Superficial Inguinal Lymph Nodes for Screening Dead Pigs for African Swine Fever
Abstract
1. Introduction
2. Materials and Methods
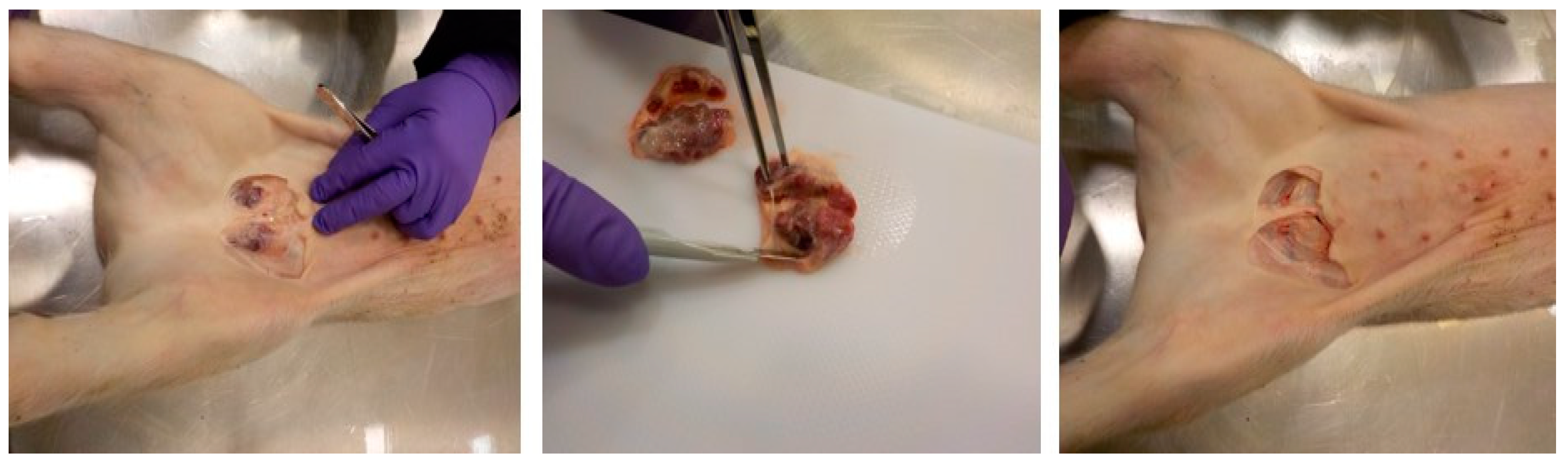
2.1. Clinical Samples
2.2. ASFV Genome Detection
2.3. Virus Isolation
2.4. Statistical Analysis
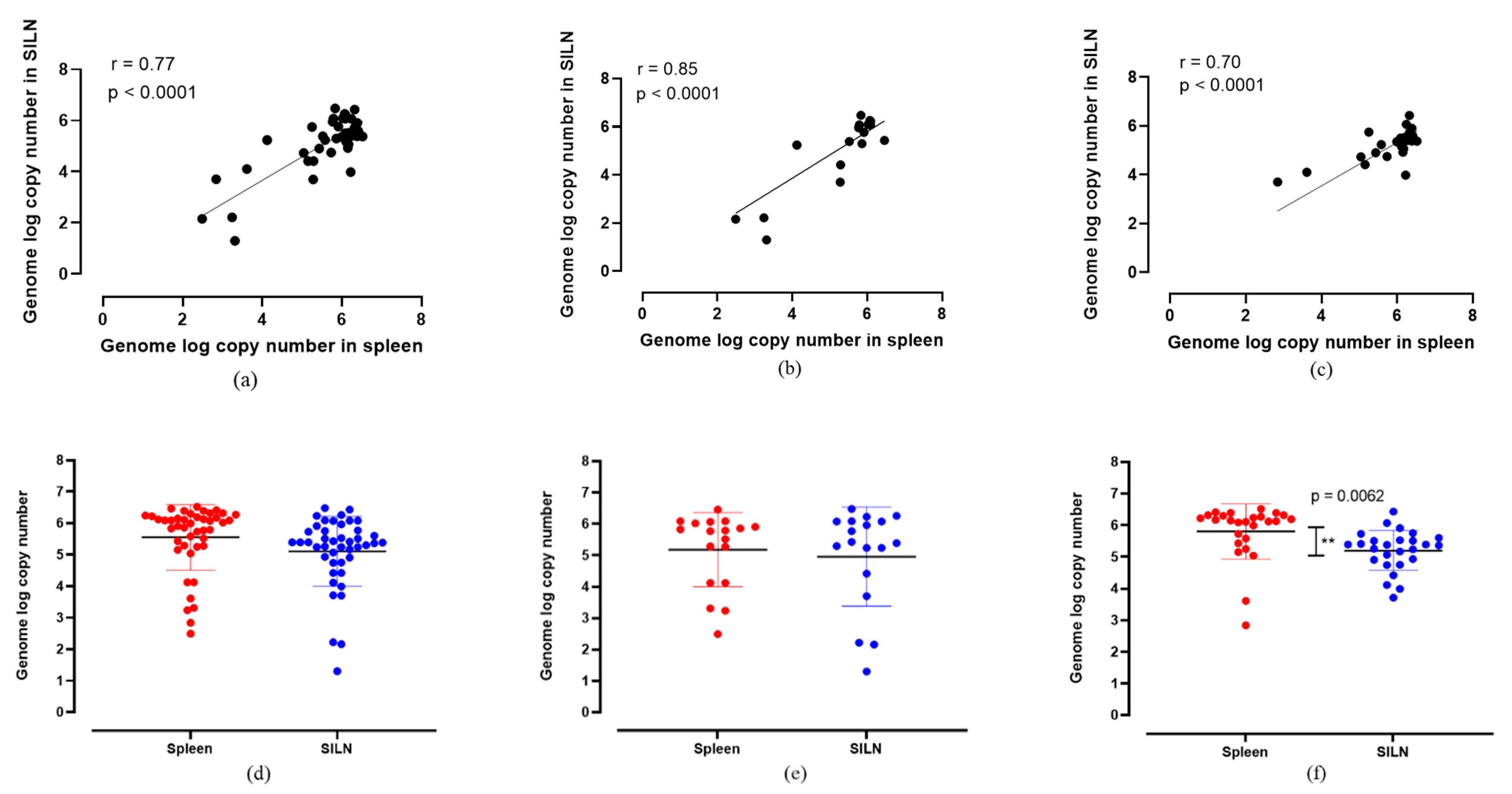
3. Results and Discussion
4. Conclusions
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Penrith, M.L. Current status of African swine fever. CABI Agric. Biosci. 2020, 1, 11. [Google Scholar] [CrossRef]
- Gonzales, W.; Moreno, C.; Duran, U.; Henao, N.; Bencosme, M.; Lora, P.; Reyes, R.; Núñez, R.; De Gracia, A.; Perez, A.M. African swine fever in the Dominican Republic. Transbound. Emerg. Dis. 2021, 68, 3018–3019. [Google Scholar] [CrossRef]
- OIE-WAHIS Immediate Notification: African Swine Fever Virus Haiti, Reported on 2021-09-19, report ID IN_151732. Available online: https://wahis.oie.int/#/report-info?reportId=39928 (accessed on 10 November 2021).
- JAVMA. African Swine Fever Reaches the Caribbean; Infections in the Region a Source of Caution, Concern in U.S. Available online: https://www.avma.org/javma-news/2021-09-15/african-swine-fever-reaches-caribbean (accessed on 10 November 2021).
- Gaudreault, N.N.; Madden, D.W.; Wilson, W.C.; Trujillo, J.D.; Richt, J.A. African Swine Fever Virus: An Emerging DNA Arbovirus. Front. Vet. Sci. 2020, 7, 215. [Google Scholar] [CrossRef] [PubMed]
- Gervasi, V.; Marcon, A.; Bellini, S.; Guberti, V. Evaluation of the Efficiency of Active and Passive Surveillance in the Detection of African Swine Fever in Wild Boar. Vet. Sci. 2019, 7, 5. [Google Scholar] [CrossRef] [PubMed]
- Cappai, S.; Rolesu, S.; Feliziani, F.; Desini, P.; Guberti, V.; Loi, F. Standardized Methodology for Target Surveillance against African Swine Fever. Vaccines 2020, 8, 723. [Google Scholar] [CrossRef] [PubMed]
- European Food Safety Authority; Boklund, A.; Cay, B.; Depner, K.; Foldi, Z.; Guberti, V.; Masiulis, M.; Miteva, A.; More, S.; Olsevskis, E.; et al. Epidemiological analyses of African swine fever in the European Union (November 2017 until November 2018). EFSA J. 2018, 16, e05494. [Google Scholar] [CrossRef]
- Busch, F.; Haumont, C.; Penrith, M.-L.; Laddomada, A.; Dietze, K.; Globig, A.; Guberti, V.; Zani, L.; Depner, K. Evidence-Based African Swine Fever Policies: Do We Address Virus and Host Adequately? Front. Vet. Sci. 2021, 8, 224. [Google Scholar] [CrossRef]
