Development and Validation of Indirect Enzyme-Linked Immunosorbent Assays for Detecting Antibodies to SARS-CoV-2 in Cattle, Swine, and Chicken
Abstract
:1. Introduction
2. Materials and Methods
2.1. Preparation of Recombinant RBD
2.2. Serum Samples
2.3. Raising Hyperimmune Sera
2.4. Virus Neutralization Test
2.5. Indirect ELISAs
2.6. Cut-Off Determination
2.7. Statistical Analysis
3. Results
3.1. Determination of P/N Threshold
3.2. Validation of iELISAs
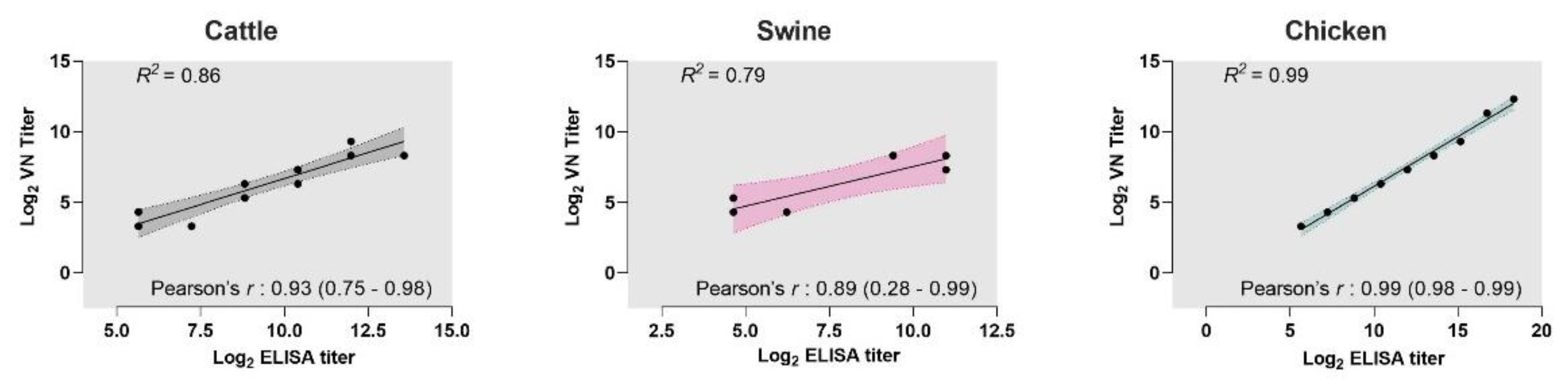
3.3. Correlation between VNT and iELISAs
3.4. Cross-Neutralization of SARS-CoV-2 Variants by HIS Raised to Wuhan RBD
4. Discussion
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- WHO. WHO Coronavirus (COVID-19) Dashboard. Available online: https://covid19.who.int/ (accessed on 19 April 2022).
- Andersen, K.G.; Rambaut, A.; Lipkin, W.I.; Holmes, E.C.; Garry, R.F. The proximal origin of SARS-CoV-2. Nat. Med. 2020, 26, 450–452. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Delahay, R.J.; de la Fuente, J.; Smith, G.C.; Sharun, K.; Snary, E.L.; Flores Girón, L.; Nziza, J.; Fooks, A.R.; Brookes, S.M.; Lean, F.Z.X.; et al. Assessing the risks of SARS-CoV-2 in wildlife. One Health Outlook 2021, 3, 7. [Google Scholar] [CrossRef] [PubMed]
- Holmes, E.C.; Goldstein, S.A.; Rasmussen, A.L.; Robertson, D.L.; Crits-Christoph, A.; Wertheim, J.O.; Anthony, S.J.; Barclay, W.S.; Boni, M.F.; Doherty, P.C.; et al. The origins of SARS-CoV-2: A critical review. Cell 2021, 184, 4848–4856. [Google Scholar] [CrossRef] [PubMed]
- Temmam, S.; Vongphayloth, K.; Baquero, E.; Munier, S.; Bonomi, M.; Regnault, B.; Douangboubpha, B.; Karami, Y.; Chretien, D.; Sanamxay, D.; et al. Bat coronaviruses related to SARS-CoV-2 and infectious for human cells. Nature 2022, 604, 330–336. [Google Scholar] [CrossRef]
- Parrish, C.R.; Holmes, E.C.; Morens, D.M.; Park, E.C.; Burke, D.S.; Calisher, C.H.; Laughlin, C.A.; Saif, L.J.; Daszak, P. Cross-species virus transmission and the emergence of new epidemic diseases. Microbiol. Mol. Biol. Rev. 2008, 72, 457–470. [Google Scholar] [CrossRef] [Green Version]
- Hale, V.L.; Dennis, P.M.; McBride, D.S.; Nolting, J.M.; Madden, C.; Huey, D.; Ehrlich, M.; Grieser, J.; Winston, J.; Lombardi, D.; et al. SARS-CoV-2 infection in free-ranging white-tailed deer. Nature 2021, 602, 481–486. [Google Scholar] [CrossRef]
- Shriner, S.A.; Ellis, J.W.; Root, J.J.; Roug, A.; Stopak, S.R.; Wiscomb, G.W.; Zierenberg, J.R.; Ip, H.S.; Torchetti, M.K.; DeLiberto, T.J. SARS-CoV-2 Exposure in Escaped Mink, Utah, USA. Emerg. Infect. Dis. 2021, 27, 988–990. [Google Scholar] [CrossRef]
- Bosco-Lauth, A.M.; Hartwig, A.E.; Porter, S.M.; Gordy, P.W.; Nehring, M.; Byas, A.D.; VandeWoude, S.; Ragan, I.K.; Maison, R.M.; Bowen, R.A. Experimental infection of domestic dogs and cats with SARS-CoV-2: Pathogenesis, transmission, and response to reexposure in cats. Proc. Natl. Acad. Sci. USA 2020, 117, 26382–26388. [Google Scholar] [CrossRef]
