Specific Recognition of the 5′-Untranslated Region of West Nile Virus Genome by Human Innate Immune System
Abstract
:1. Introduction
2. Materials and Methods
2.1. Systems’ Setup and Docking
2.2. Molecular Dynamics Simulations
3. Results
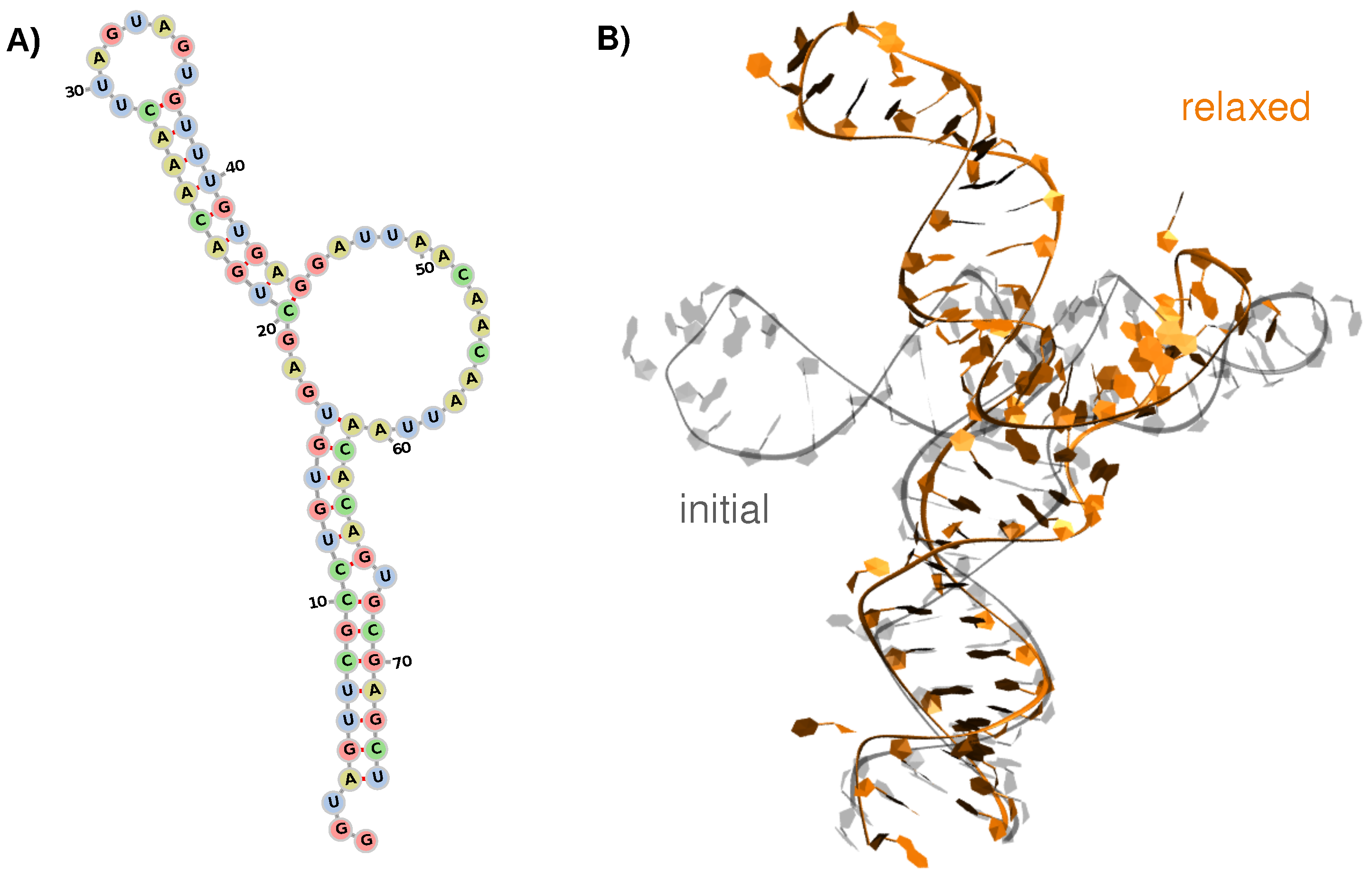
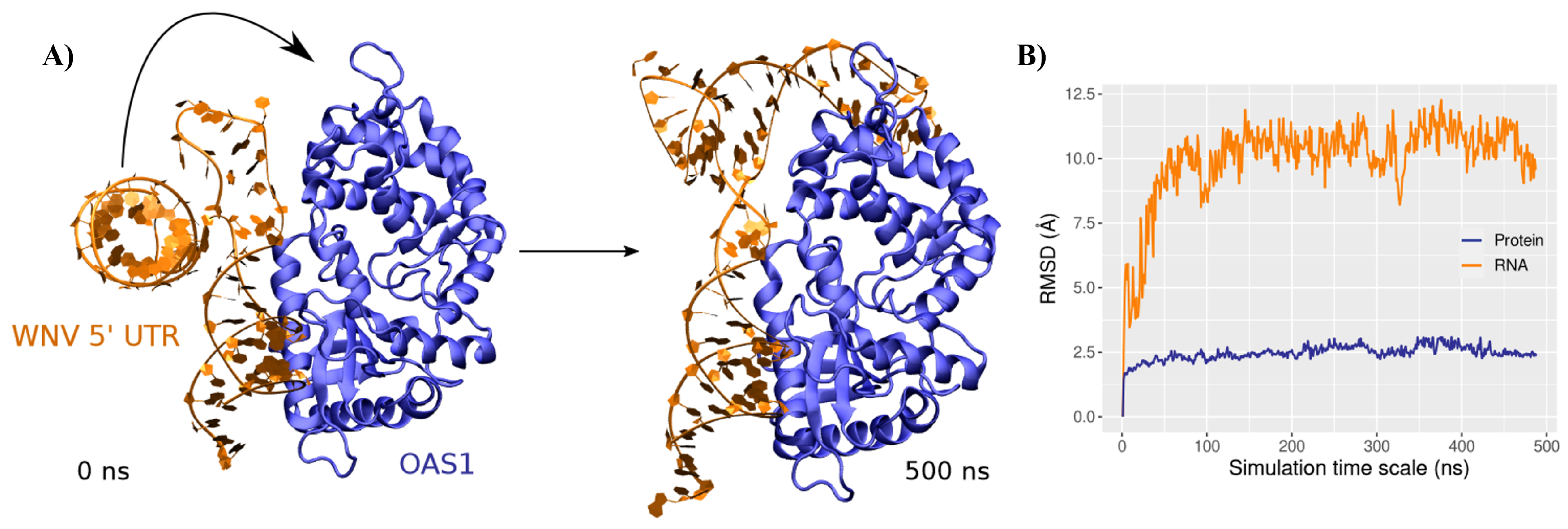
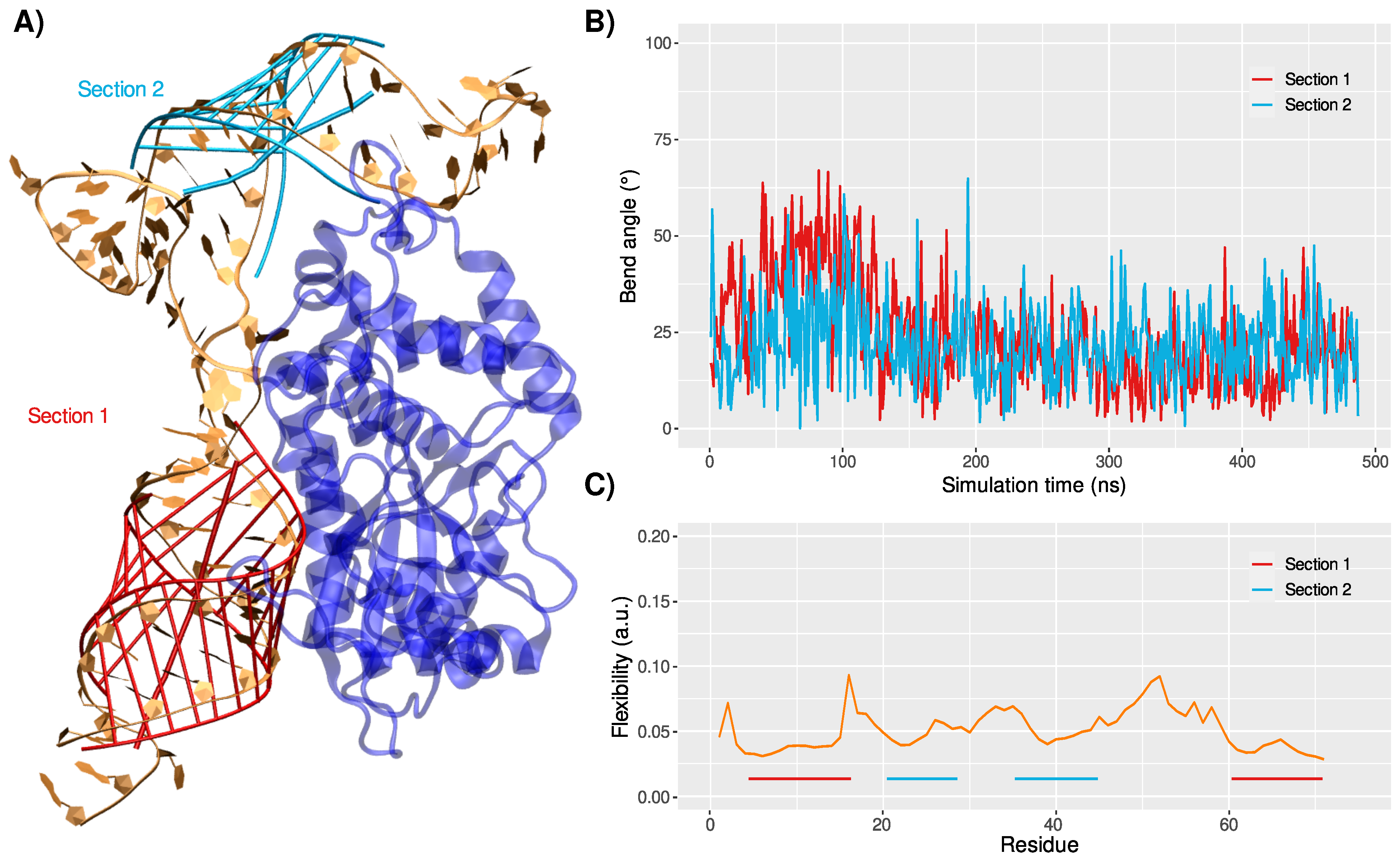
3.1. Structure and Dynamics of the West Nile 5-Untranslated Region
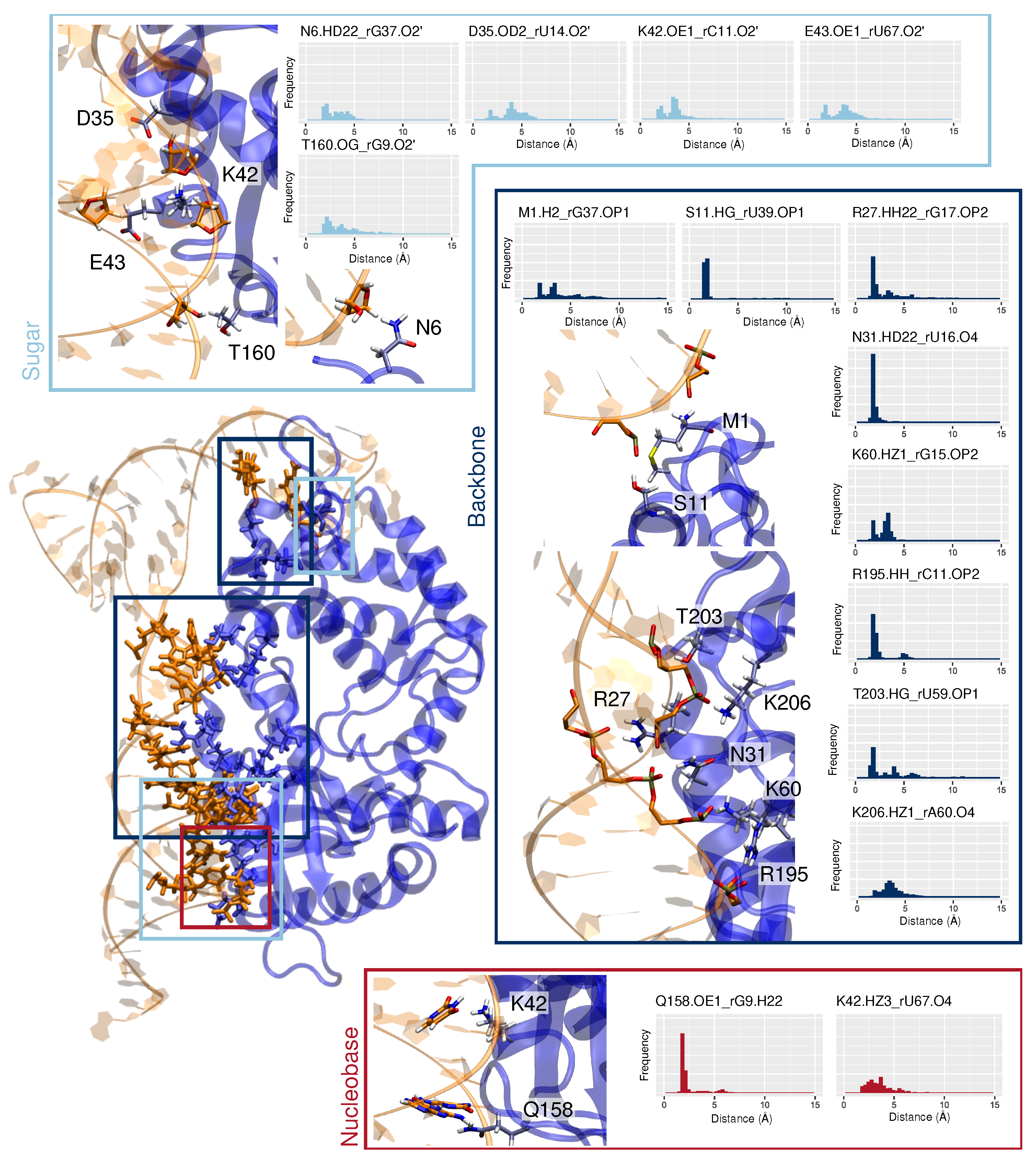
3.2. Binding Mode to OAS1
4. Discussion and Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Suthar, M.S.; Diamond, M.S.; Gale, M., Jr. West Nile virus infection and immunity. Nat. Rev. Microbiol. 2013, 11, 115–128. [Google Scholar] [CrossRef] [PubMed]
- Ronca, S.E.; Ruff, J.C.; Murray, K.O. A 20-year historical review of West Nile virus since its initial emergence in North America: Has West Nile virus become a neglected tropical disease? PLoS Negl. Trop. Dis. 2021, 15, e0009190. [Google Scholar] [CrossRef] [PubMed]
- Byas, A.D.; Ebel, G.D. Comparative pathology of west nile virus in humans and non-human animals. Pathogens 2020, 9, 48. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Colpitts, T.M.; Conway, M.J.; Montgomery, R.R.; Fikrig, E. West Nile Virus: Biology, transmission, and human infection. Clin. Microbiol. Rev. 2012, 25, 635–648. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Martin, M.F.; Nisole, S. West nile virus restriction in mosquito and human cells: A virus under confinement. Vaccines 2020, 8, 256. [Google Scholar] [CrossRef]
