1. Introduction
Bluetongue virus (BTV), as part of the virus family
Reoviridae and the genus
Orbivirus, is characterised by a double-stranded RNA genome divided into ten segments of different sizes [
1]. Among the seven structural (VP1–7) and six non-structural proteins (NS1–NS5, NS3a), VP2 plays the most important role for serotype specificity by carrying the neutralising epitopes [
2,
3,
4]. Serotype affiliation to 1 of the 24 notifiable classical BTV serotypes needs to be examined by the reference method—the virus neutralisation test [
5]. However, traditional serotyping is impeded in the case of atypical BTV, and molecular typing of segment 2/VP2 leads to the classification of putative novel serotypes. Up to now, 36 (putative) serotypes—24 serotyped notifiable serotypes, 3 more or less ‘serotyped’ atypical serotypes and 9 atypical putative novel serotypes—have been described [
6,
7]. The arthropod-borne virus (arbovirus) is transmitted via infected female
Culicoides and causes bluetongue disease in wild and domestic ruminants [
8,
9]. The clinical outcome of a BTV infection is highly variable depending not only on the BTV strain but also on the host’s breed, genetics, age and immune status [
10]. Bluetongue disease, known as a major disease of sheep, particularly affects European wool sheep breeds, with high mortality and morbidity rates [
11,
12]. Bluetongue disease is a systemic haemorrhagic viral fever, and clinical symptoms derive from direct viral endothelial cell damage and host cell response. Oedema in the head region is common, and pulmonary oedema can be fatal for the affected animal [
13,
14,
15]. In contrast to sheep, cattle and goats are mostly known to develop subclinical disease [
16]; nevertheless, the morbidity in cattle during the first BTV-8 incursion was estimated to be around 10% [
14].
In northern European countries, two important BTV incursions have been reported up to now, both caused by BTV-8 strains. With the help of animal movement restrictions and large-scale vaccination campaigns, it could be possible to successfully eradicate the disease from northern European [
17,
18,
19]. Thus, Germany was declared officially BTV-free from February 2012 until December 2018 [
20,
21]. Then, the first BTV-8 case to re-emerge in Germany was detected. Over the following years, only a few additional BTV-8 outbreaks were noted in healthy cattle and calves. The BTV-8 strain that re-emerged caused mostly mild or no clinical symptoms [
22]. This is in line with the findings of an experimental study in sheep, which showed that this BTV-8 strain was less pathogenic than the BTV-8 strain circulating during the first BTV-8 epidemic of 2006–2009 [
23]. The accidental release of frozen material contaminated by bull semen infected with the first BTV-8 epidemic strain was suggested to have caused this re-emergence [
24].
Atypical BTV, also known as small ruminant or goat-associated BTV, differs from the classical BTV serotypes in terms of both molecular and biological features [
15]. None of the atypical strains have been reported in cattle [
25]. For the classical BTV strains, a prolonged viraemia of <60 days has been reported, whereas for the Swiss BTV-25-TOV (Toggenburg virus) and German BTV-25-GER2018 strain, a viraemia of several years has been described [
26,
27]. Moreover, infection with classical BTV strains led to a long-lasting and protective humoral response, with antibodies formed against VP2 and VP7. In the virus neutralisation test (VNT), neutralisation of the classical virus strain with the respective reactive serum can be observed. BTV-25-GER2018 and the three Mongolian strains showed only partial neutralisation in the VNT in contrast to the classical strains. Several atypical BTV strains do not grow on vector-derived
Culicoides cell lines, and animal experiments have suggested vector-independent direct contact transmission for BTV-26, -27 and -28 [
28,
29]. Field infections with atypical BTV strains mostly remained without clinical disease. Mild clinical disease was observed after the experimental infection of sheep with BTV-25 and BTV-26 [
30], and moderate clinical signs were observed in sheep experimentally infected with BTV-28 [
29].
We performed an animal experiment in goats, as they represent the natural host of atypical BTVs. With the high number of in-contact goats, we could further analyse the contact transmission as one route of horizontal transmission. Two novel atypical BTV strains (BTV-25-GER2018 and the putative novel BTV-33-MNG3/2016), both isolated from clinically healthy goats, were chosen for the animal trial. BTV-25-GER2018 originated from southern Germany [
27], and the BTV-33-MNG3/2016 strain originated from Mongolia [
7]. The classical BTV strain—the BTV-8-GER2018 strain that re-emerged—was isolated in Germany in 2018 from cattle [
21] and was, for the first time, tested in an animal experiment with goats.
