Clinical and Histopathologic Features of a Feline SARS-CoV-2 Infection Model Are Analogous to Acute COVID-19 in Humans
Abstract
:1. Introduction
2. Materials and Methods
2.1. Virus
2.2. Animals
2.3. Virus Challenge
2.4. Clinical Observations and Scoring
2.5. Sampling
2.6. Histopathology
2.7. Viral RNA Analysis
2.8. Feline ACE2 Analysis
2.9. Immunohistochemistry
2.10. Genomic Sequencing
2.11. Genome Assembly
2.12. Statistical Analyses
3. Results
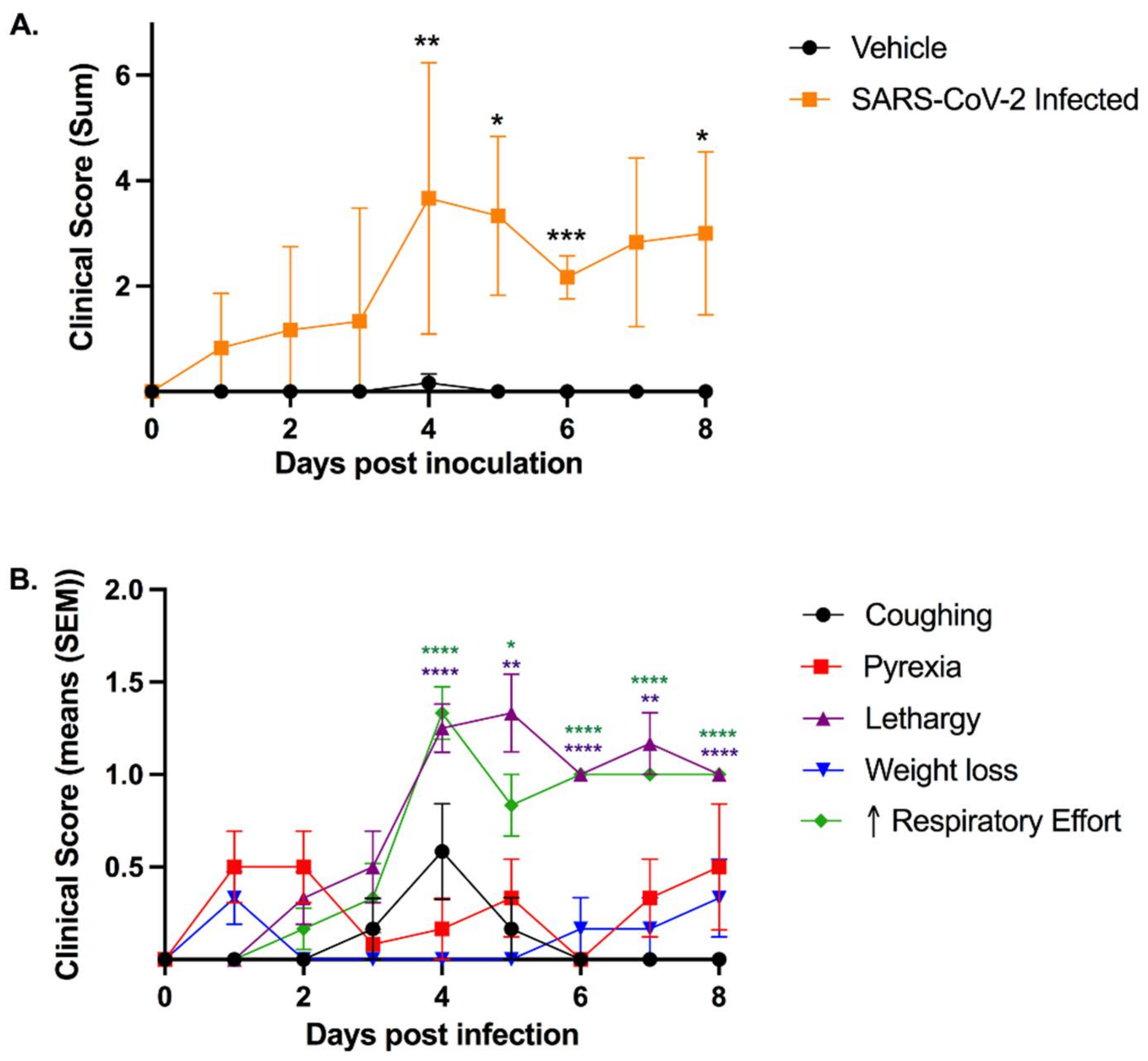
3.1. SARS-CoV-2-Infected Cats Exhibit Clinical Signs of Lower Respiratory Disease