- Brinton, M.A. Replication cycle and molecular biology of the west nile virus. Viruses 2013, 6, 13–53. [Google Scholar] [CrossRef] [Green Version]
- Pérez-Ramírez, E.; Llorente, F.; Jiménez-Clavero, M.Á. Experimental infections of wild birds with West Nile virus. Viruses 2014, 6, 752–781. [Google Scholar] [CrossRef] [Green Version]
- Sejvar, J.J. West Nile virus: An historical overview. Ochsner J. 2003, 5, 6–10. [Google Scholar]
- Bakonyi, T.; Haussig, J.M. West Nile virus keeps on moving up in Europe. Eurosurveillance 2020, 25, 2001938. [Google Scholar] [CrossRef]
- Ketkar, H.; Herman, D.; Wang, P. Genetic determinants of the re-emergence of arboviral diseases. Viruses 2019, 11, 150. [Google Scholar] [CrossRef] [Green Version]
- Voss, S.; Nitsche, C. Targeting the protease of West Nile virus. RSC Med. Chem. 2021, 12, 1262–1272. [Google Scholar] [CrossRef] [PubMed]
- Barrows, N.J.; Campos, R.K.; Liao, K.C.; Prasanth, K.R.; Soto-Acosta, R.; Yeh, S.C.; Schott-Lerner, G.; Pompon, J.; Sessions, O.M.; Bradrick, S.S.; et al. Biochemistry and Molecular Biology of Flaviviruses. Chem. Rev. 2018, 118, 4448–4482. [Google Scholar] [CrossRef]
- Smith, D.B.; Becher, P.; Bukh, J.; Gould, E.A.; Meyers, G.; Monath, T.; Muerhoff, A.S.; Pletnev, A.; Rico-Hesse, R.; Stapleton, J.T.; et al. Proposed update to the taxonomy of the genera Hepacivirus and Pegivirus within the Flaviviridae family. J. Gen. Virol. 2016, 97, 2894–2907. [Google Scholar] [CrossRef] [PubMed]
