Anti-SARS-CoV-2 Titers Predict the Severity of COVID-19
Abstract
:1. Introduction
2. Materials and Methods
2.1. Study Design and Participants
2.2. Serological Testing
2.3. Statistical Analysis
2.4. Ethics
3. Results
3.1. Patient Characteristics
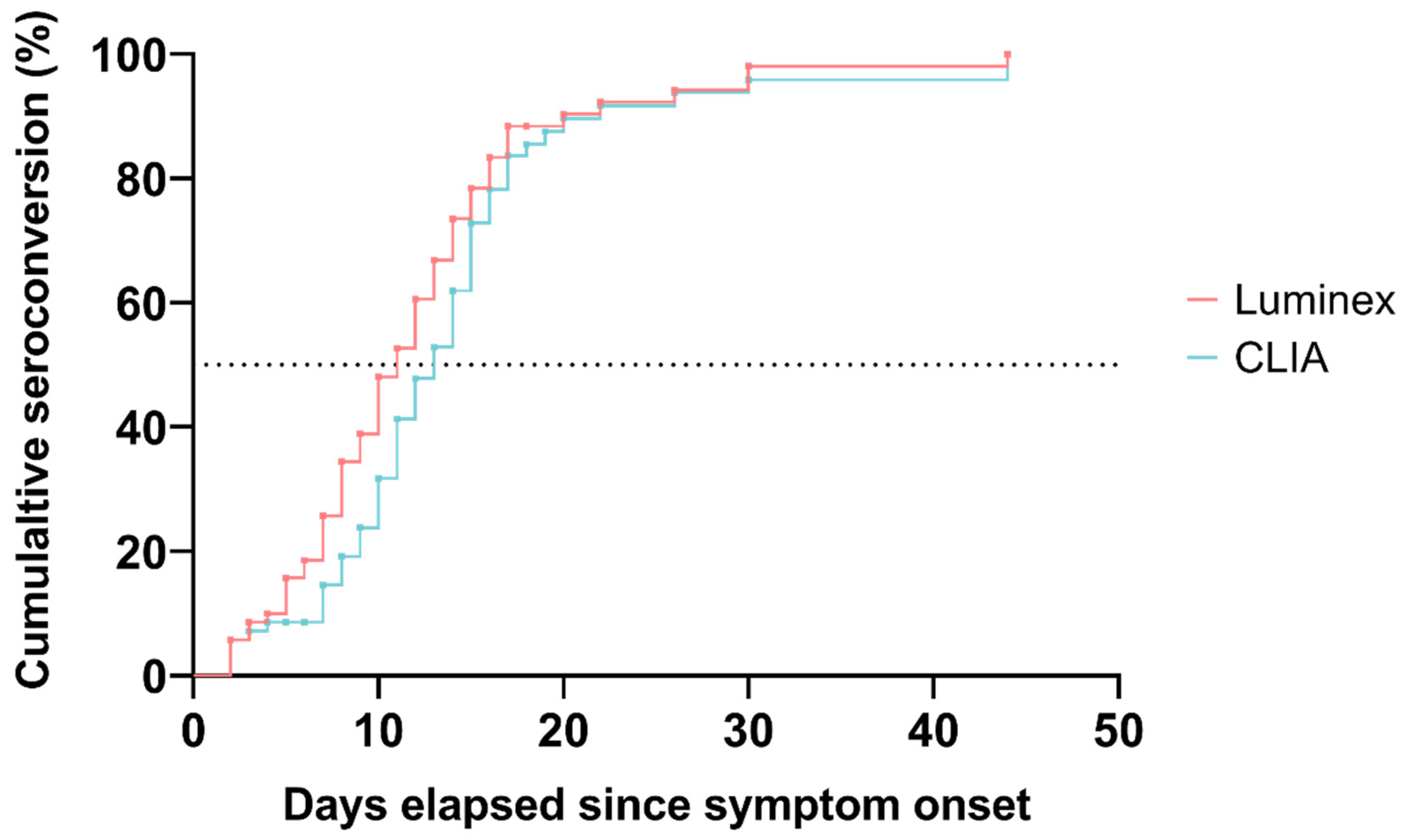
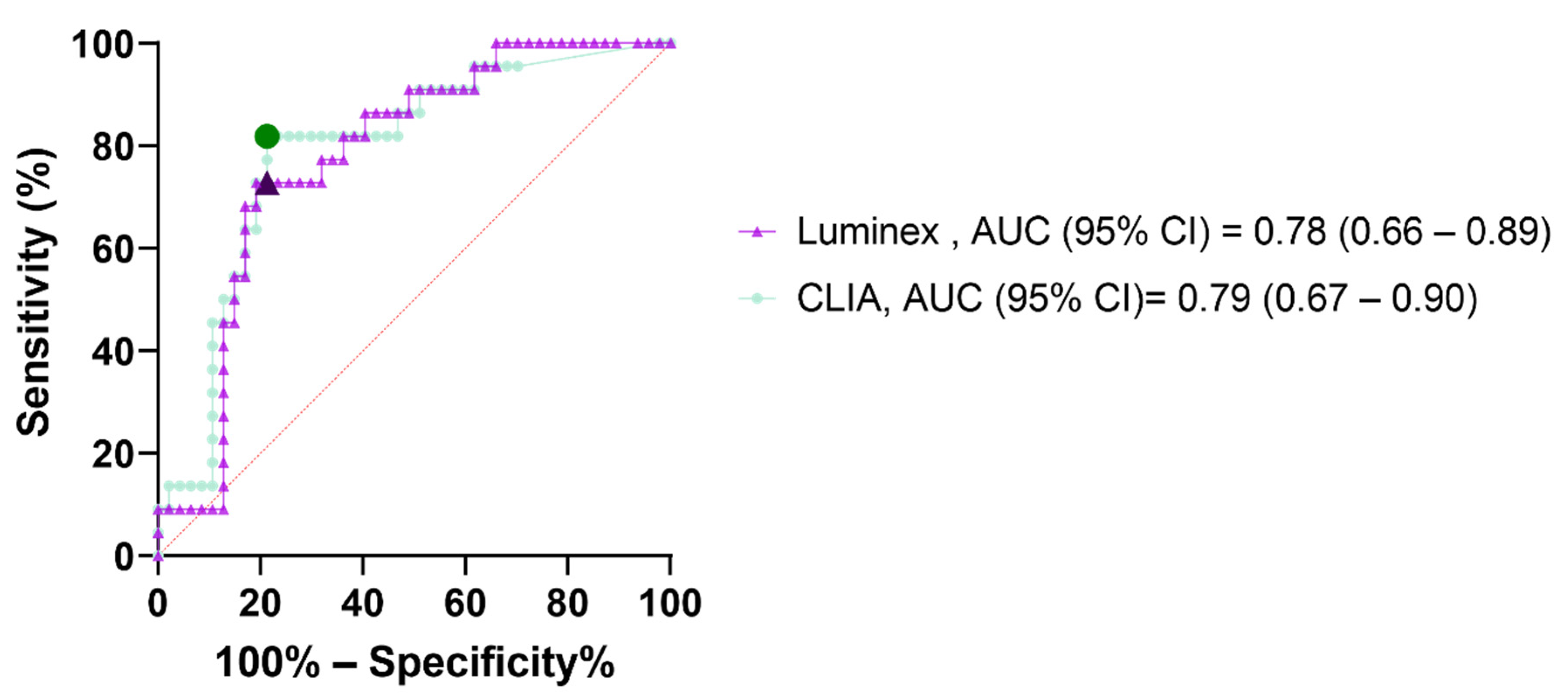
3.2. Diagnostic Performances and Kinetics of Serological Assays
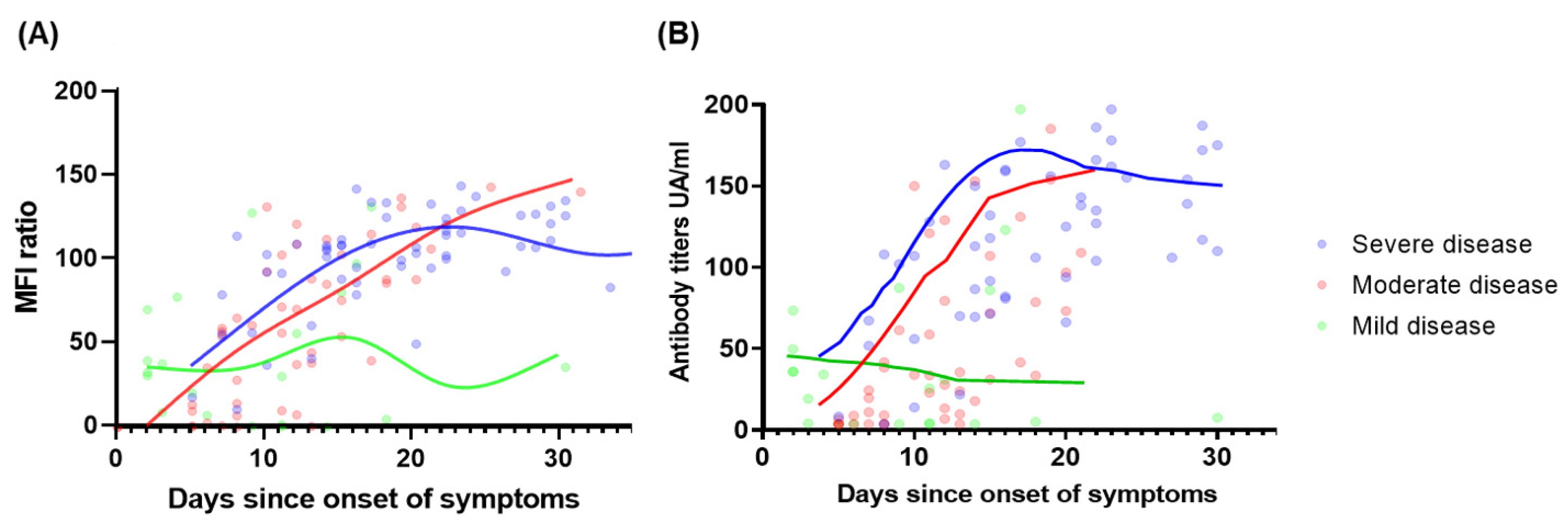
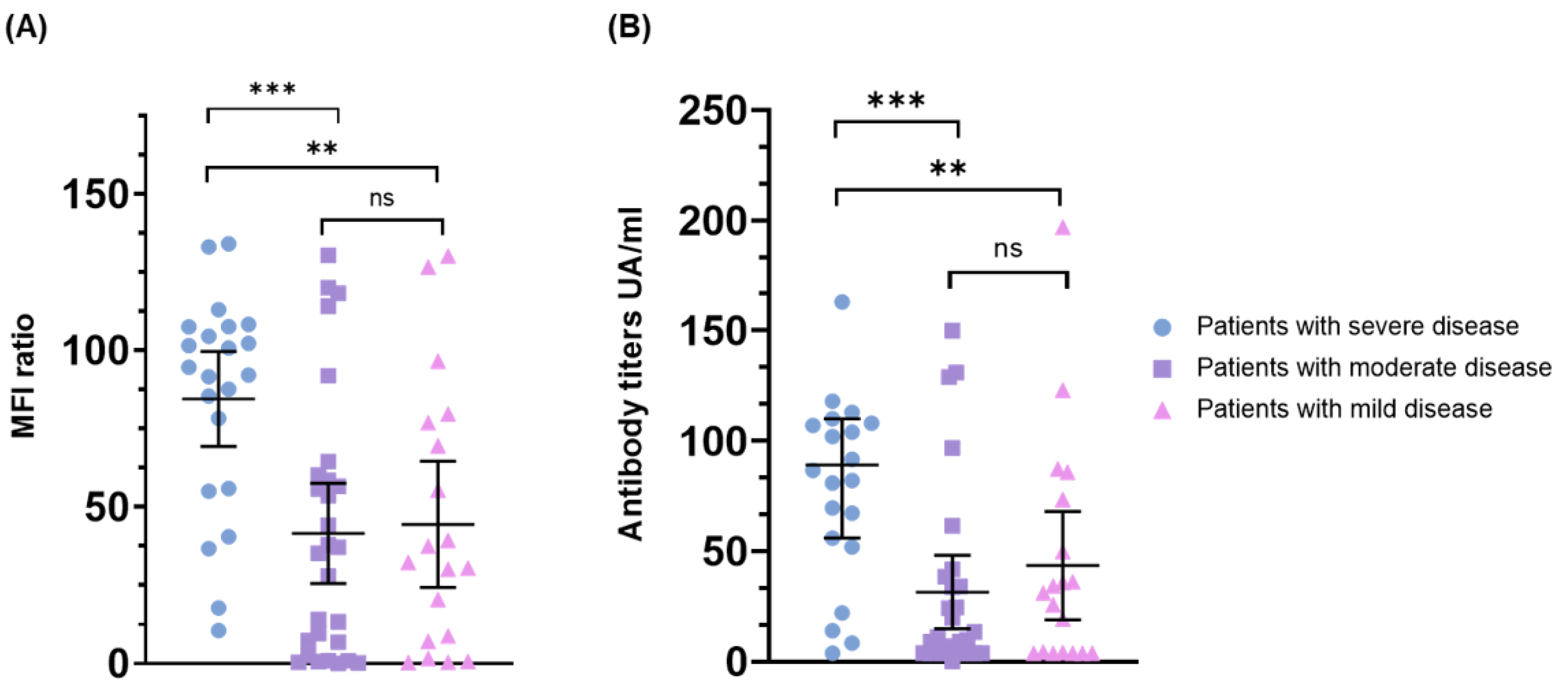
3.3. Correlation of Baseline Antibody Levels with the Severity of Disease
4. Discussion
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Hu, B.; Guo, H.; Zhou, P.; Shi, Z.-L. Characteristics of SARS-CoV-2 and COVID-19. Nat. Rev. Microbiol. 2021, 19, 141–154. [Google Scholar] [CrossRef] [PubMed]
- Huang, C.; Wang, Y.; Li, X.; Ren, L.; Zhao, J.; Hu, Y.; Zhang, L.; Fan, G.; Xu, J.; Gu, X.; et al. Clinical features of patients infected with 2019 novel coronavirus in Wuhan, China. Lancet 2020, 395, 497–506. [Google Scholar] [CrossRef] [Green Version]
- Guan, W.J.; Ni, Z.Y.; Hu, Y.; Liang, W.H.; Qu, C.Q.; He, J.X.; Liu, L.; Shan, H.; Lei, C.L.; Hui, D.S.C.; et al. Clinical Characteristics of coronavirus disease in China. N. Engl. J. Med. 2020, 382, 1708–1720. [Google Scholar] [CrossRef]
- Richardson, S.; Hirsch, J.S.; Narasimhan, M.; Crawford, J.M.; McGinn, T.; Davidson, K.W.; The Northwell COVID-19 Research Consortium. Presenting Characteristics, Comorbidities, and Outcomes Among 5700 Patients Hospitalized with COVID-19 in the New York City Area. JAMA 2020, 323, 2052–2059. [Google Scholar] [CrossRef] [PubMed]
- Garcia-Beltran, W.F.; Lam, E.C.; Astudillo, M.G.; Yang, D.; Miller, T.E.; Feldman, J.; Hauser, B.M.; Caradonna, T.M.; Clayton, K.L.; Nitido, A.D.; et al. COVID-19-neutralizing antibodies predict disease severity and survival. Cell 2021, 184, 476–488.e11. [Google Scholar] [CrossRef] [PubMed]
- Arvin, A.M.; Fink, K.; Schmid, M.A.; Cathcart, A.; Spreafico, R.; Havenar-Daughton, C.; Lanzavecchia, A.; Corti, D.; Virgin, H.W. A perspective on potential antibody-dependent enhancement of SARS-CoV-2. Nature 2020, 584, 353–363. [Google Scholar] [CrossRef]
- Iwasaki, A.; Yang, Y. The potential danger of suboptimal antibody responses in COVID-19. Nat. Rev. Immunol. 2020, 20, 339–341. [Google Scholar] [CrossRef] [Green Version]
