Clearance of Persistent SARS-CoV-2 RNA Detection in a NFκB-Deficient Patient in Association with the Ingestion of Human Breast Milk: A Case Report
Abstract
:1. Introduction
2. Materials and Methods
2.1. Ethics
2.2. Patient
2.3. COVID-19 Convalescent Plasma
2.4. Human Breast Milk
2.5. RNA Extraction and RT-qPCR
2.6. SARS-CoV-2 Genomic Sequencing and Phylogenetic and Genomic Analyses
2.7. Flow Cytometry Analysis
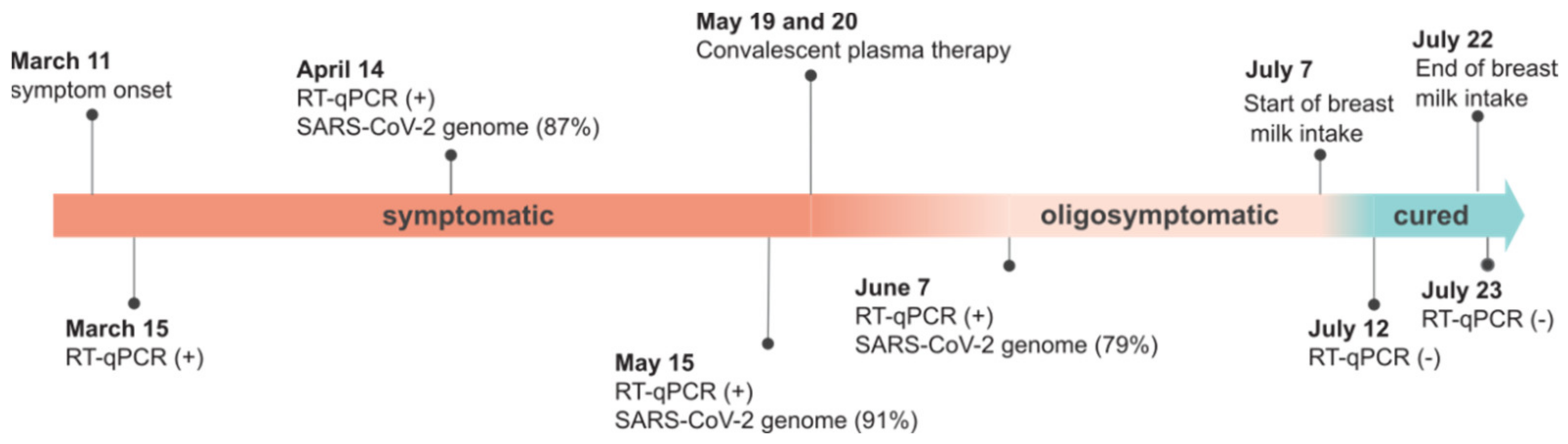
3. Case Report
4. Discussion
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Brown, K.; Yahyouche, A.; Haroon, S.; Camaradou, J.; Turner, G. Long COVID and self-management. Lancet 2022, 399, 355. [Google Scholar] [CrossRef]
- Yang, B.; Fan, J.; Huang, J.; Guo, E.; Fu, Y.; Liu, S.; Xiao, R.; Liu, C.; Lu, F.; Qin, T.; et al. Clinical and molecular characteristics of COVID-19 patients with persistent SARS-CoV-2 infection. Nat. Commun. 2021, 12, 3501. [Google Scholar] [CrossRef] [PubMed]
- See, R.H.; Zakhartchouk, A.N.; Petrick, M.; Lawrence, D.J.; Mok, C.P.Y.; Hogan, R.J.; Rowe, T.; Zitzow, L.A.; Karunakaran, K.P.; Hitt, M.M.; et al. Comparative evaluation of two severe acute respiratory syndrome (SARS) vaccine candidatesin mice challenged with SARS coronavirus. J. Gen. Virol. 2006, 87, 641–650. [Google Scholar] [CrossRef] [PubMed]
- Asahi-Ozaki, Y.; Yoshikawa, T.; Iwakura, Y.; Suzuki, Y.; Tamura, S.; Kurata, T.; Sata, T. Secretory IgA Antibodies Provide Cross-ProtectionAgainst Infection with Different Strainsof Influenza B Virus. J. Med. Virol. 2004, 74, 328–335. [Google Scholar] [CrossRef]
- Sun, L.; Kallolimath, S.; Palt, R.; Stiasny, K.; Mayrhofer, P.; Maresch, D.; Eidenberger, L.; Steinkellner, H. Increased in vitro neutralizing activity of SARS-CoV-2 IgA1 dimers compared to monomers and IgG. Proc. Nat. Acad. Sci. USA 2021, 118, e2107148118. [Google Scholar] [CrossRef]
- Sterlin, D.; Mathian, A.; Miyara, M.; Mohr, A.; Anna, F.; Claër, L.; Quentric, P.; Fadlallah, J.; Devilliers, H.; Ghillani, P.; et al. IgA dominates the early neutralizing antibody response to SARS-CoV-2. Sci. Transl. Med. 2021, 13, eabd2223. [Google Scholar] [CrossRef]
- Pace, R.; Williams, J.E.; Järvinen, K.M.; Belfort, M.B.; Pace, C.D.W.; Lackey, K.A.; Gogel, A.C.; Nguyen-Contant, P.; Kanagaiah, P.; Fitzgerald, T.; et al. Characterization of SARS-CoV-2 RNA, Antibodies, and Neutralizing Capacity in Milk Produced by Women with COVID-19. mBio 2021, 12, e03192-20. [Google Scholar] [CrossRef]
