tesoriero, J. Factors Associated with SARS-CoV-2-Related Hospital Outcomes among and between Persons Living with and without Diagnosed HIV Infection in New York State. PLoS ONE 2022, 17, e0268978. [Google Scholar] [CrossRef] [PubMed]
- Ziegler, C.G.K.; Allon, S.J.; Nyquist, S.K.; Mbano, I.M.; Miao, V.N.; Tzouanas, C.N.; Cao, Y.; Yousif, A.S.; Bals, J.; Hauser, B.M.; et al. SARS-CoV-2 Receptor ACE2 Is an Interferon-Stimulated Gene in Human Airway Epithelial Cells and Is Detected in Specific Cell Subsets across Tissues. Cell 2020, 181, 1016–1035. [Google Scholar] [CrossRef]
- Killip, M.J.; Fodor, E.; Randall, R.E. Influenza Virus Activation of the Interferon System. Virus Res. 2015, 209, 11–22. [Google Scholar] [CrossRef] [PubMed]
- Swets, M.C.; Russell, C.D.; Harrison, E.M.; Docherty, A.B.; Lone, N.; Girvan, M.; Hardwick, H.E.; Visser, L.G.; Openshaw, P.J.M.; Groeneveld, G.H.; et al. SARS-CoV-2 Co-Infection with Influenza Viruses, Respiratory Syncytial Virus, or Adenoviruses. Lancet 2022, 399, 1463–1464. [Google Scholar] [CrossRef] [PubMed]
- Stowe, J.; Tessier, E.; Zhao, H.; Guy, R.; Muller-Pebody, B.; Zambon, M.; Andrews, N.; Ramsay, M.; Lopez Bernal, J. Interactions between SARS-CoV-2 and Influenza, and the Impact of Coinfection on Disease Severity: A Test-Negative Design. Int. J. Epidemiol. 2021, 50, 1124–1133. [Google Scholar] [CrossRef] [PubMed]
- Guan, Z.; Chen, C.; Li, Y.; Yan, D.; Zhang, X.; Jiang, D.; Yang, S.; Li, L. Impact of Coinfection with SARS-CoV-2 and Influenza on Disease Severity: A Systematic Review and Meta-Analysis. Front. Public Health 2021, 9, 1944. [Google Scholar] [CrossRef]
- Ma, S.; Lai, X.; Chen, Z.; Tu, S.; Qin, K. Clinical Characteristics of Critically Ill Patients Co-Infected with SARS-CoV-2 and the Influenza Virus in Wuhan, China. Int. J. Infect. Dis. 2020, 96, 683–687. [Google Scholar] [CrossRef]
- Zheng, J.; Chen, F.; Wu, K.; Wang, J.; Li, F.; Huang, S.; Lu, J.; Huang, J.; Liu, H.; Zhou, R.; et al. Clinical and Virological Impact of Single and Dual Infections with Influenza A (H1N1) and SARS-CoV-2 in Adult Inpatients. PLoS Negl. Trop. Dis. 2021, 15, e0009997. [Google Scholar] [CrossRef]
- Kinoshita, T.; Watanabe, K.; Sakurai, Y.; Nishi, K.; Yoshikawa, R.; Yasuda, J. Co-Infection of SARS-CoV-2 and Influenza Virus Causes More Severe and Prolonged Pneumonia in Hamsters. Sci. Rep. 2021, 11, 21259. [Google Scholar] [CrossRef]
- Bai, L.; Zhao, Y.; Dong, J.; Liang, S.; Guo, M.; Liu, X.; Wang, X.; Huang, Z.; Sun, X.; Zhang, Z.; et al. Coinfection with Influenza A Virus Enhances SARS-CoV-2 Infectivity. Cell Res. 2021, 31, 395–403. [Google Scholar] [CrossRef]
- Achdout, H.; Vitner, E.B.; Politi, B.; Melamed, S.; Yahalom-Ronen, Y.; Tamir, H.; Erez, N.; Avraham, R.; Weiss, S.; Cherry, L.; et al. Increased Lethality in Influenza and SARS-CoV-2 Coinfection Is Prevented by Influenza Immunity but Not SARS-CoV-2 Immunity. Nat. Commun. 2021, 12, 5819. [Google Scholar] [CrossRef]
- Fink, G.; Orlova-Fink, N.; Schindler, T.; Grisi, S.; Ferrer, A.P.S.; Daubenberger, C.; Brentani, A. Inactivated Trivalent Influenza Vaccination Is Associated with Lower Mortality among Patients with COVID-19 in Brazil. BMJ Evid.-Based Med. 2021, 26, 192–193. [Google Scholar] [CrossRef]
- Behrouzi, B.; Araujo Campoverde, M.V.; Liang, K.; Talbot, H.K.; Bogoch, I.I.; McGeer, A.; Fröbert, O.; Loeb, M.; Vardeny, O.; Solomon, S.D.; et al. Influenza Vaccination to Reduce Cardiovascular Morbidity and Mortality in Patients with COVID-19: JACC State-of-the-Art Review. J. Am. Coll. Cardiol. 2020, 76, 1777–1794. [Google Scholar] [CrossRef] [PubMed]
- Fage, C.; Hénaut, M.; Carbonneau, J.; Piret, J.; Boivin, G. Influenza A(H1N1)Pdm09 Virus but Not Respiratory Syncytial Virus Interferes with SARS-CoV-2 Replication during Sequential Infections in Human Nasal Epithelial Cells. Viruses 2022, 14, 395. [Google Scholar] [CrossRef]
- Halfmann, P.J.; Nakajima, N.; Sato, Y.; Takahashi, K.; Accola, M.; Chiba, S.; Fan, S.; Neumann, G.; Rehrauer, W.; Suzuki, T.; et al. SARS-CoV-2 Interference of Influenza Virus Replication in Syrian Hamsters. J. Infect. Dis. 2022, 225, 282–286. [Google Scholar] [CrossRef] [PubMed]
- Murphy, R.C.; Lai, Y.; Barrow, K.A.; Hamerman, J.A.; Lacy-Hulbert, A.; Piliponsky, A.M.; Ziegler, S.F.; Altemeier, W.A.; Debley, J.S.; Gharib, S.A.; et al. Effects of Asthma and Human Rhinovirus A16 on the Expression of SARS-CoV-2 Entry Factors in Human Airway Epithelium. Am. J. Respir. Cell Mol. Biol. 2020, 63, 859–863. [Google Scholar] [CrossRef] [PubMed]
- Scotta, M.C.; Kern, L.B.; Polese-Bonatto, M.; Azevedo, T.R.; Varela, F.H.; Zavaglia, G.O.; Fernandes, I.R.; de David, C.N.; Fazolo, T.; da Costa, M.S.C.; et al. Impact of Rhinovirus on Hospitalization during the COVID-19 Pandemic: A Prospective Cohort Study. J. Clin. Virol. 2022, 156, 105197. [Google Scholar] [CrossRef]
- Le Glass, E.; Hoang, V.T.; Boschi, C.; Ninove, L.; Zandotti, C.; Boutin, A.; Bremond, V.; Dubourg, G.; Ranque, S.; Lagier, J.-C.; et al. Incidence and Outcome of Coinfections with SARS-CoV-2 and Rhinovirus. Viruses 2021, 13, 2528. [Google Scholar] [CrossRef] [PubMed]
- Dee, K.; Goldfarb, D.M.; Haney, J.; Amat, J.A.R.; Herder, V.; Stewart, M.; Szemiel, A.M.; Baguelin, M.; Murcia, P.R. Human Rhinovirus Infection Blocks Severe Acute Respiratory Syndrome Coronavirus 2 Replication within the Respiratory Epithelium: Implications for COVID-19 Epidemiology. J. Infect. Dis. 2021, 224, 31–38. [Google Scholar] [CrossRef] [PubMed]
- Vanderwall, E.R.; Barrow, K.A.; Rich, L.M.; Read, D.F.; Trapnell, C.; Okoloko, O.; Ziegler, S.F.; Hallstrand, T.S.; White, M.P.; Debley, J.S. Airway Epithelial Interferon Response to SARS-CoV-2 Is Inferior to Rhinovirus and Heterologous Rhinovirus Infection Suppresses SARS-CoV-2 Replication. Sci. Rep. 2022, 12, 6972. [Google Scholar] [CrossRef]
- Soni, S.; Namdeo Pudake, R.; Jain, U.; Chauhan, N. A Systematic Review on SARS-CoV-2-Associated Fungal Coinfections. J. Med. Virol. 2022, 94, 99–109. [Google Scholar] [CrossRef]
- Amin, A.; Vartanian, A.; Poladian, N.; Voloshko, A.; Yegiazaryan, A.; Al-Kassir, A.L.; Venketaraman, V. Root Causes of Fungal Coinfections in COVID-19 Infected Patients. Infect. Dis. Rep. 2021, 13, 1018–1035. [Google Scholar] [CrossRef]
- Salmanton-García, J.; Sprute, R.; Stemler, J.; Bartoletti, M.; Dupont, D.; Valerio, M.; Garcia-Vidal, C.; Falces-Romero, I.; Machado, M.; de la Villa, S.; et al. COVID-19-Associated Pulmonary Aspergillosis, March–August 2020. Emerg. Infect. Dis. 2021, 27, 1077–1086. [Google Scholar] [CrossRef] [PubMed]
- Lai, C.-C.; Yu, W.-L. COVID-19 Associated with Pulmonary Aspergillosis: A Literature Review. J. Microbiol. Immunol. Infect. 2021, 54, 46–53. [Google Scholar] [CrossRef] [PubMed]
- Shen, H.-P.; Tang, Y.-M.; Song, H.; Xu, W.-Q.; Yang, S.-L.; Xu, X.-J. Efficiency of Interleukin 6 and Interferon Gamma in the Differentiation of Invasive Pulmonary Aspergillosis and Pneumocystis Pneumonia in Pediatric Oncology Patients. Int. J. Infect. Dis. 2016, 48, 73–77. [Google Scholar] [CrossRef] [PubMed]
- Camargo, J.F.; Bhimji, A.; Kumar, D.; Kaul, R.; Pavan, R.; Schuh, A.; Seftel, M.; Lipton, J.H.; Gupta, V.; Humar, A.; et al. Impaired T Cell Responsiveness to Interleukin-6 in Hematological Patients with Invasive Aspergillosis. PLoS ONE 2015, 10, e0123171. [Google Scholar] [CrossRef]
- Clemons, K.V.; Grunig, G.; Sobel, R.A.; Mirels, L.F.; Rennick, D.M.; Stevens, D.A. Role of IL-10 in Invasive Aspergillosis: Increased Resistance of IL-10 Gene Knockout Mice to Lethal Systemic Aspergillosis. Clin. Exp. Immunol. 2000, 122, 186–191. [Google Scholar] [CrossRef]
- Del Sero, G.; Mencacci, A.; Cenci, E.; Fè d’Ostiani, C.; Montagnoli, C.; Bacci, A.; Mosci, P.; Kopf, M.; Romani, L. Antifungal Type 1 Responses Are Upregulated in IL-10-Deficient Mice. Microbes Infect. 1999, 1, 1169–1180. [Google Scholar] [CrossRef]
- Segrelles-Calvo, G.; de S Araújo, G.R.; Llopis-Pastor, E.; Carrillo, J.; Hernández-Hernández, M.; Rey, L.; Melean, N.R.; Escribano, I.; Antón, E.; Zamarro, C.; et al. Candida Spp. Co-Infection in COVID-19 Patients with Severe Pneumonia: Prevalence Study and Associated Risk Factors. Respir. Med. 2021, 188, 106619. [Google Scholar] [CrossRef]
- Castro, M.; Bjoraker, J.A.; Rohrbach, M.S.; Limper, A.H. Candida Albicans Induces the Release of Inflammatory Mediators from Human Peripheral Blood Monocytes. Inflammation 1996, 20, 107–122. [Google Scholar] [CrossRef]
- Steinshamn, S.; Waage, A. Tumor Necrosis Factor and Interleukin-6 in Candida Albicans Infection in Normal and Granulocytopenic Mice. Infect. Immun. 1992, 60, 4003–4008. [Google Scholar] [CrossRef]
- Campbell, L.; Hepworth, M.R.; Whittingham-Dowd, J.; Thompson, S.; Bancroft, A.J.; Hayes, K.S.; Shaw, T.N.; Dickey, B.F.; Flamar, A.-L.; Artis, D.; et al. ILC2s Mediate Systemic Innate Protection by Priming Mucus Production at Distal Mucosal Sites. J. Exp. Med. 2019, 216, 2714–2723. [Google Scholar] [CrossRef]
- Wolday, D.; Gebrecherkos, T.; Arefaine, Z.G.; Kiros, Y.K.; Gebreegzabher, A.; Tasew, G.; Abdulkader, M.; Abraha, H.E.; Desta, A.A.; Hailu, A.; et al. Effect of Co-Infection with Intestinal Parasites on COVID-19 Severity: A Prospective Observational Cohort Study. EClinicalMedicine 2021, 39, 101054. [Google Scholar] [CrossRef] [PubMed]
- Cai, P.; Mu, Y.; McManus, D.P. The Fight Against Severe COVID-19: Can Parasitic Worms Contribute? Front. Immunol. 2022, 13, 849465. [Google Scholar] [CrossRef] [PubMed]
- Ademe, M.; Girma, F. The Influence of Helminth Immune Regulation on COVID-19 Clinical Outcomes: Is It Beneficial or Detrimental? Infect. Drug Resist. 2021, 14, 4421–4426. [Google Scholar] [CrossRef]
- Wolday, D.; Tasew, G.; Amogne, W.; Urban, B.; Schallig, H.D.; Harris, V.; Rinke de Wit, T.F. Interrogating the Impact of Intestinal Parasite-Microbiome on Pathogenesis of COVID-19 in Sub-Saharan Africa. Front. Microbiol. 2021, 12, 614522. [Google Scholar] [CrossRef] [PubMed]
- Paniz-Mondolfi, A.E.; Ramírez, J.D.; Delgado-Noguera, L.A.; Rodriguez-Morales, A.J.; Sordillo, E.M. COVID-19 and Helminth Infection: Beyond the Th1/Th2 Paradigm. PLoS Negl. Trop. Dis. 2021, 15, e0009402. [Google Scholar] [CrossRef]
- Hussein, R.; Guedes, M.; Ibraheim, N.; Ali, M.M.; El-Tahir, A.; Allam, N.; Abuakar, H.; Pecoits-Filho, R.; Kotanko, P. Impact of COVID-19 and Malaria Coinfection on Clinical Outcomes: A Retrospective Cohort Study. Clin. Microbiol. Infect. 2022, 28, 1152.e1–1152.e6. [Google Scholar] [CrossRef]
- Herrmann, M.; Schulte, S.; Wildner, N.H.; Wittner, M.; Brehm, T.T.; Ramharter, M.; Woost, R.; Lohse, A.W.; Jacobs, T.; Schulze zur Wiesch, J. Analysis of Co-Inhibitory Receptor Expression in COVID-19 Infection Compared to Acute Plasmodium Falciparum Malaria: LAG-3 and TIM-3 Correlate with T Cell Activation and Course of Disease. Front. Immunol. 2020, 11, 1870. [Google Scholar] [CrossRef]
- Wildner, N.H.; Ahmadi, P.; Schulte, S.; Brauneck, F.; Kohsar, M.; Lütgehetmann, M.; Beisel, C.; Addo, M.M.; Haag, F.; Schulze Zur Wiesch, J. B Cell Analysis in SARS-CoV-2 versus Malaria: Increased Frequencies of Plasmablasts and Atypical Memory B Cells in COVID-19. J. Leukoc. Biol. 2021, 109, 77–90. [Google Scholar] [CrossRef]
- Mahajan, N.N.; Gajbhiye, R.K.; Bahirat, S.; Lokhande, P.D.; Mathe, A.; Rathi, S.; Warty, N.; Mahajan, K.N.; Srivastava, V.; Kuppusamy, P.; et al. Co-Infection of Malaria and Early Clearance of SARS-CoV-2 in Healthcare Workers. J. Med. Virol. 2021, 93, 2431–2438. [Google Scholar] [CrossRef]
- Molina, I.; Marcolino, M.S.; Pires, M.C.; Ramos, L.E.F.; Silva, R.T.; Guimarães-Júnior, M.H.; de Oliveira, I.J.R.; de Carvalho, R.L.R.; Nunes, A.G.S.; de Barros, A.L.R.M.; et al. Chagas Disease and SARS-CoV-2 Coinfection Does Not Lead to Worse in-Hospital Outcomes. Sci. Rep. 2021, 11, 20289. [Google Scholar] [CrossRef]
- Fernandez, M.L.; Benchetrit, A.; Astudillo, O.G.; Garay, A.M.; De Vedia, L.; Garcia Bournissen, F.; Lloveras, S.C.; Orduna, T.A.; Gonzalez, G.D. COVID-19 and Chagas Disease in Buenos Aires, Argentina. Front. Trop. Dis. 2022, 2, 779428. [Google Scholar] [CrossRef]
- Golda, A.; Malek, N.; Dudek, B.; Zeglen, S.; Wojarski, J.; Ochman, M.; Kucewicz, E.; Zembala, M.; Potempa, J.; Pyrc, K. 2011 Infection with Human Coronavirus NL63 Enhances Streptococcal Adherence to Epithelial Cells. J. Gen. Virol. 2011, 92, 1358–1368. [Google Scholar] [CrossRef] [PubMed]
- Ramphal, R.; Small, P.M.; Shands, J.W.; Fischlschweiger, W.; Small, P.A. Adherence of Pseudomonas Aeruginosa to Tracheal Cells Injured by Influenza Infection or by Endotracheal Intubation. Infect. Immun. 1980, 27, 614–619. [Google Scholar] [CrossRef] [PubMed]
- Rowe, H.M.; Meliopoulos, V.A.; Iverson, A.; Bomme, P.; Schultz-Cherry, S.; Rosch, J.W. Direct Interactions with Influenza Promote Bacterial Adherence during Respiratory Infections. Nat. Microbiol. 2019, 4, 1328–1336. [Google Scholar] [CrossRef] [PubMed]
- Robinot, R.; Hubert, M.; de Melo, G.D.; Lazarini, F.; Bruel, T.; Smith, N.; Levallois, S.; Larrous, F.; Fernandes, J.; Gellenoncourt, S.; et al. SARS-CoV-2 Infection Induces the Dedifferentiation of Multiciliated Cells and Impairs Mucociliary Clearance. Nat. Commun. 2021, 12, 4354. [Google Scholar] [CrossRef] [PubMed]
- Avadhanula, V.; Rodriguez, C.A.; De Vincenzo, J.P.; Wang, Y.; Webby, R.J.; Ulett, G.C.; Adderson, E.E. Respiratory Viruses Augment the Adhesion of Bacterial Pathogens to Respiratory Epithelium in a Viral Species- and Cell Type-Dependent Manner. J. Virol. 2006, 80, 1629–1636. [Google Scholar] [CrossRef]
- Larriva, M.A.-D.; Martín-DeLeon, R.; Royo, B.U.; Fernández-Navamuel, I.; Velando, A.G.; García, L.N.; Clemente, C.C.; García, F.A.; Codern, A.R.; Fernández-Arias, C.; et al. The Role of Bronchoscopy in Patients with SARS-CoV-2 Pneumonia. ERJ Open Res. 2021, 7, 00165. [Google Scholar] [CrossRef]
- Meyerholz, D.K.; Reznikov, L.R. Influence of SARS-CoV-2 on Airway Mucus Production: A Review and Proposed Model. Vet. Pathol. 2022, 59, 578–585. [Google Scholar] [CrossRef]
- Østergaard, L. SARS CoV-2 Related Microvascular Damage and Symptoms during and after COVID-19: Consequences of Capillary Transit-Time Changes, Tissue Hypoxia and Inflammation. Physiol. Rep. 2021, 9, e14726. [Google Scholar] [CrossRef]
- Noel-Savina, E.; Viatgé, T.; Faviez, G.; Lepage, B.; Mhanna, L.T.; Pontier, S.; Dupuis, M.; Collot, S.; Thomas, P.; Idoate Lacasia, J.; et al. Severe SARS-CoV-2 Pneumonia: Clinical, Functional and Imaging Outcomes at 4 Months. Respir. Med. Res. 2021, 80, 100822. [Google Scholar] [CrossRef]
- Barbeta, E.; Motos, A.; Torres, A.; Ceccato, A.; Ferrer, M.; Cilloniz, C.; Bueno, L.; Badia, J.R.; Castro, P.; Ferrando, C.; et al. SARS-CoV-2–Induced Acute Respiratory Distress Syndrome: Pulmonary Mechanics and Gas-Exchange Abnormalities. Ann. Am. Thorac. Soc. 2020, 17, 1164–1168. [Google Scholar] [CrossRef]
- Ackermann, M.; Anders, H.-J.; Bilyy, R.; Bowlin, G.L.; Daniel, C.; De Lorenzo, R.; Egeblad, M.; Henneck, T.; Hidalgo, A.; Hoffmann, M.; et al. Patients with COVID-19: In the Dark-NETs of Neutrophils. Cell Death Differ. 2021, 28, 3125–3139. [Google Scholar] [CrossRef] [PubMed]
- Paludan, S.R.; Mogensen, T.H. Innate Immunological Pathways in COVID-19 Pathogenesis. Sci. Immunol. 2022, 7, eabm5505. [Google Scholar] [CrossRef] [PubMed]
- Feldman, C.; Anderson, R. The Role of Co-Infections and Secondary Infections in Patients with COVID-19. Pneumonia 2021, 13, 5. [Google Scholar] [CrossRef] [PubMed]
- He, S.; Liu, W.; Jiang, M.; Huang, P.; Xiang, Z.; Deng, D.; Chen, P.; Xie, L. Clinical Characteristics of COVID-19 Patients with Clinically Diagnosed Bacterial Co-Infection: A Multi-Center Study. PLoS ONE 2021, 16, e0249668. [Google Scholar] [CrossRef] [PubMed]
- Sulaiman, I.; Chung, M.; Angel, L.; Tsay, J.-C.J.; Wu, B.G.; Yeung, S.T.; Krolikowski, K.; Li, Y.; Duerr, R.; Schluger, R.; et al. Microbial Signatures in the Lower Airways of Mechanically Ventilated COVID-19 Patients Associated with Poor Clinical Outcome. Nat. Microbiol. 2021, 6, 1245–1258. [Google Scholar] [CrossRef] [PubMed]
- Cele, S.; Karim, F.; Lustig, G.; San, J.E.; Hermanus, T.; Tegally, H.; Snyman, J.; Moyo-Gwete, T.; Wilkinson, E.; Bernstein, M.; et al. SARS-CoV-2 Prolonged Infection during Advanced HIV Disease Evolves Extensive Immune Escape. Cell Host Microbe 2022, 30, 154–162. [Google Scholar] [CrossRef] [PubMed]
- Chen, J.; Song, J.; Dai, L.; Post, S.R.; Qin, Z. SARS-CoV-2 Infection and Lytic Reactivation of Herpesviruses: A Potential Threat in the Postpandemic Era? J. Med. Virol. 2022, 94, 5103–5111. [Google Scholar] [CrossRef] [PubMed]
- Pathak, L.; Gayan, S.; Pal, B.; Talukdar, J.; Bhuyan, S.; Sandhya, S.; Yeger, H.; Baishya, D.; Das, B. Coronavirus Activates an Altruistic Stem Cell-Mediated Defense Mechanism That Reactivates Dormant Tuberculosis: Implications in Coronavirus Disease 2019 Pandemic. Am. J. Pathol. 2021, 191, 1255–1268. [Google Scholar] [CrossRef]
- Pakzad, R.; Malekifar, P.; Shateri, Z.; Zandi, M.; Akhavan Rezayat, S.; Soleymani, M.; Karimi, M.R.; Ahmadi, S.E.; Shahbahrami, R.; Pakzad, I.; et al. Worldwide Prevalence of Microbial Agents’ Coinfection among COVID-19 Patients: A Comprehensive Updated Systematic Review and Meta-Analysis. J. Clin. Lab. Anal. 2022, 36, e24151. [Google Scholar] [CrossRef]
- Alhumaid, S.; Al Mutair, A.; Al Alawi, Z.; Alshawi, A.M.; Alomran, S.A.; Almuhanna, M.S.; Almuslim, A.A.; Bu Shafia, A.H.; Alotaibi, A.M.; Ahmed, G.Y.; et al. Coinfections with Bacteria, Fungi, and Respiratory Viruses in Patients with SARS-CoV-2: A Systematic Review and Meta-Analysis. Pathogens 2021, 10, 809. [Google Scholar] [CrossRef]
- Langford, B.J.; So, M.; Raybardhan, S.; Leung, V.; Westwood, D.; MacFadden, D.R.; Soucy, J.-P.R.; Daneman, N. Bacterial Co-Infection and Secondary Infection in Patients with COVID-19: A Living Rapid Review and Meta-Analysis. Clin. Microbiol. Infect. 2020, 26, 1622–1629. [Google Scholar] [CrossRef] [PubMed]
- Bassetti, M.; Kollef, M.H.; Timsit, J.-F. Bacterial and Fungal Superinfections in Critically Ill Patients with COVID-19. Intensive Care Med. 2020, 46, 2071–2074. [Google Scholar] [CrossRef] [PubMed]
- Malekifar, P.; Pakzad, R.; Shahbahrami, R.; Zandi, M.; Jafarpour, A.; Rezayat, S.A.; Akbarpour, S.; Shabestari, A.N.; Pakzad, I.; Hesari, E.; et al. Viral Coinfection among COVID-19 Patient Groups: An Update Systematic Review and Meta-Analysis. BioMed Res. Int. 2021, 2021, 5313832. [Google Scholar] [CrossRef] [PubMed]
- Krumbein, H.; Kümmel, L.S.; Fragkou, P.C.; Thölken, C.; Hünerbein, B.L.; Reiter, R.; Papathanasiou, K.A.; Renz, H.; Skevaki, C. Respiratory Viral Co-Infections in Patients with COVID-19 and Associated Outcomes: A Systematic Review and Meta-Analysis. Rev. Med. Virol. 2022, e2365. [Google Scholar] [CrossRef]
- Singh, V.; Upadhyay, P.; Reddy, J.; Granger, J. SARS-CoV-2 Respiratory Co-Infections: Incidence of Viral and Bacterial Co-Pathogens. Int. J. Infect. Dis. 2021, 105, 617–620. [Google Scholar] [CrossRef]
- Hoque, M.N.; Akter, S.; Mishu, I.D.; Islam, M.R.; Rahman, M.S.; Akhter, M.; Islam, I.; Hasan, M.M.; Rahaman, M.M.; Sultana, M.; et al. Microbial Co-Infections in COVID-19: Associated Microbiota and Underlying Mechanisms of Pathogenesis. Microb. Pathog. 2021, 156, 104941. [Google Scholar] [CrossRef]
- Goldberg, E.M.; Hasegawa, K.; Lawrence, A.; Kline, J.A.; Camargo, C.A. Viral Coinfection Is Associated with Improved Outcomes in Emergency Department Patients with SARS-CoV-2. West. J. Emerg. Med. 2021, 22, 1262–1269. [Google Scholar] [CrossRef]
- Chekuri, S.; Szymczak, W.A.; Goldstein, D.Y.; Nori, P.; Marrero Rolon, R.; Spund, B.; Singh-Tan, S.; Mohrmann, L.; Assa, A.; Southern, W.N.; et al. SARS-CoV-2 Coinfection with Additional Respiratory Virus Does Not Predict Severe Disease: A Retrospective Cohort Study. J. Antimicrob. Chemother. 2021, 76, iii12–iii19. [Google Scholar] [CrossRef]
- Kumar, N.; Sharma, S.; Barua, S.; Tripathi, B.N.; Rouse, B.T. Virological and Immunological Outcomes of Coinfections. Clin. Microbiol. Rev. 2018, 31, e00111-17. [Google Scholar] [CrossRef]
- Peng, J.; Wang, Q.; Mei, H.; Zheng, H.; Liang, G.; She, X.; Liu, W. Fungal Co-Infection in COVID-19 Patients: Evidence from a Systematic Review and Meta-Analysis. Aging 2021, 13, 7745–7757. [Google Scholar] [CrossRef]
- Sreenath, K.; Batra, P.; Vinayaraj, E.V.; Bhatia, R.; SaiKiran, K.; Singh, V.; Singh, S.; Verma, N.; Singh, U.B.; Mohan, A.; et al. Coinfections with Other Respiratory Pathogens among Patients with COVID-19. Microbiol. Spectr. 2021, 9, e00163-21. [Google Scholar] [CrossRef] [PubMed]
- Narayanan, S.; Chua, J.V.; Baddley, J.W. Coronavirus Disease 2019-Associated Mucormycosis: Risk Factors and Mechanisms of Disease. Clin. Infect. Dis. Off. Publ. Infect. Dis. Soc. Am. 2022, 74, 1279–1283. [Google Scholar] [CrossRef] [PubMed]
- Muthu, V.; Rudramurthy, S.M.; Chakrabarti, A.; Agarwal, R. Epidemiology and Pathophysiology of COVID-19-Associated Mucormycosis: India Versus the Rest of the World. Mycopathologia 2021, 186, 739–754. [Google Scholar] [CrossRef] [PubMed]
- Abdel-Hamed, E.F.; Ibrahim, M.N.; Mostafa, N.E.; Moawad, H.S.F.; Elgammal, N.E.; Darwiesh, E.M.; El-rafey, D.S.; ElBadawy, N.E.; Al-Khoufi, E.A.; Hindawi, S.I. Role of Interferon Gamma in SARS-CoV-2-Positive Patients with Parasitic Infections. Gut Pathog. 2021, 13, 29. [Google Scholar] [CrossRef]
- Wilairatana, P.; Masangkay, F.R.; Kotepui, K.U.; Milanez, G.D.J.; Kotepui, M. Prevalence and Characteristics of Malaria among COVID-19 Individuals: A Systematic Review, Meta-Analysis, and Analysis of Case Reports. PLoS Negl. Trop. Dis. 2021, 15, e0009766. [Google Scholar] [CrossRef]
