1. Introduction
Human adenoviruses (HAdVs) are non-enveloped viruses with a linear double-stranded DNA genome of ~36,000 bp. HAdVs are grouped into seven species (A–G) and consist of over 100 different types (consecutively numbered) based on genomic sequencing. HAdVs can infect the respiratory, gastrointestinal, and urinary tracts as well as the conjunctiva. Overall, HAdV usually results in mild, self-limited infections in immunocompetent individuals. HAdV14 is a member of the B2 subgroup of adenovirus and, as such, predominately causes kidney and urinary tract infections but can also be associated with respiratory infections. Outbreaks of emergent strains of Ad have resulted in severe and sometimes fatal infections in otherwise healthy people. In the past fifteen years, an emergent strain of HAdV14, HAdV14p1, emerged first in the U.S. and subsequently throughout the world [
1,
2,
3,
4,
5,
6]. HAdV14p1 was first identified as the causative agent of an outbreak of acute respiratory distress syndrome (ARDS) in U.S. military populations [
7]. HAdV14p1 is 99.7% identical to HAdV14 yet displays increased lung pathogenesis in the Syrian hamster that models HAdV14p1 pathogenesis in humans [
8,
9]. Why HAdV14p1 has increased pathogenesis is still unclear.
The HAdV genome encodes roughly 40 proteins. In addition to protein encoding RNAs, non-coding RNAs (ncRNA) are also present in HAdV-infected cells [
10,
11,
12]. Viral-associated RNA (VA RNA) are encoded by the HAdV genome and are the most abundant viral ncRNA found in infected cells [
13,
14]. All HAdV encode at least one VA RNA transcript ~160 nt in length, and approximately 80% of HAdV serotypes encode at least 2 VA RNAs (VA RNA-I and VA RNA-II), with VA RNA-I encoded by all HAdV serotypes [
15,
16]. VA RNAs are required for efficient viral replication, as the deletion of VA RNA-I reduces viral titer by 20-fold and the deletion of both VA RNA-I and VA RNA-II results in a 60-fold decrease in viral titer [
17,
18]. Unlike protein encoding HAdV RNAs, VA RNAs are transcribed by cellular RNA polymerase III and are expressed throughout the viral replication cycle [
13].
VA RNA forms a complex secondary structure that is critical for its biological function [
19,
20,
21]. Overall, despite their low sequence conservation among HAdV, VA RNAs share similar secondary structure and organization of the apical stem, central domain, and terminal stem [
16,
21]. The crystal structure of HAdV-2 VA RNA-I reveals a sharply bent, coaxially stacked, wobble-enriched, and pseudoknot-anchored molecule to allow VA RNA to interfere with nearly all host systems that interact with double-stranded RNA (dsRNA) [
19]. The best characterized activity of VA RNA is its inhibition of protein kinase R (PKR) through the ability of VA RNA-I to bind and inhibit PKR dimerization [
22,
23,
24]. PKR is part of the cellular innate immune response to viral infection, and, by sensing dsRNA produced during viral infection, PKR activation results in the global inhibition of cap-dependent protein synthesis. VA RNA also binds to 2′-5′ Oligoadenylate Synthetase 1 (OAS1), which results in the activation of OAS1 [
25]. While the interaction of full-length VA RNA activates OAS1, the binding of cleaved VA RNAI (produced following DICER cleavage) to OAS1 inhibits OAS1 activity [
11,
26,
27]. Interestingly, the cleaved VA RNAI has a higher affinity for OAS1 than full-length VA RNAI, suggesting that the inhibitory activity of cleaved VA RNAI may be the predominant activity of VA RNAI on OAS1 [
28]. VA RNAI binds to the retinoic acid-inducible gene I (RIG-I). The triphosphorylyated 5′-end nucleotide of VA RNA triggers RIG-I signaling, which results in an increased type I interferon response [
29,
30,
31,
32]. These roles in modulating the cellular innate immune response to HAdV have been long understood.
Recently, new roles for VA RNAI in regulating cellular miRNA processes have been elucidated. Following transcription, VA RNAI is exported from the nucleus to the cytoplasm by Exportin 5 (Exp5). Exp5 exports RNA molecules (e.g., pre-miRNA and tRNAs) that contain a ‘mini-helix’ from the nucleus [
33,
34,
35,
36]. VA RNAI contains a ‘mini-helix’ at the end of the terminal stem, which results in VA RNA being a direct competitor with cellular small RNAs for interaction with Exp5 [
37]. In addition to competing with pre-miRNA for nuclear export through Exp5, once in the cytoplasm, VA RNAs are cleaved by DICER, like pre-miRNA, and as VA RNA expression increases, more VA RNAs are associated with DICER than host pre-miRNA, resulting in decreased host miRNA expression [
27,
38,
39]. DICER processes VA RNA into small RNAs that resemble host miRNA (called mivaRNA) that can be incorporated into the RNA-induced silencing complex (RISC) [
37,
38,
40,
41,
42]. RISC association allows for mivaRNA to bind and repress target mRNA expression [
38,
40,
41,
42,
43].
