Monitoring of Enterovirus D68 Outbreak in Israel by a Parallel Clinical and Wastewater Based Surveillance
Abstract
:1. Introduction
2. Materials and Methods
2.1. Clinical Sample Collection and Processing
2.2. Wastewater Sample Preparation
2.3. Detection of EV-D68 RNA by Quantitative PCR (RT-qPCR)
2.4. Generation of EVD68 RNA Standard and Calculation of EVD68 RNA Copies
2.5. EV-D68 VP1 Sequencing and Phylogenetic Analysis
3. Results
3.1. Calibration of the Quantitative Duplex EVD68-MS2 Assay
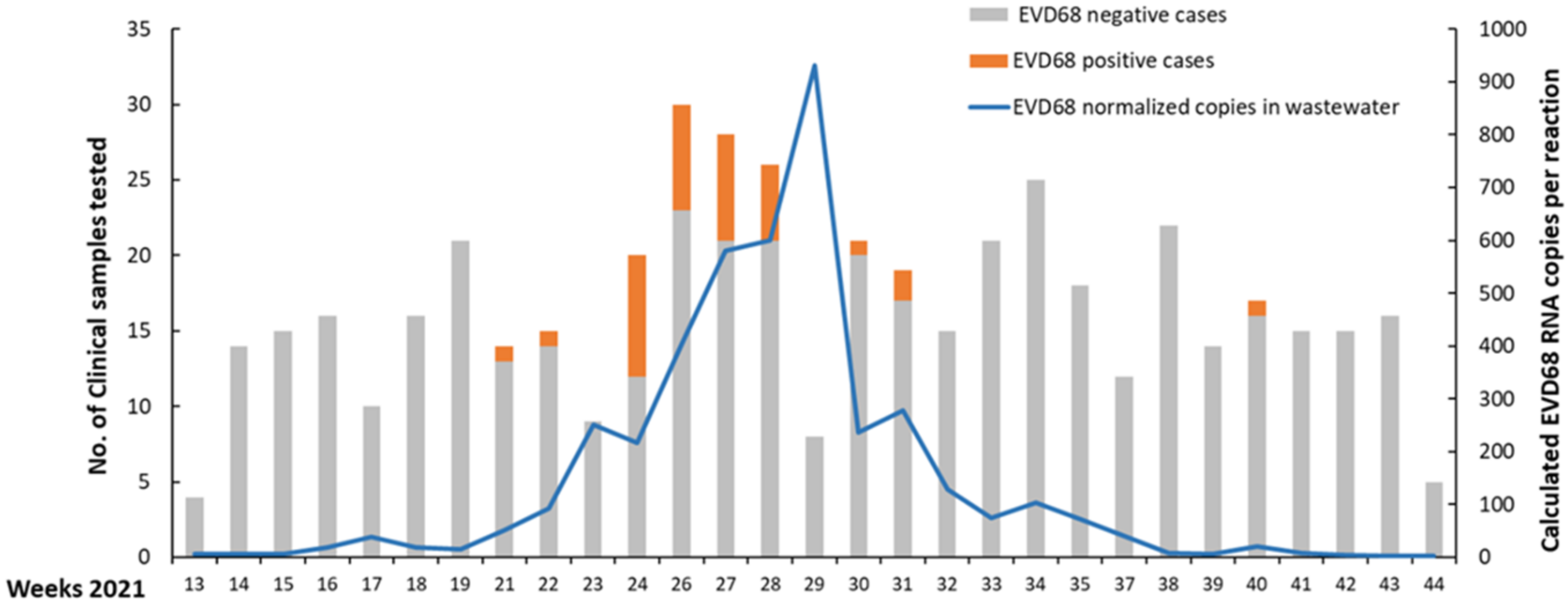
3.2. Characterization and EVD68 Infection and Monitoring of EVD68 RNA in Wastewater Samples
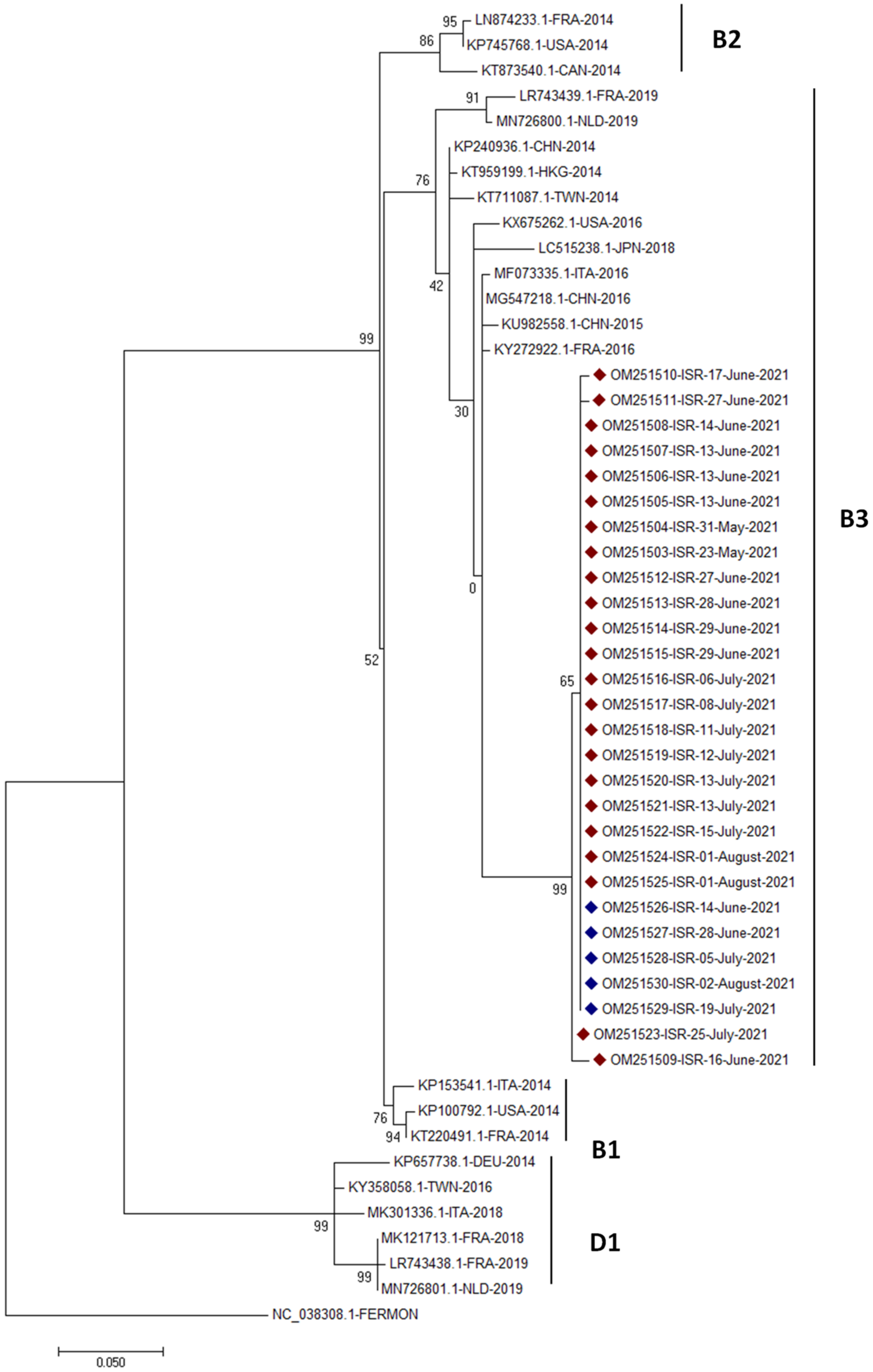
3.3. Phylogenetic Analysis
4. Discussion
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Oberste, M.S.; Maher, K.; Schnurr, D.; Flemister, M.R.; Lovchik, J.C.; Peters, H.; Sessions, W.; Kirk, C.; Chatterjee, N.; Fuller, S.; et al. Enterovirus 68 is associated with respiratory illness and shares biological features with both the enteroviruses and the rhinoviruses. J. Gen. Virol. 2004, 85 Pt 9, 2577–2584. [Google Scholar] [CrossRef] [PubMed]
- Knoester, M.; Helfferich, J.; Poelman, R.; Van Leer-Buter, C.; Brouwer, O.F.; Niesters, H.G.M. Twenty-nine Cases of Enterovi-rus-D68-associated Acute Flaccid Myelitis in Europe 2016: A Case Series and Epidemiologic Overview. Pediatric Infect. Dis. J. 2019, 38, 16–21. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Harvala, H.; Jasir, A.; Penttinen, P.; Celentano, L.P.; Greco, D.; Broberg, E. Surveillance and laboratory detection for non-polio enteroviruses in the European Union/European Economic Area, 2016. Eurosurveillance 2017, 22, 16-00807. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Shah, M.M.; Perez, A.; Lively, J.Y.; Avadhanula, V.; Boom, J.A.; Chappell, J.; Englund, J.A.; Fregoe, W.; Halasa, N.B.; Harrison, C.J.; et al. Enterovirus D68-Associated Acute Respiratory Illness horizontal line New Vaccine Surveillance Network, United States, July–November 2018–2020. MMWR Morb. Mortal Wkly. Rep. 2021, 70, 1623–1628. [Google Scholar] [CrossRef] [PubMed]
- Asghar, H.; Diop, O.M.; Weldegebriel, G.; Malik, F.; Shetty, S.; Bassioni, L.E.; Akande, A.O.; Maamoun, E.A.; Zaidi, S.; Adeniji, A.J.; et al. Environmental Surveillance for Polioviruses in the Global Polio Eradication Initiative. J. Infect. Dis. 2014, 210 (Suppl. 1), S294–S303. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Alhama, J.; Maestre, J.P.; Martin, M.A.; Michan, C. Monitoring COVID-19 through SARS-CoV-2 quantification in wastewater: Progress, challenges and prospects. Microb. Biotechnol. 2021. online ahead of print. [Google Scholar] [CrossRef] [PubMed]
- Na Ayudhya, S.S.; Laksono, B.M.; van Riel, D. The pathogenesis and virulence of enterovirus-D68 infection. Virulence 2021, 12, 2060–2072. [Google Scholar] [CrossRef] [PubMed]
- Bisseux, M.; Colombet, J.; Mirand, A.; Roque-Afonso, A.-M.; Abravanel, F.; Izopet, J.; Archimbaud, C.; Peigue-Lafeuille, H.; Debroas, D.; Bailly, J.-L.; et al. Monitoring human enteric viruses in wastewater and relevance to infections encountered in the clinical setting: A one-year experiment in central France, 2014 to 2015. Eurosurveillance 2018, 23, 17-00237. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Tedcastle, A.; Wilton, T.; Pegg, E.; Klapsa, D.; Bujaki, E.; Mate, R.; Fritzsche, M.; Majumdar, M.; Martin, J. Detection of Enterovirus D68 in Wastewater Samples from the UK between July and November 2021. Viruses 2022, 14, 143. [Google Scholar] [CrossRef] [PubMed]