- Leng, S.; Chen, X.; Mao, G. Frailty Syndrome: An Overview. Clin. Interv. Aging 2014, 9, 433. [Google Scholar] [CrossRef]
- World Health Organization. WHO Clinical Consortium on Healthy Ageing (Topic Focus: Frailty and Intrinsic Capacity); World Health Organization: Geneva, Switzerland, 2016; p. 36. [Google Scholar]
- Morley, J.E.; Vellas, B.; Kan, G.A.V.; Anker, S.D.; Bauer, J.M.; Bernabei, R.; Cesari, M.; Chumlea, W.C.; Doehner, W.; Evans, J.; et al. Frailty Consensus: A Call to Action. J. Am. Med. Dir. Assoc. 2013, 14, 392–397. [Google Scholar] [CrossRef]
- O’Caoimh, R.; Sezgin, D.; O’Donovan, M.R.; Molloy, D.W.; Clegg, A.; Rockwood, K.; Liew, A. Prevalence of Frailty in 62 Countries across the World: A Systematic Review and Meta-Analysis of Population-Level Studies. Age Ageing 2021, 50, 96–104. [Google Scholar] [CrossRef]
- Kojima, G. Prevalence of Frailty in Nursing Homes: A Systematic Review and Meta-Analysis. J. Am. Med. Dir. Assoc. 2015, 16, 940–945. [Google Scholar] [CrossRef]
- Collard, R.M.; Boter, H.; Schoevers, R.A.; Oude Voshaar, R.C. Prevalence of Frailty in Community-Dwelling Older Persons: A Systematic Review. J. Am. Geriatr. Soc. 2012, 60, 1487–1492. [Google Scholar] [CrossRef]
- Rockwood, K.; Song, X.; MacKnight, C.; Bergman, H.; Hogan, D.B.; McDowell, I.; Mitnitski, A. A Global Clinical Measure of Fitness and Frailty in Elderly People. CMAJ 2005, 173, 489–495. [Google Scholar] [CrossRef] [PubMed]
- Hewitt, J.; Carter, B.; Vilches-Moraga, A.; Quinn, T.J.; Braude, P.; Verduri, A.; Pearce, L.; Stechman, M.; Short, R.; Price, A.; et al. The Effect of Frailty on Survival in Patients with COVID-19 (COPE): A Multicentre, European, Observational Cohort Study. Lancet Public Health 2020, 5, e444–e451. [Google Scholar] [CrossRef] [PubMed]
- Sablerolles, R.S.G.; Lafeber, M.; van Kempen, J.A.L.; van de Loo, B.P.A.; Boersma, E.; Rietdijk, W.J.R.; Polinder-Bos, H.A.; Mooijaart, S.P.; van der Kuy, H.; Versmissen, J.; et al. Association between Clinical Frailty Scale Score and Hospital Mortality in Adult Patients with COVID-19 (COMET): An International, Multicentre, Retrospective, Observational Cohort Study. Lancet Healthy Longev. 2021, 2, e163–e170. [Google Scholar] [CrossRef] [PubMed]
- Kundi, H.; Çetin, E.H.Ö.; Canpolat, U.; Aras, S.; Celik, O.; Ata, N.; Birinci, S.; Çay, S.; Özeke, Ö.; Tanboğa, I.H.; et al. The Role of Frailty on Adverse Outcomes among Older Patients with COVID-19. J. Infect. 2020, 81, 944–951. [Google Scholar] [CrossRef] [PubMed]
- Izurieta, H.S.; Graham, D.J.; Jiao, Y.; Hu, M.; Lu, Y.; Wu, Y.; Chillarige, Y.; Wernecke, M.; Menis, M.; Pratt, D.; et al. Natural History of Coronavirus Disease 2019: Risk Factors for Hospitalizations and Deaths among >26 Million US Medicare Beneficiaries. J. Infect. Dis. 2021, 223, 945–956. [Google Scholar] [CrossRef] [PubMed]
- Kastora, S.; Kounidas, G.; Perrott, S.; Carter, B.; Hewitt, J.; Myint, P.K. Clinical Frailty Scale as a Point of Care Prognostic Indicator of Mortality in COVID-19: A Systematic Review and Meta-Analysis. EClinicalMedicine 2021, 36, 100896. [Google Scholar] [CrossRef] [PubMed]
- Rottler, M.; Ocskay, K.; Sipos, Z.; Görbe, A.; Virág, M.; Hegyi, P.; Molnár, T.; Erőss, B.; Leiner, T.; Molnár, Z. Clinical Frailty Scale (CFS) Indicated Frailty Is Associated with Increased in-Hospital and 30-Day Mortality in COVID-19 Patients: A Systematic Review and Meta-Analysis. Ann. Intensive Care 2022, 12, 17. [Google Scholar] [CrossRef]
- Blomaard, L.C.; van der Linden, C.M.J.; van der Bol, J.M.; Jansen, S.W.M.; Polinder-Bos, H.A.; Willems, H.C.; Festen, J.; Barten, D.G.; Borgers, A.J.; Bos, J.C.; et al. Frailty Is Associated with In-Hospital Mortality in Older Hospitalised COVID-19 Patients in the Netherlands: The COVID-OLD Study. Age Ageing 2021, 50, 631–640. [Google Scholar] [CrossRef]
- Subramaniam, A.; Anstey, C.; Curtis, J.R.; Ashwin, S.; Ponnapa Reddy, M.; Aliberti, M.J.R.; Avelino-Silva, T.J.; Welch, C.; Koduri, G.; Prowle, J.R.; et al. Characteristics and Outcomes of Patients with Frailty Admitted to ICU with Coronavirus Disease 2019: An Individual Patient Data Meta-Analysis. Crit. Care Explor. 2022, 4, e0616. [Google Scholar] [CrossRef]
- Taniguchi, L.U.; Avelino-Silva, T.J.; Dias, M.B.; Jacob-Filho, W.; Aliberti, M.J.R.; CO-FRAIL Study Group; EPICCoV Study Group; COVID HCFMUSP Study Group. Association of Frailty, Organ Support, and Long-Term Survival in Critically Ill Patients with COVID-19. Crit. Care Explor. 2022, 4, e0712. [Google Scholar] [CrossRef]
- Pranata, R.; Henrina, J.; Lim, M.A.; Lawrensia, S.; Yonas, E.; Vania, R.; Huang, I.; Lukito, A.A.; Suastika, K.; Kuswardhani, R.A.T.; et al. Clinical Frailty Scale and Mortality in COVID-19: A Systematic Review and Dose-Response Meta-Analysis. Arch. Gerontol. Geriatr. 2021, 93, 104324. [Google Scholar] [CrossRef] [PubMed]
- Kojima, G. Frailty Defined by FRAIL Scale as a Predictor of Mortality: A Systematic Review and Meta-Analysis. J. Am. Med. Dir. Assoc. 2018, 19, 480–483. [Google Scholar] [CrossRef] [PubMed]
- Kojima, G.; Iliffe, S.; Walters, K. Frailty Index as a Predictor of Mortality: A Systematic Review and Meta-Analysis. Age Ageing 2018, 47, 193–200. [Google Scholar] [CrossRef] [PubMed]
- Wanhella, K.J.; Fernandez-Patron, C. Biomarkers of Ageing and Frailty May Predict COVID-19 Severity. Ageing Res. Rev. 2022, 73, 101513. [Google Scholar] [CrossRef]
- Hussien, H.; Nastasa, A.; Apetrii, M.; Nistor, I.; Petrovic, M.; Covic, A. Different Aspects of Frailty and COVID-19: Points to Consider in the Current Pandemic and Future Ones. BMC Geriatr. 2021, 21, 389. [Google Scholar] [CrossRef]
- She, Q.; Chen, B.; Liu, W.; Li, M.; Zhao, W.; Wu, J. Frailty Pathogenesis, Assessment, and Management in Older Adults with COVID-19. Front. Med. 2021, 8, 694367. [Google Scholar] [CrossRef]
- Gu, S.; Chen, Y.; Wu, Z.; Chen, Y.; Gao, H.; Lv, L.; Guo, F.; Zhang, X.; Luo, R.; Huang, C.; et al. Alterations of the Gut Microbiota in Patients with Coronavirus Disease 2019 or H1N1 Influenza. Clin. Infect. Dis. 2020, 71, 2669–2678. [Google Scholar] [CrossRef]
- Haiminen, N.; Utro, F.; Seabolt, E.; Parida, L. Functional Profiling of COVID-19 Respiratory Tract Microbiomes. Sci. Rep. 2021, 11, 6433. [Google Scholar] [CrossRef]
- Servin, A.L. Antagonistic Activities of Lactobacilli and Bifidobacteria against Microbial Pathogens. FEMS Microbiol. Rev. 2004, 28, 405–440. [Google Scholar] [CrossRef]
- Bodera, P.; Chcialowski, A. Immunomodulatory Effect of Probiotic Bacteria. Recent Pat. Inflamm. Allergy Drug Discov. 2009, 3, 58–64. [Google Scholar] [CrossRef]
- Vallianou, N.; Stratigou, T.; Christodoulatos, G.S.; Dalamaga, M. Understanding the Role of the Gut Microbiome and Microbial Metabolites in Obesity and Obesity-Associated Metabolic Disorders: Current Evidence and Perspectives. Curr. Obes. Rep. 2019, 8, 317–332. [Google Scholar] [CrossRef] [PubMed]
- Bozkurt, H.S.; Bilen, Ö. Oral Booster Probiotic Bifidobacteria in SARS-COV-2 Patients. Int. J. Immunopathol. Pharmacol. 2021, 35, 20587384211059676. [Google Scholar] [CrossRef] [PubMed]
- Li, Q.; Cheng, F.; Xu, Q.; Su, Y.; Cai, X.; Zeng, F.; Zhang, Y. The Role of Probiotics in Coronavirus Disease-19 Infection in Wuhan: A Retrospective Study of 311 Severe Patients. Int. Immunopharmacol. 2021, 95, 107531. [Google Scholar] [CrossRef] [PubMed]
- Hegazy, M.; Ashoush, O.A.; Hegazy, M.T.; Wahba, M.; Lithy, R.M.; Abdel-Hamid, H.M.; Elshafy, S.A.A.; Abdelfatah, D.; Ibrahim, M.H.E.-D.; Abdelghani, A. Beyond Probiotic Legend: ESSAP Gut Microbiota Health Score to Delineate SARS-COV-2 Infection Severity. Br. J. Nutr. 2022, 127, 1180–1189. [Google Scholar] [CrossRef] [PubMed]
- Llor, C.; Ouchi, D.; Giner-Soriano, M.; García-Sangenís, A.; Bjerrum, L.; Morros, R. Correlation between Previous Antibiotic Exposure and COVID-19 Severity. A Population-Based Cohort Study. Antibiotics 2021, 10, 1364. [Google Scholar] [CrossRef]
- Yin, X.; Xu, X.; Li, H.; Jiang, N.; Wang, J.; Lu, Z.; Xiong, N.; Gong, Y. Evaluation of Early Antibiotic Use in Patients with Non-Severe COVID-19 without Bacterial Infection. Int. J. Antimicrob. Agents 2022, 59, 106462. [Google Scholar] [CrossRef]
- Man, W.H.; de Steenhuijsen Piters, W.A.A.; Bogaert, D. The Microbiota of the Respiratory Tract: Gatekeeper to Respiratory Health. Nat. Rev. Microbiol. 2017, 15, 259–270. [Google Scholar] [CrossRef]
- Kau, A.L.; Ahern, P.P.; Griffin, N.W.; Goodman, A.L.; Gordon, J.I. Human Nutrition, the Gut Microbiome and the Immune System. Nature 2011, 474, 327–336. [Google Scholar] [CrossRef]
- Iebba, V.; Zanotta, N.; Campisciano, G.; Zerbato, V.; Di Bella, S.; Cason, C.; Luzzati, R.; Confalonieri, M.; Palamara, A.T.; Comar, M. Profiling of Oral Microbiota and Cytokines in COVID-19 Patients. Front. Microbiol. 2021, 12, 1603. [Google Scholar] [CrossRef]
- Gupta, A.; Bhanushali, S.; Sanap, A.; Shekatkar, M.; Kharat, A.; Raut, C.; Bhonde, R.; Shouche, Y.; Kheur, S.; Sharma, A. Oral Dysbiosis and Its Linkage with SARS-CoV-2 Infection. Microbiol. Res. 2022, 261, 127055. [Google Scholar] [CrossRef]
- Tchoupou Saha, O.L.F.; Dubourg, G.; Yacouba, A.; Bossi, V.; Raoult, D.; Lagier, J.-C. Profile of the Nasopharyngeal Microbiota Affecting the Clinical Course in COVID-19 Patients. Front. Microbiol. 2022, 13, 871627. [Google Scholar] [CrossRef] [PubMed]
- Han, Y.; Jia, Z.; Shi, J.; Wang, W.; He, K. The Active Lung Microbiota Landscape of COVID-19 Patients through the Metatranscriptome Data Analysis. BioImpacts 2021, 12, 139–146. [Google Scholar] [CrossRef] [PubMed]
- Gaibani, P.; Viciani, E.; Bartoletti, M.; Lewis, R.E.; Tonetti, T.; Lombardo, D.; Castagnetti, A.; Bovo, F.; Horna, C.S.; Ranieri, M.; et al. The Lower Respiratory Tract Microbiome of Critically Ill Patients with COVID-19. Sci. Rep. 2021, 11, 10103. [Google Scholar] [CrossRef] [PubMed]
- Hurst, J.H.; McCumber, A.W.; Aquino, J.N.; Rodriguez, J.; Heston, S.M.; Lugo, D.J.; Rotta, A.T.; Turner, N.A.; Pfeiffer, T.S.; Gurley, T.C.; et al. Age-Related Changes in the Nasopharyngeal Microbiome Are Associated with Severe Acute Respiratory Syndrome Coronavirus 2 (SARS-CoV-2) Infection and Symptoms among Children, Adolescents, and Young Adults. Clin. Infect. Dis. 2022, 75, e928–e937. [Google Scholar] [CrossRef] [PubMed]
- Soffritti, I.; D’Accolti, M.; Fabbri, C.; Passaro, A.; Manfredini, R.; Zuliani, G.; Libanore, M.; Franchi, M.; Contini, C.; Caselli, E. Oral Microbiome Dysbiosis Is Associated with Symptoms Severity and Local Immune/Inflammatory Response in COVID-19 Patients: A Cross-Sectional Study. Front Microbiol 2021, 12, 687513. [Google Scholar] [CrossRef]
- Hoque, M.N.; Sarkar, M.M.H.; Rahman, M.S.; Akter, S.; Banu, T.A.; Goswami, B.; Jahan, I.; Hossain, M.S.; Shamsuzzaman, A.K.M.; Nafisa, T.; et al. SARS-CoV-2 Infection Reduces Human Nasopharyngeal Commensal Microbiome with Inclusion of Pathobionts. Sci. Rep. 2021, 11, 24042. [Google Scholar] [CrossRef]
- Ren, L.; Wang, Y.; Zhong, J.; Li, X.; Xiao, Y.; Li, J.; Yang, J.; Fan, G.; Guo, L.; Shen, Z.; et al. Dynamics of the Upper Respiratory Tract Microbiota and Its Association with Mortality in COVID-19. Am. J. Respir. Crit. Care Med. 2021, 204, 1379–1390. [Google Scholar] [CrossRef]
- Smith, N.; Goncalves, P.; Charbit, B.; Grzelak, L.; Beretta, M.; Planchais, C.; Bruel, T.; Rouilly, V.; Bondet, V.; Hadjadj, J.; et al. Distinct Systemic and Mucosal Immune Responses during Acute SARS-CoV-2 Infection. Nat. Immunol. 2021, 22, 1428–1439. [Google Scholar] [CrossRef]
- Deriu, E.; Boxx, G.M.; He, X.; Pan, C.; Benavidez, S.D.; Cen, L.; Rozengurt, N.; Shi, W.; Cheng, G. Influenza Virus Affects Intestinal Microbiota and Secondary Salmonella Infection in the Gut through Type I Interferons. PLOS Pathog. 2016, 12, e1005572. [Google Scholar] [CrossRef]
- Steed, A.L.; Christophi, G.P.; Kaiko, G.E.; Sun, L.; Goodwin, V.M.; Jain, U.; Esaulova, E.; Artyomov, M.N.; Morales, D.J.; Holtzman, M.J.; et al. The Microbial Metabolite Desaminotyrosine Protects from Influenza through Type I Interferon. Science 2017, 357, 498–502. [Google Scholar] [CrossRef]
- Ichinohe, T.; Pang, I.K.; Kumamoto, Y.; Peaper, D.R.; Ho, J.H.; Murray, T.S.; Iwasaki, A. Microbiota Regulates Immune Defense against Respiratory Tract Influenza A Virus Infection. Proc. Natl. Acad. Sci. USA 2011, 108, 5354–5359. [Google Scholar] [CrossRef] [PubMed]
- Liu, Q.; Mak, J.W.Y.; Su, Q.; Yeoh, Y.K.; Lui, G.C.-Y.; Ng, S.S.S.; Zhang, F.; Li, A.Y.L.; Lu, W.; Hui, D.S.-C.; et al. Gut Microbiota Dynamics in a Prospective Cohort of Patients with Post-Acute COVID-19 Syndrome. Gut 2022, 71, 544–552. [Google Scholar] [CrossRef] [PubMed]
- Dong, Z.; Xiang, B.-J.; Jiang, M.; Sun, M.; Dai, C. The Prevalence of Gastrointestinal Symptoms, Abnormal Liver Function, Digestive System Disease and Liver Disease in COVID-19 Infection. J. Clin. Gastroenterol. 2021, 55, 67–76. [Google Scholar] [CrossRef] [PubMed]
- Zarifian, A.; Zamiri Bidary, M.; Arekhi, S.; Rafiee, M.; Gholamalizadeh, H.; Amiriani, A.; Ghaderi, M.S.; Khadem-Rezaiyan, M.; Amini, M.; Ganji, A. Gastrointestinal and Hepatic Abnormalities in Patients with Confirmed COVID-19: A Systematic Review and Meta-Analysis. J. Med. Virol. 2021, 93, 336–350. [Google Scholar] [CrossRef] [PubMed]
- Yusuf, F.; Fahriani, M.; Mamada, S.S.; Frediansyah, A.; Abubakar, A.; Maghfirah, D.; Fajar, J.K.; Maliga, H.A.; Ilmawan, M.; Emran, T.B.; et al. Global Prevalence of Prolonged Gastrointestinal Symptoms in COVID-19 Survivors and Potential Pathogenesis: A Systematic Review and Meta-Analysis. F1000Research 2021, 10, 301. [Google Scholar] [CrossRef] [PubMed]
- Chhibber-Goel, J.; Gopinathan, S.; Sharma, A. Interplay between Severities of COVID-19 and the Gut Microbiome: Implications of Bacterial Co-Infections? Gut Pathog. 2021, 13, 14. [Google Scholar] [CrossRef]
- Zuo, T.; Zhan, H.; Zhang, F.; Liu, Q.; Tso, E.Y.K.; Lui, G.C.Y.; Chen, N.; Li, A.; Lu, W.; Chan, F.K.L.; et al. Alterations in Fecal Fungal Microbiome of Patients with COVID-19 During Time of Hospitalization until Discharge. Gastroenterology 2020, 159, 1302–1310. [Google Scholar] [CrossRef]
- Xu, X.; Zhang, W.; Guo, M.; Xiao, C.; Fu, Z.; Yu, S.; Jiang, L.; Wang, S.; Ling, Y.; Liu, F.; et al. Integrated Analysis of Gut Microbiome and Host Immune Responses in COVID-19. Front. Med. 2022, 16, 263–275. [Google Scholar] [CrossRef]
- Ventero, M.P.; Cuadrat, R.R.C.; Vidal, I.; Andrade, B.G.N.; Molina-Pardines, C.; Haro-Moreno, J.M.; Coutinho, F.H.; Merino, E.; Regitano, L.C.A.; Silveira, C.B.; et al. Nasopharyngeal Microbial Communities of Patients Infected with SARS-CoV-2 That Developed COVID-19. Front. Microbiol. 2021, 12, 560. [Google Scholar] [CrossRef]
- Albrich, W.C.; Ghosh, T.S.; Ahearn-Ford, S.; Mikaeloff, F.; Lunjani, N.; Forde, B.; Suh, N.; Kleger, G.-R.; Pietsch, U.; Frischknecht, M.; et al. A High-Risk Gut Microbiota Configuration Associates with Fatal Hyperinflammatory Immune and Metabolic Responses to SARS-CoV-2. Gut Microbes 2022, 14, 2073131. [Google Scholar] [CrossRef]
- Zuo, T.; Liu, Q.; Zhang, F.; Lui, G.C.-Y.; Tso, E.Y.; Yeoh, Y.K.; Chen, Z.; Boon, S.S.; Chan, F.K.; Chan, P.K.; et al. Depicting SARS-CoV-2 Faecal Viral Activity in Association with Gut Microbiota Composition in Patients with COVID-19. Gut 2021, 70, 276–284. [Google Scholar] [CrossRef] [PubMed]
- Zhang, F.; Wan, Y.; Zuo, T.; Yeoh, Y.K.; Liu, Q.; Zhang, L.; Zhan, H.; Lu, W.; Xu, W.; Lui, G.C.Y.; et al. Prolonged Impairment of Short-Chain Fatty Acid and L-Isoleucine Biosynthesis in Gut Microbiome in Patients with COVID-19. Gastroenterology 2022, 162, 548–561. [Google Scholar] [CrossRef] [PubMed]
- Sun, Z.; Song, Z.-G.; Liu, C.; Tan, S.; Lin, S.; Zhu, J.; Dai, F.-H.; Gao, J.; She, J.-L.; Mei, Z.; et al. Gut Microbiome Alterations and Gut Barrier Dysfunction Are Associated with Host Immune Homeostasis in COVID-19 Patients. BMC Med. 2022, 20, 24. [Google Scholar] [CrossRef] [PubMed]
- Lv, L.; Jiang, H.; Chen, Y.; Gu, S.; Xia, J.; Zhang, H.; Lu, Y.; Yan, R.; Li, L. The Faecal Metabolome in COVID-19 Patients Is Altered and Associated with Clinical Features and Gut Microbes. Anal. Chim. Acta 2021, 1152, 338267. [Google Scholar] [CrossRef] [PubMed]
- Escarcega, R.D.; Honarpisheh, P.; Colpo, G.D.; Ahnstedt, H.W.; Couture, L.; Juneja, S.; Torres, G.; Ortiz, G.J.; Sollome, J.; Tabor, N.; et al. Sex Differences in Global Metabolomic Profiles of COVID-19 Patients. Cell Death Dis. 2022, 13, 461. [Google Scholar] [CrossRef]
- Yeoh, Y.K.; Zuo, T.; Lui, G.C.-Y.; Zhang, F.; Liu, Q.; Li, A.Y.; Chung, A.C.; Cheung, C.P.; Tso, E.Y.; Fung, K.S.; et al. Gut Microbiota Composition Reflects Disease Severity and Dysfunctional Immune Responses in Patients with COVID-19. Gut 2021, 70, 698–706. [Google Scholar] [CrossRef]
- Yao, Y.; Cai, X.; Fei, W.; Ye, Y.; Zhao, M.; Zheng, C. The Role of Short-Chain Fatty Acids in Immunity, Inflammation and Metabolism. Crit. Rev. Food Sci. Nutr. 2022, 62, 1–12. [Google Scholar] [CrossRef] [PubMed]
- Sun, R.; Xu, C.; Feng, B.; Gao, X.; Liu, Z. Critical Roles of Bile Acids in Regulating Intestinal Mucosal Immune Responses. Ther. Adv. Gastroenterol. 2021, 14, 17562848211018098. [Google Scholar] [CrossRef]
- Chen, J.; Vitetta, L. The Role of Butyrate in Attenuating Pathobiont-Induced Hyperinflammation. Immune Netw. 2020, 20, e15. [Google Scholar] [CrossRef]
- Arpaia, N.; Campbell, C.; Fan, X.; Dikiy, S.; van der Veeken, J.; deRoos, P.; Liu, H.; Cross, J.R.; Pfeffer, K.; Coffer, P.J.; et al. Metabolites Produced by Commensal Bacteria Promote Peripheral Regulatory T-Cell Generation. Nature 2013, 504, 451–455. [Google Scholar] [CrossRef] [PubMed]
- Chen, J.; Zhao, K.-N.; Vitetta, L. Effects of Intestinal Microbial–Elaborated Butyrate on Oncogenic Signaling Pathways. Nutrients 2019, 11, 1026. [Google Scholar] [CrossRef] [PubMed]
- Liu, P.; Wang, Y.; Yang, G.; Zhang, Q.; Meng, L.; Xin, Y.; Jiang, X. The Role of Short-Chain Fatty Acids in Intestinal Barrier Function, Inflammation, Oxidative Stress, and Colonic Carcinogenesis. Pharmacol. Res. 2021, 165, 105420. [Google Scholar] [CrossRef] [PubMed]
- Nagata, N.; Takeuchi, T.; Masuoka, H.; Aoki, R.; Ishikane, M.; Iwamoto, N.; Sugiyama, M.; Suda, W.; Nakanishi, Y.; Terada-Hirashima, J.; et al. Human Gut Microbiota and Its Metabolites Impact Immune Responses in COVID-19 and Its Complications. Gastroenterology 2022. [Google Scholar] [CrossRef] [PubMed]
- García-Lafuente, A.; Antolín, M.; Guarner, F.; Crespo, E.; Malagelada, J.-R. Modulation of Colonic Barrier Function by the Composition of the Commensal Flora in the Rat. Gut 2001, 48, 503–507. [Google Scholar] [CrossRef]
- Guo, M.; Tao, W.; Flavell, R.A.; Zhu, S. Potential Intestinal Infection and Faecal–Oral Transmission of SARS-CoV-2. Nat. Rev. Gastroenterol. Hepatol. 2021, 18, 269–283. [Google Scholar] [CrossRef]
- Edwinson, A.; Yang, L.; Chen, J.; Grover, M. Colonic Expression of Ace2, the SARS-CoV-2 Entry Receptor, Is Suppressed by Commensal Human Microbiota. Gut Microbes 2021, 13, 1984105. [Google Scholar] [CrossRef]
- Wang, H.; Wang, H.; Sun, Y.; Ren, Z.; Zhu, W.; Li, A.; Cui, G. Potential Associations Between Microbiome and COVID-19. Front. Med. 2021, 8, 785496. [Google Scholar] [CrossRef]
- Zhang, Y.; Yan, R.; Zhou, Q. ACE2, B0AT1, and SARS-CoV-2 Spike Protein: Structural and Functional Implications. Curr. Opin. Struct. Biol. 2022, 74, 102388. [Google Scholar] [CrossRef]
- Perlot, T.; Penninger, J.M. ACE2—from the Renin-Angiotensin System to Gut Microbiota and Malnutrition. Microbes Infect. 2013, 15, 866–873. [Google Scholar] [CrossRef]
- He, Y.; Wang, J.; Li, F.; Shi, Y. Main Clinical Features of COVID-19 and Potential Prognostic and Therapeutic Value of the Microbiota in SARS-CoV-2 Infections. Front. Microbiol. 2020, 11, 1302. [Google Scholar] [CrossRef]
- Chen, T.-H.; Hsu, M.-T.; Lee, M.-Y.; Chou, C.-K. Gastrointestinal Involvement in SARS-CoV-2 Infection. Viruses 2022, 14, 1188. [Google Scholar] [CrossRef] [PubMed]
- de Almeida, V.M.; Engel, D.F.; Ricci, M.F.; Cruz, C.S.; Lopes, I.S.; Alves, D.A.; Auriol, M.D.; Magalhães, J.; Zuccoli, G.S.; Smith, B.J.; et al. Gut Microbiota from Patients with Mild COVID-19 Cause Alterations in Mice That Resemble Post-COVID Syndrome. Researchsquare 2022. [Google Scholar] [CrossRef]
- Braniste, V.; Al-Asmakh, M.; Kowal, C.; Anuar, F.; Abbaspour, A.; Tóth, M.; Korecka, A.; Bakocevic, N.; Ng, L.G.; Kundu, P.; et al. The Gut Microbiota Influences Blood-Brain Barrier Permeability in Mice. Sci. Transl. Med. 2014, 6, 263ra158. [Google Scholar] [CrossRef] [PubMed]
- Zhao, J.; Bi, W.; Xiao, S.; Lan, X.; Cheng, X.; Zhang, J.; Lu, D.; Wei, W.; Wang, Y.; Li, H.; et al. Neuroinflammation Induced by Lipopolysaccharide Causes Cognitive Impairment in Mice. Sci. Rep. 2019, 9, 5790. [Google Scholar] [CrossRef] [PubMed]
- Manosso, L.M.; Arent, C.O.; Borba, L.A.; Ceretta, L.B.; Quevedo, J.; Réus, G.Z. Microbiota-Gut-Brain Communication in the SARS-CoV-2 Infection. Cells 2021, 10, 1993. [Google Scholar] [CrossRef]
- Xu, J.; Wu, Z.; Zhang, M.; Liu, S.; Zhou, L.; Yang, C.; Liu, C. The Role of the Gastrointestinal System in Neuroinvasion by SARS-CoV-2. Front Neurosci. 2021, 15, 694446. [Google Scholar] [CrossRef] [PubMed]
- Pilz, S.; Theiler-Schwetz, V.; Trummer, C.; Krause, R.; Ioannidis, J.P.A. SARS-CoV-2 Reinfections: Overview of Efficacy and Duration of Natural and Hybrid Immunity. Environ. Res. 2022, 209, 112911. [Google Scholar] [CrossRef] [PubMed]
- Cromer, D.; Juno, J.A.; Khoury, D.; Reynaldi, A.; Wheatley, A.K.; Kent, S.J.; Davenport, M.P. Prospects for Durable Immune Control of SARS-CoV-2 and Prevention of Reinfection. Nat. Rev. Immunol. 2021, 21, 395–404. [Google Scholar] [CrossRef]