- OIE. African Swine Fever (Infection with African Swine Fever Virus). In Manual of Diagnostic Tests and Vaccines for Terrestrial Animals 2019; OIE: Paris, France, 2019. [Google Scholar]
- Rowlands, R.J.; Michaud, V.; Heath, L.; Hutchings, G.; Oura, C.; Vosloo, W.; Dwarka, R.; Onashvili, T.; Albina, E.; Dixon, L.K. African swine fever virus isolate, Georgia, 2007. Emerg. Infect. Dis. 2008, 14, 1870–1874. [Google Scholar] [CrossRef]
- Gallardo, C.; Fernández-Pinero, J.; Arias, M. African swine fever (ASF) diagnosis, an essential tool in the epidemiological investigation. Virus Res. 2019, 271, 197676. [Google Scholar] [CrossRef]
- Nurmoja, I.; Schulz, K.; Staubach, C.; Sauter-Louis, C.; Depner, K.; Conraths, F.J.; Viltrop, A. Development of African swine fever epidemic among wild boar in Estonia—Two different areas in the epidemiological focus. Sci. Rep. 2017, 7, 12562. [Google Scholar] [CrossRef]
- Zani, L.; Forth, J.H.; Forth, L.; Nurmoja, I.; Leidenberger, S.; Henke, J.; Carlson, J.; Breidenstein, C.; Viltrop, A.; Hoper, D.; et al. Deletion at the 5’-end of Estonian ASFV strains associated with an attenuated phenotype. Sci. Rep. 2018, 8, 6510. [Google Scholar] [CrossRef]
- Guinat, C.; Gogin, A.; Blome, S.; Keil, G.; Pollin, R.; Pfeiffer, D.U.; Dixon, L. Transmission routes of African swine fever virus to domestic pigs: Current knowledge and future research directions. Vet. Rec. 2016, 178, 262–267. [Google Scholar] [CrossRef]
- Greig, A. Pathogenesis of African swine fever in pigs naturally exposed to the disease. J. Comp. Pathol. 1972, 82, 73–79. [Google Scholar] [CrossRef]
- Blome, S.; Gabriel, C.; Beer, M. Pathogenesis of African swine fever in domestic pigs and European wild boar. Virus Res. 2013, 173, 122–130. [Google Scholar] [CrossRef] [PubMed]
- Heuschele, W.P. Studies on the pathogenesis of african swine fever I. Quantitative studies on the sequential development of virus in pig tissues. Arch. Die Gesamte Virusforsch. 1967, 21, 349–356. [Google Scholar] [CrossRef]
- Lacasta, A.; López Monteagudo, P.; Jiménez-Marín, A.; Accensi, F.; Ballester, M.; Argilaguet, J.; Galindo, I.; Segalés, J.; Salas, M.; Domínguez, J.; et al. Live attenuated African swine fever viruses as ideal tools to dissect the mechanisms involved in viral pathogenesis and immune protection. Vet. Res. 2015, 46, 135. [Google Scholar] [CrossRef]