Group B2 HAdV members encode only the VA RNAI gene. Bioinformatic analysis of the HAdV14 genome showed that the VA RNAI gene of Ad14 is ~98% identical to the other group B2 members HAdV11, HAdV34, and HAdV35 [
44]. In this report, we further characterize the VA RNA gene, identifying the intragenic promoter elements, transcriptional stop sites, potential OAS1 consensus sites, and GGGU/ACCC sites. Small RNA-seq was used to identify the mivaRNA produced during infection. Despite 100% sequence identity of the VA RNAs, differential expression of the mivaRNA exists between HAdV14- and HAdV14p1-infected A549 cells. We discuss the possibility that this differential mivaRNA expression could play a role in the pathogenesis of HAdV14p1.
2. Materials and Methods
2.1. Cells and Viruses
A549 cells (CCL-185, ATCC, Manassas, VA, USA) were grown at 37 °C and 5% CO
2 in DMEM supplemented with 5% heat-inactivated bovine calf serum, 100 U/mL penicillin, 100 µg/mL streptomycin, and 2 mM L-glutamine. A549 cells were monitored for mycoplasma contamination by PCR and validated by short tandem repeat markers (STR Authentication) (ATCC). HAdV14 deWit was obtained from ATCC, and HAdV14p1 (isolate 1986T) was obtained from the United States Navy [
34,
35,
36]. Viruses were propagated and plaque titered (plaque-forming units (pfu)/mL) in A549 cells.
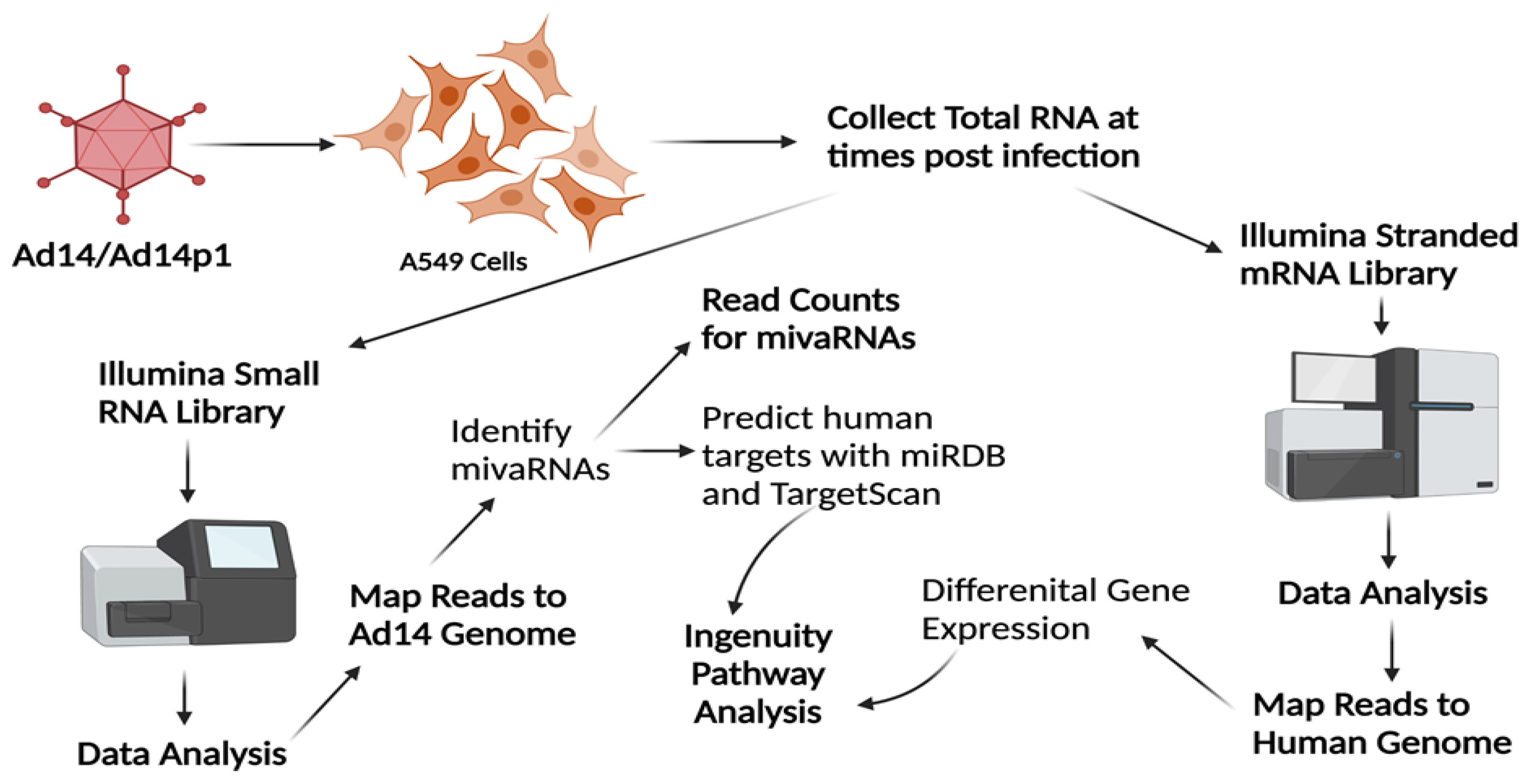
2.2. Infection of A549 Cells and Isolation of Total Cellular RNA
A549 cells were infected with either HAdV14 or HAdV14p1 at a multiplicity of infection (MOI) of 10 pfu/cell in suspension for 1 h at 37 °C, after which cells were plated and allowed to adhere until collected. Adherent and non-adherent cells were collected at 6, 12, 24, 36, and 48 h post-infection. Total RNA was isolated using miRNeasy kit (Qiagen, Germantown, MD, USA) with on column DNase treatment. The total RNA in each sample was quantified using the Qubit 2.0 Fluorometer (Invitrogen, Carlsbad, CA, USA), and quality was measured using the RNA6000 Nano chip on the Agilent 2100 Bioanalzyer (Agilent Technologies, Santa Clara, CA, USA). All samples had an RNA integrity number greater than 7.