- Goldblatt, D. SARS-CoV-2: From Herd Immunity to Hybrid Immunity. Nat. Rev. Immunol. 2022, 22, 333–334. [Google Scholar] [CrossRef]
- Grifoni, A.; Weiskopf, D.; Ramirez, S.I.; Mateus, J.; Dan, J.M.; Moderbacher, C.R.; Rawlings, S.A.; Sutherland, A.; Premkumar, L.; Jadi, R.S.; et al. Targets of T Cell Responses to SARS-CoV-2 Coronavirus in Humans with COVID-19 Disease and Unexposed Individuals. Cell 2020, 181, 1489–1501. [Google Scholar] [CrossRef]
- Qi, H.; Liu, B.; Wang, X.; Zhang, L. The Humoral Response and Antibodies against SARS-CoV-2 Infection. Nat. Immunol. 2022, 23, 1008–1020. [Google Scholar] [CrossRef]
- Castro Dopico, X.; Ols, S.; Loré, K.; Karlsson Hedestam, G.B. Immunity to SARS-CoV-2 Induced by Infection or Vaccination. J. Intern. Med. 2022, 291, 32–50. [Google Scholar] [CrossRef] [PubMed]
- Leidi, A.; Koegler, F.; Dumont, R.; Dubos, R.; Zaballa, M.-E.; Piumatti, G.; Coen, M.; Berner, A.; Darbellay Farhoumand, P.; Vetter, P.; et al. Risk of Reinfection After Seroconversion to Severe Acute Respiratory Syndrome Coronavirus 2 (SARS-CoV-2): A Population-Based Propensity-Score Matched Cohort Study. Clin. Infect. Dis. 2022, 74, 622–629. [Google Scholar] [CrossRef] [PubMed]
- Finch, E.; Lowe, R.; Fischinger, S.; Aubin, M.d.S.; Siddiqui, S.M.; Dayal, D.; Loesche, M.A.; Rhee, J.; Beger, S.; Hu, Y.; et al. SARS-CoV-2 Antibodies Protect against Reinfection for at Least 6 Months in a Multicentre Seroepidemiological Workplace Cohort. PLOS Biol. 2022, 20, e3001531. [Google Scholar] [CrossRef] [PubMed]
- Wang, Z.; Muecksch, F.; Schaefer-Babajew, D.; Finkin, S.; Viant, C.; Gaebler, C.; Hoffmann, H.-H.; Barnes, C.O.; Cipolla, M.; Ramos, V.; et al. Naturally Enhanced Neutralizing Breadth against SARS-CoV-2 One Year after Infection. Nature 2021, 595, 426–431. [Google Scholar] [CrossRef] [PubMed]
- Khoury, D.S.; Cromer, D.; Reynaldi, A.; Schlub, T.E.; Wheatley, A.K.; Juno, J.A.; Subbarao, K.; Kent, S.J.; Triccas, J.A.; Davenport, M.P. Neutralizing Antibody Levels Are Highly Predictive of Immune Protection from Symptomatic SARS-CoV-2 Infection. Nat. Med. 2021, 27, 1205–1211. [Google Scholar] [CrossRef]
- Zhang, Z.; Mateus, J.; Coelho, C.H.; Dan, J.M.; Moderbacher, C.R.; Gálvez, R.I.; Cortes, F.H.; Grifoni, A.; Tarke, A.; Chang, J.; et al. Humoral and Cellular Immune Memory to Four COVID-19 Vaccines. Cell 2022, 185, 2434–2451. [Google Scholar] [CrossRef]
- Bertoletti, A.; Le Bert, N.; Qui, M.; Tan, A.T. SARS-CoV-2-Specific T Cells in Infection and Vaccination. Cell. Mol. Immunol. 2021, 18, 2307–2312. [Google Scholar] [CrossRef] [PubMed]
- Epsi, N.J.; Richard, S.A.; Lindholm, D.A.; Mende, K.; Ganesan, A.; Huprikar, N.; Lalani, T.; Fries, A.C.; Maves, R.C.; Colombo, R.E.; et al. Understanding ‘Hybrid Immunity’: Comparison and Predictors of Humoral Immune Responses to SARS-CoV-2 Infection and COVID-19 Vaccines. Clin. Infect. Dis. 2022, ciac392. [Google Scholar] [CrossRef]
- Chen, X.; Chen, Z.; Azman, A.S.; Sun, R.; Lu, W.; Zheng, N.; Zhou, J.; Wu, Q.; Deng, X.; Zhao, Z.; et al. Neutralizing Antibodies Against Severe Acute Respiratory Syndrome Coronavirus 2 (SARS-CoV-2) Variants Induced by Natural Infection or Vaccination: A Systematic Review and Pooled Analysis. Clin. Infect. Dis. 2022, 74, 734–742. [Google Scholar] [CrossRef]
- Goldberg, Y.; Mandel, M.; Woodbridge, Y.; Fluss, R.; Novikov, I.; Yaari, R.; Ziv, A.; Freedman, L.; Huppert, A. Similarity of Protection Conferred by Previous SARS-CoV-2 Infection and by BNT162b2 Vaccine: A 3-Month Nationwide Experience from Israel. Am. J. Epidemiol. 2022, 191, kwac060. [Google Scholar] [CrossRef] [PubMed]
- Chemaitelly, H.; Ayoub, H.H.; AlMukdad, S.; Coyle, P.; Tang, P.; Yassine, H.M.; Al-Khatib, H.A.; Smatti, M.K.; Hasan, M.R.; Al-Kanaani, Z.; et al. Protection from Previous Natural Infection Compared with MRNA Vaccination against SARS-CoV-2 Infection and Severe COVID-19 in Qatar: A Retrospective Cohort Study. Lancet Microbe 2022, 3, e944–e955. [Google Scholar] [CrossRef] [PubMed]
- Bates, T.A.; McBride, S.K.; Leier, H.C.; Guzman, G.; Lyski, Z.L.; Schoen, D.; Winders, B.; Lee, J.-Y.; Lee, D.X.; Messer, W.B.; et al. Vaccination before or after SARS-CoV-2 Infection Leads to Robust Humoral Response and Antibodies That Effectively Neutralize Variants. Sci. Immunol. 2022, 7, eabn8014. [Google Scholar] [CrossRef] [PubMed]
- Qureshi, A.I.; Baskett, W.I.; Huang, W.; Lobanova, I.; Hasan Naqvi, S.; Shyu, C.-R. Reinfection with Severe Acute Respiratory Syndrome Coronavirus 2 (SARS-CoV-2) in Patients Undergoing Serial Laboratory Testing. Clin. Infect. Dis. 2022, 74, 294–300. [Google Scholar] [CrossRef] [PubMed]
- Abu-Raddad, L.J.; Chemaitelly, H.; Malek, J.A.; Ahmed, A.A.; Mohamoud, Y.A.; Younuskunju, S.; Ayoub, H.H.; Al Kanaani, Z.; Al Khal, A.; Al Kuwari, E.; et al. Assessment of the Risk of Severe Acute Respiratory Syndrome Coronavirus 2 (SARS-CoV-2) Reinfection in an Intense Reexposure Setting. Clin. Infect. Dis. 2021, 73, e1830–e1840. [Google Scholar] [CrossRef] [PubMed]
- Abu-Raddad, L.J.; Chemaitelly, H.; Coyle, P.; Malek, J.A.; Ahmed, A.A.; Mohamoud, Y.A.; Younuskunju, S.; Ayoub, H.H.; Al Kanaani, Z.; Al Kuwari, E.; et al. SARS-CoV-2 Antibody-Positivity Protects against Reinfection for at Least Seven Months with 95% Efficacy. EClinicalMedicine 2021, 35, 100861. [Google Scholar] [CrossRef] [PubMed]
- Abu-Raddad, L.J.; Chemaitelly, H.; Bertollini, R. Severity of SARS-CoV-2 Reinfections as Compared with Primary Infections. N. Engl. J. Med. 2021, 385, 2487–2489. [Google Scholar] [CrossRef]
- Mensah, A.A.; Lacy, J.; Stowe, J.; Seghezzo, G.; Sachdeva, R.; Simmons, R.; Bukasa, A.; O’Boyle, S.; Andrews, N.; Ramsay, M.; et al. Disease Severity during SARS-COV-2 Reinfection: A Nationwide Study. J. Infect. 2022, 84, 542–550. [Google Scholar] [CrossRef]
- Mohsin, M.; Mahmud, S. Omicron SARS-CoV-2 Variant of Concern: A Review on Its Transmissibility, Immune Evasion, Reinfection, and Severity. Medicine (Baltimore) 2022, 101, e29165. [Google Scholar] [CrossRef]
- Nguyen, N.N.; Houhamdi, L.; Hoang, V.T.; Delerce, J.; Delorme, L.; Colson, P.; Brouqui, P.; Fournier, P.-E.; Raoult, D.; Gautret, P. SARS-CoV-2 Reinfection and COVID-19 Severity. Emerg. Microbes Infect. 2022, 11, 894–901. [Google Scholar] [CrossRef]
- Pulliam, J.R.C.; van Schalkwyk, C.; Govender, N.; von Gottberg, A.; Cohen, C.; Groome, M.J.; Dushoff, J.; Mlisana, K.; Moultrie, H. Increased Risk of SARS-CoV-2 Reinfection Associated with Emergence of Omicron in South Africa. Science 2022, 376, eabn4947. [Google Scholar] [CrossRef]
- Altarawneh, H.N.; Chemaitelly, H.; Hasan, M.R.; Ayoub, H.H.; Qassim, S.; AlMukdad, S.; Coyle, P.; Yassine, H.M.; Al-Khatib, H.A.; Benslimane, F.M.; et al. Protection against the Omicron Variant from Previous SARS-CoV-2 Infection. N. Engl. J. Med. 2022, 386, 1288–1290. [Google Scholar] [CrossRef] [PubMed]
- Qassim, S.H.; Chemaitelly, H.; Ayoub, H.H.; AlMukdad, S.; Tang, P.; Hasan, M.R.; Yassine, H.M.; Al-Khatib, H.A.; Smatti, M.K.; Abdul-Rahim, H.F.; et al. Effects of BA.1/BA.2 Subvariant, Vaccination and Prior Infection on Infectiousness of SARS-CoV-2 Omicron Infections. J. Travel Med. 2022, 29, taac068. [Google Scholar] [CrossRef] [PubMed]
- Wolter, N.; Jassat, W.; Walaza, S.; Welch, R.; Moultrie, H.; Groome, M.J.; Amoako, D.G.; Everatt, J.; Bhiman, J.N.; Scheepers, C.; et al. Clinical Severity of SARS-CoV-2 Omicron BA.4 and BA.5 Lineages Compared to BA.1 and Delta in South Africa. Nat. Commun. 2022, 13, 5860. [Google Scholar] [CrossRef] [PubMed]
- Ferguson, N. Report 50: Hospitalisation Risk for Omicron Cases in England; Imperial College London: London, UK, 2021. [Google Scholar]
- Medić, S.; Anastassopoulou, C.; Lozanov-Crvenković, Z.; Vuković, V.; Dragnić, N.; Petrović, V.; Ristić, M.; Pustahija, T.; Gojković, Z.; Tsakris, A.; et al. Risk and Severity of SARS-CoV-2 Reinfections during 2020–2022 in Vojvodina, Serbia: A Population-Level Observational Study. Lancet Reg. Health—Eur. 2022, 20, 100453. [Google Scholar] [CrossRef] [PubMed]
- Murillo-Zamora, E.; Mendoza-Cano, O.; Delgado-Enciso, I.; Hernandez-Suarez, C.M. Predictors of Severe Symptomatic Laboratory-Confirmed SARS-CoV-2 Reinfection. Public Health 2021, 193, 113–115. [Google Scholar] [CrossRef]
- Islam, M.Z.; Riaz, B.K.; Akbar Ashrafi, S.A.; Farjana, S.; Efa, S.S.; Khan, M.A. Severity of COVID-19 Reinfection and Associated Risk Factors: Findings of a Cross-Sectional Study in Bangladesh; Infectious Diseases (except HIV/AIDS). medRxiv 2022. [Google Scholar] [CrossRef]
- Comba, I.Y.; Riestra Guiance, I.; Corsini Campioli, C.; Challener, D.; Sampathkumar, P.; Orenstein, R.; Gordon, J.; Bosch, W.; O’Horo, J.C. Clinical Characteristics and Outcomes of Patients with SARS-CoV-2 Reinfection. Mayo Clin. Proc. Innov. Qual. Outcomes 2022, 6, 361–372. [Google Scholar] [CrossRef]
- Guthmiller, J.J.; Stovicek, O.; Wang, J.; Changrob, S.; Li, L.; Halfmann, P.; Zheng, N.-Y.; Utset, H.; Stamper, C.T.; Dugan, H.L.; et al. SARS-CoV-2 Infection Severity Is Linked to Superior Humoral Immunity against the Spike. mBio 2021, 12, e02940-20. [Google Scholar] [CrossRef]
- Trinité, B.; Tarrés-Freixas, F.; Rodon, J.; Pradenas, E.; Urrea, V.; Marfil, S.; Rodríguez de la Concepción, M.L.; Ávila-Nieto, C.; Aguilar-Gurrieri, C.; Barajas, A.; et al. SARS-CoV-2 Infection Elicits a Rapid Neutralizing Antibody Response That Correlates with Disease Severity. Sci. Rep. 2021, 11, 2608. [Google Scholar] [CrossRef]
- Laing, E.D.; Epsi, N.J.; Richard, S.A.; Samuels, E.C.; Wang, W.; Vassell, R.; Ewing, D.F.; Herrup, R.; Sterling, S.L.; Lindholm, D.A.; et al. SARS-CoV-2 Antibodies Remain Detectable 12 Months after Infection and Antibody Magnitude Is Associated with Age and COVID-19 Severity. medRxiv 2021. [Google Scholar] [CrossRef]
- Legros, V.; Denolly, S.; Vogrig, M.; Boson, B.; Siret, E.; Rigaill, J.; Pillet, S.; Grattard, F.; Gonzalo, S.; Verhoeven, P.; et al. A Longitudinal Study of SARS-CoV-2-Infected Patients Reveals a High Correlation between Neutralizing Antibodies and COVID-19 Severity. Cell. Mol. Immunol. 2021, 18, 318–327. [Google Scholar] [CrossRef] [PubMed]
- Manuylov, V.; Burgasova, O.; Borisova, O.; Smetanina, S.; Vasina, D.; Grigoriev, I.; Kudryashova, A.; Semashko, M.; Cherepovich, B.; Kharchenko, O.; et al. Avidity of IgG to SARS-CoV-2 RBD as a Prognostic Factor for the Severity of COVID-19 Reinfection. Viruses 2022, 14, 617. [Google Scholar] [CrossRef] [PubMed]
- Wat, D. The Common Cold: A Review of the Literature. Eur. J. Intern. Med. 2004, 15, 79–88. [Google Scholar] [CrossRef] [PubMed]
- Kim, H.; Seiler, P.; Jones, J.C.; Ridout, G.; Camp, K.P.; Fabrizio, T.P.; Jeevan, T.; Miller, L.A.; Throm, R.E.; Ferrara, F.; et al. Antibody Responses to SARS-CoV-2 Antigens in Humans and Animals. Vaccines 2020, 8, 684. [Google Scholar] [CrossRef] [PubMed]
- Tan, C.C.S.; Owen, C.J.; Tham, C.Y.L.; Bertoletti, A.; van Dorp, L.; Balloux, F. Pre-Existing T Cell-Mediated Cross-Reactivity to SARS-CoV-2 Cannot Solely Be Explained by Prior Exposure to Endemic Human Coronaviruses. Infect. Genet. Evol. 2021, 95, 105075. [Google Scholar] [CrossRef] [PubMed]
- Mahajan, S.; Kode, V.; Bhojak, K.; Karunakaran, C.; Lee, K.; Manoharan, M.; Ramesh, A.; Hv, S.; Srivastava, A.; Sathian, R.; et al. Immunodominant T-Cell Epitopes from the SARS-CoV-2 Spike Antigen Reveal Robust Pre-Existing T-Cell Immunity in Unexposed Individuals. Sci. Rep. 2021, 11, 13164. [Google Scholar] [CrossRef]
- Braun, J.; Loyal, L.; Frentsch, M.; Wendisch, D.; Georg, P.; Kurth, F.; Hippenstiel, S.; Dingeldey, M.; Kruse, B.; Fauchere, F.; et al. SARS-CoV-2-Reactive T Cells in Healthy Donors and Patients with COVID-19. Nature 2020, 587, 270–274. [Google Scholar] [CrossRef]
- Le Bert, N.; Tan, A.T.; Kunasegaran, K.; Tham, C.Y.L.; Hafezi, M.; Chia, A.; Chng, M.H.Y.; Lin, M.; Tan, N.; Linster, M.; et al. SARS-CoV-2-Specific T Cell Immunity in Cases of COVID-19 and SARS, and Uninfected Controls. Nature 2020, 584, 457–462. [Google Scholar] [CrossRef]
- Becerra-Artiles, A.; Calvo-Calle, J.M.; Co, M.D.; Nanaware, P.P.; Cruz, J.; Weaver, G.C.; Lu, L.; Forconi, C.; Finberg, R.W.; Moormann, A.M.; et al. Broadly Recognized, Cross-Reactive SARS-CoV-2 CD4 T Cell Epitopes Are Highly Conserved across Human Coronaviruses and Presented by Common HLA Alleles. Cell Rep. 2022, 39, 110952. [Google Scholar] [CrossRef]
- Weiskopf, D. SARS-CoV-2 Specific and Cross-Reactive t Cell Responses. Top. Antivir. Med. 2021, 29, 4. [Google Scholar]
- Meckiff, B.J.; Ramírez-Suástegui, C.; Fajardo, V.; Chee, S.J.; Kusnadi, A.; Simon, H.; Grifoni, A.; Pelosi, E.; Weiskopf, D.; Sette, A.; et al. Single-Cell Transcriptomic Analysis of SARS-CoV-2 Reactive CD4+ T Cells. Cell 2020, 183, 1340–1353. [Google Scholar] [CrossRef] [PubMed]
- Ansari, A.; Arya, R.; Sachan, S.; Jha, S.N.; Kalia, A.; Lall, A.; Sette, A.; Grifoni, A.; Weiskopf, D.; Coshic, P.; et al. Immune Memory in Mild COVID-19 Patients and Unexposed Donors Reveals Persistent T Cell Responses After SARS-CoV-2 Infection. Front. Immunol. 2021, 12, 749. [Google Scholar] [CrossRef] [PubMed]
- Dykema, A.G.; Zhang, B.; Woldemeskel, B.A.; Garliss, C.C.; Cheung, L.S.; Choudhury, D.; Zhang, J.; Aparicio, L.; Bom, S.; Rashid, R.; et al. Functional Characterization of CD4+ T Cell Receptors Crossreactive for SARS-CoV-2 and Endemic Coronaviruses. J. Clin. Investig. 2021, 131, e146922. [Google Scholar] [CrossRef] [PubMed]
- Nelde, A.; Bilich, T.; Heitmann, J.S.; Maringer, Y.; Salih, H.R.; Roerden, M.; Lübke, M.; Bauer, J.; Rieth, J.; Wacker, M.; et al. SARS-CoV-2-Derived Peptides Define Heterologous and COVID-19-Induced T Cell Recognition. Nat. Immunol. 2021, 22, 74–85. [Google Scholar] [CrossRef]
- Schulien, I.; Kemming, J.; Oberhardt, V.; Wild, K.; Seidel, L.M.; Killmer, S.; Sagar; Daul, F.; Salvat Lago, M.; Decker, A.; et al. Characterization of Pre-Existing and Induced SARS-CoV-2-Specific CD8+ T Cells. Nat. Med. 2021, 27, 78–85. [Google Scholar] [CrossRef]
- Schmidt, K.G.; Nganou-Makamdop, K.; Tenbusch, M.; El Kenz, B.; Maier, C.; Lapuente, D.; Überla, K.; Spriewald, B.; Bergmann, S.; Harrer, E.G.; et al. SARS-CoV-2-Seronegative Subjects Target CTL Epitopes in the SARS-CoV-2 Nucleoprotein Cross-Reactive to Common Cold Coronaviruses. Front. Immunol. 2021, 12, 627568. [Google Scholar] [CrossRef]
- Peng, Y.; Mentzer, A.J.; Liu, G.; Yao, X.; Yin, Z.; Dong, D.; Dejnirattisai, W.; Rostron, T.; Supasa, P.; Liu, C.; et al. Broad and Strong Memory CD4+ and CD8+ T Cells Induced by SARS-CoV-2 in UK Convalescent Individuals Following COVID-19. Nat. Immunol. 2020, 21, 1336–1345. [Google Scholar] [CrossRef]
- Lineburg, K.E.; Grant, E.J.; Swaminathan, S.; Chatzileontiadou, D.S.M.; Szeto, C.; Sloane, H.; Panikkar, A.; Raju, J.; Crooks, P.; Rehan, S.; et al. CD8+ T Cells Specific for an Immunodominant SARS-CoV-2 Nucleocapsid Epitope Cross-React with Selective Seasonal Coronaviruses. Immunity 2021, 54, 1055–1065.e5. [Google Scholar] [CrossRef]
- Loyal, L.; Braun, J.; Henze, L.; Kruse, B.; Dingeldey, M.; Reimer, U.; Kern, F.; Schwarz, T.; Mangold, M.; Unger, C.; et al. Cross-Reactive CD4+ T Cells Enhance SARS-CoV-2 Immune Responses upon Infection and Vaccination. Science 2021, 374, eabh1823. [Google Scholar] [CrossRef]
- Lipsitch, M.; Grad, Y.H.; Sette, A.; Crotty, S. Cross-Reactive Memory T Cells and Herd Immunity to SARS-CoV-2. Nat. Rev. Immunol. 2020, 20, 709–713. [Google Scholar] [CrossRef]
- Moss, P. The T Cell Immune Response against SARS-CoV-2. Nat. Immunol. 2022, 23, 186–193. [Google Scholar] [CrossRef] [PubMed]
- Majdoubi, A.; Michalski, C.; O’Connell, S.E.; Dada, S.; Narpala, S.; Gelinas, J.; Mehta, D.; Cheung, C.; Winkler, D.F.H.; Basappa, M.; et al. A Majority of Uninfected Adults Show Preexisting Antibody Reactivity against SARS-CoV-2. JCI Insight 2021, 6, e146316. [Google Scholar] [CrossRef] [PubMed]
- Yuen, R.R.; Steiner, D.; Pihl, R.M.F.; Chavez, E.; Olson, A.; Smith, E.L.; Baird, L.A.; Korkmaz, F.; Urick, P.; Sagar, M.; et al. Novel ELISA Protocol Links Pre-Existing SARS-CoV-2 Reactive Antibodies with Endemic Coronavirus Immunity and Age and Reveals Improved Serologic Identification of Acute COVID-19 via Multi-Parameter Detection. Front. Immunol. 2021, 12, 614676. [Google Scholar] [CrossRef] [PubMed]
- Dalakas, M.C.; Bitzogli, K.; Alexopoulos, H. Anti-SARS-CoV-2 Antibodies within IVIg Preparations: Cross-Reactivities with Seasonal Coronaviruses, Natural Autoimmunity, and Therapeutic Implications. Front. Immunol. 2021, 12, 627285. [Google Scholar] [CrossRef]
- Guo, L.; Wang, Y.; Kang, L.; Hu, Y.; Wang, L.; Zhong, J.; Chen, H.; Ren, L.; Gu, X.; Wang, G.; et al. Cross-Reactive Antibody against Human Coronavirus OC43 Spike Protein Correlates with Disease Severity in COVID-19 Patients: A Retrospective Study. Emerg. Microbes Infect. 2021, 10, 664–676. [Google Scholar] [CrossRef] [PubMed]
- Kubota-Koketsu, R.; Terada, Y.; Yunoki, M.; Sasaki, T.; Nakayama, E.E.; Kamitani, W.; Shioda, T. Neutralizing and Binding Activities against SARS-CoV-1/2, MERS-CoV, and Human Coronaviruses 229E and OC43 by Normal Human Intravenous Immunoglobulin Derived from Healthy Donors in Japan. Transfusion 2021, 61, 356–360. [Google Scholar] [CrossRef] [PubMed]
- Anderson, D.E.; Tan, C.W.; Chia, W.N.; Young, B.E.; Linster, M.; Low, J.H.; Tan, Y.-J.; Chen, M.I.-C.; Smith, G.J.D.; Leo, Y.S.; et al. Lack of Cross-Neutralization by SARS Patient Sera towards SARS-CoV-2. Emerg. Microbes Infect. 2020, 9, 900–902. [Google Scholar] [CrossRef]
- García-Jiménez, Á.F.; Cáceres-Martell, Y.; Fernández-Soto, D.; Martínez Fleta, P.; Casasnovas, J.M.; Sánchez-Madrid, F.; Frade, J.M.R.; Valés-Gómez, M.; Reyburn, H.T. Cross-Reactive Cellular, but Not Humoral, Immunity Is Detected between OC43 and SARS-CoV-2 NPs in People Not Infected with SARS-CoV-2: Possible Role of CTFH Cells. J. Leukoc. Biol. 2022, 112, 339–346. [Google Scholar] [CrossRef]
- Song, G.; He, W.; Callaghan, S.; Anzanello, F.; Huang, D.; Ricketts, J.; Torres, J.L.; Beutler, N.; Peng, L.; Vargas, S.; et al. Cross-Reactive Serum and Memory B-Cell Responses to Spike Protein in SARS-CoV-2 and Endemic Coronavirus Infection. Nat. Commun. 2021, 12, 2938. [Google Scholar] [CrossRef]
- Hicks, J.; Klumpp-Thomas, C.; Kalish, H.; Shunmugavel, A.; Mehalko, J.; Denson, J.-P.; Snead, K.R.; Drew, M.; Corbett, K.S.; Graham, B.S.; et al. Serologic Cross-Reactivity of SARS-CoV-2 with Endemic and Seasonal Betacoronaviruses. J. Clin. Immunol. 2021, 41, 906–913. [Google Scholar] [CrossRef]
- Tso, F.Y.; Lidenge, S.J.; Peña, P.B.; Clegg, A.A.; Ngowi, J.R.; Mwaiselage, J.; Ngalamika, O.; Julius, P.; West, J.T.; Wood, C. High Prevalence of Pre-Existing Serological Cross-Reactivity against Severe Acute Respiratory Syndrome Coronavirus-2 (SARS-CoV-2) in Sub-Saharan Africa. Int. J. Infect. Dis. 2021, 102, 577–583. [Google Scholar] [CrossRef]
- Tajuelo, A.; López-Siles, M.; Más, V.; Pérez-Romero, P.; Aguado, J.M.; Briz, V.; McConnell, M.J.; Martín-Galiano, A.J.; López, D. Cross-Recognition of SARS-CoV-2 B-Cell Epitopes with Other Betacoronavirus Nucleoproteins. Int. J. Mol. Sci. 2022, 23, 2977. [Google Scholar] [CrossRef] [PubMed]
- Stoddard, C.I.; Sung, K.; Ojee, E.; Adhiambo, J.; Begnel, E.R.; Slyker, J.; Gantt, S.; Matsen, F.A.; Kinuthia, J.; Wamalwa, D.; et al. Distinct Antibody Responses to Endemic Coronaviruses Pre- and Post-SARS-CoV-2 Infection in Kenyan Infants and Mothers. Viruses 2022, 14, 1517. [Google Scholar] [CrossRef] [PubMed]
- Denninger, V.; Xu, C.K.; Meisl, G.; Morgunov, A.S.; Fiedler, S.; Ilsley, A.; Emmenegger, M.; Malik, A.Y.; Piziorska, M.A.; Schneider, M.M.; et al. Understanding the Role of Memory Re-Activation and Cross-Reactivity in the Defense against SARS-CoV-2. bioRxiv 2021. [Google Scholar] [CrossRef]
- Ng, K.W.; Faulkner, N.; Cornish, G.H.; Rosa, A.; Harvey, R.; Hussain, S.; Ulferts, R.; Earl, C.; Wrobel, A.G.; Benton, D.J.; et al. Preexisting and de Novo Humoral Immunity to SARS-CoV-2 in Humans. Science 2020, 370, 1339–1343. [Google Scholar] [CrossRef] [PubMed]
- Shrock, E.; Fujimura, E.; Kula, T.; Timms, R.T.; Lee, I.-H.; Leng, Y.; Robinson, M.L.; Sie, B.M.; Li, M.Z.; Chen, Y.; et al. Viral Epitope Profiling of COVID-19 Patients Reveals Cross-Reactivity and Correlates of Severity. Science 2020, 370, eabd4250. [Google Scholar] [CrossRef] [PubMed]
- Wang, J.; Young, B.E.; Li, D.; Seppo, A.; Zhou, Q.; Wiltse, A.; Nowak-Wegrzyn, A.; Murphy, K.; Widrick, K.; Diaz, N.; et al. Broad Cross-Reactive IgA and IgG against Human Coronaviruses in Milk Induced by COVID-19 Vaccination and Infection. Vaccines 2022, 10, 980. [Google Scholar] [CrossRef]
- Aydillo, T.; Rombauts, A.; Stadlbauer, D.; Aslam, S.; Abelenda-Alonso, G.; Escalera, A.; Amanat, F.; Jiang, K.; Krammer, F.; Carratala, J.; et al. Immunological Imprinting of the Antibody Response in COVID-19 Patients. Nat. Commun. 2021, 12, 3781. [Google Scholar] [CrossRef] [PubMed]
- Cohen, S.A.; Kellogg, C.; Equils, O. Neutralizing and Cross-Reacting Antibodies: Implications for Immunotherapy and SARS-CoV-2 Vaccine Development. Hum. Vaccines Immunother. 2021, 17, 84–87. [Google Scholar] [CrossRef]
- Aguilar-Bretones, M.; Westerhuis, B.M.; Raadsen, M.P.; Bruin, E.D.; Chandler, F.D.; Okba, N.M.A.; Haagmans, B.L.; Langerak, T.; Endeman, H.; van den Akker, J.P.C.; et al. Seasonal Coronavirus–Specific B Cells with Limited SARS-CoV-2 Cross-Reactivity Dominate the IgG Response in Severe COVID-19. J. Clin. Investig. 2021, 131, e150613. [Google Scholar] [CrossRef]