- Lorenzini, T.; Fliegauf, M.; Klammer, N.; Frede, N.; Proietti, M.; Bulashevska, A.; Camacho-Ordonez, N.; Varjosalo, M.; Kinnunen, M.; de Vries, E.; et al. Characterization of the clinical and immunologic phenotype and management of 157 individuals with 56 distinct heterozygous NFKB1 mutations. J. Allergy Clin. Immunol. 2020, 146, 901–911. [Google Scholar] [CrossRef]
- Souza, W.M.; Amorim, M.R.; Sesti-Costa, R.; Coimbra, L.D.; Brunetti, N.S.; Toledo-Teixeira, D.A.; de Souza, G.F.; Muraro, S.P.; Parise, P.L.; Barbosa, P.P.; et al. Neutralisation of SARS-CoV-2 lineage P.1 by antibodies elicited through natural SARS-CoV-2 infection or vaccination with an inactivated SARS-CoV-2 vaccine: An immunological study. Lancet Microbe 2021, 2, E527–E535. [Google Scholar]
- Khare, S.; Gurry, C.; Freitas, L.; Schultz, M.B.; Bach, G.; Diallo, A.; Akite, N.; Ho, J.; Lee, R.T.C.; Yeo, W.; et al. GISAID’s Role in Pandemic Response. China CDC Wkly. 2021, 3, 1049–1051. [Google Scholar] [CrossRef]
- Ong, D.S.Y.; Claas, E.C.J.; Breijer, S.; Vaessen, N. Comparison of the GeneFinder TM COVID-19 Plus RealAmp Kit on the sample-to-result Platform ELITeInGenius to the national reference method: An added value of N gene target detection? J. Clin. Virol. 2020, 132, 104632. [Google Scholar] [CrossRef] [PubMed]
- Corman, V.M.; Landt, O.; Kaiser, M.; Molenkamp, R.; Meijer, A.; Chu, D.K.; Bleicker, T.; Brünink, S.; Schneider, J.; Schmidt, M.L.; et al. Detection of 2019 novel coronavirus (2019-nCoV) by real-time RT-PCR. Euro Surveill. 2020, 25, 2000045. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Faria, N.R.; Mellan, T.A.; Whittaker, C.; Claro, I.M.; Candido, D.D.S.; Mishra, S.; Crispim, M.A.E.; Sales, F.C.; Hawryluk, C.; McCrone, J.T.; et al. Genomics and epidemiology of the P.1 SARS-CoV-2 lineage in Manaus, Brazil. Science 2021, 372, 815–821. [Google Scholar] [CrossRef] [PubMed]
- Candido, D.S.; Claro, I.M.; de Jesus, J.G.; Souza, W.M.; Moreira, F.R.R.; Dellicour, S.; Mellan, T.A.; du Plessis, L.; Pereira, R.H.M.; Sales, F.C.S.; et al. Evolution and epidemic spread of SARS-CoV-2 in Brazil. Science 2020, 369, 1255–1260. [Google Scholar] [CrossRef] [PubMed]
- Sahlin, K.; Mäkinen, V. Accurate spliced alignment of long RNA sequencing reads. Bioinformatics 2021, 37, 4643–4651. [Google Scholar] [CrossRef] [PubMed]
- Li, H.; Handsaker, B.; Wysoker, A.; Fennell, T.; Ruan, J.; Homer, N.; Marth, G.; Abecasis, G.; Durbin, R. The Sequence Alignment/Map format and SAMtools. Bioinformatics 2009, 25, 2078–2079. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- O’Toole, A.; Scher, E.; Underwood, A.; Jackson, B.; Hill, V.; McCrone, J.T.; Colquhoun, R.; Ruis, C.; Abu-Dahab, K.; Taylor, B.; et al. Assignment of epidemiological lineages in an emerging pandemic using the pangolin tool. Virus Evol. 2021, 7, veab064. [Google Scholar] [CrossRef]
