Strong SARS-CoV-2 N-Specific CD8+ T Immunity Induced by Engineered Extracellular Vesicles Associates with Protection from Lethal Infection in Mice
Abstract
:1. Introduction
2. Materials and Methods
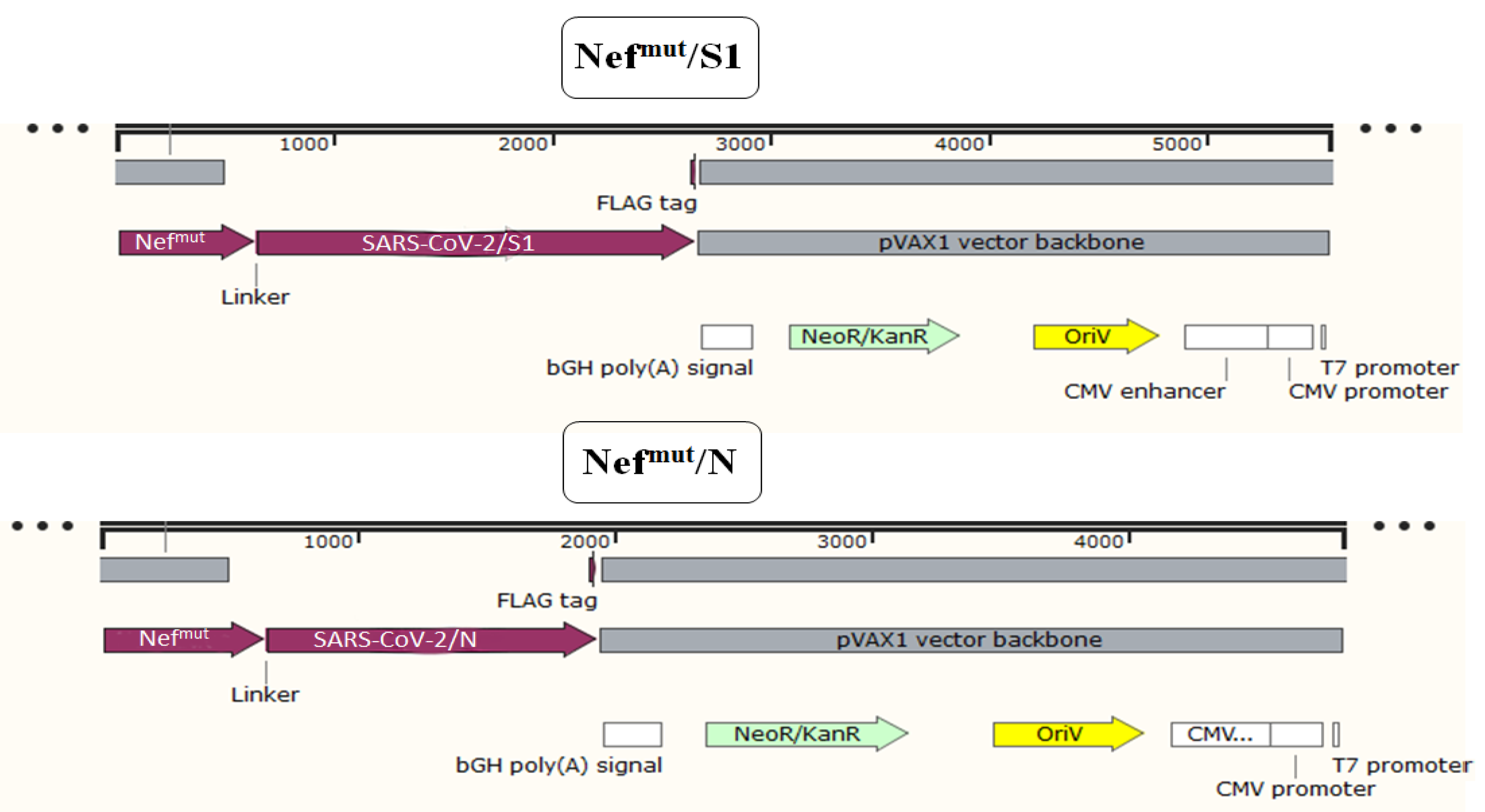
2.1. DNA Constructs
2.2. Animals and Authorizations
2.3. Mouse Immunization
2.4. Isolation of Cells from Blood, Bronchoalveolar Lavage Fluids (BALFs), Lungs, and Spleen
2.5. IFN-γ EliSpot Assay
2.6. Intracellular Cytokine Staining (ICS) and Flow Cytometry Analysis
2.7. SARS-CoV-2 Preparation and In Vitro Titration
2.8. Mouse Infection
2.9. Statistical Analysis
3. Results
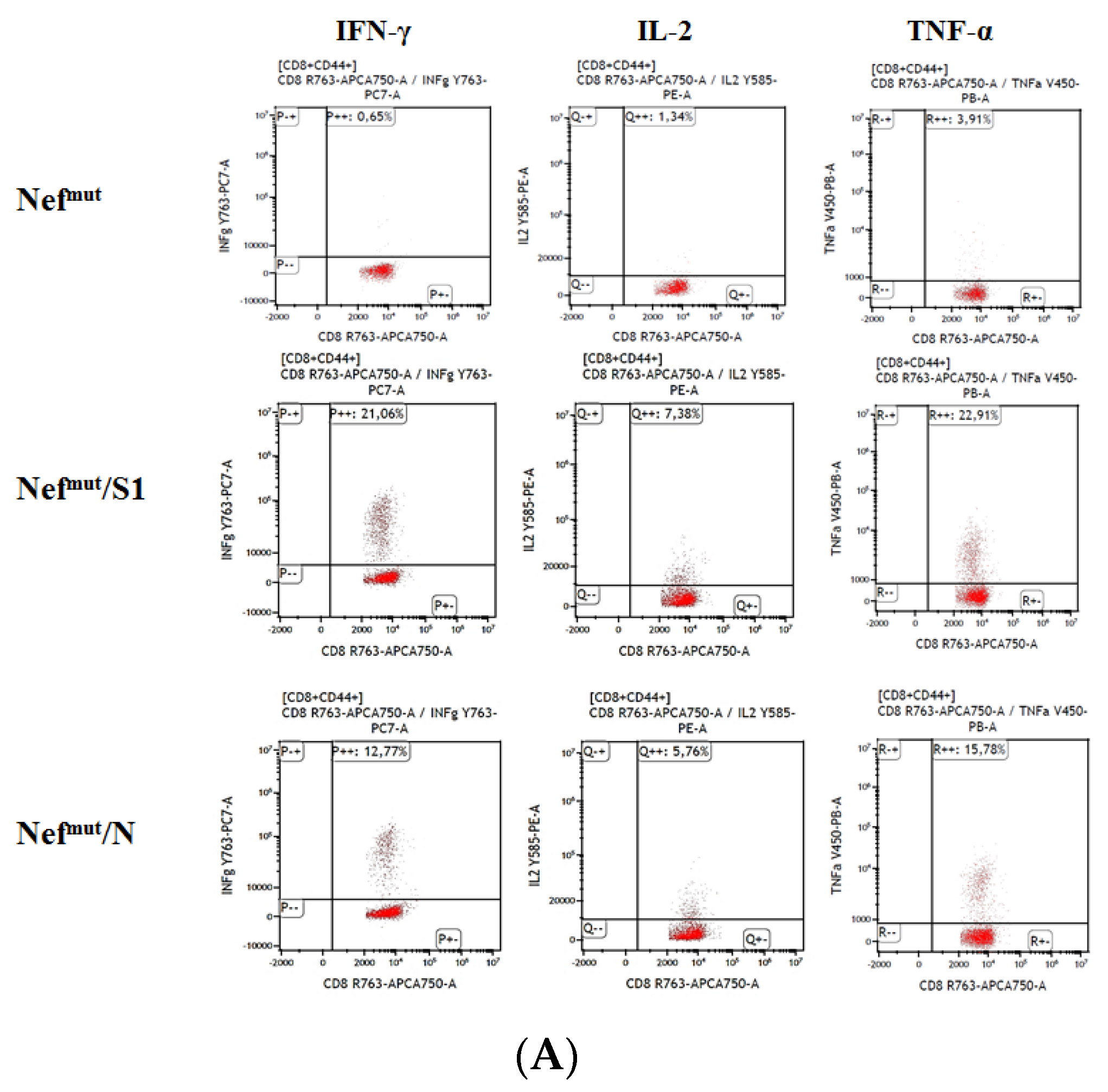
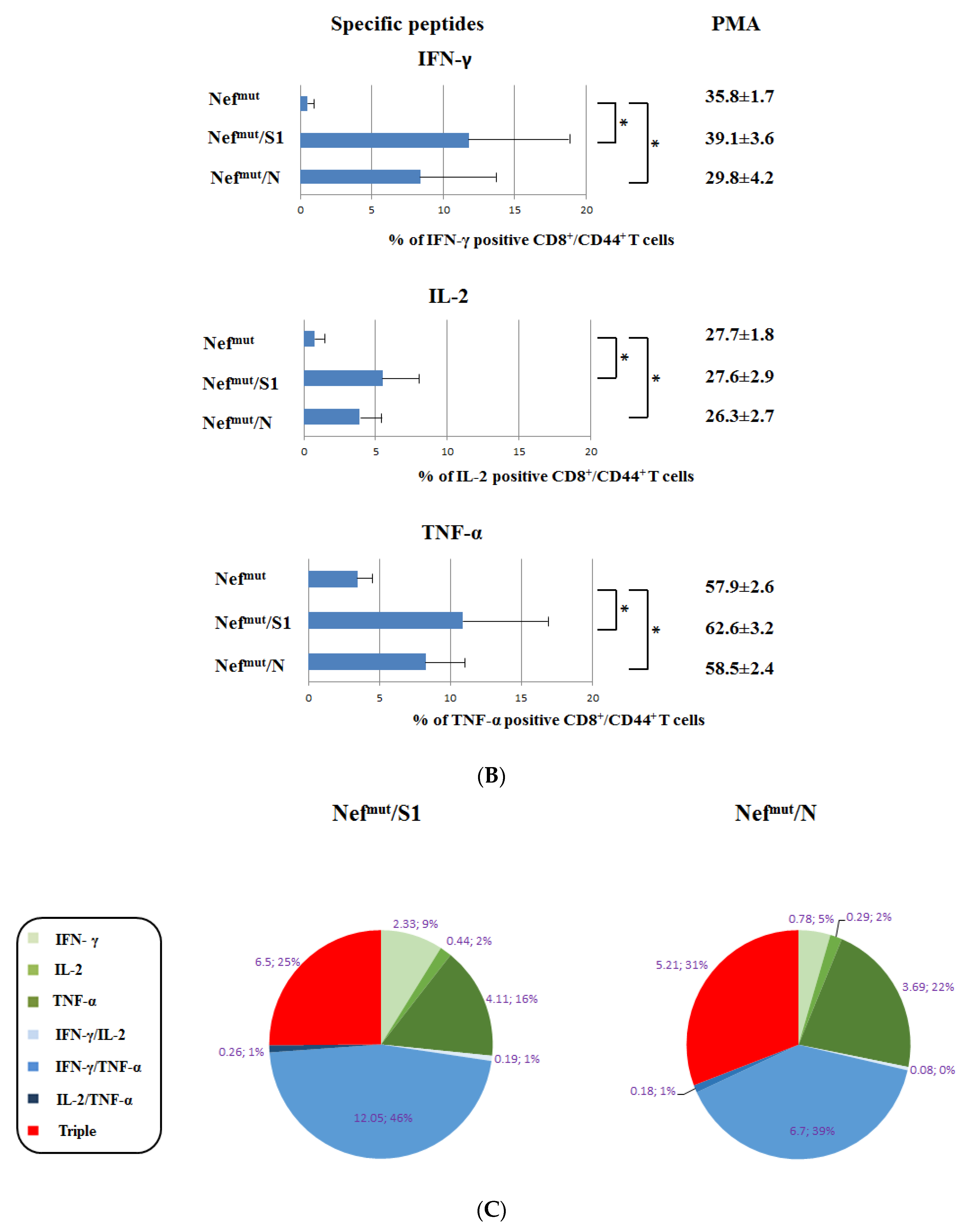
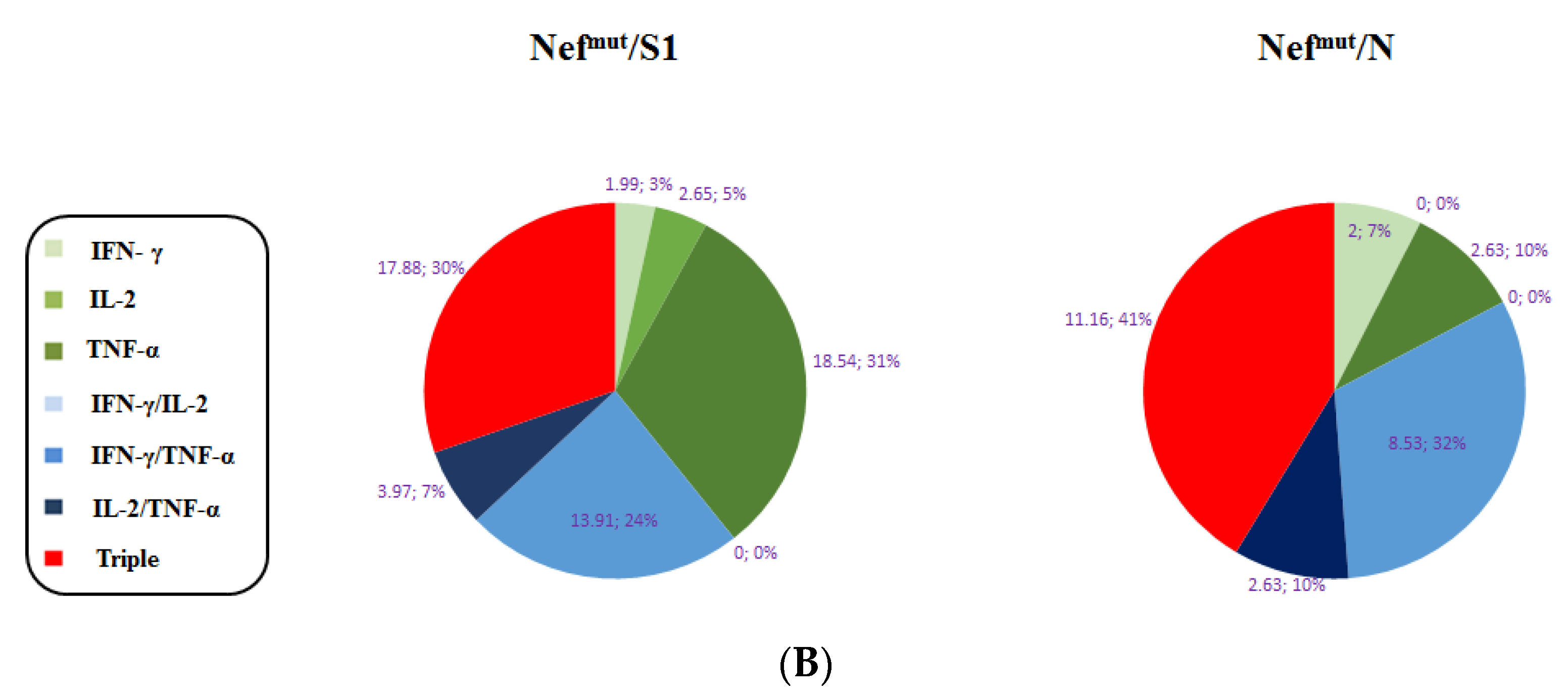
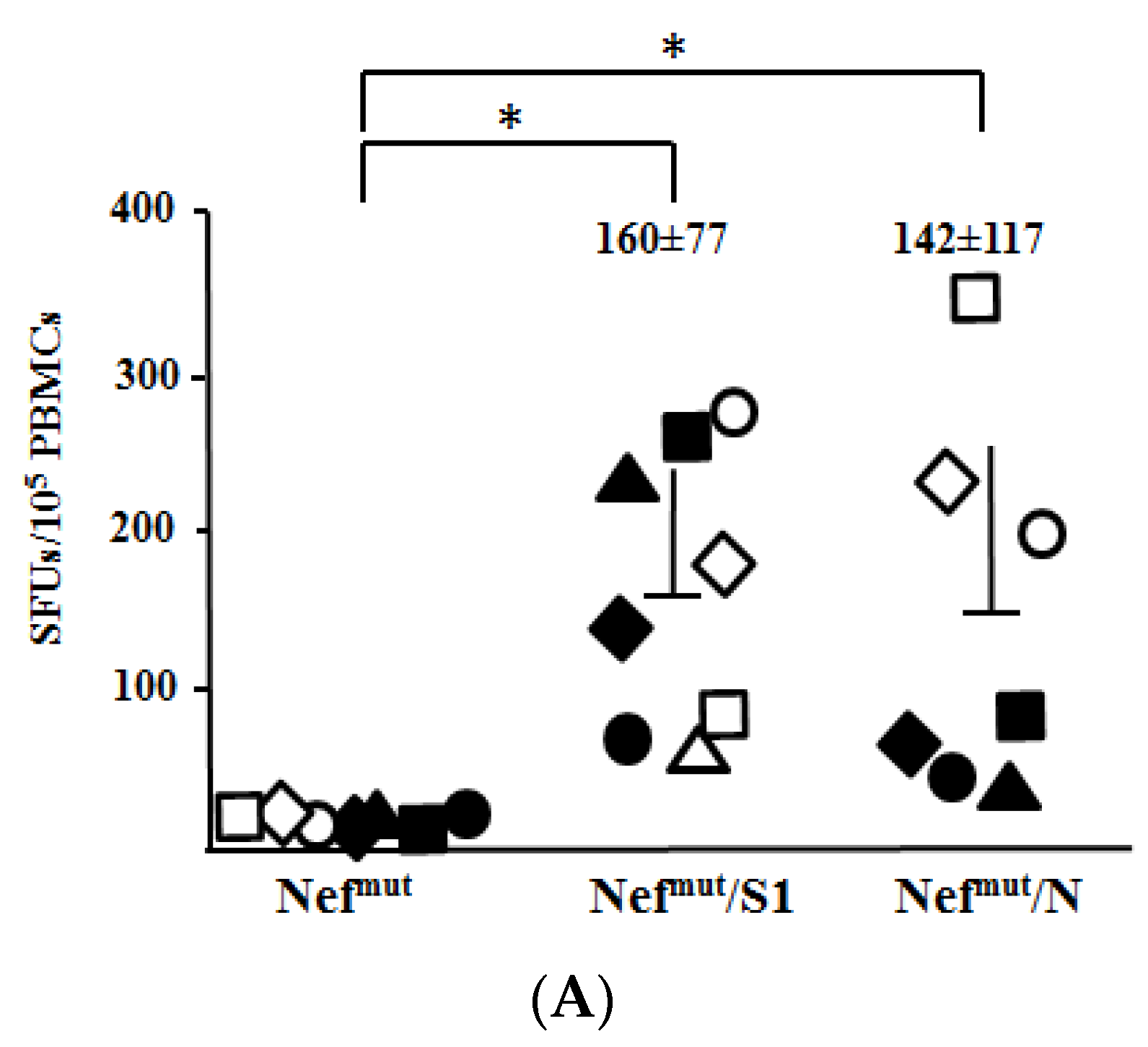
3.1. Induction of Polyfunctional Antigen-Specific CD8+ T Lymphocytes after IM Injection of DNA Vectors Expressing Either SARS-CoV-2 S1 or N Fused with Nefmut
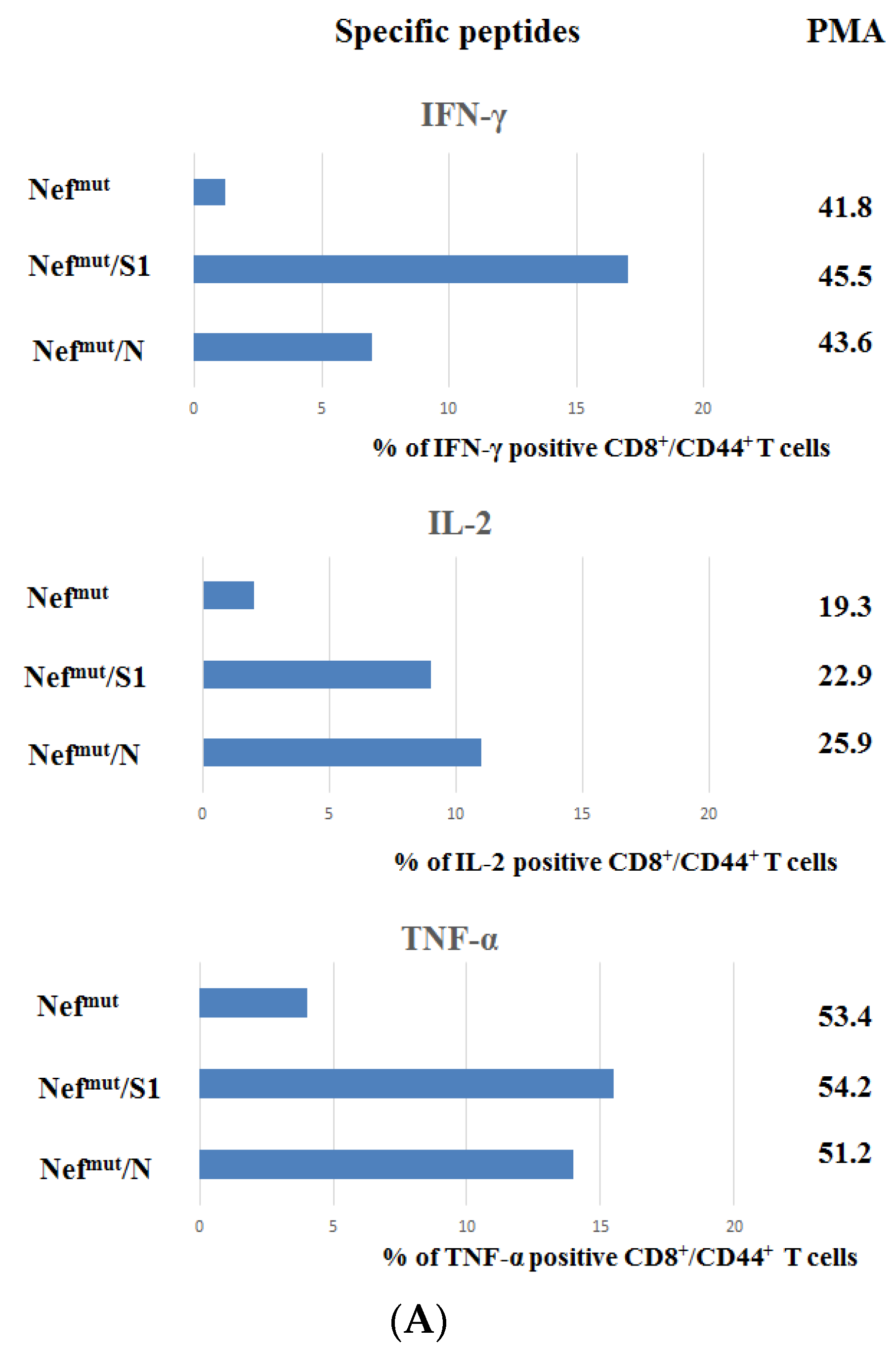
3.2. Detection of Polyfunctional Antigen-Specific CD8+ T Lymphocytes in BALFs from Immunized Mice
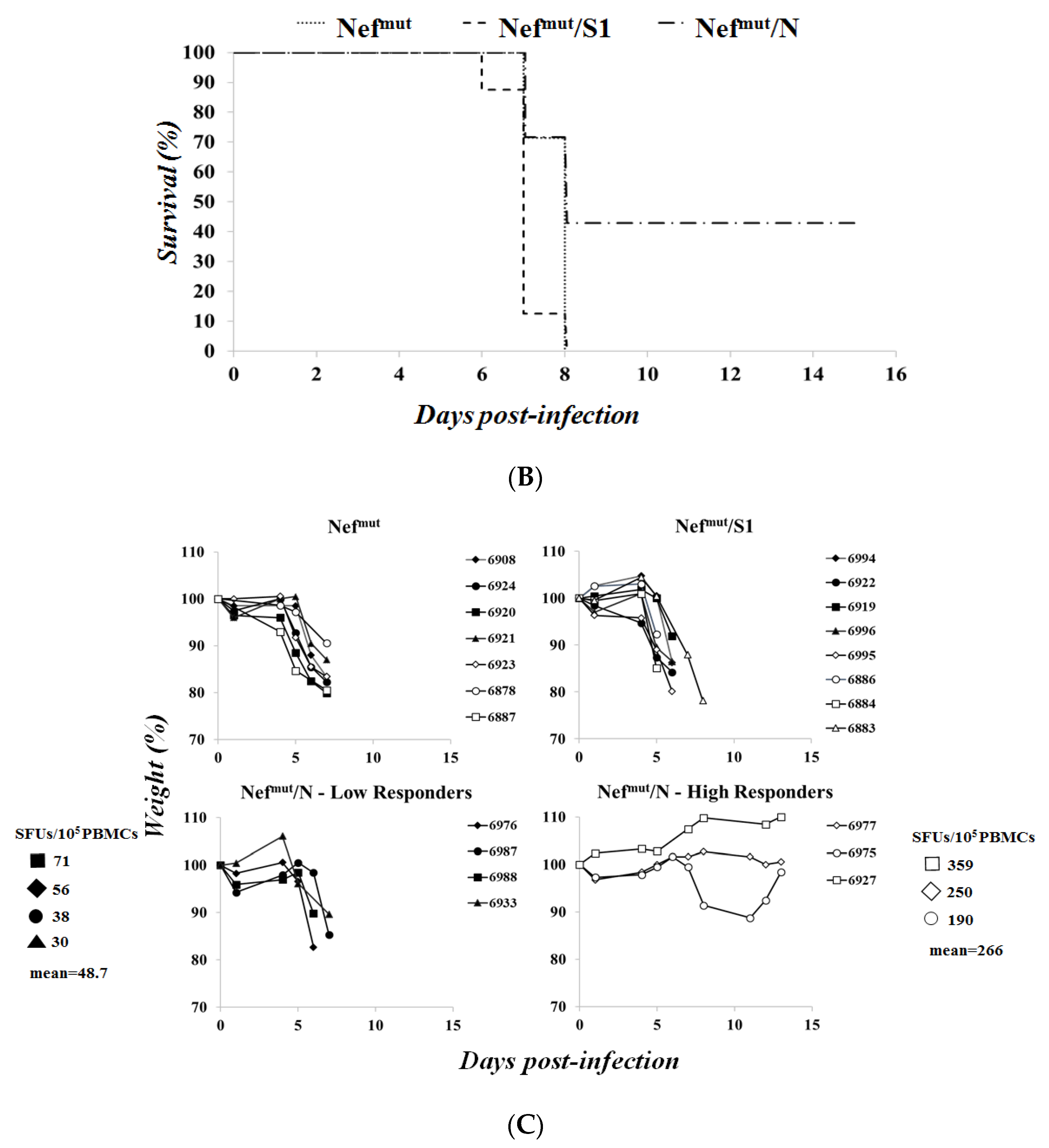
3.3. Association of High Levels of Circulating N-Specific CD8+ T Cells with Resistance to Lethal SARS-CoV-2 Infection
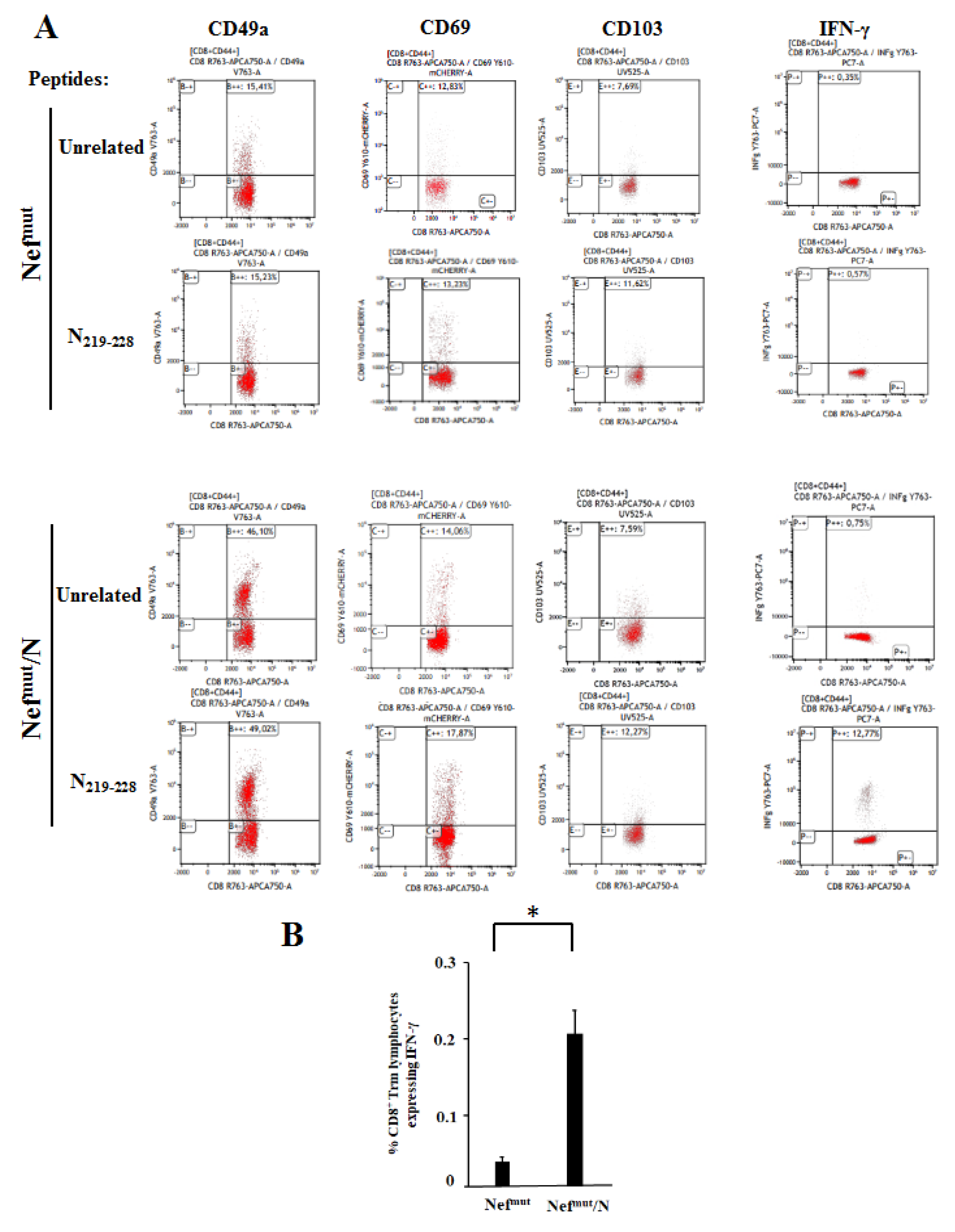
3.4. Detection of N-Specific CD8+ T-Resident Memory (Trm) Cells in Lungs of Immunized Mice
4. Discussion
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Plotkin, S.A. Vaccines: Correlates of Vaccine-Induced Immunity. Clin. Infect. Dis. 2008, 47, 401–409. [Google Scholar] [CrossRef] [Green Version]
- McMahan, K.; Yu, J.; Mercado, N.B.; Loos, C.; Tostanoski, L.H.; Chandrashekar, A.; Liu, J.; Peter, L.; Atyeo, C.; Zhu, A.; et al. Correlates of protection against SARS-CoV-2 in rhesus macaques. Nature 2021, 590, 630–634. [Google Scholar] [CrossRef] [PubMed]
