mRNA Vaccine Development for Emerging Animal and Zoonotic Diseases
Abstract
:1. Introduction
2. Forms of mRNA Vaccines
2.1. Non-Amplifying mRNA Vaccines
2.2. Self-Amplifying mRNA Vaccines
3. Delivery Platform and Formulation of mRNA Vaccines
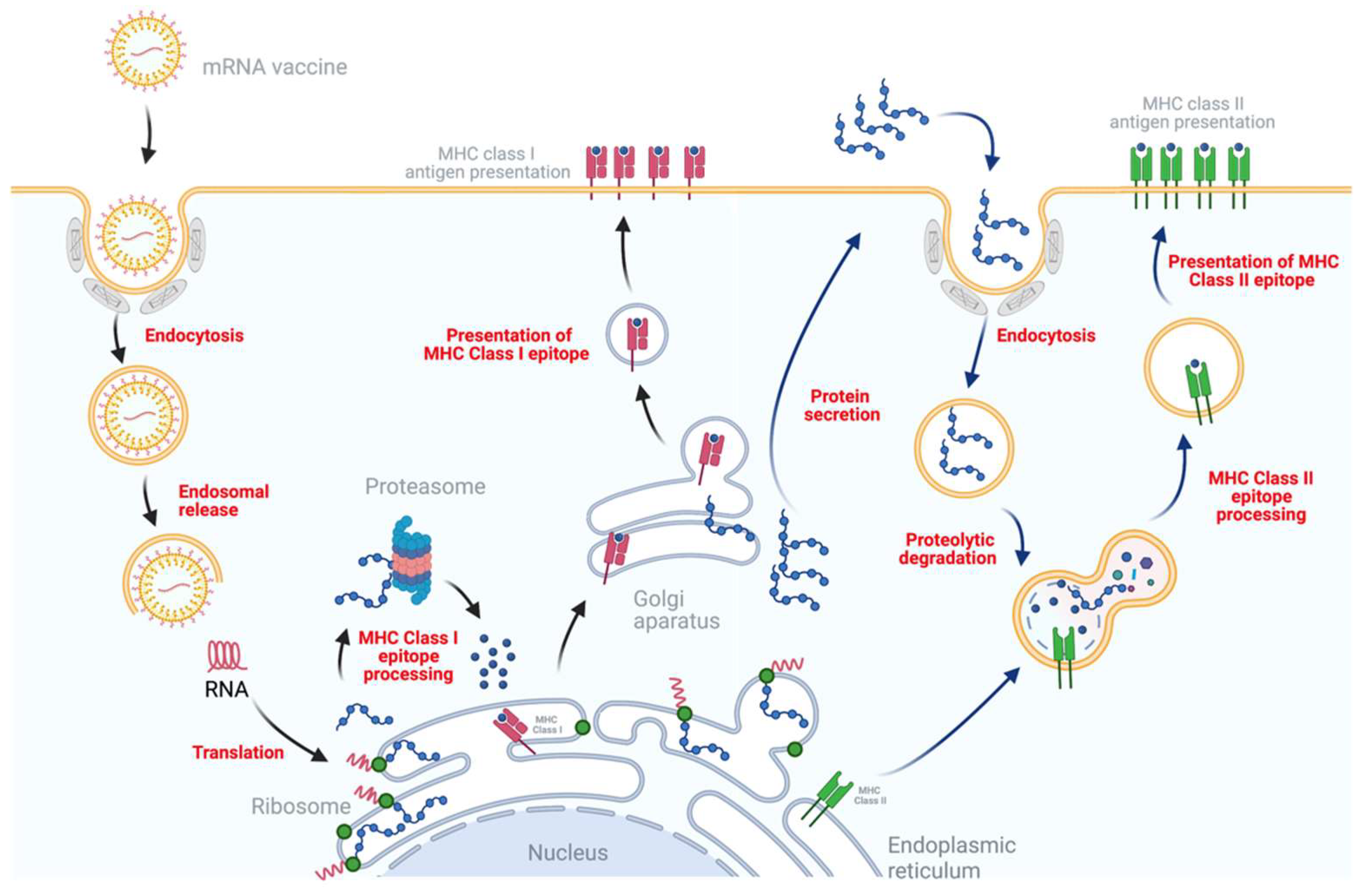
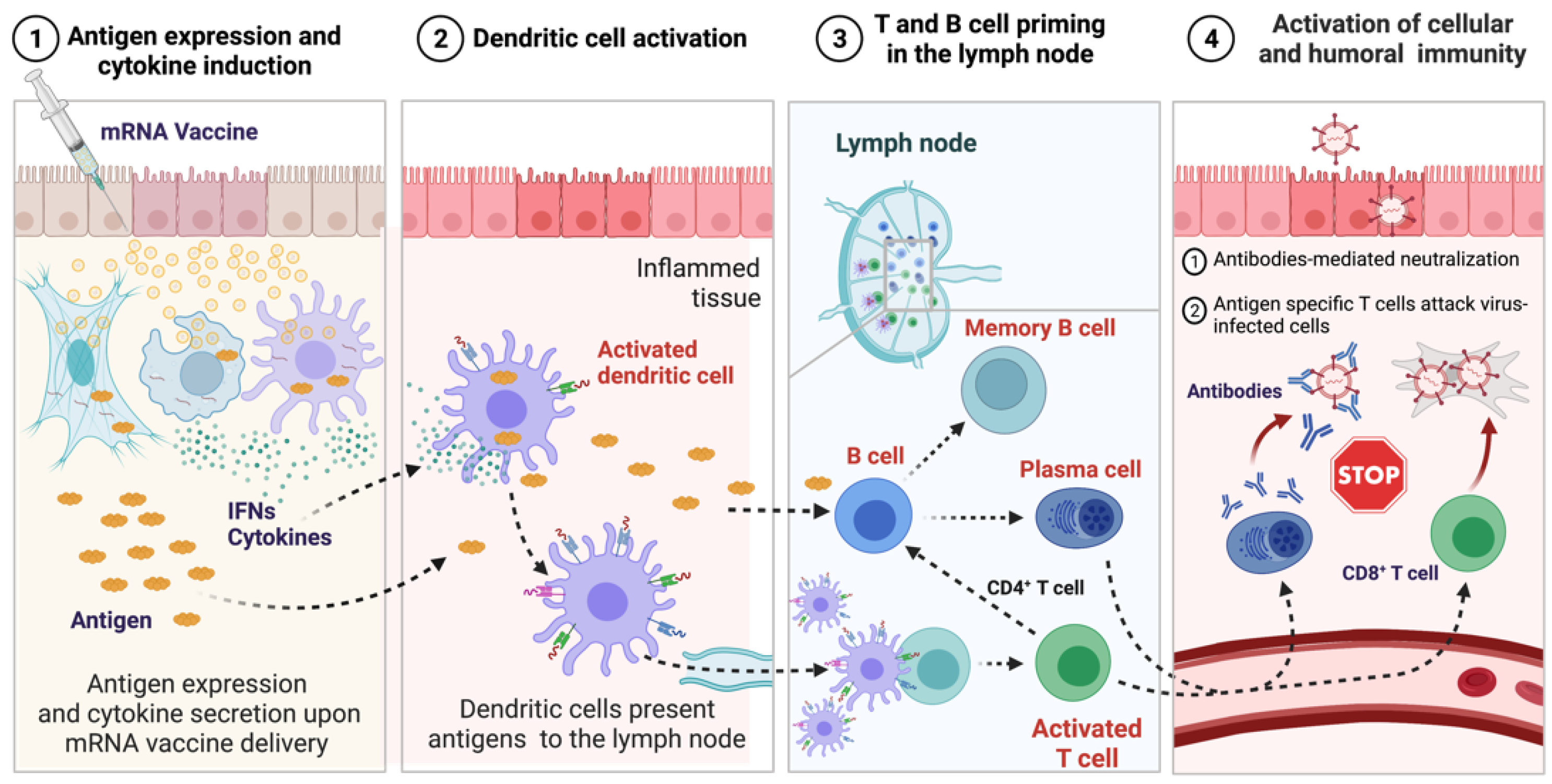
4. Mechanism of Immune Response Induced by mRNA Vaccines
5. Development of mRNA Vaccines for Prevention of Animal Infectious Diseases and Zoonoses
5.1. mRNA Vaccines for Foot-and-Mouth Disease
5.2. mRNA Vaccines for Rabies Disease
5.3. mRNA Vaccines for Influenza
5.4. mRNA Vaccines for Zoonotic Mosquito-Borne Flaviviruses
6. The Future Potential and Challenges of mRNA Vaccines in the Veterinary Field
6.1. T-Cell-Directed mRNA Vaccine for ASFV
6.2. Polyvalent Mosaic mRNA Vaccine for PRRSV
6.3. RBD-Based mRNA Vaccine for Animal Coronavirus
6.4. E2-Based mRNA Vaccine for Pestiviruses
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
References
- Excler, J.L.; Saville, M.; Berkley, S.; Kim, J.H. Vaccine Development for Emerging Infectious Diseases. Nat. Med. 2021, 27, 591–600. [Google Scholar] [CrossRef] [PubMed]
- Belongia, E.A.; Naleway, A.L. Smallpox Vaccine: The Good, the Bad, and the Ugly. Clin. Med. Res. 2003, 1, 87–92. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Roeder, P.; Mariner, J.; Kock, R. Rinderpest: The Veterinary Perspective on Eradication. Philos. Trans. R. Soc. Lond. Ser. B Biol. Sci. 2013, 368, 1–12. [Google Scholar] [CrossRef] [PubMed]
- Zhang, Y.; Zeng, G.; Pan, H.; Li, C.; Hu, Y.; Chu, K.; Han, W.; Chen, Z.; Tang, R.; Yin, W.; et al. Safety, Tolerability, and Immunogenicity of an Inactivated SARS-CoV-2 Vaccine in Healthy Adults Aged 18-59 Years: A Randomised, Double-Blind, Placebo-Controlled, Phase 1/2 Clinical Trial. Lancet Infect. Dis. 2021, 21, 181–192. [Google Scholar] [CrossRef]
- al Kaabi, N.; Zhang, Y.; Xia, S.; Yang, Y.; al Qahtani, M.M.; Abdulrazzaq, N.; Al Nusair, M.; Hassany, M.; Jawad, J.S.; Abdalla, J.; et al. Effect of 2 Inactivated SARS-CoV-2 Vaccines on Symptomatic COVID-19 Infection in Adults: A Randomized Clinical Trial. JAMA 2021, 326, 35–45. [Google Scholar] [CrossRef]
- Wu, S.; Huang, J.; Zhang, Z.; Wu, J.; Zhang, J.; Hu, H.; Zhu, T.; Zhang, J.; Luo, L.; Fan, P.; et al. Safety, Tolerability, and Immunogenicity of an Aerosolised Adenovirus Type-5 Vector-Based COVID-19 Vaccine (Ad5-NCoV) in Adults: Preliminary Report of an Open-Label and Randomised Phase 1 Clinical Trial. Lancet Infect. Dis. 2021, 21, 1654–1664. [Google Scholar] [CrossRef]
- Barda, N.; Dagan, N.; Ben-Shlomo, Y.; Kepten, E.; Waxman, J.; Ohana, R.; Hernán, M.A.; Lipsitch, M.; Kohane, I.; Netzer, D.; et al. Safety of the BNT162b2 MRNA Covid-19 Vaccine in a Nationwide Setting. N. Engl. J. Med. 2021, 385, 1078–1090. [Google Scholar] [CrossRef]
- Polack, F.P.; Thomas, S.J.; Kitchin, N.; Absalon, J.; Gurtman, A.; Lockhart, S.; Perez, J.L.; Pérez Marc, G.; Moreira, E.D.; Zerbini, C.; et al. Safety and Efficacy of the BNT162b2 MRNA Covid-19 Vaccine. N. Engl. J. Med. 2020, 383, 2603–2615. [Google Scholar] [CrossRef]
- Trimpert, J.; Dietert, K.; Firsching, T.C.; Ebert, N.; Thi Nhu Thao, T.; Vladimirova, D.; Kaufer, S.; Labroussaa, F.; Abdelgawad, A.; Conradie, A.; et al. Development of Safe and Highly Protective Live-Attenuated SARS-CoV-2 Vaccine Candidates by Genome Recoding. Cell Rep. 2021, 36, 109493. [Google Scholar] [CrossRef]
- Wang, Y.; Yang, C.; Song, Y.; Coleman, J.R.; Stawowczyk, M.; Tafrova, J.; Tasker, S.; Boltz, D.; Baker, R.; Garcia, L.; et al. Scalable Live-Attenuated SARS-CoV-2 Vaccine Candidate Demonstrates Preclinical Safety and Efficacy. Proc. Natl. Acad. Sci. USA 2021, 118, e2102775118. [Google Scholar] [CrossRef]
- Walter, E.B.; Talaat, K.R.; Sabharwal, C.; Gurtman, A.; Lockhart, S.; Paulsen, G.C.; Barnett, E.D.; Muñoz, F.M.; Maldonado, Y.; Pahud, B.A.; et al. Evaluation of the BNT162b2 Covid-19 Vaccine in Children 5 to 11 Years of Age. N. Engl. J. Med. 2022, 386, 35–46. [Google Scholar] [CrossRef] [PubMed]
- Sharif, N.; Alzahrani, K.J.; Ahmed, S.N.; Dey, S.K. Efficacy, Immunogenicity and Safety of COVID-19 Vaccines: A Systematic Review and Meta-Analysis. Front. Immunol. 2021, 12, 4149. [Google Scholar] [CrossRef] [PubMed]
- Baden, L.R.; el Sahly, H.M.; Essink, B.; Kotloff, K.; Frey, S.; Novak, R.; Diemert, D.; Spector, S.A.; Rouphael, N.; Creech, C.B.; et al. Efficacy and Safety of the MRNA-1273 SARS-CoV-2 Vaccine. N. Engl. J. Med. 2021, 384, 403–416. [Google Scholar] [CrossRef] [PubMed]