- Liu, L.; Wei, Q.; Lin, Q.; Fang, J.; Wang, H.; Kwok, H.; Tang, H.; Nishiura, K.; Peng, J.; Tan, Z.; et al. Anti–spike IgG causes severe acute lung injury by skewing macrophage responses during acute SARS-CoV infection. JCI Insight 2019, 4, 4. [Google Scholar] [CrossRef]
- Darmon, M.; Dumas, G. Anticipating outcomes for patients with COVID-19 and identifying prognosis patterns. Lancet Infect. Dis. 2021, 21, 744–745. [Google Scholar] [CrossRef]
- Gutiérrez-Gutiérrez, B.; del Toro, M.D.; Borobia, A.M.; Carcas, A.; Jarrín, I.; Yllescas, M.; Ryan, P.; Pachón, J.; Carratalà, J.; Berenguer, J.; et al. Identification and validation of clinical phenotypes with prognostic implications in patients admitted to hospital with COVID-19: A multicentre cohort study. Lancet Infect. Dis. 2021, 21, 783–792. [Google Scholar] [CrossRef]
- Dupont, T.; Caillat-Zucman, S.; Fremeaux-Bacchi, V.; Morin, F.; Lengliné, E.; Darmon, M.; de Latour, R.P.; Zafrani, L.; Azoulay, E.; Dumas, G. Identification of Distinct Immunophenotypes in Critically Ill Coronavirus Disease 2019 Patients. Chest 2020, 159, 1884–1893. [Google Scholar] [CrossRef] [PubMed]
- Wynants, L.; Van Calster, B.; Collins, G.S.; Riley, R.D.; Heinze, G.; Schuit, E.; Bonten, M.M.J.; Dahly, D.L.; Damen, J.A.; Debray, T.P.A.; et al. Prediction models for diagnosis and prognosis of covid-19: Systematic review and critical appraisal. BMJ 2020, 369, m1328. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Knight, S.R.; Ho, A.; Pius, R.; Buchan, I.; Carson, G.; Drake, T.M.; Dunning, J.; Fairfield, C.J.; Gamble, C.; Green, C.A.; et al. Risk stratification of patients admitted to hospital with covid-19 using the ISARIC WHO Clinical Characterisation Protocol: Development and validation of the 4C Mortality Score. BMJ 2020, 370, m3339. [Google Scholar] [CrossRef] [PubMed]
- Post, N.; Eddy, D.; Huntley, C.; van Schalkwyk, M.C.I.; Shrotri, M.; Leeman, D.; Rigby, S.; Williams, S.V.; Bermingham, W.H.; Kellam, P.; et al. Antibody response to SARS-CoV-2 infection in humans: A systematic review. PLoS ONE 2020, 15, e0244126. [Google Scholar] [CrossRef]