- Moderbacher, C.R.; Ramirez, S.I.; Dan, J.M.; Grifoni, A.; Hastie, K.M.; Weiskopf, D.; Belanger, S.; Abbott, R.K.; Kim, C.; Choi, J.; et al. Antigen-Specific Adaptive Immunity to SARS-CoV-2 in Acute COVID-19 and Associations with Age and Disease Severity. Cell 2020, 183, 996–1012.e19. [Google Scholar] [CrossRef] [PubMed]
- Peng, Y.; Mentzer, A.J.; Liu, G.; Yao, X.; Yin, Z.; Dong, D.; Dejnirattisai, W.; Rostron, T.; Supasa, P.; Liu, C.; et al. Broad and strong memory CD4+ and CD8+ T cells induced by SARS-CoV-2 in UK convalescent individuals following COVID-19. Nat. Immunol. 2020, 21, 1336–1345. [Google Scholar] [CrossRef]
- Westmeier, J.; Paniskaki, K.; Karaköse, Z.; Werner, T.; Sutter, K.; Dolff, S.; Overbeck, M.; Limmer, A.; Liu, J.; Zheng, X.; et al. Impaired cytotoxic CD8+ T cell response in elderly COVID-19 patients. mBio 2020, 11, 11. [Google Scholar] [CrossRef] [PubMed]
- Tan, A.T.; Linster, M.; Tan, C.W.; Le Bert, N.; Ni Chia, W.; Kunasegaran, K.; Zhuang, Y.; Tham, C.Y.L.; Chia, A.; Smith, G.J.D.; et al. Early induction of functional SARS-CoV-2-specific T cells associates with rapid viral clearance and mild disease in COVID-19 patients. Cell Rep. 2021, 34, 108728. [Google Scholar] [CrossRef]
- Schulien, I.; Kemming, J.; Oberhardt, V.; Wild, K.; Seidel, L.M.; Killmer, S.; Daul, F.; Lago, M.S.; Decker, A.; Luxenburger, H.; et al. Characterization of pre-existing and induced SARS-CoV-2-specific CD8+ T cells. Nat. Med. 2020, 27, 78–85. [Google Scholar] [CrossRef]
