Clinical Application of Detecting COVID-19 Risks: A Natural Language Processing Approach
Abstract
1. Background
- We develop a biomedical NER pipeline to identify clinical as well as non-clinical named entities from the COVID-19 texts. We attempt to consolidate and explain data science best practices through this pipeline, with numerous convenient features that can be used as it is or as a starting point for further customization and improvement.
- We develop a new dataset by curating a large number of scientific publications and case reports on COVID-19, and we scientifically parse the text from these scientific articles and prepare a dataset from it. We annotate a part of this dataset on biomedical-named entities to prepare a gold-standard dataset to train the NER pipeline. A portion of the gold-standard dataset is also reserved as a test set.
- We de-identify the patients’ personal information after identifying the named entities, thus adhering to the Health Insurance Portability and Accountability Act (HIPAA) [13].
- We demonstrate the efficacy and utility of this pipeline by comparing it with the state-of-the-art methods on public benchmark datasets. We also show the key findings related to COVID-19 in the analysis.
2. Previous Work
3. Materials and Methods
3.1. Data Cohort
- We specify the timeline between November 2021 and March 2022 for data collection.
- We specify English as the language to get the publications.
- We exclude many early-pandemic scientific articles, the intuition being that the disease symptoms and diagnosis, drugs and vaccination information were not clear during that time.
- We specify the population groups in adults: 19–44 years, middle-aged: 45–64 years, aged: 65+ years, during data collection.
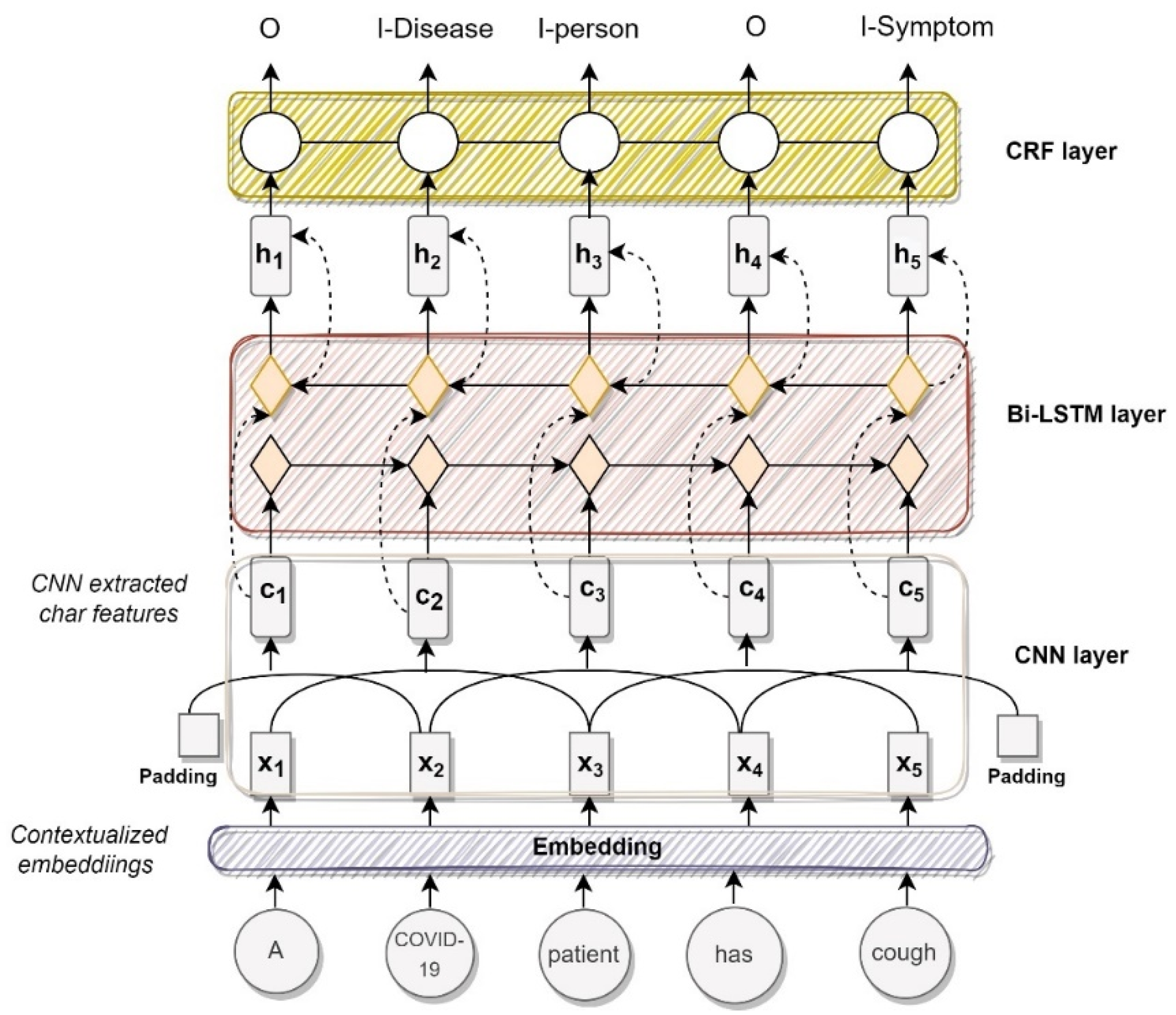
3.2. Biomedical Named Entity Recognition Pipeline Structure
3.3. Evaluation
- -
- Micro-average F1 to measures the F1-score of aggregated contributions of all classes.
- -
- Macro-average F1that adds all the measures (Precision, Recall, or F-Measure) and divides with the number of labels, which is more like an average.