- Ma, H.; Zeng, W.; He, H.; Zhao, D.; Jiang, D.; Zhou, P.; Cheng, L.; Li, Y.; Ma, X.; Jin, T. Serum IgA, IgM, and IgG responses in COVID-19. Cell. Mol. Immunol. 2020, 17, 773–775. [Google Scholar] [CrossRef]
- Huang, J.; Mao, T.; Li, S.; Wu, L.; Xu, X.; Li, H.; Xu, C.; Su, F.; Dai, J.; Shi, J.; et al. Long period dynamics of viral load and antibodies for SARS-CoV-2 infection: An observational cohort study. medRxiv 2020. [Google Scholar] [CrossRef]
- Lynch, K.L.; Whitman, J.D.; Lacanienta, N.P.; Beckerdite, E.W.; Kastner, S.A.; Shy, B.R.; Goldgof, G.M.; Levine, A.G.; Bapat, S.P.; Stramer, S.L.; et al. Magnitude and kinetics of anti-SARS-CoV-2 antibody responses and their relationship to disease severity. medRxiv 2020. [Google Scholar] [CrossRef]
- Ou, J.; Tan, M.; He, H.; Tan, H.; Mai, J.; Long, Y.; Jiang, X.; He, Q.; Huang, Y.; Li, Y.; et al. Study on the expression levels of antibodies against SARS-CoV-2 at different period of disease and its related factors in 192 cases of COVID-19 patients. medRxiv 2020. [Google Scholar] [CrossRef]
- Tan, W.-T.; Lu, Y.; Zhang, J.; Wang, J.; Dan, Y.; Tan, Z.; He, X.; Qian, C.; Sun, Q.; Hu, Q.; et al. Viral Kinetics and Antibody Responses in Patients with COVID-19. medRxiv 2020. [Google Scholar] [CrossRef] [Green Version]
- Xie, J.; Ding, C.; Li, J.; Wang, Y.; Guo, H.; Lu, Z.; Wang, J.; Zheng, C.; Jin, T.; Gao, Y.; et al. Characteristics of patients with coronavirus disease (COVID-19) confirmed using an IgM-IgG antibody test. J. Med. Virol. 2020, 92, 2004–2010. [Google Scholar] [CrossRef]
- Perera, R.A.; Mok, C.K.; Tsang, O.T.; Lv, H.; Ko, R.L.; Wu, N.C.; Yuan, M.; Leung, W.S.; Mc Chan, J.; Chik, T.S.; et al. Serological assays for severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2), March 2020. Eurosurveillance 2020, 25, 2000421. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Rijkers, G.; Murk, J.-L.; Wintermans, B.; van Looy, B.; van den Berge, M.; Veenemans, J.; Stohr, J.; Reusken, C.; van der Pol, P.; Reimerink, J. Differences in antibody kinetics and functionality between severe and mild SARS-CoV-2 infections. medRxiv 2020. [Google Scholar] [CrossRef]
- Cervia, C.; Nilsson, J.; Zurbuchen, Y.; Valaperti, A.; Schreiner, J.; Wolfensberger, A.; Raeber, M.E.; Adamo, S.; Weigang, S.; Emmenegger, M.; et al. Systemic and mucosal antibody responses specific to SARS-CoV-2 during mild versus severe COVID-19. J. Allergy Clin. Immunol. 2021, 147, 545–557.e9. [Google Scholar] [CrossRef] [PubMed]