- Bosco-Lauth, A.M.; Walker, A.; Guilbert, L.; Porter, S.; Hartwig, A.; McVicker, E.; Bielefeldt-Ohmann, H.; Bowen, R.A. Susceptibility of livestock to SARS-CoV-2 infection. Emerg. Microbes Infect. 2021, 10, 2199–2201. [Google Scholar] [CrossRef]
- Chandler, J.C.; Bevins, S.N.; Ellis, J.W.; Linder, T.J.; Tell, R.M.; Jenkins-Moore, M.; Root, J.J.; Lenoch, J.B.; Robbe-Austerman, S.; DeLiberto, T.J.; et al. SARS-CoV-2 exposure in wild white-tailed deer (Odocoileus virginianus). Proc. Natl. Acad. Sci. USA 2021, 118, e2114828118. [Google Scholar] [CrossRef]
- Fritz, M.; Rosolen, B.; Krafft, E.; Becquart, P.; Elguero, E.; Vratskikh, O.; Denolly, S.; Boson, B.; Vanhomwegen, J.; Gouilh, M.A.; et al. High prevalence of SARS-CoV-2 antibodies in pets from COVID-19+ households. One Health 2020, 11, 100192. [Google Scholar] [CrossRef] [PubMed]
- Hobbs, E.C.; Reid, T.J. Animals and SARS-CoV-2: Species susceptibility and viral transmission in experimental and natural conditions, and the potential implications for community transmission. Transbound. Emerg. Dis. 2021, 68, 1850–1867. [Google Scholar] [CrossRef] [PubMed]
- Kuchipudi, S.V.; Surendran-Nair, M.; Ruden, R.M.; Yon, M.; Nissly, R.H.; Vandegrift, K.J.; Nelli, R.K.; Li, L.; Jayarao, B.M.; Maranas, C.D.; et al. Multiple spillovers from humans and onward transmission of SARS-CoV-2 in white-tailed deer. Proc. Natl. Acad. Sci. USA 2022, 119, e2121644119. [Google Scholar] [CrossRef] [PubMed]
- Mahdy, M.A.A.; Younis, W.; Ewaida, Z. An Overview of SARS-CoV-2 and Animal Infection. Front. Vet. Sci. 2020, 7, 596391. [Google Scholar] [CrossRef]
- Vandegrift, K.J.; Yon, M.; Surendran-Nair, M.; Gontu, A.; Amirthalingam, S.; Nissly, R.H.; Levine, N.; Stuber, T.; DeNicola, A.J.; Boulanger, J.R.; et al. Detection of SARS-CoV-2 Omicron variant (B.1.1.529) infection of white-tailed deer. bioRxiv 2022, 2022.02.04.479189. [Google Scholar] [CrossRef]
- Damas, J.; Hughes, G.M.; Keough, K.C.; Painter, C.A.; Persky, N.S.; Corbo, M.; Hiller, M.; Koepfli, K.-P.; Pfenning, A.R.; Zhao, H.; et al. Broad host range of SARS-CoV-2 predicted by comparative and structural analysis of ACE2 in vertebrates. Proc. Natl. Acad. Sci. USA 2020, 117, 22311. [Google Scholar] [CrossRef]
- Huang, X.; Zhang, C.; Pearce, R.; Omenn, G.S.; Zhang, Y. Identifying the Zoonotic Origin of SARS-CoV-2 by Modeling the Binding Affinity between the Spike Receptor-Binding Domain and Host ACE2. J. Proteome Res. 2020, 19, 4844–4856. [Google Scholar] [CrossRef]
- Singh, J.; Pandit, P.; McArthur, A.G.; Banerjee, A.; Mossman, K. Evolutionary trajectory of SARS-CoV-2 and emerging variants. Virol. J. 2021, 18, 166. [Google Scholar] [CrossRef]
- Ulrich, L.; Wernike, K.; Hoffmann, D.; Mettenleiter, T.C.; Beer, M. Experimental Infection of Cattle with SARS-CoV-2. Emerg. Infect. Dis. 2020, 26, 2979–2981. [Google Scholar] [CrossRef]