- Hadfield, J.; Megill, C.; Bell, S.M.; Huddleston, J.; Potter, B.; Callender, C.; Sagulenko, P.; Bedford, T.; Neher, R.A. Nextstrain: Real-time tracking of pathogen evolution. Bioinformatics 2018, 34, 4121–4123. [Google Scholar] [CrossRef]
- Kadriyan, H.; Dirja, B.T.; Suryani, D.; Yudhanto, D. COVID-19 infection in the palatine tonsil tissue and detritus: The detection of the virus compartment withRT-PCR. BMJ Case Rep 2021, 14, e239108. [Google Scholar] [CrossRef]
- Lehmann, M.; Allers, K.; Heldt, C.; Meinhardt, J.; Schmidt, F.; Rodriguez-Sillke, Y. Human small intestinal infection by SARS-CoV-2 is characterized by a mucosal infiltration with activated CD8+T cells. Mucosal Immunol. 2021, 14, 1381–1392. [Google Scholar] [CrossRef]
- Lee, S.; Yoon, G.Y.; Myoung, J.; Kim, S.; Ahn, D. Robust and persistent SARS-CoV-2 infection in the human intestinal brush border expressing cells. Emerg. Microbes Infect. 2020, 9, 2169–2179. [Google Scholar] [CrossRef] [PubMed]
- Mishra, M.; Zahra, A.; Chauhan, L.V.; Thakkar, R.; Ng, J.; Joshi, S.; Spitzer, E.D.; Marcos, L.A.; Lipkin, W.I.; Mishra, N. A Short Series of Case Reports of COVID-19 in Immunocompromised Patients. Viruses 2022, 14, 934. [Google Scholar] [CrossRef]
- Yang, Y.; Iwasaki, A. Impact of Chronic HIV Infection on SARS-CoV-2 Infection, COVID-19 Disease and Vaccines. Curr. HIV/AIDS Rep. 2022, 19, 5–16. [Google Scholar] [CrossRef] [PubMed]
- Perl, S.H.; Uzan-Yulzari, A.; Klainer, H.; Asiskovich, L.; Youngster, M.; Rinott, E.; Youngster, I. SARS-CoV-2-Specific Antibodies in Breast Milk After COVID-19 Vaccination of Breastfeeding Women. JAMA 2021, 325, 2013–2014. [Google Scholar] [CrossRef]
- Atyeo, C.; Alter, G. The multifaceted roles of breast milk antibodies. Cell 2021, 184, 1486–1499. [Google Scholar] [CrossRef]
- Henkle, E.; Steinhoff, M.C.; Omer, S.B.; Roy, E.; Arifeen, S.E.; Raqib, R. The Effect of Exclusive Breast-feeding on Respiratory Illness in Young Infants in a Maternal Immunization Trial in Bangladesh. Pediatr. Infect. Dis. J. 2013, 32, 431–435. [Google Scholar] [CrossRef]
- Antunes, K.H.; Fachi, J.L.; de Paula, R.; da Silva, E.F.; Pral, L.P.; dos Santos, A.A.; Dias, G.B.M.; Vargas, J.E.; Puga, R.; Mayer, E.Q.; et al. Microbiota-derived acetate protects against respiratory syncytial virus infection through a GPR43-type 1 interferon response. Nat. Commun. 2019, 10, 3273. [Google Scholar] [CrossRef] [Green Version]
- Ramos-Casals, M.; Brito-Zerón, P.; Mariette, X. Systemic and organ-specific immune-related manifestations of COVID-19. Nat. Rev. Rheumatol. 2021, 17, 315–332. [Google Scholar] [CrossRef]