- Kraitchman, D.; Kamel, I.; Weiss, C.; Georgiades, C. Mapping of the swine superficial lymph nodes: Developing a large animal model for lymphatic interventional research. J. Vasc. Interv. Radiol. 2017, 28, S182. [Google Scholar] [CrossRef][Green Version]
- Lee, H.S.; Bui, V.N.; Dao, D.T.; Bui, N.A.; Le, T.D.; Kieu, M.A.; Nguyen, Q.H.; Tran, L.H.; Roh, J.-H.; So, K.-M.; et al. Pathogenicity of an African swine fever virus strain isolated in Vietnam and alternative diagnostic specimens for early detection of viral infection. Porc. Health Manag. 2021, 7, 36. [Google Scholar] [CrossRef]
- Pikalo, J.; Deutschmann, P.; Fischer, M.; Roszyk, H.; Beer, M.; Blome, S. African Swine Fever Laboratory Diagnosis-Lessons Learned from Recent Animal Trials. Pathogens 2021, 10, 177. [Google Scholar] [CrossRef]
- Adedeji, A.J.; Luka, P.D.; Atai, R.B.; Olubade, T.A.; Hambolu, D.A.; Ogunleye, M.A.; Muwanika, V.B.; Masembe, C. First-Time Presence of African Swine Fever Virus Genotype II in Nigeria. Microbiol. Resour. Announc. 2021, 10, e0035021. [Google Scholar] [CrossRef] [PubMed]
- Goonewardene, K.B.; Chung, C.J.; Goolia, M.; Blakemore, L.; Fabian, A.; Mohamed, F.; Nfon, C.; Clavijo, A.; Dodd, K.A.; Ambagala, A. Evaluation of oral fluid as an aggregate sample for early detection of African swine fever virus using four independent pen-based experimental studies. Transbound. Emerg. Dis. 2021, 68, 2867–2877. [Google Scholar] [CrossRef]
- Senthilkumaran, C.; Bittner, H.; Ambagala, A.; Lung, O.; Babiuk, S.; Yang, M.; Zimmerman, J.; Gimenez-Lirola, L.G.; Nfon, C. Use of Oral Fluids for Detection of Virus and Antibodies in Pigs Infected with Swine Vesicular Disease Virus. Transbound. Emerg. Dis. 2017, 64, 1762–1770. [Google Scholar] [CrossRef]
- Tignon, M.; Gallardo, C.; Iscaro, C.; Hutet, E.; Van der Stede, Y.; Kolbasov, D.; De Mia, G.M.; Le Potier, M.F.; Bishop, R.P.; Arias, M.; et al. Development and inter-laboratory validation study of an improved new real-time PCR assay with internal control for detection and laboratory diagnosis of African swine fever virus. J. Virol. Methods 2011, 178, 161–170. [Google Scholar] [CrossRef] [PubMed]
- Moniwa, M.; Clavijo, A.; Li, M.; Collignon, B.; Kitching, P.R. Performance of a foot-and-mouth disease virus reverse transcription-polymerase chain reaction with amplification controls between three real-time instruments. J. Vet. Diagn. Investig. 2007, 19, 9–20. [Google Scholar] [CrossRef] [PubMed]
