Novel Coronavirus and Common Pneumonia Detection from CT Scans Using Deep Learning-Based Extracted Features
Abstract
:1. Introduction
2. Recent Studies
3. Deep Convolutional Neural Networks
4. Experimental Dataset
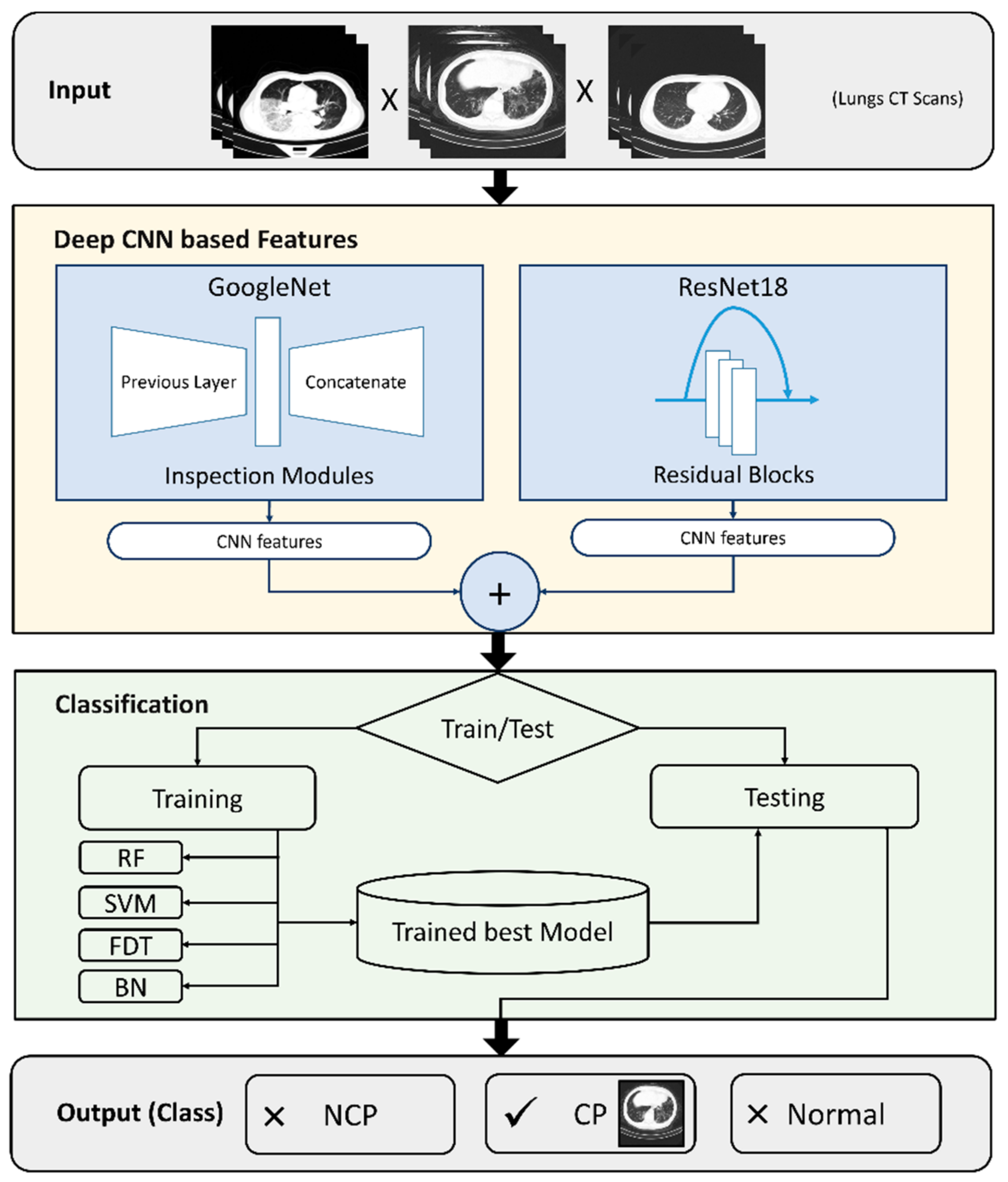
5. Method
5.1. Deep Learning Features
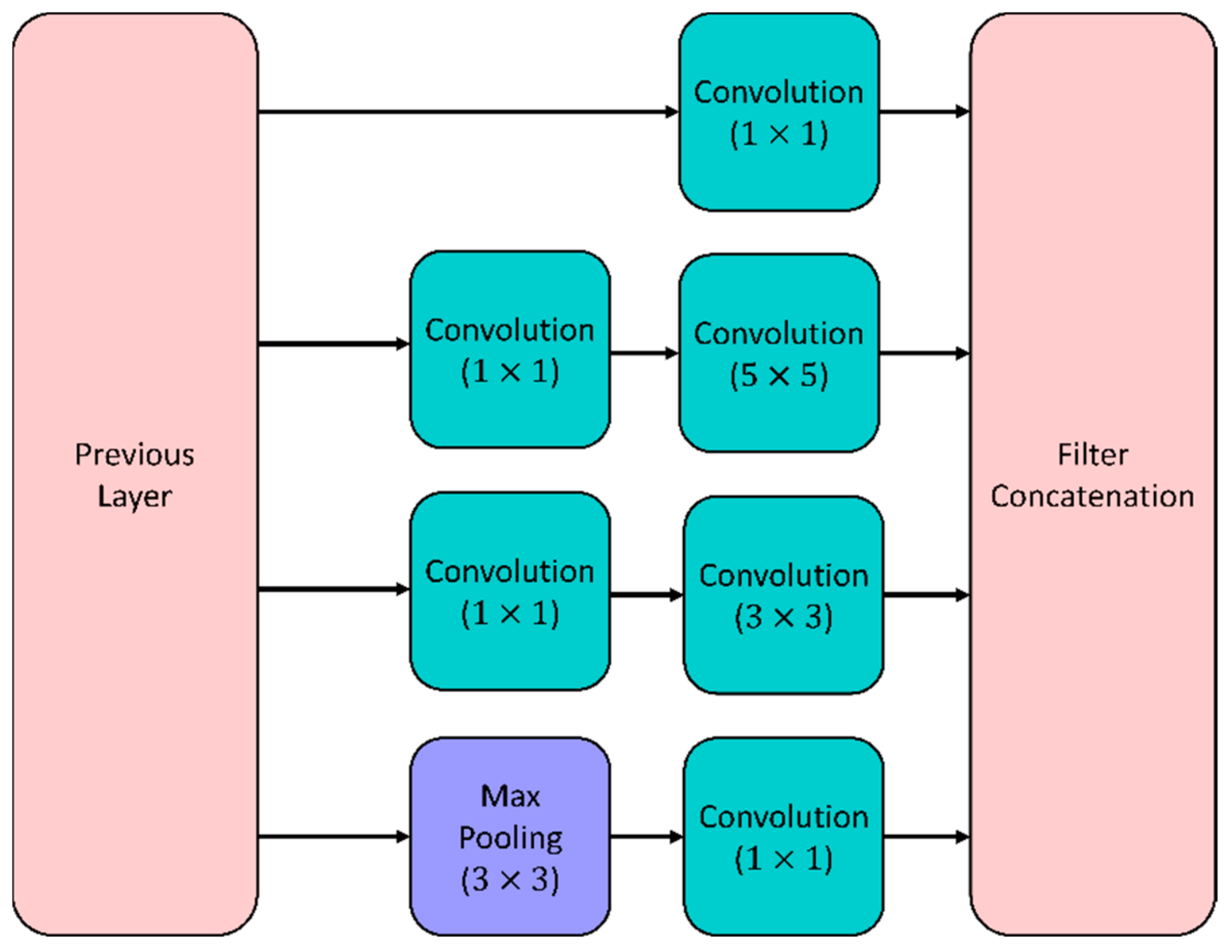
5.1.1. GoogleNet
5.1.2. ResNet18
