Are Bordetella bronchiseptica Siphoviruses (Genus Vojvodinavirus) Appropriate for Phage Therapy—Bacterial Allies or Foes?
Abstract
1. Introduction
2. Materials and Methods
2.1. Bacterial Strains
2.2. Phages
2.3. Phage Genome Sequencing and Analysis
2.4. Structural Protein Analysis
2.5. Phage Sensitivity to Environmental Factors
2.6. In Vitro Inhibition of Bacterial Growth and Biofilm Formation by Phages
2.7. Phage Effect on B. bronchiseptica Biofilm Formation
2.8. Phage Effect on 24-h Old B. bronchiseptica Biofilm
2.9. Phage Genome Presence in Bacteria upon Infection
2.10. Phenotypic Changes of lysogenic B. bronchiseptica
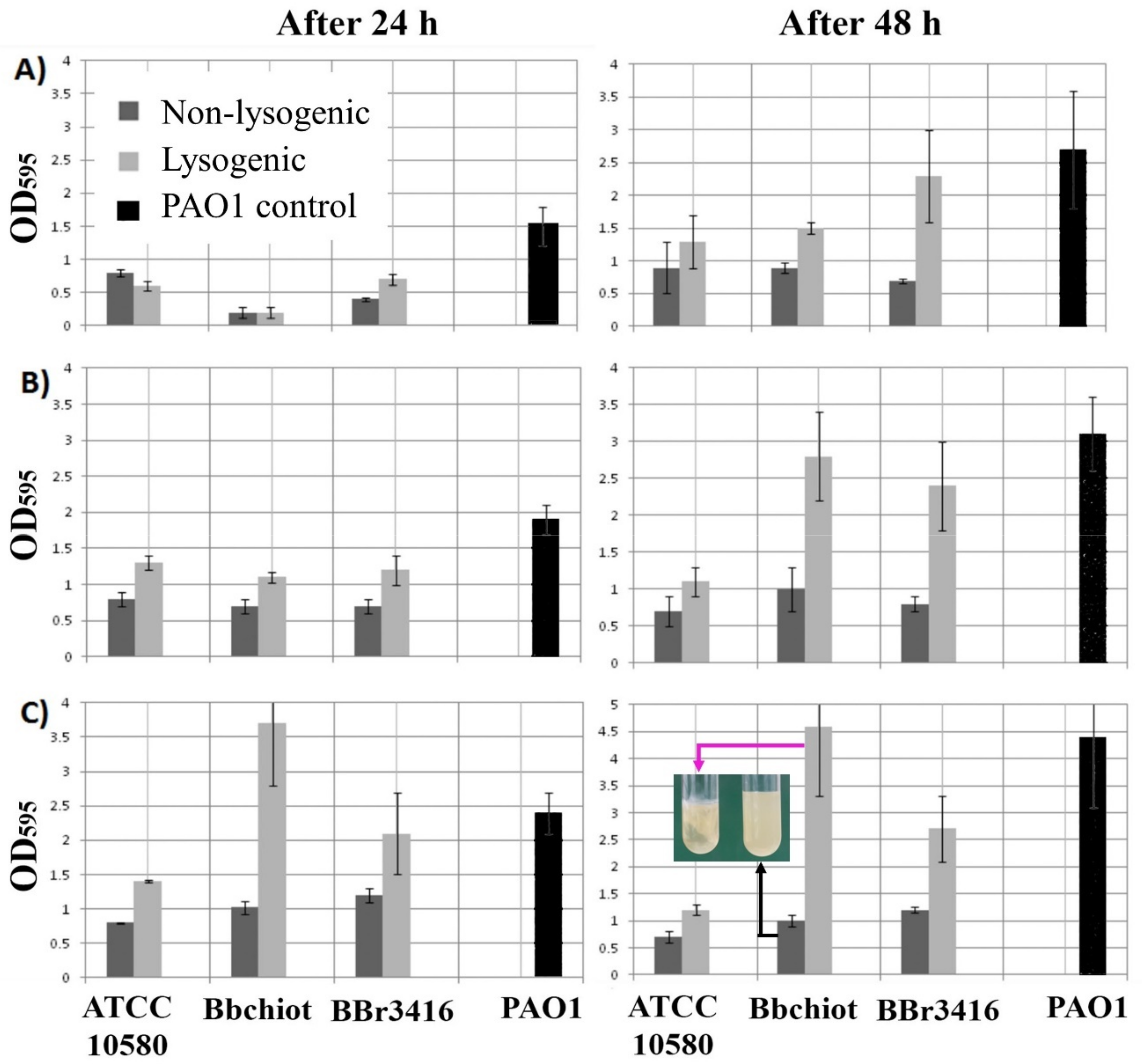
2.10.1. Biofilm Production of Lysogenic Bacteria
2.10.2. Hemolytic Activity of Lysogenic Bacteria
2.10.3. Susceptibility of Lysogenic Bacteria to Antibiotics
2.10.4. Swimming and Twitching Motility of Lysogenic Bacteria
2.10.5. Fimbria Production
3. Results
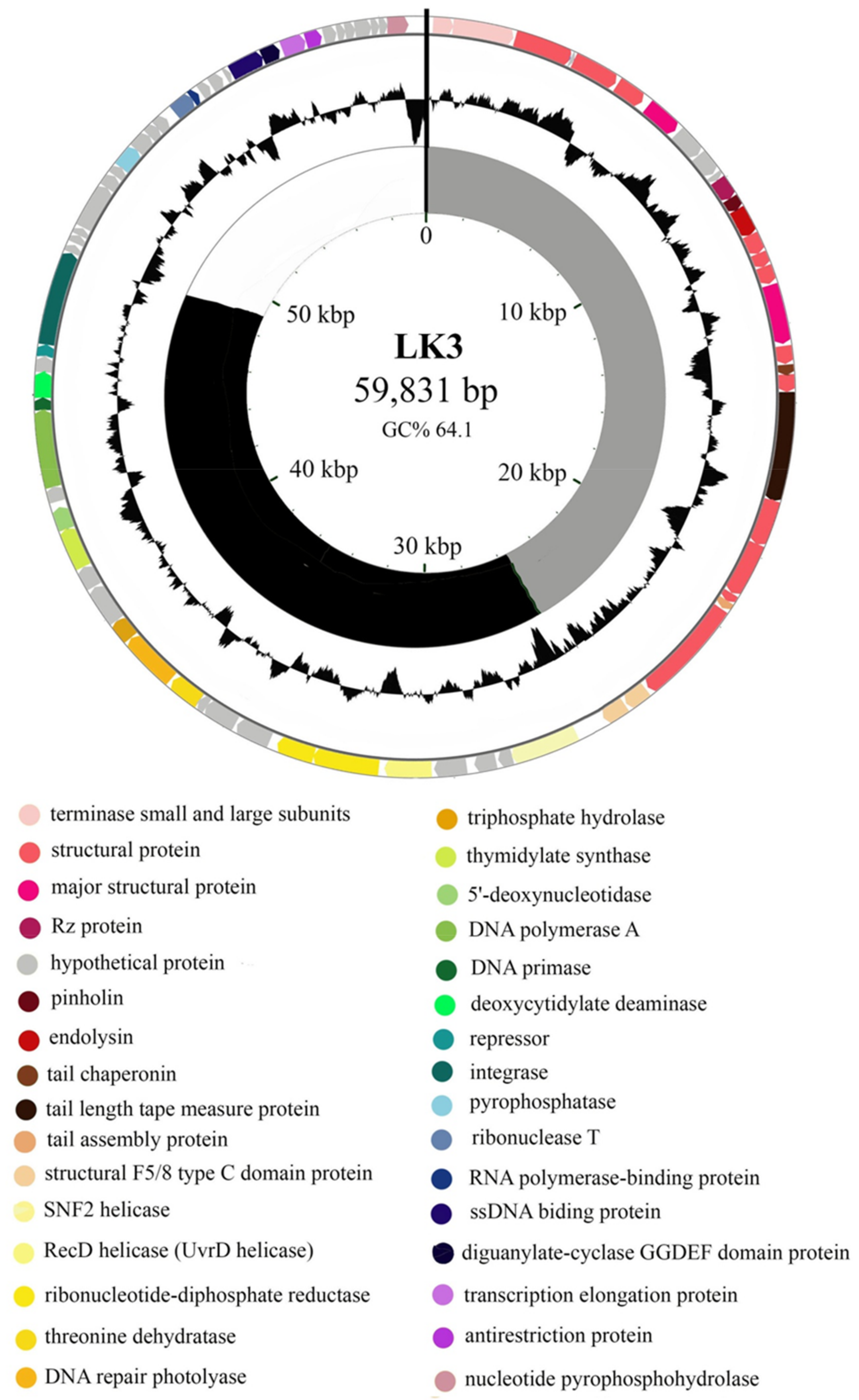
3.1. Bordetella Phage LK3 Genome Properties
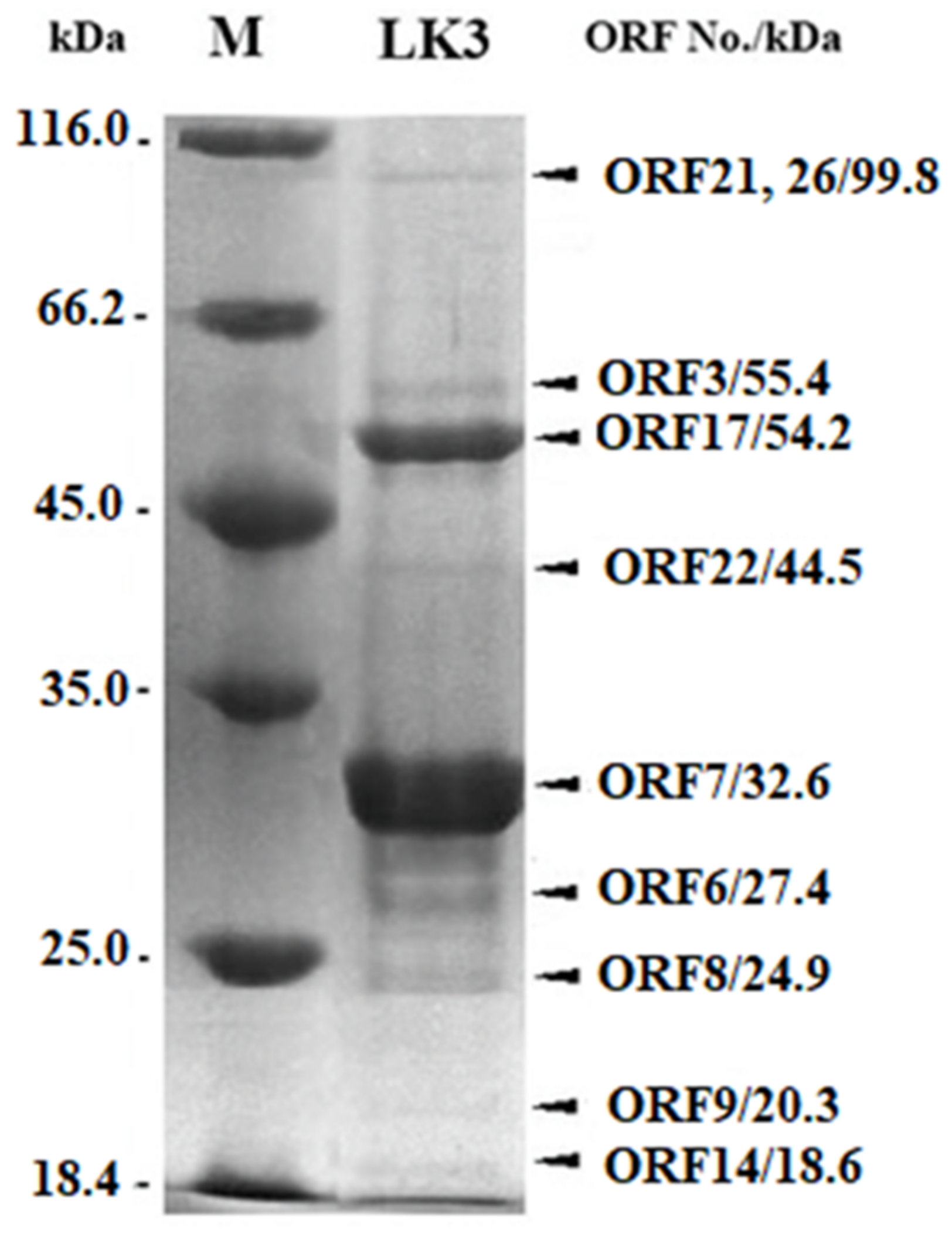
3.2. Proteomic Characterization of LK3 Phage
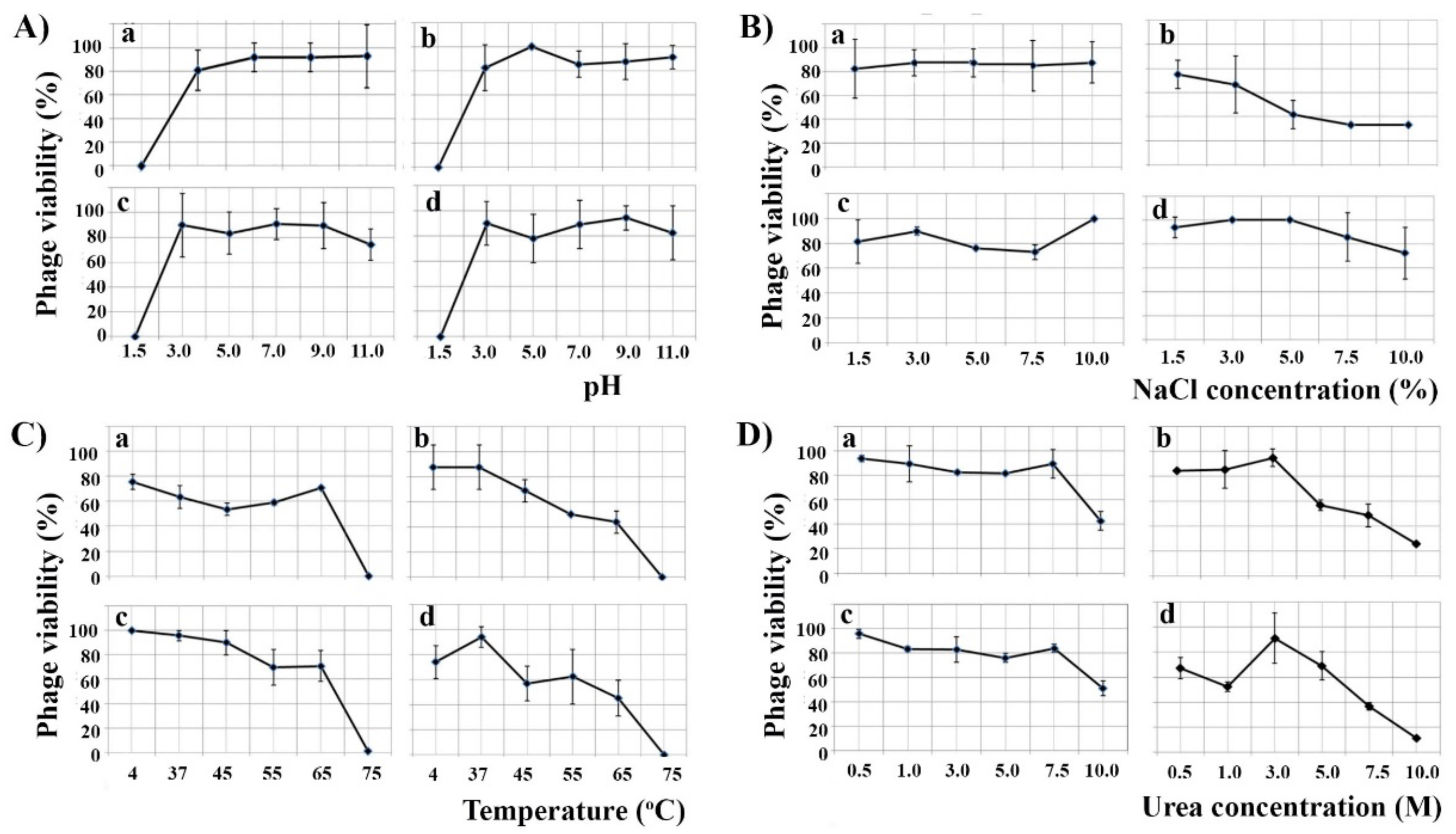
3.3. Phage Resistance to Environmental Factors
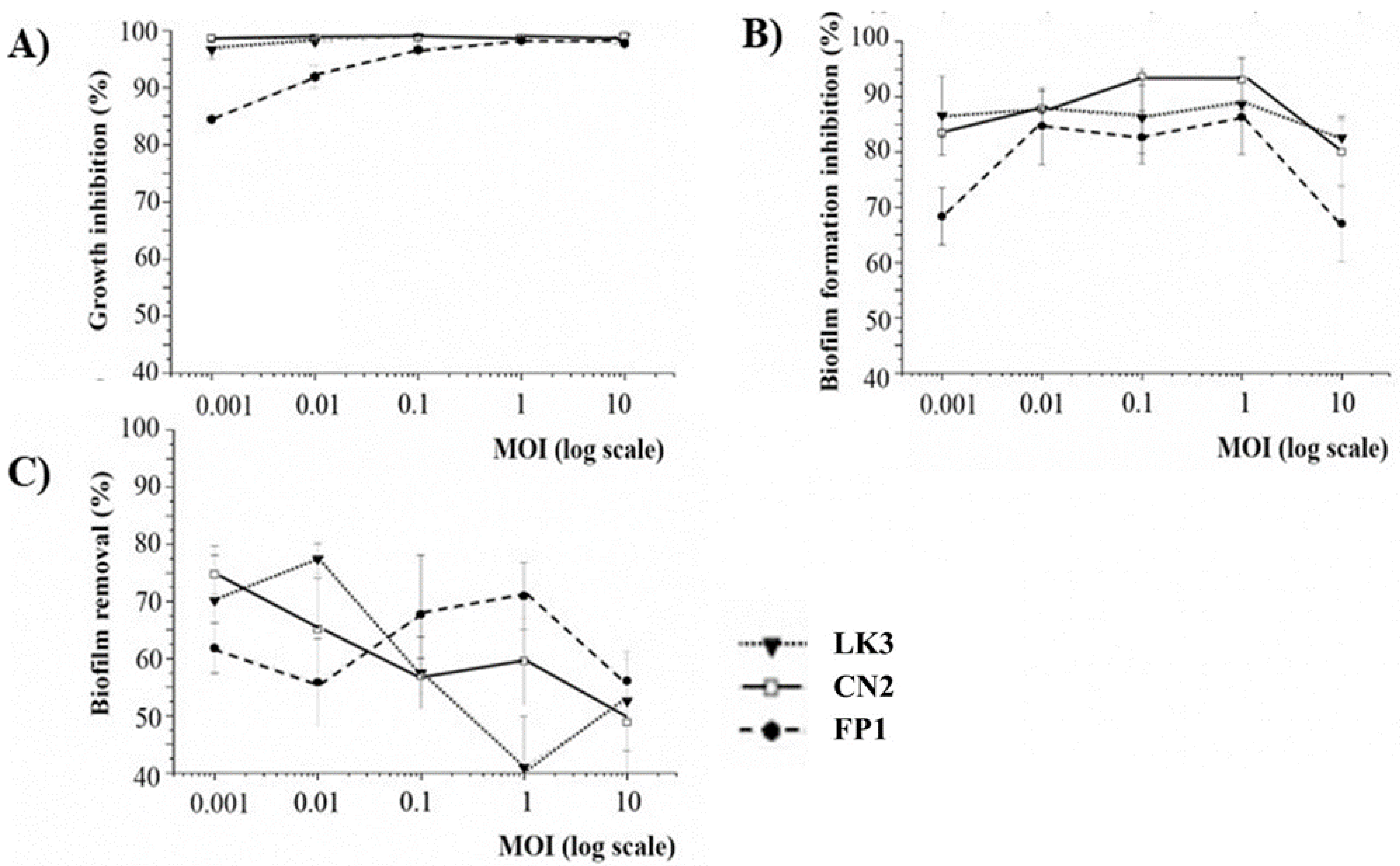
3.4. Phage Inhibition of Bacterial Growth and Biofilm Formation
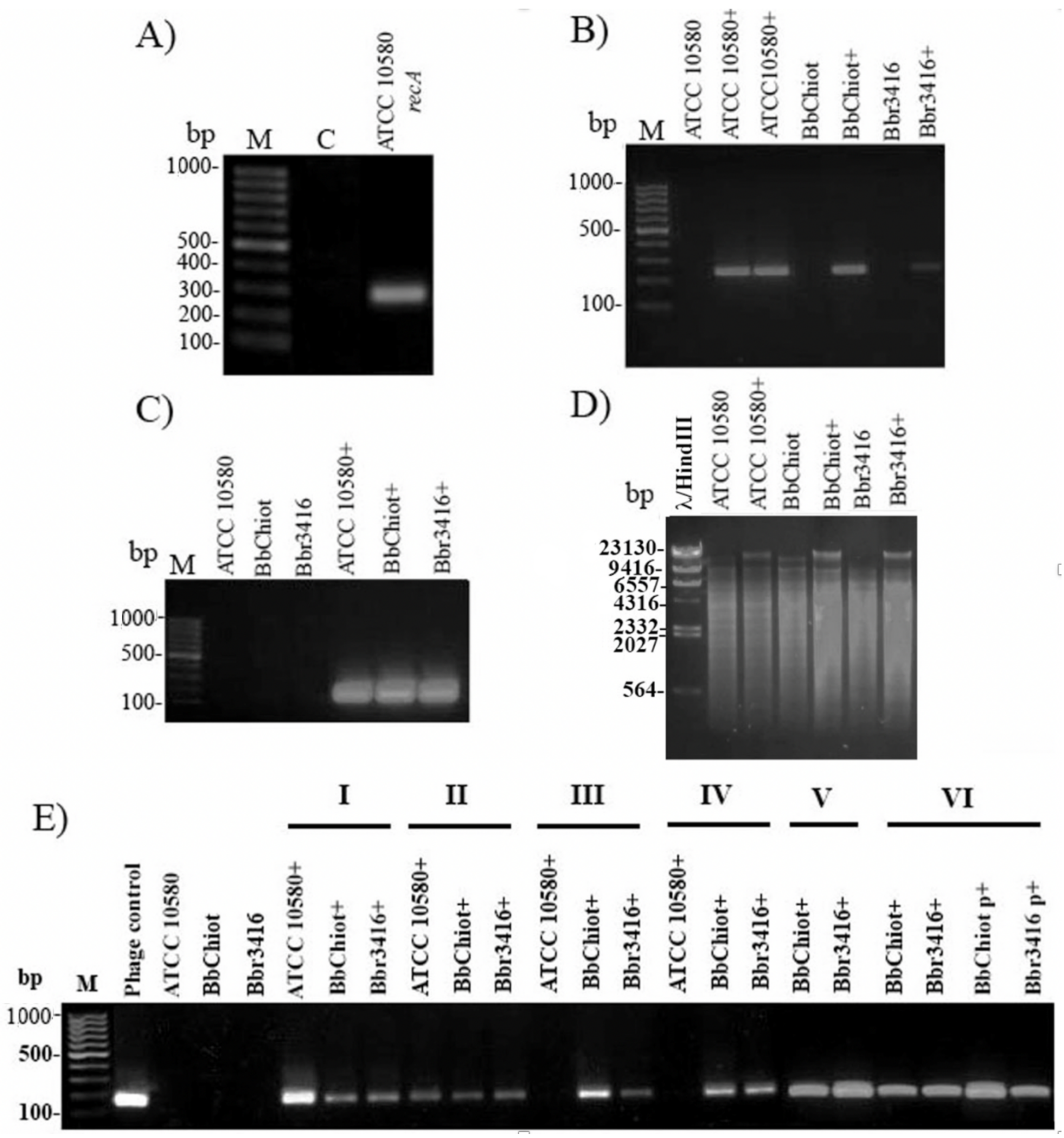
3.5. Confirmation of LK3 Infection and Stability of Lysogens
3.6. Lysogenic Conversion of B. bronchiseptica by LK3 Siphovirus
4. Discussion
5. Conclusions
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Garcia-Cantu, M.C.; Hartmann, F.A.; Brown, C.M.; Darien, B.J. Bordetella bronchiseptica and equine respiratory infections: A review of 30 cases. Equine. Vet. Educ. 2000, 12, 45–50. [Google Scholar] [CrossRef]
- Parkhill, J.; Sebaihia, M.; Preston, A.; Murphy, L.D.; Thomson, N.; Harris, D.E.; Holden, M.T.G.; Churcher, C.M.; Bentley, S.D.; Mungall, K.L.; et al. Comparative analysis of the genome sequences of Bordetella pertussis, Bordetella parapertussis and Bordetella bronchiseptica. Nat. Genet. 2003, 35, 32–40. [Google Scholar] [CrossRef]
- Deeb, B.J.; Digiacomo, R.F.; Bernard, B.L.; Silbernagel, S.M. Pasteurella-Multocida and Bordetella-Bronchiseptica Infections in Rabbits. J. Clin. Microbiol. 1990, 28, 70–75. [Google Scholar] [CrossRef] [PubMed]
- Goodnow, R.A. Biology of Bordetella-Bronchiseptica. Microbiol. Rev. 1980, 44, 722–738. [Google Scholar] [CrossRef]