- Redd, A.D.; Nardin, A.; Kared, H.; Bloch, E.M.; Pekosz, A.; Laeyendecker, O.; Abel, B.; Fehlings, M.; Quinn, T.C.; Tobian, A.A.R. CD8+ T-Cell Responses in COVID-19 Convalescent Individuals Target Conserved Epitopes from Multiple Prominent SARS-CoV-2 Circulating Variants. Open Forum Infect. Dis. 2021, 8, ofab143. [Google Scholar] [CrossRef]
- Tarke, A.; Sidney, J.; Methot, N.; Yu, E.D.; Zhang, Y.; Dan, J.M.; Goodwin, B.; Rubiro, P.; Sutherland, A.; Wang, E.; et al. Impact of SARS-CoV-2 variants on the total CD4+ and CD8+ T cell reactivity in infected or vaccinated individuals. Cell Rep. Med. 2021, 2, 100355. [Google Scholar] [CrossRef]
- Ferretti, A.P.; Kula, T.; Wang, Y.; Nguyen, D.M.; Weinheimer, A.; Dunlap, G.S.; Xu, Q.; Nabilsi, N.; Perullo, C.R.; Cristofaro, A.W.; et al. Unbiased Screens Show CD8+ T Cells of COVID-19 Patients Recognize Shared Epitopes in SARS-CoV-2 that Largely Reside outside the Spike Protein. Immunity 2020, 53, 1095–1107.e3. [Google Scholar] [CrossRef]
- Le Bert, N.; Tan, A.T.; Kunasegaran, K.; Tham, C.Y.L.; Hafezi, M.; Chia, A.; Chng, M.H.Y.; Lin, M.; Tan, N.; Linster, M.; et al. SARS-CoV-2-specific T cell immunity in cases of COVID-19 and SARS, and uninfected controls. Nature 2020, 584, 457–462. [Google Scholar] [CrossRef] [PubMed]
- Ferrantelli, F.; Manfredi, F.; Chiozzini, C.; Leone, P.; Giovannelli, A.; Olivetta, E.; Federico, M. Long-Term Antitumor CD8+ T Cell Immunity Induced by Endogenously Engineered Extracellular Vesicles. Cancers 2021, 13, 2263. [Google Scholar] [CrossRef] [PubMed]
- Anticoli, S.; Aricò, E.; Arenaccio, C.; Manfredi, F.; Chiozzini, C.; Olivetta, E.; Ferrantelli, F.; Lattanzi, L.; D’Urso, M.T.; Proietti, E.; et al. Engineered exosomes emerging from muscle cells break immune tolerance to HER2 in transgenic mice and induce antigen-specific CTLs upon challenge by human dendritic cells. Klin. Wochenschr. 2018, 96, 211–221. [Google Scholar] [CrossRef] [PubMed]