- Richter, J.; Tryfonos, C.; Tourvas, A.; Floridou, D.; Paphitou, N.I.; Christodoulou, C. Complete genome sequence of West Nile Virus (WNV) from the first human case of neuroinvasive WNV infection in Cyprus. Genome Announc. 2017, 5. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Deo, S.; Patel, T.R.; Dzananovic, E.; Booy, E.P.; Zeid, K.; McEleney, K.; Harding, S.E.; McKenna, S.A. Activation of 2′ 5′-oligoadenylate synthetase by stem-loops at the 5′-end of the West Nile virus genome. PLoS ONE 2014, 9, e92545. [Google Scholar]
- Guillemin, A.; Kumar, A.; Wencker, M.; Ricci, E.P. Shaping the Innate Immune Response Through Post-Transcriptional Regulation of Gene Expression Mediated by RNA-Binding Proteins. Front. Immunol. 2022, 12, 796012. [Google Scholar] [CrossRef]
- Sadler, A.J.; Williams, B.R. Interferon-inducible antiviral effectors. Nat. Rev. Immunol. 2008, 8, 559–568. [Google Scholar] [CrossRef]
- Choi, U.Y.; Kang, J.S.; Hwang, Y.S.; Kim, Y.J. Oligoadenylate synthase-like (OASL) proteins: Dual functions and associations with diseases. Exp. Mol. Med. 2015, 47, e144. [Google Scholar] [CrossRef] [Green Version]
- Schwartz, S.L.; Conn, G.L. RNA regulation of the antiviral protein 2′-5′-oligoadenylate synthetase. Wiley Interdiscip. Rev. RNA 2019, 10, e1534. [Google Scholar] [CrossRef]
- Bignon, E.; Miclot, T.; Terenzi, A.; Barone, G.; Monari, A. Structure of the 5′-untranslated region in SARS-CoV-2 genome and its specific recognition by innate immune system via the human oligoadenylate synthase 1. Chem. Commun. 2022. [Google Scholar] [CrossRef]
- Huffman, J.E.; Butler-Laporte, G.; Khan, A.; Pairo-Castineira, E.; Drivas, T.G.; Peloso, G.M.; Nakanishi, T.; Ganna, A.; Verma, A.; Baillie, J.K.; et al. Multi-ancestry fine mapping implicates OAS1 splicing in risk of severe COVID-19. Nat. Genet. 2022, 54, 125–127. [Google Scholar] [CrossRef]