4. Results
4.1. Comparison with Baseline Methods
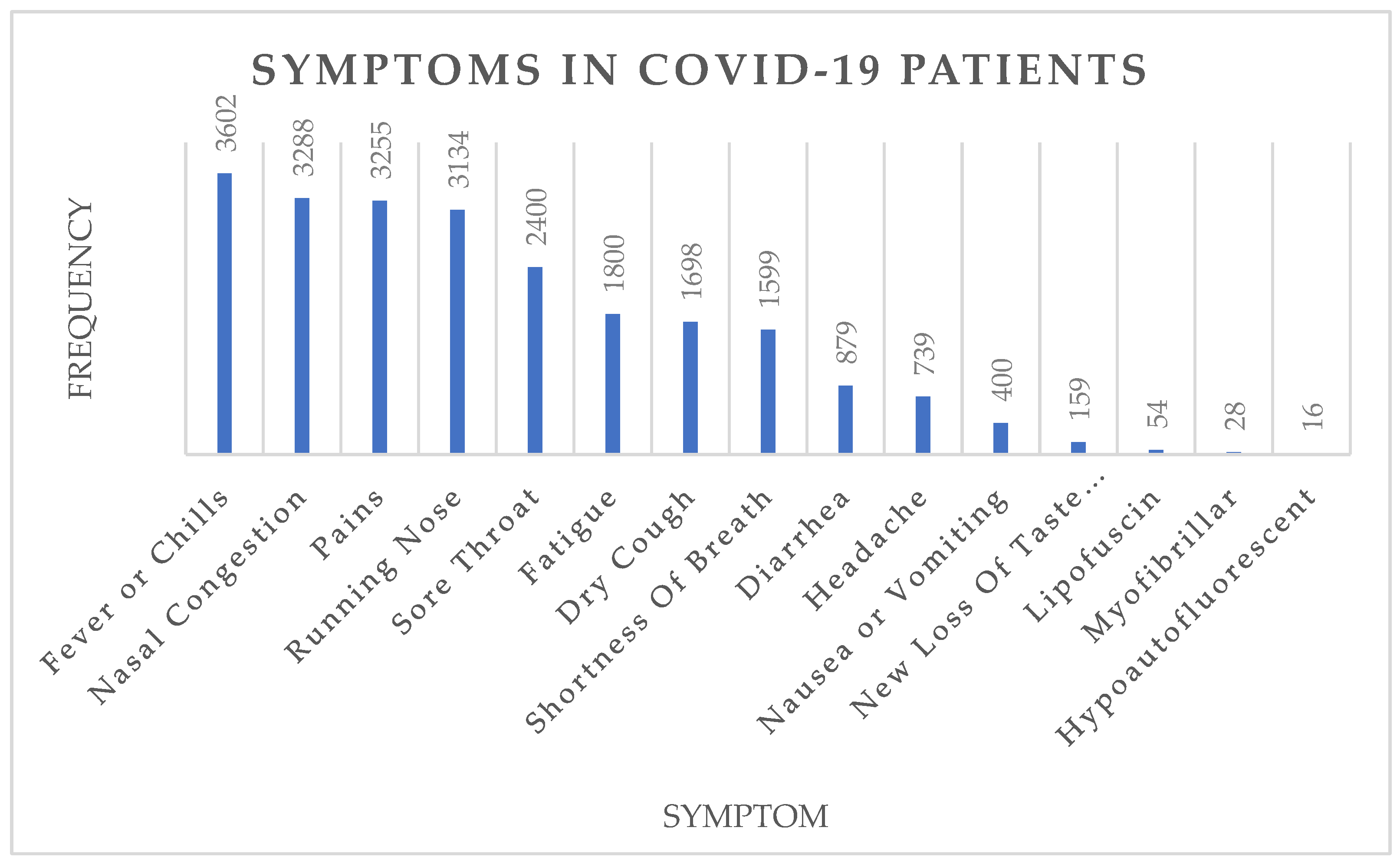
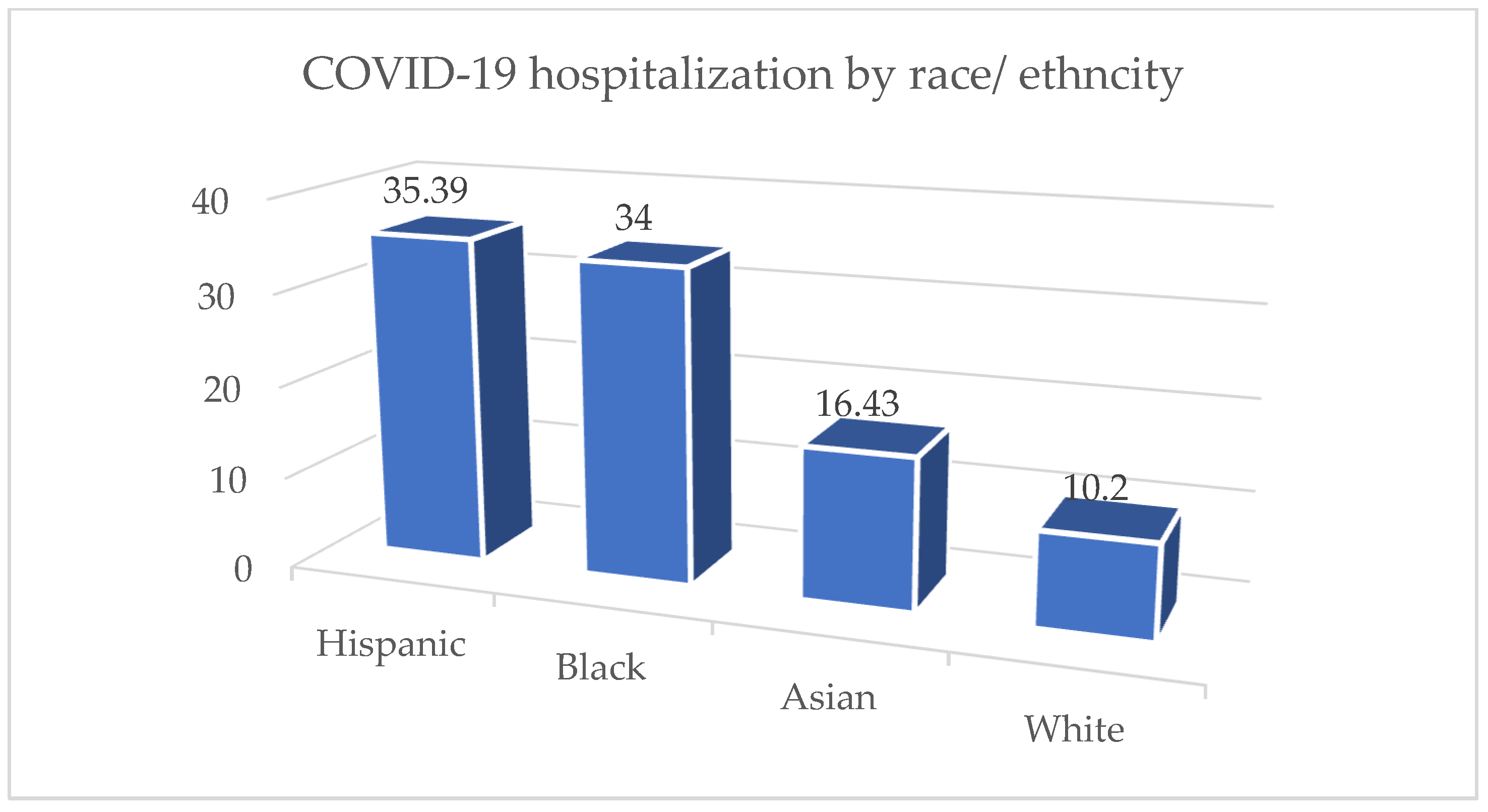
4.2. Pandemic Surveillance
5. Discussion
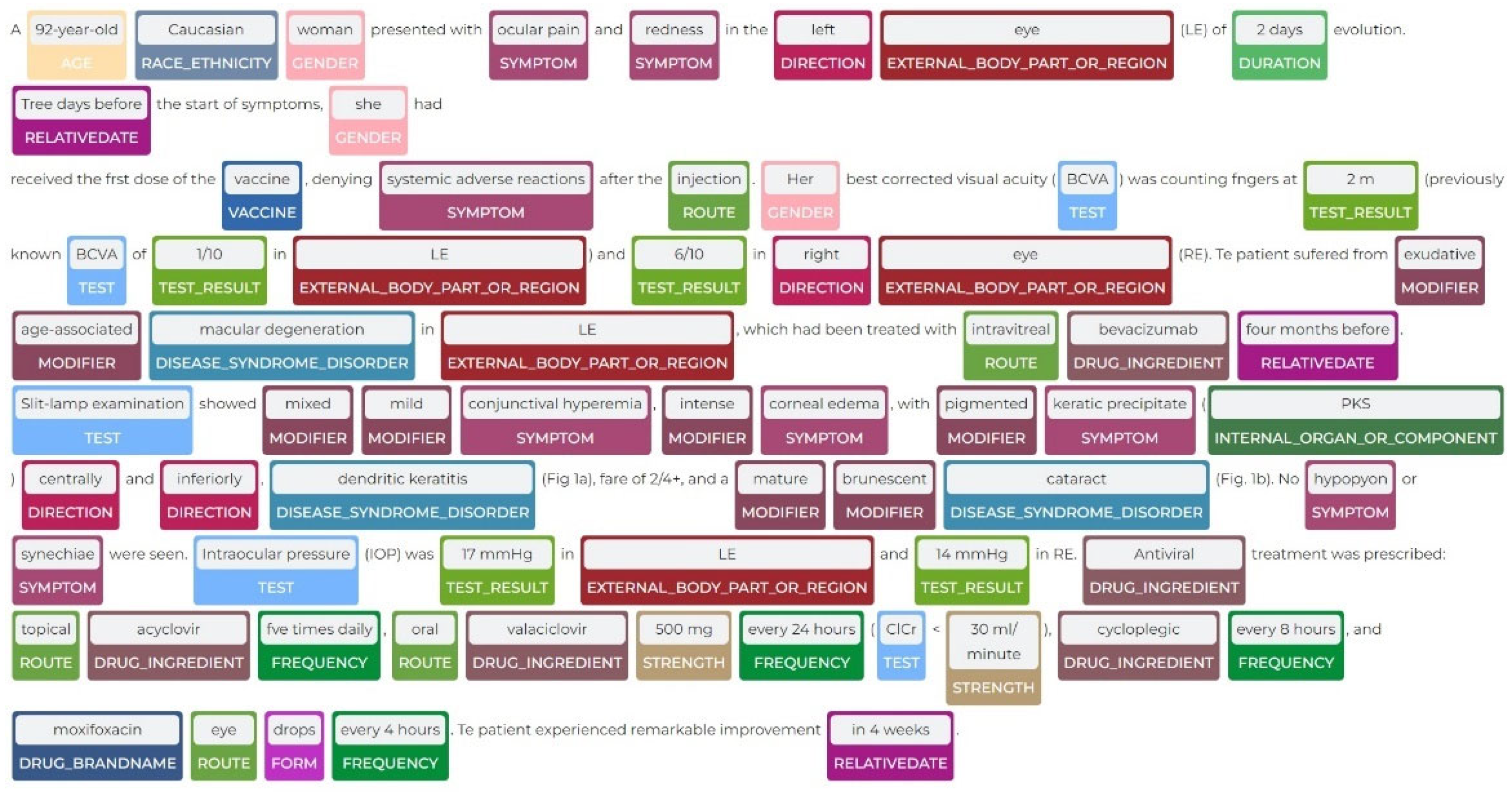
5.1. Implications in Healthcare
5.2. Transfer Learning
5.3. Limitations
6. Conclusions
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Chen, Q.; Leaman, R.; Allot, A.; Luo, L.; Wei, C.-H.; Yan, S.; Lu, Z. Artificial Intelligence in Action: Addressing the COVID-19 Pandemic with Natural Language Processing. Annu. Rev. Biomed. Data Sci. 2021, 4, 313–339. [Google Scholar] [CrossRef] [PubMed]
- Raza, S.; Schwartz, B.; Rosella, L.C. CoQUAD: A COVID-19 Question Answering Dataset System, Facilitating Research, Benchmarking, and Practice. BMC Bioinform. 2022, 23, 210. [Google Scholar] [CrossRef] [PubMed]
- Allen Institute. COVID-19 Open Research Dataset Challenge (CORD-19). 2020. Available online: https://www.kaggle.com/datasets/allen-institute-for-ai/CORD-19-research-challenge (accessed on 27 November 2022).
- Chen, Q.; Allot, A.; Lu, Z. LitCovid: An Open Database of COVID-19 Literature. Nucleic Acids Res. 2021, 49, D1534–D1540. [Google Scholar] [CrossRef] [PubMed]
- Wang, L.L.; Lo, K. Text Mining Approaches for Dealing with the Rapidly Expanding Literature on COVID-19. Brief. Bioinform. 2021, 22, 781–799. [Google Scholar] [CrossRef]
- Reeves, R.M.; Christensen, L.; Brown, J.R.; Conway, M.; Levis, M.; Gobbel, G.T.; Shah, R.U.; Goodrich, C.; Rickets, I.; Minter, F.; et al. Adaptation of an NLP System to a New Healthcare Environment to Identify Social Determinants of Health. J. Biomed. Inform. 2021, 120, 103851. [Google Scholar] [CrossRef]
- Nadeau, D.; Sekine, S. A Survey of Named Entity Recognition and Classification. Lingvisticae Investig. 2007, 30, 3–26. [Google Scholar] [CrossRef]
- Boudjellal, N.; Zhang, H.; Khan, A.; Ahmad, A.; Naseem, R.; Shang, J.; Dai, L. ABioNER: A BERT-Based Model for Arabic Biomedical Named-Entity Recognition. Complexity 2021, 2021, 6633213. [Google Scholar] [CrossRef]
- Dmis-Lab. DMIS-Lab/Biobert: Bioinformatics’2020: BioBERT: A Pre-Trained Biomedical Language Representation Model for Biomedical Text Mining. GitHub. 2020. Available online: https://github.com/dmis-lab/biobert (accessed on 27 November 2022).