- Steensels, D.; Pierlet, N.; Penders, J.; Mesotten, D.; Heylen, L. Comparison of SARS-CoV-2 Antibody Response Following Vaccination with BNT162b2 and mRNA-1273. JAMA 2021, 326, 1533–1535. [Google Scholar] [CrossRef]
- Barin, B.; Kasap, U.; Selçuk, F.; Volkan, E.; Uluçkan, Ö. Comparison of SARS-CoV-2 anti-spike receptor binding domain IgG antibody responses after CoronaVac, BNT162b2, ChAdOx1 COVID-19 vaccines, and a single booster dose: A prospective, longitudinal population-based study. Lancet Microbe 2022, 4, e274–e283. [Google Scholar] [CrossRef]
- Zhong, D.; Xiao, S.; Debes, A.K.; Egbert, E.R.; Caturegli, P.; Colantuoni, E.; Milstone, A.M. Durability of Antibody Levels After Vaccination With mRNA SARS-CoV-2 Vaccine in Individuals with or Without Prior Infection. JAMA 2021, 326, 2524–2526. [Google Scholar] [CrossRef] [PubMed]
- Turner, J.S.; O’Halloran, J.A.; Kalaidina, E.; Kim, W.; Schmitz, A.J.; Zhou, J.Q. SARS-CoV-2 mRNA vaccines induce persistent human germinal centre responses. Nature 2021, 596, 109–113. [Google Scholar] [CrossRef] [PubMed]
- Trofin, F.; Nastase, E.V.; Iancu, L.S.; Constantinescu, D.; Cianga, C.M.; Lunca, C. Anti-RBD IgA and IgG Response and Transmission in Breast Milk of Anti-SARS-CoV-2 Vaccinated Mothers. Pathogens 2022, 11, 286. [Google Scholar] [CrossRef] [PubMed]
- Walls, A.C.; Sprouse, K.R.; Bowen, J.E.; Joshi, A.; Franko, N.; Navarro, M.J.; Stewart, C.; Cameroni, E.; McCallum, M.; Goecker, E.A.; et al. SARS-CoV-2 breakthrough infections elicit potent, broad, and durable neutralizing antibody responses. Cell 2022, 185, 872–880. [Google Scholar] [CrossRef] [PubMed]
- Yu, J.; Collier, A.Y.; Rowe, M.; Mardas, F.; Ventura, J.D.; Wan, H. Neutralization of the SARS-CoV-2 Omicron BA.1 and BA.2 Variants. N. Engl. J. Med. 2022, 386, 1579–1580. [Google Scholar] [CrossRef]
- Choi, B.; Choudhary, M.C.; Regan, J.; Sparks, J.A.; Padera, R.F.; Qiu, X.; Solomon, I.H.; Kuo, H.-H.; Boucau, J.; Bowman, K.; et al. Persistence and Evolution of SARS-CoV-2 in an Immunocompromised Host. N. Engl. J. Med. 2020, 383, 2291–2293. [Google Scholar] [CrossRef]
- Corey, L.; Beyrer, C.; Cohen, M.S.; Michael, N.L.; Bedford, T.; Rolland, M. SARS-CoV-2 Variants in Patients with Immunosuppression. N. Engl. J. Med. 2021, 385, 562–566. [Google Scholar] [CrossRef]
- Kemp, S.A.; Collier, D.A.; Datir, R.P.; Ferreira, I.A.T.M.; Gayed, S.; Jahun, A.; Hosmillo, M.; Rees-Spear, C.; Mlcochova, P.; Lumb, I.U.; et al. SARS-CoV-2 evolution during treatment of chronic infection. Nature 2021, 592, 277–282. [Google Scholar] [CrossRef]
| Parameters | 19 May 2021 | 27 May 2021 | 7 June 2021 | 22 July 2021 | Reference Values |