- Zhou, S.; Butler-Laporte, G.; Nakanishi, T.; Morrison, D.R.; Afilalo, J.; Afilalo, M.; Laurent, L.; Pietzner, M.; Kerrison, N.; Zhao, K.; et al. A Neanderthal OAS1 isoform protects individuals of European ancestry against COVID-19 susceptibility and severity. Nat. Med. 2021, 27, 659–667. [Google Scholar] [CrossRef] [PubMed]
- Mashimo, T.; Lucas, M.; Simon-Chazottes, D.; Frenkiel, M.P.; Montagutelli, X.; Ceccaldi, P.E.; Deubel, V.; Guénet, J.L.; Desprès, P. A nonsense mutation in the gene encoding 2′-5′-oligoadenylate synthetase/L1 isoform is associated with West Nile virus susceptibility in laboratory mice. Proc. Natl. Acad. Sci. USA 2002, 99, 11311–11316. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- McMurtrey, C.P.; Lelic, A.; Piazza, P.; Chakrabarti, A.K.; Yablonsky, E.J.; Wahl, A.; Bardet, W.; Eckerd, A.; Cook, R.L.; Hess, R.; et al. Epitope discovery in West Nile virus infection: Identification and immune recognition of viral epitopes. Proc. Natl. Acad. Sci. USA 2008, 105, 2981–2986. [Google Scholar] [CrossRef] [Green Version]
- Yakub, I.; Lillibridge, K.M.; Moran, A.; Gonzalez, O.Y.; Belmont, J.; Gibbs, R.A.; Tweardy, D.J. Single nucleotide polymorphisms in genes for 2′-5′-oligoadenylate synthetase and RNase L in patients hospitalized with West Nile virus infection. J. Infect. Dis. 2005, 192, 1741–1748. [Google Scholar] [CrossRef]