2. Materials and Methods
2.1. Animals
Thirty 4- to 6-month-old Thuringian goats (twenty-six female and four castrated male goats) were kept at the facilities of the Friedrich-Loeffler-Institut, Insel Riems, Germany, under biosafety level 3 conditions. The competent authority (State Office for Agriculture, Food Safety and Fisheries of Mecklenburg-Vorpommern, Rostock, Germany; Ref. No. LALLF 7221.3-1-048/19; date of approval 7 November 2019) approved the animal experiment. For the study, goats from a restriction-free area with no history of BTV vaccination were chosen. Before housing at the FLI, all goats tested negative for the presence of BTV antibodies in cELISA and for BTV genome by RT-qPCR. All animals were in good health.
2.2. Virus Preparations
Two atypical BTV strains (BTV-25-GER2018 and BTV-33-MNG3/2016), both isolated from clinically healthy goats, as well as BTV-8-GER2018, were chosen for the animal trial. The full-length sequences of the coding regions for all three virus isolates are available (BTV-25: LR798441-50; BTV-33: LR877358-67; BTV-8: OM523087-96). The BTV-33-MNG3/2016 strain was isolated on BHK-21 (CT) cells (FLI cell culture collection number RIE0164) and then passaged five times on BHK-21 (BSR/5) cells (FLI cell culture collection number RIE0194). BHK-21 (BSR/5) cells are a clone of BHK-21 cells. The BTV-25-GER2018 strain was isolated on BHK-21 (BSR/5) cells and passaged a total of six times on BHK-21 (BSR/5) cells, whereas the BTV-8-GER2018 strain was isolated on BHK-21 (BSR/5) cells and then passaged again twice on BHK-21 (BSR/5) cells. All the virus stocks contained the infected cell–supernatant mixture of the last previously described cell passage and were stored at −80 °C until usage for the animal trial.
2.3. Experimental Design and Sample Collection
Three groups (A, B, C), each with ten randomly assigned animals, were kept in separate rooms without contact. In each group, five of the ten goats were inoculated subcutaneously in the shoulder neck region with 4 mL of the respective virus preparation at two different injection sites. The other five goats were kept in direct contact within the same housing unit as transmission controls. In group A, five goats were infected with BTV-33-MNG3/2016 virus, in group B, five goats were infected with BTV-25-GER2018 virus and in group C, five goats were infected with BTV-8-GER2018 virus. Back titration on BSR cells revealed the following titres:
| For group A, BTV-33-MNG3/2016, 104.83 CCID50/mL (per goat 2.7*105 CCID50); |
| For group B, BTV-25-GER2018, 103.3 CCID50/mL (per goat 8.0 * 103 CCID50); |
| For group C, BTV-8-GER2018, 103.67 CCID50/mL (per goat 1.9*104 CCID50). |
Body temperature was taken daily, and the goats were clinically scored throughout the experiment, applying a modification of the Clinical Score system [
31] described in an earlier study [
7]. On the sampling days (0, 3, 5, 7, 10, 12, 14, 17, 21, 24, 28, 31, 35 and 42 dpi), nasal, oral, ocular and rectal swabs were taken as well as blood from the jugular vein by using an adapter system (Kabe Labortechnik GmbH, Nümbrecht, Germany). We used plane tubes containing ethylenediaminetetraacetic acid (EDTA) for EDTA blood collection and serum tubes without anticoagulants for serum collection (Kabe Labortechnik GmbH, Nümbrecht, Germany). Serum was collected after centrifugation at 2000 rpm for 15 min. After euthanising, lung, liver, spleen, mediastinal and mesenterial lymph nodes were taken from the goats positive for the BTV genome in the EDTA blood by RT-qPCR. (Once a goat was noted as positive in the study, it remained classified as positive until the end of the study.)