- Pickering, B.S.; Smith, G.; Pinette, M.M.; Embury-Hyatt, C.; Moffat, E.; Marszal, P.; Lewis, C.E. Susceptibility of Domestic Swine to Experimental Infection with Severe Acute Respiratory Syndrome Coronavirus 2. Emerg. Infect. Dis. 2021, 27, 104–112. [Google Scholar] [CrossRef]
- Di Teodoro, G.; Valleriani, F.; Puglia, I.; Monaco, F.; Di Pancrazio, C.; Luciani, M.; Krasteva, I.; Petrini, A.; Marcacci, M.; D’Alterio, N.; et al. SARS-CoV-2 replicates in respiratory ex vivo organ cultures of domestic ruminant species. Vet. Microbiol. 2021, 252, 108933. [Google Scholar] [CrossRef] [PubMed]
- Bashor, L.; Gagne, R.B.; Bosco-Lauth, A.; Bowen, R.; Stenglein, M.; VandeWoude, S. SARS-CoV-2 evolution in animals suggests mechanisms for rapid variant selection. Proc. Natl. Acad. Sci. USA. 2021, 118, e2105253118. [Google Scholar] [CrossRef] [PubMed]
- Daszak, P.; Cunningham, A.A.; Hyatt, A.D. Emerging infectious diseases of wildlife--threats to biodiversity and human health. Science 2000, 287, 443–449. [Google Scholar] [CrossRef] [PubMed]
- Kerstin, W.; Jens, B.; Silke, A.; Kerstin, A.; Tanja, G.; Karsten, D.; Martin, B. Serological screening suggests single SARS-CoV-2 spillover events to cattle. bioRxiv 2022. [Google Scholar] [CrossRef]
- Santini, J.M.; Edwards, S.J.L. Host range of SARS-CoV-2 and implications for public health. Lancet Microbe 2020, 1, e141–e142. [Google Scholar] [CrossRef]
- Drain, P.K. Rapid Diagnostic Testing for SARS-CoV-2. N. Engl. J. Med. 2022, 386, 264–272. [Google Scholar] [CrossRef]
- Reynard, C.; Allen, J.A.; Shinkins, B.; Prestwich, G.; Goves, J.; Davies, K.; Body, R. COVID-19 rapid diagnostics: Practice review. Emerg. Med. J. 2022, 39, 70. [Google Scholar] [CrossRef]
- Wang, W.; Xu, Y.; Gao, R.; Lu, R.; Han, K.; Wu, G.; Tan, W. Detection of SARS-CoV-2 in Different Types of Clinical Specimens. JAMA 2020, 323, 1843–1844. [Google Scholar] [CrossRef] [Green Version]
- Meekins, D.A.; Gaudreault, N.N.; Richt, J.A. Natural and Experimental SARS-CoV-2 Infection in Domestic and Wild Animals. Viruses 2021, 13, 1993. [Google Scholar] [CrossRef]
- Palmer Mitchell, V.; Martins, M.; Falkenberg, S.; Buckley, A.; Caserta Leonardo, C.; Mitchell Patrick, K.; Cassmann Eric, D.; Rollins, A.; Zylich Nancy, C.; Renshaw Randall, W.; et al. Susceptibility of White-Tailed Deer (Odocoileus virginianus) to SARS-CoV-2. J. Virol. 2021, 95, e00083-21. [Google Scholar] [CrossRef]
- Cool, K.; Gaudreault, N.N.; Morozov, I.; Trujillo, J.D.; Meekins, D.A.; McDowell, C.; Carossino, M.; Bold, D.; Mitzel, D.; Kwon, T.; et al. Infection and transmission of ancestral SARS-CoV-2 and its alpha variant in pregnant white-tailed deer. Emerg. Microbes Infect. 2022, 11, 95–112. [Google Scholar] [CrossRef] [PubMed]
- Haveri, A.; Ekström, N.; Solastie, A.; Virta, C.; Österlund, P.; Isosaari, E.; Nohynek, H.; Palmu, A.A.; Melin, M. Persistence of neutralizing antibodies a year after SARS-CoV-2 infection in humans. Eur. J. Immunol. 2021, 51, 3202–3213. [Google Scholar] [CrossRef] [PubMed]