2.3. Small RNA Library Preparation, Sequencing and Data Analysis
TruSeq Small RNA library prep kit (Illumina, San Diego, CA, USA) was used to create sequencing libraries. Specifically, adapters were ligated using the 5′ phosphate and 3′ hydroxyl groups common to most mature miRNAs. After adapter ligation, samples were reverse transcribed and amplified. Finally, the libraries were size selected using a 6% polyacrylamide gel and concentrated using ethanol precipitation. Purified libraries were normalized and pooled to create a double-stranded cDNA library and were sequenced on the Illumina MiSeq to render 50 base pair single end reads at the Loyola Stritch School of Medicine Genomics Facility. Adapter sequences were removed, and low-quality reads were trimmed from raw sequencing reads using Cutadapt (v. 1.11). The resulting reads were mapped to the HAdV14 deWit genome (GenBank accession number AY803294) in CLC Genomics Workbench (Qiagen).
2.4. Differential Expression Analysis
Differential expression was conducted with CLC Genomics Workbench. The VA RNA sequence was added to miRbase 22 and annotated to contain either total 5′ and 3′ reads or mivaRNA 5′A, 5′G, 3′A and 3′C seeds. Quantify miRNA 1.2 (CLC) was used with the following settings: allow for length-based isomiRs, no additional upstream bases, 5 additional downstream bases, no missing or mismatch bases, minimum sequence length = 18, maximum sequence length 25. Differential Expression for RNA-Seq 2.6 (CLC) was used for differential expression analysis with normalization set to Trimmed Mean of Means (TMM).
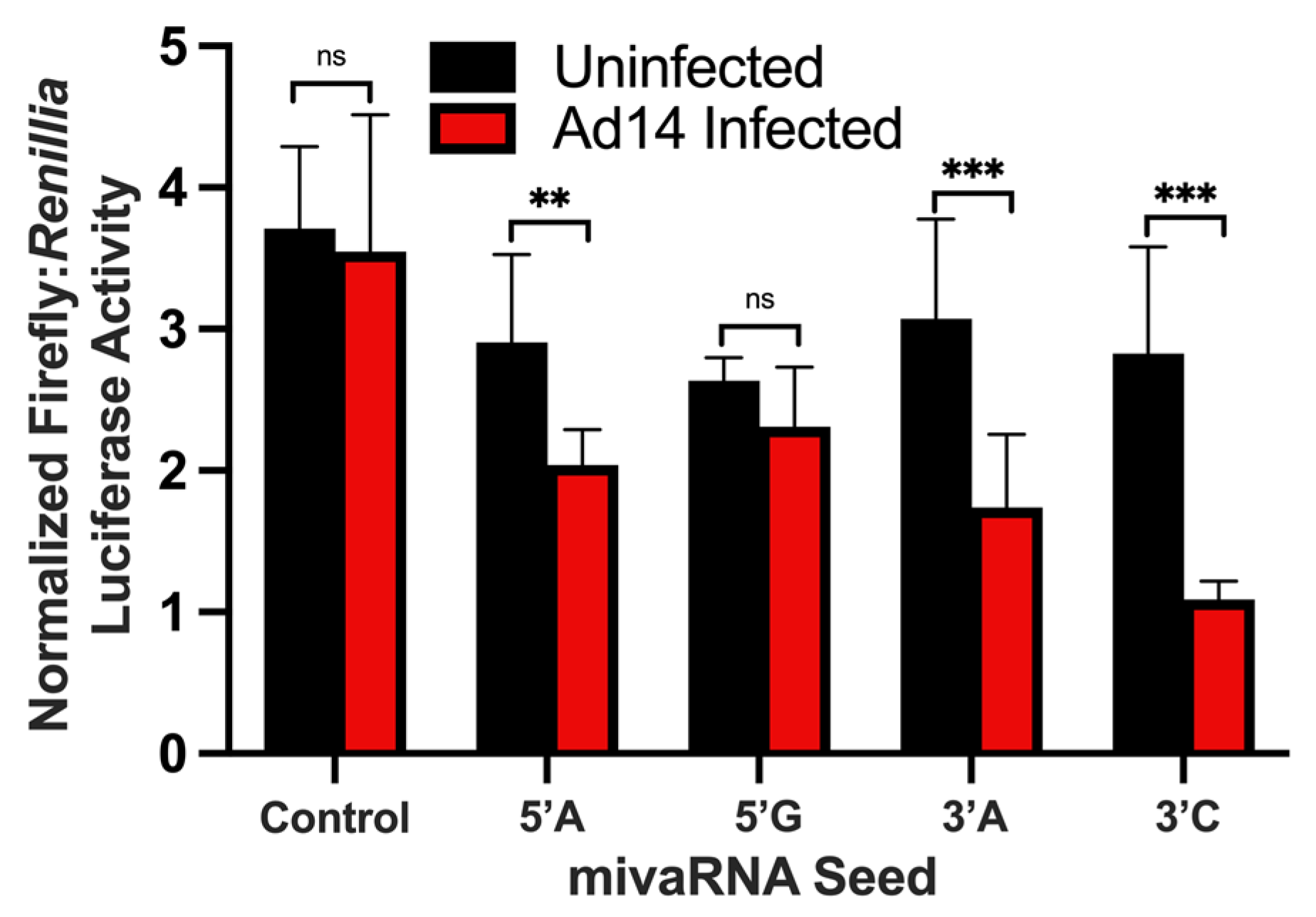
2.5. HAdV-14 mivaRNA 3′ UTR Luciferase Reporter Assays
The complement of the either the 5′ or 3′ seed sequences of Ad14 VA RNA were cloned into the multiple cloning site of pmirGLO Dual Luciferase Vector (Promega, Madison, WI, USA). Constructs were sequenced to confirm the correct orientation of the complimentary HAdV14 VA RNA seed sequences. Plasmids were purified using PureLink HiPure Plasmid Maxi Prep (Invitrogen) and transfected into A549 cells using Lipofectamine LTX (Invitrogen). Twenty-four hours after transfection, cells were infected with Ad14 at an MOI of 10. After an additional 24 h, luciferase activity was determined using a dual-luciferase assay (Promega). Firefly luciferase activity was normalized to Renilla luciferase activity. One-way ANOVA was followed by the Holm-Sidák test with p < 0.05 considered significant (Prism 9; GraphPad Software, San Diego, CA, USA).