- Perera, N.; Dehmer, M.; Emmert-Streib, F. Named Entity Recognition and Relation Detection for Biomedical Information Extraction. Front. Cell Dev. Biol. 2020, 8, 673. [Google Scholar] [CrossRef]
- Cho, H.; Lee, H. Biomedical Named Entity Recognition Using Deep Neural Networks with Contextual Information. BMC Bioinform. 2019, 20, 735. [Google Scholar] [CrossRef]
- Raza, S.; Schwartz, B. Detecting Biomedical Named Entities in COVID-19 Texts. In Proceedings of the 1st Workshop on Healthcare AI and COVID-19, ICML 2022, Baltimore, MA, USA, 17–23 July 2022. [Google Scholar]
- Nosowsky, R.; Giordano, T.J. The Health Insurance Portability and Accountability Act of 1996 (HIPAA) Privacy Rule: Implications for Clinical Research. Annu. Rev. Med. 2006, 57, 575–590. [Google Scholar] [CrossRef]
- Sang, E.F.; De Meulder, F. Introduction to the CoNLL-2003 shared task: Language-independent named entity recognition. arXiv 2003, arXiv:cs/0306050. [Google Scholar]
- Doğan, R.I.; Leaman, R.; Lu, Z. NCBI Disease Corpus: A Resource for Disease Name Recognition and Concept Normalization. J. Biomed. Inform. 2014, 47, 1–10. [Google Scholar] [CrossRef] [PubMed]
- Li, J.; Sun, Y.; Johnson, R.J.; Sciaky, D.; Wei, C.-H.; Leaman, R.; Davis, A.P.; Mattingly, C.J.; Wiegers, T.C.; Lu, Z. BioCreative V CDR Task Corpus: A Resource for Chemical Disease Relation Extraction. Database 2016, 2016, baw068. [Google Scholar] [CrossRef] [PubMed]
- Krallinger, M.; Rabal, O.; Leitner, F.; Vazquez, M.; Salgado, D.; Lu, Z.; Leaman, R.; Lu, Y.; Ji, D.; Lowe, D.M.; et al. The CHEMDNER Corpus of Chemicals and Drugs and Its Annotation Principles. J. Cheminform. 2015, 7, S2. [Google Scholar] [CrossRef] [PubMed]
- Smith, L.; Tanabe, L.K.; Kuo, C.-J.; Chung, I.; Hsu, C.-N.; Lin, Y.-S.; Klinger, R.; Friedrich, C.M.; Ganchev, K.; Torii, M.; et al. Overview of BioCreative II Gene Mention Recognition. Genome Biol. 2008, 9, S2. [Google Scholar] [CrossRef]
- Collier, N.; Kim, J.-D. Introduction to the Bio-Entity Recognition Task at JNLPBA. In Proceedings of the International Joint Workshop on Natural Language Processing in Biomedicine and its Applications (NLPBA/BioNLP), Geneva, Switzerland, 28–29 August 2004; pp. 73–78. [Google Scholar]