- Van Niel, G.; D’Angelo, G.; Raposo, G. Shedding light on the cell biology of extracellular vesicles. Nat. Rev. Mol. Cell Biol. 2018, 19, 213–228. [Google Scholar] [CrossRef]
- Lattanzi, L.; Federico, M. A strategy of antigen incorporation into exosomes: Comparing cross-presentation levels of antigens delivered by engineered exosomes and by lentiviral virus-like particles. Vaccine 2012, 30, 7229–7237. [Google Scholar] [CrossRef]
- Anticoli, S.; Manfredi, F.; Chiozzini, C.; Arenaccio, C.; Olivetta, E.; Ferrantelli, F.; Capocefalo, A.; Falcone, E.; Ruggieri, A.; Federico, M. An Exosome-Based Vaccine Platform Imparts Cytotoxic T Lymphocyte Immunity Against Viral Antigens. Biotechnol. J. 2018, 13, e1700443. [Google Scholar] [CrossRef]
- Federico, M. Virus-Induced CD8+ T-Cell Immunity and Its Exploitation to Contain the SARS-CoV-2 Pandemic. Vaccines 2021, 9, 922. [Google Scholar] [CrossRef]
- Ferrantelli, F.; Chiozzini, C.; Manfredi, F.; Giovannelli, A.; Leone, P.; Federico, M. Simultaneous CD8+ T-Cell Immune Response against SARS-Cov-2 S, M, and N Induced by Endogenously Engineered Extracellular Vesicles in Both Spleen and Lungs. Vaccines 2021, 9, 240. [Google Scholar] [CrossRef]
- McCray, P.B., Jr.; Pewe, L.; Wohlford-Lenane, C.; Hickey, M.; Manzel, L.; Shi, L.; Netland, J.; Jia, H.P.; Halabi, C.; Sigmund, C.D.; et al. Lethal Infection of K18-hACE2 Mice Infected with Severe Acute Respiratory Syndrome Coronavirus. J. Virol. 2007, 81, 813–821. [Google Scholar] [CrossRef] [Green Version]
- Zhi, Y.; Kobinger, G.P.; Jordan, H.; Suchma, K.; Weiss, S.R.; Shen, H.; Schumer, G.; Gao, G.; Boyer, J.L.; Crystal, R.G.; et al. Identification of murine CD8 T cell epitopes in codon-optimized SARS-associated coronavirus spike protein. Virology 2005, 335, 34–45. [Google Scholar] [CrossRef] [Green Version]