- Anand, P.; Stahel, V.P. Correction to: The Safety of Covid-19 MRNA Vaccines: A Review. Patient Saf. Surg. 2021, 15, 22. [Google Scholar] [CrossRef] [PubMed]
- Pardi, N.; Hogan, M.J.; Porter, F.W.; Weissman, D. MRNA Vaccines—A New Era in Vaccinology. Nat. Rev. Drug Discov. 2018, 17, 261–279. [Google Scholar] [CrossRef] [Green Version]
- Chaudhary, N.; Weissman, D.; Whitehead, K.A. MRNA Vaccines for Infectious Diseases: Principles, Delivery and Clinical Translation. Nat. Rev. Drug Discov. 2021, 20, 817–838. [Google Scholar] [CrossRef]
- Teijaro, J.R.; Farber, D.L. COVID-19 Vaccines: Modes of Immune Activation and Future Challenges. Nat. Rev. Immunol. 2021, 21, 195–197. [Google Scholar] [CrossRef]
- Wang, Y.; Zhang, Z.; Luo, J.; Han, X.; Wei, Y.; Wei, X. MRNA Vaccine: A Potential Therapeutic Strategy. Mol. Cancer 2021, 20, 33. [Google Scholar] [CrossRef]
- Tomley, F.M.; Shirley, M.W. Livestock Infectious Diseases and Zoonoses. Philos. Trans. R. Soc. Lond. Ser. B Biol. Sci. 2009, 364, 2637–2642. [Google Scholar] [CrossRef] [Green Version]
- Wang, T.; Sun, Y.; Qiu, H.J. African Swine Fever: An Unprecedented Disaster and Challenge to China. Infect. Dis. Poverty 2018, 7, 111. [Google Scholar] [CrossRef]
- Liu, J.; Liu, B.; Shan, B.; Wei, S.; An, T.; Shen, G.; Chen, Z. Prevalence of African Swine Fever in China, 2018–2019. J. Med. Virol. 2020, 92, 1023–1034. [Google Scholar] [CrossRef]
- Ge, S.; Li, J.; Fan, X.; Liu, F.; Li, L.; Wang, Q.; Ren, W.; Bao, J.; Liu, C.; Wang, H.; et al. Molecular Characterization of African Swine Fever Virus, China, 2018. Emerg. Infect. Dis. 2018, 24, 2131–2133. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Kumar, N.; Chander, Y.; Kumar, R.; Khandelwal, N.; Riyesh, T.; Chaudhary, K.; Shanmugasundaram, K.; Kumar, S.; Kumar, A.; Gupta, M.K.; et al. Isolation and Characterization of Lumpy Skin Disease Virus from Cattle in India. PLoS ONE 2021, 16, e0241022. [Google Scholar] [CrossRef] [PubMed]
- Sprygin, A.; Pestova, Y.; Wallace, D.B.; Tuppurainen, E.; Kononov, A.V. Transmission of Lumpy Skin Disease Virus: A Short Review. Virus Res. 2019, 269, 197637. [Google Scholar] [CrossRef]
- Jiang, Y.; Li, G.; Yu, L.; Li, L.; Zhang, Y.; Zhou, Y.; Tong, W.; Liu, C.; Gao, F.; Tong, G. Genetic Diversity of Porcine Reproductive and Respiratory Syndrome Virus (PRRSV) From 1996 to 2017 in China. Front. Microbiol. 2020, 11, 1–17. [Google Scholar] [CrossRef] [PubMed]
- Montaner-Tarbes, S.; del Portillo, H.A.; Montoya, M.; Fraile, L. Key Gaps in the Knowledge of the Porcine Respiratory Reproductive Syndrome Virus (PRRSV). Front. Vet. Sci. 2019, 6, 38. [Google Scholar] [CrossRef] [Green Version]
- Robin, M.; Page, P.; Archer, D.; Baylis, M. African Horse Sickness: The Potential for an Outbreak in Disease-Free Regions and Current Disease Control and Elimination Techniques. Equine Vet. J. 2016, 48, 659–669. [Google Scholar] [CrossRef] [Green Version]
- Rahman, M.T.; Sobur, M.A.; Islam, M.S.; Ievy, S.; Hossain, M.J.; Zowalaty, M.E.E.; Rahman, A.M.M.T.; Ashour, H.M. Zoonotic Diseases: Etiology, Impact, and Control. Microorganisms 2020, 8, 1405. [Google Scholar] [CrossRef]
- Kennedy, R.B.; Ovsyannikova, I.G.; Palese, P.; Poland, G.A. Current Challenges in Vaccinology. Front. Immunol. 2020, 11, 1181. [Google Scholar] [CrossRef]
- Pardi, N. MRNA Innovates the Vaccine Field. Vaccines 2021, 9, 486. [Google Scholar] [CrossRef]
- Choi, J.; Indrisiunaite, G.; Demirci, H.; Ieong, K.W.; Wang, J.; Petrov, A.; Prabhakar, A.; Rechavi, G.; Dominissini, D.; He, C.; et al. 2′-O-Methylation in MRNA Disrupts TRNA Decoding during Translation Elongation. Nat. Struct. Mol. Biol. 2018, 25, 208–216. [Google Scholar] [CrossRef] [PubMed]
- Elliott, B.A.; Ho, H.T.; Ranganathan, S.V.; Vangaveti, S.; Ilkayeva, O.; Abou Assi, H.; Choi, A.K.; Agris, P.F.; Holley, C.L. Modification of Messenger RNA by 2′-O-Methylation Regulates Gene Expression in Vivo. Nat. Commun. 2019, 10, 3401. [Google Scholar] [CrossRef] [Green Version]
- Assi, H.A.; Rangadurai, A.K.; Shi, H.; Liu, B.; Clay, M.C.; Erharter, K.; Kreutz, C.; Holley, C.L.; Al-Hashimi, H.M. 2′-O-Methylation Can Increase the Abundance and Lifetime of Alternative RNA Conformational States. Nucleic Acids Res. 2020, 48, 12365–12379. [Google Scholar] [CrossRef]