- Kadlec, K.; Wiegand, I.; Kehrenberg, C.; Schwarz, S. Studies on the mechanisms of beta-lactam resistance in Bordetella bronchiseptica. J. Antimicrob. Chemother. 2007, 59, 396–402. [Google Scholar] [CrossRef] [PubMed]
- Egberink, H.; Addie, D.; Belak, S.; Boucraut-Baralon, C.; Frymus, T.; Gruffydd-Jones, T.; Hartmann, K.; Hosie, M.J.; Lloret, A.; Lutz, H.; et al. Bordetella bronchiseptica infection in cats. ABCD guidelines on prevention and management. J. Feline Med. Surg. 2009, 11, 610–614. [Google Scholar] [CrossRef] [PubMed]
- Viitanen, S.J.; Lappalainen, A.; Rajamaki, M.M. Co-infections with Respiratory Viruses in Dogs with Bacterial Pneumonia. J. Vet. Intern. Med. 2015, 29, 544–551. [Google Scholar] [CrossRef]
- Berglof, A.; Norlander, T.; Feinstein, R.; Otori, N.; Stierna, P.; Sandstedt, K. Association of bronchopneumonia with sinusitis due to Bordetella bronchiseptica in an experimental rabbit model. Am. J. Rhinol. 2000, 14, 125–130. [Google Scholar] [CrossRef]
- Kumar, S.; Singh, B.R.; Bhardwaj, M.; Singh, V. Occurrence of Bordetella infection in pigs in northern India. Int. J. Microbiol. 2014, 2014, 238575. [Google Scholar] [CrossRef]
- Woolfrey, B.F.; Moody, J.A. Human Infections Associated with Bordetella-Bronchiseptica. Clin. Microbiol. Rev. 1991, 4, 243–255. [Google Scholar] [CrossRef]
- Ellis, J.A.; Gow, S.P.; Waldner, C.L.; Shields, S.; Wappel, S.; Bowers, A.; Lacoste, S.; Xu, Z.; Ball, E. Comparative efficacy of intranasal and oral vaccines against Bordetella bronchiseptica in dogs. Vet. J. 2016, 212, 71–77. [Google Scholar] [CrossRef] [PubMed]
- Ellis, J.A. How well do vaccines for Bordetella bronchiseptica work in dogs? A critical review of the literature 1977–2014. Vet. J. 2015, 204, 5–16. [Google Scholar] [CrossRef]
- Speakman, A.J.; Binns, S.H.; Dawson, S.; Hart, C.A.; Gaskell, R.M. Antimicrobial susceptibility of Bordetella bronchiseptica isolates from cats and a comparison of the agar dilution and E-test methods. Vet. Microbiol. 1997, 54, 63–72. [Google Scholar] [CrossRef]
- Pruller, S.; Rensch, U.; Meemken, D.; Kaspar, H.; Kopp, P.A.; Klein, G.; Kehrenberg, C. Antimicrobial Susceptibility of Bordetella bronchiseptica Isolates from Swine and Companion Animals and Detection of Resistance Genes. PLoS ONE 2015, 10, e0135703. [Google Scholar] [CrossRef]
- Zhang, Y.; Yang, H.; Guo, L.; Zhao, M.F.; Wang, F.; Song, W.B.; Hua, L.; Wang, L.; Liang, W.; Tang, X.B.A.; et al. Isolation, Antimicrobial Resistance Phenotypes, and Virulence Genes of Bordetella bronchiseptica From Pigs in China, 2018–2020. Front. Vet. Sci. 2021, 8, 672716. [Google Scholar] [CrossRef]