- Uzuner, Ö.; South, B.R.; Shen, S.; DuVall, S.L. 2010 I2b2/VA Challenge on Concepts, Assertions, and Relations in Clinical Text. J. Am. Med. Inform. Assoc. 2011, 18, 552–556. [Google Scholar] [CrossRef]
- Sun, W.; Rumshisky, A.; Uzuner, O. Evaluating Temporal Relations in Clinical Text: 2012 I2b2 Challenge. J. Am. Med. Inform. Assoc. 2013, 20, 806–813. [Google Scholar] [CrossRef]
- Lample, G.; Ballesteros, M.; Subramanian, S.; Kawakami, K.; Dyer, C. Neural Architectures for Named Entity Recognition. arXiv 2016, arXiv:1603.01360. [Google Scholar]
- Chiu, J.P.C.; Nichols, E. Named Entity Recognition with Bidirectional LSTM-CNNs. Trans. Assoc. Comput. Linguist. 2016, 4, 357–370. [Google Scholar] [CrossRef]
- Wang, X.; Zhang, Y.; Ren, X.; Zhang, Y.; Zitnik, M.; Shang, J.; Langlotz, C.; Han, J. Cross-Type Biomedical Named Entity Recognition with Deep Multi-Task Learning. Bioinformatics 2019, 35, 1745–1752. [Google Scholar] [CrossRef]
- Luo, L.; Yang, Z.; Yang, P.; Zhang, Y.; Wang, L.; Lin, H.; Wang, J. An Attention-Based BiLSTM-CRF Approach to Document-Level Chemical Named Entity Recognition. Bioinformatics 2018, 34, 1381–1388. [Google Scholar] [CrossRef] [PubMed]
- Xu, K.; Yang, Z.; Kang, P.; Wang, Q.; Liu, W. Document-Level Attention-Based BiLSTM-CRF Incorporating Disease Dictionary for Disease Named Entity Recognition. Comput. Biol. Med. 2019, 108, 122–132. [Google Scholar] [CrossRef] [PubMed]
- Zhao, Z.; Yang, Z.; Luo, L.; Wang, L.; Zhang, Y.; Lin, H.; Wang, J. Disease Named Entity Recognition from Biomedical Literature Using a Novel Convolutional Neural Network. BMC Med. Genom. 2017, 10, 75–83. [Google Scholar] [CrossRef] [PubMed]
- Yoon, W.; So, C.H.; Lee, J.; Kang, J. Collabonet: Collaboration of Deep Neural Networks for Biomedical Named Entity Recognition. BMC Bioinform. 2019, 20, 55–65. [Google Scholar] [CrossRef] [PubMed]
- Beltagy, I.; Lo, K.; Cohan, A. SCIBERT: A Pretrained Language Model for Scientific Text. In Proceedings of the 2019 Conference on Empirical Methods in Natural Language Processing and the 9th International Joint Conference on Natural Language Processing (EMNLP-IJCNLP), Hong Kong, China, 3–7 November 2020; pp. 3615–3620. [Google Scholar]