- Leppek, K.; Byeon, G.W.; Kladwang, W.; Wayment-Steele, H.K.; Kerr, C.H.; Xu, A.F.; Kim, D.S.; Topkar, V.V.; Choe, C.; Rothschild, D.; et al. Combinatorial Optimization of MRNA Structure, Stability, and Translation for RNA-Based Therapeutics. bioRxiv 2021, 29, 437587. [Google Scholar] [CrossRef]
- Zlotorynski, E. RNA Metabolism: The Short Tail That Wags the MRNA. Nature reviews. Mol. Cell Biol. 2018, 19, 2–3. [Google Scholar] [CrossRef]
- Kim, S.C.; Sekhon, S.S.; Shin, W.R.; Ahn, G.; Cho, B.K.; Ahn, J.Y.; Kim, Y.H. Modifications of MRNA Vaccine Structural Elements for Improving MRNA Stability and Translation Efficiency. Mol. Cell. Toxicol. 2021, 18, 1–8. [Google Scholar] [CrossRef] [PubMed]
- Jalkanen, A.L.; Coleman, S.J.; Wilusz, J. Determinants and Implications of MRNA Poly(A) Tail Size—Does This Protein Make My Tail Look Big? Semin. Cell Dev. Biol. 2014, 34, 24–32. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Eyler, D.E.; Franco, M.K.; Batool, Z.; Wu, M.Z.; Dubuke, M.L.; Dobosz-Bartoszek, M.; Jones, J.D.; Polikanov, Y.S.; Roy, B.; Koutmou, K.S. Pseudouridinylation of MRNA Coding Sequences Alters Translation. Proc. Natl. Acad. Sci. USA 2019, 116, 23068–23074. [Google Scholar] [CrossRef]
- Nelson, J.; Sorensen, E.W.; Mintri, S.; Rabideau, A.E.; Zheng, W.; Besin, G.; Khatwani, N.; Su, S.V.; Miracco, E.J.; Issa, W.J.; et al. Impact of MRNA Chemistry and Manufacturing Process on Innate Immune Activation. Sci. Adv. 2020, 6, eaaz6893. [Google Scholar] [CrossRef]
- Kauffman, K.J.; Mir, F.F.; Jhunjhunwala, S.; Kaczmarek, J.C.; Hurtado, J.E.; Yang, J.H.; Webber, M.J.; Kowalski, P.S.; Heartlein, M.W.; DeRosa, F.; et al. Efficacy and Immunogenicity of Unmodified and Pseudouridine-Modified MRNA Delivered Systemically with Lipid Nanoparticles in Vivo. Biomaterials 2016, 109, 78–87. [Google Scholar] [CrossRef] [Green Version]
- Wayment-Steele, H.K.; Kim, D.S.; Choe, C.A.; Nicol, J.J.; Wellington-Oguri, R.; Watkins, A.M.; Parra Sperberg, R.A.; Huang, P.S.; Participants, E.; Das, R. Theoretical Basis for Stabilizing Messenger RNA through Secondary Structure Design. Nucleic Acids Res. 2021, 49, 10604–10617. [Google Scholar] [CrossRef] [PubMed]
- Bloom, K.; van den Berg, F.; Arbuthnot, P. Self-Amplifying RNA Vaccines for Infectious Diseases. Gene Ther. 2021, 28, 117–129. [Google Scholar] [CrossRef]
- Blakney, A. The next Generation of RNA Vaccines: Self-Amplifying RNA. Biochemist 2021, 43, 14–17. [Google Scholar] [CrossRef]
- Blakney, A.K.; Ip, S.; Geall, A.J. An Update on Self-Amplifying MRNA Vaccine Development. Vaccines 2021, 9, 97. [Google Scholar] [CrossRef]
- Beissert, T.; Perkovic, M.; Vogel, A.; Erbar, S.; Walzer, K.C.; Hempel, T.; Brill, S.; Haefner, E.; Becker, R.; Türeci, Ö.; et al. A Trans-Amplifying RNA Vaccine Strategy for Induction of Potent Protective Immunity. Mol. Ther. 2020, 28, 119–128. [Google Scholar] [CrossRef] [PubMed]
- Fuller, D.H.; Berglund, P. Amplifying RNA Vaccine Development. N. Engl. J. Med. 2020, 382, 2469–2471. [Google Scholar] [CrossRef]
- Minnaert, A.K.; Vanluchene, H.; Verbeke, R.; Lentacker, I.; de Smedt, S.C.; Raemdonck, K.; Sanders, N.N.; Remaut, K. Strategies for Controlling the Innate Immune Activity of Conventional and Self-Amplifying MRNA Therapeutics: Getting the Message Across. Adv. Drug Deliv. Rev. 2021, 176, 113900. [Google Scholar] [CrossRef]
- Blakney, A.K.; McKay, P.F.; Bouton, C.R.; Hu, K.; Samnuan, K.; Shattock, R.J. Innate Inhibiting Proteins Enhance Expression and Immunogenicity of Self-Amplifying RNA. Mol. Ther. 2021, 29, 1174–1185. [Google Scholar] [CrossRef]
- Li, Y.; Teague, B.; Zhang, Y.; Su, Z.; Porter, E.; Dobosh, B.; Wagner, T.; Irvine, D.J.; Weiss, R. In Vitro Evolution of Enhanced RNA Replicons for Immunotherapy. Sci. Rep. 2019, 9, 6932. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Beissert, T.; Koste, L.; Perkovic, M.; Walzer, K.C.; Erbar, S.; Selmi, A.; Diken, M.; Kreiter, S.; Türeci, Ö.; Sahin, U. Improvement of In Vivo Expression of Genes Delivered by Self-Amplifying RNA Using Vaccinia Virus Immune Evasion Proteins. Hum. Gene Ther. 2017, 28, 1138–1146. [Google Scholar] [CrossRef] [Green Version]
- El-Jesr, M.; Teir, M.; de Motes, C.M. Vaccinia Virus Activation and Antagonism of Cytosolic DNA Sensing. Front. Immunol. 2020, 11, 2319. [Google Scholar] [CrossRef] [PubMed]
- Qiu, P.; Ziegelhoffer, P.; Sun, J.; Yang, N.S. Gene Gun Delivery of MRNA in Situ Results in Efficient Transgene Expression and Genetic Immunization. Gene Ther. 1996, 3, 262–268. [Google Scholar]
