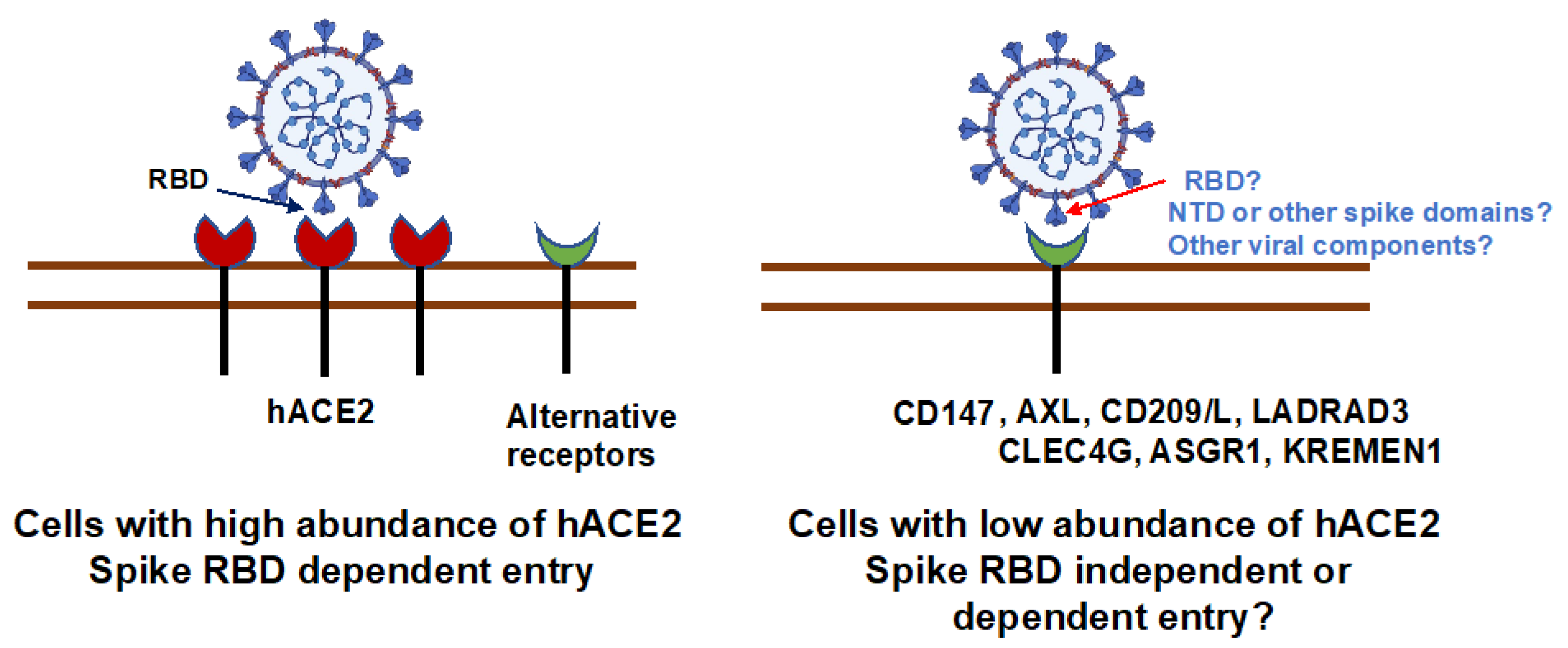
ACE2-Independent Alternative Receptors for SARS-CoV-2
Abstract
1. Introduction
2. CD147
| Receptor | Viruses | Reference |
|---|---|---|
| CD147 | HIV, HBV, HCV, measles virus, CMV, KSHV, and SARS-CoV | [55,56,57,58,59,60,61,62,63,64] |
| AXL | Dengue, zika, Ebola, Lassa, Marburg virus, Hantaan virus, and Andes virus | [65,66,67,68,69,70,71] |
| CD209L and CD209 | Sindbis virus, Ebola virus, Japanese encephalitis virus, HIV, HCV, influenza A virus, SARS-CoV | [71,72,73,74,75,76,77,78,79] |
| CLEC4G/LSECtin | Ebloa, Lassa, Marburg virus, filovirus Japanese encephalitis virus, SARS-CoV | [71,74,80,81,82,83] |
| KREMEN1 | Coxsackievirus A10 and other human type A enteroviruses | [84,85,86] |
| ASGR1/CLEC4H1 | Hepatitis E virus | [87] |
| LDLRAD3 | Venezuelan equine encephalitis virus | [88] |
| TMEM30A/CD50A | Lujo virus | [89] |
3. AXL
4. C-Type Lectins
5. LDLRAD3 and TMEM30A
6. KREMEN1 and ASGR1
7. Conclusions and Perspective
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Acknowledgments
Conflicts of Interest
References
- Pak, A.; Adegboye, O.A.; Adekunle, A.I.; Rahman, K.M.; McBryde, E.S.; Eisen, D.P. Economic Consequences of the COVID-19 Outbreak: The Need for Epidemic Preparedness. Front. Public Health 2020, 8, 241. [Google Scholar] [CrossRef] [PubMed]
- Rahman, S.; Rahman, M.M.; Miah, M.; Begum, M.N.; Sarmin, M.; Mahfuz, M.; Hossain, M.E.; Rahman, M.Z.; Chisti, M.J.; Ahmed, T.; et al. COVID-19 reinfections among naturally infected and vaccinated individuals. Sci. Rep. 2022, 12, 1438. [Google Scholar] [CrossRef] [PubMed]
- Jeffery-Smith, A.; Rowland, T.A.J.; Patel, M.; Whitaker, H.; Iyanger, N.; Williams, S.V.; Giddings, R.; Thompson, L.; Zavala, M.; Aiano, F.; et al. Reinfection with new variants of SARS-CoV-2 after natural infection: A prospective observational cohort in, 13 care homes in England. Lancet Healthy Longev. 2021, 2, e811–e819. [Google Scholar] [CrossRef]
- Nguyen, N.N.; Houhamdi, L.; Hoang, V.T.; Delerce, J.; Delorme, L.; Colson, P.; Brouqui, P.; Fournier, P.E.; Raoult, D.; Gautret, P. SARS-CoV-2 reinfection and COVID-19 severity. Emerg. Microbes Infect 2022, 11, 894–901. [Google Scholar] [CrossRef] [PubMed]
- Jackson, C.B.; Farzan, M.; Chen, B.; Choe, H. Mechanisms of SARS-CoV-2 entry into cells. Nat. Rev. Mol. Cell Biol. 2022, 23, 3–20. [Google Scholar] [CrossRef]
- Hansen, J.; Baum, A.; Pascal, K.E.; Russo, V.; Giordano, S.; Wloga, E.; Fulton, B.O.; Yan, Y.; Koon, K.; Patel, K.; et al. Studies in humanized mice and convalescent humans yield a SARS-CoV-2 antibody cocktail. Science 2020, 369, 1010–1014. [Google Scholar] [CrossRef]
- Hoffmann, M.; Kleine-Weber, H.; Pöhlmann, S. A Multibasic Cleavage Site in the Spike Protein of SARS-CoV-2 Is Essential for Infection of Human Lung Cells. Mol. Cell 2020, 78, 779–784.e775. [Google Scholar] [CrossRef]
- Bestle, D.; Heindl, M.R.; Limburg, H.; Van Lam van, T.; Pilgram, O.; Moulton, H.; Stein, D.A.; Hardes, K.; Eickmann, M.; Dolnik, O.; et al. TMPRSS2 and furin are both essential for proteolytic activation of SARS-CoV-2 in human airway cells. Life Sci. Alliance 2020, 3, e202000786. [Google Scholar] [CrossRef]
- Hoffmann, M.; Kleine-Weber, H.; Schroeder, S.; Kruger, N.; Herrler, T.; Erichsen, S.; Schiergens, T.S.; Herrler, G.; Wu, N.H.; Nitsche, A.; et al. SARS-CoV-2 Cell Entry Depends on ACE2 and TMPRSS2 and Is Blocked by a Clinically Proven Protease Inhibitor. Cell 2020, 181, 271–280. [Google Scholar] [CrossRef]
- Zang, R.; Gomez Castro, M.F.; McCune, B.T.; Zeng, Q.; Rothlauf, P.W.; Sonnek, N.M.; Liu, Z.; Brulois, K.F.; Wang, X.; Greenberg, H.B.; et al. TMPRSS2 and TMPRSS4 promote SARS-CoV-2 infection of human small intestinal enterocytes. Sci. Immunol. 2020, 5, eabc3582. [Google Scholar] [CrossRef]
- Kim, Y.; Jang, G.; Lee, D.; Kim, N.; Seon, J.W.; Kim, Y.-H.; Lee, C. Trypsin enhances SARS-CoV-2 infection by facilitating viral entry. Arch. Virol. 2022, 167, 441–458. [Google Scholar] [CrossRef]