- Weil, M.; Mandelboim, M.; Mendelson, E.; Manor, Y.; Shulman, L.; Ram, D.; Barkai, G.; Shemer, Y.; Wolf, D.; Kra-Oz, Z.; et al. Human enterovirus D68 in clinical and sewage samples in Israel. J. Clin. Virol. 2016, 86, 52–55. [Google Scholar] [CrossRef] [PubMed]
- Seegen RV7 Kit. Available online: https://www.seegene.com/assays/allplex_rv_essential_assay (accessed on 10 April 2022).
- Bar-Or, I.; Weil, M.; Indenbaum, V.; Bucris, E.; Bar-Ilan, D.; Elul, M.; Levia, N.; Aguvaeva, I.; Cohen, Z.; Shirazi, R.; et al. Detection of SARS-CoV-2 variants by genomic analysis of wastewater samples in Israel. Sci. Total Environ. 2021, 789, 148002. [Google Scholar] [CrossRef] [PubMed]
- Ikuse, T.; Aizawa, Y.; Takihara, H.; Okuda, S.; Watanabe, K.; Saitoh, A. Development of Novel PCR Assays for Improved Detection of Enterovirus D68. J. Clin. Microbiol. 2021, 59, e0115121. [Google Scholar] [CrossRef] [PubMed]
- Dreier, J.; Störmer, M.; Kleesiek, K. Use of Bacteriophage MS2 as an Internal Control in Viral Reverse Transcription-PCR Assays. J. Clin. Microbiol. 2005, 43, 4551–4557. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Bioline SensiFast Mix. Available online: https://www.bioline.com/sensifast-probe-no-rox-one-step-kit.html (accessed on 10 April 2022).
- Reliance Master Mix. Available online: www.bio-rad.com/en-il/product/ (accessed on 10 April 2022).
- Megascript Kit. Available online: https://www.thermofisher.com/order/catalog/product/AM1334 (accessed on 10 April 2022).
- PSS MagLead. Available online: https://www.pss.co.jp/english/ (accessed on 10 April 2022).
- Primer Conversion Calculator. Available online: http://www.scienceprimer.com (accessed on 10 April 2022).
- ABI 3500 Sequencer. Available online: https://www.thermofisher.com/il/en/home/life-science/ (accessed on 10 April 2022).