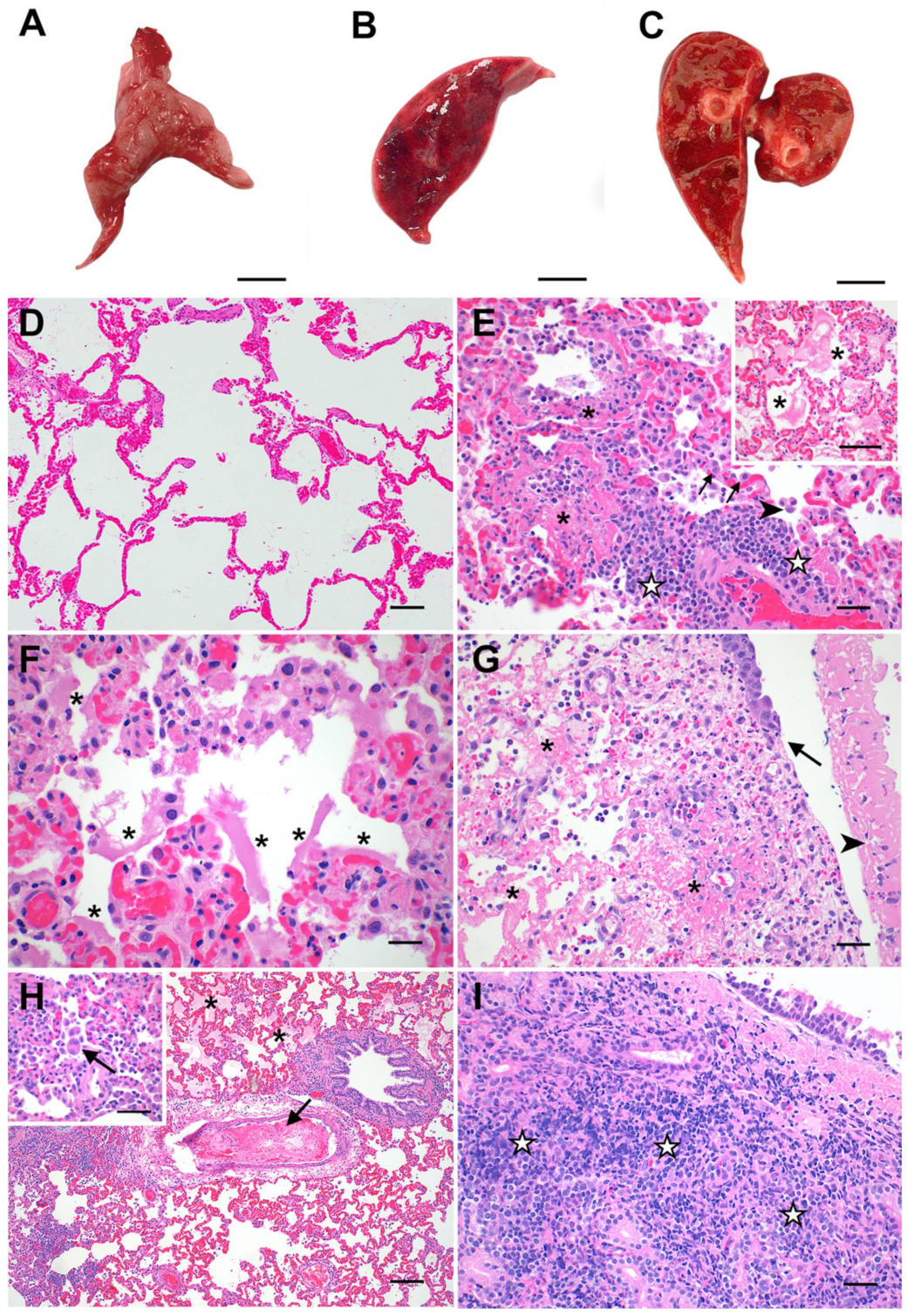
3.2. Feline SARS-CoV-2 Infection Pathology Mirrors Acute COVID-19
3.3. Viral Sequencing and Genome Assembly
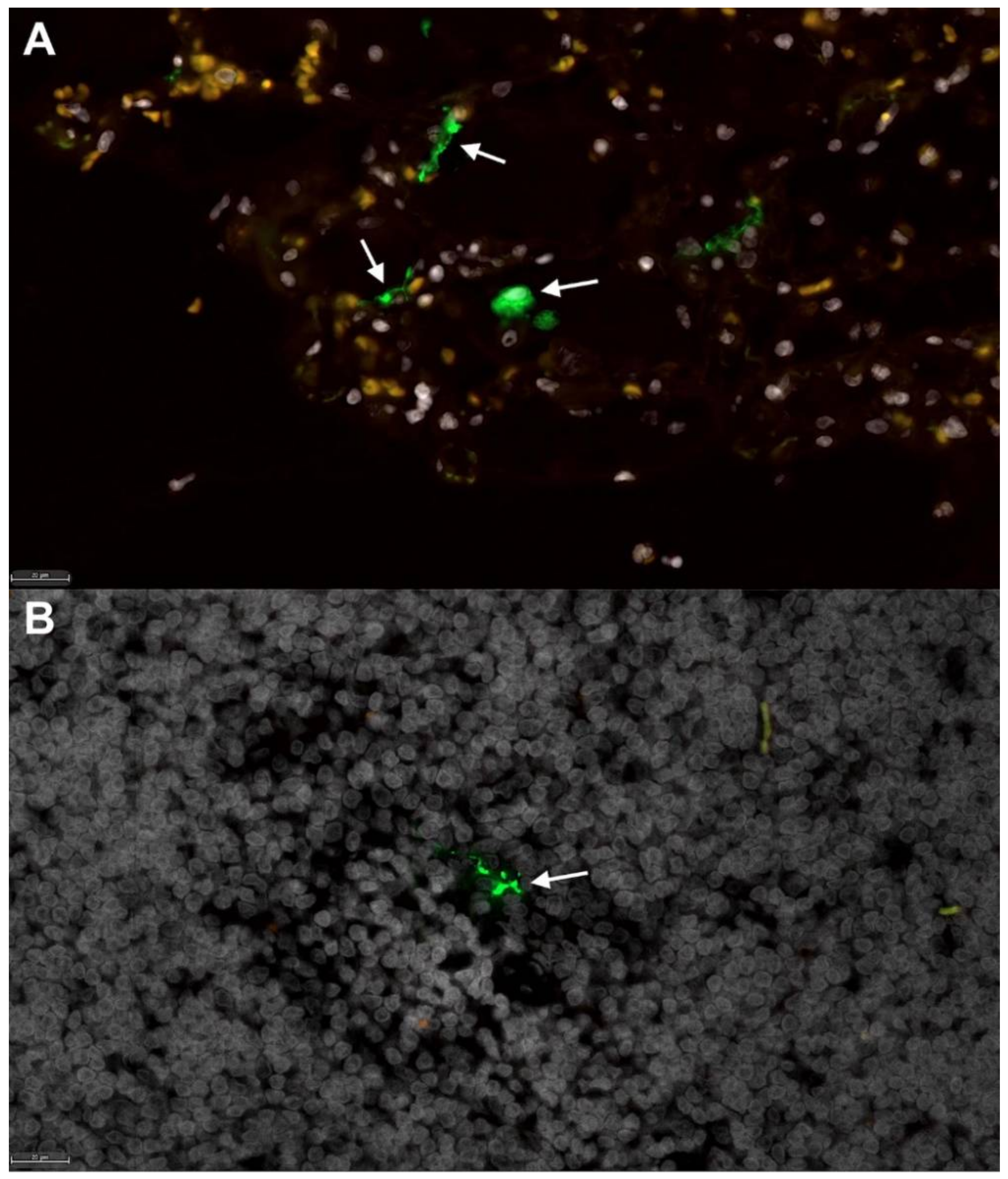
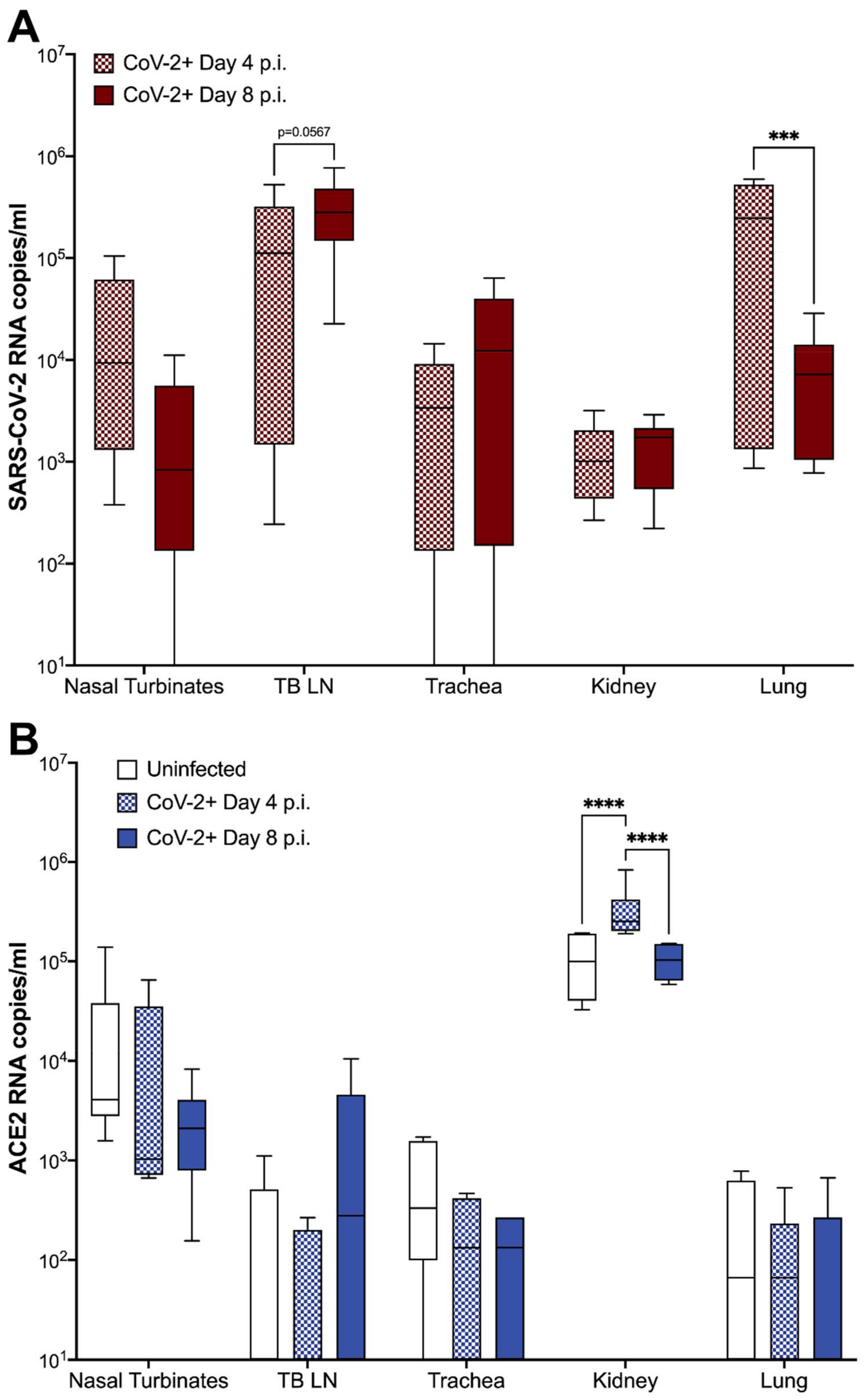
3.4. ACE2 Expression and Viral RNA in Feline Tissues during SARS-CoV-2 Infection
4. Discussion
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- WHO. World Health Organization Coronavirus (COVID-19) Dashboard. Available online: https://covid19.who.int (accessed on 8 July 2021).