- Lin, C.-Y.; Wolf, J.; Brice, D.C.; Sun, Y.; Locke, M.; Cherry, S.; Castellaw, A.H.; Wehenkel, M.; Crawford, J.C.; Zarnitsyna, V.I.; et al. Pre-Existing Humoral Immunity to Human Common Cold Coronaviruses Negatively Impacts the Protective SARS-CoV-2 Antibody Response. Cell Host Microbe 2022, 30, 83–96.e4. [Google Scholar] [CrossRef]
- Anderson, E.M.; Goodwin, E.C.; Verma, A.; Arevalo, C.P.; Bolton, M.J.; Weirick, M.E.; Gouma, S.; McAllister, C.M.; Christensen, S.R.; Weaver, J.; et al. Seasonal Human Coronavirus Antibodies Are Boosted upon SARS-CoV-2 Infection but Not Associated with Protection. Cell 2021, 184, 1858–1864.e10. [Google Scholar] [CrossRef] [PubMed]
- Nguyen-Contant, P.; Embong, A.K.; Kanagaiah, P.; Chaves, F.A.; Yang, H.; Branche, A.R.; Topham, D.J.; Sangster, M.Y. S Protein-Reactive IgG and Memory B Cell Production after Human SARS-CoV-2 Infection Includes Broad Reactivity to the S2 Subunit. mBio 2020, 11, e01991-20. [Google Scholar] [CrossRef] [PubMed]
- Prévost, J.; Gasser, R.; Beaudoin-Bussières, G.; Richard, J.; Duerr, R.; Laumaea, A.; Anand, S.P.; Goyette, G.; Benlarbi, M.; Ding, S.; et al. Cross-Sectional Evaluation of Humoral Responses against SARS-CoV-2 Spike. Cell Rep. Med. 2020, 1, 100126. [Google Scholar] [CrossRef] [PubMed]
- Crowley, A.R.; Natarajan, H.; Hederman, A.P.; Bobak, C.A.; Weiner, J.A.; Wieland-Alter, W.; Lee, J.; Bloch, E.M.; Tobian, A.A.; Redd, A.D.; et al. Boosting of Cross-Reactive Antibodies to Endemic Coronaviruses by SARS-CoV-2 Infection but Not Vaccination with Stabilized Spike. eLife 2022, 11, e75228. [Google Scholar] [CrossRef]
- Becker, M.; Strengert, M.; Junker, D.; Kaiser, P.D.; Kerrinnes, T.; Traenkle, B.; Dinter, H.; Häring, J.; Ghozzi, S.; Zeck, A.; et al. Exploring beyond Clinical Routine SARS-CoV-2 Serology Using MultiCoV-Ab to Evaluate Endemic Coronavirus Cross-Reactivity. Nat. Commun. 2021, 12, 1152. [Google Scholar] [CrossRef] [PubMed]
- Ortega, N.; Ribes, M.; Vidal, M.; Rubio, R.; Aguilar, R.; Williams, S.; Barrios, D.; Alonso, S.; Hernández-Luis, P.; Mitchell, R.A.; et al. Seven-Month Kinetics of SARS-CoV-2 Antibodies and Role of Pre-Existing Antibodies to Human Coronaviruses. Nat. Commun. 2021, 12, 4740. [Google Scholar] [CrossRef]
- Geanes, E.S.; LeMaster, C.; Fraley, E.R.; Khanal, S.; McLennan, R.; Grundberg, E.; Selvarangan, R.; Bradley, T. Cross-Reactive Antibodies Elicited to Conserved Epitopes on SARS-CoV-2 Spike Protein after Infection and Vaccination. Sci. Rep. 2022, 12, 6496. [Google Scholar] [CrossRef]
- Dugas, M.; Grote-Westrick, T.; Vollenberg, R.; Lorentzen, E.; Brix, T.; Schmidt, H.; Tepasse, P.-R.; Kühn, J. Less Severe Course of COVID-19 Is Associated with Elevated Levels of Antibodies against Seasonal Human Coronaviruses OC43 and HKU1 (HCoV OC43, HCoV HKU1). Int. J. Infect. Dis. 2021, 105, 304–306. [Google Scholar] [CrossRef]
- Dugas, M.; Grote-Westrick, T.; Merle, U.; Fontenay, M.; Kremer, A.E.; Hanses, F.; Vollenberg, R.; Lorentzen, E.; Tiwari-Heckler, S.; Duchemin, J.; et al. Lack of Antibodies against Seasonal Coronavirus OC43 Nucleocapsid Protein Identifies Patients at Risk of Critical COVID-19. J. Clin. Virol. 2021, 139, 104847. [Google Scholar] [CrossRef]
- Henss, L.; Scholz, T.; von Rhein, C.; Wieters, I.; Borgans, F.; Eberhardt, F.J.; Zacharowski, K.; Ciesek, S.; Rohde, G.; Vehreschild, M.; et al. Analysis of Humoral Immune Responses in Patients with Severe Acute Respiratory Syndrome Coronavirus 2 Infection. J. Infect. Dis. 2021, 223, 56–61. [Google Scholar] [CrossRef]
- Sagar, M.; Reifler, K.; Rossi, M.; Miller, N.S.; Sinha, P.; White, L.F.; Mizgerd, J.P. Recent Endemic Coronavirus Infection Is Associated with Less-Severe COVID-19. J. Clin. Investig. 2021, 131, e143380. [Google Scholar] [CrossRef]
- Kaplonek, P.; Wang, C.; Bartsch, Y.; Fischinger, S.; Gorman, M.J.; Bowman, K.; Kang, J.; Dayal, D.; Martin, P.; Nowak, R.P.; et al. Early Cross-Coronavirus Reactive Signatures of Humoral Immunity against COVID-19. Sci. Immunol. 2021, 6, eabj2901. [Google Scholar] [CrossRef] [PubMed]
- Gombar, S.; Bergquist, T.; Pejaver, V.; Hammarlund, N.E.; Murugesan, K.; Mooney, S.; Shah, N.; Pinsky, B.A.; Banaei, N. SARS-CoV-2 Infection and COVID-19 Severity in Individuals with Prior Seasonal Coronavirus Infection. Diagn. Microbiol. Infect. Dis. 2021, 100, 115338. [Google Scholar] [CrossRef] [PubMed]
- Loos, C.; Atyeo, C.; Fischinger, S.; Burke, J.; Slein, M.D.; Streeck, H.; Lauffenburger, D.; Ryan, E.T.; Charles, R.C.; Alter, G. Evolution of Early SARS-CoV-2 and Cross-Coronavirus Immunity. mSphere 2020, 5, e00622-20. [Google Scholar] [CrossRef] [PubMed]
- Dispinseri, S.; Marzinotto, I.; Brigatti, C.; Pirillo, M.F.; Tolazzi, M.; Bazzigaluppi, E.; Canitano, A.; Borghi, M.; Gallinaro, A.; Caccia, R.; et al. Seasonal Betacoronavirus Antibodies’ Expansion Post-BNT161b2 Vaccination Associates with Reduced SARS-CoV-2 VoC Neutralization. J. Clin. Immunol. 2022, 42, 448–458. [Google Scholar] [CrossRef] [PubMed]
- Tetro, J.A. Is COVID-19 Receiving ADE from Other Coronaviruses? Microbes Infect. 2020, 22, 72–73. [Google Scholar] [CrossRef]
- Borrega, R.; Nelson, D.K.S.; Koval, A.P.; Bond, N.G.; Heinrich, M.L.; Rowland, M.M.; Lathigra, R.; Bush, D.J.; Aimukanova, I.; Phinney, W.N.; et al. Cross-Reactive Antibodies to SARS-CoV-2 and MERS-CoV in Pre-COVID-19 Blood Samples from Sierra Leoneans. Viruses 2021, 13, 2325. [Google Scholar] [CrossRef]
- Galipeau, Y.; Siragam, V.; Laroche, G.; Marion, E.; Greig, M.; McGuinty, M.; Booth, R.A.; Durocher, Y.; Cuperlovic-Culf, M.; Bennett, S.A.L.; et al. Relative Ratios of Human Seasonal Coronavirus Antibodies Predict the Efficiency of Cross-Neutralization of SARS-CoV-2 Spike Binding to ACE2. eBioMedicine 2021, 74, 103700. [Google Scholar] [CrossRef]
- Ou, X.; Liu, Y.; Lei, X.; Li, P.; Mi, D.; Ren, L.; Guo, L.; Guo, R.; Chen, T.; Hu, J.; et al. Characterization of Spike Glycoprotein of SARS-CoV-2 on Virus Entry and Its Immune Cross-Reactivity with SARS-CoV. Nat. Commun. 2020, 11, 1620. [Google Scholar] [CrossRef]
- Wells, D.A.; Cantoni, D.; Mayora-Neto, M.; Di Genova, C.; Sampson, A.; Ferrari, M.; Carnell, G.; Nadesalingam, A.; Smith, P.; Chan, A.; et al. Human Seasonal Coronavirus Neutralisation and COVID-19 Severity. J. Med. Virol. 2022, 94, 4820–4829. [Google Scholar] [CrossRef]
- Poston, D.; Weisblum, Y.; Wise, H.; Templeton, K.; Jenks, S.; Hatziioannou, T.; Bieniasz, P. Absence of Severe Acute Respiratory Syndrome Coronavirus 2 Neutralizing Activity in Prepandemic Sera from Individuals with Recent Seasonal Coronavirus Infection. Clin. Infect. Dis. 2021, 73, e1208–e1211. [Google Scholar] [CrossRef]
- Schwaiger, J.; Karbiener, M.; Aberham, C.; Farcet, M.R.; Kreil, T.R. No SARS-CoV-2 Neutralization by Intravenous Immunoglobulins Produced from Plasma Collected Before the 2020 Pandemic. J. Infect. Dis. 2020, 222, 1960–1964. [Google Scholar] [CrossRef]
- Sallis, R.; Young, D.R.; Tartof, S.Y.; Sallis, J.F.; Sall, J.; Li, Q.; Smith, G.N.; Cohen, D.A. Physical Inactivity Is Associated with a Higher Risk for Severe COVID-19 Outcomes: A Study in 48 440 Adult Patients. Br. J. Sports Med. 2021, 55, 1099–1105. [Google Scholar] [CrossRef] [PubMed]
- Lee, S.W.; Lee, J.; Moon, S.Y.; Jin, H.Y.; Yang, J.M.; Ogino, S.; Song, M.; Hong, S.H.; Ghayda, R.A.; Kronbichler, A.; et al. Physical Activity and the Risk of SARS-CoV-2 Infection, Severe COVID-19 Illness and COVID-19 Related Mortality in South Korea: A Nationwide Cohort Study. Br. J. Sports Med. 2022, 56, 901–912. [Google Scholar] [CrossRef]
- Salgado-Aranda, R.; Pérez-Castellano, N.; Núñez-Gil, I.; Orozco, A.J.; Torres-Esquivel, N.; Flores-Soler, J.; Chamaisse-Akari, A.; Mclnerney, A.; Vergara-Uzcategui, C.; Wang, L.; et al. Influence of Baseline Physical Activity as a Modifying Factor on COVID-19 Mortality: A Single-Center, Retrospective Study. Infect. Dis. Ther. 2021, 10, 801–814. [Google Scholar] [CrossRef] [PubMed]
- Cho, D.-H.; Lee, S.J.; Jae, S.Y.; Kim, W.J.; Ha, S.J.; Gwon, J.G.; Choi, J.; Kim, D.W.; Kim, J.Y. Physical Activity and the Risk of COVID-19 Infection and Mortality: A Nationwide Population-Based Case-Control Study. J. Clin. Med. 2021, 10, 1539. [Google Scholar] [CrossRef] [PubMed]
- Brawner, C.A.; Ehrman, J.K.; Bole, S.; Kerrigan, D.J.; Parikh, S.S.; Lewis, B.K.; Gindi, R.M.; Keteyian, C.; Abdul-Nour, K.; Keteyian, S.J. Inverse Relationship of Maximal Exercise Capacity to Hospitalization Secondary to Coronavirus Disease 2019. Mayo Clin. Proc. 2021, 96, 32–39. [Google Scholar] [CrossRef]
- Ekblom-Bak, E.; Väisänen, D.; Ekblom, B.; Blom, V.; Kallings, L.V.; Hemmingsson, E.; Andersson, G.; Wallin, P.; Salier Eriksson, J.; Holmlund, T.; et al. Cardiorespiratory Fitness and Lifestyle on Severe COVID-19 Risk in 279,455 Adults: A Case Control Study. Int. J. Behav. Nutr. Phys. Act. 2021, 18, 135. [Google Scholar] [CrossRef]
- Christensen, R.A.G.; Arneja, J.; Cyr, K.S.; Sturrock, S.L.; Brooks, J.D. The Association of Estimated Cardiorespiratory Fitness with COVID-19 Incidence and Mortality: A Cohort Study. PLoS ONE 2021, 16, e0250508. [Google Scholar] [CrossRef]
- af Geijerstam, A.; Mehlig, K.; Börjesson, M.; Robertson, J.; Nyberg, J.; Adiels, M.; Rosengren, A.; Åberg, M.; Lissner, L. Fitness, Strength and Severity of COVID-19: A Prospective Register Study of 1 559 187 Swedish Conscripts. BMJ Open 2021, 11, e051316. [Google Scholar] [CrossRef]
- Wong, C.-M.; Lai, H.-K.; Ou, C.-Q.; Ho, S.-Y.; Chan, K.-P.; Thach, T.-Q.; Yang, L.; Chau, Y.-K.; Lam, T.-H.; Hedley, A.J.; et al. Is Exercise Protective Against Influenza-Associated Mortality? PLoS ONE 2008, 3, e2108. [Google Scholar] [CrossRef]
- Matthews, C.E.; Ockene, I.S.; Freedson, P.S.; Rosal, M.C.; Merriam, P.A.; Hebert, J.R. Moderate to Vigorous Physical Activity and Risk of Upper-Respiratory Tract Infection. Med. Sci. Sports Exerc. 2002, 34, 1242–1248. [Google Scholar] [CrossRef] [PubMed]
- Olivo, C.R.; Miyaji, E.N.; Oliveira, M.L.S.; Almeida, F.M.; Lourenço, J.D.; Abreu, R.M.; Arantes, P.M.M.; Lopes, F.D.; Martins, M.A. Aerobic Exercise Attenuates Pulmonary Inflammation Induced by Streptococcus Pneumoniae. J. Appl. Physiol. 2014, 117, 998–1007. [Google Scholar] [CrossRef] [PubMed]
- Huh, J.Y. The Role of Exercise-Induced Myokines in Regulating Metabolism. Arch. Pharm. Res. 2018, 41, 14–29. [Google Scholar] [CrossRef]
- Bruunsgaard, H. Physical Activity and Modulation of Systemic Low-Level Inflammation. J. Leukoc. Biol. 2005, 78, 819–835. [Google Scholar] [CrossRef] [PubMed]
- Woods, J.A.; Vieira, V.J.; Keylock, K.T. Exercise, Inflammation, and Innate Immunity. Immunol. Allergy Clin. N. Am. 2009, 29, 381–393. [Google Scholar] [CrossRef]
- Spielmann, G.; McFarlin, B.K.; O’Connor, D.P.; Smith, P.J.W.; Pircher, H.; Simpson, R.J. Aerobic Fitness Is Associated with Lower Proportions of Senescent Blood T-Cells in Man. Brain. Behav. Immun. 2011, 25, 1521–1529. [Google Scholar] [CrossRef]
- Simpson, R.J.; Kunz, H.; Agha, N.; Graff, R. Chapter Fifteen—Exercise and the Regulation of Immune Functions. In Progress in Molecular Biology and Translational Science; Bouchard, C., Ed.; Molecular and Cellular Regulation of Adaptation to Exercise; Academic Press: Cambridge, MA, USA, 2015; Volume 135, pp. 355–380. [Google Scholar]
- Collie, S.; Saggers, R.T.; Bandini, R.; Steenkamp, L.; Champion, J.; Gray, G.; Bekker, L.-G.; Goga, A.; Garrett, N.; Patricios, J. Association between Regular Physical Activity and the Protective Effect of Vaccination against SARS-CoV-2 in a South African Case–Control Study. Br. J. Sports Med. 2022. [Google Scholar] [CrossRef]
- Scheffer, D.d.L.; Latini, A. Exercise-Induced Immune System Response: Anti-Inflammatory Status on Peripheral and Central Organs. Biochim. Biophys. Acta BBA—Mol. Basis Dis. 2020, 1866, 165823. [Google Scholar] [CrossRef]
- Mathot, E.; Liberman, K.; Cao Dinh, H.; Njemini, R.; Bautmans, I. Systematic Review on the Effects of Physical Exercise on Cellular Immunosenescence-Related Markers—An Update. Exp. Gerontol. 2021, 149, 111318. [Google Scholar] [CrossRef]
- Landi, F.; Marzetti, E.; Martone, A.M.; Bernabei, R.; Onder, G. Exercise as a Remedy for Sarcopenia. Curr. Opin. Clin. Nutr. Metab. Care 2014, 17, 25–31. [Google Scholar] [CrossRef]
- Gomes-Neto, M.; Saquetto, M.B.; Silva, C.M.; Carvalho, V.O.; Ribeiro, N.; Conceição, C.S. Effects of Respiratory Muscle Training on Respiratory Function, Respiratory Muscle Strength, and Exercise Tolerance in Patients Poststroke: A Systematic Review with Meta-Analysis. Arch. Phys. Med. Rehabil. 2016, 97, 1994–2001. [Google Scholar] [CrossRef] [PubMed]
- Stanford, K.I.; Goodyear, L.J. Exercise Regulation of Adipose Tissue. Adipocyte 2016, 5, 153–162. [Google Scholar] [CrossRef] [PubMed]
- Fiuza-Luces, C.; Santos-Lozano, A.; Joyner, M.; Carrera-Bastos, P.; Picazo, O.; Zugaza, J.L.; Izquierdo, M.; Ruilope, L.M.; Lucia, A. Exercise Benefits in Cardiovascular Disease: Beyond Attenuation of Traditional Risk Factors. Nat. Rev. Cardiol. 2018, 15, 731–743. [Google Scholar] [CrossRef] [PubMed]
- Bouassida, A.; Chamari, K.; Zaouali, M.; Feki, Y.; Zbidi, A.; Tabka, Z. Review on Leptin and Adiponectin Responses and Adaptations to Acute and Chronic Exercise. Br. J. Sports Med. 2010, 44, 620–630. [Google Scholar] [CrossRef] [PubMed]
- Filgueira, T.O.; Castoldi, A.; Santos, L.E.R.; de Amorim, G.J.; de Sousa Fernandes, M.S.; Anastácio, W.d.L.d.N.; Campos, E.Z.; Santos, T.M.; Souto, F.O. The Relevance of a Physical Active Lifestyle and Physical Fitness on Immune Defense: Mitigating Disease Burden, with Focus on COVID-19 Consequences. Front. Immunol. 2021, 12, 587146. [Google Scholar] [CrossRef] [PubMed]
- Rehm, J.; Samokhvalov, A.V.; Neuman, M.G.; Room, R.; Parry, C.; Lönnroth, K.; Patra, J.; Poznyak, V.; Popova, S. The Association between Alcohol Use, Alcohol Use Disorders and Tuberculosis (TB). A Systematic Review. BMC Public Health 2009, 9, 450. [Google Scholar] [CrossRef]
- Simou, E.; Leonardi-Bee, J.; Britton, J. The Effect of Alcohol Consumption on the Risk of ARDS: A Systematic Review and Meta-Analysis. Chest 2018, 154, 58–68. [Google Scholar] [CrossRef]
- Liu, M.; Gao, Y.; Shi, S.; Chen, Y.; Yang, K.; Tian, J. Drinking No-Links to the Severity of COVID-19: A Systematic Review and Meta-Analysis. J. Infect. 2020, 81, e126–e127. [Google Scholar] [CrossRef]
- Hamer, M.; Kivimäki, M.; Gale, C.R.; Batty, G.D. Lifestyle Risk Factors, Inflammatory Mechanisms, and COVID-19 Hospitalization: A Community-Based Cohort Study of 387,109 Adults in UK. Brain. Behav. Immun. 2020, 87, 184–187. [Google Scholar] [CrossRef]
- Dai, M.; Tao, L.; Chen, Z.; Tian, Z.; Guo, X.; Allen-Gipson, D.S.; Tan, R.; Li, R.; Chai, L.; Ai, F.; et al. Influence of Cigarettes and Alcohol on the Severity and Death of COVID-19: A Multicenter Retrospective Study in Wuhan, China. Front. Physiol. 2020, 11, 588553. [Google Scholar] [CrossRef]
- Bhalla, S.; Sharma, B.; Smith, D.; Boley, R.; McCluskey, C.; Ilyas, Y.; Afshar, M.; Balk, R.; Karnik, N.; Keshavarzian, A. Investigating Unhealthy Alcohol Use as an Independent Risk Factor for Increased COVID-19 Disease Severity: Observational Cross-Sectional Study. JMIR Public Health Surveill. 2021, 7, e33022. [Google Scholar] [CrossRef] [PubMed]
- Lassen, M.C.H.; Skaarup, K.G.; Sengeløv, M.; Iversen, K.; Ulrik, C.S.; Jensen, J.U.S.; Biering-Sørensen, T. Alcohol Consumption and the Risk of Acute Respiratory Distress Syndrome in COVID-19. Ann. Am. Thorac. Soc. 2021, 18, 1074–1076. [Google Scholar] [CrossRef] [PubMed]
- Alberca, R.W.; Rigato, P.O.; Ramos, Y.Á.L.; Teixeira, F.M.E.; Branco, A.C.C.; Fernandes, I.G.; Pietrobon, A.J.; Duarte, A.J.d.S.; Aoki, V.; Orfali, R.L.; et al. Clinical Characteristics and Survival Analysis in Frequent Alcohol Consumers with COVID-19. Front. Nutr. 2021, 8, 260. [Google Scholar] [CrossRef] [PubMed]
- Wendt, F.R.; De Lillo, A.; Pathak, G.A.; De Angelis, F.; Polimanti, R. Host Genetic Liability for Severe COVID-19 Associates with Alcohol Drinking Behavior and Diabetic Outcomes in Participants of European Descent. Front. Genet. 2021, 12, 765247. [Google Scholar] [CrossRef] [PubMed]
- Huang, W.; Zhou, H.; Hodgkinson, C.; Montero, A.; Goldman, D.; Chang, S.L. Network Meta-Analysis on the Mechanisms Underlying Alcohol Augmentation of COVID-19 Pathologies. Alcohol. Clin. Exp. Res. 2021, 45, 675–688. [Google Scholar] [CrossRef]
- Sureshchandra, S.; Raus, A.; Jankeel, A.; Ligh, B.J.K.; Walter, N.A.R.; Newman, N.; Grant, K.A.; Messaoudi, I. Dose-Dependent Effects of Chronic Alcohol Drinking on Peripheral Immune Responses. Sci. Rep. 2019, 9, 7847. [Google Scholar] [CrossRef]
- Yeligar, S.M.; Chen, M.M.; Kovacs, E.J.; Sisson, J.H.; Burnham, E.L.; Brown, L.A.S. Alcohol and Lung Injury and Immunity. Alcohol 2016, 55, 51–59. [Google Scholar] [CrossRef]
- Ben-Eliyahu, S.; Page, G.G.; Yirmiya, R.; Taylor, A.N. Acute Alcohol Intoxication Suppresses Natural Killer Cell Activity and Promotes Tumor Metastasis. Nat. Med. 1996, 2, 457–460. [Google Scholar] [CrossRef]
- Sisson, J.H. Alcohol and Airways Function in Health and Disease. Alcohol 2007, 41, 293–307. [Google Scholar] [CrossRef]
- Arcavi, L.; Benowitz, N.L. Cigarette Smoking and Infection. Arch. Intern. Med. 2004, 164, 2206–2216. [Google Scholar] [CrossRef]
- Lacedonia, D.; Scioscia, G.; Santomasi, C.; Fuso, P.; Carpagnano, G.E.; Portacci, A.; Mastroianni, F.; Larizza, G.; Sabato, E.; Profilo, E.; et al. Impact of Smoking, COPD and Comorbidities on the Mortality of COVID-19 Patients. Sci. Rep. 2021, 11, 19251. [Google Scholar] [CrossRef] [PubMed]
- Rao, S.; Baranova, A.; Cao, H.; Chen, J.; Zhang, X.; Zhang, F. Genetic Mechanisms of COVID-19 and Its Association with Smoking and Alcohol Consumption. Brief. Bioinform. 2021, 22, bbab284. [Google Scholar] [CrossRef] [PubMed]
- Clift, A.K.; von Ende, A.; Tan, P.S.; Sallis, H.M.; Lindson, N.; Coupland, C.A.C.; Munafò, M.R.; Aveyard, P.; Hippisley-Cox, J.; Hopewell, J.C. Smoking and COVID-19 Outcomes: An Observational and Mendelian Randomisation Study Using the UK Biobank Cohort. Thorax 2022, 77, 65–73. [Google Scholar] [CrossRef]
- Young-Wolff, K.C.; Slama, N.; Alexeeff, S.E.; Sakoda, L.C.; Fogelberg, R.; Myers, L.C.; Campbell, C.I.; Adams, A.S.; Prochaska, J.J. Tobacco Smoking and Risk of SARS-CoV-2 Infection and Disease Severity among Adults in an Integrated Healthcare System in California. Nicotine Tob. Res. 2023, 25, 211–220. [Google Scholar] [CrossRef]
- Puebla Neira, D.; Watts, A.; Seashore, J.; Polychronopoulou, E.; Kuo, Y.-F.; Sharma, G. Smoking and Risk of COVID-19 Hospitalization. Respir. Med. 2021, 182, 106414. [Google Scholar] [CrossRef] [PubMed]
- Simons, D.; Shahab, L.; Brown, J.; Perski, O. The Association of Smoking Status with SARS-CoV-2 Infection, Hospitalization and Mortality from COVID-19: A Living Rapid Evidence Review with Bayesian Meta-Analyses (Version 7). Addiction 2021, 116, 1319–1368. [Google Scholar] [CrossRef] [PubMed]
- Piasecki, T.M.; Smith, S.S.; Baker, T.B.; Slutske, W.S.; Adsit, R.T.; Bolt, D.M.; Conner, K.L.; Bernstein, S.L.; Eng, O.D.; Lazuk, D.; et al. Smoking Status, Nicotine Medication, Vaccination, and COVID-19 Hospital Outcomes: Findings from the COVID EHR Cohort at the University of Wisconsin (CEC-UW) Study. Nicotine Tob. Res. 2022, ntac201. [Google Scholar] [CrossRef]
- Hou, H.; Li, Y.; Zhang, P.; Wu, J.; Shi, L.; Xu, J.; Diao, J.; Wang, Y.; Yang, H. Smoking Is Independently Associated with an Increased Risk for COVID-19 Mortality: A Systematic Review and Meta-Analysis Based on Adjusted Effect Estimates. Nicotine Tob. Res. 2021, 23, 1947–1951. [Google Scholar] [CrossRef]
- Reddy, R.K.; Charles, W.N.; Sklavounos, A.; Dutt, A.; Seed, P.T.; Khajuria, A. The Effect of Smoking on COVID-19 Severity: A Systematic Review and Meta-Analysis. J. Med. Virol. 2021, 93, 1045–1056. [Google Scholar] [CrossRef]
- Patanavanich, R.; Glantz, S.A. Smoking Is Associated with Worse Outcomes of COVID-19 Particularly among Younger Adults: A Systematic Review and Meta-Analysis. BMC Public Health 2021, 21, 1554. [Google Scholar] [CrossRef] [PubMed]
- Poudel, R.; Daniels, L.B.; DeFilippis, A.P.; Hamburg, N.M.; Khan, Y.; Keith, R.J.; Kumar, R.S.; Strokes, A.C.; Robertson, R.M.; Bhatnagar, A. Smoking Is Associated with Increased Risk of Cardiovascular Events, Disease Severity, and Mortality among Patients Hospitalized for SARS-CoV-2 Infections. PLoS ONE 2022, 17, e0270763. [Google Scholar] [CrossRef] [PubMed]
- Farsalinos, K.; Barbouni, A.; Poulas, K.; Polosa, R.; Caponnetto, P.; Niaura, R. Current Smoking, Former Smoking, and Adverse Outcome among Hospitalized COVID-19 Patients: A Systematic Review and Meta-Analysis. Ther. Adv. Chronic Dis. 2020, 11, 2040622320935765. [Google Scholar] [CrossRef] [PubMed]
- Umnuaypornlert, A.; Kanchanasurakit, S.; Lucero-Prisno, D.E.I.; Saokaew, S. Smoking and Risk of Negative Outcomes among COVID-19 Patients: A Systematic Review and Meta-Analysis. Tob. Induc. Dis. 2021, 19, 9. [Google Scholar] [CrossRef] [PubMed]
- Saadatian-Elahi, M.; Amour, S.; Elias, C.; Henaff, L.; Dananché, C.; Vanhems, P. Tobacco Smoking and Severity of COVID-19: Experience from a Hospital-Based Prospective Cohort Study in Lyon, France. J. Med. Virol. 2021, 93, 6822–6827. [Google Scholar] [CrossRef]
- Benowitz, N.L.; Goniewicz, M.L.; Halpern-Felsher, B.; Krishnan-Sarin, S.; Ling, P.M.; O’Connor, R.J.; Pentz, M.A.; Robertson, R.M.; Bhatnagar, A. Tobacco Product Use and the Risks of SARS-CoV-2 Infection and COVID-19: Current Understanding and Recommendations for Future Research. Lancet Respir. Med. 2022, 10, 900–915. [Google Scholar] [CrossRef]
- Assad, N.A.; Kapoor, V.; Sood, A. Biomass Smoke Exposure and Chronic Lung Disease. Curr. Opin. Pulm. Med. 2016, 22, 150–157. [Google Scholar] [CrossRef]