5.2. Classification
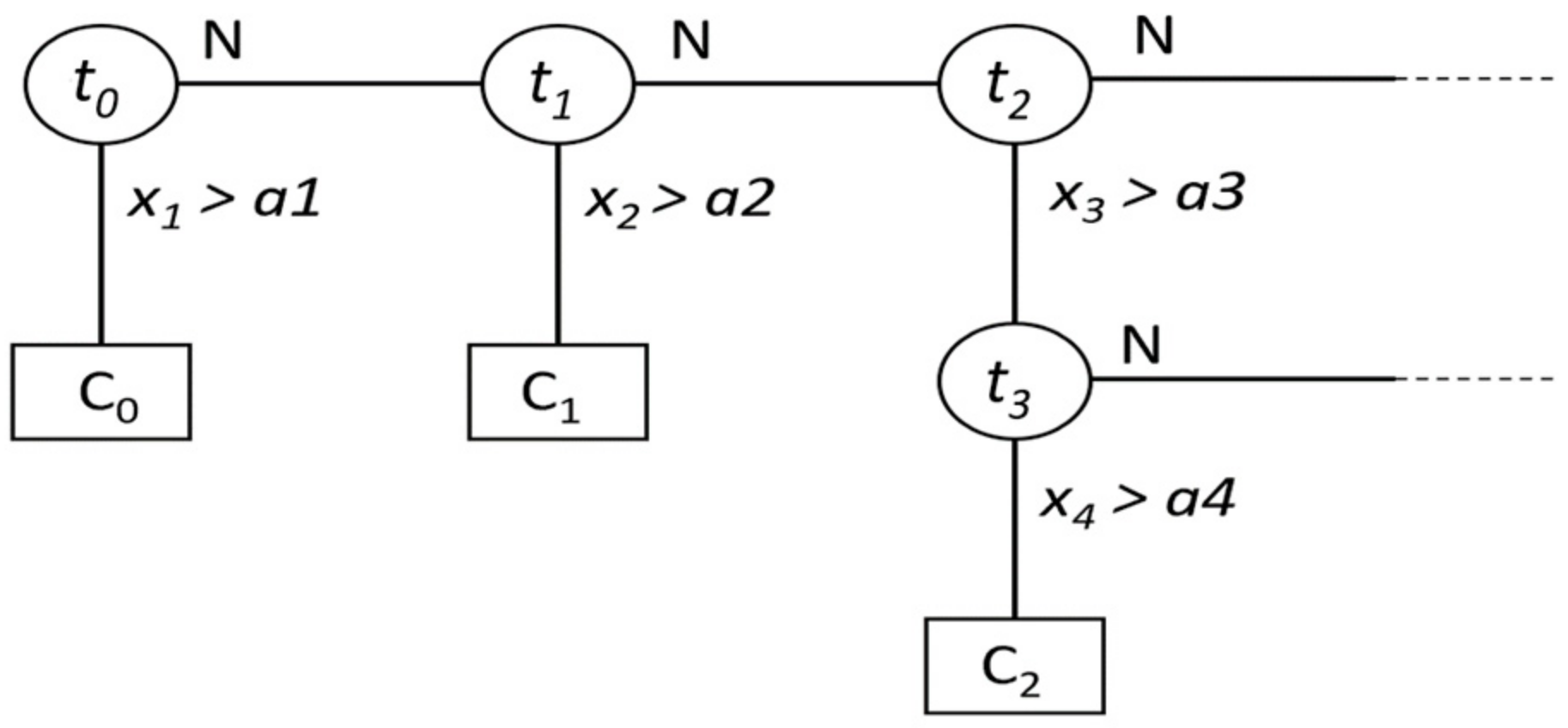
5.2.1. Fast Decision Tree (FDT)
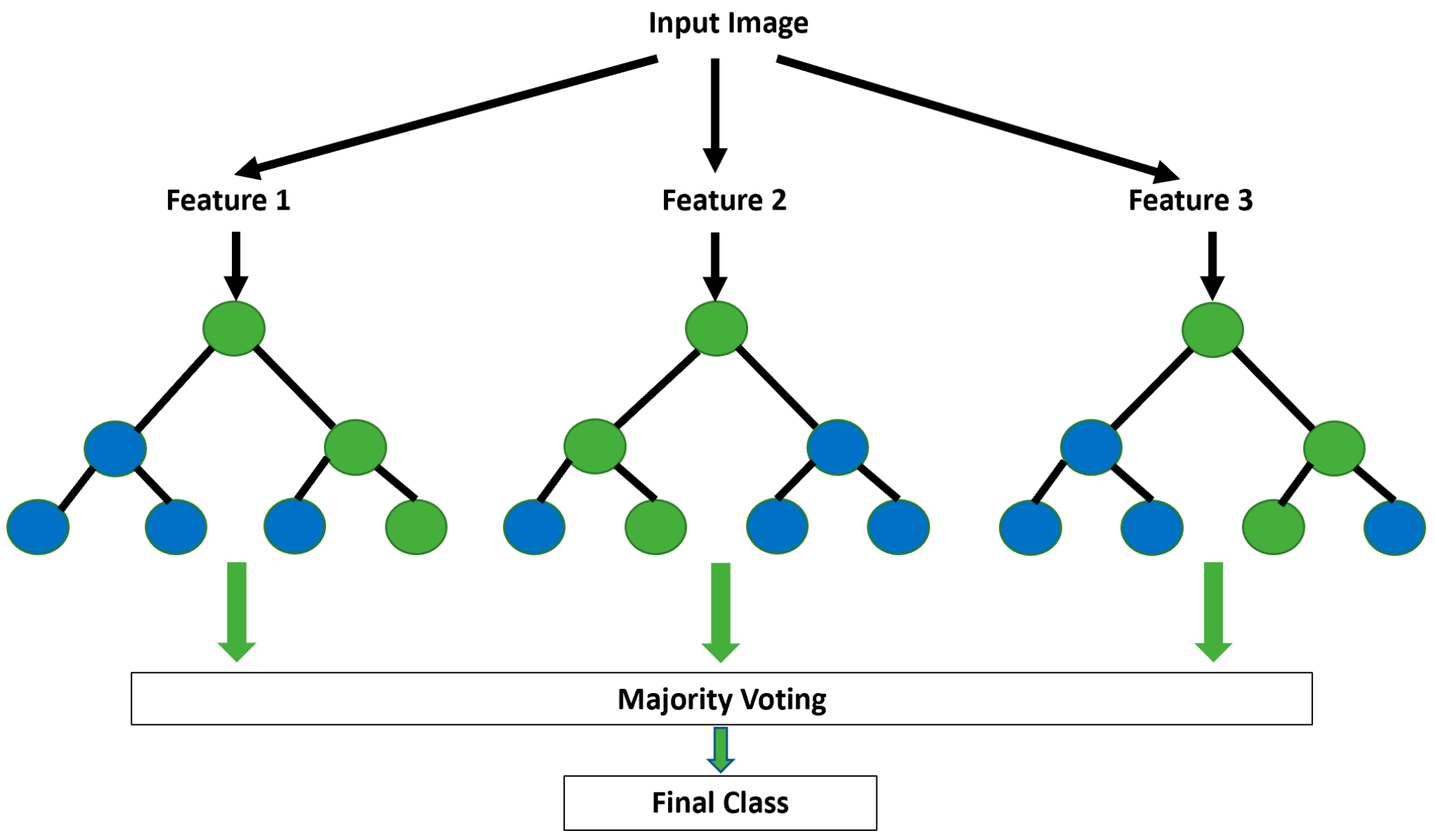
5.2.2. Random Forest (RF)
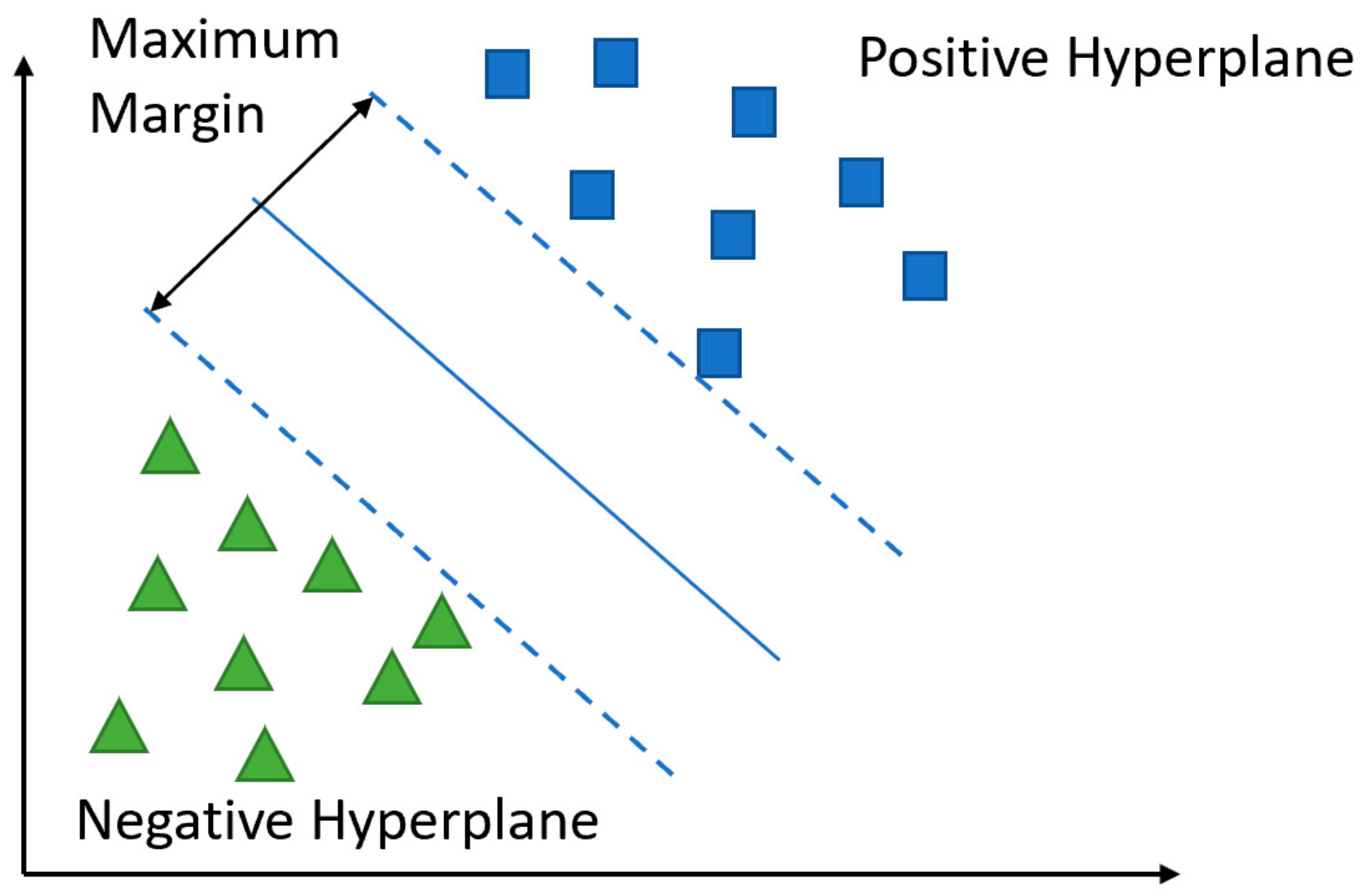
5.2.3. Support Vector Machine (SVM)
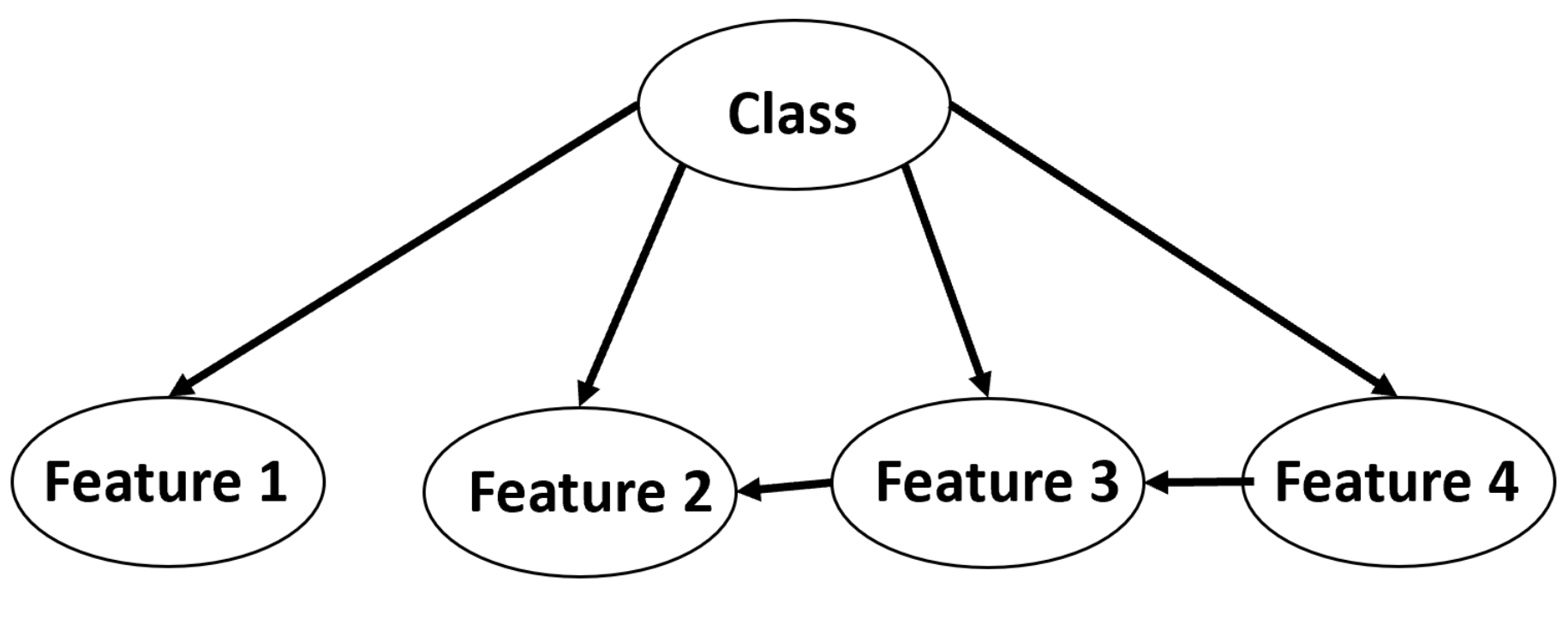
5.2.4. Bayesian Network (BN)
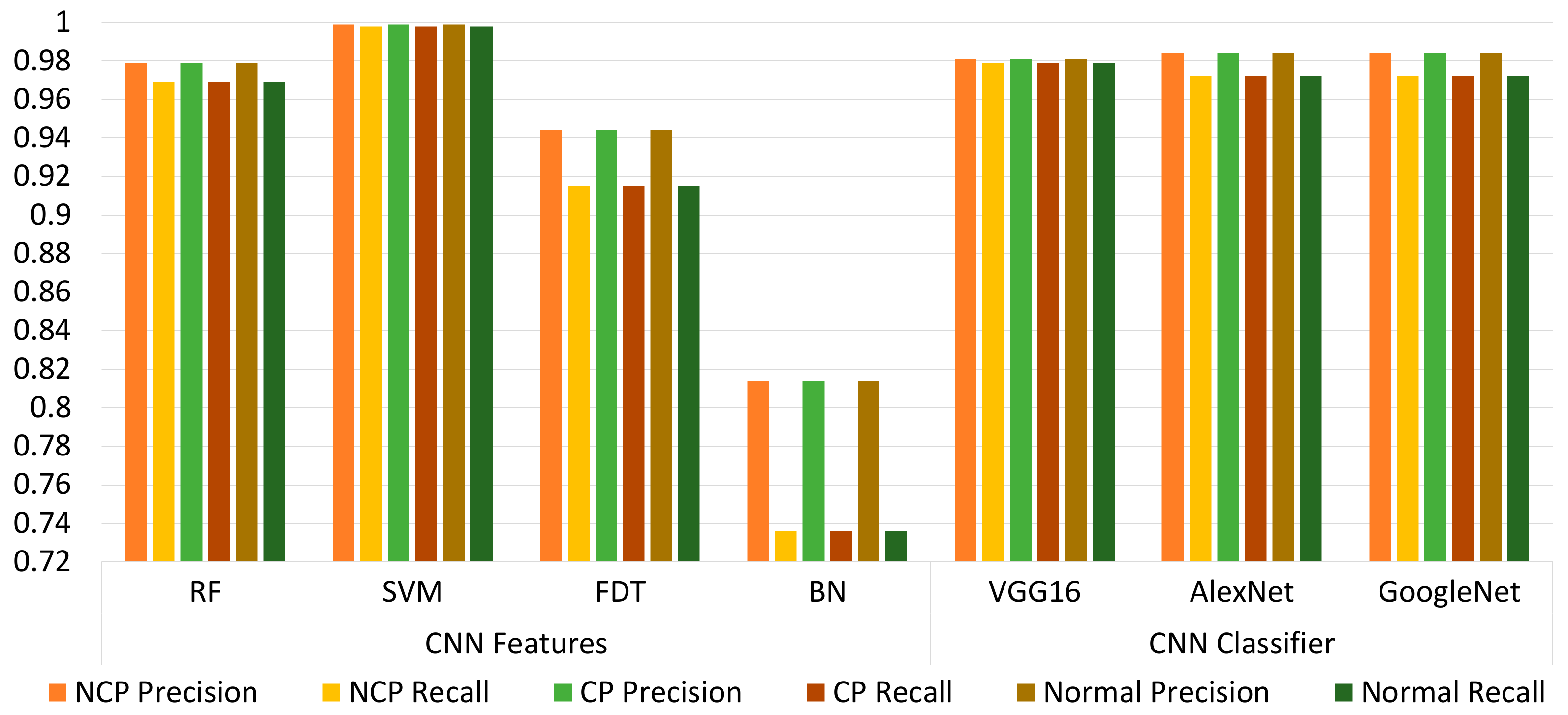
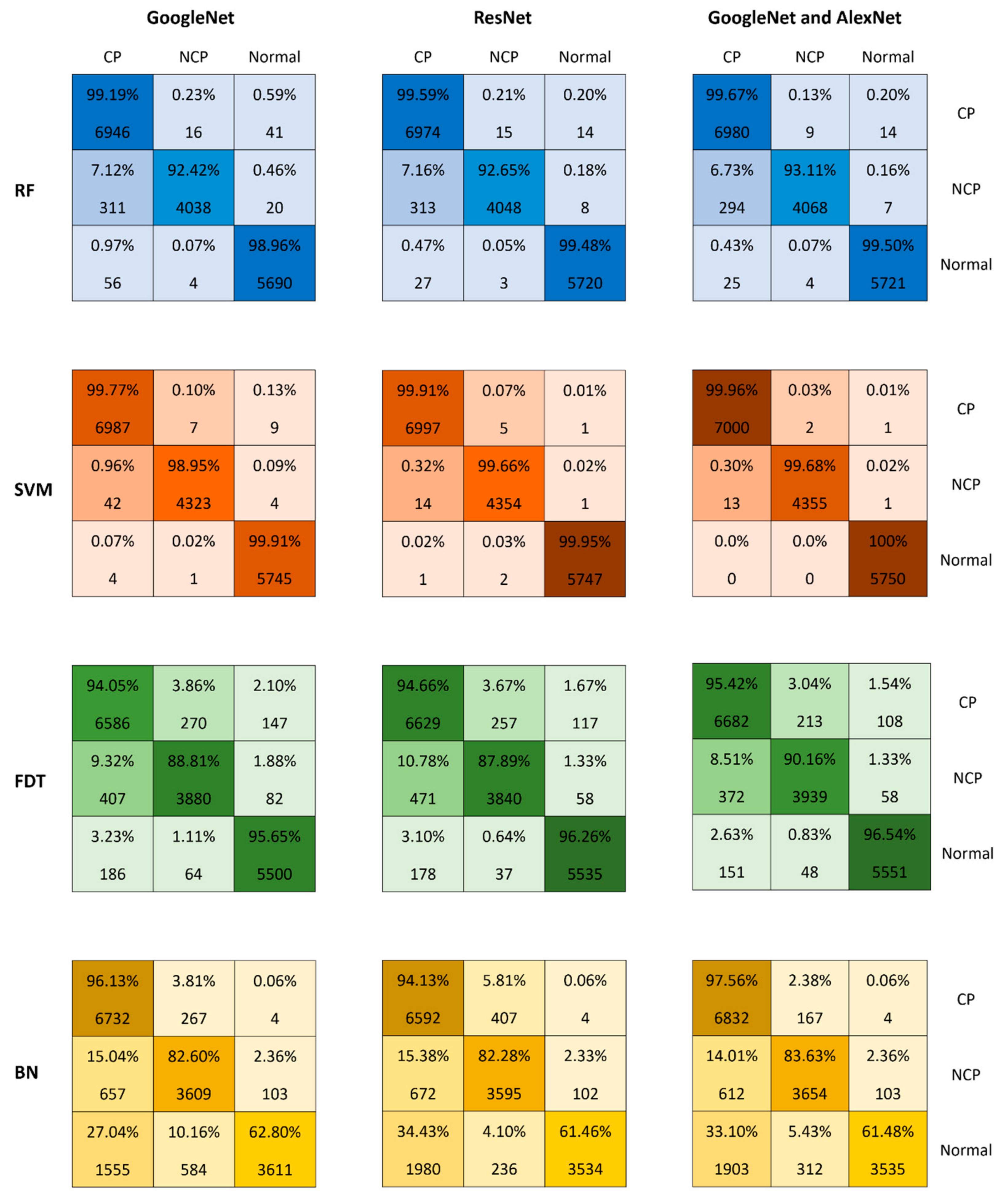