|---|---|---|---|---|---|
| WBC (×103/μL) | 6.78 | 13.09 | 10.96 | 9.57 | 4–10 |
| RBC (×106/μL) | 4.14 | 4.09 | 4.55 | 5.40 | 4.2–5.4 |
| Hb (g/dl) | 12.10 | 12.40 | 13.80 | 15.30 | 12–16 |
| Ht (%) | 37.10 | 37.90 | 42.90 | 46.20 | 37–47 |
| MCH (fl) | 89.60 | 92.70 | 94.30 | 85.50 | 80–99 |
| MCHC (pg) | 29.20 | 30.30 | 30.30 | 28.30 | 27–32 |
| CHCM (g/dl) | 32.60 | 32.70 | 32.20 | 34.20 | 32–37 |
| RDW (%) | 13.50 | 15.40 | 16.80 | 15.20 | 10–15 |
| PLT (×103/μL) | 260 | 292 | 206 | 299 | 150–400 |
| MPV (fl) | 12.20 | 12.40 | 12.20 | 10.80 | 6–10 |
| Neutrophil/mm3 | 4170 | 7880 | 9070 | 5010 | 2000–8000 |
| Lymphocyte/mm3 | 1320 | 2400 | 1320 | 3730 | 1000–4000 |
| Monocyte/mm3 | 1170 | 2400 | 550 | 680 | 200–800 |
| Eosinophil/mm3 | 20 | 130 | 0 | 80 | 0–450 |
| Basophil/mm3 | 20 | 0 | 0 | 7 | 0–200 |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2022 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Sabino, J.S.; Amorim, M.R.; de Souza, W.M.; Marega, L.F.; Mofatto, L.S.; Toledo-Teixeira, D.A.; Forato, J.; Stabeli, R.G.; Costa, M.L.; Spilki, F.R.; et al. Clearance of Persistent SARS-CoV-2 RNA Detection in a NFκB-Deficient Patient in Association with the Ingestion of Human Breast Milk: A Case Report. Viruses 2022, 14, 1042. https://doi.org/10.3390/v14051042
Sabino JS, Amorim MR, de Souza WM, Marega LF, Mofatto LS, Toledo-Teixeira DA, Forato J, Stabeli RG, Costa ML, Spilki FR, et al. Clearance of Persistent SARS-CoV-2 RNA Detection in a NFκB-Deficient Patient in Association with the Ingestion of Human Breast Milk: A Case Report. Viruses. 2022; 14(5):1042. https://doi.org/10.3390/v14051042
Chicago/Turabian StyleSabino, Janine S., Mariene R. Amorim, William M. de Souza, Lia F. Marega, Luciana S. Mofatto, Daniel A. Toledo-Teixeira, Julia Forato, Rodrigo G. Stabeli, Maria Laura Costa, Fernando R. Spilki, and et al. 2022. "Clearance of Persistent SARS-CoV-2 RNA Detection in a NFκB-Deficient Patient in Association with the Ingestion of Human Breast Milk: A Case Report" Viruses 14, no. 5: 1042. https://doi.org/10.3390/v14051042
APA StyleSabino, J. S., Amorim, M. R., de Souza, W. M., Marega, L. F., Mofatto, L. S., Toledo-Teixeira, D. A., Forato, J., Stabeli, R. G., Costa, M. L., Spilki, F. R., Sabino, E. C., Faria, N. R., Benites, B. D., Addas-Carvalho, M., Stucchi, R. S. B., Vasconcelos, D. M., Weaver, S. C., Granja, F., Proenca-Modena, J. L., & Vilela, M. M. d. S. (2022). Clearance of Persistent SARS-CoV-2 RNA Detection in a NFκB-Deficient Patient in Association with the Ingestion of Human Breast Milk: A Case Report. Viruses, 14(5), 1042. https://doi.org/10.3390/v14051042