2.6. Prediction of mivaRNA Host Gene Targets and Functional Enrichment Analysis
TargetScan and miRDB were used to predict cellular targets for mivaRNAs by seed sequences and filtered to remove duplicate targets. KEGG (
https://www.genome.jp/kegg/, accessed on 2 May 2021) was used to predict the cellular pathways and processes potentially regulated by mivaRNA–target mRNA interaction.
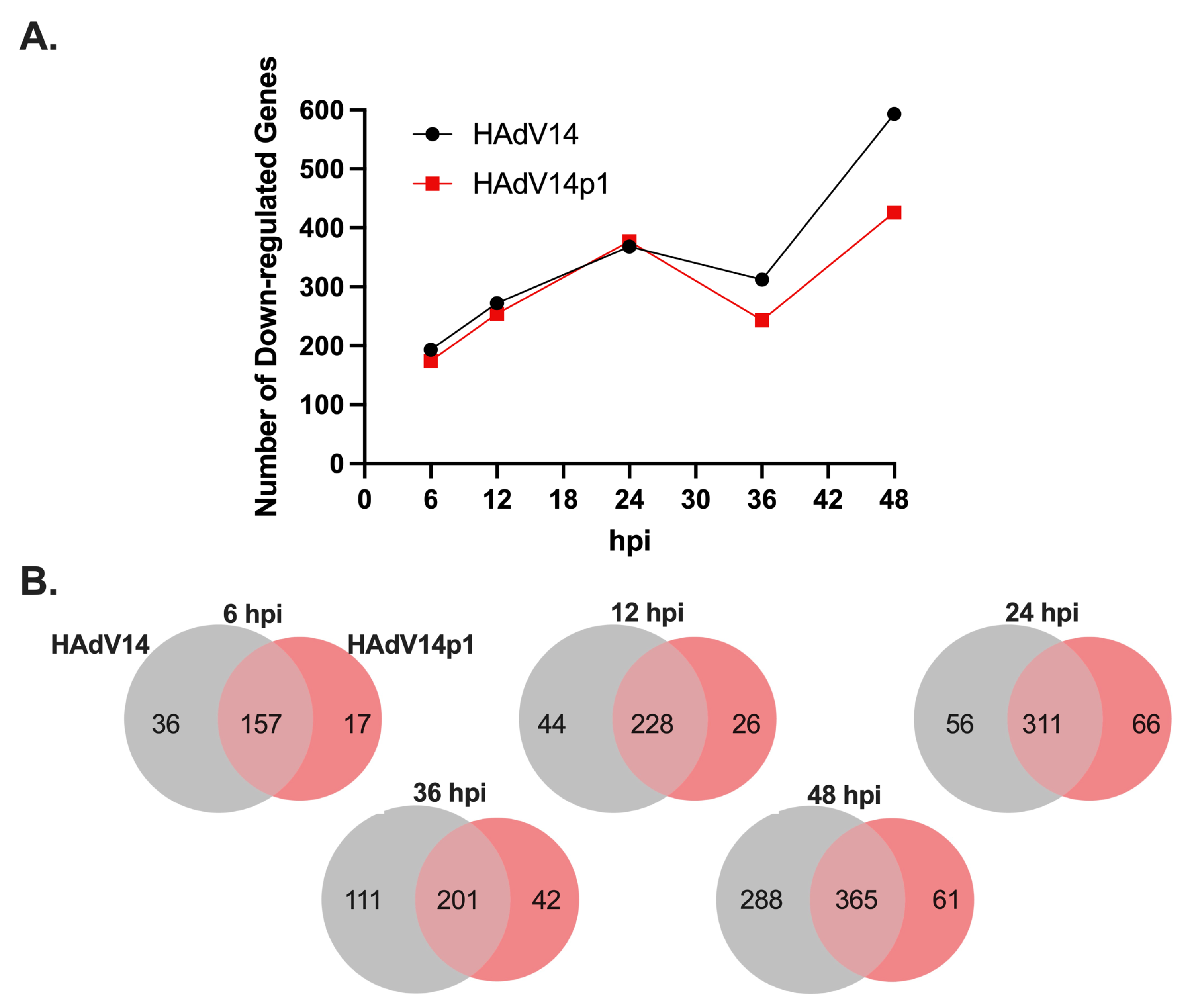
2.7. RNA-Seq Library Preparation, Sequencing, Differential Expression Analysis, and Bioinformatic Analysis
Illumina stranded mRNA libraries were made from the total RNA preps from A549-infected cells. Sequencing was performed at University of Oregon Genomics & Cell Characterization Core (Eugene, OR, USA) on an Illumina HiSeq4000 set for pair ends 100 bp reads and demultiplexed. Reads were trimmed with Trim Reads 2.4 (CLC) before RNA-seq Analysis 2.21 (CLC) was performed, mapping to the human genome (Hg38) with a mismatch cost of 2, insertion cost of 3, deletion cost of 3, and length/similarity fractions set to 0.8. Paired reads were counted as a single read and expression value was set to reads per kilobase million (RPKM). Differential expression for RNA-Seq 2.4 (CLC) using gene expression values from RNA-seq analysis was performed and the results were filtered to the predicted mivaRNA targeted genes. Ingenuity Pathway Analysis (IPA) (Qiagen) was used for functional enrichment with the differential RNA-seq analysis from HAdV14-infected cells at 36 hpi vs. uninfected cells. FDR was set to 0.05 with absolute expression fold changes being >1.5 and expression intensity being >1.