2.4. RNA Extraction and RT-qPCR
Lentil-sized organ samples (approx. 30 mg) were homogenised in 500 µL of a serum-free medium using the TissueLyser II tissue homogeniser (QIAGEN, Hilden, Germany). For RNA extraction, 100 µL of the liquid starting material was used (cell culture material, EDTA blood, serum, swab samples collected in 2 mL serum-free media and the liquid part of the homogenised organ material) and eluted in 100 µL of elution buffer. Extraction was performed with the NucleoMagVET kit (Macherey-Nagel, Düren, Germany) using the half-automated KingFisher platform (King-Fisher Flex magnetic particle processor, Thermo Fisher Scientific, Darmstadt, Germany). As a control for successful RNA extraction, 10 µL of internal control RNA (IC-2 RNA) was added during the extraction process [
32]. The RNA was amplified with the Pan-BTV-S10-RT-qPCR recommended by the OIE [
33], adapted with an additional probe and named BTV-S10-primer-probe-mix-v2. Comparative evaluation data for the BTV-S10-OIE assay and the modified BTV-S10-primer-probe-mix-v2 are summarised in the
supplemental material (Table S1). For the generation of 200 µL of BTV-S10-primer-probe-mix-v2, the following oligos were mixed: 20.0 µL of BTV_IVI_F (5’-TGG AYA AAG CRA TGT CAA A-3´), 20.0 µL of BTV_IVI_R (5´-ACR TCA TCA CGA AAC GCT TC-3´), 3.75 µL of BTV_IVI_FAM (5´-FAM-ARG CTG CAT TCG CAT CGT ACG C-BHQ1-3´), 2.50 µL of BTV_IVI_FAM_v2 (5´-FAM-AGG CTG CAT ACG CAT CRT ACG C-BHQ1-3´) and 153.75µL of 0.1x TE (pH 8.0). The final composition of the RT-qPCR reactions was 1.25 μL of RNase-free water, 6.25 μL of 2x RT-PCR buffer, 0.5 μL of RT-PCR Enzyme Mix, 1 μL of BTV-S10-primer-probe-mix-v2-FAM, 1 μL of EGFP-mix1-HEX and 2.5 µL of the heat-denatured template RNA (at 95 °C for 5 min). All RT-qPCRs were run on the CFX 96 real-time PCR cycler (Bio-Rad, Hercules, CA, USA) with the AgPath-ID™ One-Step RT-PCR Reagents from Applied Biosystems™ (Waltham, MA, USA). The temperature profile used was 10 min at 45 °C (reverse transcription) and 10 min at 95 °C (inactivation of the reverse transcriptase/activation Taq polymerase) followed by 42 cycles of 15 s at 95 °C (denaturation), 20 s at 56 °C (annealing) and 30 s at 72 °C (elongation). Fluorescence values (FAM, HEX) were collected during the annealing step. Samples were considered positive when the quantification cycle (Cq) values were <40.
In every run, a generated BTV standard series produced by droplet PCR (QX200 Droplet Digital PCR System, Bio Rad, Hercules, CA, USA) was included for calculating the genome copy numbers.
2.5. Serological Analysis
2.5.1. ELISA
Serum samples of 0, 7, 14, 21, 28 and 42 dpi were screened for BTV-group-specific antibodies (VP7) using a competitiveELISA (ID Screen® Bluetongue Competition, ID-Vet, France) and plate reader reading at a wavelength of 450 nm according to the manufacturer’s instructions. The results were expressed as the percent of negativity compared to the negative kit control (% S/N = optical density (OD) of the sample/OD of the negative control multiplied by 100) and denoted as a positive or negative result (<50% S/N were considered as positive, samples with ≥50% S/N as negative).
2.5.2. VNT
Serum samples of 0, 7, 14, 21, 28 and 42 dpi of goats positive for BTV RNA in the EDTA blood were screened by VNT for the presence of neutralising antibodies.
Briefly, the serum was diluted in log2 steps starting from 1:10 to 1:1280 and titrated against 100 CCID50 of the respective virus (BTV-25-GER2018, BTV-33-MNG3/2016 or BTV-8-GER2018). The plates were incubated for 1 h at 37 °C before overnight incubation at 4 °C. The following day, 100 µL of a BHK-21 (BSR/5) cell suspension of approximately 30,000 cells/100 µL was added per well. After incubation for 3–5 days at 37 °C, the plates were screened with a stereomicroscope for cytopathic effect (CpE). The neutralisation titre was determined as the highest dilution of serum with 100% neutralisation (no CpE). The Spearman and Kärber methods were used for the calculations.