- Popenda, M.; Szachniuk, M.; Antczak, M.; Purzycka, K.J.; Lukasiak, P.; Bartol, N.; Blazewicz, J.; Adamiak, R.W. Automated 3D structure composition for large RNAs. Nucleic Acids Res. 2012, 40, e112. [Google Scholar] [CrossRef] [PubMed]
- Donovan, J.; Dufner, M.; Korennykh, A. Structural basis for cytosolic double-stranded RNA surveillance by human oligoadenylate synthetase 1. Proc. Natl. Acad. Sci. USA 2013, 110, 1652–1657. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Yan, Y.; Tao, H.; He, J.; Huang, S.Y. The HDOCK server for integrated protein–protein docking. Nat. Protoc. 2020, 15, 1829–1852. [Google Scholar] [CrossRef]
- Yan, Y.; Zhang, D.; Zhou, P.; Li, B.; Huang, S.Y. HDOCK: A web server for protein–protein and protein–DNA/RNA docking based on a hybrid strategy. Nucleic Acids Res. 2017, 45, W365–W373. [Google Scholar] [CrossRef]
- Phillips, J.C.; Hardy, D.J.; Maia, J.D.; Stone, J.E.; Ribeiro, J.V.; Bernardi, R.C.; Buch, R.; Fiorin, G.; Hénin, J.; Jiang, W.; et al. Scalable molecular dynamics on CPU and GPU architectures with NAMD. J. Chem. Phys. 2020, 153, 044130. [Google Scholar] [CrossRef]
- Aytenfisu, A.H.; Spasic, A.; Grossfield, A.; Stern, H.A.; Mathews, D.H. Revised RNA Dihedral Parameters for the Amber Force Field Improve RNA Molecular Dynamics. J. Chem. Theory Comput. 2017, 13, 900–915. [Google Scholar] [CrossRef] [PubMed]
- Hopkins, C.W.; Le Grand, S.; Walker, R.C.; Roitberg, A.E. Long-time-step molecular dynamics through hydrogen mass repartitioning. J. Chem. Theory Comput. 2015, 11, 1864–1874. [Google Scholar] [CrossRef] [PubMed]