- Kowalski, P.S.; Rudra, A.; Miao, L.; Anderson, D.G. Delivering the Messenger: Advances in Technologies for Therapeutic MRNA Delivery. Mol. Ther. 2019, 27, 710–728. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Schoenmaker, L.; Witzigmann, D.; Kulkarni, J.A.; Verbeke, R.; Kersten, G.; Jiskoot, W.; Crommelin, D.J.A. MRNA-Lipid Nanoparticle COVID-19 Vaccines: Structure and Stability. Int. J. Pharm. 2021, 601, 120586. [Google Scholar] [CrossRef] [PubMed]
- Hou, X.; Zaks, T.; Langer, R.; Dong, Y. Lipid Nanoparticles for MRNA Delivery. Nat. Rev. Mater. 2021, 6, 1078–1094. [Google Scholar] [CrossRef]
- Reichmuth, A.M.; Oberli, M.A.; Jeklenec, A.; Langer, R.; Blankschtein, D. MRNA Vaccine Delivery Using Lipid Nanoparticles. Ther. Deliv. 2016, 7, 319–334. [Google Scholar] [CrossRef] [Green Version]
- Aditya, N.P.; Patankar, S.; Madhusudhan, B.; Murthy, R.S.R.; Souto, E.B. Arthemeter-Loaded Lipid Nanoparticles Produced by Modified Thin-Film Hydration: Pharmacokinetics, Toxicological and in Vivo Anti-Malarial Activity. Eur. J. Pharm. Sci. 2010, 40, 448–455. [Google Scholar] [CrossRef]
- Zhang, H. Thin-Film Hydration Followed by Extrusion Method for Liposome Preparation. Methods Mol. Biol. 2017, 1522, 17–22. [Google Scholar] [CrossRef]
- Otake, K.; Shimomura, T.; Goto, T.; Imura, T.; Furuya, T.; Yoda, S.; Takebayashi, Y.; Sakai, H.; Abe, M. Preparation of Liposomes Using an Improved Supercritical Reverse Phase Evaporation Method. Langmuir ACS J. Surf. Colloids 2006, 22, 2543–2550. [Google Scholar] [CrossRef]
- Huang, X.; Lee, R.J.; Qi, Y.; Li, Y.; Lu, J.; Meng, Q.; Teng, L.; Xie, J. Microfluidic Hydrodynamic Focusing Synthesis of Polymer-Lipid Nanoparticles for SiRNA Delivery. Oncotarget 2017, 8, 96826–96836. [Google Scholar] [CrossRef] [Green Version]
- Lu, M.; Ozcelik, A.; Grigsby, C.L.; Zhao, Y.; Guo, F.; Leong, K.W.; Huang, T.J. Microfluidic Hydrodynamic Focusing for Synthesis of Nanomaterials. Nano Today 2016, 11, 778–792. [Google Scholar] [CrossRef] [PubMed]
- Buschmann, M.D.; Carrasco, M.J.; Alishetty, S.; Paige, M.; Alameh, M.G.; Weissman, D. Nanomaterial Delivery Systems for MRNA Vaccines. Vaccines 2021, 9, 65. [Google Scholar] [CrossRef] [PubMed]
- Zhang, C.; Maruggi, G.; Shan, H.; Li, J. Advances in MRNA Vaccines for Infectious Diseases. Front. Immunol. 2019, 10, 594. [Google Scholar] [CrossRef] [Green Version]
- Saunders, K.O.; Pardi, N.; Parks, R.; Santra, S.; Mu, Z.; Sutherland, L.; Scearce, R.; Barr, M.; Eaton, A.; Hernandez, G.; et al. Lipid Nanoparticle Encapsulated Nucleoside-Modified MRNA Vaccines Elicit Polyfunctional HIV-1 Antibodies Comparable to Proteins in Nonhuman Primates. NPJ Vaccines 2021, 6, 50. [Google Scholar] [CrossRef] [PubMed]
- Pardi, N.; Parkhouse, K.; Kirkpatrick, E.; McMahon, M.; Zost, S.J.; Mui, B.L.; Tam, Y.K.; Karikó, K.; Barbosa, C.J.; Madden, T.D.; et al. Nucleoside-Modified MRNA Immunization Elicits Influenza Virus Hemagglutinin Stalk-Specific Antibodies. Nat. Commun. 2018, 9, 3361. [Google Scholar] [CrossRef] [Green Version]
- Richner, J.M.; Himansu, S.; Dowd, K.A.; Butler, S.L.; Salazar, V.; Fox, J.M.; Julander, J.G.; Tang, W.W.; Shresta, S.; Pierson, T.C.; et al. Modified MRNA Vaccines Protect against Zika Virus Infection. Cell 2017, 168, 1114–1125. [Google Scholar] [CrossRef] [Green Version]
- Pardi, N.; Hogan, M.J.; Naradikian, M.S.; Parkhouse, K.; Cain, D.W.; Jones, L.; Moody, M.A.; Verkerke, H.P.; Myles, A.; Willis, E.; et al. Nucleoside-Modified MRNA Vaccines Induce Potent T Follicular Helper and Germinal Center B Cell Responses. J. Exp. Med. 2018, 215, 1571–1588. [Google Scholar] [CrossRef] [PubMed]
- Alameh, M.G.; Tombácz, I.; Bettini, E.; Lederer, K.; Sittplangkoon, C.; Wilmore, J.R.; Gaudette, B.T.; Soliman, O.Y.; Pine, M.; Hicks, P.; et al. Lipid Nanoparticles Enhance the Efficacy of MRNA and Protein Subunit Vaccines by Inducing Robust T Follicular Helper Cell and Humoral Responses. Immunity 2021, 54, 2877–2892. [Google Scholar] [CrossRef]
- Frederickson, R.; Herzog, R.W. RNA-Based Vaccines and Innate Immune Activation: Not Too Hot and Not Too Cold. Mol. Ther. 2021, 29, 1365–1366. [Google Scholar] [CrossRef]
- Linares-Fernández, S.; Lacroix, C.; Exposito, J.Y.; Verrier, B. Tailoring MRNA Vaccine to Balance Innate/Adaptive Immune Response. Trends Mol. Med. 2020, 26, 311–323. [Google Scholar] [CrossRef]