- Bayati, A.; Kumar, R.; Francis, V.; McPherson, P.S. SARS-CoV-2 infects cells after viral entry via clathrin-mediated endocytosis. J. Biol. Chem. 2021, 296, 100306. [Google Scholar] [CrossRef]
- Inoue, Y.; Tanaka, N.; Tanaka, Y.; Inoue, S.; Morita, K.; Zhuang, M.; Hattori, T.; Sugamura, K. Clathrin-dependent entry of severe acute respiratory syndrome coronavirus into target cells expressing ACE2 with the cytoplasmic tail deleted. J. Virol. 2007, 81, 8722–8729. [Google Scholar] [CrossRef]
- Daly, J.L.; Simonetti, B.; Klein, K.; Chen, K.-E.; Williamson, M.K.; Antón-Plágaro, C.; Shoemark, D.K.; Simón-Gracia, L.; Bauer, M.; Hollandi, R.; et al. Neuropilin-1 is a host factor for SARS-CoV-2 infection. Science 2020, 370, 861–865. [Google Scholar] [CrossRef]
- Cantuti-Castelvetri, L.; Ojha, R.; Pedro, L.D.; Djannatian, M.; Franz, J.; Kuivanen, S.; van der Meer, F.; Kallio, K.; Kaya, T.; Anastasina, M.; et al. Neuropilin-1 facilitates SARS-CoV-2 cell entry and infectivity. Science 2020, 370, 856–860. [Google Scholar] [CrossRef]
- Nguyen, L.; McCord, K.A.; Bui, D.T.; Bouwman, K.M.; Kitova, E.N.; Elaish, M.; Kumawat, D.; Daskhan, G.C.; Tomris, I.; Han, L.; et al. Sialic acid-containing glycolipids mediate binding and viral entry of SARS-CoV-2. Nat. Chem. Biol. 2022, 18, 81–90. [Google Scholar] [CrossRef]
- Arrindell, J.; Abou Atmeh, P.; Jayet, L.; Sereme, Y.; Mege, J.L.; Desnues, B. Vimentin is an important ACE2 co-receptor for SARS-CoV-2 in epithelial cells. iScience 2022, 25, 105463. [Google Scholar] [CrossRef]
- Amraei, R.; Xia, C.; Olejnik, J.; White, M.R.; Napoleon, M.A.; Lotfollahzadeh, S.; Hauser, B.M.; Schmidt, A.G.; Chitalia, V.; Muhlberger, E.; et al. Extracellular vimentin is an attachment factor that facilitates SARS-CoV-2 entry into human endothelial cells. Proc. Natl. Acad. Sci. USA 2022, 119, e2113874119. [Google Scholar] [CrossRef]
- Clausen, T.M.; Sandoval, D.R.; Spliid, C.B.; Pihl, J.; Perrett, H.R.; Painter, C.D.; Narayanan, A.; Majowicz, S.A.; Kwong, E.M.; McVicar, R.N.; et al. SARS-CoV-2 Infection Depends on Cellular Heparan Sulfate and ACE2. Cell 2020, 183, 1043–1057.e1015. [Google Scholar] [CrossRef]
- Bohan, D.; Van Ert, H.; Ruggio, N.; Rogers, K.J.; Badreddine, M.; Aguilar Briseño, J.A.; Elliff, J.M.; Rojas Chavez, R.A.; Gao, B.; Stokowy, T.; et al. Phosphatidylserine receptors enhance SARS-CoV-2 infection. PLoS Pathog. 2021, 17, e1009743. [Google Scholar] [CrossRef]
- Maemura, T.; Kuroda, M.; Armbrust, T.; Yamayoshi, S.; Halfmann, P.J.; Kawaoka, Y. Antibody-Dependent Enhancement of SARS-CoV-2 Infection Is Mediated by the IgG Receptors FcγRIIA and FcγRIIIA but Does Not Contribute to Aberrant Cytokine Production by Macrophages. mBio 2021, 12, e0198721. [Google Scholar] [CrossRef] [PubMed]
- Ibrahim, I.M.; Abdelmalek, D.H.; Elshahat, M.E.; Elfiky, A.A. COVID-19 spike-host cell receptor GRP78 binding site prediction. J. Infect. 2020, 80, 554–562. [Google Scholar] [CrossRef] [PubMed]
- Shahriari Felordi, M.; Memarnejadian, A.; Najimi, M.; Vosough, M. Is There any Alternative Receptor for SARS-CoV-2? Cell J. 2021, 23, 247–250. [Google Scholar] [PubMed]
- Cui, C.; Huang, C.; Zhou, W.; Ji, X.; Zhang, F.; Wang, L.; Zhou, Y.; Cui, Q. AGTR2, One Possible Novel Key Gene for the Entry of SARS-CoV-2 Into Human Cells. IEEE/ACM Trans. Comput. Biol. Bioinform. 2021, 18, 1230–1233. [Google Scholar] [CrossRef] [PubMed]
- Masre, S.F.; Jufri, N.F.; Ibrahim, F.W.; Abdul Raub, S.H. Classical and alternative receptors for SARS-CoV-2 therapeutic strategy. Rev. Med. Virol. 2021, 31, e2207. [Google Scholar] [CrossRef]
- Sun, G.; Cui, Q.; Garcia, G., Jr.; Wang, C.; Zhang, M.; Arumugaswami, V.; Riggs, A.D.; Shi, Y. Comparative transcriptomic analysis of SARS-CoV-2 infected cell model systems reveals differential innate immune responses. Sci. Rep. 2021, 11, 17146. [Google Scholar] [CrossRef]