- Cisek, A.A.; Dabrowska, I.; Gregorczyk, K.P.; Wyzewski, Z. Phage Therapy in Bacterial Infections Treatment: One Hundred Years After the Discovery of Bacteriophages. Curr. Microbiol. 2017, 74, 277–283. [Google Scholar] [CrossRef] [PubMed]
- Petrovic Fabijan, A.; Lin, R.C.Y.; Ho, J.; Maddocks, S.; Ben Zakour, N.L.; Iredell, J.R.; Westmead Bacteriophage Therapy Team. Safety of bacteriophage therapy in severe Staphylococcus aureus infection. Nat. Microbiol. 2020, 5, 465–472. [Google Scholar] [CrossRef] [PubMed]
- Petrovic Fabijan, A.; Khalid, A.; Maddocks, S.; Ho, J.; Gilbey, T.; Sandaradura, I.; Lin, R.C.; Ben Zakour, N.; Venturini, C.; Bowring, B.; et al. Phage therapy for severe bacterial infections: A narrative review. Med. J. Aust. 2020, 212, 279–285. [Google Scholar] [CrossRef] [PubMed]
- Knezevic, P.; Hoyle, N.S.; Matsuzaki, S.; Gorski, A. Editorial: Advances in Phage Therapy: Present Challenges and Future Perspectives. Front. Microbiol. 2021, 12, 1390. [Google Scholar] [CrossRef] [PubMed]
- Liu, M.S.; Gingery, M.; Doulatov, S.R.; Liu, Y.C.; Hodes, A.; Baker, S.; Davis, P.; Simmonds, M.; Churcher, C.; Mungall, K.; et al. Genomic and genetic analysis of Bordetella bacteriophages encoding reverse transcriptase-mediated tropism-switching cassettes. J. Bacteriol. 2004, 186, 1503–1517. [Google Scholar] [CrossRef] [PubMed]
- Chen, Y.B.; Yang, L.; Sun, E.C.; Song, J.Y.; Wu, B. Characterisation of a newly detected bacteriophage infecting Bordetella bronchiseptica in swine. Arch. Virol. 2019, 164, 33–40. [Google Scholar] [CrossRef] [PubMed]
- Szymczak, M.; Grygorcewicz, B.; Karczewska-Golec, J.; Decewicz, P.; Pankowski, J.A.; Orszigh-Szturo, H.; Bacal, P.; Dolegowska, B.; Golec, P. Characterization of a Unique Bordetella bronchiseptica vB_BbrP_BB8 Bacteriophage and Its Application as an Antibacterial Agent. Int. J. Mol. Sci. 2020, 21, 1403. [Google Scholar] [CrossRef]
- Petrovic, A.; Kostanjsek, R.; Rakhely, G.; Knezevic, P. The First Siphoviridae Family Bacteriophages Infecting Bordetella bronchiseptica Isolated from Environment. Microb. Ecol. 2017, 73, 368–377. [Google Scholar] [CrossRef]
- Knezevic, P.K.A.; Adriaenssens, E.M. ICTV Taxonomy History: Vojvodinavirus; ICTV, 2020. [Google Scholar]