- Niederwerder, M.C.; Stoian, A.M.M.; Rowland, R.R.R.; Dritz, S.S.; Petrovan, V.; Constance, L.A.; Gebhardt, J.T.; Olcha, M.; Jones, C.K.; Woodworth, J.C.; et al. Infectious Dose of African Swine Fever Virus When Consumed Naturally in Liquid or Feed. Emerg. Infect. Dis. 2019, 25, 891–897. [Google Scholar] [CrossRef] [PubMed]
- Fischer, M.; Hühr, J.; Blome, S.; Conraths, F.J.; Probst, C. Stability of African Swine Fever Virus in Carcasses of Domestic Pigs and Wild Boar Experimentally Infected with the ASFV “Estonia 2014” Isolate. Viruses 2020, 12, 1118. [Google Scholar] [CrossRef]
- Mazur-Panasiuk, N.; Żmudzki, J.; Woźniakowski, G. African Swine Fever Virus—Persistence in Different Environmental Conditions and the Possibility of its Indirect Transmission. J. Vet. Res. 2019, 63, 303–310. [Google Scholar] [CrossRef]
- Gallardo, C.; Soler, A.; Rodze, I.; Nieto, R.; Cano-Gómez, C.; Fernandez-Pinero, J.; Arias, M. Attenuated and non-haemadsorbing (non-HAD) genotype II African swine fever virus (ASFV) isolated in Europe, Latvia 2017. Transbound. Emerg. Dis. 2019, 66, 1399–1404. [Google Scholar] [CrossRef]
| Category | Experiment | ASFV Strain | Pig # | DPI/DPE | Dead or Euthanized | Genome Log Copy # in Blood | Genome Log Copy # in SILNs | Genome Log Copy # in Spleen |
|---|---|---|---|---|---|---|---|---|
| Moderately virulent | 1 | ASFV Estonia 2014 | 1 | 10 | Euthanized | 4.02 | 1.30 | 3.31 |
| 2 | 9 | Euthanized | 3.50 | 2.22 | 3.24 | |||
| 3 | 17 | Euthanized | 3.59 | 2.16 | 2.49 | |||
| 2 | ASFV Estonia 2014 | 4 | 11 | Euthanized | 4.27 | 5.24 | 4.12 | |
| 5 | 10 | Euthanized | 5.76 | 6.08 | 6.09 | |||
| 6 | 9 | Euthanasia | 5.92 | 6.09 | 6.02 | |||
| 7 | 8 | * Dead | Not collected | 6.23 | 6.09 | |||
| 8 | 11 | Euthanized | 5.70 | 3.70 | 5.28 | |||
| 9 | 8 | Euthanized | 5.30 | 6.08 | 5.79 | |||
| 10 | 11 | * Dead | Not collected | 5.24 | 4.12 | |||
| 11 | 8 | Euthanized | 5.46 | 5.77 | 5.91 | |||
| 12 | 9 | Euthanized | 6.03 | 5.43 | 6.46 | |||
| 13 | 8 | Euthanized | 5.93 | 5.96 | 5.77 | |||
| 14 | 9 | Euthanized | 5.92 | 5.30 | 5.86 | |||
| 15 | 8 | Euthanized | 5.70 | 6.26 | 6.07 | |||
| 16 | 10 | Euthanized | 4.40 | 4.42 | 5.29 | |||
| 17 | 10 | * Dead | Not collected | 5.39 | 5.52 | |||
| 18 | 7 | * Dead | Not collected | 6.48 | 5.83 | |||