- Blanco-Melo, D.; Nilsson-Payant, B.E.; Liu, W.C.; Uhl, S.; Hoagland, D.; Møller, R.; Jordan, T.X.; Oishi, K.; Panis, M.; Sachs, D.; et al. Imbalanced Host Response to SARS-CoV-2 Drives Development of COVID-19. Cell 2020, 181, 1036–1045.e1039. [Google Scholar] [CrossRef]
- Hikmet, F.; Méar, L.; Edvinsson, Å.; Micke, P.; Uhlén, M.; Lindskog, C. The protein expression profile of ACE2 in human tissues. Mol. Syst. Biol. 2020, 16, e9610. [Google Scholar] [CrossRef]
- Singh, M.; Bansal, V.; Feschotte, C. A Single-Cell RNA Expression Map of Human Coronavirus Entry Factors. Cell Rep 2020, 32, 108175. [Google Scholar] [CrossRef]
- Xu, C.; Wang, A.; Geng, K.; Honnen, W.; Wang, X.; Bruiners, N.; Singh, S.; Ferrara, F.; D’Angelo, S.; Bradbury, A.R.M.; et al. Human Immunodeficiency Viruses Pseudotyped with SARS-CoV-2 Spike Proteins Infect a Broad Spectrum of Human Cell Lines through Multiple Entry Mechanisms. Viruses 2021, 13, 953. [Google Scholar] [CrossRef]
- Hoffmann, M.; Sidarovich, A.; Arora, P.; Krüger, N.; Nehlmeier, I.; Kempf, A.; Graichen, L.; Winkler, M.S.; Niemeyer, D.; Goffinet, C.; et al. Evidence for an ACE2-Independent Entry Pathway That Can Protect from Neutralization by an Antibody Used for COVID-19 Therapy. mBio 2022, 13, e0036422. [Google Scholar] [CrossRef]
- Beyerstedt, S.; Casaro, E.B.; Rangel É, B. COVID-19: Angiotensin-converting enzyme, 2 (ACE2) expression and tissue susceptibility to SARS-CoV-2 infection. Eur. J. Clin. Microbiol. Infect Dis. 2021, 40, 905–919. [Google Scholar] [CrossRef]
- Meng, B.; Abdullahi, A.; Ferreira, I.; Goonawardane, N.; Saito, A.; Kimura, I.; Yamasoba, D.; Gerber, P.P.; Fatihi, S.; Rathore, S.; et al. Altered TMPRSS2 usage by SARS-CoV-2 Omicron impacts infectivity and fusogenicity. Nature 2022, 603, 706–714. [Google Scholar] [CrossRef]
- Willett, B.J.; Grove, J.; MacLean, O.A.; Wilkie, C.; De Lorenzo, G.; Furnon, W.; Cantoni, D.; Scott, S.; Logan, N.; Ashraf, S.; et al. SARS-CoV-2 Omicron is an immune escape variant with an altered cell entry pathway. Nat. Microbiol. 2022, 7, 1161–1179. [Google Scholar] [CrossRef]
- Pia, L.; Rowland-Jones, S. Omicron entry route. Nat. Rev. Immunol. 2022, 22, 144. [Google Scholar] [CrossRef]
- Peacock, T.P.; Brown, J.C.; Zhou, J.; Thakur, N.; Sukhova, K.; Newman, J.; Kugathasan, R.; Yan, A.W.C.; Furnon, W.; De Lorenzo, G.; et al. The altered entry pathway and antigenic distance of the SARS-CoV-2 Omicron variant map to separate domains of spike protein. bioRxiv 2022, 15, e0241955. [Google Scholar]
- Shilts, J.; Crozier, T.W.M.; Greenwood, E.J.D.; Lehner, P.J.; Wright, G.J. No evidence for basigin/CD147 as a direct SARS-CoV-2 spike binding receptor. Sci. Rep. 2021, 11, 413. [Google Scholar] [CrossRef]
- Ragotte, R.J.; Pulido, D.; Donnellan, F.R.; Hill, M.L.; Gorini, G.; Davies, H.; Brun, J.; McHugh, K.; King, L.D.W.; Skinner, K.; et al. Human Basigin (CD147) Does Not Directly Interact with SARS-CoV-2 Spike Glycoprotein. mSphere 2021, 6, e00647-21. [Google Scholar] [CrossRef]
- Wang, K.; Chen, W.; Zhang, Z.; Deng, Y.; Lian, J.-Q.; Du, P.; Wei, D.; Zhang, Y.; Sun, X.-X.; Gong, L.; et al. CD147-spike protein is a novel route for SARS-CoV-2 infection to host cells. Signal Transduct. Target. Ther. 2020, 5, 283. [Google Scholar] [CrossRef]
- Wang, S.; Qiu, Z.; Hou, Y.; Deng, X.; Xu, W.; Zheng, T.; Wu, P.; Xie, S.; Bian, W.; Zhang, C.; et al. AXL is a candidate receptor for SARS-CoV-2 that promotes infection of pulmonary and bronchial epithelial cells. Cell Res. 2021, 31, 126–140. [Google Scholar] [CrossRef]
- Amraei, R.; Yin, W.; Napoleon, M.A.; Suder, E.L.; Berrigan, J.; Zhao, Q.; Olejnik, J.; Chandler, K.B.; Xia, C.; Feldman, J.; et al. CD209L/L-SIGN and CD209/DC-SIGN Act as Receptors for SARS-CoV-2. ACS Cent. Sci. 2021, 7, 1156–1165. [Google Scholar] [CrossRef] [PubMed]
- Gu, Y.; Cao, J.; Zhang, X.; Gao, H.; Wang, Y.; Wang, J.; He, J.; Jiang, X.; Zhang, J.; Shen, G.; et al. Receptome profiling identifies KREMEN1 and ASGR1 as alternative functional receptors of SARS-CoV-2. Cell Res. 2022, 32, 24–37. [Google Scholar] [CrossRef]