- Lau, E.H.Y.; Hui, D.S.C.; Tsang, O.T.Y.; Chan, W.-H.; Kwan, M.Y.W.; Chiu, S.S.; Cheng, S.M.S.; Ko, R.L.W.; Li, J.K.C.; Chaothai, S.; et al. Long-term persistence of SARS-CoV-2 neutralizing antibody responses after infection and estimates of the duration of protection. EClinicalMedicine 2021, 41, 101174. [Google Scholar] [CrossRef] [PubMed]
- Mehdi, F.; Chattopadhyay, S.; Thiruvengadam, R.; Yadav, S.; Kumar, M.; Sinha, S.K.; Goswami, S.; Kshetrapal, P.; Wadhwa, N.; Chandramouli Natchu, U.; et al. Development of a Fast SARS-CoV-2 IgG ELISA, Based on Receptor-Binding Domain, and Its Comparative Evaluation Using Temporally Segregated Samples From RT-PCR Positive Individuals. Front. Microbiol. 2020, 11, 618097. [Google Scholar] [CrossRef]
- Stadlbauer, D.; Amanat, F.; Chromikova, V.; Jiang, K.; Strohmeier, S.; Arunkumar, G.A.; Tan, J.; Bhavsar, D.; Capuano, C.; Kirkpatrick, E.; et al. SARS-CoV-2 Seroconversion in Humans: A Detailed Protocol for a Serological Assay, Antigen Production, and Test Setup. Curr. Protoc. Microbiol. 2020, 57, e100. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Gontu, A.; Srinivasan, S.; Salazar, E.; Nair, M.S.; Nissly, R.H.; Greenawalt, D.; Bird, I.M.; Herzog, C.M.; Ferrari, M.J.; Poojary, I.; et al. Limited window for donation of convalescent plasma with high live-virus neutralizing antibody titers for COVID-19 immunotherapy. Commun. Biol. 2021, 4, 267. [Google Scholar] [CrossRef]
- Salazar, E.; Kuchipudi, S.V.; Christensen, P.A.; Eagar, T.; Yi, X.; Zhao, P.; Jin, Z.; Long, S.W.; Olsen, R.J.; Chen, J.; et al. Convalescent plasma anti–SARS-CoV-2 spike protein ectodomain and receptor-binding domain IgG correlate with virus neutralization. J. Clin. Investig. 2020, 130, 6728–6738. [Google Scholar] [CrossRef]
- Grodzki, A.C.; Berenstein, E. Antibody Purification: Affinity Chromatography—Protein A and Protein G Sepharose. In Immunocytochemical Methods and Protocols; Oliver, C., Jamur, M.C., Eds.; Humana Press: Totowa, NJ, USA, 2010; pp. 33–41. [Google Scholar] [CrossRef]
- Gruell, H.; Vanshylla, K.; Tober-Lau, P.; Hillus, D.; Schommers, P.; Lehmann, C.; Kurth, F.; Sander, L.E.; Klein, F. mRNA booster immunization elicits potent neutralizing serum activity against the SARS-CoV-2 Omicron variant. Nat. Med. 2022, 28, 477–480. [Google Scholar] [CrossRef]
- Farag, E.A.; Islam, M.M.; Enan, K.; El-Hussein, A.-R.M.; Bansal, D.; Haroun, M. SARS-CoV-2 at the human-animal interface: A review. Heliyon 2021, 7, e08496. [Google Scholar] [CrossRef]
- Sharun, K.; Dhama, K.; Pawde, A.M.; Gortazar, C.; Tiwari, R.; Bonilla-Aldana, D.K.; Rodriguez-Morales, A.J.; de la Fuente, J.; Michalak, I.; Attia, Y.A. SARS-CoV-2 in animals: Potential for unknown reservoir hosts and public health implications. Vet. Q. 2021, 41, 181–201. [Google Scholar] [CrossRef]
- Sun, Y.; Lin, W.; Dong, W.; Xu, J. Origin and evolutionary analysis of the SARS-CoV-2 Omicron variant. J. Biosaf. Biosecur. 2022, 4, 33–37. [Google Scholar] [CrossRef] [PubMed]