- He, R.; Lu, Z.; Zhang, L.; Fan, T.; Xiong, R.; Shen, X.; Feng, H.; Meng, H.; Lin, W.; Jiang, W.; et al. The clinical course and its correlated immune status in COVID-19 pneumonia. J. Clin. Virol. 2020, 127, 104361. [Google Scholar] [CrossRef] [PubMed]
- Laing, A.G.; Lorenc, A.; Del Molino Del Barrio, I.; Das, A.; Fish, M.; Monin, L.; Muñoz-Ruiz, M.; McKenzie, D.R.; Hayday, T.S.; Francos-Quijorna, I.; et al. A consensus Covid-19 immune signature combines immuno-protection with discrete sepsis-like traits associated with poor prognosis. medRxiv 2020. [Google Scholar] [CrossRef]
- Minervina, A.A.; Komech, E.E.; Titov, A.; Koraichi, M.B.; Rosati, E.; Mamedov, I.Z.; Franke, A.; Efimov, G.A.; Chudakov, D.M.; Mora, T.; et al. Longitudinal high-throughput TCR repertoire profiling reveals the dynamics of T-cell memory formation after mild COVID-19 infection. eLife 2021, 10, e63502. [Google Scholar] [CrossRef]
- Sun, B.; Feng, Y.; Mo, X.; Zheng, P.; Wang, Q.; Li, P.; Peng, P.; Liu, X.; Chen, Z.; Huang, H.; et al. Kinetics of SARS-CoV-2 specific IgM and IgG responses in COVID-19 patients. Emerg. Microbes Infect. 2020, 9, 940–948. [Google Scholar] [CrossRef]
- Bao, J.; Li, C.; Zhang, K.; Kang, H.; Chen, W.; Gu, B. Comparative analysis of laboratory indexes of severe and non-severe patients infected with COVID-19. Clin. Chim. Acta 2020, 509, 180–194. [Google Scholar] [CrossRef]
- Qu, J.; Wu, C.; Li, X.; Zhang, G.; Jiang, Z.; Li, X.; Zhu, Q.; Liu, L. Profile of Immunoglobulin G and IgM Antibodies Against Severe Acute Respiratory Syndrome Coronavirus 2 (SARS-CoV-2). Clin. Infect. Dis. 2020, 71, 2255–2258. [Google Scholar] [CrossRef]
- Grein, J.; Ohmagari, N.; Shin, D.; Diaz, G.; Asperges, E.; Castagna, A.; Feldt, T.; Green, G.; Green, M.L.; Lescure, F.X.; et al. Compassionate Use of Remdesivir for Patients with Severe Covid-19. N. Engl. J. Med. 2020, 382, 2327–2336. [Google Scholar] [CrossRef]
- WHO; R&D Blueprint. COVID-19 Theurapeutic Trial Synopsis; WHO: Geneva, Switzerland, 2020. [Google Scholar]