- Lee, J.; Yoon, W.; Kim, S.; Kim, D.; Kim, S.; So, C.H.; Kang, J. BioBERT: A Pre-Trained Biomedical Language Representation Model for Biomedical Text Mining. Bioinformatics 2020, 36, 1234–1240. [Google Scholar] [CrossRef]
- Office of Disease Prevention and Health Promotion Social Determinants of Health. Healthy People 2030. Available online: https://health.gov/healthypeople/objectives-and-data/social-determinants-health (accessed on 7 October 2021).
- Toscano, F.; O’Donnell, E.; Unruh, M.A.; Golinelli, D.; Carullo, G.; Messina, G.; Casalino, L.P. Electronic Health Records Implementation: Can the European Union Learn from the United States? Eur. J. Public Health 2018, 28, cky213.401. [Google Scholar] [CrossRef]
- Fernández-Calienes, R. Health Insurance Portability and Accountability Act of 1996. In Encyclopedia of the Fourth Amendment; CQ Press: Washington, DC, USA, 2013. [Google Scholar]
- Meystre, S.M.; Friedlin, F.J.; South, B.R.; Shen, S.; Samore, M.H. Automatic De-Identification of Textual Documents in the Electronic Health Record: A Review of Recent Research. BMC Med. Res. Methodol. 2010, 10, 70. [Google Scholar] [CrossRef]
- Lafferty, J.; Mccallum, A.; Pereira, F. Conditional Random Fields: Probabilistic Models for Segmenting and Labeling Sequence Data Abstract. In Proceedings of the Eighteenth International Conference on Machine Learning (ICML-2001), Williamstown, MA, USA, June 28–1 July 2001; pp. 282–289. [Google Scholar]
- Tsochantaridis, I.; Joachims, T.; Hofmann, T.; Altun, Y.; Singer, Y. Large Margin Methods for Structured and Interdependent Output Variables. J. Mach. Learn. Res. 2005, 6, 1453–1484. [Google Scholar]
- Yang, X.; Lyu, T.; Li, Q.; Lee, C.Y.; Bian, J.; Hogan, W.R.; Wu, Y. A Study of Deep Learning Methods for De-Identification of Clinical Notes in Cross-Institute Settings. BMC Med. Inform. Decis. Mak. 2019, 19, 232. [Google Scholar] [CrossRef]
- John Snow Labs. Spark OCR. Available online: https://nlp.johnsnowlabs.com/docs/en/ocr (accessed on 27 November 2022).