- de Beuckelaer, A.; Pollard, C.; van Lint, S.; Roose, K.; van Hoecke, L.; Naessens, T.; Udhayakumar, V.K.; Smet, M.; Sanders, N.; Lienenklaus, S.; et al. Type I Interferons Interfere with the Capacity of MRNA Lipoplex Vaccines to Elicit Cytolytic T Cell Responses. Mol. Ther. 2016, 24, 2012–2020. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- de Beuckelaer, A.; Grooten, J.; de Koker, S. Type I Interferons Modulate CD8 + T Cell Immunity to MRNA Vaccines. Trends Mol. Med. 2017, 23, 216–226. [Google Scholar] [CrossRef] [PubMed]
- Pollard, C.; Rejman, J.; de Haes, W.; Verrier, B.; van Gulck, E.; Naessens, T.; de Smedt, S.; Bogaert, P.; Grooten, J.; Vanham, G.; et al. Type I IFN Counteracts the Induction of Antigen-Specific Immune Responses by Lipid-Based Delivery of MRNA Vaccines. Mol. Ther. J. Am. Soc. Gene Ther. 2013, 21, 251–259. [Google Scholar] [CrossRef] [Green Version]
- Nitika; Wei, J.; Hui, A.M. The Development of MRNA Vaccines for Infectious Diseases: Recent Updates. Infect. Drug Resist. 2021, 14, 5271–5285. [Google Scholar] [CrossRef] [PubMed]
- Oreskovic, Z.; Nechvatalova, K.; Krejci, J.; Kummer, V.; Faldyna, M. Aspects of Intradermal Immunization with Different Adjuvants: The Role of Dendritic Cells and Th1/Th2 Response. PLoS ONE 2019, 14, e0211896. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Zeng, C.; Zhang, C.; Walker, P.G.; Dong, Y. Formulation and Delivery Technologies for MRNA Vaccines. Curr. Top. Microbiol. Immunol. 2020, 6, 1–40. [Google Scholar] [CrossRef]
- Jamal, S.M.; Belsham, G.J. Foot-and-Mouth Disease: Past, Present and Future. Vet. Res. 2013, 44, 116. [Google Scholar] [CrossRef] [Green Version]
- Belsham, G.J. Towards Improvements in Foot-and-Mouth Disease Vaccine Performance. Acta Vet. Scand. 2020, 62, 20. [Google Scholar] [CrossRef]
- Pulido, M.R.; Sobrino, F.; Borrego, B.; Sáiz, M. RNA Immunization Can Protect Mice against Foot-and-Mouth Disease Virus. Antivir. Res. 2010, 85, 556–558. [Google Scholar] [CrossRef]
- Rodríguez-Pulido, M.; Sobrino, F.; Borrego, B.; Sáiz, M. Inoculation of Newborn Mice with Non-Coding Regions of Foot-and-Mouth Disease Virus RNA Can Induce a Rapid, Solid and Wide-Range Protection against Viral Infection. Antivir. Res. 2011, 92, 500–504. [Google Scholar] [CrossRef]
- Borrego, B.; Rodríguez-Pulido, M.; Mateos, F.; de la Losa, N.; Sobrino, F.; Sáiz, M. Delivery of Synthetic RNA Can Enhance the Immunogenicity of Vaccines against Foot-and-Mouth Disease Virus (FMDV) in Mice. Vaccine 2013, 31, 4375–4381. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Newman, J.; Rowlands, D.J.; Tuthill, T.J. An Engineered Maturation Cleavage Provides a Recombinant Mimic of Foot-and-Mouth Disease Virus Capsid Assembly-Disassembly. Life 2021, 11, 500. [Google Scholar] [CrossRef] [PubMed]
- Xiao, Y.; Zhang, S.; Yan, H.; Geng, X.; Wang, Y.; Xu, X.; Wang, M.; Zhang, H.; Huang, B.; Pang, W.; et al. The High Immunity Induced by the Virus-Like Particles of Foot-and-Mouth Disease Virus Serotype O. Front. Vet. Sci. 2021, 8, 87. [Google Scholar] [CrossRef] [PubMed]
- Kaur, M.; Garg, R.; Singh, S.; Bhatnagar, R. Rabies Vaccines: Where Do We Stand, Where Are We Heading? Expert Rev. Vaccines 2015, 14, 369–381. [Google Scholar] [CrossRef]
- Armbruster, N.; Jasny, E.; Petsch, B. Advances in RNA Vaccines for Preventive Indications: A Case Study of A Vaccine Against Rabies. Vaccines 2019, 7, 132. [Google Scholar] [CrossRef] [Green Version]
- Alberer, M.; Gnad-Vogt, U.; Hong, H.S.; Mehr, K.T.; Backert, L.; Finak, G.; Gottardo, R.; Bica, M.A.; Garofano, A.; Koch, S.D.; et al. Safety and Immunogenicity of a MRNA Rabies Vaccine in Healthy Adults: An Open-Label, Non-Randomised, Prospective, First-in-Human Phase 1 Clinical Trial. Lancet 2017, 390, 1511–1520. [Google Scholar] [CrossRef]
- Stitz, L.; Vogel, A.; Schnee, M.; Voss, D.; Rauch, S.; Mutzke, T.; Ketterer, T.; Kramps, T.; Petsch, B. A Thermostable Messenger RNA Based Vaccine against Rabies. PLoS Negl. Trop. Dis. 2017, 11, e0006108. [Google Scholar] [CrossRef] [Green Version]
- Schnee, M.; Vogel, A.B.; Voss, D.; Petsch, B.; Baumhof, P.; Kramps, T.; Stitz, L. An MRNA Vaccine Encoding Rabies Virus Glycoprotein Induces Protection against Lethal Infection in Mice and Correlates of Protection in Adult and Newborn Pigs. PLoS Negl. Trop. Dis. 2016, 10, e0004746. [Google Scholar] [CrossRef]