2.6. Isolation in Cell Culture
All the blood samples from the experimentally infected goats were processed identically for the virus-isolation experiments: 500 µL of EDTA blood was centrifuged (8000 rpm) for 2 min, and the red blood cells were washed twice in 1 mL of PBS and finally diluted in 500 µL of PBS. In cases of no sufficient cell pellet being generated after initial centrifugation, the unwashed blood was used for virus isolation experiments. In both cases, blood preparations were lysed using 20 s ultrasound treatment at 30 W (Sonifier 450, Branson Ultrasonics, Danbury, CT, USA). BHK-21 (BSR/5) cells in T25 cm² cell flasks were initially incubated for three hours at 37 °C using the cultivation medium with 5.32 g of Hanks salts, 4.76 g of Earle’s salts, 1.25 g of NaHCO3, 10 mL of non-essential amino acids and 120 mg of sodium pyruvate per litre of medium (FLI intern medium number ZB5d) supplemented with 10% foetal calf serum (FCS). Afterwards, the cells were inoculated with 500 µL of the blood preparation for two hours and incubated on a tilt shaker in an incubator at 37 °C and 5% CO2. Then, the blood inoculum was removed, and the flasks were refilled with the medium supplemented with 10% FCS and antibiotics at double the standard concentration (20,000 µg/mL Penicillin, 20,000 units/mL Streptomycin, 10 mg/mL Gentamicin, 250 µg/mL Amphotericin B). After 3 to 4 days of incubation at 37 °C, the infected BSR cell monolayer was split by using 1 mL of trypsin and mixed with 5 mL of the supernatant. In the next step, 3 mL of the cell–trypsin–supernatant suspension was transferred to a new T75 cm² cell flask with fresh BSR cells grown for 3 h. Three passages were performed, and the success of virus replication was confirmed by the genomic load estimated by RT-qPCR.
2.7. Blood Analysis
For the blood analysis, the EDTA blood was analysed within the same day of sampling using the ProCyte Dx (Idexx, Westbrook, ME, USA) according to the manufacturer’s instructions. Several blood parameters were measured including the total white blood cell count, lymphocytes, neutrophils, monocytes, basophils, thrombocytes, erythrocytes and haematocrit.
2.8. Statistical Analysis
For statistical analysis, we used the Mann–Whitney U test with Bonferroni correction for the individual bleeding time points in GraphPad Prism 9.0.0. We compared the viraemic animals of groups A and B infected with the atypical BTV strains (n = 6) and the viraemic animals of group C infected with the classical BTV strain (n = 2).
Viraemia was analysed by comparing the dpi of its onset, the maximum viral load in log10 genome copies/mL and the total virus production with the area under the curve (AUC). Furthermore, we analysed the total WBC count, the lymphocyte count, the monocyte count, basophils, thrombocytes and erythrocytes in K/µL (1000 cells per µL) and haematocrit in %. Non-parametric tests were preferred because of the small group sizes and the potential non-normality of the data.
4. Discussion
Atypical BTV has been on the rise in the last few years since the first discovery of BTV-25-TOV in 2008. Investigations on the pathogenesis of different BTV strains were conducted over the years, for classical as well as atypical BTV strains. In our study, we investigated the comparative pathogenesis and immune response of classical and atypical BTV strains using three different goat groups. To represent the classical BTV, we chose the re-emerged BTV-8 strain isolated from cattle in Germany, whose kinetics and pathogenesis in goats have never been tested before. To represent atypical BTV strains, we chose BTV-25 isolated from goats in Germany and BTV-33 isolated from goats in Mongolia. All three strains were isolated from asymptomatic ruminants naturally infected in the field. The pathogenesis and immunological study revealed clear differences concerning the measured blood parameters such as total WBC, lymphocyte and monocyte count. Furthermore, only the animals infected (two of the five inoculated goats) with the classical BTV strains showed a humoral response. However, the atypical and classical strains did not diverge regarding the dpi at which viraemia began, the maximum viral load in log10 genome copies/mL and the total virus production.