- Fenwick, C.; Croxatto, A.; Coste, A.T.; Pojer, F.; André, C.; Pellaton, C.; Farina, A.; Campos, J.; Hacker, D.; Lau, K.; et al. Changes in SARS-CoV-2 Spike versus Nucleoprotein Antibody Responses Impact the Estimates of Infections in Population-Based Seroprevalence Studies. J. Virol. 2021, 95, e01828-20. [Google Scholar] [CrossRef] [PubMed]
- Hanson, K.E.; Caliendo, A.M.; Arias, C.A.; Englund, J.A.; Hayden, M.K.; Lee, M.J.; Loeb, M.; Patel, R.; Altayar, O.; El Alayli, A.; et al. Infectious Diseases Society of America Guidelines on the Diagnosis of COVID-19:Serologic Testing. Clin. Infect. Dis. 2020. [Google Scholar] [CrossRef]
- Winter, A.K.; Hegde, S.T. The important role of serology for COVID-19 control. Lancet Infect. Dis. 2020, 20, 758–759. [Google Scholar] [CrossRef]
- Caruana, G.; Croxatto, A.; Coste, A.T.; Opota, O.; Lamoth, F.; Jaton, K.; Greub, G. Diagnostic strategies for SARS-CoV-2 infection and interpretation of microbiological results. Clin. Microbiol. Infect. 2020, 26, 1178–1182. [Google Scholar] [CrossRef] [PubMed]
- Coste, A.T.; Jaton, K.; Papadimitriou-Olivgeris, M.; Croxatto, A.; Greub, G. Indication for SARS-CoV-2 serology: First month follow-up. Clin. Microbiol. Infect. Dis. 2020, 5, 1–4. [Google Scholar] [CrossRef]
- Greub, G.; Coste, A.; Croxatto, A. Sérologie SARS-CoV-2: Quand la prescrire? Pipette-Swiss Laboratoy Med. 2020, 2, 14–15. [Google Scholar]
- Kritikos, A.; Caruana, G.; Brouillet, R.; Miroz, J.-P.; Abed-Maillard, S.; Stieger, G.; Opota, O.; Croxatto, A.; Vollenweider, P.; Bart, P.-A.; et al. Sensitivity of Rapid Antigen Testing and RT-PCR Performed on Nasopharyngeal Swabs versus Saliva Samples in COVID-19 Hospitalized Patients: Results of a Prospective Comparative Trial (RESTART). Microorganisms 2021, 9, 1910. [Google Scholar] [CrossRef]
- Miller, T.E.; Beltran, W.F.G.; Bard, A.Z.; Gogakos, T.; Anahtar, M.N.; Astudillo, M.G.; Yang, D.; Thierauf, J.; Fisch, A.S.; Mahowald, G.K.; et al. Clinical sensitivity and interpretation of PCR and serological COVID-19 diagnostics for patients presenting to the hospital. FASEB J. 2020, 34, 13877–13884. [Google Scholar] [CrossRef]
- Coste, A.T.; Jaton, K.; Papadimitriou-Olivgeris, M.; Greub, G.; Croxatto, A. Comparison of SARS-CoV-2 serological tests with different antigen targets. J. Clin. Virol. 2020, 134, 104690. [Google Scholar] [CrossRef]