6. Results
7. Discussion
8. Conclusions
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Acknowledgments
Conflicts of Interest
References
- Narin, A.; Kaya, C.; Pamuk, Z. Automatic Detection of Coronavirus Disease (COVID-19) Using X-Ray Images and Deep Convolutional Neural Networks. Pattern Anal. Appl. 2021, 24, 1207–1220. [Google Scholar] [CrossRef] [PubMed]
- Roosa, K.; Lee, Y.; Luo, R.; Kirpich, A.; Rothenberg, R.; Hyman, J.M.; Yan, P.; Chowell, G. Real-Time Forecasts of the COVID-19 Epidemic in China from February 5th to February 24th, 2020. Infect. Dis. Model. 2020, 5, 256–263. [Google Scholar] [CrossRef] [PubMed]
- Wang, L.-F.; Shi, Z.; Zhang, S.; Field, H.; Daszak, P.; Eaton, B. Review of Bats and SARS. Emerg. Infect. Dis. 2006, 12, 1834–1840. [Google Scholar] [CrossRef] [PubMed]
- Mahase, E. Coronavirus: Covid-19 Has Killed More People than SARS and MERS Combined, despite Lower Case Fatality Rate. BMJ 2020, m641. [Google Scholar] [CrossRef] [Green Version]
- Covid-19 Pandemic Statistics. Available online: https://www.worldometers.info/coronavirus/ (accessed on 6 April 2022).