The clinical picture of BTV depends not only on the virus strain but also on the affected host species [
10]. Severe disease is reported for sheep, whereas for other domestic ruminants, such as goats and cattle, subclinical manifestation is stated [
15]. The atypical virus strains chosen for the animal trial were isolated from asymptomatic goats. This is in line with the findings of the animal trial performed here, where all goats infected with BTV-25-GER2018 and BTV-33-MNG3/2016 showed an asymptomatic BTV infection. For BTV-8, however, clinical disease was seen in cattle and goats during the first outbreak from 2006 to 2009 and with less severity in cattle in the second BTV-8 epidemic in 2015–2019 [
11,
34]. During the first epidemic, goats showed a high seroprevalence rate in northwestern Europe (25% in Germany) [
19], and mild clinical disease could be seen in one experimental BTV-8-infected goat with fever and generalised illness including apathy, dysphagia, diarrhoea and lameness [
35]. Furthermore, BTV-8 was able to cross the caprine placenta, as reported for cattle [
36]. To our knowledge, we have tested the newly emerged BTV-8 strain (BTV-8-GER2018) for the first time in goat as the host species. The two infected goats in our study showed neither clinical signs nor fever, which is line with reports about subclinical disease in goats and a reduced pathogenicity of the newly emerged BTV-8 strain [
23].
In our animal experiment, we observed a difference between the inoculated and infected goats. For BTV-33-MNG3/2016, five out of five inoculated goats became infected, whereas for BTV-8-GER2018, two became infected, and for BTV-25-GER2018, only one goat became infected out of five. An experimental study with BTV-8 in cattle did not find a correlation between the amount of inoculum and the kinetics of viraemia [
37]. In this study, the length and the intensity of viraemia and the neutralising antibody response did not differ between the different infectious doses, varying from just 10 TCID
50 to 10
6 TCID
50 [
37]. Through the natural inoculation route by BTV-positive
Culicoides midges, only one bite can transmit around 0.32 to 7.79 TCID
50 and establish infection [
38,
39]. The probability of virus transmission from an infectious vector to a susceptible ruminant host was described as close to 100% [
40]. The inoculation titres used in our study are all in the range of other animal trials, and lower infection titres more likely resemble the natural vector-borne transmission cycle. However, we did not use the same infectious dose for all groups, and it cannot be completely ruled out that it had an influence on the study outcome. The reduced infectivity of BTV-8-GER2018 after subcutaneous infection described in our study (two of five goats became BTV-positive) might be in line with the findings of Flannery et al. 2019, who described the re-emerged BTV-8 strain as having a reduced vector competence as well as the relatively slow spread of the re-emerged BTV strain [