- Kondo, T.; Nakano, Y.; Adachi, S.; Murohara, T. Effects of Tobacco Smoking on Cardiovascular Disease. Circ. J. 2019, 83, 1980–1985. [Google Scholar] [CrossRef]
- Zhu, P.; Pan, X.-F.; Sheng, L.; Chen, H.; Pan, A. Cigarette Smoking, Diabetes, and Diabetes Complications: Call for Urgent Action. Curr. Diab. Rep. 2017, 17, 78. [Google Scholar] [CrossRef]
- Habesoglu, M.; Demir, K.; Yumusakhuylu, A.C.; Sahin Yilmaz, A.; Oysu, C. Does Passive Smoking Have an Effect on Nasal Mucociliary Clearance? Otolaryngol. Neck Surg. 2012, 147, 152–156. [Google Scholar] [CrossRef]
- Morrison, D.; Rahman, I.; Lannan, S.; MacNEE, W. Epithelial Permeability, Inflammation, and Oxidant Stress in the Air Spaces of Smokers. Am. J. Respir. Crit. Care Med. 1999, 159, 473–479. [Google Scholar] [CrossRef]
- Johnson, J.D.; Houchens, D.P.; Kluwe, W.M.; Craig, D.K.; Fisher, G.L. Effects of Mainstream and Environmental Tobacco Smoke on the Immune System in Animals and Humans: A Review. Crit. Rev. Toxicol. 1990, 20, 369–395. [Google Scholar] [CrossRef] [PubMed]
- Purkayastha, A.; Sen, C.; Garcia, G.; Langerman, J.; Shia, D.W.; Meneses, L.K.; Vijayaraj, P.; Durra, A.; Koloff, C.R.; Freund, D.R.; et al. Direct Exposure to SARS-CoV-2 and Cigarette Smoke Increases Infection Severity and Alters the Stem Cell-Derived Airway Repair Response. Cell Stem Cell 2020, 27, 869–875. [Google Scholar] [CrossRef] [PubMed]
- Burke, A.; FitzGerald, G.A. Oxidative Stress and Smoking-Induced Vascular Injury. Prog. Cardiovasc. Dis. 2003, 46, 79–90. [Google Scholar] [CrossRef] [PubMed]
- Gonçalves, R.B.; Coletta, R.D.; Silvério, K.G.; Benevides, L.; Casati, M.Z.; da Silva, J.S.; Nociti, F.H. Impact of Smoking on Inflammation: Overview of Molecular Mechanisms. Inflamm. Res. 2011, 60, 409–424. [Google Scholar] [CrossRef] [PubMed]
- Liu, H.; Xin, J.; Cai, S.; Jiang, X. Mendelian Randomization Analysis Provides Causality of Smoking on the Expression of ACE2, a Putative SARS-CoV-2 Receptor. eLife 2021, 10, e64188. [Google Scholar] [CrossRef] [PubMed]
- Smith, J.C.; Sausville, E.L.; Girish, V.; Yuan, M.L.; Vasudevan, A.; John, K.M.; Sheltzer, J.M. Cigarette Smoke Exposure and Inflammatory Signaling Increase the Expression of the SARS-CoV-2 Receptor ACE2 in the Respiratory Tract. Dev. Cell 2020, 53, 514–529.e3. [Google Scholar] [CrossRef]
- Akerström, S.; Mousavi-Jazi, M.; Klingström, J.; Leijon, M.; Lundkvist, A.; Mirazimi, A. Nitric Oxide Inhibits the Replication Cycle of Severe Acute Respiratory Syndrome Coronavirus. J. Virol. 2005, 79, 1966–1969. [Google Scholar] [CrossRef] [PubMed]
- Wang, H.; Yu, M.; Ochani, M.; Amella, C.A.; Tanovic, M.; Susarla, S.; Li, J.H.; Wang, H.; Yang, H.; Ulloa, L.; et al. Nicotinic Acetylcholine Receptor Alpha7 Subunit Is an Essential Regulator of Inflammation. Nature 2003, 421, 384–388. [Google Scholar] [CrossRef]
- Pavlov, V.A.; Tracey, K.J. The Cholinergic Anti-Inflammatory Pathway. Brain. Behav. Immun. 2005, 19, 493–499. [Google Scholar] [CrossRef]
- Wells, J.C.; Sawaya, A.L.; Wibaek, R.; Mwangome, M.; Poullas, M.S.; Yajnik, C.S.; Demaio, A. The Double Burden of Malnutrition: Aetiological Pathways and Consequences for Health. The Lancet 2020, 395, 75–88. [Google Scholar] [CrossRef]
- Foolchand, A.; Ghazi, T.; Chuturgoon, A.A. Malnutrition and Dietary Habits Alter the Immune System Which May Consequently Influence SARS-CoV-2 Virulence: A Review. Int. J. Mol. Sci. 2022, 23, 2654. [Google Scholar] [CrossRef] [PubMed]
- Söderström, L.; Rosenblad, A.; Adolfsson, E.T.; Bergkvist, L. Malnutrition Is Associated with Increased Mortality in Older Adults Regardless of the Cause of Death. Br. J. Nutr. 2017, 117, 532–540. [Google Scholar] [CrossRef] [PubMed]
- Ponce, J.; Anzalone, A.J.; Bailey, K.; Sayles, H.; Timmerman, M.; Jackson, M.; McClay, J.; Hanson, C.; National COVID Cohort Collaborative (N3C) Consortium. Impact of Malnutrition on Clinical Outcomes in Patients Diagnosed with COVID-19. J. Parenter. Enter. Nutr. 2022, 46, 1797–1807. [Google Scholar] [CrossRef]
- Kurtz, A.; Grant, K.; Marano, R.; Arrieta, A.; Grant, K.; Feaster, W.; Steele, C.; Ehwerhemuepha, L. Long-Term Effects of Malnutrition on Severity of COVID-19. Sci. Rep. 2021, 11, 14974. [Google Scholar] [CrossRef] [PubMed]
- Zhao, X.; Li, Y.; Ge, Y.; Shi, Y.; Lv, P.; Zhang, J.; Fu, G.; Zhou, Y.; Jiang, K.; Lin, N.; et al. Evaluation of Nutrition Risk and Its Association with Mortality Risk in Severely and Critically Ill COVID-19 Patients. J. Parenter. Enter. Nutr. 2021, 45, 32–42. [Google Scholar] [CrossRef]
- Yu, Y.; Ye, J.; Chen, M.; Jiang, C.; Lin, W.; Lu, Y.; Ye, H.; Li, Y.; Wang, Y.; Liao, Q.; et al. Malnutrition Prolongs the Hospitalization of Patients with COVID-19 Infection: A Clinical Epidemiological Analysis. J. Nutr. Health Aging 2021, 25, 369–373. [Google Scholar] [CrossRef]
- Yue, Y.; Ma, W.; Accorsi, E.K.; Ding, M.; Hu, F.; Willett, W.C.; Chan, A.T.; Sun, Q.; Edwards, J.R.; Smith-Warner, S.A.; et al. Long-Term Diet and Risk of SARS-CoV-2 Infection and Coronavirus Disease 2019 (COVID-19) Severity. Am. J. Clin. Nutr. 2022, 116, 1672–1681. [Google Scholar] [CrossRef]
- Hou, Y.-C.; Su, W.-L.; Chao, Y.-C. COVID-19 Illness Severity in the Elderly in Relation to Vegetarian and Non-Vegetarian Diets: A Single-Center Experience. Front. Nutr. 2022, 9, 837458. [Google Scholar] [CrossRef]
- Kim, H.; Rebholz, C.M.; Hegde, S.; LaFiura, C.; Raghavan, M.; Lloyd, J.F.; Cheng, S.; Seidelmann, S.B. Plant-Based Diets, Pescatarian Diets and COVID-19 Severity: A Population-Based Case–Control Study in Six Countries. BMJ Nutr. Prev. Health 2021, 4, 257–266. [Google Scholar] [CrossRef]
- Merino, J.; Joshi, A.D.; Nguyen, L.H.; Leeming, E.R.; Mazidi, M.; Drew, D.A.; Gibson, R.; Graham, M.S.; Lo, C.-H.; Capdevila, J.; et al. Diet Quality and Risk and Severity of COVID-19: A Prospective Cohort Study. Gut 2021, 70, 2096–2104. [Google Scholar] [CrossRef]
- Trujillo-Mayol, I.; Guerra-Valle, M.; Casas-Forero, N.; Sobral, M.M.C.; Viegas, O.; Alarcón-Enos, J.; Ferreira, I.M.; Pinho, O. Western Dietary Pattern Antioxidant Intakes and Oxidative Stress: Importance During the SARS-CoV-2/COVID-19 Pandemic. Adv. Nutr. 2021, 12, 670–681. [Google Scholar] [CrossRef]
- Chrysohoou, C.; Panagiotakos, D.B.; Pitsavos, C.; Das, U.N.; Stefanadis, C. Adherence to the Mediterranean Diet Attenuates Inflammation and Coagulation Process in Healthy Adults. J. Am. Coll. Cardiol. 2004, 44, 152–158. [Google Scholar] [CrossRef] [PubMed]
- Ponzo, V.; Pellegrini, M.; D’Eusebio, C.; Bioletto, F.; Goitre, I.; Buscemi, S.; Frea, S.; Ghigo, E.; Bo, S. Mediterranean Diet and SARS-COV-2 Infection: Is There Any Association? A Proof-of-Concept Study. Nutrients 2021, 13, 1721. [Google Scholar] [CrossRef] [PubMed]
- Perez-Araluce, R.; Martinez-Gonzalez, M.A.; Fernández-Lázaro, C.I.; Bes-Rastrollo, M.; Gea, A.; Carlos, S. Mediterranean Diet and the Risk of COVID-19 in the ‘Seguimiento Universidad de Navarra’ Cohort. Clin. Nutr. 2021, 41, 3061–3068. [Google Scholar] [CrossRef] [PubMed]
- Horne, B.D.; Muhlestein, J.B.; May, H.T.; Le, V.T.; Bair, T.L.; Knowlton, K.U.; Anderson, J.L.; INSPIRE Registry Investigators. Association of Periodic Fasting with Lower Severity of COVID-19 Outcomes in the SARS-CoV-2 Pre-Vaccine Era: An Observational Cohort from the INSPIRE Registry. medRxiv 2022. [Google Scholar] [CrossRef]
- Sukkar, S.G.; Cogorno, L.; Pisciotta, L.; Pasta, A.; Vena, A.; Gradaschi, R.; Dentone, C.; Guiddo, E.; Martino, E.; Beltramini, S.; et al. Clinical Efficacy of Eucaloric Ketogenic Nutrition in the COVID-19 Cytokine Storm: A Retrospective Analysis of Mortality and Intensive Care Unit Admission. Nutrition 2021, 89, 111236. [Google Scholar] [CrossRef]
- Gangitano, E.; Tozzi, R.; Gandini, O.; Watanabe, M.; Basciani, S.; Mariani, S.; Lenzi, A.; Gnessi, L.; Lubrano, C. Ketogenic Diet as a Preventive and Supportive Care for COVID-19 Patients. Nutrients 2021, 13, 1004. [Google Scholar] [CrossRef]
- Ling, V.; Zabetakis, I. The Role of an Anti-Inflammatory Diet in Conjunction to COVID-19. Diseases 2021, 9, 76. [Google Scholar] [CrossRef]
- Julkunen, H.; Cichońska, A.; Slagboom, P.E.; Würtz, P. Nightingale Health UK Biobank Initiative Metabolic Biomarker Profiling for Identification of Susceptibility to Severe Pneumonia and COVID-19 in the General Population. eLife 2021, 10, e63033. [Google Scholar] [CrossRef]
- Zapata, B.R.; Müller, J.M.; Vásquez, J.E.; Ravera, F.; Lago, G.; Cañón, E.; Castañeda, D.; Pradenas, M.; Ramírez-Santana, M. Omega-3 Index and Clinical Outcomes of Severe COVID-19: Preliminary Results of a Cross-Sectional Study. Int. J. Environ. Res. Public. Health 2021, 18, 7722. [Google Scholar] [CrossRef]
- Mazidimoradi, A.; Alemzadeh, E.; Alemzadeh, E.; Salehiniya, H. The Effect of Polyunsaturated Fatty Acids on the Severity and Mortality of COVID Patients: A Systematic Review. Life Sci. 2022, 299, 120489. [Google Scholar] [CrossRef]
- Bejan, C.A.; Cahill, K.N.; Staso, P.J.; Choi, L.; Peterson, J.F.; Phillips, E.J. DrugWAS: Drug-Wide Association Studies for COVID-19 Drug Repurposing. Clin. Pharmacol. Ther. 2021, 110, 1537–1546. [Google Scholar] [CrossRef]
- Sedighiyan, M.; Abdollahi, H.; Karimi, E.; Badeli, M.; Erfanian, R.; Raeesi, S.; Hashemi, R.; Vahabi, Z.; Asanjarani, B.; Mansouri, F.; et al. Omega-3 Polyunsaturated Fatty Acids Supplementation Improve Clinical Symptoms in Patients with COVID-19: A Randomised Clinical Trial. Int. J. Clin. Pract. 2021, 75, e14854. [Google Scholar] [CrossRef] [PubMed]
- Langlois, P.L.; D’Aragon, F.; Hardy, G.; Manzanares, W. Omega-3 Polyunsaturated Fatty Acids in Critically Ill Patients with Acute Respiratory Distress Syndrome: A Systematic Review and Meta-Analysis. Nutrition 2019, 61, 84–92. [Google Scholar] [CrossRef] [PubMed]
- Serhan, C.N.; Chiang, N. Resolution Phase Lipid Mediators of Inflammation: Agonists of Resolution. Curr. Opin. Pharmacol. 2013, 13, 632–640. [Google Scholar] [CrossRef] [PubMed]
- Serhan, C.N. Pro-Resolving Lipid Mediators Are Leads for Resolution Physiology. Nature 2014, 510, 92–101. [Google Scholar] [CrossRef] [PubMed]
- Zaloga, G.P. Narrative Review of N-3 Polyunsaturated Fatty Acid Supplementation upon Immune Functions, Resolution Molecules and Lipid Peroxidation. Nutrients 2021, 13, 662. [Google Scholar] [CrossRef]
- Schwarz, B.; Sharma, L.; Roberts, L.; Peng, X.; Bermejo, S.; Leighton, I.; Casanovas-Massana, A.; Minasyan, M.; Farhadian, S.; Ko, A.I.; et al. Cutting Edge: Severe SARS-CoV-2 Infection in Humans Is Defined by a Shift in the Serum Lipidome, Resulting in Dysregulation of Eicosanoid Immune Mediators. J. Immunol. 2021, 206, 329–334. [Google Scholar] [CrossRef]
- Goc, A.; Niedzwiecki, A.; Rath, M. Polyunsaturated ω-3 Fatty Acids Inhibit ACE2-Controlled SARS-CoV-2 Binding and Cellular Entry. Sci. Rep. 2021, 11, 5207. [Google Scholar] [CrossRef]
- Brockman-Schneider, R.A.; Pickles, R.J.; Gern, J.E. Effects of Vitamin D on Airway Epithelial Cell Morphology and Rhinovirus Replication. PLoS ONE 2014, 9, e86755. [Google Scholar] [CrossRef]
- Hansdottir, S.; Monick, M.M.; Lovan, N.; Powers, L.; Gerke, A.; Hunninghake, G.W. Vitamin D Decreases Respiratory Syncytial Virus Induction of NF-ΚB–Linked Chemokines and Cytokines in Airway Epithelium While Maintaining the Antiviral State. J. Immunol. 2010, 184, 965–974. [Google Scholar] [CrossRef]
- Cannell, J.J.; Vieth, R.; Umhau, J.C.; Holick, M.F.; Grant, W.B.; Madronich, S.; Garland, C.F.; Giovannucci, E. Epidemic Influenza and Vitamin D. Epidemiol. Infect. 2006, 134, 1129–1140. [Google Scholar] [CrossRef]
- Martineau, A.R.; Cantorna, M.T. Vitamin D for COVID-19: Where Are We Now? Nat. Rev. Immunol. 2022, 22, 529–530. [Google Scholar] [CrossRef] [PubMed]
- Eggersdorfer, M.; Berger, M.M.; Calder, P.C.; Gombart, A.F.; Ho, E.; Laviano, A.; Meydani, S.N. Perspective: Role of Micronutrients and Omega-3 Long-Chain Polyunsaturated Fatty Acids for Immune Outcomes of Relevance to Infections in Older Adults—A Narrative Review and Call for Action. Adv. Nutr. 2022, 13, 1415–1430. [Google Scholar] [CrossRef] [PubMed]
- Orchard, L.; Baldry, M.; Nasim-Mohi, M.; Monck, C.; Saeed, K.; Grocott, M.P.W.; Ahilanandan, D. Vitamin-D Levels and Intensive Care Unit Outcomes of a Cohort of Critically Ill COVID-19 Patients. Clin. Chem. Lab. Med. CCLM 2021, 59, 1155–1163. [Google Scholar] [CrossRef] [PubMed]
- Campi, I.; Gennari, L.; Merlotti, D.; Mingiano, C.; Frosali, A.; Giovanelli, L.; Torlasco, C.; Pengo, M.F.; Heilbron, F.; Soranna, D.; et al. Vitamin D and COVID-19 Severity and Related Mortality: A Prospective Study in Italy. BMC Infect. Dis. 2021, 21, 566. [Google Scholar] [CrossRef]
- Ramirez-Sandoval, J.C.; Castillos-Ávalos, V.J.; Paz-Cortés, A.; Santillan-Ceron, A.; Hernandez-Jimenez, S.; Mehta, R.; Correa-Rotter, R. Very Low Vitamin D Levels Are a Strong Independent Predictor of Mortality in Hospitalized Patients with Severe COVID-19. Arch. Med. Res. 2022, 53, 215–222. [Google Scholar] [CrossRef]
- Dissanayake, H.A.; de Silva, N.L.; Sumanatilleke, M.; de Silva, S.D.N.; Gamage, K.K.K.; Dematapitiya, C.; Kuruppu, D.C.; Ranasinghe, P.; Pathmanathan, S.; Katulanda, P. Prognostic and Therapeutic Role of Vitamin D in COVID-19: Systematic Review and Meta-Analysis. J. Clin. Endocrinol. Metab. 2022, 107, 1484–1502. [Google Scholar] [CrossRef]
- D’Ecclesiis, O.; Gavioli, C.; Martinoli, C.; Raimondi, S.; Chiocca, S.; Miccolo, C.; Bossi, P.; Cortinovis, D.; Chiaradonna, F.; Palorini, R.; et al. Vitamin D and SARS-CoV2 Infection, Severity and Mortality: A Systematic Review and Meta-Analysis. PLoS ONE 2022, 17, e0268396. [Google Scholar] [CrossRef]
- Loucera, C.; Peña-Chilet, M.; Esteban-Medina, M.; Muñoyerro-Muñiz, D.; Villegas, R.; Lopez-Miranda, J.; Rodriguez-Baño, J.; Túnez, I.; Bouillon, R.; Dopazo, J.; et al. Real World Evidence of Calcifediol or Vitamin D Prescription and Mortality Rate of COVID-19 in a Retrospective Cohort of Hospitalized Andalusian Patients. Sci. Rep. 2021, 11, 23380. [Google Scholar] [CrossRef]
- Nielsen, N.M.; Junker, T.G.; Cohen, A.S.; Munger, K.L.; Stenager, E.; Ascherio, A.; Boding, L.; Hviid, A. Vitamin D Status and Severity of COVID-19. Sci. Rep. 2022, 12, 19823. [Google Scholar] [CrossRef]
- Butler-Laporte, G.; Nakanishi, T.; Mooser, V.; Morrison, D.R.; Abdullah, T.; Adeleye, O.; Mamlouk, N.; Kimchi, N.; Afrasiabi, Z.; Rezk, N.; et al. Vitamin D and COVID-19 Susceptibility and Severity in the COVID-19 Host Genetics Initiative: A Mendelian Randomization Study. PLOS Med. 2021, 18, e1003605. [Google Scholar] [CrossRef]
- Freitas, A.T.; Calhau, C.; Antunes, G.; Araújo, B.; Bandeira, M.; Barreira, S.; Bazenga, F.; Braz, S.; Caldeira, D.; Santos, S.C.R.; et al. Vitamin D-Related Polymorphisms and Vitamin D Levels as Risk Biomarkers of COVID-19 Disease Severity. Sci. Rep. 2021, 11, 20837. [Google Scholar] [CrossRef] [PubMed]
- Hafezi, S.; Saheb Sharif-Askari, F.; Saheb Sharif-Askari, N.; Ali Hussain Alsayed, H.; Alsafar, H.; Al Anouti, F.; Hamid, Q.; Halwani, R. Vitamin D Enhances Type I IFN Signaling in COVID-19 Patients. Sci. Rep. 2022, 12, 17778. [Google Scholar] [CrossRef] [PubMed]
- Müller, V.; Fraser, C.; Herbeck, J.T. A Strong Case for Viral Genetic Factors in HIV Virulence. Viruses 2011, 3, 204–216. [Google Scholar] [CrossRef]
- Challen, R.; Brooks-Pollock, E.; Read, J.M.; Dyson, L.; Tsaneva-Atanasova, K.; Danon, L. Risk of Mortality in Patients Infected with SARS-CoV-2 Variant of Concern 202012/1: Matched Cohort Study. BMJ 2021, 372, n579. [Google Scholar] [CrossRef] [PubMed]
- Davies, N.G.; Jarvis, C.I.; Edmunds, W.J.; Jewell, N.P.; Diaz-Ordaz, K.; Keogh, R.H. Increased Mortality in Community-Tested Cases of SARS-CoV-2 Lineage B.1.1.7. Nature 2021, 593, 270–274. [Google Scholar] [CrossRef]
- Fisman, D.N.; Tuite, A.R. Evaluation of the Relative Virulence of Novel SARS-CoV-2 Variants: A Retrospective Cohort Study in Ontario, Canada. CMAJ 2021, 193, E1619–E1625. [Google Scholar] [CrossRef] [PubMed]
- Funk, T.; Pharris, A.; Spiteri, G.; Bundle, N.; Melidou, A.; Carr, M.; Gonzalez, G.; Garcia-Leon, A.; Crispie, F.; O’Connor, L.; et al. Characteristics of SARS-CoV-2 Variants of Concern B.1.1.7, B.1.351 or P.1: Data from Seven EU/EEA Countries, Weeks 38/2020 to 10/2021. Euro Surveill. Bull. Eur. Sur Mal. Transm. Eur. Commun. Dis. Bull. 2021, 26, 2100348. [Google Scholar] [CrossRef] [PubMed]
- Abu-Raddad, L.J.; Chemaitelly, H.; Ayoub, H.H.; Yassine, H.M.; Benslimane, F.M.; Al Khatib, H.A.; Tang, P.; Hasan, M.R.; Coyle, P.; AlMukdad, S.; et al. Severity, Criticality, and Fatality of the Severe Acute Respiratory Syndrome Coronavirus 2 (SARS-CoV-2) Beta Variant. Clin. Infect. Dis. 2021, 75, e1188–e1191. [Google Scholar] [CrossRef]
- Twohig, K.A.; Nyberg, T.; Zaidi, A.; Thelwall, S.; Sinnathamby, M.A.; Aliabadi, S.; Seaman, S.R.; Harris, R.J.; Hope, R.; Lopez-Bernal, J.; et al. Hospital Admission and Emergency Care Attendance Risk for SARS-CoV-2 Delta (B.1.617.2) Compared with Alpha (B.1.1.7) Variants of Concern: A Cohort Study. Lancet Infect. Dis. 2022, 22, 35–42. [Google Scholar] [CrossRef]
- Sheikh, A.; McMenamin, J.; Taylor, B.; Robertson, C. Public Health Scotland and the EAVE II Collaborators SARS-CoV-2 Delta VOC in Scotland: Demographics, Risk of Hospital Admission, and Vaccine Effectiveness. Lancet Lond. Engl. 2021, 397, 2461–2462. [Google Scholar] [CrossRef]
- Peralta-Santos, A.; Rodrigues, E.F.; Moreno, J.; Ricoca, V.; Casaca, P.; Fernandes, E.; Gomes, J.P.; Ferreira, R.; Isidro, J.; Pinto, M.; et al. Omicron (BA.1) SARS-CoV-2 Variant Is Associated with Reduced Risk of Hospitalization and Length of Stay Compared with Delta (B.1.617.2). medRxiv 2022. [Google Scholar] [CrossRef]
- Lewnard, J.A.; Hong, V.X.; Patel, M.M.; Kahn, R.; Lipsitch, M.; Tartof, S.Y. Clinical Outcomes among Patients Infected with Omicron (B.1.1.529) SARS-CoV-2 Variant in Southern California. medRxiv 2022. [Google Scholar] [CrossRef]
- Davies, M.-A.; Kassanjee, R.; Rousseau, P.; Morden, E.; Johnson, L.; Solomon, W.; Hsiao, N.-Y.; Hussey, H.; Meintjes, G.; Paleker, M.; et al. Outcomes of Laboratory-Confirmed SARS-CoV-2 Infection in the Omicron-Driven Fourth Wave Compared with Previous Waves in the Western Cape Province, South Africa. Trop. Med. Int. Health 2022, 27, 564–573. [Google Scholar] [CrossRef] [PubMed]
- Strasser, Z.H.; Greifer, N.; Hadavand, A.; Murphy, S.N.; Estiri, H. Estimates of SARS-CoV-2 Omicron BA.2 Subvariant Severity in New England. JAMA Netw. Open 2022, 5, e2238354. [Google Scholar] [CrossRef] [PubMed]
- Wolter, N.; Jassat, W.; Walaza, S.; Welch, R.; Moultrie, H.; Groome, M.; Amoako, D.G.; Everatt, J.; Bhiman, J.N.; Scheepers, C.; et al. Early Assessment of the Clinical Severity of the SARS-CoV-2 Omicron Variant in South Africa: A Data Linkage Study. Lancet 2022, 399, 437–446. [Google Scholar] [CrossRef] [PubMed]
- Romero, P.E.; Dávila-Barclay, A.; Salvatierra, G.; González, L.; Cuicapuza, D.; Solís, L.; Marcos-Carbajal, P.; Huancachoque, J.; Maturrano, L.; Tsukayama, P. The Emergence of Sars-CoV-2 Variant Lambda (C.37) in South America. Microbiol. Spectr. 2021, 9, e00789-21. [Google Scholar] [CrossRef] [PubMed]
- Bhattacharyya, R.P.; Hanage, W.P. Challenges in Inferring Intrinsic Severity of the SARS-CoV-2 Omicron Variant. N. Engl. J. Med. 2022, 386, e14. [Google Scholar] [CrossRef] [PubMed]
- Hui, K.P.Y.; Ho, J.C.W.; Cheung, M.-C.; Ng, K.-C.; Ching, R.H.H.; Lai, K.-L.; Kam, T.T.; Gu, H.; Sit, K.-Y.; Hsin, M.K.Y.; et al. SARS-CoV-2 Omicron Variant Replication in Human Bronchus and Lung Ex Vivo. Nature 2022, 603, 715–720. [Google Scholar] [CrossRef] [PubMed]
- Abdelnabi, R.; Foo, C.S.; Zhang, X.; Lemmens, V.; Maes, P.; Slechten, B.; Raymenants, J.; André, E.; Weynand, B.; Dallemier, K.; et al. The Omicron (B.1.1.529) SARS-CoV-2 Variant of Concern Does Not Readily Infect Syrian Hamsters. Microbiology 2022, 198, 105253. [Google Scholar] [CrossRef]
- Chen, D.-Y.; Kenney, D.; Chin, C.V.; Tavares, A.H.; Khan, N.; Conway, H.L.; Liu, G.; Choudhary, M.C.; Gertje, H.P.; O’Connell, A.K.; et al. Role of Spike in the Pathogenic and Antigenic Behavior of SARS-CoV-2 BA.1 Omicron. bioRxiv 2022. [Google Scholar] [CrossRef]
- Kudriavtsev, A.V.; Vakhrusheva, A.V.; Novoseletsky, V.N.; Bozdaganyan, M.E.; Shaitan, K.V.; Kirpichnikov, M.P.; Sokolova, O.S. Immune Escape Associated with RBD Omicron Mutations and SARS-CoV-2 Evolution Dynamics. Viruses 2022, 14, 1603. [Google Scholar] [CrossRef]
- Butt, A.A.; Dargham, S.R.; Coyle, P.; Yassine, H.M.; Al-Khal, A.; Abou-Samra, A.-B.; Abu-Raddad, L.J. COVID-19 Disease Severity in Persons Infected with Omicron BA.1 and BA.2 Sublineages and Association with Vaccination Status. JAMA Intern. Med. 2022, 182, 1097–1099. [Google Scholar] [CrossRef] [PubMed]
- Sievers, C.; Zacher, B.; Ullrich, A.; Huska, M.; Fuchs, S.; Buda, S.; Haas, W.; Diercke, M.; Heiden, M.A.D.; Kröger, S. SARS-CoV-2 Omicron Variants BA.1 and BA.2 Both Show Similarly Reduced Disease Severity of COVID-19 Compared to Delta, Germany, 2021 to 2022. Eurosurveillance 2022, 27, 2200396. [Google Scholar] [CrossRef] [PubMed]
- Davies, M.-A.; Morden, E.; Rosseau, P.; Arendse, J.; Bam, J.-L.; Boloko, L.; Cloete, K.; Cohen, C.; Chetty, N.; Dane, P.; et al. Outcomes of Laboratory-Confirmed SARS-CoV-2 Infection during Resurgence Driven by Omicron Lineages BA.4 and BA.5 Compared with Previous Waves in the Western Cape Province, South Africa. Int. J. Infect. Dis. 2023, 127, 63–68. [Google Scholar] [CrossRef] [PubMed]
- Lewnard, J.A.; Hong, V.; Tartof, S.Y. Association of SARS-CoV-2 BA.4/BA.5 Omicron Lineages with Immune Escape and Clinical Outcome. medRxiv 2022. [Google Scholar] [CrossRef]