- Bai, Y.; Yao, L.; Wei, T.; Tian, F.; Jin, D.-Y.; Chen, L.; Wang, M. Presumed Asymptomatic Carrier Transmission of COVID-19. JAMA 2020, 323, 1406. [Google Scholar] [CrossRef] [Green Version]
- Vaishya, R.; Javaid, M.; Khan, I.H.; Haleem, A. Artificial Intelligence (AI) Applications for COVID-19 Pandemic. Diabetes Metab. Syndr. Clin. Res. Rev. 2020, 14, 337–339. [Google Scholar] [CrossRef]
- Cohen, J.P.; Morrison, P.; Dao, L. COVID-19 Image Data Collection. arXiv 2020, arXiv:2003.11597. [Google Scholar]
- Tabik, S.; Gomez-Rios, A.; Martin-Rodriguez, J.L.; Sevillano-Garcia, I.; Rey-Area, M.; Charte, D.; Guirado, E.; Suarez, J.L.; Luengo, J.; Valero-Gonzalez, M.A.; et al. COVIDGR Dataset and COVID-SDNet Methodology for Predicting COVID-19 Based on Chest X-ray Images. IEEE J. Biomed. Health Inform. 2020, 24, 3595–3605. [Google Scholar] [CrossRef]
- Radiological Society of North America (RSNA) Pneumonia Detection Challenge Dataset. Available online: https://www.kaggle.com/c/snapneumonia-detection-challenge/data (accessed on 6 April 2022).
- Wang, X.; Peng, Y.; Lu, L.; Lu, Z.; Bagheri, M.; Summers, R.M. ChestX-ray8: Hospital-Scale Chest X-ray Database and Benchmarks on Weakly-Supervised Classification and Localization of Common Thorax Diseases. In Proceedings of the IEEE Conference on Computer Vision and Pattern Recognition, Honolulu, HI, USA, 21–26 July 2017; pp. 2097–2106. [Google Scholar]
- Alistair, J.; Pollard, T.J.; Greenbaum, N.R.; Lungren, M.P.; Deng, C.; Peng, Y.; Lu, Z.; Mark, R.G.; Berkowitz, S.J.; Horng, S. MIMIC-CXR-JPG, a Large Publicly Available Database of Labeled Chest Radiographs. arXiv 2019, arXiv:1901.07042. [Google Scholar]
- Bustos, A.; Pertusa, A.; Salinas, J.-M.; de la Iglesia-Vayá, M. PadChest: A Large Chest X-ray Image Dataset with Multi-Label Annotated Reports. Med. Image Anal. 2020, 66, 101797. [Google Scholar] [CrossRef]
- Lamsal, R. Design and Analysis of a Large-Scale COVID-19 Tweets Dataset. Appl. Intell. 2020, 51, 2790–2804. [Google Scholar] [CrossRef]
- Mbunge, E.; Simelane, S.; Fashoto, S.G.; Akinnuwesi, B.; Metfula, A.S. Application of Deep Learning and Machine Learning Models to Detect COVID-19 Face Masks—A Review. Sustain. Oper. Comput. 2021, 2, 235–245. [Google Scholar] [CrossRef]
- Horry, M.J.; Chakraborty, S.; Paul, M.; Ulhaq, A.; Pradhan, B.; Saha, M.; Shukla, N. COVID-19 Detection through Transfer Learning Using Multimodal Imaging Data. IEEE Access 2020, 8, 149808–149824. [Google Scholar] [CrossRef]
- Cohen, J.P.; Morrison, P.; Dao, L.; Roth, K.; Duong, T.Q.; Ghassemi, M. COVID-19 Image Data Collection: Prospective Predictions Are the Future. arXiv 2020, arXiv:2006.11988. [Google Scholar]
- Ismael, A.M.; Şengür, A. Deep Learning Approaches for COVID-19 Detection Based on Chest X-ray Images. Expert Syst. Appl. 2021, 164, 114054. [Google Scholar] [CrossRef]
- Alazab, M.; Awajan, A.; Mesleh, A.; Abraham, A.; Jatana, V.; Alhyari, S. COVID-19 Prediction and Detection Using Deep Learning. Int. J. Comput. Inf. Syst. Ind. Manag. Appl. 2020, 12, 168–181. [Google Scholar]
- Mangal, A.; Kalia, S.; Rajgopal, H.; Rangarajan, K.; Namboodiri, V.; Banerjee, S.; Arora, C. CovidAID: COVID-19 Detection Using Chest X-ray. arXiv 2020, arXiv:2004.09803. [Google Scholar]
- Alom, M.Z.; Rahman, M.M.; Nasrin, M.S.; Taha, T.M.; Asari, V.K. COVID_MTNet: COVID-19 Detection with Multi-Task Deep Learning Approaches. arXiv 2020, arXiv:2004.03747. [Google Scholar]
- Waheed, A.; Goyal, M.; Gupta, D.; Khanna, A.; Al-Turjman, F.; Pinheiro, P.R. CovidGAN: Data Augmentation Using Auxiliary Classifier GAN for Improved Covid-19 Detection. IEEE Access 2020, 8, 91916–91923. [Google Scholar] [CrossRef]
- Hussain, E.; Hasan, M.; Rahman, M.A.; Lee, I.; Tamanna, T.; Parvez, M.Z. CoroDet: A Deep Learning Based Classification for COVID-19 Detection Using Chest X-ray Images. Chaos Solitons Fractals 2021, 142, 110495. [Google Scholar] [CrossRef]
- Jain, R.; Gupta, M.; Taneja, S.; Hemanth, D.J. Deep Learning Based Detection and Analysis of COVID-19 on Chest X-ray Images. Appl. Intell. 2020, 51, 1690–1700. [Google Scholar] [CrossRef]
- Alshazly, H.; Linse, C.; Barth, E.; Martinetz, T. Explainable COVID-19 Detection Using Chest CT Scans and Deep Learning. Sensors 2021, 21, 455. [Google Scholar] [CrossRef]
- Alam, N.-A.-A.; Ahsan, M.; Based, M.A.; Haider, J.; Kowalski, M. COVID-19 Detection from Chest X-ray Images Using Feature Fusion and Deep Learning. Sensors 2021, 21, 1480. [Google Scholar] [CrossRef]