23]. For BTV-25-GER2018, a moderate seroprevalence of 18% to 23% was described in a naturally infected goat flock over a prolonged period of time (4.5 years), whereas 31% to 38% were BTV RNA-positive [
27]. These results of only moderate seroprevalence over a longer time period, together with the findings of our study that only one of five goats became BTV-positive after inoculation, suggest a low infectivity of BTV-25-GER2018. For BTV-33-MNG3/2016, no data are available about the seroprevalence or infectivity of this atypical BTV strain in the field. Hence, our study indicates a potentially high infectivity rate of BTV-33-MNG3/2016 in goats.
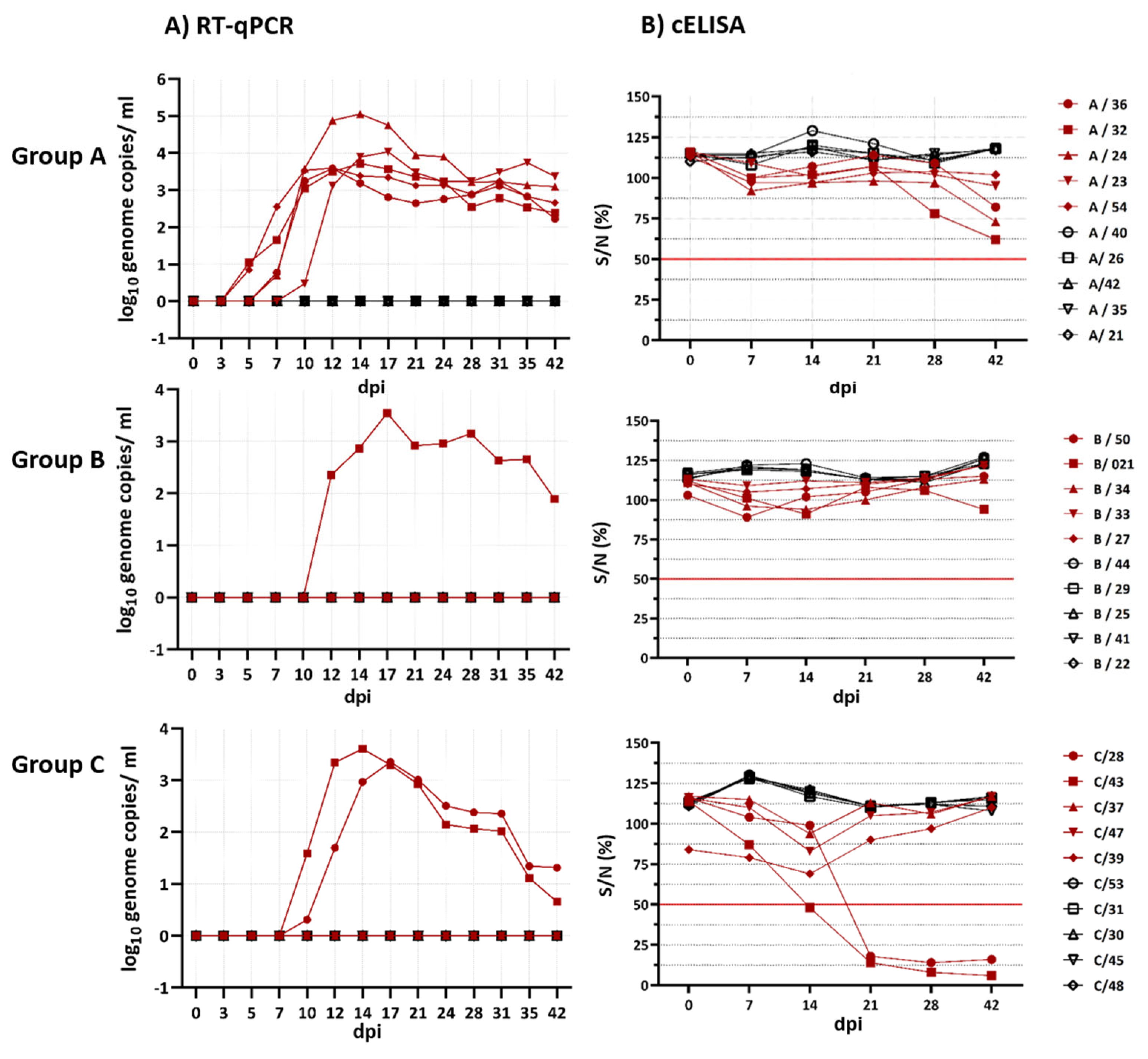
Interestingly, BTV-8-GER2018 and BTV-25-GER2018 showed moderate Cq values with peaks between 27.3 and 28.2 (3.4–3.6 log
10 genome copies/mL) in the EDTA blood. These relatively high Cq value peaks are not surprising for atypical BTV strains but are surprising for BTV-8. However, our results of a reduced viraemia of the German re-emerged BTV-8 strain are in line with the observed lower genome concentrations seen in BTV-8-FRA2017-infected sheep in comparison to the UKG2007 strain. Remarkably, BTV-33-MNG3/2016 showed the lowest peak Cq values (two of the five goats, 22.5 and 25.9). Nevertheless, BTV-33-MNG3/2016 was not the only atypical BTV strain that showed lower Cq values during infection, comparable to classical BTV strains; this has also been described for BTV-26, BTV-27 and BTV-28 [
25,
28,
29]. The high ΔCq value differences among the respective serum samples throughout the study period indicate that BTV is connected to the red blood cells, and only few virus particles circulate freely. For the classical BTV strains, this phenomenon is well known [
41], and our study confirms it for the two atypical BTV strains tested here as well. The time point of infection varied little in the groups, starting with the earliest from 5 to 10 dpi with BTV-33-MNG3/2016 followed by BTV-8 at 10 dpi and BTV-25 at 12 dpi. For the BTV-8 strains FRA2017 and UKG2007, earlier infection time points starting at 2 dpi have been described in sheep [
23]. Hence, our findings for BTV-8-GER2018 presenting a late onset of viraemia are surprising and prove that BTV can vary strongly within the same serotype. However, no significant difference regarding the onset of viraemia between the atypical and classical strains was found in our study. For all three groups, no contact transmission was observed, and, therefore, this does not seem to play a role for BTV-25-GER2018, BTV-33-MNG3/2016 and BTV-8-GER2018. We were able to reisolate all three virus strains from the RT-qPCR animals. For successful virus isolation, the Cq value seems to play a crucial role. This could explain the shorter time periods of virus isolation for BTV-8 (7 days; SD 1.4) and BTV-25 (1 day), with higher Cq values, compared to BTV-33-MNG3/2016 (13.6 days; SD 5.2), with lower Cq values. The virus isolation length of BTV-8-GER2018 is consistent with the findings of related BTV-8 strains [
23].