- Hoffmann, D.; Mereiter, S.; Jin Oh, Y.; Monteil, V.; Elder, E.; Zhu, R.; Canena, D.; Hain, L.; Laurent, E.; Grünwald-Gruber, C.; et al. Identification of lectin receptors for conserved SARS-CoV-2 glycosylation sites. EMBO J. 2021, 40, e108375. [Google Scholar] [CrossRef] [PubMed]
- Zhu, S.; Liu, Y.; Zhou, Z.; Zhang, Z.; Xiao, X.; Liu, Z.; Chen, A.; Dong, X.; Tian, F.; Chen, S.; et al. Genome-wide CRISPR activation screen identifies candidate receptors for SARS-CoV-2 entry. Sci. China Life Sci. 2022, 65, 701–717. [Google Scholar] [CrossRef] [PubMed]
- Grass, G.D.; Toole, B.P. How, with whom and when: An overview of CD147-mediated regulatory networks influencing matrix metalloproteinase activity. Biosci. Rep. 2015, 36, e00283. [Google Scholar] [CrossRef]
- Available online: https://www.proteinatlas.org/ENSG00000172270-BSG/tissue (accessed on 15 August 2022).
- Landras, A.; Reger de Moura, C.; Jouenne, F.; Lebbe, C.; Menashi, S.; Mourah, S. CD147 Is a Promising Target of Tumor Progression and a Prognostic Biomarker. Cancers 2019, 11, 1803. [Google Scholar] [CrossRef]
- Xin, X.; Zeng, X.; Gu, H.; Li, M.; Tan, H.; Jin, Z.; Hua, T.; Shi, R.; Wang, H. CD147/EMMPRIN overexpression and prognosis in cancer: A systematic review and meta-analysis. Sci. Rep. 2016, 6, 32804. [Google Scholar] [CrossRef]
- Wang, S.-J.; Chao, D.; Wei, W.; Nan, G.; Li, J.-Y.; Liu, F.-L.; Li, L.; Jiang, J.-L.; Cui, H.-Y.; Chen, Z.-N. CD147 promotes collective invasion through cathepsin B in hepatocellular carcinoma. J. Exp. Clin. Cancer Res. 2020, 39, 145. [Google Scholar] [CrossRef]
- Yang, H.; Chen, B. CD147 in Ovarian and Other Cancers. Int. J. Gynecol. Cancer 2013, 23, 2–8. [Google Scholar] [CrossRef]
- Li, F.; Zhang, J.; Guo, J.; Jia, Y.; Han, Y.; Wang, Z. RNA interference targeting CD147 inhibits metastasis and invasion of human breast cancer MCF-7 cells by downregulating MMP-9/VEGF expression. Acta Biochim. Et Biophys. Sin. 2018, 50, 676–684. [Google Scholar] [CrossRef]
- Iacono, K.T.; Brown, A.L.; Greene, M.I.; Saouaf, S.J. CD147 immunoglobulin superfamily receptor function and role in pathology. Exp. Mol. Pathol. 2007, 83, 283–295. [Google Scholar] [CrossRef]
- Xiong, L.; Edwards, C.K., 3rd; Zhou, L. The biological function and clinical utilization of CD147 in human diseases: A review of the current scientific literature. Int. J. Mol. Sci. 2014, 15, 17411–17441. [Google Scholar] [CrossRef]
- Ali, M.M.; Mirza, I.; Naquiallah, D.; Hassan, C.; Masrur, M.; Bianco, F.M.; Mahmoud, A.M. CD147 Levels in Blood and Adipose Tissues Correlate with Vascular Dysfunction in Obese Diabetic Adults. J. Cardiovasc. Dev. Dis. 2021, 9, 7. [Google Scholar] [CrossRef]
- Watanabe, A.; Yoneda, M.; Ikeda, F.; Terao-Muto, Y.; Sato, H.; Kai, C. CD147/EMMPRIN Acts as a Functional Entry Receptor for Measles Virus on Epithelial Cells. J. Virol. 2010, 84, 4183–4193. [Google Scholar] [CrossRef]
- Muramatsu, T. Basigin (CD147), a multifunctional transmembrane glycoprotein with various binding partners. J. Biochem. 2016, 159, 481–490. [Google Scholar] [CrossRef]
- Pushkarsky, T.; Yurchenko, V.; Laborico, A.; Bukrinsky, M. CD147 stimulates HIV-1 infection in a signal-independent fashion. Biochem. Biophys. Res. Commun. 2007, 363, 495–499. [Google Scholar] [CrossRef]
- Chen, Z.; Mi, L.; Xu, J.; Yu, J.; Wang, X.; Jiang, J.; Xing, J.; Shang, P.; Qian, A.; Li, Y.; et al. Function of HAb18G/CD147 in invasion of host cells by severe acute respiratory syndrome coronavirus. J. Infect. Dis. 2005, 191, 755–760. [Google Scholar] [CrossRef]
- Dai, L.; Bai, L.; Lu, Y.; Xu, Z.; Reiss, K.; Del Valle, L.; Kaleeba, J.; Toole, B.P.; Parsons, C.; Qin, Z. Emmprin and KSHV: New partners in viral cancer pathogenesis. Cancer Lett. 2013, 337, 161–166. [Google Scholar] [CrossRef]
- Tian, X.; Zhao, C.; Zhu, H.; She, W.; Zhang, J.; Liu, J.; Li, L.; Zheng, S.; Wen, Y.M.; Xie, Y. Hepatitis B virus (HBV) surface antigen interacts with and promotes cyclophilin a secretion: Possible link to pathogenesis of HBV infection. J. Virol. 2010, 84, 3373–3381. [Google Scholar] [CrossRef]