- Bashar, A.; Latif, G.; Ben Brahim, G.; Mohammad, N.; Alghazo, J. COVID-19 Pneumonia Detection Using Optimized Deep Learning Techniques. Diagnostics 2021, 11, 1972. [Google Scholar] [CrossRef]
- Subramanian, N.; Elharrouss, O.; Al-Maadeed, S.; Chowdhury, M. A Review of Deep Learning-Based Detection Methods for COVID-19. Comput. Biol. Med. 2022, 143, 105233. [Google Scholar] [CrossRef]
- Das, D.; Biswas, S.K.; Bandyopadhyay, S. Perspective of AI System for COVID-19 Detection Using Chest Images: A Review. Multimed. Tools Appl. 2022, 81, 21471–21501. [Google Scholar] [CrossRef]
- Latif, G.; Iskandar, D.N.F.A.; Alghazo, J.; Jaffar, A. Improving Brain MR Image Classification for Tumor Segmentation Using Phase Congruency. Curr. Med. Imaging Rev. 2018, 14, 914–922. [Google Scholar] [CrossRef]
- Butt, M.M.; Latif, G.; Iskandar, D.N.F.A.; Alghazo, J.; Khan, A.H. Multi-Channel Convolutions Neural Network Based Diabetic Retinopathy Detection from Fundus Images. Procedia Comput. Sci. 2019, 163, 283–291. [Google Scholar] [CrossRef]
- Ahmad, M.; Qadri, S.F.; Ashraf, M.U.; Subhi, K.; Khan, S.; Zareen, S.S.; Qadri, S. Efficient Liver Segmentation from Computed Tomography Images Using Deep Learning. Comput. Intell. Neurosci. 2022, 2022, 2665283. [Google Scholar] [CrossRef]
- Qadri, S.F.; Shen, L.; Ahmad, M.; Qadri, S.; Zareen, S.S.; Akbar, M.A. SVseg: Stacked sparse autoencoder-based patch classification modeling for vertebrae segmentation. Mathematics 2022, 10, 796. [Google Scholar] [CrossRef]
- Gozes, O.; Frid-Adar, M.; Greenspan, H.; Browning, P.D.; Zhang, H.; Ji, W.; Bernheim, A.; Siegel, E. Rapid AI Development Cycle for the Coronavirus (COVID-19) Pandemic: Initial Results for Automated Detection & Patient Monitoring Using Deep Learning CT Image Analysis. arXiv 2020, arXiv:2003.05037. [Google Scholar]
- Shan, F.; Gao, Y.; Wang, J.; Shi, W.; Shi, N.; Han, M.; Xue, Z.; Shen, D.; Shi, Y. Abnormal Lung Quantification in Chest CT Images of COVID-19 Patients with Deep Learning and Its Application to Severity Prediction. Med. Phys. 2021, 48, 1633–1645. [Google Scholar] [CrossRef] [PubMed]
- El-Rashidy, N.; Abdelrazik, S.; Abuhmed, T.; Amer, E.; Ali, F.; Hu, J.-W.; El-Sappagh, S. Comprehensive Survey of Using Machine Learning in the COVID-19 Pandemic. Diagnostics 2021, 11, 1155. [Google Scholar] [CrossRef] [PubMed]
- Nwankpa, C.; Ijomah, W.; Gachagan, A.; Marshall, S. Activation Functions: Comparison of Trends in Practice and Research for Deep Learning. arXiv 2018, arXiv:1811.03378. [Google Scholar]
- China Consortium of Chest CT Image Investigation (CC-CCII) Dataset. Available online: http://ncov-ai.big.ac.cn/download?lang=en (accessed on 17 December 2021).
- Linka, K.; Hillgärtner, M.; Abdolazizi, K.P.; Aydin, R.C.; Itskov, M.; Cyron, C.J. Constitutive Artificial Neural Networks: A Fast and General Approach to Predictive Data-Driven Constitutive Modeling by Deep Learning. J. Comput. Phys. 2021, 429, 110010. [Google Scholar] [CrossRef]
- Latif, G.; Alghazo, J.; Maheswar, R.; Vijayakumar, V.; Butt, M. Deep Learning Based Intelligence Cognitive Vision Drone for Automatic Plant Diseases Identification and Spraying. J. Intell. Fuzzy Syst. 2020, 39, 8103–8114. [Google Scholar] [CrossRef]
- Shaikh, E.; Mohiuddin, I.; Manzoor, A.; Latif, G.; Mohammad, N. Automated Grading for Handwritten Answer Sheets Using Convolutional Neural Networks. In Proceedings of the 2019 2nd International Conference on new Trends in Computing Sciences (ICTCS), Amman, Jordan, 9–11 October 2019. [Google Scholar] [CrossRef]
- Alzubaidi, L.; Fadhel, M.A.; Al-Shamma, O.; Zhang, J.; Santamaría, J.; Duan, Y.; Oleiwi, S.R. Towards a Better Understanding of Transfer Learning for Medical Imaging: A Case Study. Appl. Sci. 2020, 10, 4523. [Google Scholar] [CrossRef]
- Latif, G.; Al Anezi, F.Y.; Sibai, F.N.; Alghazo, J. Lung Opacity Pneumonia Detection with Improved Residual Networks. J. Med. Biol. Eng. 2021, 41, 581–591. [Google Scholar] [CrossRef]
- Sanghvi, K.; Aralkar, A.; Sanghvi, S.; Saha, I. A Survey on Image Classification Techniques. SSRN Electron. J. 2020. [Google Scholar] [CrossRef]
- Purdila, V.; Pentiuc, S.-G. Fast Decision Tree Algorithm. Adv. Electr. Comput. Eng. 2014, 14, 65–68. [Google Scholar] [CrossRef]
- Latif, G.; Iskandar, D.A.; Alghazo, J. Multiclass Brain Tumor Classification Using Region Growing Based Tumor Segmentation and Ensemble Wavelet Features. In Proceedings of the 2018 International Conference on Computing and Big Data, Seattle, WA, USA, 25–30 June 2018. [Google Scholar] [CrossRef]