4. Discussion
The adenoviral VA RNA gene is a multifunctional non-coding RNA molecule. VA RNAs are processed by DICER into mivaRNA that resemble and function like cellular miRNAs. While all human adenoviruses encode at least one VA RNA gene, the amount of sequence similarity between VA RNA genes from all human adenoviruses is limited between all human Ad, but is somewhat conserved within Ad groups. In this study, we sought to identify the mivaRNA produced during prototype HAdV14 deWit infection, assess predicted target gene expression, and compare mivaRNA expression with that of the emergent pathogenic strain HAdV14p1, using A549 cells that are permissive for HAdV14 and HAdV14p1 infection.
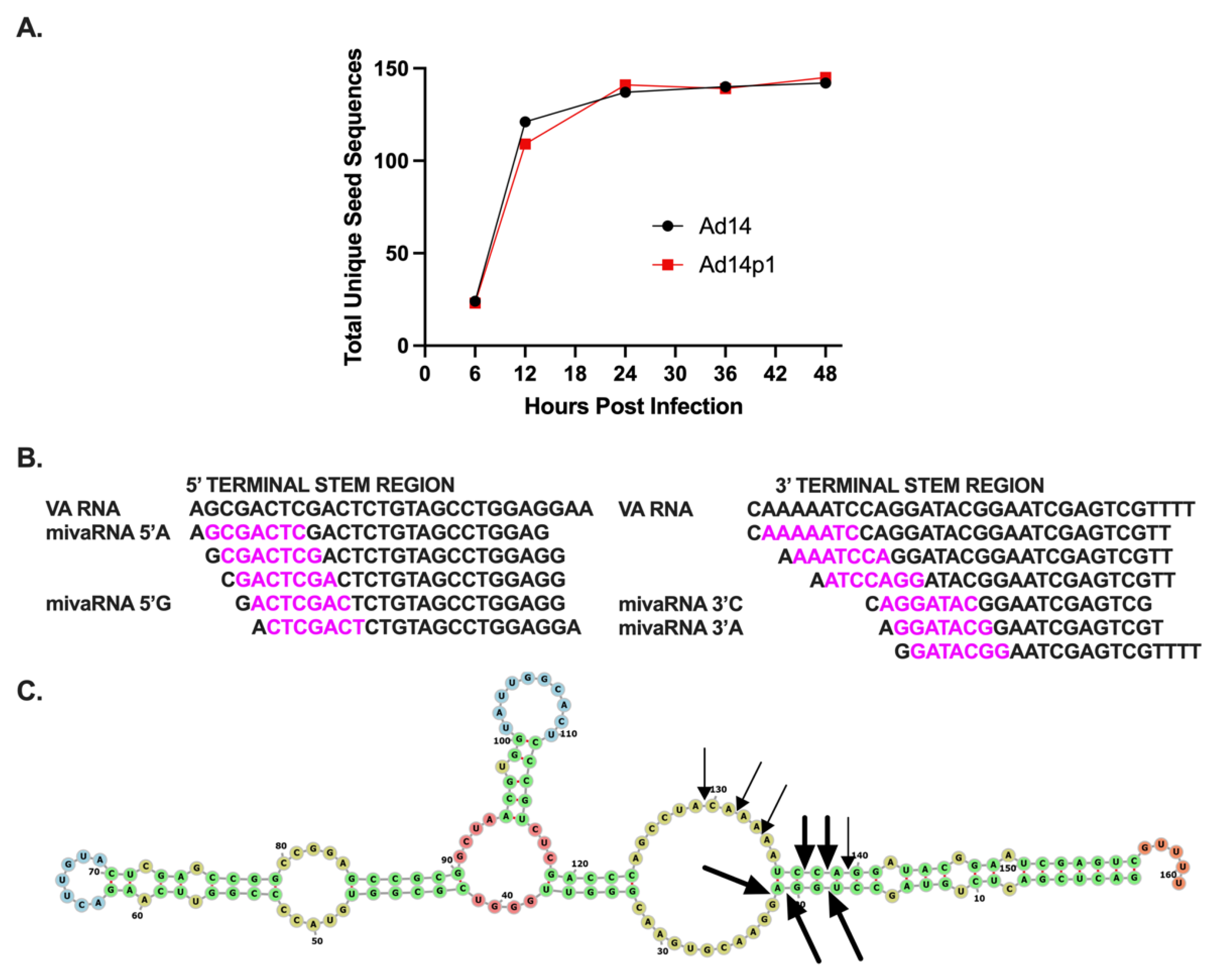
HAdV14, like other group B2 HAdV, only encode one VA RNA gene [
44]. In HAdVs that only encode one VA RNA, that VA RNA resembles group C HAdV VA RNAI. Here, further bioinformatic analysis of the HAdV14 VA RNA gene has shown that the VA RNA gene contains both RNA Pol III Box A and B promoter sites as well as two internal terminator sequences and a backup terminator sequence. The HAdV14 VA RNA gene also encodes two potential OAS1 consensus activation sequences. Based on the predicted secondary structure of the HAdV14 VA RNA molecule, it includes a terminal stem, central domain, and apical stem. Overall, the bioinformatic results suggest that, like other HAdVs that only encode one single VA RNA gene, HAdV14 VA RNA resembles that of VA RNAI, rather than VA RNAII.