- Annotation Lab; John Snow Labs: Lewes, DE, USA, 2022.
- Ogren, P.V.; Savova, G.K.; Chute, C.G. Constructing Evaluation Corpora for Automated Clinical Named Entity Recognition. In Proceedings of the Sixth International Conference on Language Resources and Evaluation (LREC’08), Marrakech, Morocco, 26 May–1 June 2008; pp. 3143–3150. [Google Scholar]
- Chen, Y.; Lasko, T.A.; Mei, Q.; Denny, J.C.; Xu, H. A Study of Active Learning Methods for Named Entity Recognition in Clinical Text. J. Biomed. Inform. 2015, 58, 11–18. [Google Scholar] [CrossRef]
- Snow, R.; O’Connor, B.; Jurafsky, D.; Ng, A.Y. Cheap and Fast—But Is It Good? Evaluating Non-Expert Annotations for Natural Language Tasks. In Proceedings of the 2008 Conference on Empirical Methods in Natural Language Processing, Honolulu, HI, USA, 25–27 October 2008; pp. 254–263. [Google Scholar]
- ML Pipelines—Documentation. Available online: https://spark.apache.org/docs/latest/ml-pipeline.html (accessed on 27 November 2022).
- Kocaman, V.; Talby, D. Spark NLP: Natural Language Understanding at Scale. Softw. Impacts 2021, 8, 100058. [Google Scholar] [CrossRef]
- Webster, J.J.; Kit, C. Tokenization as the Initial Phase in NLP. In Proceedings of the COLING 1992. The 14th International Conference on Computational Linguistics, Nantes, France, 23–28 August 1992; Volume 4. [Google Scholar]
- Peng, Y.; Yan, S.; Lu, Z. Transfer Learning in Biomedical Natural Language Processing: An Evaluation of BERT and ELMo on Ten Benchmarking Datasets. arXiv 2019, arXiv:1906.05474. [Google Scholar]
- Johnson, A.E.W.; Pollard, T.J.; Shen, L.; Lehman, L.W.H.; Feng, M.; Ghassemi, M.; Moody, B.; Szolovits, P.; Anthony Celi, L.; Mark, R.G. MIMIC-III, a Freely Accessible Critical Care Database. Sci. Data 2016, 3, 160035. [Google Scholar] [CrossRef] [PubMed]
- Huang, Z.; Xu, W.; Yu, K. Bidirectional LSTM-CRF Models for Sequence Tagging. arXiv 2015, arXiv:1508.01991. [Google Scholar]
- Ma, X.; Hovy, E. End-to-End Sequence Labeling via Bi-Directional Lstm-Cnns-Crf. arXiv 2016, arXiv:1603.01354. [Google Scholar]
- Bakken, D.E.; Rarameswaran, R.; Blough, D.M.; Franz, A.A.; Palmer, T.J. Data Obfuscation: Anonymity and Desensitization of Usable Data Sets. IEEE Secur. Priv. 2004, 2, 34–41. [Google Scholar] [CrossRef]
- Medical Data De-Identification—John Snow Labs; John Snow Labs: Lewes, DE, USA, 2022.
- GitHub. ay94 NER-Datasets. 2022. Available online: https://github.com/ay94/NER-datasets (accessed on 27 November 2022).