- Zhao, J.; Zhao, J.; Perlman, S. T Cell Responses Are Required for Protection from Clinical Disease and for Virus Clearance in Severe Acute Respiratory Syndrome Coronavirus-Infected Mice. J. Virol. 2010, 84, 9318–9325. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Michelini, Z.; Mazzei, C.; Magurano, F.; Baggieri, M.; Marchi, A.; Andreotti, M.; Cara, A.; Gaudino, A.; Mazzalupi, M.; Antonelli, F.; et al. UltraViolet SANitizing System for Sterilization of Ambulances Fleets and for Real-Time Monitoring of Their Sterilization Level. Int. J. Environ. Res. Public Health 2021, 19, 331. [Google Scholar] [CrossRef] [PubMed]
- Di Bonito, P.; Chiozzini, C.; Arenaccio, C.; Anticoli, S.; Manfredi, F.; Olivetta, E.; Ferrantelli, F.; Falcone, E.; Ruggieri, A.; Federico, M. Antitumor HPV E7-specific CTL activity elicited by in vivo engineered exosomes produced through DNA inoculation. Int. J. Nanomed. 2017, 12, 4579–4591. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Takamura, S.; Kato, S.; Motozono, C.; Shimaoka, T.; Ueha, S.; Matsuo, K.; Miyauchi, K.; Masumoto, T.; Katsushima, A.; Nakayama, T.; et al. Interstitial-resident memory CD8+ T cells sustain frontline epithelial memory in the lung. J. Exp. Med. 2019, 216, 2736–2747. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Oladunni, F.S.; Park, J.-G.; Pino, P.A.; Gonzalez, O.; Akhter, A.; Allué-Guardia, A.; Olmo-Fontánez, A.; Gautam, S.; Garcia-Vilanova, A.; Ye, C.; et al. Lethality of SARS-CoV-2 infection in K18 human angiotensin-converting enzyme 2 transgenic mice. Nat. Commun. 2020, 11, 6122. [Google Scholar] [CrossRef]