- Aldrich, C.; Leroux–Roels, I.; Huang, K.B.; Bica, M.A.; Loeliger, E.; Schoenborn-Kellenberger, O.; Walz, L.; Leroux-Roels, G.; von Sonnenburg, F.; Oostvogels, L. Proof-of-Concept of a Low-Dose Unmodified MRNA-Based Rabies Vaccine Formulated with Lipid Nanoparticles in Human Volunteers: A Phase 1 Trial. Vaccine 2021, 39, 1310–1318. [Google Scholar] [CrossRef]
- Stokes, A.; Pion, J.; Binazon, O.; Laffont, B.; Bigras, M.; Dubois, G.; Blouin, K.; Young, J.K.; Ringenberg, M.A.; ben Abdeljelil, N.; et al. Nonclinical Safety Assessment of Repeated Administration and Biodistribution of a Novel Rabies Self-Amplifying MRNA Vaccine in Rats. Regul. Toxicol. Pharmacol. 2020, 113, 104648. [Google Scholar] [CrossRef]
- Shrestha, S.S.; Swerdlow, D.L.; Borse, R.H.; Prabhu, V.S.; Finelli, L.; Atkins, C.Y.; Owusu-Edusei, K.; Bell, B.; Mead, P.S.; Biggerstaff, M.; et al. Estimating the Burden of 2009 Pandemic Influenza A (H1N1) in the United States (April 2009–April 2010). Clin. Infect. Dis. 2011, 52 (Suppl. 1), S75–S82. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Jester, B.J.; Uyeki, T.M.; Jernigan, D.B. Fifty Years of Influenza A(H3N2) Following the Pandemic of 1968. Am. J. Public Health 2020, 110, 669–676. [Google Scholar] [CrossRef] [PubMed]
- Vogel, A.B.; Lambert, L.; Kinnear, E.; Busse, D.; Erbar, S.; Reuter, K.C.; Wicke, L.; Perkovic, M.; Beissert, T.; Haas, H.; et al. Self-Amplifying RNA Vaccines Give Equivalent Protection against Influenza to MRNA Vaccines but at Much Lower Doses. Mol. Ther. 2018, 26, 446–455. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Magini, D.; Giovani, C.; Mangiavacchi, S.; MacCari, S.; Cecchi, R.; Ulmer, J.B.; de Gregorio, E.; Geall, A.J.; Brazzoli, M.; Bertholet, S. Self-Amplifying MRNA Vaccines Expressing Multiple Conserved Influenza Antigens Confer Protection against Homologous and Heterosubtypic Viral Challenge. PLoS ONE 2016, 11, e0161193. [Google Scholar] [CrossRef]
- McCullough, K.C.; Bassi, I.; Milona, P.; Suter, R.; Thomann-Harwood, L.; Englezou, P.; Démoulins, T.; Ruggli, N. Self-Replicating Replicon-RNA Delivery to Dendritic Cells by Chitosan-Nanoparticles for Translation In Vitro and In Vivo. Molecular therapy. Nucleic Acids 2014, 3, E173. [Google Scholar] [CrossRef]
- Freyn, A.W.; Ramos da Silva, J.; Rosado, V.C.; Bliss, C.M.; Pine, M.; Mui, B.L.; Tam, Y.K.; Madden, T.D.; de Souza Ferreira, L.C.; Weissman, D.; et al. A Multi-Targeting, Nucleoside-Modified MRNA Influenza Virus Vaccine Provides Broad Protection in Mice. Mol. Ther. 2020, 28, 1569–1584. [Google Scholar] [CrossRef] [PubMed]
- Abbasi, J. Moderna’s MRNA Vaccine for Seasonal Flu Enters Clinical Trials. JAMA 2021, 326, 1365. [Google Scholar] [CrossRef]
- Collins, N.D.; Barrett, A.D.T. Live Attenuated Yellow Fever 17D Vaccine: A Legacy Vaccine Still Controlling Outbreaks In Modern Day. Curr. Infect. Dis. Rep. 2017, 19, 14. [Google Scholar] [CrossRef]
- Mandl, C.W.; Aberle, J.H.; Aberle, S.W.; Holzmann, H.; Allison, S.L.; Heinz, F.X. In Vitro-Synthesized Infectious RNA as an Attenuated Live Vaccine in a Flavivirus Model. Nat. Med. 1998, 4, 1438–1440. [Google Scholar] [CrossRef]
- Aberle, J.H.; Aberle, S.W.; Kofler, R.M.; Mandl, C.W. Humoral and Cellular Immune Response to RNA Immunization with Flavivirus Replicons Derived from Tick-Borne Encephalitis Virus. J. Virol. 2005, 79, 15107–15113. [Google Scholar] [CrossRef] [Green Version]
- Kofler, R.M.; Aberle, J.H.; Aberle, S.W.; Allison, S.L.; Heinz, F.X.; Mandl, C.W. Mimicking Live Flavivirus Immunization with a Noninfectious RNA Vaccine. Proc. Natl. Acad. Sci. USA 2004, 101, 1951–1956. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Fleeton, M.N.; Chen, M.; Berglund, P.; Rhodes, G.; Parker, S.E.; Murphy, M.; Atkins, G.J.; Liljeström, P. Self-Replicative RNA Vaccines Elicit Protection against Influenza A Virus, Respiratory Syncytial Virus, and a Tickborne Encephalitis Virus. J. Infect. Dis. 2001, 183, 1395–1398. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Richner, J.M.; Jagger, B.W.; Shan, C.; Fontes, C.R.; Dowd, K.A.; Cao, B.; Himansu, S.; Caine, E.A.; Nunes, B.T.D.; Medeiros, D.B.A.; et al. Vaccine Mediated Protection Against Zika Virus-Induced Congenital Disease. Cell 2017, 170, 273–283. [Google Scholar] [CrossRef] [Green Version]
- Pattnaik, A.; Sahoo, B.R.; Pattnaik, A.K. Current Status of Zika Virus Vaccines: Successes and Challenges. Vaccines 2020, 8, 266. [Google Scholar] [CrossRef]