- Sexton, T. IOB Format Intro; Nestor: Gaithersburg, MA, USA, 2022. [Google Scholar]
- Akbik, A.; Blythe, D.; Vollgraf, R. Contextual String Embeddings for Sequence Labeling. In Proceedings of the 27th international Conference on Computational Linguistics, Santa Fe, NM, USA, 20–26 August 2018; pp. 1638–1649. [Google Scholar]
- Müller, M.; Salathé, M.; Kummervold, P.E. Covid-Twitter-Bert: A Natural Language Processing Model to Analyse COVID-19 Content on Twitter. arXiv 2020, arXiv:2005.07503. [Google Scholar]
- Tsai, R.T.-H.; Wu, S.-H.; Chou, W.-C.; Lin, Y.-C.; He, D.; Hsiang, J.; Sung, T.-Y.; Hsu, W.-L. Various Criteria in the Evaluation of Biomedical Named Entity Recognition. BMC Bioinform. 2006, 7, 92. [Google Scholar] [CrossRef]
- Perone, C.S.; Silveira, R.; Paula, T.S. Evaluation of Sentence Embeddings in Downstream and Linguistic Probing Tasks. arXiv 2018, arXiv:1806.06259. [Google Scholar]
- Abdi, S.; Bennett-AbuAyyash, C.; MacDonald, L.; Hohenadel, K.; Johnson, K.O.; Leece, P. Provincial Implementation Supports for Socio-Demographic Data Collection during COVID-19 in Ontario’s Public Health System. Can. J. Public Health 2021, 112, 853–861. [Google Scholar] [CrossRef]
- Ortiz-Egea, J.M.; Sánchez, C.G.; López-Jiménez, A.; Navarro, O.D. Herpetic Anterior Uveitis Following Pfizer–BioNTech Coronavirus Disease 2019 Vaccine: Two Case Reports. J. Med. Case Rep. 2022, 16, 127. [Google Scholar] [CrossRef] [PubMed]
- Raza, S. A Machine Learning Model for Predicting, Diagnosing, and Mitigating Health Disparities in Hospital Readmission. Healthc. Anal. 2022, 2, 100100. [Google Scholar] [CrossRef]

| Benchmark Datasets | ||
|---|---|---|
| Corpus | Entity Types | Data Size |
| NCBI-Disease [15] | Diseases | 793 PubMed abstracts |
| BC5CDR [16] | Diseases | 1500 PubMed articles |
| BC5CDR [16] | Chemicals | 1500 PubMed articles |
| BC4CHEMD [17] | Chemicals | 10,000 PubMed abstracts |
| BC2GM [18] | Gene/Proteins | 20,000 sentences |
| JNLPBA [19] | Genes, proteins | 2404 abstracts |
| i2b2-Clinical [20] | Problem, Treatment, and Test. | 426 discharge summaries |
| I2b2 2012 [21] | Clinical (problems, tests, treatments, clinical departments, occurrences (admission, discharge) and evidence). | 310 discharge summaries |
| Benchmark methods | ||
| Method | Description | |
| BiLSTM-CRF [22] | Bidirectional Long short-term memory (LSTM) and Conditional random field (CRF) architecture for NER. | |
| BiLSTM-CNN-Char [23] | A hybrid LSTM and Convolutional Neural Network (CNN) architecture that learns both character-level and word-level features for the NER task. | |
| BiLSTM-CRF-MTL [24] | A multi-task learning (MTL) framework with a BiLSTM-CRF model to collectively use the training data of different types of entities. | |
| Att-BiLSTM-CRF [25], | Attention (Att) based BiLSTM model with a CRF layer for chemical NER task. | |
| Doc-Att-BiLSTM-CRF [26] | Document (Doc)-level Attention (Att)-based BiLSTM-CRF network for disease NER task. | |
| MCNN [27] | A multiple (M) label CNN-based network for disease NER from biomedical literature. | |
| CollaboNet [28] | A collaboration of deep neural networks, i.e., BiLSTM-CRF with a single task model trained for each specific entity type. | |
| SciBERT [29] | A pre-trained language model based on Bidirectional Encoder Representations from Transformers (BERT) pretrained on a large multi-domain corpus of scientific publications to improve performance on downstream scientific tasks including NER. | |
| BioBERT [30] | A pre-trained biomedical language representation model based on BERT for biomedical text mining | |
| Entity Type | Entities |
|---|---|
| Clinical name entities | Admission (patient admission status), oncology (tumor/cancer), blood pressure, respiration (e.g., shortness of breath), dosage (amount of medicine/drug taken), vital signs, symptoms, kidney disease, temperature (body), diabetes, vaccine, time (days, weeks or so), obesity (status), BMI, height (of patient), heart disease, pulse, hypertension, drug name, cerebrovascular disease, disease, treatment, clinical department, weight (of patient), admission/discharge (from hospital), modifier (modifies the current state), external body part, test, strength, route, test result. |
| Non-clinical entities | Name (of patient), location, date, relative date, duration, relationship status, social status, family history (family members, alone, with family, homeless), employment status, race/ethnicity, gender, social history, sexual orientation, diet (food type, nutrients, minerals), alcohol, smoking. |
| Hyperparameter | Optimal Value (Values Used) |
|---|---|
| Learning rate | 1 × 10−3 (1 × 10−2, 1 × 10−3, 1 × 10−5, 2 × 10−5, 5 × 10−5, 3 × 10−4) |
| Batch size | 64 (8, 16, 32, 64, 128) |
| Epochs | 30 ({2, 3, …, 30}) |