- Zhao, J.; Yuan, Q.; Wang, H.; Liu, W.; Liao, X.; Su, Y.; Wang, X.; Yuan, J.; Li, T.; Li, J.; et al. Antibody Responses to SARS-CoV-2 in Patients with Novel Coronavirus Disease 2019. Clin. Infect. Dis. 2020, 71, 2027–2034. [Google Scholar] [CrossRef]
- Béné, M.C.; Bittencourt, M.D.C.; Eveillard, M.; Le Bris, Y. Good IgA Bad IgG in SARS-CoV-2 Infection? Clin. Infect. Dis. 2020, 71, 897–898. [Google Scholar] [CrossRef] [Green Version]
- Recovery Collaborative Group. Tocilizumab in patients admitted to hospital with COVID-19 (RECOVERY): A randomised, controlled, open-label, platform trial. Lancet 2021, 397, 1637–1645. [Google Scholar] [CrossRef]
- Recovery Collaborative Group. Casirivimab and imdevimab in patients admitted to hospital with COVID-19 (RECOVERY): A randomised, controlled, open-label, platform trial. Lancet 2022, 399, 665–676. [Google Scholar] [CrossRef]
- ACTIV-3/TICO Bamlanivimab Study Group; Lundgren, J.D.; Grund, B.; Barkauskas, C.E.; Holland, T.L.; Gottlieb, R.L.; Sandkovsky, U.; Brown, S.M.; Knowlton, K.U.; Self, W.H.; et al. Responses to a Neutralizing Monoclonal Antibody for Hospitalized Patients With COVID-19 According to Baseline Antibody and Antigen Levels: A Randomized Controlled Trial. Ann. Intern. Med. 2022, 175, 234–243. [Google Scholar] [CrossRef]
- Mudatsir, M.; Fajar, J.K.; Wulandari, L.; Soegiarto, G.; Ilmawan, M.; Purnamasari, Y.; Mahdi, B.A.; Jayanto, G.D.; Suhendra, S.; Setianingsih, Y.A.; et al. Predictors of COVID-19 severity: A systematic review and meta-analysis. F1000Research 2020, 9, 1107. [Google Scholar] [CrossRef] [PubMed]
- Marin, B.G.; Aghagoli, G.; Lavine, K.; Yang, L.; Siff, E.J.; Chiang, S.S.; Salazar-Mather, T.P.; Dumenco, L.; Savaria, M.C.; Aung, S.N.; et al. Predictors of COVID -19 severity: A literature review. Rev. Med. Virol. 2021, 31, 1–10. [Google Scholar] [CrossRef] [PubMed]
| Characteristics, n (%) | Total (n = 70) | Severe Disease (n = 22) | Moderate Disease (n = 29) | Mild Disease (n = 19) | p_Value |
| Male sex | 40 (57) | 14 (64) | 16 (55) | 10 (53) | 0.74 |
| Median age, year (IQR) | 60 (51–73) | 60 (54–64) | 60 (51–74) | 59 (43–78) | 0.80 |
| Obesity (BMI ≥ 30) | 25 (36) | 8 (36) | 15 (52) | 2 (11) | 0.01 |
| Active or prior smoker | 12 (17) | 2 (9) | 8 (28) | 2 (11) | 0.14 |