- D’Aloja, P.; Santarcangelo, A.C.; Arold, S.T.; Baur, A.; Federico, M. Genetic and functional analysis of the human immunodeficiency virus (HIV) type 1-inhibiting F12-HIVnef allele. J. Gen. Virol. 2001, 82 Pt 11, 2735–2745. [Google Scholar] [CrossRef] [Green Version]
- Allie, S.R.; Bradley, J.E.; Mudunuru, U.; Schultz, M.D.; Graf, B.A.; Lund, F.E.; Randall, T.D. The establishment of resident memory B cells in the lung requires local antigen encounter. Nat. Immunol. 2019, 20, 97–108. [Google Scholar] [CrossRef] [PubMed]
- Gangadaran, P.; Li, X.J.; Lee, H.W.; Oh, J.M.; Kalimuthu, S.; Rajendran, R.L.; Son, S.H.; Baek, S.H.; Singh, T.D.; Zhu, L.; et al. A new bioluminescent reporter system to study the biodistribution of systematically injected tumor-derived bioluminescent extracellular vesicles in mice. Oncotarget 2017, 8, 109894–109914. [Google Scholar] [CrossRef] [Green Version]
- Lai, C.P.; Mardini, O.; Ericsson, M.; Prabhakar, S.; Maguire, C.A.; Chen, J.W.; Tannous, B.A.; Breakefield, X.O. Dynamic Biodistribution of Extracellular Vesicles in Vivo Using a Multimodal Imaging Reporter. ACS Nano 2014, 8, 483–494. [Google Scholar] [CrossRef] [Green Version]
- Wiklander, O.P.B.; Nordin, J.Z.; O’Loughlin, A.; Gustafsson, Y.; Corso, G.; Mäger, I.; Vader, P.; Lee, Y.; Sork, H.; Seow, Y.; et al. Extracellular vesicle in vivo biodistribution is determined by cell source, route of administration and targeting. J. Extracell. Vesicles 2015, 4, 26316. [Google Scholar] [CrossRef] [Green Version]
- Takahashi, Y.; Nishikawa, M.; Shinotsuka, H.; Matsui, Y.; Ohara, S.; Imai, T.; Takakura, Y. Visualization and in vivo tracking of the exosomes of murine melanoma B16-BL6 cells in mice after intravenous injection. J. Biotechnol. 2013, 165, 77–84. [Google Scholar] [CrossRef] [PubMed]