- Roth, C.; Cantaert, T.; Colas, C.; Prot, M.; Casadémont, I.; Levillayer, L.; Thalmensi, J.; Langlade-Demoyen, P.; Gerke, C.; Bahl, K.; et al. A Modified MRNA Vaccine Targeting Immunodominant NS Epitopes Protects Against Dengue Virus Infection in HLA Class I Transgenic Mice. Front. Immunol. 2019, 10, 1424. [Google Scholar] [CrossRef]
- Zhang, M.; Sun, J.; Li, M.; Jin, X. Modified MRNA-LNP Vaccines Confer Protection against Experimental DENV-2 Infection in Mice. Molecular therapy. Methods Clin. Dev. 2020, 18, 702–712. [Google Scholar] [CrossRef] [PubMed]
- VanBlargan, L.A.; Himansu, S.; Foreman, B.M.; Ebel, G.D.; Pierson, T.C.; Diamond, M.S. An MRNA Vaccine Protects Mice against Multiple Tick-Transmitted Flavivirus Infections. Cell Rep. 2018, 25, 3382–3392.e3. [Google Scholar] [CrossRef] [Green Version]
- Huber, S.R.; van Beek, J.; de Jonge, J.; Luytjes, W.; van Baarle, D. T Cell Responses to Viral Infections—Opportunities for Peptide Vaccination. Front. Immunol. 2014, 5, 171. [Google Scholar] [CrossRef]
- Zhao, D.; Liu, R.; Zhang, X.; Li, F.; Wang, J.; Zhang, J.; Liu, X.; Wang, L.; Zhang, J.; Wu, X.; et al. Replication and Virulence in Pigs of the First African Swine Fever Virus Isolated in China. Emerg. Microbes Infect. 2019, 8, 438–447. [Google Scholar] [CrossRef] [Green Version]
- Chen, W.; Zhao, D.; He, X.; Liu, R.; Wang, Z.; Zhang, X.; Li, F.; Shan, D.; Chen, H.; Zhang, J.; et al. A Seven-Gene-Deleted African Swine Fever Virus Is Safe and Effective as a Live Attenuated Vaccine in Pigs. Science China. Life Sci. 2020, 63, 623–634. [Google Scholar] [CrossRef]
- Borca, M.V.; Ramirez-Medina, E.; Silva, E.; Vuono, E.; Rai, A.; Pruitt, S.; Espinoza, N.; Velazquez-Salinas, L.; Gay, C.G.; Gladue, D.P. ASFV-G-∆I177L as an Effective Oral Nasal Vaccine against the Eurasia Strain of Africa Swine Fever. Viruses 2021, 13, 765. [Google Scholar] [CrossRef] [PubMed]
- Borca, M.V.; Ramirez-Medina, E.; Silva, E.; Vuono, E.; Rai, A.; Pruitt, S.; Holinka, L.G.; Velazquez-Salinas, L.; Zhu, J.; Gladue, D.P. Development of a Highly Effective African Swine Fever Virus Vaccine by Deletion of the I177L Gene Results in Sterile Immunity against the Current Epidemic Eurasia Strain. J. Virol. 2020, 94, e02017-19. [Google Scholar] [CrossRef] [PubMed]
- Ros-Lucas, A.; Correa-Fiz, F.; Bosch-Camós, L.; Rodriguez, F.; Alonso-Padilla, J. Computational Analysis of African Swine Fever Virus Protein Space for the Design of an Epitope-Based Vaccine Ensemble. Pathogens 2020, 9, 1078. [Google Scholar] [CrossRef] [PubMed]
- Bosch-Camós, L.; López, E.; Navas, M.J.; Pina-Pedrero, S.; Accensi, F.; Correa-Fiz, F.; Park, C.; Carrascal, M.; Domínguez, J.; Salas, M.L.; et al. Identification of Promiscuous African Swine Fever Virus T-Cell Determinants Using a Multiple Technical Approach. Vaccines 2021, 9, 29. [Google Scholar] [CrossRef]
- Burmakina, G.; Malogolovkin, A.; Tulman, E.R.; Xu, W.; Delhon, G.; Kolbasov, D.; Rock, D.L. Identification of T-Cell Epitopes in African Swine Fever Virus CD2v and C-Type Lectin Proteins. J. Gen. Virol. 2019, 100, 259–265. [Google Scholar] [CrossRef]
- Moise, L.; Gutiérrez, A.H.; Khan, S.; Tan, S.; Ardito, M.; Martin, W.D.; de Groot, A.S. New Immunoinformatics Tools for Swine: Designing Epitope-Driven Vaccines, Predicting Vaccine Efficacy, and Making Vaccines on Demand. Front. Immunol. 2020, 11, 2531. [Google Scholar] [CrossRef]
- Pérez-Núñez, D.; Sunwoo, S.Y.; Sánchez, E.G.; Haley, N.; García-Belmonte, R.; Nogal, M.; Morozov, I.; Madden, D.; Gaudreault, N.N.; Mur, L.; et al. Evaluation of a Viral DNA-Protein Immunization Strategy against African Swine Fever in Domestic Pigs. Vet. Immunol. Immunopathol. 2019, 208, 34–43. [Google Scholar] [CrossRef]
- Wei, C.; Dai, A.; Fan, J.; Li, Y.; Chen, A.; Zhou, X.; Luo, M.; Yang, X.; Liu, J. Efficacy of Type 2 PRRSV Vaccine against Challenge with the Chinese Lineage 1 (NADC30-like) PRRSVs in Pigs. Sci. Rep. 2019, 9, 10781. [Google Scholar] [CrossRef]
- Zhao, H.; Han, Q.; Zhang, L.; Zhang, Z.; Wu, Y.; Shen, H.; Jiang, P. Emergence of Mosaic Recombinant Strains Potentially Associated with Vaccine JXA1-R and Predominant Circulating Strains of Porcine Reproductive and Respiratory Syndrome Virus in Different Provinces of China. Virol. J. 2017, 14, 67. [Google Scholar] [CrossRef]
- Eclercy, J.; Renson, P.; Lebret, A.; Hirchaud, E.; Normand, V.; Andraud, M.; Paboeuf, F.; Blanchard, Y.; Rose, N.; Bourry, O. A Field Recombinant Strain Derived from Two Type 1 Porcine Reproductive and Respiratory Syndrome Virus (PRRSV-1) Modified Live Vaccines Shows Increased Viremia and Transmission in SPF Pigs. Viruses 2019, 11, 296. [Google Scholar] [CrossRef] [Green Version]