| Hypertension | 34 (49) | 11 (50) | 16 (55) | 7 (37) | 0.45 |
| History of cardiac rhythm or conduction disorder | 8 (11) | 1 (5) | 6 (21) | 1 (5) | 0.12 |
| Coronary disease | 5 (7) | 0 (0) | 4 (14) | 1 (5) | 0.40 |
| Heart failure | 8 (11) | 1 (5) | 6 (21) | 1 (5) | 0.12 |
| Stroke | 3 (4) | 0 (0) | 2 (7) | 1 (5) | 0.92 |
| Chronic kidney disease (eGFR < 60) | 10 (14) | 1 (5) | 5 (17) | 4 (21) | 0.27 |
| Chronic pulmonary disease * | 10 (14) | 3 (14) | 6 (21) | 1 (5) | 0.32 |
| Dyslipidemia | 17 (24) | 3 (14) | 13 (45) | 1 (5) | 0.01 |
| History of thromboembolic event | 5 (7) | 1 (5) | 4 (14) | 0 (0) | 0.40 |
| Malignancy | 8 (11) | 2 (9) | 2 (7) | 4 (21) | 0.29 |
| Diabetes | 17 (24) | 4 (18) | 10 (34) | 3 (16) | 0.24 |
| Connective tissue disease | 2 (3) | 0 (0) | 2 (7) | 0 (0) | 0.91 |
| Transplantation (SOT and HSCT) | 3 (4) | 1 (5) | 1 (3) | 1 (5) | 0.95 |
| Characteristics | Total (n = 70) | Severe Disease (n = 22) | Moderate Disease (n = 29) | Mild Disease (n = 19) | p_Value |
|---|---|---|---|---|---|
| Median duration of symptoms before admission (IQR) | 10 (6–13) | 13 (9–15) | 8 (7–12) | 9 (3–11) | 0.55 |
| Fever, n (%) | 48 (69) | 19 (86) | 19 (66) | 10 (53) | 0.06 |
| Angina, n (%) | 7 (10) | 2 (9) | 4 (14) | 1 (5) | 0.61 |
| Rhinorrhea, n (%) | 5 (7) | 1 (5) | 3 (10) | 1 (5) | 0.67 |
| Cough (productive or not), n (%) | 49 (70) | 18 (82) | 22 (76) | 9 (47) | 0.03 |
| Anosmia/dysgeusia, n (%) | 9 (13) | 1 (5) | 6 (21) | 2 (11) | 0.21 |
| Dyspnea, n (%) | 42 (60) | 18 (82) | 20 (69) | 4 (21) | <0.01 |
| Diarrhea, n (%) | 16 (23) | 4 (18) | 8 (28) | 4 (21) | 0.71 |
| Radiological infiltrate, n (%) | 49/61 (80) | 20/21 (95) | 24/27 (89) | 5/13 (38) | <0.01 |
| White Blood cell count, G/L | 7.6 (5.5–9.2) | 8 (6.9–10.5) | 7.4 (5.2–8.8) | 6.9 (5.1–9.1) | 0.10 |
| Lymphocytes count, G/L | 0.9 (0.7–1.2) | 0.9 (0.5–1.1) | 0.9 (0.7–1.2) | 1.1 (0.7–1.8) | 0.40 |
| Platelets count, G/L | 216 (161–308) | 209 (158–291) | 202 (156–280) | 244 (212–323) | 0.53 |
| D-dimer, ng/ml | 1016 (521–1799) | 1127 (828–2677) | 752 (507–1656) | 881 (468–1419) | 0.03 |
| Creatinine, µmol/L | 88 (71–119) | 93 (81–105) | 88 (77–124) | 74 (69–115) | 0.66 |
| CRP, mg/L | 86 (38–151) | 156 (83–273) | 88 (53–125) | 11 (4–71) | <0.01 |