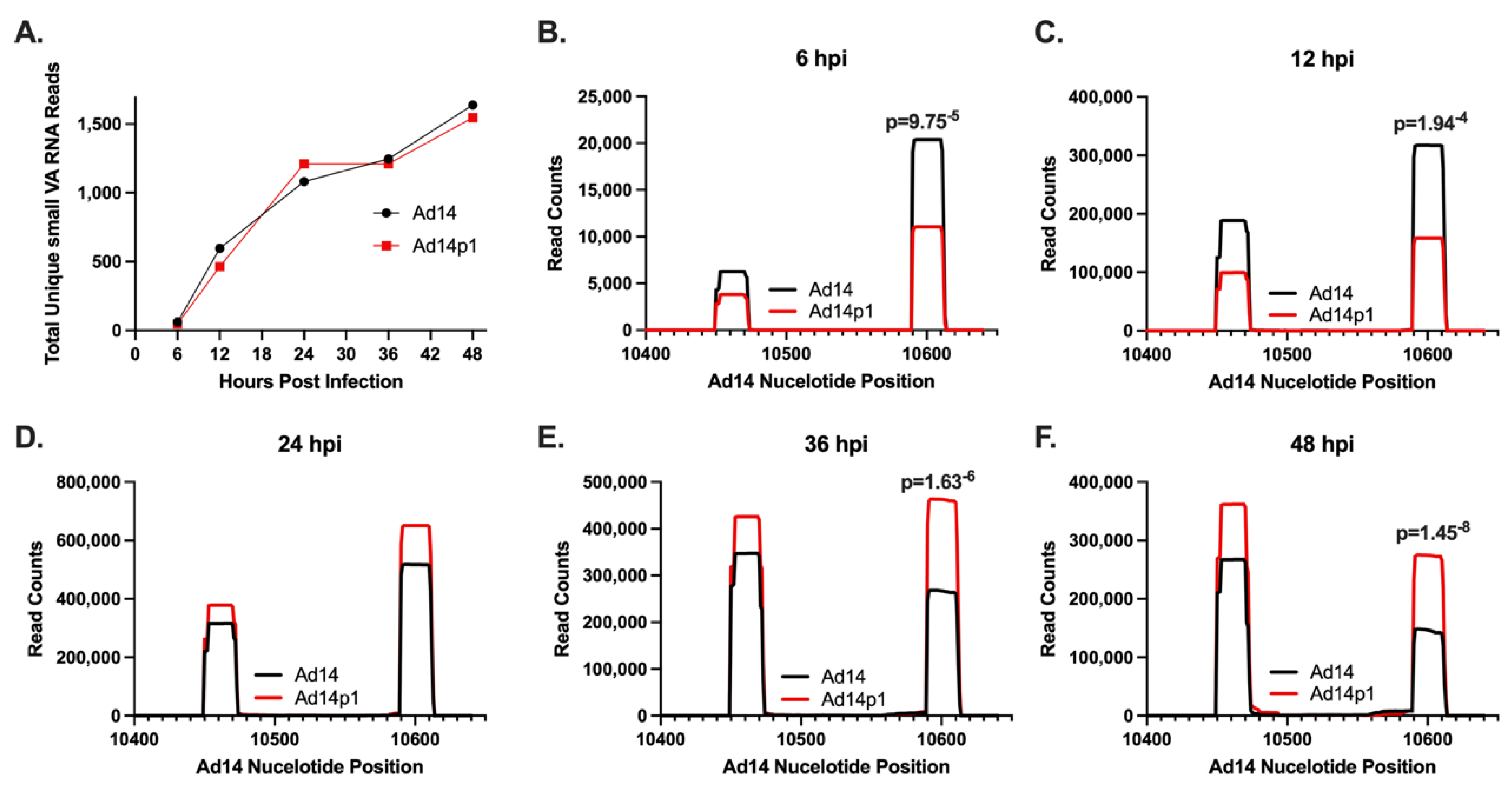
To characterize the mivaRNA produced from the HAdV14 VA RNA gene, we infected A549 cells and sampled mivaRNA expression through all phases of the Ad14 infectious cycle from very early (6 hpi) all the way through to full CPE (48 hpi). At all times, >90% of all small RNA reads that mapped to the HAdV14 genome mapped specifically to the VA RNA gene. These results are consistent with those seen during Ad5 infection of IMR-90 cells [
48]. Zhao and colleagues first observed that there was temporal expression of mivaRNA molecules during the infection cycle [
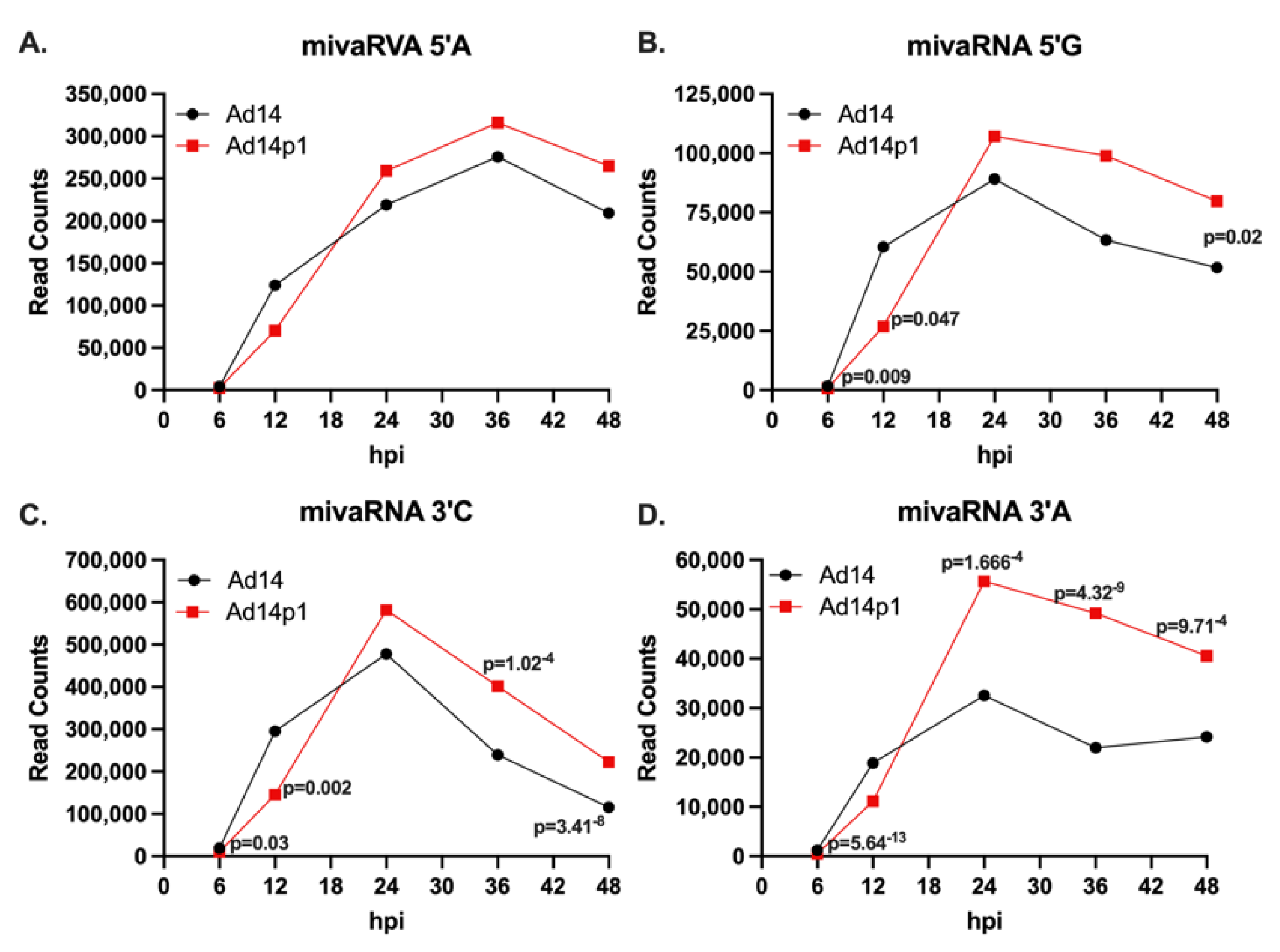
48]. We also observed temporal expression of mivaRNAs during HAdV14 infection. Our data showed that early during infection, the 3′ mivaRNAs are the most abundant HAdV14 mivaRNAs, while between 24 and 36 hpi, the 5′ mivaRNAs begin to be more abundant. This is opposite of what was observed following HAdV2-infected IMR-90 cells. The reason for this difference is unclear. The difference could be virus-specific (HAdV2 vs. HAdV14), cell type-specific (lung fibroblast, IMR-90, vs. lung epithelial, A549), or a combination. Supporting this theory are studies by Kamel et al., which showed that infection of 293 cells with HAdV5, HAdV11 (a group B2 HAdV), and HAdV37 resulted in more 3′ mivaRNA expression early during infection [
27]. The same temporal expression patterns were seen during HAdV14p1 infection, as was observed by Zhao [
48]. When we normalized mivaRNA expression, we observed that there was a significant difference in the expression of total mivaRNAs between HAdV14 and HAdV14p1 over time. Specifically, early in the infection cycle, HAdV14 mivaRNAs showed higher expression than HAdV14p1. After 24 hpi, this pattern is flipped, with HAdV14p1 mivaRNAs expressed at a higher level than HAdV14. Previous studies in our lab and others have shown that there is no difference in the replication rates of HAdV14 or HAdV14p1 in either tissue culture cells or in vivo in the lungs of Syrian hamsters, thus eliminating differences in viral replication as the reason for the differential expression of mivaRNAs [
8,
49]. The studies here were not designed to test whether this mivaRNA expression pattern has any role in the increased pathogenesis seen during HAdV14p1 infection. Studies are underway to test this possibility using a Syrian hamster model of differences in HAdV14p1 vs. HAdV14 lung pathogenesis [
8,
50].