- Wei, C.; Shan, K.-J.; Wang, W.; Zhang, S.; Huan, Q.; Qian, W. Evidence for a mouse origin of the SARS-CoV-2 Omicron variant. J. Genet. Genom. 2021, 48, 1111–1121. [Google Scholar] [CrossRef] [PubMed]
- Berhane, Y.; Suderman, M.; Babiuk, S.; Pickering, B. Susceptibility of turkeys, chickens and chicken embryos to SARS-CoV-2. Transbound. Emerg. Dis. 2021, 68, 3038–3042. [Google Scholar] [CrossRef] [PubMed]
- Vijgen, L.; Keyaerts, E.; Lemey, P.; Maes, P.; Van Reeth, K.; Nauwynck, H.; Pensaert, M.; Van Ranst, M. Evolutionary history of the closely related group 2 coronaviruses: Porcine hemagglutinating encephalomyelitis virus, bovine coronavirus, and human coronavirus OC43. J. Virol. 2006, 80, 7270–7274. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Zhou, P.; Fan, H.; Lan, T.; Yang, X.-L.; Shi, W.-F.; Zhang, W.; Zhu, Y.; Zhang, Y.-W.; Xie, Q.-M.; Mani, S.; et al. Fatal swine acute diarrhoea syndrome caused by an HKU2-related coronavirus of bat origin. Nature 2018, 556, 255–258. [Google Scholar] [CrossRef] [PubMed]
- Cui, J.; Li, F.; Shi, Z.-L. Origin and evolution of pathogenic coronaviruses. Nat. Rev. Microbiol. 2019, 17, 181–192. [Google Scholar] [CrossRef] [Green Version]
- Vergara-Alert, J.; Rodon, J.; Carrillo, J.; Te, N.; Izquierdo-Useros, N.; Rodriguez de la Concepcion, M.L.; Avila-Nieto, C.; Guallar, V.; Valencia, A.; Cantero, G.; et al. Pigs are not susceptible to SARS-CoV-2 infection but are a model for viral immunogenicity studies. Transbound. Emerg. Dis. 2021, 68, 1721–1725. [Google Scholar] [CrossRef]
- Wernike, K.; Aebischer, A.; Michelitsch, A.; Hoffmann, D.; Freuling, C.; Balkema-Buschmann, A.; Graaf, A.; Muller, T.; Osterrieder, N.; Rissmann, M.; et al. Multi-species ELISA for the detection of antibodies against SARS-CoV-2 in animals. Transbound. Emerg. Dis. 2021, 68, 1779–1785. [Google Scholar] [CrossRef]
- Matusali, G.; Colavita, F.; Lapa, D.; Meschi, S.; Bordi, L.; Piselli, P.; Gagliardini, R.; Corpolongo, A.; Nicastri, E.; Antinori, A.; et al. SARS-CoV-2 Serum Neutralization Assay: A Traditional Tool for a Brand-New Virus. Viruses 2021, 13, 655. [Google Scholar] [CrossRef]
- Premkumar, L.; Segovia-Chumbez, B.; Jadi, R.; Martinez, D.R.; Raut, R.; Markmann, A.; Cornaby, C.; Bartelt, L.; Weiss, S.; Park, Y.; et al. The receptor binding domain of the viral spike protein is an immunodominant and highly specific target of antibodies in SARS-CoV-2 patients. Sci. Immunol. 2020, 5, eabc8413. [Google Scholar] [CrossRef]
- Sancilio, A.E.; D’Aquila, R.T.; McNally, E.M.; Velez, M.P.; Ison, M.G.; Demonbreun, A.R.; McDade, T.W. A surrogate virus neutralization test to quantify antibody-mediated inhibition of SARS-CoV-2 in finger stick dried blood spot samples. Sci. Rep. 2021, 11, 15321. [Google Scholar] [CrossRef] [PubMed]
- Tan, C.W.; Chia, W.N.; Qin, X.; Liu, P.; Chen, M.I.; Tiu, C.; Hu, Z.; Chen, V.C.; Young, B.E.; Sia, W.R.; et al. A SARS-CoV-2 surrogate virus neutralization test based on antibody-mediated blockage of ACE2-spike protein-protein interaction. Nat. Biotechnol. 2020, 38, 1073–1078. [Google Scholar] [CrossRef] [PubMed]