| Procalcitonin, µg/L | 0.2 (0.1–0.5) | 0.5 (0.2–1.3) | 0.2 (0.1–0.3) | 0.2 (0.1–0.2) | 0.39 |
| Serum ferritin, µg/L | 997 (434–1690) | 1296 (673–2117) | 976 (365–1639) | 823 (397–1198) | 0.10 |
| Troponin, ng/ml | 12 (6–40) | 15 (9–125) | 10 (6–30) | 11 (6–20) | 0.06 |
| AST, U/L | 57 (33–89) | 71 (53–88) | 54 (31–92) | 36 (25–53) | 0.50 |
| ALT, U/L | 38 (22–65) | 50 (37–69) | 37 (20–58) | 26 (17–32) | 0.49 |
| Bilirubin, µmol/L | 9 (7–13) | 9 (6–13) | 9.5 (6–12) | 8 (7–10) | 0.60 |
| ABO Group O, n (%) | 22/38 (58) | 6/16 (38) | 8/10 (80) | 8/12 (67) | 0.25 |
| Selected Test (Threshold Value) | Sensitivity % (95% CI) | Specificity % (95% CI) | PPV % (95% CI) | NPV % (95% CI) |
|---|---|---|---|---|
| CLIA (>50.8 UA/mL) | 82 (61–93) | 79 (65–88) | 63 (49–76) | 91 (80–96) |
| Luminex (MFI ratio >77.6) | 72 (52–87) | 81 (67–90) | 64 (48–77) | 86 (76–93) |
| D-dimer (>1000 ng/mL) | 71 (48–89) | 59 (42–75) | 44 (33–56) | 82 (69–91) |
| CRP (>100 mg/L) | 68 (45–86) | 73 (57–85) | 56 (42–69) | 82 (71–90) |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2022 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Kritikos, A.; Gabellon, S.; Pagani, J.-L.; Monti, M.; Bochud, P.-Y.; Manuel, O.; Coste, A.; Greub, G.; Perreau, M.; Pantaleo, G.; et al. Anti-SARS-CoV-2 Titers Predict the Severity of COVID-19. Viruses 2022, 14, 1089. https://doi.org/10.3390/v14051089
Kritikos A, Gabellon S, Pagani J-L, Monti M, Bochud P-Y, Manuel O, Coste A, Greub G, Perreau M, Pantaleo G, et al. Anti-SARS-CoV-2 Titers Predict the Severity of COVID-19. Viruses. 2022; 14(5):1089. https://doi.org/10.3390/v14051089
Chicago/Turabian StyleKritikos, Antonios, Sophie Gabellon, Jean-Luc Pagani, Matteo Monti, Pierre-Yves Bochud, Oriol Manuel, Alix Coste, Gilbert Greub, Matthieu Perreau, Giuseppe Pantaleo, and et al. 2022. "Anti-SARS-CoV-2 Titers Predict the Severity of COVID-19" Viruses 14, no. 5: 1089. https://doi.org/10.3390/v14051089
APA StyleKritikos, A., Gabellon, S., Pagani, J.-L., Monti, M., Bochud, P.-Y., Manuel, O., Coste, A., Greub, G., Perreau, M., Pantaleo, G., Croxatto, A., & Lamoth, F. (2022). Anti-SARS-CoV-2 Titers Predict the Severity of COVID-19. Viruses, 14(5), 1089. https://doi.org/10.3390/v14051089