Our results identified the core seed sequences produced during Ad14/Ad14p1 infection as coming from both the 5′ and 3′ terminal stems. Minor variants are also produced. How these arise is unclear. One possibility is that RNA pol III synthesis of VA RNAs might have some minor wobble in its exact initiation site, as these minor mivaRNA species show very slight variations in either the 5′ or 3′ ends. It is possible that this could have an effect on expression, as small differences in the 5′ start can result in a completely different mivaRNA seed sequence. We identified a group of long mivaRNAs produced from the 3′ terminal stem that start between 8 and 4 nucleotides upstream of the predominant 3′ mivaRNA. Interestingly, the longest mivaRNA produced from the 3′ terminal stem has the exact same seed sequencing as hsa-miR-584c-3p. It has been well documented that HAdV mivaRNAs can regulate gene expression, like gene-encoded miRNAs [
26,
40,
42]. Using a luciferase reporter vector, we have shown that both the 5′ and 3′ HAdV-14 mivaRNAs can suppress gene expression during viral infection. Our results confirm that the 3′ mivaRNAs are more abundantly expressed than 5′ mivaRNAs early during infection, as the 3′ mivaRNAs showed stronger repression of luciferase activity than the 5′ mivaRNAs. mivaRNAs have been shown to associate with RISC complexes, but not all mivaRNAs associate with RISC complexes at the same rate [
27,
38,
40,
42]. This could account for the difference in repressive activity seen between the mivaRNAs in our reporter assay. The association of HAdV14 mivaRNAs with the RISC complex was not a focus of these studies, but is under investigation.
The ability of mivaRNAs to target and regulate host gene expression is an attractive theory for potential roles in HAdV replication. As a result, numerous studies have been conducted to determine potential target genes for mivaRNAs. Bioinformatic, microarray studies, and RNA-seq analysis have shown that many of the potential cellular gene targets for mivaRNAs are involved in DNA repair, cell growth, cell death, RNA metabolism, cell signaling pathways, and metabolism [
26,
42,
48]. Our bioinformatic analysis (
Table 6) on the predicted HAdV14 mivaRNA targets show similar cellular pathways to be potential targets, despite the lack of sequencing identity between HAdV14 seed sequences and HAdV2/5 seed sequences. Our IPA analysis (
Table 7) also shows functional enrichment of the same cellular pathways from our RNA-seq data on HAdV14 mivaRNA target genes. This suggests that there is some conserved biological reason for mivaRNA expression during viral infection. However, studies have raised questions as to the role of mivaRNAs in viral replication. The mutation of the seed sequences of HAdV5 mivaRNAs had no impact on viral replication in HEK293 cells [
51]. One underlying problem with studies (including these studies) on HAdV mivaRNAs is that they have been conducted in tissue culture cell lines rather than relevant primary cells or in a permissive animal model, which might clarify the role of mivaRNAs during infection in vivo. HAdV mivaRNAs have also been found in persistently infected lymphoid cells, suggesting a potential role in regulating lytic and persistent infection [
52,
53]. In addition to mivaRNAs from the VA RNA gene, another viral small RNA that maps to the Major Late Promoter region has been identified that regulates viral gene expression; we are in the process of exploring if there are other HAdV14 small viral RNAs [
54].
In this study, we characterized the HAdV14 VA RNA gene and the mivaRNAs produced from it. We identified the predominant seed sequences and the cellular genes that the mivaRNAs can target. Our bioinformatic analysis using RNA-seq data of the expression of mivaRNA target genes showed that HAdV14 mivaRNA can regulate similar pathways seen from HAdV2/5 studies despite the lack of identity of the mivaRNA seed sequences. Despite 100% sequence identity later in infection, HAdV14p1-infected cells express significantly more mivaRNA than HAdV14-infected cells. Whether this difference has an impact on viral replication or pathogenesis is unknown. Due to a lack of studies in immunocompetent animal models, the role of HAdV mivaRNA in viral pathogenesis is largely unexplored. To answer questions about viral factors, such as mivaRNAs, we have developed an immunocompetent Syrian hamster model of HAdV14p1 lung pathogenesis that displays pathology similar to what is seen in the lungs of severely infected human patients [
8,
50]. We propose that the Syrian hamster model of HAdV14p1 lung pathogenesis could provide the model system to explore the roles of mivaRNA in viral replication and pathogenesis in vivo.