- Crawford, K.H.D.; Eguia, R.; Dingens, A.S.; Loes, A.N.; Malone, K.D.; Wolf, C.R.; Chu, H.Y.; Tortorici, M.A.; Veesler, D.; Murphy, M.; et al. Protocol and Reagents for Pseudotyping Lentiviral Particles with SARS-CoV-2 Spike Protein for Neutralization Assays. Viruses 2020, 12, 513. [Google Scholar] [CrossRef] [PubMed]
- Neerukonda, S.N.; Vassell, R.; Herrup, R.; Liu, S.; Wang, T.; Takeda, K.; Yang, Y.; Lin, T.L.; Wang, W.; Weiss, C.D. Establishment of a well-characterized SARS-CoV-2 lentiviral pseudovirus neutralization assay using 293T cells with stable expression of ACE2 and TMPRSS2. PLoS ONE 2021, 16, e0248348. [Google Scholar] [CrossRef] [PubMed]

| VNT Status | Total | Positive | Negative | Sensitivity/Specificity |
|---|---|---|---|---|
| a. Cattle | ||||
| Positive sera * | 13 | 13 | 0 | 100% (13/13; sensitivity) |
| Negative sera | 150 | 0 | 150 | 100% (150/150; specificity) |
| b. Swine | ||||
| Positive sera | 6 | 6 | 0 | 100% (6/6; sensitivity) |
| Negative sera | 150 | 0 | 150 | 100% (150/150; specificity) |
| c. Chicken | ||||
| Positive sera * | 24 | 24 | 0 | 100% (24/24; sensitivity) |
| Negative sera | 150 | 0 | 150 | 100% (150/150; specificity) |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2022 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Gontu, A.; Marlin, E.A.; Ramasamy, S.; Neerukonda, S.; Anil, G.; Morgan, J.; Quraishi, M.; Chen, C.; Boorla, V.S.; Nissly, R.H.; et al. Development and Validation of Indirect Enzyme-Linked Immunosorbent Assays for Detecting Antibodies to SARS-CoV-2 in Cattle, Swine, and Chicken. Viruses 2022, 14, 1358. https://doi.org/10.3390/v14071358
Gontu A, Marlin EA, Ramasamy S, Neerukonda S, Anil G, Morgan J, Quraishi M, Chen C, Boorla VS, Nissly RH, et al. Development and Validation of Indirect Enzyme-Linked Immunosorbent Assays for Detecting Antibodies to SARS-CoV-2 in Cattle, Swine, and Chicken. Viruses. 2022; 14(7):1358. https://doi.org/10.3390/v14071358
Chicago/Turabian StyleGontu, Abhinay, Erika A. Marlin, Santhamani Ramasamy, Sabarinath Neerukonda, Gayatri Anil, Jasmine Morgan, Meysoon Quraishi, Chen Chen, Veda Sheersh Boorla, Ruth H. Nissly, and et al. 2022. "Development and Validation of Indirect Enzyme-Linked Immunosorbent Assays for Detecting Antibodies to SARS-CoV-2 in Cattle, Swine, and Chicken" Viruses 14, no. 7: 1358. https://doi.org/10.3390/v14071358
APA StyleGontu, A., Marlin, E. A., Ramasamy, S., Neerukonda, S., Anil, G., Morgan, J., Quraishi, M., Chen, C., Boorla, V. S., Nissly, R. H., Jakka, P., Chothe, S. K., Ravichandran, A., Kodali, N., Amirthalingam, S., LaBella, L., Kelly, K., Natesan, P., Minns, A. M., ... Kuchipudi, S. V. (2022). Development and Validation of Indirect Enzyme-Linked Immunosorbent Assays for Detecting Antibodies to SARS-CoV-2 in Cattle, Swine, and Chicken. Viruses, 14(7), 1358. https://doi.org/10.3390/v14071358