- Anderson, K.G.; Sung, H.; Skon, C.N.; Lefrancois, L.; Deisinger, A.; Vezys, V.; Masopust, D. Cutting Edge: Intravascular Staining Redefines Lung CD8 T Cell Responses. J. Immunol. 2012, 189, 2702–2706. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Park, S.; Gebhardt, T.; Mackay, L.K. Tissue-Resident Memory T Cells in Cancer Immunosurveillance. Trends Immunol. 2019, 40, 735–747. [Google Scholar] [CrossRef] [PubMed]
- Cheuk, S.; Schlums, H.; Sérézal, I.G.; Martini, E.; Chiang, S.C.; Marquardt, N.; Gibbs, A.; Detlofsson, E.; Introini, A.; Forkel, M.; et al. CD49a Expression Defines Tissue-Resident CD8+ T Cells Poised for Cytotoxic Function in Human Skin. Immunity 2017, 46, 287–300. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Franciszkiewicz, K.; Le Floc’H, A.; Boutet, M.; Vergnon, I.; Schmitt, A.; Mami-Chouaib, F. CD103 or LFA-1 Engagement at the Immune Synapse between Cytotoxic T Cells and Tumor Cells Promotes Maturation and Regulates T-cell Effector Functions. Cancer Res. 2013, 73, 617–628. [Google Scholar] [CrossRef] [Green Version]
- Le Floc’H, A.; Jalil, A.; Franciszkiewicz, K.; Validire, P.; Vergnon, I.; Mami-Chouaib, F.; Floc’H, A.L. Minimal Engagement of CD103 on Cytotoxic T Lymphocytes with an E-Cadherin-Fc Molecule Triggers Lytic Granule Polarization via a Phospholipase Cγ–Dependent Pathway. Cancer Res. 2011, 71, 328–338. [Google Scholar] [CrossRef] [Green Version]
- Tatsi, E.-B.; Filippatos, F.; Michos, A. SARS-CoV-2 variants and effectiveness of vaccines: A review of current evidence. Epidemiology Infect. 2021, 149, e237. [Google Scholar] [CrossRef]
- Wei, J.; Stoesser, N.; Matthews, P.C.; Ayoubkhani, D.; Studley, R.; Bell, I.; Bell, J.I.; Newton, J.N.; Farrar, J.; Diamond, I.; et al. Antibody responses to SARS-CoV-2 vaccines in 45,965 adults from the general population of the United Kingdom. Nat. Microbiol. 2021, 6, 1140–1149. [Google Scholar] [CrossRef]