- Feng, X.; Xiu, B.; Xu, L.; Yang, X.; He, J.; Leong, D.; He, F.; Zhang, H. Hepatitis C virus core protein promotes the migration and invasion of hepatocyte via activating transcription of extracellular matrix metalloproteinase inducer. Virus Res. 2011, 158, 146–153. [Google Scholar] [CrossRef]
- Qin, Z.; Dai, L.; Slomiany, M.G.; Toole, B.P.; Parsons, C. Direct activation of emmprin and associated pathogenesis by an oncogenic herpesvirus. Cancer Res. 2010, 70, 3884–3889. [Google Scholar] [CrossRef] [PubMed]
- Pushkarsky, T.; Zybarth, G.; Dubrovsky, L.; Yurchenko, V.; Tang, H.; Guo, H.; Toole, B.; Sherry, B.; Bukrinsky, M. CD147 facilitates HIV-1 infection by interacting with virus-associated cyclophilin A. Proc. Natl. Acad. Sci. USA 2001, 98, 6360–6365. [Google Scholar] [CrossRef] [PubMed]
- Vanarsdall, A.L.; Pritchard, S.R.; Wisner, T.W.; Liu, J.; Jardetzky, T.S.; Johnson, D.C. CD147 Promotes Entry of Pentamer-Expressing Human Cytomegalovirus into Epithelial and Endothelial Cells. mBio 2018, 9, e00781-18. [Google Scholar] [CrossRef]
- Xie, S.; Zhang, H.; Liang, Z.; Yang, X.; Cao, R. AXL, an Important Host Factor for DENV and ZIKV Replication. Front. Cell Infect Microbiol. 2021, 11, 575346. [Google Scholar] [CrossRef] [PubMed]
- Richard, A.S.; Shim, B.S.; Kwon, Y.C.; Zhang, R.; Otsuka, Y.; Schmitt, K.; Berri, F.; Diamond, M.S.; Choe, H. AXL-dependent infection of human fetal endothelial cells distinguishes Zika virus from other pathogenic flaviviruses. Proc. Natl. Acad. Sci. USA 2017, 114, 2024–2029. [Google Scholar] [CrossRef] [PubMed]
- Strange, D.P.; Jiyarom, B.; Zarandi, N.P.; Xie, X.; Baker, C.; Sadri-Ardekani, H.; Shi, P.-Y.; Verma, S. Axl Promotes Zika Virus Entry and Modulates the Antiviral State of Human Sertoli Cells. mBio 2019, 10, e01372-19. [Google Scholar] [CrossRef] [PubMed]
- Zwernik, S.D.; Adams, B.H.; Raymond, D.A.; Warner, C.M.; Kassam, A.B.; Rovin, R.A.; Akhtar, P. AXL receptor is required for Zika virus strain MR-766 infection in human glioblastoma cell lines. Mol. Ther. Oncolytics 2021, 23, 447–457. [Google Scholar] [CrossRef]
- Liu, S.; DeLalio, L.J.; Isakson, B.E.; Wang, T.T. AXL-Mediated Productive Infection of Human Endothelial Cells by Zika Virus. Circ. Res. 2016, 119, 1183–1189. [Google Scholar] [CrossRef]
- Shimojima, M.; Takada, A.; Ebihara, H.; Neumann, G.; Fujioka, K.; Irimura, T.; Jones, S.; Feldmann, H.; Kawaoka, Y. Tyro3 family-mediated cell entry of Ebola and Marburg viruses. J. Virol. 2006, 80, 10109–10116. [Google Scholar] [CrossRef]
- Shimojima, M.; Stroher, U.; Ebihara, H.; Feldmann, H.; Kawaoka, Y. Identification of cell surface molecules involved in dystroglycan-independent Lassa virus cell entry. J. Virol. 2012, 86, 2067–2078. [Google Scholar] [CrossRef]
- Klimstra, W.B.; Nangle, E.M.; Smith, M.S.; Yurochko, A.D.; Ryman, K.D. DC-SIGN and L-SIGN can act as attachment receptors for alphaviruses and distinguish between mosquito cell- and mammalian cell-derived viruses. J. Virol. 2003, 77, 12022–12032. [Google Scholar] [CrossRef]
- Khoo, U.-S.; Chan, K.Y.K.; Chan, V.S.F.; Lin, C.L.S. DC-SIGN and L-SIGN: The SIGNs for infection. J. Mol. Med. 2008, 86, 861–874. [Google Scholar] [CrossRef]
- Shimojima, M.; Takenouchi, A.; Shimoda, H.; Kimura, N.; Maeda, K. Distinct usage of three C-type lectins by Japanese encephalitis virus: DC-SIGN, DC-SIGNR, and LSECtin. Arch. Virol. 2014, 159, 2023–2031. [Google Scholar] [CrossRef]
- Fayssel, N.; Bensghir, R.; Ouladlahsen, A.; Abdelghaffar, H.; Sodqi, M.; Lahlou, K.; Benjelloun, S.; Marhoum El Filali, K.; Ezzikouri, S.; Wakrim, L. Association of CD209L tandem repeats polymorphism with susceptibility to human immunodeficiency virus-1 infection, disease progression, and treatment outcomes: A Moroccan cohort study. Clin. Microbiol. Infect. 2015, 21, 513.e1–513.e5. [Google Scholar] [CrossRef]