For all three virus strains, the spleens at 42 dpi showed the lowest Cq values. Lung, liver and mediastinal lymph nodes were positive for BTV RNA for the two atypical BTV strains but not for BTV-8. One hypothesis might be the ascending humoral immune response of BTV-8, starting to clear BTV infection in the case of BTV-8. In detail, the two goats infected with BTV-8, which were positive in the RT-qPCR, developed BTV group- and serotype-specific antibodies starting at 21 dpi, whereas the goats infected with BTV-25 and BTV-33-MNG3/2016 did not develop group- and serotype-specific antibodies, despite positivity in the RT-qPCR. However, at 42 dpi, there was a very slight tendency of goats positive for BTV-25 and BTV-33-MNG3/2016 in the RT-qPCR to approach the cut-off value in the cELISA. A longer study period than 42 dpi might have shown whether these goats infected with atypical BTV would have become cELISA-positive. However, the humoral immune response of goats infected with the atypical BTV strains was not comparable to the fast and strong reaction of goats infected with BTV-8 in our study. We can conclude that there is a difference in the immunopathogenesis of the classical BTV strains and the atypical BTV strains BTV-25-GER2018 and BTV-33-MNG3/2016 tested here.
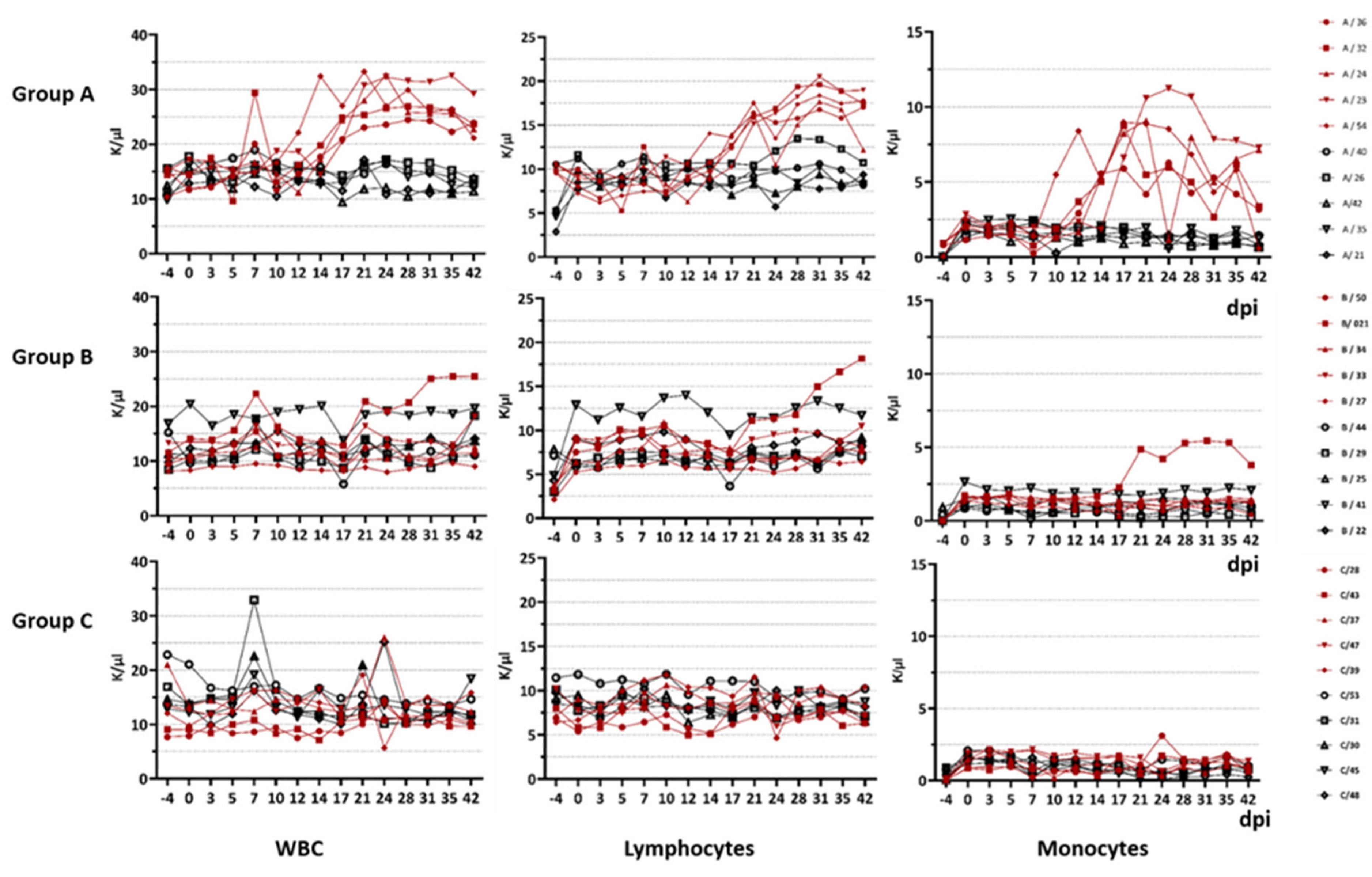
The difference in immunopathogenesis is underlined by the blood parameters measured during this study. For goats infected with BTV-33-MNG3/2016 and BTV-25-GER2018, we stated a significant difference in the WBC count compared to the goats infected with the classical strain. In general, the WBC count of goats is described as 9000 cells/µL with a range of 4000 to 14,000. Leukocytosis starts at a WBC count of more than 13,000/µL, and leukopenia starts at less than 4000 /µL. However, total WBC counts and differential cell counts vary significantly with age: for 3-month-old goats, increased WBC counts of 18.18 ± 3.84 × 10
3 /µL were reported, whereas for adult goats at an age of 2 years, the WBC count decreased to 8.08 ± 2.51 × 10
3/µL [