- Essmann, U.; Perera, L.; Berkowitz, M.L.; Darden, T.; Lee, H.; Pedersen, L.G. A smooth particle mesh Ewald method. J. Chem. Phys. 1995, 103, 8577–8593. [Google Scholar] [CrossRef] [Green Version]
- Lavery, R.; Moakher, M.; Maddocks, J.H.; Petkeviciute, D.; Zakrzewska, K. Conformational analysis of nucleic acids revisited: Curves+. Nucleic Acids Res. 2009, 37, 5917–5929. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Fleetwood, O.; Kasimova, M.A.; Westerlund, A.M.; Delemotte, L. Molecular Insights from Conformational Ensembles via Machine Learning. Biophys. J. 2020, 118, 765–780. [Google Scholar] [CrossRef] [Green Version]
- Bignon, E.; Gillet, N.; Chan, C.H.; Jiang, T.; Monari, A.; Dumont, E. Recognition of a Tandem Lesion by DNA bacterial formamidopyrimidine Glycosylases Explored Combining Molecular Dynamics and Machine Learning. Comput. Struct. Biotechnol. J. 2021, 19, 2861–2869. [Google Scholar] [CrossRef] [PubMed]
- Jiang, T.; Monari, A.; Dumont, E.; Bignon, E. Molecular mechanisms associated with clustered lesion-induced impairment of 8-oxoG recognition by the human glycosylase OGG1. Molecules 2021, 26, 6465. [Google Scholar] [CrossRef]
- Bignon, E.; Gillet, N.; Jiang, T.; Morell, C.; Dumont, E. A Dynamic View of the Interaction of Histone Tails with Clustered Abasic Sites in a Nucleosome Core Particle. J. Phys. Chem. Lett. 2021, 12, 6014–6019. [Google Scholar] [CrossRef]
- Humphrey, W.; Dalke, A.; Schulten, K. VMD: Visual molecular dynamics. J. Mol. Graph. 1996, 14, 33–38. [Google Scholar] [CrossRef]
- Inkscape Project. Inkscape. Available online: https://inkscape.org/ (accessed on 20 March 2022).