- Gardner, J.P.; Durso, R.J.; Arrigale, R.R.; Donovan, G.P.; Maddon, P.J.; Dragic, T.; Olson, W.C. L-SIGN (CD, 209L) is a liver-specific capture receptor for hepatitis C virus. Proc. Natl. Acad. Sci. USA 2003, 100, 4498–4503. [Google Scholar] [CrossRef]
- Alvarez, C.P.; Lasala, F.; Carrillo, J.; Muniz, O.; Corbi, A.L.; Delgado, R. C-type lectins DC-SIGN and L-SIGN mediate cellular entry by Ebola virus in cis and in trans. J. Virol. 2002, 76, 6841–6844. [Google Scholar] [CrossRef]
- Ludwig, I.S.; Lekkerkerker, A.N.; Depla, E.; Bosman, F.; Musters, R.J.; Depraetere, S.; van Kooyk, Y.; Geijtenbeek, T.B. Hepatitis C virus targets DC-SIGN and L-SIGN to escape lysosomal degradation. J. Virol. 2004, 78, 8322–8332. [Google Scholar] [CrossRef] [PubMed]
- Londrigan, S.L.; Turville, S.G.; Tate, M.D.; Deng, Y.M.; Brooks, A.G.; Reading, P.C. N-linked glycosylation facilitates sialic acid-independent attachment and entry of influenza A viruses into cells expressing DC-SIGN or L-SIGN. J. Virol. 2011, 85, 2990–3000. [Google Scholar] [CrossRef]
- Gramberg, T.; Hofmann, H.; Möller, P.; Lalor, P.F.; Marzi, A.; Geier, M.; Krumbiegel, M.; Winkler, T.; Kirchhoff, F.; Adams, D.H.; et al. LSECtin interacts with filovirus glycoproteins and the spike protein of SARS coronavirus. Virology 2005, 340, 224–236. [Google Scholar] [CrossRef]
- Pipirou, Z.; Powlesland, A.S.; Steffen, I.; Pohlmann, S.; Taylor, M.E.; Drickamer, K. Mouse LSECtin as a model for a human Ebola virus receptor. Glycobiology 2011, 21, 806–812. [Google Scholar] [CrossRef] [PubMed]
- Powlesland, A.S.; Fisch, T.; Taylor, M.E.; Smith, D.F.; Tissot, B.; Dell, A.; Pohlmann, S.; Drickamer, K. A novel mechanism for LSECtin binding to Ebola virus surface glycoprotein through truncated glycans. J. Biol. Chem. 2008, 283, 593–602. [Google Scholar] [CrossRef] [PubMed]
- Zhao, D.; Han, X.; Zheng, X.; Wang, H.; Yang, Z.; Liu, D.; Han, K.; Liu, J.; Wang, X.; Yang, W.; et al. The Myeloid LSECtin Is a DAP12-Coupled Receptor That Is Crucial for Inflammatory Response Induced by Ebola Virus Glycoprotein. PLoS Pathog. 2016, 12, e1005487. [Google Scholar]
- Staring, J.; van den Hengel, L.G.; Raaben, M.; Blomen, V.A.; Carette, J.E.; Brummelkamp, T.R. KREMEN1 Is a Host Entry Receptor for a Major Group of Enteroviruses. Cell Host Microbe 2018, 23, 636–643.e5. [Google Scholar] [CrossRef] [PubMed]
- Cui, Y.; Peng, R.; Song, H.; Tong, Z.; Qu, X.; Liu, S.; Zhao, X.; Chai, Y.; Wang, P.; Gao, G.F.; et al. Molecular basis of Coxsackievirus A10 entry using the two-in-one attachment and uncoating receptor KRM1. Proc. Natl. Acad. Sci. USA 2020, 117, 18711–18718. [Google Scholar] [CrossRef]
- Zhao, Y.; Zhou, D.; Ni, T.; Karia, D.; Kotecha, A.; Wang, X.; Rao, Z.; Jones, E.Y.; Fry, E.E.; Ren, J.; et al. Hand-foot-and-mouth disease virus receptor KREMEN1 binds the canyon of Coxsackie Virus A10. Nat. Commun. 2020, 11, 38. [Google Scholar] [CrossRef]
- Zhang, L.; Tian, Y.; Wen, Z.; Zhang, F.; Qi, Y.; Huang, W.; Zhang, H.; Wang, Y. Asialoglycoprotein receptor facilitates infection of PLC/PRF/5 cells by HEV through interaction with ORF2. J. Med Virol. 2016, 88, 2186–2195. [Google Scholar] [CrossRef]
- Ma, H.; Kim, A.S.; Kafai, N.M.; Earnest, J.T.; Shah, A.P.; Case, J.B.; Basore, K.; Gilliland, T.C.; Sun, C.; Nelson, C.A.; et al. LDLRAD3 is a receptor for Venezuelan equine encephalitis virus. Nature 2020, 588, 308–314. [Google Scholar] [CrossRef]
- Raaben, M.; Jae, L.T.; Herbert, A.S.; Kuehne, A.I.; Stubbs, S.H.; Chou, Y.Y.; Blomen, V.A.; Kirchhausen, T.; Dye, J.M.; Brummelkamp, T.R.; et al. NRP2 and CD63 Are Host Factors for Lujo Virus Cell Entry. Cell Host Microbe 2017, 22, 688–696.e5. [Google Scholar] [CrossRef]
- Zhang, M.Y.; Zhang, Y.; Wu, X.D.; Zhang, K.; Lin, P.; Bian, H.J.; Qin, M.M.; Huang, W.; Wei, D.; Zhang, Z.; et al. Disrupting CD147-RAP2 interaction abrogates erythrocyte invasion by Plasmodium falciparum. Blood 2018, 131, 1111–1121. [Google Scholar] [CrossRef]