- Hansen, C.H.; Friis, N.U.; Bager, P.; Stegger, M.; Fonager, J.; Fomsgaard, A.; Gram, M.A.; Christiansen, L.E.; Ethelberg, S.; Legarth, R.; et al. Risk of Reinfection, Vaccine Protection, and Severity of Infection with the BA.5 Omicron Subvariant: A Nation-Wide Population-Based Study in Denmark. Lancet Infect. Dis. 2022. [Google Scholar] [CrossRef]
- Markov, P.V.; Katzourakis, A.; Stilianakis, N.I. Antigenic Evolution Will Lead to New SARS-CoV-2 Variants with Unpredictable Severity. Nat. Rev. Microbiol. 2022, 20, 251–252. [Google Scholar] [CrossRef] [PubMed]
- Mourier, T.; Shuaib, M.; Hala, S.; Mfarrej, S.; Alofi, F.; Naeem, R.; Alsomali, A.; Jorgensen, D.; Subudhi, A.K.; Ben Rached, F.; et al. SARS-CoV-2 Genomes from Saudi Arabia Implicate Nucleocapsid Mutations in Host Response and Increased Viral Load. Nat. Commun. 2022, 13, 601. [Google Scholar] [CrossRef] [PubMed]
- Dumonteil, E.; Fusco, D.; Drouin, A.; Herrera, C. Genomic Signatures of SARS-CoV-2 Associated with Patient Mortality. Viruses 2021, 13, 227. [Google Scholar] [CrossRef]
- Volz, E.; Hill, V.; McCrone, J.T.; Price, A.; Jorgensen, D.; O’Toole, Á.; Southgate, J.; Johnson, R.; Jackson, B.; Nascimento, F.F.; et al. Evaluating the Effects of SARS-CoV-2 Spike Mutation D614G on Transmissibility and Pathogenicity. Cell 2021, 184, 64–75.e11. [Google Scholar] [CrossRef]
- Lee, A.C.-Y.; Zhang, A.J.; Chan, J.F.-W.; Li, C.; Fan, Z.; Liu, F.; Chen, Y.; Liang, R.; Sridhar, S.; Cai, J.-P.; et al. Oral SARS-CoV-2 Inoculation Establishes Subclinical Respiratory Infection with Virus Shedding in Golden Syrian Hamsters. Cell Rep. Med. 2020, 1, 100121. [Google Scholar] [CrossRef]
- Ryan, K.A.; Bewley, K.R.; Fotheringham, S.A.; Slack, G.S.; Brown, P.; Hall, Y.; Wand, N.I.; Marriott, A.C.; Cavell, B.E.; Tree, J.A.; et al. Dose-Dependent Response to Infection with SARS-CoV-2 in the Ferret Model and Evidence of Protective Immunity. Nat. Commun. 2021, 12, 81. [Google Scholar] [CrossRef]
- Imai, M.; Iwatsuki-Horimoto, K.; Hatta, M.; Loeber, S.; Halfmann, P.J.; Nakajima, N.; Watanabe, T.; Ujie, M.; Takahashi, K.; Ito, M.; et al. Syrian Hamsters as a Small Animal Model for SARS-CoV-2 Infection and Countermeasure Development. Proc. Natl. Acad. Sci. USA 2020, 117, 16587–16595. [Google Scholar] [CrossRef]
- Gandhi, M.; Beyrer, C.; Goosby, E. Masks Do More Than Protect Others During COVID-19: Reducing the Inoculum of SARS-CoV-2 to Protect the Wearer. J. Gen. Intern. Med. 2020, 35, 3063–3066. [Google Scholar] [CrossRef] [PubMed]
- Spinelli, M.A.; Glidden, D.V.; Gennatas, E.D.; Bielecki, M.; Beyrer, C.; Rutherford, G.; Chambers, H.; Goosby, E.; Gandhi, M. Importance of Non-Pharmaceutical Interventions in Lowering the Viral Inoculum to Reduce Susceptibility to Infection by SARS-CoV-2 and Potentially Disease Severity. Lancet Infect. Dis. 2021, 21, e296–e301. [Google Scholar] [CrossRef] [PubMed]
- Guallar, M.P.; Meiriño, R.; Donat-Vargas, C.; Corral, O.; Jouvé, N.; Soriano, V. Inoculum at the Time of SARS-CoV-2 Exposure and Risk of Disease Severity. Int. J. Infect. Dis. 2020, 97, 290–292. [Google Scholar] [CrossRef] [PubMed]
- Bielecki, M.; Züst, R.; Siegrist, D.; Meyerhofer, D.; Crameri, G.A.G.; Stanga, Z.; Stettbacher, A.; Buehrer, T.W.; Deuel, J.W. Social Distancing Alters the Clinical Course of COVID-19 in Young Adults: A Comparative Cohort Study. Clin. Infect. Dis. 2021, 72, 598–603. [Google Scholar] [CrossRef] [PubMed]
- Van Damme, W.; Dahake, R.; van de Pas, R.; Vanham, G.; Assefa, Y. COVID-19: Does the Infectious Inoculum Dose-Response Relationship Contribute to Understanding Heterogeneity in Disease Severity and Transmission Dynamics? Med. Hypotheses 2021, 146, 110431. [Google Scholar] [CrossRef] [PubMed]
- Han, A.; Czajkowski, L.M.; Donaldson, A.; Baus, H.A.; Reed, S.M.; Athota, R.S.; Bristol, T.; Rosas, L.A.; Cervantes-Medina, A.; Taubenberger, J.K.; et al. A Dose-Finding Study of a Wild-Type Influenza A(H3N2) Virus in a Healthy Volunteer Human Challenge Model. Clin. Infect. Dis. Off. Publ. Infect. Dis. Soc. Am. 2019, 69, 2082–2090. [Google Scholar] [CrossRef]
- Memoli, M.J.; Czajkowski, L.; Reed, S.; Athota, R.; Bristol, T.; Proudfoot, K.; Fargis, S.; Stein, M.; Dunfee, R.L.; Shaw, P.A.; et al. Validation of the Wild-Type Influenza A Human Challenge Model H1N1pdMIST: An A(H1N1)Pdm09 Dose-Finding Investigational New Drug Study. Clin. Infect. Dis. Off. Publ. Infect. Dis. Soc. Am. 2015, 60, 693–702. [Google Scholar] [CrossRef]
- Yu, I.T.S.; Li, Y.; Wong, T.W.; Tam, W.; Chan, A.T.; Lee, J.H.W.; Leung, D.Y.C.; Ho, T. Evidence of Airborne Transmission of the Severe Acute Respiratory Syndrome Virus. N. Engl. J. Med. 2004, 350, 1731–1739. [Google Scholar] [CrossRef]
- Mills, J.; Van Kirk, J.E.; Wright, P.F.; Chanock, R.M. Experimental Respiratory Syncytial Virus Infection of Adults. Possible Mechanisms of Resistance to Infection and Illness. J. Immunol. Baltim. Md 1950 1971, 107, 123–130. [Google Scholar]
- Little, P.; Read, R.C.; Amlôt, R.; Chadborn, T.; Rice, C.; Bostock, J.; Yardley, L. Reducing Risks from Coronavirus Transmission in the Home—The Role of Viral Load. BMJ 2020, 369, m1728. [Google Scholar] [CrossRef]
- Bixler, S.L.; Stefan, C.P.; Jay, A.N.; Rossi, F.D.; Ricks, K.M.; Shoemaker, C.J.; Moreau, A.M.; Zeng, X.; Hooper, J.W.; Dyer, D.N.; et al. Exposure Route Influences Disease Severity in the COVID-19 Cynomolgus Macaque Model. Viruses 2022, 14, 1013. [Google Scholar] [CrossRef] [PubMed]
- Port, J.R.; Yinda, C.K.; Owusu, I.O.; Holbrook, M.; Fischer, R.; Bushmaker, T.; Avanzato, V.A.; Schulz, J.E.; Martens, C.; van Doremalen, N.; et al. SARS-CoV-2 Disease Severity and Transmission Efficiency Is Increased for Airborne Compared to Fomite Exposure in Syrian Hamsters. Nat. Commun. 2021, 12, 4985. [Google Scholar] [CrossRef] [PubMed]
- Brosseau, L.M.; Escandón, K.; Ulrich, A.K.; Rasmussen, A.L.; Roy, C.J.; Bix, G.J.; Popescu, S.V.; Moore, K.A.; Osterholm, M.T. Severe Acute Respiratory Syndrome Coronavirus 2 (SARS-CoV-2) Dose, Infection, and Disease Outcomes for Coronavirus Disease 2019 (COVID-19): A Review. Clin. Infect. Dis. 2022, 75, e1195–e1201. [Google Scholar] [CrossRef] [PubMed]
- University of Oxford. A Dose Finding Human Experimental Infection Study with SARS-CoV-2 in Healthy Volunteers with Immunologically Sensitised with Either Previous, Microbiologically Confirmed, SARS-CoV-2 Infection and/or Vaccination against SARS-CoV2; University of Oxford: Oxford, UK, 2022. Available online: https://clinicaltrials.gov/ct2/show/NCT04864548 (accessed on 11 December 2022).
- Bilal, U.; Cainzos-Achirica, M.; Cleries, M.; Santaeugènia, S.; Corbella, X.; Comin-Colet, J.; Vela, E. Socioeconomic Status, Life Expectancy and Mortality in a Universal Healthcare Setting: An Individual-Level Analysis of >6 Million Catalan Residents. Prev. Med. 2019, 123, 91–94. [Google Scholar] [CrossRef] [PubMed]
- Sanzenbacher, G.T.; Webb, A.; Cosgrove, C.M.; Orlova, N. Rising Inequality in Life Expectancy by Socioeconomic Status. N. Am. Actuar. J. 2021, 25, S566–S581. [Google Scholar] [CrossRef]
- Alkire, S.; Dirksen, J.; Nogales, R.; Oldiges, C. Multidimensional Poverty and COVID-19 Risk Factors: A Rapid Overview of Interlinked Deprivations across 5.8 Billion People. In Oxford Poverty and Human Development Initiative (OPHI); University of Oxford: Oxford, UK, 2021; Volume 53a, pp. 1–12. [Google Scholar]
- Desmet, K.; Wacziarg, R. JUE Insight: Understanding Spatial Variation in COVID-19 across the United States. J. Urban Econ. 2022, 127, 103332. [Google Scholar] [CrossRef]
- Sun, Y.; Hu, X.; Xie, J. Spatial Inequalities of COVID-19 Mortality Rate in Relation to Socioeconomic and Environmental Factors across England. Sci. Total Environ. 2021, 758, 143595. [Google Scholar] [CrossRef]
- Abedi, V.; Olulana, O.; Avula, V.; Chaudhary, D.; Khan, A.; Shahjouei, S.; Li, J.; Zand, R. Racial, Economic, and Health Inequality and COVID-19 Infection in the United States. J. Racial Ethn. Health Disparities 2021, 8, 732–742. [Google Scholar] [CrossRef]
- Wachtler, B.; Michalski, N.; Nowossadeck, E.; Diercke, M.; Wahrendorf, M.; Santos-Hövener, C.; Lampert, T.; Hoebel, J. Socioeconomic Inequalities and COVID-19—A Review of the Current International Literature. J. Health Monit. 2020, 5, 3–17. [Google Scholar] [CrossRef]
- Martín-Sánchez, F.J.; Valls Carbó, A.; Miró, Ò.; Llorens, P.; Jiménez, S.; Piñera, P.; Burillo-Putze, G.; Martín, A.; García-Lamberechts, J.E.; Jacob, J.; et al. Socio-Demographic Health Determinants Are Associated with Poor Prognosis in Spanish Patients Hospitalized with COVID-19. J. Gen. Intern. Med. 2021, 36, 3737–3742. [Google Scholar] [CrossRef]
- Vaughan, L.; Veruttipong, D.; Shaw, J.G.; Levy, N.; Edwards, L.; Winget, M. Relationship of Socio-Demographics, Comorbidities, Symptoms and Healthcare Access with Early COVID-19 Presentation and Disease Severity. BMC Infect. Dis. 2021, 21, 40. [Google Scholar] [CrossRef] [PubMed]
- Ingraham, N.E.; Purcell, L.N.; Karam, B.S.; Dudley, R.A.; Usher, M.G.; Warlick, C.A.; Allen, M.L.; Melton, G.B.; Charles, A.; Tignanelli, C.J. Racial and Ethnic Disparities in Hospital Admissions from COVID-19: Determining the Impact of Neighborhood Deprivation and Primary Language. J. Gen. Intern. Med. 2021, 36, 3462–3470. [Google Scholar] [CrossRef] [PubMed]
- Little, C.; Alsen, M.; Barlow, J.; Naymagon, L.; Tremblay, D.; Genden, E.; Trosman, S.; Iavicoli, L.; van Gerwen, M. The Impact of Socioeconomic Status on the Clinical Outcomes of COVID-19; a Retrospective Cohort Study. J. Community Health 2021, 46, 794–802. [Google Scholar] [CrossRef] [PubMed]
- Moreno-Montoya, J.; Ballesteros, S.M.; Idrovo, A.J. COVID-19 Distribution in Bogotá, Colombia: Effect of Poverty during the First 2 Months of Pandemic. J. Epidemiol. Community Health 2022, 76, 116–120. [Google Scholar] [CrossRef]
- Broad, J.; Forman, J.; Brighouse, J.; Sobande, A.; McIntosh, A.; Watterson, C.; Boot, E.; Montgomery, F.; Gilmour, I.; Tan, J.; et al. Post-COVID-19 Paediatric Inflammatory Multisystem Syndrome: Association of Ethnicity, Key Worker and Socioeconomic Status with Risk and Severity. Arch. Dis. Child. 2021, 106, 1218–1225. [Google Scholar] [CrossRef]
- Ríos, V.; Denova-Gutiérrez, E.; Barquera, S. Association between Living in Municipalities with High Crowding Conditions and Poverty and Mortality from COVID-19 in Mexico. PLoS ONE 2022, 17, e0264137. [Google Scholar] [CrossRef]
- Millán-Guerrero, R.O.; Caballero-Hoyos, R.; Monárrez-Espino, J. Poverty and Survival from COVID-19 in Mexico. J. Public Health 2021, 43, 437–444. [Google Scholar] [CrossRef]
- Sesé, L.; Nguyen, Y.; Leprieur, E.G.; Annesi-Maesano, I.; Cavalin, C.; Bouillé, J.G.d.; Demestier, L.; Dhote, R.; Tandjaoui-Lambiotte, Y.; Bauvois, A.; et al. Impact of Socioeconomic Status in Patients Hospitalised for COVID-19 in the Greater Paris Area. Eur. Respir. J. 2020, 56, 2002364. [Google Scholar] [CrossRef]
- Lone, N.I.; McPeake, J.; Stewart, N.I.; Blayney, M.C.; Seem, R.C.; Donaldson, L.; Glass, E.; Haddow, C.; Hall, R.; Martin, C.; et al. Influence of Socioeconomic Deprivation on Interventions and Outcomes for Patients Admitted with COVID-19 to Critical Care Units in Scotland: A National Cohort Study. Lancet Reg. Health—Eur. 2021, 1, 100005. [Google Scholar] [CrossRef]
- Chung, G.K.-K.; Chan, S.-M.; Chan, Y.-H.; Yip, T.C.-F.; Ma, H.-M.; Wong, G.L.-H.; Chung, R.Y.-N.; Wong, H.; Wong, S.Y.-S.; Yeoh, E.K.; et al. Differential Impacts of Multimorbidity on COVID-19 Severity across the Socioeconomic Ladder in Hong Kong: A Syndemic Perspective. Int. J. Environ. Res. Public. Health 2021, 18, 8168. [Google Scholar] [CrossRef]
- Zakeri, R.; Bendayan, R.; Ashworth, M.; Bean, D.M.; Dodhia, H.; Durbaba, S.; O’Gallagher, K.; Palmer, C.; Curcin, V.; Aitken, E.; et al. A Case-Control and Cohort Study to Determine the Relationship between Ethnic Background and Severe COVID-19. EClinicalMedicine 2020, 28, 100574. [Google Scholar] [CrossRef]
- Patel, A.P.; Paranjpe, M.D.; Kathiresan, N.P.; Rivas, M.A.; Khera, A.V. Race, Socioeconomic Deprivation, and Hospitalization for COVID-19 in English Participants of a National Biobank. Int. J. Equity Health 2020, 19, 114. [Google Scholar] [CrossRef]
- Razieh, C.; Zaccardi, F.; Islam, N.; Gillies, C.L.; Chudasama, Y.V.; Rowlands, A.; Kloecker, D.E.; Davies, M.J.; Khunti, K.; Yates, T. Ethnic Minorities and COVID-19: Examining Whether Excess Risk Is Mediated through Deprivation. Eur. J. Public Health 2021, 31, 630–634. [Google Scholar] [CrossRef]
- Agyemang, C.; Richters, A.; Jolani, S.; Hendriks, S.; Zalpuri, S.; Yu, E.; Pijls, B.; Prins, M.; Stronks, K.; Zeegers, M.P. Ethnic Minority Status as Social Determinant for COVID-19 Infection, Hospitalisation, Severity, ICU Admission and Deaths in the Early Phase of the Pandemic: A Meta-Analysis. BMJ Glob. Health 2021, 6, e007433. [Google Scholar] [CrossRef]
- Mackey, K.; Ayers, C.K.; Kondo, K.K.; Saha, S.; Advani, S.M.; Young, S.; Spencer, H.; Rusek, M.; Anderson, J.; Veazie, S.; et al. Racial and Ethnic Disparities in COVID-19–Related Infections, Hospitalizations, and Deaths: A Systematic Review. Ann. Intern. Med. 2021, 174, 362–373. [Google Scholar] [CrossRef]
- Arasteh, K. Prevalence of Comorbidities and Risks Associated with COVID-19 among Black and Hispanic Populations in New York City: An Examination of the 2018 New York City Community Health Survey. J. Racial Ethn. Health Disparities 2021, 8, 863–869. [Google Scholar] [CrossRef]
- Lee, H.; Shin, S.H.; Gu, S.; Zhao, D.; Kang, D.; Joi, Y.R.; Suh, G.Y.; Pastor-Barriuso, R.; Guallar, E.; Cho, J.; et al. Racial Differences in Comorbidity Profile among Patients with Chronic Obstructive Pulmonary Disease. BMC Med. 2018, 16, 178. [Google Scholar] [CrossRef]
- Daw, J. Contribution of Four Comorbid Conditions to Racial/Ethnic Disparities in Mortality Risk. Am. J. Prev. Med. 2017, 52, S95–S102. [Google Scholar] [CrossRef]
- Cossrow, N.; Falkner, B. Race/Ethnic Issues in Obesity and Obesity-Related Comorbidities. J. Clin. Endocrinol. Metab. 2004, 89, 2590–2594. [Google Scholar] [CrossRef]
- Paeratakul, S.; Lovejoy, J.C.; Ryan, D.H.; Bray, G.A. The Relation of Gender, Race and Socioeconomic Status to Obesity and Obesity Comorbidities in a Sample of US Adults. Int. J. Obes. 2002, 26, 1205–1210. [Google Scholar] [CrossRef]
- Williams, D.R. Race/Ethnicity and Socioeconomic Status: Measurement and Methodological Issues. Int. J. Health Serv. 1996, 26, 483–505. [Google Scholar] [CrossRef] [PubMed]
- Fowler, D.; Brimblecombe, P.; Burrows, J.; Heal, M.R.; Grennfelt, P.; Stevenson, D.S.; Jowett, A.; Nemitz, E.; Coyle, M.; Liu, X.; et al. A Chronology of Global Air Quality. Philos. Trans. R. Soc. Math. Phys. Eng. Sci. 2020, 378, 20190314. [Google Scholar] [CrossRef]
- Rzymski, P.; Poniedziałek, B.; Rosińska, J.; Rogalska, M.; Zarębska-Michaluk, D.; Rorat, M.; Moniuszko-Malinowska, A.; Lorenc, B.; Kozielewicz, D.; Piekarska, A.; et al. The Association of Airborne Particulate Matter and Benzo[a]Pyrene with the Clinical Course of COVID-19 in Patients Hospitalized in Poland. Environ. Pollut. 2022, 306, 119469. [Google Scholar] [CrossRef] [PubMed]
- Kogevinas, M.; Castaño-Vinyals, G.; Karachaliou, M.; Espinosa, A.; de Cid, R.; Garcia-Aymerich, J.; Carreras, A.; Cortés, B.; Pleguezuelos, V.; Jiménez, A.; et al. Ambient Air Pollution in Relation to SARS-CoV-2 Infection, Antibody Response, and COVID-19 Disease: A Cohort Study in Catalonia, Spain (COVICAT Study). Environ. Health Perspect. 2021, 129, 117003. [Google Scholar] [CrossRef]
- Bozack, A.; Pierre, S.; DeFelice, N.; Colicino, E.; Jack, D.; Chillrud, S.N.; Rundle, A.; Astua, A.; Quinn, J.W.; McGuinn, L.; et al. Long-Term Air Pollution Exposure and COVID-19 Mortality: A Patient-Level Analysis from New York City. Am. J. Respir. Crit. Care Med. 2022, 205, 651–662. [Google Scholar] [CrossRef]
- Di Ciaula, A.; Bonfrate, L.; Portincasa, P.; Appice, C.; Belfiore, A.; Binetti, M.; Cafagna, G.; Campanale, G.; Carrieri, A.; Cascella, G.; et al. Nitrogen Dioxide Pollution Increases Vulnerability to COVID-19 through Altered Immune Function. Environ. Sci. Pollut. Res. 2022, 29, 44404–44412. [Google Scholar] [CrossRef]
- López-Feldman, A.; Heres, D.; Marquez-Padilla, F. Air Pollution Exposure and COVID-19: A Look at Mortality in Mexico City Using Individual-Level Data. Sci. Total Environ. 2021, 756, 143929. [Google Scholar] [CrossRef]
- Mendy, A.; Wu, X.; Keller, J.L.; Fassler, C.S.; Apewokin, S.; Mersha, T.B.; Xie, C.; Pinney, S.M. Long-Term Exposure to Fine Particulate Matter and Hospitalization in COVID-19 Patients. Respir. Med. 2021, 178, 106313. [Google Scholar] [CrossRef]
- Mendy, A.; Wu, X.; Keller, J.L.; Fassler, C.S.; Apewokin, S.; Mersha, T.B.; Xie, C.; Pinney, S.M. Air Pollution and the Pandemic: Long-Term PM2.5 Exposure and Disease Severity in COVID-19 Patients. Respirology 2021, 26, 1181–1187. [Google Scholar] [CrossRef]
- Chen, C.; Wang, J.; Kwong, J.; Kim, J.; van Donkelaar, A.; Martin, R.V.; Hystad, P.; Su, Y.; Lavigne, E.; Kirby-McGregor, M.; et al. Association between Long-Term Exposure to Ambient Air Pollution and COVID-19 Severity: A Prospective Cohort Study. CMAJ 2022, 194, E693–E700. [Google Scholar] [CrossRef]
- Bergamaschi, R.; Ponzano, M.; Schiavetti, I.; Carmisciano, L.; Cordioli, C.; Filippi, M.; Radaelli, M.; Immovilli, P.; Capobianco, M.; De Rossi, N.; et al. The Effect of Air Pollution on COVID-19 Severity in a Sample of Patients with Multiple Sclerosis. Eur. J. Neurol. 2022, 29, 535–542. [Google Scholar] [CrossRef]
- Bowe, B.; Xie, Y.; Gibson, A.K.; Cai, M.; van Donkelaar, A.; Martin, R.V.; Burnett, R.; Al-Aly, Z. Ambient Fine Particulate Matter Air Pollution and the Risk of Hospitalization among COVID-19 Positive Individuals: Cohort Study. Environ. Int. 2021, 154, 106564. [Google Scholar] [CrossRef] [PubMed]
- Elliott, J.; Bodinier, B.; Whitaker, M.; Delpierre, C.; Vermeulen, R.; Tzoulaki, I.; Elliott, P.; Chadeau-Hyam, M. COVID-19 Mortality in the UK Biobank Cohort: Revisiting and Evaluating Risk Factors. Eur. J. Epidemiol. 2021, 36, 299–309. [Google Scholar] [CrossRef] [PubMed]
- Konstantinoudis, G.; Padellini, T.; Bennett, J.; Davies, B.; Ezzati, M.; Blangiardo, M. Long-Term Exposure to Air-Pollution and COVID-19 Mortality in England: A Hierarchical Spatial Analysis. Environ. Int. 2021, 146, 106316. [Google Scholar] [CrossRef] [PubMed]
- Pegoraro, V.; Heiman, F.; Levante, A.; Urbinati, D.; Peduto, I. An Italian Individual-Level Data Study Investigating on the Association between Air Pollution Exposure and COVID-19 Severity in Primary-Care Setting. BMC Public Health 2021, 21, 902. [Google Scholar] [CrossRef]
- Marquès, M.; Correig, E.; Ibarretxe, D.; Anoro, E.; Antonio Arroyo, J.; Jericó, C.; Borrallo, R.M.; Miret, M.; Näf, S.; Pardo, A.; et al. Long-Term Exposure to PM10 above WHO Guidelines Exacerbates COVID-19 Severity and Mortality. Environ. Int. 2022, 158, 106930. [Google Scholar] [CrossRef]
- Chen, Z.; Huang, B.Z.; Sidell, M.A.; Chow, T.; Eckel, S.P.; Pavlovic, N.; Martinez, M.P.; Lurmann, F.; Thomas, D.C.; Gilliland, F.D.; et al. Near-Roadway Air Pollution Associated with COVID-19 Severity and Mortality—Multiethnic Cohort Study in Southern California. Environ. Int. 2021, 157, 106862. [Google Scholar] [CrossRef]
- Gangwar, R.S.; Bevan, G.H.; Palanivel, R.; Das, L.; Rajagopalan, S. Oxidative Stress Pathways of Air Pollution Mediated Toxicity: Recent Insights. Redox Biol. 2020, 34, 101545. [Google Scholar] [CrossRef]
- Yan, Z.; Jin, Y.; An, Z.; Liu, Y.; Samet, J.M.; Wu, W. Inflammatory Cell Signaling Following Exposures to Particulate Matter and Ozone. Biochim. Biophys. Acta BBA—Gen. Subj. 2016, 1860, 2826–2834. [Google Scholar] [CrossRef]
- Manivannan, J.; Sundaresan, L. Systems Level Insights into the Impact of Airborne Exposure on SARS-CoV-2 Pathogenesis and COVID-19 Outcome—A Multi-Omics Big Data Study. Gene Rep. 2021, 25, 101312. [Google Scholar] [CrossRef]
- Woodby, B.; Arnold, M.M.; Valacchi, G. SARS-CoV-2 Infection, COVID-19 Pathogenesis, and Exposure to Air Pollution: What Is the Connection? Ann. N. Y. Acad. Sci. 2021, 1486, 15–38. [Google Scholar] [CrossRef]
- Signorini, A.; Segre, A.M.; Polgreen, P.M. The Use of Twitter to Track Levels of Disease Activity and Public Concern in the U.S. during the Influenza A H1N1 Pandemic. PLoS ONE 2011, 6, e19467. [Google Scholar] [CrossRef] [PubMed]
- Lin, C.-I.; Tsai, C.-H.; Sun, Y.-L.; Hsieh, W.-Y.; Lin, Y.-C.; Chen, C.-Y.; Lin, C.-S. Instillation of Particulate Matter 2.5 Induced Acute Lung Injury and Attenuated the Injury Recovery in ACE2 Knockout Mice. Int. J. Biol. Sci. 2018, 14, 253–265. [Google Scholar] [CrossRef] [PubMed]
- Li, H.-H.; Liu, C.-C.; Hsu, T.-W.; Lin, J.-H.; Hsu, J.-W.; Li, A.F.-Y.; Yeh, Y.-C.; Hung, S.-C.; Hsu, H.-S. Upregulation of ACE2 and TMPRSS2 by Particulate Matter and Idiopathic Pulmonary Fibrosis: A Potential Role in Severe COVID-19. Part. Fibre Toxicol. 2021, 18, 11. [Google Scholar] [CrossRef] [PubMed]
- Steptoe, A.; Feldman, P.J. Neighborhood Problems as Sources of Chronic Stress: Development of a Measure of Neighborhood Problems, and Associations with Socioeconomic Status and Health. Ann. Behav. Med. 2001, 23, 177–185. [Google Scholar] [CrossRef] [PubMed]
- Zajacova, A.; Dowd, J.B.; Aiello, A.E. Socioeconomic and Race/Ethnic Patterns in Persistent Infection Burden among U.S. Adults. J. Gerontol. Ser. A 2009, 64, 272–279. [Google Scholar] [CrossRef] [PubMed]
- Dowd, J.B.; Zajacova, A.; Aiello, A. Early Origins of Health Disparities: Burden of Infection, Health, and Socioeconomic Status in U.S. Children. Soc. Sci. Med. 2009, 68, 699–707. [Google Scholar] [CrossRef]