42]. Hence, our 6-month-old goats in the animal trial showed higher WBC counts than adults, but the values are comparable to those of the 3-month-old goats described in the literature. Nevertheless, the goats infected with atypical BTV strains developed a significant increase in WBCs over the course of infection, which can be constituted as leukocytosis from approx. 17 dpi until the end of the animal trial at 42 dpi. In detail, we observed lymphocytosis and monocytosis in all of the viraemic animals infected with the atypical strains. BTV infection was previously described to be followed by pan-leukopenia, with its peak at 7–8 dpi [
5]. Transient leukopenia and lymphopenia were seen in previous studies with BTV-1 and BTV-8 in sheep, where leukopenia occurred during the very first days of infection, resulting from a depletion of T-lymphocytes. However, in the same study, a proliferation in the B cell population was seen, possibly linked to the upcoming antibody production [
43]. However, in our study, we found leukopenia neither during the first days after inoculation nor over the whole time period. It remains unknown which lymphocyte fraction increased in our animal study, and it is difficult to link the increase to the B cell fraction, as the antibody levels did not rise in groups A and B. An increase in monocytes has not been commonly described in other BTV infection studies [
43]. In future studies, more details about the alterations in the lymphocyte populations, as well as various cytokines, should be collected to understand the differences in immunopathogenesis between classical and atypical BTV serotypes.
In conclusion, our animal study showed differences in the immunological reaction between the two atypical BTV strains BTV-25-GER2018 and BTV-33-MNG3/2016 and the classical BTV strain BTV-8-GER2018. We found differences in the humoral response and also in the total WBC count, which indicate a possible difference in the cellular immune response. However, regarding the infectivity rate and peak Cq values, the atypical and classical BTV strains did not differ. Our experiment underlines that BTV has a highly variable disease outcome depending on the strains, the host and the individual differences between animals, which makes the generalisation of our results (atypical versus classical BTV strains) difficult. Nevertheless, our animal study strongly suggests that the phylogenetically distinct atypical BTV strains differ from the classical BTV strains in terms of immunopathogenesis. Furthermore, no horizontal transmission was seen for BTV-25, BTV-33 and BTV-8.