| LSTM state size | 200 (200, 250) |
| Dropout rate | 0.5 ({0.3, 0.35, …, 0.7}) |
| Optimizer | Adam |
| CNN filters | 2 (2, 3, 4, 5) |
| Hidden Size | 768 |
| Embedding Size | 128 |
| Max Seq Length | 512 |
| Warmup Steps | 3000 |
| Methods/ Dataset | Metric | NCBI | BC5CDR | BC2GM | JNLPBA | i2b2-Clinical | Our Dataset |
|---|---|---|---|---|---|---|---|
| BiLSTM-CRF | micro | 85.80 | 84.22 | 78.46 | 74.29 | 83.66 | 87.10 |
| macro | 86.12 | 85.09 | 80.01 | 75.10 | 84.01 | 88.01 | |
| BiLSTM-CRF-MTL | micro | 86.46 | 84.94 | 80.34 | 77.03 | 82.38 | 88.39 |
| macro | 88.01 | 85.00 | 81.12 | 77.14 | 83.96 | 88.97 | |
| CT-BERT | micro | 77.50 | 76.85 | 74.10 | 68.00 | 77.07 | 78.10 |
| macro | 78.50 | 77.96 | 75.37 | 68.98 | 78.01 | 78.98 | |
| SciBERT | micro | 82.88 | 82.94 | 84.08 | 75.77 | 78.19 | 80.95 |
| macro | 83.32 | 83.13 | 85.84 | 77.01 | 79.10 | 81.14 | |
| BioBERT-Base v1.0 | micro | 84.01 | 86.56 | 78.68 | 86.28 | 85.87 | 84.01 |
| macro | 79.10 | 78.90 | 79.00 | 78.13 | 72.18 | 79.10 | |
| BioBERT-Base v1.1 | micro | 88.52 | 87.15 | 79.39 | 76.16 | 86.27 | 88.52 |
| macro | 85.89 | 87.10 | 87.18 | 75.45 | 87.78 | 85.89 | |
| BioBERT-Base v1.2 | micro | 89.12 | 87.81 | 83.34 | 76.45 | 86.88 | 89.12 |
| macro | 86.78 | 87.89 | 86.07 | 75.15 | 86.98 | 86.78 | |
| Our approach | micro | 90.58 | 89.90 | 89.15 | 79.92 | 89.10 | 94.78 |
| macro | 91.83 | 90.34 | 90.38 | 80.94 | 90.48 | 95.37 |
| Model | Macro |
|---|---|
| BiLSTM-CNN-CRF | 94.18 ± 0.12 |
| BiLSTM-CNN | 87.37 ± 0.02 |
| Map-CNN-CRF | 80.55 ± 0.03 |
| Map-CNN | 69.25 ± 0.04 |
| Entity | TP | FP | FN | Prec | Recall | F1 |
|---|---|---|---|---|---|---|
| Disease | 818 | 98 | 112 | 0.89 | 0.88 | 0.89 |
| Gender | 390 | 78 | 101 | 0.83 | 0.79 | 0.81 |
| Employment | 234 | 29 | 132 | 0.89 | 0.64 | 0.74 |
| Race_Ethnicity | 334 | 65 | 96 | 0.84 | 0.78 | 0.81 |
| Smoking | 309 | 24 | 97 | 0.93 | 0.76 | 0.84 |
| Psychological_Condition | 218 | 29 | 58 | 0.88 | 0.79 | 0.83 |
| Death_Entity | 387 | 34 | 103 | 0.92 | 0.79 | 0.85 |
| BMI | 146 | 12 | 29 | 0.92 | 0.83 | 0.88 |
| Diabetes | 157 | 10 | 28 | 0.94 | 0.85 | 0.89 |
| Macro-average | 2993 | 379 | 756 | 0.89 | 0.79 | 0.84 |
| Micro-average | 2993 | 379 | 756 | 0.89 | 0.80 | 0.84 |
| Drugs | Vaccine | Non-Medical Treatments |
|---|---|---|
| Hydroxychloroquine | Pfizer-BioNTech | Isolation |
| Paxlovid | Moderna | Wear masks |
| Actemra | AstraZeneca | Vaccination |
| Immunomodulators | CoronaVac | Oxygen support |
| Steroid | BBIBP-CorV | Medication |
| Amoxicillin | Janssen | Hand sanitization |
| Sentence | Begin | End | Chunks | Biomedical Entity | Confidence |
|---|---|---|---|---|---|
| 0 | 2 | 12 | 73-year-old | Age | 1.00 |
| 0 | 14 | 18 | woman | Gender | 1.00 |
| 0 | 32 | 43 | Fever Clinic | Clinical Department | 0.98 |
| 0 | 52 | 65 | First Hospital | Clinical Department | 0.51 |
| 0 | 109 | 134 | Fever, temperature | Symptom | 0.80 |
| 0 | 156 | 160 | Cough | Symptom | 0.99 |
| 0 | 163 | 175 | Expectoration | Symptom | 1.00 |
| 0 | 178 | 196 | Shortness of breath | Symptom | 0.39 |
| 0 | 203 | 218 | General weakness | Symptom | 0.77 |
| 0 | 233 | 244 | Prior 5 days | Relative Date | 0.42 |
| 1 | 247 | 249 | She | Gender | 1.00 |
| 1 | 261 | 264 | Mild | Modifier | 0.90 |
| 1 | 266 | 273 | Diarrhea | Symptom | 1.00 |
| 1 | 280 | 289 | Stools/day | Symptom | 0.85 |
| 1 | 292 | 303 | 2 days prior | Relative Date | 0.68 |
| 1 | 322 | 329 | Hospital | Clinical Department | 1.00 |
| 1 | 386 | 402 | COVID-19 positive | Disease Syndrome | 0.90 |
| 1 | 436 | 454 | Healthcare provider | Employment | 0.94 |
| 2 | 486 | 494 | Cirrhosis | Disease Syndrome | 0.96 |
| 2 | 500 | 514 | Type 2 diabetes | Diabetes | 0.95 |
| 2 | 535 | 541 | Smoking | Smoking | 1.00 |
| 2 | 546 | 553 | Drinking | Alcohol | 0.93 |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2022 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Bashir, S.R.; Raza, S.; Kocaman, V.; Qamar, U. Clinical Application of Detecting COVID-19 Risks: A Natural Language Processing Approach. Viruses 2022, 14, 2761. https://doi.org/10.3390/v14122761
Bashir SR, Raza S, Kocaman V, Qamar U. Clinical Application of Detecting COVID-19 Risks: A Natural Language Processing Approach. Viruses. 2022; 14(12):2761. https://doi.org/10.3390/v14122761
Chicago/Turabian StyleBashir, Syed Raza, Shaina Raza, Veysel Kocaman, and Urooj Qamar. 2022. "Clinical Application of Detecting COVID-19 Risks: A Natural Language Processing Approach" Viruses 14, no. 12: 2761. https://doi.org/10.3390/v14122761
APA StyleBashir, S. R., Raza, S., Kocaman, V., & Qamar, U. (2022). Clinical Application of Detecting COVID-19 Risks: A Natural Language Processing Approach. Viruses, 14(12), 2761. https://doi.org/10.3390/v14122761