- Wu, C.; Chen, X.; Cai, Y.; Zhou, X.; Xu, S.; Huang, H.; Zhang, L.; Zhou, X.; Du, C.; Zhang, Y. Risk factors associated with acute respiratory distress syndrome and death in patients with coronavirus disease 2019 pneumonia in Wuhan, China. JAMA Intern. Med. 2020, 80, 934–943. [Google Scholar] [CrossRef] [Green Version]
- Chen, G.; Wu, D.; Guo, W.; Cao, Y.; Huang, D.; Wang, H.; Wang, T.; Zhang, X.; Chen, H.; Yu, H.; et al. Clinical and immunologic features in severe and moderate Coronavirus Disease 2019. J. Clin. Investig. 2020, 130, 2620–2629. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Pedersen, S.F.; Ho, Y.-C. SARS-CoV-2: A storm is raging. J. Clin. Investig. 2020, 130, 2202–2205. [Google Scholar] [CrossRef] [PubMed]
- Kumar, S.; Yadav, P.K.; Srinivasan, R.; Perumal, N. Selection of animal models for COVID-19 research. VirusDisease 2020, 31, 453–458. [Google Scholar] [CrossRef]
- Gretebeck, L.M.; Subbarao, K. Animal models for SARS and MERS coronaviruses. Curr. Opin. Virol. 2015, 13, 123–129. [Google Scholar] [CrossRef] [PubMed]
- Hikmet, F.; Méar, L.; Edvinsson, Å.; Micke, P.; Uhlén, M.; Lindskog, C. The protein expression profile of ACE2 in human tissues. Mol. Syst. Biol. 2020, 16, e9610. [Google Scholar] [CrossRef]
- Kiros, M.; Andualem, H.; Kiros, T.; Hailemichael, W.; Getu, S.; Geteneh, A.; Alemu, D.; Abegaz, W.E. COVID-19 pandemic: Current knowledge about the role of pets and other animals in disease transmission. Virol. J. 2020, 17, 143. [Google Scholar] [CrossRef]
- Oreshkova, N.; Molenaar, R.J.; Vreman, S.; Harders, F.; Oude Munnink, B.B.; Hakze-van der Honing, R.W.; Gerhards, N.; Tolsma, P.; Bouwstra, R.; Sikkema, R.S.; et al. SARS-CoV-2 infection in farmed minks, the Netherlands, April and May 2020. Eurosurveillance 2020, 25, 2001005. [Google Scholar] [CrossRef]
- Newman, A.; Smith, D.; Ghai, R.R.; Wallace, R.M.; Torchetti, M.K.; Loiacono, C.; Murrell, L.S.; Carpenter, A.; Moroff, S.; Rooney, J.A.; et al. First reported cases of SARS-CoV-2 infection in companion animals—New York, March–April 2020. Morb. Mortal. Wkly. Rep. 2020, 69, 710–713. [Google Scholar] [CrossRef]
- Molenaar, R.J.; Vreman, S.; Hakze-van der Honing, R.W.; Zwart, R.; de Rond, J.; Weesendorp, E.; Smit, L.A.M.; Koopmans, M.; Bouwstra, R.; Stegeman, A.; et al. Clinical and Pathological Findings in SARS-CoV-2 Disease Outbreaks in Farmed Mink (Neovison vison). Vet. Pathol. 2020, 57, 653–657. [Google Scholar] [CrossRef]
- Luan, J.; Lu, Y.; Jin, X.; Zhang, L. Spike protein recognition of mammalian ACE2 predicts the host range and an optimized ACE2 for SARS-CoV-2 infection. Biochem. Biophys. Res. Commun. 2020, 526, 165–169. [Google Scholar] [CrossRef]