- Badeti, S.; Jiang, Q.; Naghizadeh, A.; Tseng, H.-C.; Bushkin, Y.; Marras, S.A.E.; Nisa, A.; Tyagi, S.; Chen, F.; Romanienko, P.; et al. Development of a novel human CD147 knock-in NSG mouse model to test SARS-CoV-2 viral infection. Cell Biosci. 2022, 12, 88. [Google Scholar] [CrossRef]
- Williamson, C.D.; Donaldson, J.G. Arf6, JIP3, and dynein shape and mediate macropinocytosis. Mol. Biol. Cell 2019, 30, 1477–1489. [Google Scholar] [CrossRef]
- Mercer, J.; Helenius, A. Virus entry by macropinocytosis. Nat. Cell Biol. 2009, 11, 510–520. [Google Scholar] [CrossRef]
- Janssen, J.W.; Schulz, A.S.; Steenvoorden, A.C.; Schmidberger, M.; Strehl, S.; Ambros, P.F.; Bartram, C.R. A novel putative tyrosine kinase receptor with oncogenic potential. Oncogene 1991, 6, 2113–2120. [Google Scholar]
- Wium, M.; Ajayi-Smith, A.F.; Paccez, J.D.; Zerbini, L.F. The Role of the Receptor Tyrosine Kinase Axl in Carcinogenesis and Development of Therapeutic Resistance: An Overview of Molecular Mechanisms and Future Applications. Cancers 2021, 13, 1521. [Google Scholar] [CrossRef]
- Lemke, G. Biology of the TAM receptors. Cold Spring Harb. Perspect. Biol. 2013, 5, a009076. [Google Scholar] [CrossRef]
- Gay, C.M.; Balaji, K.; Byers, L.A. Giving AXL the axe: Targeting AXL in human malignancy. Br. J. Cancer 2017, 116, 415–423. [Google Scholar] [CrossRef]
- Brenig, R.; Pop, O.T.; Triantafyllou, E.; Geng, A.; Singanayagam, A.; Perez-Shibayama, C.; Besse, L.; Cupovic, J.; Künzler, P.; Boldanova, T.; et al. Expression of AXL receptor tyrosine kinase relates to monocyte dysfunction and severity of cirrhosis. Life Sci. Alliance 2019, 3, e201900465. [Google Scholar] [CrossRef]
- Neubauer, A.; Fiebeler, A.; Graham, D.K.; O’Bryan, J.P.; Schmidt, C.A.; Barckow, P.; Serke, S.; Siegert, W.; Snodgrass, H.R.; Huhn, D.; et al. Expression of axl, a transforming receptor tyrosine kinase, in normal and malignant hematopoiesis. Blood 1994, 84, 1931–1941. [Google Scholar] [CrossRef]
- Available online: https://www.proteinatlas.org/ENSG00000167601-AXL/tissue (accessed on 15 August 2022).
- Agrelli, A.; de Moura, R.R.; Crovella, S.; Brandão, L.A.C. ZIKA virus entry mechanisms in human cells. Infect. Genet. Evol. 2019, 69, 22–29. [Google Scholar] [CrossRef]
- Zhang, L.; Richard, A.S.; Jackson, C.B.; Ojha, A.; Choe, H. Phosphatidylethanolamine and Phosphatidylserine Synergize To Enhance GAS6/AXL-Mediated Virus Infection and Efferocytosis. J. Virol. 2020, 95, e02079-20. [Google Scholar] [CrossRef]
- Den Dunnen, J.; Gringhuis, S.I.; Geijtenbeek, T.B.H. Innate signaling by the C-type lectin DC-SIGN dictates immune responses. Cancer Immunol. Immunother. 2009, 58, 1149–1157. [Google Scholar] [CrossRef] [PubMed]
- Rahimi, N. C-type Lectin CD209L/L-SIGN and CD209/DC-SIGN: Cell Adhesion Molecules Turned to Pathogen Recognition Receptors. Biology 2020, 10, 1. [Google Scholar] [CrossRef] [PubMed]
- Zhang, F.; Ren, S.; Zuo, Y. DC-SIGN, DC-SIGNR and LSECtin: C-Type Lectins for Infection. Int. Rev. Immunol. 2014, 33, 54–66. [Google Scholar] [CrossRef] [PubMed]
- Dominguez-Soto, A.; Aragoneses-Fenoll, L.; Martin-Gayo, E.; Martinez-Prats, L.; Colmenares, M.; Naranjo-Gomez, M.; Borras, F.E.; Munoz, P.; Zubiaur, M.; Toribio, M.L.; et al. The DC-SIGN–related lectin LSECtin mediates antigen capture and pathogen binding by human myeloid cells. Blood 2007, 109, 5337–5345. [Google Scholar] [CrossRef]
- Tang, L.; Yang, J.; Liu, W.; Tang, X.; Chen, J.; Zhao, D.; Wang, M.; Xu, F.; Lu, Y.; Liu, B.; et al. Liver sinusoidal endothelial cell lectin, LSECtin, negatively regulates hepatic T-cell immune response. Gastroenterology 2009, 137, 1498–1508.e5. [Google Scholar] [CrossRef]
- Tang, L.; Yang, J.; Tang, X.; Ying, W.; Qian, X.; He, F. The DC-SIGN family member LSECtin is a novel ligand of CD44 on activated T cells. Eur. J. Immunol. 2010, 40, 1185–1191. [Google Scholar] [CrossRef]
- Available online: https://www.proteinatlas.org/ENSG00000179241-LDLRAD3 (accessed on 20 August 2022).