- Latif, G.; Ben Brahim, G.; Iskandar, D.N.F.A.; Bashar, A.; Alghazo, J. Glioma Tumors’ Classification Using Deep-Neural-Network-Based Features with SVM Classifier. Diagnostics 2022, 12, 1018. [Google Scholar] [CrossRef]
- Kruse, R.; Borgelt, C.; Braune, C.; Mostaghim, S.; Steinbrecher, M. Introduction to Bayes Networks. Texts Comput. Sci. 2016, 459–463. [Google Scholar] [CrossRef]
- Zhang, K.; Liu, X.; Shen, J.; Li, Z.; Sang, Y.; Wu, X.; Zha, Y.; Liang, W.; Wang, C.; Wang, K.; et al. Clinically Applicable AI System for Accurate Diagnosis, Quantitative Measurements, and Prognosis of COVID-19 Pneumonia Using Computed Tomography. Cell 2020, 181, 1423–1433.e11. [Google Scholar] [CrossRef]
- Wu, X.; Chen, C.; Zhong, M.; Wang, J.; Shi, J. COVID-AL: The Diagnosis of COVID-19 with Deep Active Learning. Med. Image Anal. 2021, 68, 101913. [Google Scholar] [CrossRef]
- Li, Z.; Zhang, J.; Li, B.; Gu, X.; Luo, X. COVID-19 Diagnosis on CT Scan Images Using a Generative Adversarial Network and Concatenated Feature Pyramid Network with an Attention Mechanism. Med. Phys. 2021, 48, 4334–4349. [Google Scholar] [CrossRef]
- Fu, Y.; Xue, P.; Dong, E. Densely Connected Attention Network for Diagnosing COVID-19 Based on Chest CT. Comput. Biol. Med. 2021, 137, 104857. [Google Scholar] [CrossRef]

| Class | Total of Patients | Total Images | Sample CT Scan Images | ||
|---|---|---|---|---|---|
| CP | 932 | 35,191 | 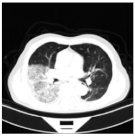 | 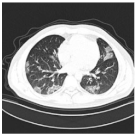 | 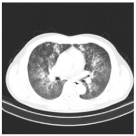 |
| NCP | 929 | 21,872 | 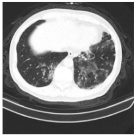 | 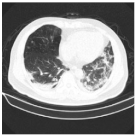 | 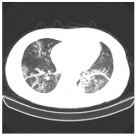 |
| Normal | 850 | 28,548 |  |  | 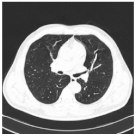 |
| Method | Accuracy | Precision | Recall | F1-Measure | Training Time |
|---|---|---|---|---|---|
| AlexNet | 98.49 | 0.986 | 0.977 | 0.982 | 25,568 |
| VGG16 | 97.32 | 0.971 | 0.960 | 0.972 | 73,756 |
| GoogleNet | 98.71 | 0.984 | 0.972 | 0.978 | 54,866 |
| Method | Accuracy | Precision | Recall | F1-Measure | Training Time |
|---|---|---|---|---|---|
| Random forest | 93.25 | 0.932 | 0.897 | 0.979 | 162 |
| Support vector machine | 99.61 | 0.996 | 0.994 | 0.997 | 6027 |
| Fast decision tree | 93.25 | 0.932 | 0.897 | 0.979 | 162 |
| Bayesian network | 81.49 | 0.81 | 0.729 | 0.937 | 178 |
| Method | Accuracy | Precision | Recall | F1-Measure | Training Time |
|---|---|---|---|---|---|
| Random forest | 97.78 | 0.978 | 0.967 | 0.999 | 202 |
| Support vector machine | 99.86 | 0.999 | 0.998 | 0.999 | 7513 |
| Fast decision tree | 93.47 | 0.999 | 0.998 | 0.999 | 133 |
| Bayesian network | 80.14 | 0.798 | 0.708 | 0.93 | 210 |
| Method | Accuracy | Precision | Recall | F1-Measure | Training Time |
|---|---|---|---|---|---|
| Random forest | 97.93 | 0.979 | 0.969 | 0.999 | 241 |
| Support vector machine | 99.90 | 0.999 | 0.998 | 0.999 | 12,382 |
| Fast decision tree | 94.45 | 0.944 | 0.915 | 0.984 | 302 |
| Bayesian network | 81.88 | 0.814 | 0.736 | 0.931 | 404 |
| Reference | Method | Data | Accuracy |
|---|---|---|---|
| Proposed method | Hybrid ResNet18 and GoogleNet 2000 features with SVM | CC-CCII dataset | 99.91% |
| Kang et al. (2020) [49] | A custom-designed 7-layered 3D CNN model | CC-CCII dataset | 88.94% |
| Xing et al. (2020) [50] | Hybrid active learning with 2D U-Net and residual network | CC-CCII dataset | 95% |
| Li et al. (2021) [51] | Hybrid generative adversarial network and DenseNet | CC-CCII dataset | 85% |
| Fu et al. (2021) [52] | Densely connected attention network (DenseNet) | CC-CCII dataset | 96.06% |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2022 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Latif, G.; Morsy, H.; Hassan, A.; Alghazo, J. Novel Coronavirus and Common Pneumonia Detection from CT Scans Using Deep Learning-Based Extracted Features. Viruses 2022, 14, 1667. https://doi.org/10.3390/v14081667
Latif G, Morsy H, Hassan A, Alghazo J. Novel Coronavirus and Common Pneumonia Detection from CT Scans Using Deep Learning-Based Extracted Features. Viruses. 2022; 14(8):1667. https://doi.org/10.3390/v14081667
Chicago/Turabian StyleLatif, Ghazanfar, Hamdy Morsy, Asmaa Hassan, and Jaafar Alghazo. 2022. "Novel Coronavirus and Common Pneumonia Detection from CT Scans Using Deep Learning-Based Extracted Features" Viruses 14, no. 8: 1667. https://doi.org/10.3390/v14081667
APA StyleLatif, G., Morsy, H., Hassan, A., & Alghazo, J. (2022). Novel Coronavirus and Common Pneumonia Detection from CT Scans Using Deep Learning-Based Extracted Features. Viruses, 14(8), 1667. https://doi.org/10.3390/v14081667