- Kumar, S.; Stecher, G.; Tamura, K. MEGA7: Molecular Evolutionary Genetics Analysis Version 7.0 for Bigger Datasets. Mol. Biol. Evol. 2016, 33, 1870–1874. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Vircell Standard Control. Available online: https://en.vircell.com/products/amplirun-total-sars-cov-2-control (accessed on 10 April 2022).
- NCBI Database. Available online: https://www.ncbi.nlm.nih.gov/genbank/ (accessed on 10 April 2022).
- Benschop, K.S.; Albert, J.; Anton, A.; Andrés, C.; Aranzamendi, M.; Armannsdóttir, B.; Bailly, J.-L.; Baldanti, F.; Baldvinsdóttir, G.E.; Beard, S.; et al. Re-emergence of enterovirus D68 in Europe after easing the COVID-19 lockdown, September 2021. Eurosurveillance 2021, 26, 2100998. [Google Scholar] [CrossRef] [PubMed]
- Midgley, S.E.; Benschop, K.; Dyrdak, R.; Mirand, A.; Bailly, J.-L.; Bierbaum, S.; Buderus, S.; Böttcher, S.; Eis-Hübinger, A.-M.; Hönemann, M.; et al. Co-circulation of multiple enterovirus D68 subclades, including a novel B3 cluster, across Europe in a season of expected low prevalence, 2019/20. Eurosurveillance 2020, 25, 1900749. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Ikner, L.A.; Gerba, C.P.; Bright, K.R. Concentration and Recovery of Viruses from Water: A Comprehensive Review. Food Environ. Virol. 2012, 4, 41–67. [Google Scholar] [CrossRef] [PubMed]
- Schrader, C.; Schielke, A.; Ellerbroek, L.; Johne, R. PCR inhibitors-occurrence, properties and removal. J. Appl. Microbiol. 2012, 113, 1014–1026. [Google Scholar] [CrossRef] [PubMed]
- Hasan, S.W.; Ibrahim, Y.; Daou, M.; Kannout, H.; Jan, N.; Lopes, A.; Alsafar, H.; Yousef, A.F. Detection and quantification of SARS-CoV-2 RNA in wastewater and treated effluents: Surveillance of COVID-19 epidemic in the United Arab Emirates. Sci. Total Environ. 2021, 764, 142929. [Google Scholar] [CrossRef] [PubMed]
- Lu, D.; Huang, Z.; Luo, J.; Zhang, X.; Sha, S. Primary concentration-The critical step in implementing the wastewater based epi-demiology for the COVID-19 pandemic: A mini-review. Sci. Total Environ. 2020, 747, 141245. [Google Scholar] [CrossRef] [PubMed]

| Case No. | Sample Date | Age (Years) | Gender | Pre-Existing Disease (Asthma or Other) | Fever | Enteric Symptoms | Respiratory Symptoms | Neurological Symptoms | Dermatological Symptoms | Co-Infection |
|---|---|---|---|---|---|---|---|---|---|---|
| 1 | 23 May 2021 | 6 | M | Y (Other) | Y | N | Cough, shortness of breath | N | N | N |
| 2 | 5 May 2021 | 2 | F | Y | N | N | Asthma exacerbation | N | N | N |
| 3 | 13 June 2021 | 2 | M | Y (Asthma) | Y | N | Rhinorrhea, asthma exacerbation | N | N | N |
| 4 | 13 June 2021 | 3 | M | N | N | N | Rhinorrhea, asthma exacerbation | N | N | N |
| 5 | 13 June 2021 | 3 | M | Y (Asthma) | Y | N | Asthma exacerbation | N | N | N |
| 6 | 13 June 2021 | 2 | F | Y (Asthma) | Y | N | Asthma exacerbation | N | N | N |
| 7 | 14 June 2021 | 14 | M | Y (Other) | Y | N | Sore throat, weakness, tonsilitis, asthma exacerbation | N | N | N |
| 8 | 16 June 2021 | 0 | M | Y (Other) | Y | N | N | Aseptic meningitis | N | N |
| 9 | 17 June 2021 | 0 | M | Y (Other) | Y | N | Respiratory distress | N | N | MSSA |
| 10 | 17 June 2021 | 0 | M | N | Y | N | Stridor, bronchiolitis, asthma exacerbation | N | N | Adeno in stool |