- Liu, Y.; Hu, G.; Wang, Y.; Ren, W.; Zhao, X.; Ji, F.; Zhu, Y.; Feng, F.; Gong, M.; Ju, X.; et al. Functional and genetic analysis of viral receptor ACE2 orthologs reveals a broad potential host range of SARS-CoV-2. Proc. Natl. Acad. Sci. USA 2021, 118, e2025373118. [Google Scholar] [CrossRef]
- Hayashi, T.; Abiko, K.; Mandai, M.; Yaegashi, N.; Konishi, I. Highly conserved binding region of ACE2 as a receptor for SARS-CoV-2 between humans and mammals. Vet. Q. 2020, 40, 243–249. [Google Scholar] [CrossRef] [PubMed]
- McAloose, D.; Laverack, M.; Wang, L.; Killian, M.L.; Caserta, L.C.; Yuan, F.; Mitchell, P.K.; Queen, K.; Mauldin, M.R.; Cronk, B.D.; et al. From people to panthera: Natural SARS-CoV-2 infection in tigers and lions at the Bronx Zoo. mBio 2020, 11, e02220-20. [Google Scholar] [CrossRef] [PubMed]
- Shi, J.; Wen, Z.; Zhong, G.; Yang, H.; Wang, C.; Huang, B.; Liu, R.; He, X.; Shuai, L.; Sun, Z.; et al. Susceptibility of ferrets, cats, dogs, and other domesticated animals to SARS-coronavirus 2. Science 2020, 358, 1016–1020. [Google Scholar] [CrossRef] [Green Version]
- Zhang, Q.; Zhang, H.; Gao, J.; Huang, K.; Yang, Y.; Hui, X.; He, X.; Li, C.; Gong, W.; Zhang, Y.; et al. A serological survey of SARS-CoV-2 in cat in Wuhan. Emerg. Microbes Infect. 2020, 9, 2013–2019. [Google Scholar] [CrossRef] [PubMed]
- Lakdawala, S.S.; Menachery, V.D. The search for a COVID-19 animal model. Science 2020, 368, 942–943. [Google Scholar] [CrossRef] [PubMed]
- Hernández, M.; Abad, D.; Eiros, J.M.; Rodríguez-Lázaro, D. Are animals a neglected transmission route of SARS-CoV-2? Pathogens 2020, 9, 480. [Google Scholar] [CrossRef]
- Van de Velde, H.; Janssens, G.P.; de Rooster, H.; Polis, I.; Peters, I.; Ducatelle, R.; Nguyen, P.; Buyse, J.; Rochus, K.; Xu, J.; et al. The cat as a model for human obesity: Insights into depot-specific inflammation associated with feline obesity. Br. J. Nutr. 2013, 110, 1326–1335. [Google Scholar] [CrossRef] [Green Version]
- Nelson, R.W.; Reusch, C.E. Animal models of disease: Classification and etiology of diabetes in dogs and cats. J. Endocrinol. 2014, 222, T1–T9. [Google Scholar] [CrossRef] [Green Version]
- Samaha, G.; Beatty, J.; Wade, C.M.; Haase, B. The Burmese cat as a genetic model of type 2 diabetes in humans. Anim. Genet. 2019, 50, 319–325. [Google Scholar] [CrossRef]
- Hoenig, M. The cat as a model for human obesity and diabetes. J. Diabetes Sci. Technol. 2012, 6, 525–533. [Google Scholar] [CrossRef] [Green Version]