- Tizolova, A.; Guiso, N.; Guillot, S. Insertion sequences shared by Bordetella species and implications for the biological diagnosis of pertussis syndrome. Eur. J. Clin. Microbiol. Infect. Dis. 2013, 32, 89–96. [Google Scholar] [CrossRef]
- Tizolova, A.; Brun, D.; Guiso, N.; Guillot, S. Development of real-time PCR assay for differential detection of Bordetella bronchiseptica and Bordetella parapertussis. Diagn. Microbiol. Infect. Dis. 2014, 78, 347–351. [Google Scholar] [CrossRef]
- Sambrook, J.; Russell, D.W. Molecular Cloning: A Laboratory Manual, 3rd ed.; Cold Spring Harbor Laboratory Press: New York, NY, USA, 2001. [Google Scholar]
- Besemer, J.; Lomsadze, A.; Borodovsky, M. GeneMarkS: A self-training method for prediction of gene starts in microbial genomes. Implications for finding sequence motifs in regulatory regions. Nucleic Acids Res. 2001, 29, 2607–2618. [Google Scholar] [CrossRef]
- Altschul, S.F.; Gish, W.; Miller, W.; Myers, E.W.; Lipman, D.J. Basic local alignment search tool. J. Mol. Biol. 1990, 215, 403–410. [Google Scholar] [CrossRef]
- Lowe, T.M.; Chan, P.P. tRNAscan-SE On-line: Integrating search and context for analysis of transfer RNA genes. Nucleic Acids Res. 2016, 44, W54–W57. [Google Scholar] [CrossRef]
- Walker, P.J.; Siddell, S.G.; Lefkowitz, E.J.; Mushegian, A.R.; Adriaenssens, E.M.; Dempsey, D.M.; Dutilh, B.E.; Harrach, B.; Harrison, R.L.; Hendrickson, R.C.; et al. Changes to virus taxonomy and the Statutes ratified by the International Committee on Taxonomy of Viruses (2020). Arch. Virol. 2020, 165, 2737–2748. [Google Scholar] [CrossRef] [PubMed]
- McLaughlin, M.R. Simple colorimetric microplate test of phage lysis in Salmonella enterica. J. Microbiol. Methods 2007, 69, 394–398. [Google Scholar] [CrossRef] [PubMed]
- Knezevic, P.; Petrovic, O. A colorimetric microtiter plate method for assessment of phage effect on Pseudomonas aeruginosa biofilm. J. Microbiol. Methods 2008, 74, 114–118. [Google Scholar] [CrossRef]
- Knezevic, P.; Obreht, D.; Curcin, S.; Petrusic, M.; Aleksic, V.; Kostanjsek, R.; Petrovic, O. Phages of Pseudomonas aeruginosa: Response to environmental factors and in vitro ability to inhibit bacterial growth and biofilm formation. J. Appl. Microbiol. 2011, 111, 245–254. [Google Scholar] [CrossRef]
- Stepanovic, S.; Vukovic, D.; Dakic, I.; Savic, B.; Svabic-Vlahovic, M. A modified microtiter-plate test for quantification of staphylococcal biofilm formation. J. Microbiol. Methods 2000, 40, 175–179. [Google Scholar] [CrossRef]