- Yaya, S.; Bishwajit, G. Burden of Acute Respiratory Infections among Under-Five Children in Relation to Household Wealth and Socioeconomic Status in Bangladesh. Trop. Med. Infect. Dis. 2019, 4, 36. [Google Scholar] [CrossRef]
- Singleton, C.R.; Winkler, M.; Houghtaling, B.; Adeyemi, O.S.; Roehll, A.M.; Pionke, J.J.; Anderson Steeves, E. Understanding the Intersection of Race/Ethnicity, Socioeconomic Status, and Geographic Location: A Scoping Review of U.S. Consumer Food Purchasing. Int. J. Environ. Res. Public. Health 2020, 17, 7677. [Google Scholar] [CrossRef]
- Belanger, M.J.; Hill, M.A.; Angelidi, A.M.; Dalamaga, M.; Sowers, J.R.; Mantzoros, C.S. COVID-19 and Disparities in Nutrition and Obesity. N. Engl. J. Med. 2020, 383, e69. [Google Scholar] [CrossRef]
- Rehm, C.D.; Peñalvo, J.L.; Afshin, A.; Mozaffarian, D. Dietary Intake among US Adults, 1999-2012. JAMA 2016, 315, 2542–2553. [Google Scholar] [CrossRef]
- Wallace, J.M.; Vaughn, M.G.; Bachman, J.G.; O’Malley, P.M.; Johnston, L.D.; Schulenberg, J.E. Race/Ethnicity, Socioeconomic Factors, and Smoking among Early Adolescent Girls in the United States. Drug Alcohol Depend. 2009, 104, S42–S49. [Google Scholar] [CrossRef] [PubMed]
- Wagenknecht, L.E.; Perkins, L.L.; Cutter, G.R.; Sidney, S.; Burke, G.L.; Manolio, T.A.; Jacobs, D.R.; Liu, K.A.; Friedman, G.D.; Hughes, G.H. Cigarette Smoking Behavior Is Strongly Related to Educational Status: The CARDIA Study. Prev. Med. 1990, 19, 158–169. [Google Scholar] [CrossRef] [PubMed]
- Dieteren, C.; Bonfrer, I. Socioeconomic Inequalities in Lifestyle Risk Factors across Low- and Middle-Income Countries. BMC Public Health 2021, 21, 951. [Google Scholar] [CrossRef] [PubMed]
- Harrison, C.A.; Taren, D. How Poverty Affects Diet to Shape the Microbiota and Chronic Disease. Nat. Rev. Immunol. 2018, 18, 279–287. [Google Scholar] [CrossRef]
- Miller, G.E.; Engen, P.A.; Gillevet, P.M.; Shaikh, M.; Sikaroodi, M.; Forsyth, C.B.; Mutlu, E.; Keshavarzian, A. Lower Neighborhood Socioeconomic Status Associated with Reduced Diversity of the Colonic Microbiota in Healthy Adults. PLoS ONE 2016, 11, e0148952. [Google Scholar] [CrossRef]
- Bowyer, R.C.E.; Jackson, M.A.; Le Roy, C.I.; Ni Lochlainn, M.; Spector, T.D.; Dowd, J.B.; Steves, C.J. Socioeconomic Status and the Gut Microbiome: A TwinsUK Cohort Study. Microorganisms 2019, 7, 17. [Google Scholar] [CrossRef]
- Foster, H.M.; Celis-Morales, C.A.; Nicholl, B.I.; Petermann-Rocha, F.; Pell, J.P.; Gill, J.M.; O’Donnell, C.A.; Mair, F.S. The Effect of Socioeconomic Deprivation on the Association between an Extended Measurement of Unhealthy Lifestyle Factors and Health Outcomes: A Prospective Analysis of the UK Biobank Cohort. Lancet Public Health 2018, 3, e576–e585. [Google Scholar] [CrossRef]
- Assari, S.; Chalian, H.; Bazargan, M. Race, Ethnicity, Socioeconomic Status, and Chronic Lung Disease in the U.S. Res. Health Sci. 2020, 5, 48–63. [Google Scholar] [CrossRef]
- Bello, A.K.; Peters, J.; Rigby, J.; Rahman, A.A.; Nahas, M.E. Socioeconomic Status and Chronic Kidney Disease at Presentation to a Renal Service in the United Kingdom. Clin. J. Am. Soc. Nephrol. 2008, 3, 1316–1323. [Google Scholar] [CrossRef]
- Guariguata, L.; Whiting, D.R.; Hambleton, I.; Beagley, J.; Linnenkamp, U.; Shaw, J.E. Global Estimates of Diabetes Prevalence for 2013 and Projections for 2035. Diabetes Res. Clin. Pract. 2014, 103, 137–149. [Google Scholar] [CrossRef]
- Gordon-Larsen, P.; Adair, L.S.; Popkin, B.M. The Relationship of Ethnicity, Socioeconomic Factors, and Overweight in U.S. Adolescents. Obes. Res. 2003, 11, 121–129. [Google Scholar] [CrossRef] [PubMed]
- Ward, E.; Jemal, A.; Cokkinides, V.; Singh, G.K.; Cardinez, C.; Ghafoor, A.; Thun, M. Cancer Disparities by Race/Ethnicity and Socioeconomic Status. CA. Cancer J. Clin. 2004, 54, 78–93. [Google Scholar] [CrossRef] [PubMed]
- Brown, T.; Dassonville, C.; Derbez, M.; Ramalho, O.; Kirchner, S.; Crump, D.; Mandin, C. Relationships between Socioeconomic and Lifestyle Factors and Indoor Air Quality in French Dwellings. Environ. Res. 2015, 140, 385–396. [Google Scholar] [CrossRef] [PubMed]
- Vanker, A.; Barnett, W.; Nduru, P.M.; Gie, R.P.; Sly, P.D.; Zar, H.J. Home Environment and Indoor Air Pollution Exposure in an African Birth Cohort Study. Sci. Total Environ. 2015, 536, 362–367. [Google Scholar] [CrossRef] [PubMed]
- Gordon, S.B.; Bruce, N.G.; Grigg, J.; Hibberd, P.L.; Kurmi, O.P.; Lam, K.H.; Mortimer, K.; Asante, K.P.; Balakrishnan, K.; Balmes, J.; et al. Respiratory Risks from Household Air Pollution in Low and Middle Income Countries. Lancet Respir. Med. 2014, 2, 823–860. [Google Scholar] [CrossRef]
- Singleton, R.; Salkoski, A.J.; Bulkow, L.; Fish, C.; Dobson, J.; Albertson, L.; Skarada, J.; Kovesi, T.; McDonald, C.; Hennessy, T.W.; et al. Housing Characteristics and Indoor Air Quality in Households of Alaska Native Children with Chronic Lung Conditions. Indoor Air 2017, 27, 478–486. [Google Scholar] [CrossRef]
- Ginsburg, A.S.; Izadnegahdar, R.; Berkley, J.A.; Walson, J.L.; Rollins, N.; Klugman, K.P. Undernutrition and Pneumonia Mortality. Lancet Glob. Health 2015, 3, e735–e736. [Google Scholar] [CrossRef]
- Holuka, C.; Merz, M.P.; Fernandes, S.B.; Charalambous, E.G.; Seal, S.V.; Grova, N.; Turner, J.D. The COVID-19 Pandemic: Does Our Early Life Environment, Life Trajectory and Socioeconomic Status Determine Disease Susceptibility and Severity? Int. J. Mol. Sci. 2020, 21, 5094. [Google Scholar] [CrossRef]
- Vliegenthart, J.; Noppe, G.; van Rossum, E.F.C.; Koper, J.W.; Raat, H.; van den Akker, E.L.T. Socioeconomic Status in Children Is Associated with Hair Cortisol Levels as a Biological Measure of Chronic Stress. Psychoneuroendocrinology 2016, 65, 9–14. [Google Scholar] [CrossRef]
- Cohen, S.; Doyle, W.J.; Baum, A. Socioeconomic Status Is Associated with Stress Hormones. Psychosom. Med. 2006, 68, 414–420. [Google Scholar] [CrossRef]
- Baum, A.; Garofalo, J.P.; Yali, A.M. Socioeconomic Status and Chronic Stress: Does Stress Account for SES Effects on Health? Ann. N. Y. Acad. Sci. 1999, 896, 131–144. [Google Scholar] [CrossRef] [PubMed]
- Öhlin, B.; Nilsson, P.M.; Nilsson, J.-Å.; Berglund, G. Chronic Psychosocial Stress Predicts Long-Term Cardiovascular Morbidity and Mortality in Middle-Aged Men. Eur. Heart J. 2004, 25, 867–873. [Google Scholar] [CrossRef] [PubMed]
- Kopp, M.S.; Réthelyi, J. Where Psychology Meets Physiology: Chronic Stress and Premature Mortality—The Central-Eastern European Health Paradox. Brain Res. Bull. 2004, 62, 351–367. [Google Scholar] [CrossRef]
- Roth, G.A.; Abate, D.; Abate, K.H.; Abay, S.M.; Abbafati, C.; Abbasi, N.; Abbastabar, H.; Abd-Allah, F.; Abdela, J.; Abdelalim, A.; et al. Global, Regional, and National Age-Sex-Specific Mortality for 282 Causes of Death in 195 Countries and Territories, 1980–2017: A Systematic Analysis for the Global Burden of Disease Study 2017. Lancet 2018, 392, 1736–1788. [Google Scholar] [CrossRef] [PubMed]
- Mozaffarian, D.; Wilson, P.W.F.; Kannel, W.B. Beyond Established and Novel Risk Factors. Circulation 2008, 117, 3031–3038. [Google Scholar] [CrossRef]
- Barbaresko, J.; Rienks, J.; Nöthlings, U. Lifestyle Indices and Cardiovascular Disease Risk: A Meta-Analysis. Am. J. Prev. Med. 2018, 55, 555–564. [Google Scholar] [CrossRef]
- Shigeta, H.; Shigeta, M.; Nakazawa, A.; Nakamura, N.; Yoshikawa, T. Lifestyle, Obesity, and Insulin Resistance. Diabetes Care 2001, 24, 608. [Google Scholar] [CrossRef]
- Hu, F.B. Globalization of Diabetes: The Role of Diet, Lifestyle, and Genes. Diabetes Care 2011, 34, 1249–1257. [Google Scholar] [CrossRef]
- Stengel, B.; Tarver-Carr, M.E.; Powe, N.R.; Eberhardt, M.S.; Brancati, F.L. Lifestyle Factors, Obesity and the Risk of Chronic Kidney Disease. Epidemiol. Camb. Mass 2003, 14, 479–487. [Google Scholar] [CrossRef]
- Hartman, J.E.; Boezen, H.M.; de Greef, M.H.; Bossenbroek, L.; ten Hacken, N.H. Consequences of Physical Inactivity in Chronic Obstructive Pulmonary Disease. Expert Rev. Respir. Med. 2010, 4, 735–745. [Google Scholar] [CrossRef]
- Gobbens, R.J.J.; van Assen, M.A.L.M.; Luijkx, K.G.; Wijnen-Sponselee, M.T.; Schols, J.M.G.A. Determinants of Frailty. J. Am. Med. Dir. Assoc. 2010, 11, 356–364. [Google Scholar] [CrossRef] [PubMed]
- Brinkman, S.; Voortman, T.; Kiefte-de Jong, J.C.; van Rooij, F.J.A.; Ikram, M.A.; Rivadeneira, F.; Franco, O.H.; Schoufour, J.D. The Association between Lifestyle and Overall Health, Using the Frailty Index. Arch. Gerontol. Geriatr. 2018, 76, 85–91. [Google Scholar] [CrossRef] [PubMed]
- Pasolli, E.; Asnicar, F.; Manara, S.; Zolfo, M.; Karcher, N.; Armanini, F.; Beghini, F.; Manghi, P.; Tett, A.; Ghensi, P.; et al. Extensive Unexplored Human Microbiome Diversity Revealed by Over 150,000 Genomes from Metagenomes Spanning Age, Geography, and Lifestyle. Cell 2019, 176, 649–662.e20. [Google Scholar] [CrossRef] [PubMed]
- Babszky, G.; Torma, F.; Aczel, D.; Bakonyi, P.; Gombos, Z.; Feher, J.; Szabó, D.; Ligeti, B.; Pongor, S.; Balogh, L.; et al. COVID-19 Infection Alters the Microbiome: Elite Athletes and Sedentary Patients Have Similar Bacterial Flora. Genes 2021, 12, 1577. [Google Scholar] [CrossRef] [PubMed]
- Barraza-Villarreal, A.; Sunyer, J.; Hernandez-Cadena, L.; Escamilla-Nuñez, M.C.; Sienra-Monge, J.J.; Ramírez-Aguilar, M.; Cortez-Lugo, M.; Holguin, F.; Diaz-Sánchez, D.; Olin, A.C.; et al. Air Pollution, Airway Inflammation, and Lung Function in a Cohort Study of Mexico City Schoolchildren. Environ. Health Perspect. 2008, 116, 832–838. [Google Scholar] [CrossRef]
- Abramson, M.J.; Wigmann, C.; Altug, H.; Schikowski, T. Ambient Air Pollution Is Associated with Airway Inflammation in Older Women: A Nested Cross-Sectional Analysis. BMJ Open Respir. Res. 2020, 7, e000549. [Google Scholar] [CrossRef]
- Guarnieri, M.; Balmes, J.R. Outdoor Air Pollution and Asthma. Lancet 2014, 383, 1581–1592. [Google Scholar] [CrossRef]
- Li, J.; Sun, S.; Tang, R.; Qiu, H.; Huang, Q.; Mason, T.G.; Tian, L. Major Air Pollutants and Risk of COPD Exacerbations: A Systematic Review and Meta-Analysis. Int. J. Chron. Obstruct. Pulmon. Dis. 2016, 11, 3079–3091. [Google Scholar] [CrossRef]
- Shah, A.S.V.; Langrish, J.P.; Nair, H.; McAllister, D.A.; Hunter, A.L.; Donaldson, K.; Newby, D.E.; Mills, N.L. Global Association of Air Pollution and Heart Failure: A Systematic Review and Meta-Analysis. Lancet Lond. Engl. 2013, 382, 1039–1048. [Google Scholar] [CrossRef]
- Yorifuji, T.; Kashima, S.; Tsuda, T.; Ishikawa-Takata, K.; Ohta, T.; Tsuruta, K.; Doi, H. Long-Term Exposure to Traffic-Related Air Pollution and the Risk of Death from Hemorrhagic Stroke and Lung Cancer in Shizuoka, Japan. Sci. Total Environ. 2013, 443, 397–402. [Google Scholar] [CrossRef]
- Park, S.K.; Adar, S.D.; O’Neill, M.S.; Auchincloss, A.H.; Szpiro, A.; Bertoni, A.G.; Navas-Acien, A.; Kaufman, J.D.; Diez-Roux, A.V. Long-Term Exposure to Air Pollution and Type 2 Diabetes Mellitus in a Multiethnic Cohort. Am. J. Epidemiol. 2015, 181, 327–336. [Google Scholar] [CrossRef]
- Vogiatzis, I.; Zakynthinos, G.; Andrianopoulos, V. Mechanisms of Physical Activity Limitation in Chronic Lung Diseases. Pulm. Med. 2012, 2012, e634761. [Google Scholar] [CrossRef]
- Dhillon, S.S.; Sima, C.A.; Kirkham, A.R.; Syed, N.; Camp, P.G. Physical Activity Measurement Accuracy in Individuals with Chronic Lung Disease: A Systematic Review with Meta-Analysis of Method Comparison Studies. Arch. Phys. Med. Rehabil. 2015, 96, 2079–2088. [Google Scholar] [CrossRef] [PubMed]
- Gallant, J.; Hsue, P.Y.; Shreay, S.; Meyer, N. Comorbidities among US Patients with Prevalent HIV Infection—A Trend Analysis. J. Infect. Dis. 2017, 216, 1525–1533. [Google Scholar] [CrossRef]
- Perz, J.F.; Armstrong, G.L.; Farrington, L.A.; Hutin, Y.J.F.; Bell, B.P. The Contributions of Hepatitis B Virus and Hepatitis C Virus Infections to Cirrhosis and Primary Liver Cancer Worldwide. J. Hepatol. 2006, 45, 529–538. [Google Scholar] [CrossRef] [PubMed]
- Jenkins, T.P.; Rathnayaka, Y.; Perera, P.K.; Peachey, L.E.; Nolan, M.J.; Krause, L.; Rajakaruna, R.S.; Cantacessi, C. Infections by Human Gastrointestinal Helminths Are Associated with Changes in Faecal Microbiota Diversity and Composition. PLoS ONE 2017, 12, e0184719. [Google Scholar] [CrossRef]
- Stensvold, C.R.; van der Giezen, M. Associations between Gut Microbiota and Common Luminal Intestinal Parasites. Trends Parasitol. 2018, 34, 369–377. [Google Scholar] [CrossRef] [PubMed]
- Giacomin, P.; Croese, J.; Krause, L.; Loukas, A.; Cantacessi, C. Suppression of Inflammation by Helminths: A Role for the Gut Microbiota? Philos. Trans. R. Soc. B Biol. Sci. 2015, 370, 20140296. [Google Scholar] [CrossRef] [PubMed]
- Zaiss, M.M.; Rapin, A.; Lebon, L.; Dubey, L.K.; Mosconi, I.; Sarter, K.; Piersigilli, A.; Menin, L.; Walker, A.W.; Rougemont, J.; et al. The Intestinal Microbiota Contributes to the Ability of Helminths to Modulate Allergic Inflammation. Immunity 2015, 43, 998–1010. [Google Scholar] [CrossRef] [PubMed]
- Gurung, M.; Li, Z.; You, H.; Rodrigues, R.; Jump, D.B.; Morgun, A.; Shulzhenko, N. Role of Gut Microbiota in Type 2 Diabetes Pathophysiology. EBioMedicine 2020, 51, 102590. [Google Scholar] [CrossRef] [PubMed]
- Qin, J.; Li, Y.; Cai, Z.; Li, S.; Zhu, J.; Zhang, F.; Liang, S.; Zhang, W.; Guan, Y.; Shen, D.; et al. A Metagenome-Wide Association Study of Gut Microbiota in Type 2 Diabetes. Nature 2012, 490, 55–60. [Google Scholar] [CrossRef]
- Ley, R.E.; Turnbaugh, P.J.; Klein, S.; Gordon, J.I. Human Gut Microbes Associated with Obesity. Nature 2006, 444, 1022–1023. [Google Scholar] [CrossRef] [PubMed]
- Turnbaugh, P.J.; Ley, R.E.; Mahowald, M.A.; Magrini, V.; Mardis, E.R.; Gordon, J.I. An Obesity-Associated Gut Microbiome with Increased Capacity for Energy Harvest. Nature 2006, 444, 1027–1031. [Google Scholar] [CrossRef] [PubMed]
- Tang, W.H.W.; Bäckhed, F.; Landmesser, U.; Hazen, S.L. Intestinal Microbiota in Cardiovascular Health and Disease: JACC State-of-the-Art Review. J. Am. Coll. Cardiol. 2019, 73, 2089–2105. [Google Scholar] [CrossRef] [PubMed]
- Trøseid, M.; Andersen, G.Ø.; Broch, K.; Hov, J.R. The Gut Microbiome in Coronary Artery Disease and Heart Failure: Current Knowledge and Future Directions. EBioMedicine 2020, 52, 102649. [Google Scholar] [CrossRef]
- Wilck, N.; Matus, M.G.; Kearney, S.M.; Olesen, S.W.; Forslund, K.; Bartolomaeus, H.; Haase, S.; Mähler, A.; Balogh, A.; Markó, L.; et al. Salt-Responsive Gut Commensal Modulates TH17 Axis and Disease. Nature 2017, 551, 585–589. [Google Scholar] [CrossRef]
- Yang, T.; Santisteban, M.M.; Rodriguez, V.; Li, E.; Ahmari, N.; Carvajal, J.M.; Zadeh, M.; Gong, M.; Qi, Y.; Zubcevic, J.; et al. Gut Dysbiosis Is Linked to Hypertension. Hypertension 2015, 65, 1331–1340. [Google Scholar] [CrossRef]
- Wypych, T.P.; Wickramasinghe, L.C.; Marsland, B.J. The Influence of the Microbiome on Respiratory Health. Nat. Immunol. 2019, 20, 1279–1290. [Google Scholar] [CrossRef]
- Zhang, D.; Li, S.; Wang, N.; Tan, H.-Y.; Zhang, Z.; Feng, Y. The Cross-Talk Between Gut Microbiota and Lungs in Common Lung Diseases. Front. Microbiol. 2020, 11, 301. [Google Scholar] [CrossRef]
- Qin, N.; Yang, F.; Li, A.; Prifti, E.; Chen, Y.; Shao, L.; Guo, J.; Le Chatelier, E.; Yao, J.; Wu, L.; et al. Alterations of the Human Gut Microbiome in Liver Cirrhosis. Nature 2014, 513, 59–64. [Google Scholar] [CrossRef]
- Mouries, J.; Brescia, P.; Silvestri, A.; Spadoni, I.; Sorribas, M.; Wiest, R.; Mileti, E.; Galbiati, M.; Invernizzi, P.; Adorini, L.; et al. Microbiota-Driven Gut Vascular Barrier Disruption Is a Prerequisite for Non-Alcoholic Steatohepatitis Development. J. Hepatol. 2019, 71, 1216–1228. [Google Scholar] [CrossRef]
- Ren, Z.; Fan, Y.; Li, A.; Shen, Q.; Wu, J.; Ren, L.; Lu, H.; Ding, S.; Ren, H.; Liu, C.; et al. Alterations of the Human Gut Microbiome in Chronic Kidney Disease. Adv. Sci. 2020, 7, 2001936. [Google Scholar] [CrossRef]
- Dinan, T.G.; Cryan, J.F. Gut Instincts: Microbiota as a Key Regulator of Brain Development, Ageing and Neurodegeneration. J. Physiol. 2017, 595, 489–503. [Google Scholar] [CrossRef]
- Liang, S.; Wu, X.; Hu, X.; Wang, T.; Jin, F. Recognizing Depression from the Microbiota–Gut–Brain Axis. Int. J. Mol. Sci. 2018, 19, 1592. [Google Scholar] [CrossRef]
- Lloyd-Price, J.; Abu-Ali, G.; Huttenhower, C. The Healthy Human Microbiome. Genome Med. 2016, 8, 51. [Google Scholar] [CrossRef] [PubMed]
- Jackson, M.A.; Jeffery, I.B.; Beaumont, M.; Bell, J.T.; Clark, A.G.; Ley, R.E.; O’Toole, P.W.; Spector, T.D.; Steves, C.J. Signatures of Early Frailty in the Gut Microbiota. Genome Med. 2016, 8, 8. [Google Scholar] [CrossRef] [PubMed]
- Shin, H.E.; Kwak, S.E.; Lee, J.-H.; Zhang, D.; Bae, J.H.; Song, W. Exercise, the Gut Microbiome, and Frailty. Ann. Geriatr. Med. Res. 2019, 23, 105–114. [Google Scholar] [CrossRef] [PubMed]
- Padilha de Lima, A.; Macedo Rogero, M.; Araujo Viel, T.; Garay-Malpartida, H.M.; Aprahamian, I.; Lima Ribeiro, S.M. Interplay between Inflammaging, Frailty and Nutrition in COVID-19: Preventive and Adjuvant Treatment Perspectives. J. Nutr. Health Aging 2022, 26, 67–76. [Google Scholar] [CrossRef]
- Bosco, N.; Noti, M. The Aging Gut Microbiome and Its Impact on Host Immunity. Genes Immun. 2021, 22, 289–303. [Google Scholar] [CrossRef]
- Ganesan, K.; Chung, S.K.; Vanamala, J.; Xu, B. Causal Relationship between Diet-Induced Gut Microbiota Changes and Diabetes: A Novel Strategy to Transplant Faecalibacterium Prausnitzii in Preventing Diabetes. Int. J. Mol. Sci. 2018, 19, 3720. [Google Scholar] [CrossRef]
- de Groot, P.F.; Belzer, C.; Aydin, Ö.; Levin, E.; Levels, J.H.; Aalvink, S.; Boot, F.; Holleman, F.; van Raalte, D.H.; Scheithauer, T.P.; et al. Distinct Fecal and Oral Microbiota Composition in Human Type 1 Diabetes, an Observational Study. PLoS ONE 2017, 12, e0188475. [Google Scholar] [CrossRef] [PubMed]
- Santisteban, M.M.; Qi, Y.; Zubcevic, J.; Kim, S.; Yang, T.; Shenoy, V.; Cole-Jeffrey, C.T.; Lobaton, G.O.; Stewart, D.C.; Rubiano, A.; et al. Hypertension-Linked Pathophysiological Alterations in the Gut. Circ. Res. 2017, 120, 312–323. [Google Scholar] [CrossRef]
- Jayashree, B.; Bibin, Y.S.; Prabhu, D.; Shanthirani, C.S.; Gokulakrishnan, K.; Lakshmi, B.S.; Mohan, V.; Balasubramanyam, M. Increased Circulatory Levels of Lipopolysaccharide (LPS) and Zonulin Signify Novel Biomarkers of Proinflammation in Patients with Type 2 Diabetes. Mol. Cell. Biochem. 2014, 388, 203–210. [Google Scholar] [CrossRef]
- Fulop, T.; Larbi, A.; Witkowski, J.M.; McElhaney, J.; Loeb, M.; Mitnitski, A.; Pawelec, G. Aging, Frailty and Age-Related Diseases. Biogerontology 2010, 11, 547–563. [Google Scholar] [CrossRef]
- Sewo Sampaio, P.Y.; Sampaio, R.A.C.; Coelho Júnior, H.J.; Teixeira, L.F.M.; Tessutti, V.D.; Uchida, M.C.; Arai, H. Differences in Lifestyle, Physical Performance and Quality of Life between Frail and Robust Brazilian Community-Dwelling Elderly Women. Geriatr. Gerontol. Int. 2016, 16, 829–835. [Google Scholar] [CrossRef] [PubMed]
- Wong, C.H.; Weiss, D.; Sourial, N.; Karunananthan, S.; Quail, J.M.; Wolfson, C.; Bergman, H. Frailty and Its Association with Disability and Comorbidity in a Community-Dwelling Sample of Seniors in Montreal: A Cross-Sectional Study. Aging Clin. Exp. Res. 2010, 22, 54–62. [Google Scholar] [CrossRef]
- Espinoza, S.E.; Quiben, M.; Hazuda, H.P. Distinguishing Comorbidity, Disability, and Frailty. Curr. Geriatr. Rep. 2018, 7, 201–209. [Google Scholar] [CrossRef]
- Foley, M.K.; Searle, S.D.; Toloue, A.; Booth, R.; Falkenham, A.; Falzarano, D.; Rubino, S.; Francis, M.E.; McNeil, M.; Richardson, C.; et al. Centenarians and Extremely Old People Living with Frailty Can Elicit Durable SARS-CoV-2 Spike Specific IgG Antibodies with Virus Neutralization Functions Following Virus Infection as Determined by Serological Study. eClinicalMedicine 2021, 37, 100975. [Google Scholar] [CrossRef]
- Seiffert, P.; Konka, A.; Kasperczyk, J.; Kawa, J.; Lejawa, M.; Maślanka-Seiffert, B.; Zembala-John, J.; Bugdol, M.; Romanik, M.; Bułdak, R.; et al. Immunogenicity of the BNT162b2 MRNA COVID-19 Vaccine in Older Residents of a Long-Term Care Facility: Relation with Age, Frailty and Prior Infection Status. Biogerontology 2022, 23, 53–64. [Google Scholar] [CrossRef] [PubMed]
- Salmerón Ríos, S.; Mas Romero, M.; Cortés Zamora, E.B.; Tabernero Sahuquillo, M.T.; Romero Rizos, L.; Sánchez-Jurado, P.M.; Sánchez-Nievas, G.; Señalada, J.J.B.; García Nogueras, I.; Estrella Cazalla, J. de D.; et al. Immunogenicity of the BNT162b2 Vaccine in Frail or Disabled Nursing Home Residents: COVID-A Study. J. Am. Geriatr. Soc. 2021, 69, 1441–1447. [Google Scholar] [CrossRef]
- Dyer, A.H.; Noonan, C.; McElheron, M.; Batten, I.; Reddy, C.; Connolly, E.; Pierpoint, R.; Murray, C.; Leonard, A.; Higgins, C.; et al. Previous SARS-CoV-2 Infection, Age, and Frailty Are Associated with 6-Month Vaccine-Induced Anti-Spike Antibody Titer in Nursing Home Residents. J. Am. Med. Dir. Assoc. 2022, 23, 434–439. [Google Scholar] [CrossRef] [PubMed]
- Shapiro, J.R.; Sitaras, I.; Park, H.S.; Aytenfisu, T.Y.; Caputo, C.; Li, M.; Lee, J.; Johnston, T.S.; Li, H.; Wouters, C.; et al. Association of Frailty, Age, and Biological Sex with Severe Acute Respiratory Syndrome Coronavirus 2 Messenger RNA Vaccine-Induced Immunity in Older Adults. Clin. Infect. Dis. Off. Publ. Infect. Dis. Soc. Am. 2022, 75, S61–S71. [Google Scholar] [CrossRef] [PubMed]
- Frailty and Age Impact Immune Responses to Moderna COVID-19 MRNA Vaccine. Available online: https://www.researchsquare.com (accessed on 18 August 2022).