- Noyes, N.C.; Hampton, B.; Migliorini, M.; Strickland, D.K. Regulation of Itch and Nedd4 E3 Ligase Activity and Degradation by LRAD3. Biochemistry 2016, 55, 1204–1213. [Google Scholar] [CrossRef]
- Available online: https://www.proteinatlas.org/ENSG00000112697-TMEM30A/tissue (accessed on 25 August 2022).
- Lopez-Marques, R.L.; Theorin, L.; Palmgren, M.G.; Pomorski, T.G. P4-ATPases: Lipid flippases in cell membranes. Pflügers Arch.-Eur. J. Physiol. 2014, 466, 1227–1240. [Google Scholar] [CrossRef]
- Van der Mark, V.A.; Elferink, R.P.; Paulusma, C.C. P4 ATPases: Flippases in health and disease. Int J Mol Sci 2013, 14, 7897–7922. [Google Scholar] [CrossRef]
- Paulusma, C.C.; Folmer, D.E.; Ho-Mok, K.S.; de Waart, D.R.; Hilarius, P.M.; Verhoeven, A.J.; Oude Elferink, R.P. ATP8B1 requires an accessory protein for endoplasmic reticulum exit and plasma membrane lipid flippase activity. Hepatology 2008, 47, 268–278. [Google Scholar] [CrossRef]
- Li, N.; Yang, Y.; Liang, C.; Qiu, Q.; Pan, C.; Li, M.; Yang, S.; Chen, L.; Zhu, X.; Hu, Y. Tmem30a Plays Critical Roles in Ensuring the Survival of Hematopoietic Cells and Leukemia Cells in Mice. Am. J. Pathol. 2018, 188, 1457–1468. [Google Scholar] [CrossRef]
- Ennishi, D.; Healy, S.; Bashashati, A.; Saberi, S.; Hother, C.; Mottok, A.; Chan, F.C.; Chong, L.; Abraham, L.; Kridel, R.; et al. TMEM30A loss-of-function mutations drive lymphomagenesis and confer therapeutically exploitable vulnerability in B-cell lymphoma. Nat. Med. 2020, 26, 577–588. [Google Scholar] [CrossRef]
- Rothbächer, U.; Lemaire, P. Crème de la Kremen of Wnt signalling inhibition. Nat. Cell Biol. 2002, 4, E172. [Google Scholar] [CrossRef]
- Osada, M.; Ito, E.; Fermin, H.A.; Vazquez-Cintron, E.; Venkatesh, T.; Friedel, R.H.; Pezzano, M. The Wnt signaling antagonist Kremen1 is required for development of thymic architecture. Clin. Dev. Immunol. 2006, 13, 299–319. [Google Scholar] [CrossRef]
- Causeret, F.; Sumia, I.; Pierani, A. Kremen1 and Dickkopf1 control cell survival in a Wnt-independent manner. Cell Death Differ. 2016, 23, 323–332. [Google Scholar] [CrossRef]
- Hoober, J.K. ASGR1 and Its Enigmatic Relative, CLEC10A. Int. J. Mol. Sci. 2020, 21, 4818. [Google Scholar] [CrossRef]
- Liu, Z.; VanBlargan, L.A.; Bloyet, L.-M.; Rothlauf, P.W.; Chen, R.E.; Stumpf, S.; Zhao, H.; Errico, J.M.; Theel, E.S.; Liebeskind, M.J.; et al. Identification of SARS-CoV-2 spike mutations that attenuate monoclonal and serum antibody neutralization. Cell Host Microbe 2021, 29, 477–488.e4. [Google Scholar] [CrossRef]
| Receptor | Viral Components | Reference |
|---|---|---|
| CD147 | NA, anti-RBD mAb resistant | [30] |
| No full-length spike or RBD binding | [37,38] | |
| RBD | [39] | |
| AXL | PS in virions (ACE2 dependent) | [20] |
| NTD | [40] | |
| CD209L/L-SIGN/CLEC4M | RBD (N-glycans) | [41] |
| NTD (high affinity), RBD, S2 | [42] | |
| CD209/DCSIGN/CLEC4L | spike trimer | [43] |
| CLEC4G/LSECtin | RBD (N-glycans) | [43] |
| NTD | [44] | |
| KREMEN1 | RBD (high affinity), NTD | [42] |
| ASGR1/CLEC4H1 | RBD (high affinity), NTD | [42] |
| LDLRAD3 | NTD | [44] |
| TMEM30A/CD50A | NTD | [44] |
| Clec4g (mouse) | RBD (N-glycans) | [43] |
| CD209c (mouse) | RBD (N-glycans) | [43] |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2022 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Lim, S.; Zhang, M.; Chang, T.L. ACE2-Independent Alternative Receptors for SARS-CoV-2. Viruses 2022, 14, 2535. https://doi.org/10.3390/v14112535
Lim S, Zhang M, Chang TL. ACE2-Independent Alternative Receptors for SARS-CoV-2. Viruses. 2022; 14(11):2535. https://doi.org/10.3390/v14112535
Chicago/Turabian StyleLim, Suhyeon, Monica Zhang, and Theresa L. Chang. 2022. "ACE2-Independent Alternative Receptors for SARS-CoV-2" Viruses 14, no. 11: 2535. https://doi.org/10.3390/v14112535
APA StyleLim, S., Zhang, M., & Chang, T. L. (2022). ACE2-Independent Alternative Receptors for SARS-CoV-2. Viruses, 14(11), 2535. https://doi.org/10.3390/v14112535