- Cui, J.; O’connell, C.M.; Hagen, C.; Sawicki, K.; Smyth, J.A.; Verardi, P.H.; van Kruiningen, H.J.; Garmendia, A.E. Broad Protection of Pigs against Heterologous PRRSV Strains by a GP5-Mosaic DNA Vaccine Prime/GP5-Mosaic RVaccinia (VACV) Vaccine Boost. Vaccines 2020, 8, 106. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Cui, J.; O’Connell, C.M.; Costa, A.; Pan, Y.; Smyth, J.A.; Verardi, P.H.; Burgess, D.J.; van Kruiningen, H.J.; Garmendia, A.E. A PRRSV GP5-Mosaic Vaccine: Protection of Pigs from Challenge and Ex Vivo Detection of IFNγ Responses against Several Genotype 2 Strains. PLoS ONE 2019, 14, e0208801. [Google Scholar] [CrossRef] [Green Version]
- Zhou, P.; Yang, X.L.; Wang, X.G.; Hu, B.; Zhang, L.; Zhang, W.; Si, H.R.; Zhu, Y.; Li, B.; Huang, C.L.; et al. A Pneumonia Outbreak Associated with a New Coronavirus of Probable Bat Origin. Nature 2020, 579, 270–273. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Shi, Z.; Hu, Z. A Review of Studies on Animal Reservoirs of the SARS Coronavirus. Virus Res. 2008, 133, 74–87. [Google Scholar] [CrossRef] [PubMed]
- Sahin, U.; Muik, A.; Derhovanessian, E.; Vogler, I.; Kranz, L.M.; Vormehr, M.; Baum, A.; Pascal, K.; Quandt, J.; Maurus, D.; et al. COVID-19 Vaccine BNT162b1 Elicits Human Antibody and T H 1 T Cell Responses. Nature 2020, 586, 594–599. [Google Scholar] [CrossRef]
- Mulligan, M.J.; Lyke, K.E.; Kitchin, N.; Absalon, J.; Gurtman, A.; Lockhart, S.; Neuzil, K.; Raabe, V.; Bailey, R.; Swanson, K.A.; et al. Phase I/II Study of COVID-19 RNA Vaccine BNT162b1 in Adults. Nature 2020, 586, 589–593. [Google Scholar] [CrossRef]
- Greaney, A.J.; Loes, A.N.; Gentles, L.E.; Crawford, K.H.D.; Starr, T.N.; Malone, K.D.; Chu, H.Y.; Bloom, J.D. Antibodies Elicited by MRNA-1273 Vaccination Bind More Broadly to the Receptor Binding Domain than Do Those from SARS-CoV-2 Infection. Sci. Transl. Med. 2021, 13, eabi9915. [Google Scholar] [CrossRef] [PubMed]
- Endsley, J.J.; Ridpath, J.F.; Neill, J.D.; Sandbulte, M.R.; Roth, J.A. Induction of T Lymphocytes Specific for Bovine Viral Diarrhea Virus in Calves with Maternal Antibody. Viral Immunol. 2004, 17, 13–23. [Google Scholar] [CrossRef]
- Franzoni, G.; Kurkure, N.V.; Edgar, D.S.; Everett, H.E.; Gerner, W.; Bodman-Smith, K.B.; Crooke, H.R.; Grahama, S.P. Assessment of the Phenotype and Functionality of Porcine CD8 T Cell Responses Following Vaccination with Live Attenuated Classical Swine Fever Virus (CSFV) and Virulent CSFV Challenge. Clin. Vaccine Immunol. 2013, 20, 1604–1616. [Google Scholar] [CrossRef] [Green Version]


| Types | Inactivated Vaccines | Live Attenuated Vaccines | Recombinant Subunit Vaccines or Synthetic Peptide Vaccines | Recombinant Live Vectored Vaccines | DNA Vaccines | mRNA Vaccines | |
|---|---|---|---|---|---|---|---|
| Characteristics | |||||||
| Advantages | Strong humoral immune responses | + | ++ | + | ++ | + | + |
| Strong cellular immune responses | - | ++ | - | ++ | + | ++ | |
| Rapid development and production | + | - | + | + | ++ | ++ | |
| Discrimination between infected and vaccinated animals | + | - | ++ | ++ | ++ | ++ | |
| Good stability | + | + | + | + | ++ | - | |
| Convenient storage conditions | + | - | + | - | ++ | - | |
| Disadvantages | Infectious | N | Y | N | Y | N | N |
| Reversion or mutation | - | ++ | - | + | - | - | |
| Risk at genetic integration | - | + | - | + | ++ | - | |
| Adjuvants required | Y | N | Y | N | N | N |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2022 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Le, T.; Sun, C.; Chang, J.; Zhang, G.; Yin, X. mRNA Vaccine Development for Emerging Animal and Zoonotic Diseases. Viruses 2022, 14, 401. https://doi.org/10.3390/v14020401
Le T, Sun C, Chang J, Zhang G, Yin X. mRNA Vaccine Development for Emerging Animal and Zoonotic Diseases. Viruses. 2022; 14(2):401. https://doi.org/10.3390/v14020401
Chicago/Turabian StyleLe, Ting, Chao Sun, Jitao Chang, Guijie Zhang, and Xin Yin. 2022. "mRNA Vaccine Development for Emerging Animal and Zoonotic Diseases" Viruses 14, no. 2: 401. https://doi.org/10.3390/v14020401
APA StyleLe, T., Sun, C., Chang, J., Zhang, G., & Yin, X. (2022). mRNA Vaccine Development for Emerging Animal and Zoonotic Diseases. Viruses, 14(2), 401. https://doi.org/10.3390/v14020401