- Ferrucci, L.; Fabbri, E. Inflammageing: Chronic Inflammation in Ageing, Cardiovascular Disease, and Frailty. Nat. Rev. Cardiol. 2018, 15, 505–522. [Google Scholar] [CrossRef] [PubMed]
- Rea, I.M.; Gibson, D.S.; McGilligan, V.; McNerlan, S.E.; Alexander, H.D.; Ross, O.A. Age and Age-Related Diseases: Role of Inflammation Triggers and Cytokines. Front. Immunol. 2018, 9, 586. [Google Scholar] [CrossRef]
- Dowell, A.C.; Butler, M.S.; Jinks, E.; Tut, G.; Lancaster, T.; Sylla, P.; Begum, J.; Bruton, R.; Pearce, H.; Verma, K.; et al. Children Develop Robust and Sustained Cross-Reactive Spike-Specific Immune Responses to SARS-CoV-2 Infection. Nat. Immunol. 2022, 23, 40–49. [Google Scholar] [CrossRef]
- Cӑtoi, A.F.; Corina, A.; Katsiki, N.; Vodnar, D.C.; Andreicuț, A.D.; Stoian, A.P.; Rizzo, M.; Pérez-Martínez, P. Gut Microbiota and Aging-A Focus on Centenarians. Biochim. Biophys. Acta BBA—Mol. Basis Dis. 2020, 1866, 165765. [Google Scholar] [CrossRef]
- Nagpal, R.; Mainali, R.; Ahmadi, S.; Wang, S.; Singh, R.; Kavanagh, K.; Kitzman, D.W.; Kushugulova, A.; Marotta, F.; Yadav, H. Gut Microbiome and Aging: Physiological and Mechanistic Insights. Nutr. Healthy Aging 2018, 4, 267–285. [Google Scholar] [CrossRef]
- Ghosh, S.; Lertwattanarak, R.; Garduño, J.d.J.; Galeana, J.J.; Li, J.; Zamarripa, F.; Lancaster, J.L.; Mohan, S.; Hussey, S.; Musi, N. Elevated Muscle TLR4 Expression and Metabolic Endotoxemia in Human Aging. J. Gerontol. Ser. A 2015, 70, 232–246. [Google Scholar] [CrossRef]
- Franceschi, C.; Garagnani, P.; Parini, P.; Giuliani, C.; Santoro, A. Inflammaging: A New Immune-Metabolic Viewpoint for Age-Related Diseases. Nat. Rev. Endocrinol. 2018, 14, 576–590. [Google Scholar] [CrossRef]
- Shintouo, C.M.; Mets, T.; Beckwee, D.; Bautmans, I.; Ghogomu, S.M.; Souopgui, J.; Leemans, L.; Meriki, H.D.; Njemini, R. Is Inflammageing Influenced by the Microbiota in the Aged Gut? A Systematic Review. Exp. Gerontol. 2020, 141, 111079. [Google Scholar] [CrossRef]
- Vaiserman, A.M.; Koliada, A.K.; Marotta, F. Gut Microbiota: A Player in Aging and a Target for Anti-Aging Intervention. Ageing Res. Rev. 2017, 35, 36–45. [Google Scholar] [CrossRef] [PubMed]
- MacSwain, K.L.H.; Sherry, S.B.; Stewart, S.H.; Watt, M.C.; Hadjistavropoulos, H.D.; Graham, A.R. Gender Differences in Health Anxiety: An Investigation of the Interpersonal Model of Health Anxiety. Personal. Individ. Differ. 2009, 47, 938–943. [Google Scholar] [CrossRef]
- Özdin, S.; Bayrak Özdin, Ş. Levels and Predictors of Anxiety, Depression and Health Anxiety during COVID-19 Pandemic in Turkish Society: The Importance of Gender. Int. J. Soc. Psychiatry 2020, 66, 504–511. [Google Scholar] [CrossRef] [PubMed]
- Hiza, H.A.B.; Casavale, K.O.; Guenther, P.M.; Davis, C.A. Diet Quality of Americans Differs by Age, Sex, Race/Ethnicity, Income, and Education Level. J. Acad. Nutr. Diet. 2013, 113, 297–306. [Google Scholar] [CrossRef]
- Pivonello, R.; Auriemma, R.S.; Pivonello, C.; Isidori, A.M.; Corona, G.; Colao, A.; Millar, R.P. Sex Disparities in COVID-19 Severity and Outcome: Are Men Weaker or Women Stronger? Neuroendocrinology 2021, 111, 1066–1085. [Google Scholar] [CrossRef]
- Thompson, A.E.; Anisimowicz, Y.; Miedema, B.; Hogg, W.; Wodchis, W.P.; Aubrey-Bassler, K. The Influence of Gender and Other Patient Characteristics on Health Care-Seeking Behaviour: A QUALICOPC Study. BMC Fam. Pract. 2016, 17, 38. [Google Scholar] [CrossRef]
- Solomou, I.; Constantinidou, F. Prevalence and Predictors of Anxiety and Depression Symptoms during the COVID-19 Pandemic and Compliance with Precautionary Measures: Age and Sex Matter. Int. J. Environ. Res. Public. Health 2020, 17, 4924. [Google Scholar] [CrossRef]
- Tharakan, T.; Khoo, C.C.; Giwercman, A.; Jayasena, C.N.; Sofikitis, N.; Salonia, A.; Minhas, S. Are Sex Disparities in COVID-19 a Predictable Outcome of Failing Men’s Health Provision? Nat. Rev. Urol. 2022, 19, 47–63. [Google Scholar] [CrossRef]
- Cai, H. Sex Difference and Smoking Predisposition in Patients with COVID-19. Lancet Respir. Med. 2020, 8, e20. [Google Scholar] [CrossRef]
- Wilsnack, R.W.; Vogeltanz, N.D.; Wilsnack, S.C.; Harris, T.R. Gender Differences in Alcohol Consumption and Adverse Drinking Consequences: Cross-Cultural Patterns. Addiction 2000, 95, 251–265. [Google Scholar] [CrossRef]
- Aryal, S.; Diaz-Guzman, E.; Mannino, D.M. COPD and Gender Differences: An Update. Transl. Res. 2013, 162, 208–218. [Google Scholar] [CrossRef] [PubMed]
- Kanter, R.; Caballero, B. Global Gender Disparities in Obesity: A Review. Adv. Nutr. 2012, 3, 491–498. [Google Scholar] [CrossRef] [PubMed]
- Mauvais-Jarvis, F. Gender Differences in Glucose Homeostasis and Diabetes. Physiol. Behav. 2018, 187, 20–23. [Google Scholar] [CrossRef] [PubMed]
- Song, J.-J.; Ma, Z.; Wang, J.; Chen, L.-X.; Zhong, J.-C. Gender Differences in Hypertension. J Cardiovasc. Transl. Res. 2020, 13, 47–54. [Google Scholar] [CrossRef] [PubMed]
- Thom, T.; Haase, N.; Rosamond, W.; Howard, V.J.; Rumsfeld, J.; Manolio, T.; Zheng, Z.-J.; Flegal, K.; O’Donnell, C.; Kittner, S.; et al. Heart Disease and Stroke Statistics--2006 Update: A Report from the American Heart Association Statistics Committee and Stroke Statistics Subcommittee. Circulation 2006, 113, e85–e151. [Google Scholar] [CrossRef]
- Wray, S.; Arrowsmith, S. The Physiological Mechanisms of the Sex-Based Difference in Outcomes of COVID19 Infection. Front. Physiol. 2021, 12, 627260. [Google Scholar] [CrossRef]
- Jazwinski, S.M. Longevity, Genes, and Aging. Science 1996, 273, 54–59. [Google Scholar] [CrossRef]
- Brooks-Wilson, A.R. Genetics of Healthy Aging and Longevity. Hum. Genet. 2013, 132, 1323–1338. [Google Scholar] [CrossRef]
- Forgetta, V.; Manousaki, D.; Istomine, R.; Ross, S.; Tessier, M.-C.; Marchand, L.; Li, M.; Qu, H.-Q.; Bradfield, J.P.; Grant, S.F.A.; et al. Rare Genetic Variants of Large Effect Influence Risk of Type 1 Diabetes. Diabetes 2020, 69, 784–795. [Google Scholar] [CrossRef]
- Hyttinen, V.; Kaprio, J.; Kinnunen, L.; Koskenvuo, M.; Tuomilehto, J. Genetic Liability of Type 1 Diabetes and the Onset Age among 22,650 Young Finnish Twin Pairs: A Nationwide Follow-Up Study. Diabetes 2003, 52, 1052–1055. [Google Scholar] [CrossRef]
- Diabetes Genetics Initiative of Broad Institute of Harvard and MIT, Lund University, and Novartis Institutes of BioMedical Research; Saxena, R.; Voight, B.F.; Lyssenko, V.; Burtt, N.P.; de Bakker, P.I.W.; Chen, H.; Roix, J.J.; Kathiresan, S.; Hirschhorn, J.N.; et al. Genome-Wide Association Analysis Identifies Loci for Type 2 Diabetes and Triglyceride Levels. Science 2007, 316, 1331–1336. [Google Scholar] [CrossRef]
- Scott, L.J.; Mohlke, K.L.; Bonnycastle, L.L.; Willer, C.J.; Li, Y.; Duren, W.L.; Erdos, M.R.; Stringham, H.M.; Chines, P.S.; Jackson, A.U.; et al. A Genome-Wide Association Study of Type 2 Diabetes in Finns Detects Multiple Susceptibility Variants. Science 2007, 316, 1341–1345. [Google Scholar] [CrossRef]
- Walley, A.J.; Asher, J.E.; Froguel, P. The Genetic Contribution to Non-Syndromic Human Obesity. Nat. Rev. Genet. 2009, 10, 431–442. [Google Scholar] [CrossRef]
- Yang, W.; Kelly, T.; He, J. Genetic Epidemiology of Obesity. Epidemiol. Rev. 2007, 29, 49–61. [Google Scholar] [CrossRef] [PubMed]
- Keating, M.T.; Sanguinetti, M.C. Molecular Genetic Insights into Cardiovascular Disease. Science 1996, 272, 681–685. [Google Scholar] [CrossRef]
- Genetic Variants in Novel Pathways Influence Blood Pressure and Cardiovascular Disease Risk. Nature 2011, 478, 103–109. [CrossRef] [PubMed]
- Kim, D.-K.; Weller, B.; Lin, C.-W.; Sheykhkarimli, D.; Knapp, J.J.; Dugied, G.; Zanzoni, A.; Pons, C.; Tofaute, M.J.; Maseko, S.B.; et al. A Proteome-Scale Map of the SARS-CoV-2–Human Contactome. Nat. Biotechnol. 2022, 1–10. [Google Scholar] [CrossRef] [PubMed]
- Planas, D.; Saunders, N.; Maes, P.; Guivel-Benhassine, F.; Planchais, C.; Buchrieser, J.; Bolland, W.-H.; Porrot, F.; Staropoli, I.; Lemoine, F.; et al. Considerable Escape of SARS-CoV-2 Omicron to Antibody Neutralization. Nature 2022, 602, 671–675. [Google Scholar] [CrossRef] [PubMed]
- Satterfield, B.A.; Bhatt, D.L.; Gersh, B.J. Cardiac Involvement in the Long-Term Implications of COVID-19. Nat. Rev. Cardiol. 2022, 19, 332–341. [Google Scholar] [CrossRef] [PubMed]
- Xie, Y.; Xu, E.; Bowe, B.; Al-Aly, Z. Long-Term Cardiovascular Outcomes of COVID-19. Nat. Med. 2022, 28, 583–590. [Google Scholar] [CrossRef] [PubMed]
- Liao, T.; Meng, D.; Xiong, L.; Wu, S.; Yang, L.; Wang, S.; Zhou, M.; He, X.; Cao, X.; Xiong, H.; et al. Long-Term Effects of COVID-19 on Health Care Workers 1-Year Post-Discharge in Wuhan. Infect. Dis. Ther. 2022, 11, 145–163. [Google Scholar] [CrossRef] [PubMed]
- Desai, A.D.; Lavelle, M.; Boursiquot, B.C.; Wan, E.Y. Long-Term Complications of COVID-19. Am. J. Physiol.-Cell Physiol. 2022, 322, C1–C11. [Google Scholar] [CrossRef]
- Meyer, P.T.; Hellwig, S.; Blazhenets, G.; Hosp, J.A. Molecular Imaging Findings on Acute and Long-Term Effects of COVID-19 on the Brain: A Systematic Review. J. Nucl. Med. 2022, 63, 971–980. [Google Scholar] [CrossRef]
- Mehandru, S.; Merad, M. Pathological Sequelae of Long-Haul COVID. Nat. Immunol. 2022, 23, 194–202. [Google Scholar] [CrossRef]
- Taquet, M.; Sillett, R.; Zhu, L.; Mendel, J.; Camplisson, I.; Dercon, Q.; Harrison, P.J. Neurological and Psychiatric Risk Trajectories after SARS-CoV-2 Infection: An Analysis of 2-Year Retrospective Cohort Studies Including 1 284 437 Patients. Lancet Psychiatry 2022, 9, 815–827. [Google Scholar] [CrossRef]
- Unnikrishnan, R.; Misra, A. Diabetes and COVID19: A Bidirectional Relationship. Eur. J. Clin. Nutr. 2021, 75, 1332–1336. [Google Scholar] [CrossRef]
- Rey-Reñones, C.; Martinez-Torres, S.; Martín-Luján, F.M.; Pericas, C.; Redondo, A.; Vilaplana-Carnerero, C.; Dominguez, A.; Grau, M. Type 2 Diabetes Mellitus and COVID-19: A Narrative Review. Biomedicines 2022, 10, 2089. [Google Scholar] [CrossRef]
- Hsu, C. Yes, AKI Truly Leads to CKD. J. Am. Soc. Nephrol. 2012, 23, 967–969. [Google Scholar] [CrossRef] [PubMed]
- Al-Aly, Z.; Xie, Y.; Bowe, B. High-Dimensional Characterization of Post-Acute Sequelae of COVID-19. Nature 2021, 594, 259–264. [Google Scholar] [CrossRef] [PubMed]
- Raman, B.; Bluemke, D.A.; Lüscher, T.F.; Neubauer, S. Long COVID: Post-Acute Sequelae of COVID-19 with a Cardiovascular Focus. Eur. Heart J. 2022, 43, 1157–1172. [Google Scholar] [CrossRef]
- Carrillo-Garcia, P.; Garmendia-Prieto, B.; Cristofori, G.; Montoya, I.L.; Hidalgo, J.J.; Feijoo, M.Q.; Cortés, J.J.B.; Gómez-Pavón, J. Health Status in Survivors Older than 70 Years after Hospitalization with COVID-19: Observational Follow-up Study at 3 Months. Eur. Geriatr. Med. 2021, 12, 1091–1094. [Google Scholar] [CrossRef] [PubMed]
- Greco, G.I.; Noale, M.; Trevisan, C.; Zatti, G.; Dalla Pozza, M.; Lazzarin, M.; Haxhiaj, L.; Ramon, R.; Imoscopi, A.; Bellon, S.; et al. Increase in Frailty in Nursing Home Survivors of Coronavirus Disease 2019: Comparison with Noninfected Residents. J. Am. Med. Dir. Assoc. 2021, 22, 943–947. [Google Scholar] [CrossRef] [PubMed]
- Taniguchi, L.U.; Avelino-Silva, T.J.; Dias, M.B.; Jacob-Filho, W.; Aliberti, M.J.R.; on behalf of COVID-19 and Frailty (CO-FRAIL) Study Group and EPIdemiology of Critical COVID-19 (EPICCoV) Study Group, for COVID Hospital das Clinicas, University of Sao Paulo Medical School (HCFMUSP) Study Group. Patient-Centered Outcomes Following COVID-19: Frailty and Disability Transitions in Critical Care Survivors. Crit. Care Med. 2022, 50, 955–963. [Google Scholar] [CrossRef]
- Stockwell, S.; Trott, M.; Tully, M.; Shin, J.; Barnett, Y.; Butler, L.; McDermott, D.; Schuch, F.; Smith, L. Changes in Physical Activity and Sedentary Behaviours from before to during the COVID-19 Pandemic Lockdown: A Systematic Review. BMJ Open Sport Exerc. Med. 2021, 7, e000960. [Google Scholar] [CrossRef]
- Bu, F.; Bone, J.K.; Mitchell, J.J.; Steptoe, A.; Fancourt, D. Longitudinal Changes in Physical Activity during and after the First National Lockdown Due to the COVID-19 Pandemic in England. Sci. Rep. 2021, 11, 17723. [Google Scholar] [CrossRef]
- Ross, J.A.; Malone, P.K.; Levy, S. The Impact of the Severe Acute Respiratory Syndrome Coronavirus 2 (SARS-CoV-2) Pandemic on Substance Use in the United States. Clin. Infect. Dis. 2022, 75, S81–S85. [Google Scholar] [CrossRef]
- Calina, D.; Hartung, T.; Mardare, I.; Mitroi, M.; Poulas, K.; Tsatsakis, A.; Rogoveanu, I.; Docea, A.O. COVID-19 Pandemic and Alcohol Consumption: Impacts and Interconnections. Toxicol. Rep. 2021, 8, 529–535. [Google Scholar] [CrossRef]
- Sallie, S.N.; Ritou, V.; Bowden-Jones, H.; Voon, V. Assessing International Alcohol Consumption Patterns during Isolation from the COVID-19 Pandemic Using an Online Survey: Highlighting Negative Emotionality Mechanisms. BMJ Open 2020, 10, e044276. [Google Scholar] [CrossRef]
- McPhee, M.D.; Keough, M.T.; Rundle, S.; Heath, L.M.; Wardell, J.D.; Hendershot, C.S. Depression, Environmental Reward, Coping Motives and Alcohol Consumption During the COVID-19 Pandemic. Front. Psychiatry 2020, 11, 574676. [Google Scholar] [CrossRef] [PubMed]
- Josephson, A.; Kilic, T.; Michler, J.D. Socioeconomic Impacts of COVID-19 in Low-Income Countries. Nat. Hum. Behav. 2021, 5, 557–565. [Google Scholar] [CrossRef]
- Hamadani, J.D.; Hasan, M.I.; Baldi, A.J.; Hossain, S.J.; Shiraji, S.; Bhuiyan, M.S.A.; Mehrin, S.F.; Fisher, J.; Tofail, F.; Tipu, S.M.M.U.; et al. Immediate Impact of Stay-at-Home Orders to Control COVID-19 Transmission on Socioeconomic Conditions, Food Insecurity, Mental Health, and Intimate Partner Violence in Bangladeshi Women and Their Families: An Interrupted Time Series. Lancet Glob. Health 2020, 8, e1380–e1389. [Google Scholar] [CrossRef] [PubMed]
- Faghri, P.D.; Dobson, M.; Landsbergis, P.; Schnall, P.L. COVID-19 Pandemic: What Has Work Got to Do with It? J. Occup. Environ. Med. 2021, 63, e245. [Google Scholar] [CrossRef] [PubMed]
- Qian, Y.; Fan, W. Who Loses Income during the COVID-19 Outbreak? Evidence from China. Res. Soc. Stratif. Mobil. 2020, 68, 100522. [Google Scholar] [CrossRef]
- Picchioni, F.; Goulao, L.F.; Roberfroid, D. The Impact of COVID-19 on Diet Quality, Food Security and Nutrition in Low and Middle Income Countries: A Systematic Review of the Evidence. Clin. Nutr. 2021, 43, 2955–2964. [Google Scholar] [CrossRef]
- Krumer-Nevo, M.; Refaeli, T. COVID-19: A Poverty-Aware Perspective. Am. J. Orthopsychiatr. 2021, 91, 423–431. [Google Scholar] [CrossRef]
- Madai Boukar, A.; Mbock, O.; Kilolo, J.-M.M. The Impacts of the COVID-19 Pandemic on Employment in Cameroon: A General Equilibrium Analysis. Afr. Dev. Rev. 2021, 33, S88–S101. [Google Scholar] [CrossRef]
- Selden, T.M.; Berdahl, T.A. COVID-19 And Racial/Ethnic Disparities In Health Risk, Employment, And Household Composition: Study Examines Potential Explanations for Racial-Ethnic Disparities in COVID-19 Hospitalizations and Mortality. Health Aff. 2020, 39, 1624–1632. [Google Scholar] [CrossRef]
- Béné, C.; Bakker, D.; Chavarro, M.J.; Even, B.; Melo, J.; Sonneveld, A. Global Assessment of the Impacts of COVID-19 on Food Security. Glob. Food Secur. 2021, 31, 100575. [Google Scholar] [CrossRef]
- Agberotimi, S.F.; Akinsola, O.S.; Oguntayo, R.; Olaseni, A.O. Interactions Between Socioeconomic Status and Mental Health Outcomes in the Nigerian Context Amid COVID-19 Pandemic: A Comparative Study. Front. Psychol. 2020, 11, 559819. [Google Scholar] [CrossRef]
- Iob, E.; Frank, P.; Steptoe, A.; Fancourt, D. Levels of Severity of Depressive Symptoms among At-Risk Groups in the UK During the COVID-19 Pandemic. JAMA Netw. Open 2020, 3, e2026064. [Google Scholar] [CrossRef]
- Kikuchi, H.; Machida, M.; Nakamura, I.; Saito, R.; Odagiri, Y.; Kojima, T.; Watanabe, H.; Inoue, S. Development of Severe Psychological Distress among Low-Income Individuals during the COVID-19 Pandemic: Longitudinal Study. BJPsych Open 2021, 7, e50. [Google Scholar] [CrossRef] [PubMed]
- Fang, D.; Thomsen, M.R.; Nayga, R.M. The Association between Food Insecurity and Mental Health during the COVID-19 Pandemic. BMC Public Health 2021, 21, 607. [Google Scholar] [CrossRef] [PubMed]
- Naylor-Wardle, J.; Rowland, B.; Kunadian, V. Socioeconomic Status and Cardiovascular Health in the COVID-19 Pandemic. Heart 2021, 107, 358–365. [Google Scholar] [CrossRef] [PubMed]
- George, A.; Stead, T.S.; Ganti, L. What’s the Risk: Differentiating Risk Ratios, Odds Ratios, and Hazard Ratios? Cureus 2020, 12, e10047. [Google Scholar] [CrossRef]
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2023 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).