- Janda, J.M.; Abbott, S.L. Expression of an iron-regulated hemolysin by Edwardsiella tarda. FEMS Microbiol. Lett. 1993, 111, 275–280. [Google Scholar] [CrossRef]
- Kearns, D.B. A field guide to bacterial swarming motility. Nat. Rev. Microbiol. 2010, 8, 634–644. [Google Scholar] [CrossRef] [PubMed]
- Jain, S.; Chen, J. Attachment and Biofilm formation by various serotypes of salmonella as influenced by cellulose production and thin aggregative fimbriae biosynthesis. J. Food Protect. 2007, 70, 2473–2479. [Google Scholar] [CrossRef]
- Rosenberg, I.M. Protein Analysis and Purification: Benchtop Techniques; Springer Science & Business Media: Berlin, Germany, 2006. [Google Scholar]
- Azeredo, J.; Sutherland, I.W. The use of phages for the removal of infectious biofilms. Curr. Pharm. Biotechno. 2008, 9, 261–266. [Google Scholar] [CrossRef] [PubMed]
- Harper, D.R.; Enright, M.C. Bacteriophages for the treatment of Pseudomonas aeruginosa infections. J. Appl. Microbiol. 2011, 111, 1–7. [Google Scholar] [CrossRef]
- Sutherland, I.W.; Hughes, K.A.; Skillman, L.C.; Tait, K. The interaction of phage and biofilms. FEMS Microbiol. Lett. 2004, 232, 1–6. [Google Scholar] [CrossRef]
- Irie, Y.; Mattoo, S.; Yuk, M.H. The Bvg virulence control system regulates biofilm formation in Bordetella bronchiseptica. J. Bacteriol. 2004, 186, 5692–5698. [Google Scholar] [CrossRef] [PubMed]
- Ceyssens, P.J.; Mesyanzhinov, V.; Sykilinda, N.; Briers, Y.; Roucourt, B.; Lavigne, R.; Robben, J.; Domashin, A.; Miroshnikov, K.; Volckaert, G.; et al. The genome and structural Proteome of YuA, a new Pseudomonas aeruginosa phage resembling M6. J. Bacteriol. 2008, 190, 1429–1435. [Google Scholar] [CrossRef]
- Davies, E.V.; Winstanley, C.; Fothergill, J.L.; James, C.E. The role of temperate bacteriophages in bacterial infection. Fems Microbiol. Lett. 2016, 363, fnw015. [Google Scholar] [CrossRef]
- Karatan, E.; Watnick, P. Signals, Regulatory Networks, and Materials That Build and Break Bacterial Biofilms. Microbiol. Mol. Biol. Rev. 2009, 73, 310. [Google Scholar] [CrossRef]
- Conover, M.S.; Mishra, M.; Deora, R. Extracellular DNA Is Essential for Maintaining Bordetella Biofilm Integrity on Abiotic Surfaces and in the Upper Respiratory Tract of Mice. PLoS ONE 2011, 6, e16861. [Google Scholar] [CrossRef] [PubMed]
- Abedon, S.T. Phage Evolution and Ecology. Adv. Appl. Microbiol. 2009, 67, 1–45. [Google Scholar] [CrossRef] [PubMed]