- Wallner, M.; Eaton, D.M.; Berretta, R.M.; Borghetti, G.; Wu, J.; Baker, S.T.; Feldsott, E.A.; Sharp, T.E., 3rd; Mohsin, S.; Oyama, M.A.; et al. A Feline HFpEF Model with Pulmonary Hypertension and Compromised Pulmonary Function. Sci. Rep. 2017, 7, 16587. [Google Scholar] [CrossRef] [PubMed]
- Prat, V.; Rozec, B.; Gauthier, C.; Lauzier, B. Human heart failure with preserved ejection versus feline cardiomyopathy: What can we learn from both veterinary and human medicine? Heart Fail. Rev. 2017, 22, 783–794. [Google Scholar] [CrossRef]
- Gaudreault, N.N.; Trujillo, J.D.; Carossino, M.; Meekins, D.A.; Morozov, I.; Madden, D.W.; Indran, S.V.; Bold, D.; Balaraman, V.; Kwon, T.; et al. SARS-CoV-2 infection, disease and transmission in domestic cats. Emerg. Microbes Infect. 2020, 9, 2322–2332. [Google Scholar] [CrossRef] [PubMed]
- Bosco-Lauth, A.M.; Hartwig, A.E.; Porter, S.M.; Gordy, P.W.; Nehring, M.; Byas, A.D.; VandeWoude, S.; Ragan, I.K.; Maison, R.M.; Bowen, R.A. Experimental infection of domestic dogs and cats with SARS-CoV-2: Pathogenesis, transmission, and response to reexposure in cats. Proc. Natl. Acad. Sci. USA 2020, 117, 26382–26388. [Google Scholar] [CrossRef]
- Gaudreault, N.N.; Carossino, M.; Morozov, I.; Trujillo, J.D.; Meekins, D.A.; Madden, D.W.; Cool, K.; Artiaga, B.L.; McDowell, C.; Bold, D.; et al. Experimental re-infected cats do not transmit SARS-CoV-2. Emerg. Microbes Infect. 2021, 10, 638–650. [Google Scholar] [CrossRef] [PubMed]
- Van den Brand, J.M.; Haagmans, B.L.; Leijten, L.; van Riel, D.; Martina, B.E.; Osterhaus, A.D.; Kuiken, T. Pathology of experimental SARS coronavirus infection in cats and ferrets. Vet. Pathol. 2008, 45, 551–562. [Google Scholar] [CrossRef] [Green Version]
- Van den Brand, J.M.; Haagmans, B.L.; van Riel, D.; Osterhaus, A.D.; Kuiken, T. The pathology and pathogenesis of experimental severe acute respiratory syndrome and influenza in animal models. J. Comp. Pathol. 2014, 151, 83–112. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Coleman, C.M.; Frieman, M.B. Growth and quantification of MERS-CoV infection. Curr. Protoc. Microbiol. 2015, 37, 1521–1529. [Google Scholar] [CrossRef] [PubMed]
- Reed, L.J.; Muench, H. A simple method of estimating fifty per cent endpoints. Am. J. Epidemiol. 1938, 27, 493–497. [Google Scholar] [CrossRef]
- Ritchey, J.W.; Levy, J.K.; Bliss, S.K.; Tompkins, W.A.; Tompkins, M.B. Constitutive expression of types 1 and 2 cytokines by alveolar macrophages from feline immunodeficiency virus-infected cats. Vet. Immunol. Immunopathol. 2001, 79, 83–100. [Google Scholar] [CrossRef]
- Polak, S.B.; Van Gool, I.C.; Cohen, D.; Jan, H.; van Paassen, J. A systematic review of pathological findings in COVID-19: A pathophysiological timeline and possible mechanisms of disease progression. Mod. Pathol. 2020, 33, 2128–2138. [Google Scholar] [CrossRef] [PubMed]