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2022 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Ferrantelli, F.; Chiozzini, C.; Manfredi, F.; Leone, P.; Spada, M.; Di Virgilio, A.; Giovannelli, A.; Sanchez, M.; Cara, A.; Michelini, Z.; et al. Strong SARS-CoV-2 N-Specific CD8+ T Immunity Induced by Engineered Extracellular Vesicles Associates with Protection from Lethal Infection in Mice. Viruses 2022, 14, 329. https://doi.org/10.3390/v14020329
Ferrantelli F, Chiozzini C, Manfredi F, Leone P, Spada M, Di Virgilio A, Giovannelli A, Sanchez M, Cara A, Michelini Z, et al. Strong SARS-CoV-2 N-Specific CD8+ T Immunity Induced by Engineered Extracellular Vesicles Associates with Protection from Lethal Infection in Mice. Viruses. 2022; 14(2):329. https://doi.org/10.3390/v14020329
Chicago/Turabian StyleFerrantelli, Flavia, Chiara Chiozzini, Francesco Manfredi, Patrizia Leone, Massimo Spada, Antonio Di Virgilio, Andrea Giovannelli, Massimo Sanchez, Andrea Cara, Zuleika Michelini, and et al. 2022. "Strong SARS-CoV-2 N-Specific CD8+ T Immunity Induced by Engineered Extracellular Vesicles Associates with Protection from Lethal Infection in Mice" Viruses 14, no. 2: 329. https://doi.org/10.3390/v14020329
APA StyleFerrantelli, F., Chiozzini, C., Manfredi, F., Leone, P., Spada, M., Di Virgilio, A., Giovannelli, A., Sanchez, M., Cara, A., Michelini, Z., & Federico, M. (2022). Strong SARS-CoV-2 N-Specific CD8+ T Immunity Induced by Engineered Extracellular Vesicles Associates with Protection from Lethal Infection in Mice. Viruses, 14(2), 329. https://doi.org/10.3390/v14020329