- Broussard, G.W.; Oldfield, L.M.; Villanueva, V.M.; Lunt, B.L.; Shine, E.E.; Hatfull, G.F. Integration-Dependent Bacteriophage Immunity Provides Insights into the Evolution of Genetic Switches. Mol. Cell 2013, 49, 237–248. [Google Scholar] [CrossRef] [PubMed]
- Golding, I. Decision Making in Living Cells: Lessons from a Simple System. Annu. Rev. Biophys. 2011, 40, 63–80. [Google Scholar] [CrossRef]
- Amgarten, D.; Martins, L.F.; Lombardi, K.C.; Antunes, L.P.; de Souza, A.P.S.; Nicastro, G.G.; Kitajima, E.W.; Quaggio, R.B.; Upton, C.; Setubal, J.C.; et al. Three novel Pseudomonas phages isolated from composting provide insights into the evolution and diversity of tailed phages. BMC Genom. 2017, 18, 1–18. [Google Scholar] [CrossRef]
- Bae, H.W.; Chung, I.Y.; Sim, N.; Cho, Y.H. Complete genome sequence of Pseudomonas aeruginosa siphophage MP1412. J. Virol. 2012, 86, 9537. [Google Scholar] [CrossRef][Green Version]
- Casjens, S.R.; Gilcrease, E.B. Determining DNA packaging strategy by analysis of the termini of the chromosomes in tailed-bacteriophage virions. Methods Mol. Biol. 2009, 502, 91–111. [Google Scholar] [CrossRef]
- Lee, J.; Chastain, P.D., II; Kusakabe, T.; Griffith, J.D.; Richardson, C.C. Coordinated leading and lagging strand DNA synthesis on a minicircular template. Mol. Cell 1998, 1, 1001–1010. [Google Scholar] [CrossRef]
- Wallen, J.R.; Majka, J.; Ellenberger, T. Discrete Interactions between Bacteriophage T7 Primase-Helicase and DNA Polymerase Drive the Formation of a Priming Complex Containing Two Copies of DNA Polymerase. Biochemistry-Us 2013, 52, 4026–4036. [Google Scholar] [CrossRef] [PubMed][Green Version]
- Liu, M.; Deora, R.; Doulatov, S.R.; Gingery, M.; Eiserling, F.A.; Preston, A.; Maskell, D.J.; Simons, R.W.; Cotter, P.A.; Parkhill, J.; et al. Reverse transcriptase-mediated tropism switching in Bordetella bacteriophage. Science 2002, 295, 2091–2094. [Google Scholar] [CrossRef] [PubMed]
- Camire, R.M.; Bos, M.H.A. The molecular basis of factor V and VIII procofactor activation. J. Thromb. Haemost. 2009, 7, 1951–1961. [Google Scholar] [CrossRef]
- Briers, Y.; Walmagh, M.; Lavigne, R. Use of bacteriophage endolysin EL188 and outer membrane permeabilizers against Pseudomonas aeruginosa. J. Appl. Microbiol. 2011, 110, 778–785. [Google Scholar] [CrossRef] [PubMed]
- Fernandes, S.; Sao-Jose, C. Enzymes and Mechanisms Employed by Tailed Bacteriophages to Breach the Bacterial Cell Barriers. Viruses 2018, 10, 396. [Google Scholar] [CrossRef]