- Wickham, H. ggplot2: Elegant Graphics for Data Analysis; Springer: New York, NY, USA, 2009. [Google Scholar]
- RStudio Team. RStudio: Integrated Development Environment for R; RStudio, PBC: Boston, MA, USA, 2020. [Google Scholar]
- Deo, S.; Patel, T.R.; Chojnowski, G.; Koul, A.; Dzananovic, E.; McEleney, K.; Bujnicki, J.M.; McKenna, S.A. Characterization of the termini of the West Nile virus genome and their interactions with the small isoform of the 2′ 5′-oligoadenylate synthetase family. J. Struct. Biol. 2015, 190, 236–249. [Google Scholar] [CrossRef] [Green Version]
- Li, X.F.; Jiang, T.; Yu, X.D.; Deng, Y.Q.; Zhao, H.; Zhu, Q.Y.; Qin, E.D.; Qin, C.F. RNA elements within the 5′-untranslated region of the West Nile virus genome are critical for RNA synthesis and virus replication. J. Gen. Virol. 2010, 91, 1218–1223. [Google Scholar] [CrossRef] [PubMed]
- Vachon, V.K.; Calderon, B.M.; Conn, G.L. A novel RNA molecular signature for activation of 2′-5′ oligoadenylate synthetase-1. Nucleic Acids Res. 2015, 43, 544–552. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Schwartz, S.L.; Park, E.N.; Vachon, V.K.; Danzy, S.; Lowen, A.C.; Conn, G.L. Human OAS1 activation is highly dependent on both RNA sequence and context of activating RNA motifs. Nucleic Acids Res. 2020, 48, 7520–7531. [Google Scholar] [CrossRef]
- Kodym, R.; Kodym, E.; Story, M.D. 2′-5′-Oligoadenylate synthetase is activated by a specific RNA sequence motif. Biochem. Biophys. Res. Commun. 2009, 388, 317–322. [Google Scholar] [CrossRef] [PubMed]
- Feng, R.; Patil, S.; Zhao, X.; Miao, Z.; Qian, A. RNA Therapeutics - Research and Clinical Advancements. Front. Mol. Biosci. 2021, 8, 913. [Google Scholar] [CrossRef] [PubMed]
- Kim, Y.K. RNA therapy: Rich history, various applications and unlimited future prospects. Exp. Mol. Med. 2022, 54, 455–465. [Google Scholar] [CrossRef] [PubMed]

Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2022 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Bignon, E.; Marazzi, M.; Miclot, T.; Barone, G.; Monari, A. Specific Recognition of the 5′-Untranslated Region of West Nile Virus Genome by Human Innate Immune System. Viruses 2022, 14, 1282. https://doi.org/10.3390/v14061282
Bignon E, Marazzi M, Miclot T, Barone G, Monari A. Specific Recognition of the 5′-Untranslated Region of West Nile Virus Genome by Human Innate Immune System. Viruses. 2022; 14(6):1282. https://doi.org/10.3390/v14061282
Chicago/Turabian StyleBignon, Emmanuelle, Marco Marazzi, Tom Miclot, Giampaolo Barone, and Antonio Monari. 2022. "Specific Recognition of the 5′-Untranslated Region of West Nile Virus Genome by Human Innate Immune System" Viruses 14, no. 6: 1282. https://doi.org/10.3390/v14061282
APA StyleBignon, E., Marazzi, M., Miclot, T., Barone, G., & Monari, A. (2022). Specific Recognition of the 5′-Untranslated Region of West Nile Virus Genome by Human Innate Immune System. Viruses, 14(6), 1282. https://doi.org/10.3390/v14061282