| Highly virulent | 3 | ASFV Ghana 20 | 19 | 7 | Dead | 5.13 | 4.75 | 5.73 |
| 20 | 7 | Euthanized | 5.88 | 5.73 | 6.32 | |||
| 4 | ASFV Nigeria RV502 | 21 | 7 | Euthanized | 6.03 | 6.43 | 6.32 | |
| 22 | 7 | Euthanized | 5.58 | 5.38 | 6.52 | |||
| 5 | ASFV Vietnam2561 | 23 | 9 | Dead | 5.37 | 5.51 | 6.30 | |
| 24 | 12 | Euthanized | 3.49 | 4.11 | 3.61 | |||
| 25 | 12 | Euthanized | 3.15 | 3.71 | 2.84 | |||
| 6 | ASFV Georgia 2007/1 | 26 | 15 | Dead | 5.80 | 5.75 | 5.25 | |
| 27 | 22 | Dead | 3.64 | 5.60 | 6.41 | |||
| 28 | 17 | Euthanized | 5.64 | 5.24 | 5.58 | |||
| 29 | 18 | Dead | 6.09 | 5.07 | 6.17 | |||
| 30 | 23 | Euthanized | 5.52 | 3.99 | 6.22 | |||
| 31 | 14 | Dead | 5.90 | 5.36 | 5.99 | |||
| 32 | 16 | Euthanized | 5.52 | 4.91 | 5.43 | |||
| 33 | 23 | Euthanized | 3.87 | 5.39 | 6.39 | |||
| 34 | 23 | Euthanized | 5.79 | 5.26 | 6.13 | |||
| ** 35 | 7 | Dead | 5.39 | 6.07 | 6.24 | |||
| 36 | 14 | Euthanized | 5.54 | 5.39 | 6.12 | |||
| 37 | 14 | Euthanized | 5.48 | 5.51 | 6.09 | |||
| 38 | 13 | Euthanized | 5.89 | 4.93 | 6.15 | |||
| 39 | 22 | Dead | 5.34 | 5.91 | 6.39 | |||
| 40 | 25 | Euthanized | 6.09 | 5.17 | 6.11 | |||
| 41 | 14 | Euthanized | 5.20 | 4.74 | 5.04 | |||
| 42 | 13 | Dead | 5.20 | 5.41 | 6.27 | |||
| 43 | 15 | Euthanized | 4.77 | 4.42 | 5.15 | |||
| 44 | 25 | Euthanized | 6.00 | 5.53 | 6.19 |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2022 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Goonewardene, K.B.; Onyilagha, C.; Goolia, M.; Le, V.P.; Blome, S.; Ambagala, A. Superficial Inguinal Lymph Nodes for Screening Dead Pigs for African Swine Fever. Viruses 2022, 14, 83. https://doi.org/10.3390/v14010083
Goonewardene KB, Onyilagha C, Goolia M, Le VP, Blome S, Ambagala A. Superficial Inguinal Lymph Nodes for Screening Dead Pigs for African Swine Fever. Viruses. 2022; 14(1):83. https://doi.org/10.3390/v14010083
Chicago/Turabian StyleGoonewardene, Kalhari Bandara, Chukwunonso Onyilagha, Melissa Goolia, Van Phan Le, Sandra Blome, and Aruna Ambagala. 2022. "Superficial Inguinal Lymph Nodes for Screening Dead Pigs for African Swine Fever" Viruses 14, no. 1: 83. https://doi.org/10.3390/v14010083
APA StyleGoonewardene, K. B., Onyilagha, C., Goolia, M., Le, V. P., Blome, S., & Ambagala, A. (2022). Superficial Inguinal Lymph Nodes for Screening Dead Pigs for African Swine Fever. Viruses, 14(1), 83. https://doi.org/10.3390/v14010083