- Lohr, J.E.; Chen, F.; Hill, R.T. Genomic analysis of bacteriophage Phi JL001: Insights into its interaction with a sponge-associated alpha-proteobacterium. Appl. Environ. Microb. 2005, 71, 1598–1609. [Google Scholar] [CrossRef]
- Seguritan, V.; Feng, I.W.; Rohwer, F.; Swift, M.; Segall, A.M. Genome sequences of two closely related Vibrio parahaemolyticus phages, VP16T and VP16C. J. Bacteriol. 2003, 185, 6434–6447. [Google Scholar] [CrossRef]
- Dyson, Z.A.; Seviour, R.J.; Tucci, J.; Petrovski, S. Genome Sequences of Pseudomonas oryzihabitans Phage POR1 and Pseudomonas aeruginosa Phage PAE1. Microbiol. Resour. Ann. 2016, 4, e01515-15. [Google Scholar] [CrossRef]
- Forti, F.; Roach, D.R.; Cafora, M.; Pasini, M.E.; Horner, D.S.; Fiscarelli, E.V.; Rossitto, M.; Cariani, L.; Briani, F.; Debarbieux, L.; et al. Design of a Broad-Range Bacteriophage Cocktail That Reduces Pseudomonas aeruginosa Biofilms and Treats Acute Infections in Two Animal Models. Antimicrob. Agents Chemother. 2018, 62, e02573-17. [Google Scholar] [CrossRef]
- Essoh, C.; Latino, L.; Midoux, C.; Blouin, Y.; Loukou, G.; Nguetta, S.P.A.; Lathro, S.; Cablanmian, A.; Kouassi, A.K.; Vergnaud, G.; et al. Investigation of a Large Collection of Pseudomonas aeruginosa Bacteriophages Collected from a Single Environmental Source in Abidjan, Cote d’Ivoire. PLoS ONE 2015, 10, e0130548. [Google Scholar] [CrossRef]
- Fortier, L.C.; Sekulovic, O. Importance of prophages to evolution and virulence of bacterial pathogens. Virulence 2013, 4, 354–365. [Google Scholar] [CrossRef] [PubMed]
- Gestal, M.C.; Howard, L.K.; Dewan, K.; Johnson, H.M.; Barbier, M.; Bryant, C.; Soumana, I.H.; Rivera, I.; Linz, B.; Blas-Machado, U.; et al. Enhancement of immune response against Bordetella spp. by disrupting immunomodulation. Sci. Rep. 2019, 9, 20261. [Google Scholar] [CrossRef]
- Cieslik, M.; Baginska, N.; Jonczyk-Matysiak, E.; Wegrzyn, A.; Wegrzyn, G.; Gorski, A. Temperate Bacteriophages-The Powerful Indirect Modulators of Eukaryotic Cells and Immune Functions. Viruses 2021, 13, 1013. [Google Scholar] [CrossRef] [PubMed]
- Cattelan, N.; Dubey, P.; Arnal, L.; Yantorno, O.M.; Deora, R. Bordetella biofilms: A lifestyle leading to persistent infections. Pathog. Dis. 2016, 74, ftv108. [Google Scholar] [CrossRef]
- Jonczyk, E.; Klak, M.; Miedzybrodzki, R.; Gorski, A. The influence of external factors on bacteriophages—Review. Folia Microbiol. (Praha) 2011, 56, 191–200. [Google Scholar] [CrossRef] [PubMed]