| 11 | 25 June 2021 | 3 | M | Y (Other) | Y | N | Rhinorrhea | N | N | GAS mastoiditis |
| 12 | 27 June 2021 | 0 | M | Y (Asthma) | N | N | Asthma exacerbation | N | N | N |
| 13 | 27 June 2021 | 3 | F | N | N | N | Asthma exacerbation | N | N | N |
| 14 | 27 June 2021 | 7 | F | N | N | N | Asthma exacerbation | N | N | N |
| 15 | 28 June 2021 | 3 | M | Y (Other) | Y | N | Rhinorrhea | N | N | N |
| 16 | 29 June 2021 | 3 | F | Y (Other) | N | N | Cough, rhinorrhea | N | N | N |
| 17 | 29 June 2021 | 0 | F | N | N | N | Cough, rhinorrhea | Y | N | N |
| 18 | 1 July 2021 | 6 | F | Y (Other) | Y | N | Asthma exacerbation | N | N | N |
| 19 | 5 July 2021 | 3 | F | Y (Other) | N | N | Asthma exacerbation | N | N | N |
| 20 | 6 July 2021 | 0 | M | N | Y | N | N | Viral encephalitis | N | HHV6 positive in CSF |
| 21 | 6 July 2021 | 6 | M | N | Y | N | Throat pain, asthma exacerbation | N | N | N |
| 22 | 8 July 2021 | 0 | F | Y (Other) | Y | N | Cough, rhinorrhea | N | N | N |
| 23 | 8 July 2021 | 0 | F | N | N | N | N | N | N | N |
| 24 | 8 July 2021 | 2 | M | N | Y | N | Cough, rhinorrhea; Tonsilitis | N | N | rt pleuropneumonia |
| 25 | 11 July 2021 | 1 | M | Y (Asthma) | Y | N | Asthma exacerbation | N | N | N |
| 26 | 12 July 2021 | 1 | M | N | N | N | Rhinorrhea | N | N | N |
| 27 | 13 July 2021 | 2 | M | Y (Asthma) | Y | N | Asthma exacerbation | N | N | N |
| 28 | 13 July 2021 | 3 | F | Y (Asthma) | N | N | Asthma exacerbation | N | N | N |
| 29 | 15 July 2021 | 2 | F | Y (Other) | N | intussusception | ARDS | N | N | N |
| 30 | 25 July 2021 | 3 | F | N | Y | N | Weakness, asthma exacerbation | N | N | N |
| 31 | 1 August 2021 | 1 | M | Y (Other) | Y | N | Asthma exacerbation | N | N | N |
| 32 | 1 August 2021 | 3 | F | Y (Asthma) | Y | N | Asthma exacerbation | N | N | N |
| 33 | 10 October 2021 | 2 | M | Y (Other) | N | N | N | N | N | N |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2022 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Erster, O.; Bar-Or, I.; Levy, V.; Shatzman-Steuerman, R.; Sofer, D.; Weiss, L.; Vasserman, R.; Fratty, I.S.; Kestin, K.; Elul, M.; et al. Monitoring of Enterovirus D68 Outbreak in Israel by a Parallel Clinical and Wastewater Based Surveillance. Viruses 2022, 14, 1010. https://doi.org/10.3390/v14051010
Erster O, Bar-Or I, Levy V, Shatzman-Steuerman R, Sofer D, Weiss L, Vasserman R, Fratty IS, Kestin K, Elul M, et al. Monitoring of Enterovirus D68 Outbreak in Israel by a Parallel Clinical and Wastewater Based Surveillance. Viruses. 2022; 14(5):1010. https://doi.org/10.3390/v14051010
Chicago/Turabian StyleErster, Oran, Itay Bar-Or, Virginia Levy, Rachel Shatzman-Steuerman, Danit Sofer, Leah Weiss, Rinat Vasserman, Ilana S. Fratty, Klil Kestin, Michal Elul, and et al. 2022. "Monitoring of Enterovirus D68 Outbreak in Israel by a Parallel Clinical and Wastewater Based Surveillance" Viruses 14, no. 5: 1010. https://doi.org/10.3390/v14051010
APA StyleErster, O., Bar-Or, I., Levy, V., Shatzman-Steuerman, R., Sofer, D., Weiss, L., Vasserman, R., Fratty, I. S., Kestin, K., Elul, M., Levi, N., Alkrenawi, R., Mendelson, E., Mandelboim, M., & Weil, M. (2022). Monitoring of Enterovirus D68 Outbreak in Israel by a Parallel Clinical and Wastewater Based Surveillance. Viruses, 14(5), 1010. https://doi.org/10.3390/v14051010