| Label of Primer | Sequence (5′→3′) | Expected Product (bp) | Reference |
|---|---|---|---|
| Bbr-F | TGA CTT CAT GGT TGC CGT TC | 241 | [23] |
| Bbr-R | TCG GGA GCG TGA TTT CAG TA | ||
| Hem-F | ACG CCC GAA CCG TTA TTT GG | 170 | This study |
| Hem-R | CAT TTC CCC GCA ACT CGA CA | ||
| RecA-F | ATG GCG ACA ACG AGG TCG AA | 263 | [23] |
| RecA-R | CAG CAG GTC GGT CAG GTT GA | ||
| Strains | |||||
|---|---|---|---|---|---|
| Characteristics | ATCC10580 | ATCC10580+ | Bbchiot | Bbchiot+ | |
| Hemolysis (%) | Sheep blood | 14.3 ± 3.7 | 26 ± 0.28 * | N.T. | N.T. |
| Rat blood | 47 ± 6.8 | 38.1 ± 6.1 | N.T. | N.T. | |
| Cattle blood | 66.95 ± 7 | 59.1 ± 12.02 | N.T. | N.T. | |
| Motility (mm) | Swimming | 14.6 ± 0.7 | 14.4 ± 1.6 | 66.9 ± 2.5 | 78.2 ± 1.8 |
| Twitching | 4.0 ± 0.0 | 4.3 ± 0.2 | 5.0 ± 0.3 | 4.1 ± 0.2 | |
| Fimbria production | Congo-red reduction (%) | 10.0 ± 2.1 | 11,8 ± 2 | 9.1 ± 1.3 | 8.5 ± 2.3 |
| Susceptibility to antibiotics 1 (mm) | Amoxicillin/clavulanic acid | 15.8 ± 2.1 | 12.1 ± 0.9 * | 18.8 ± 1.0 | 15.3 ± 0.9 * |
| Ceftazidime | 17.3 ± 3.0 | 16.0 ± 2.0 | 20.0 ± 1.0 | 13.5 ± 0.5 * | |
| Doxycycline | 43.0 ± 1.2 | 38.3 ± 1.0 | 33.5 ± 1.7 | 35.9 ± 3.6 | |
| Sulfamethoxazole/Trimethoprim | 25.5 ± 4.1 | 19.6 ± 1.1 | 0 | 0 | |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Petrovic Fabijan, A.; Aleksic Sabo, V.; Gavric, D.; Doffkay, Z.; Rakhely, G.; Knezevic, P. Are Bordetella bronchiseptica Siphoviruses (Genus Vojvodinavirus) Appropriate for Phage Therapy—Bacterial Allies or Foes? Viruses 2021, 13, 1732. https://doi.org/10.3390/v13091732
Petrovic Fabijan A, Aleksic Sabo V, Gavric D, Doffkay Z, Rakhely G, Knezevic P. Are Bordetella bronchiseptica Siphoviruses (Genus Vojvodinavirus) Appropriate for Phage Therapy—Bacterial Allies or Foes? Viruses. 2021; 13(9):1732. https://doi.org/10.3390/v13091732
Chicago/Turabian StylePetrovic Fabijan, Aleksandra, Verica Aleksic Sabo, Damir Gavric, Zsolt Doffkay, Gábor Rakhely, and Petar Knezevic. 2021. "Are Bordetella bronchiseptica Siphoviruses (Genus Vojvodinavirus) Appropriate for Phage Therapy—Bacterial Allies or Foes?" Viruses 13, no. 9: 1732. https://doi.org/10.3390/v13091732
APA StylePetrovic Fabijan, A., Aleksic Sabo, V., Gavric, D., Doffkay, Z., Rakhely, G., & Knezevic, P. (2021). Are Bordetella bronchiseptica Siphoviruses (Genus Vojvodinavirus) Appropriate for Phage Therapy—Bacterial Allies or Foes? Viruses, 13(9), 1732. https://doi.org/10.3390/v13091732