- Tian, S.; Xiong, Y.; Liu, H.; Niu, L.; Guo, J.; Liao, M.; Xiao, S.-Y. Pathological study of the 2019 novel coronavirus disease (COVID-19) through postmortem core biopsies. Mod. Pathol. 2020, 33, 1007–1014. [Google Scholar] [CrossRef] [Green Version]
- Von der Thüsen, J.; van der Eerden, M. Histopathology and genetic susceptibility in COVID-19 pneumonia. Eur. J. Clin. Investig. 2020, 50, e13259. [Google Scholar] [CrossRef]
- Xu, Z.; Shi, L.; Wang, Y.; Zhang, J.; Huang, L.; Zhang, C.; Liu, S.; Zhao, P.; Liu, H.; Zhu, L.; et al. Pathological findings of COVID-19 associated with acute respiratory distress syndrome. Lancet Respir. Med. 2020, 8, 420–422. [Google Scholar] [CrossRef]
- Miller, C.; Bielefeldt-Ohmann, H.; MacMillan, M.; Huitron-Resendiz, S.; Henriksen, S.; Elder, J.; VandeWoude, S. Strain-specific viral distribution and neuropathology of feline immunodeficiency virus. Vet. Immunol. Immunopathol. 2011, 143, 282–291. [Google Scholar] [CrossRef] [Green Version]
- Miller, C.; Boegler, K.; Carver, S.; MacMillan, M.; Bielefeldt-Ohmann, H.; VandeWoude, S. Pathogenesis of oral FIV infection. PLoS ONE 2017, 12, e0185138. [Google Scholar] [CrossRef] [Green Version]
- Deiana, M.; Mori, A.; Piubelli, C.; Scarso, S.; Favarato, M.; Pomari, E. Assessment of the direct quantitation of SARS-CoV-2 by droplet digital PCR. Sci. Rep. 2020, 10, 18764. [Google Scholar] [CrossRef] [PubMed]
- O’Driscoll, A.; Belogrudov, V.; Carroll, J.; Kropp, K.; Walsh, P.; Ghazal, P.; Sleator, R.D. HBLAST: Parallelised sequence similarity—A Hadoop MapReducable basic local alignment search tool. J. Biomed. Inf. 2015, 54, 58–64. [Google Scholar] [CrossRef]
- Lu, J.; du Plessis, L.; Liu, Z.; Hill, V.; Kang, M.; Lin, H.; Sun, J.; François, S.; Kraemer, M.U.G.; Faria, N.R.; et al. Genomic epidemiology of SARS-CoV-2 in Guangdong province, China. Cell 2020, 181, 997–1003. [Google Scholar] [CrossRef]
- Koren, S.; Walenz, B.P.; Berlin, K.; Miller, J.R.; Bergman, N.H.; Phillippy, A.M. Canu: Scalable and accurate long-read assembly via adaptive k-mer weighting and repeat separation. Genome Res. 2017, 27, 722–736. [Google Scholar] [CrossRef] [Green Version]
- Danecek, P.; Bonfield, J.K.; Liddle, J.; Marshall, J.; Ohan, V.; Pollard, M.O.; Whitwham, A.; Keane, T.; McCarthy, S.A.; Davies, R.M.; et al. Twelve years of SAMtools and BCFtools. Gigascience 2021, 10, giab008. [Google Scholar] [CrossRef]
- Li, H. Minimap2: Pairwise alignment for nucleotide sequences. Bioinformatics 2018, 34, 3094–3100. [Google Scholar] [CrossRef] [PubMed]
- Loman, N.J.; Quick, J.; Simpson, J.T. A complete bacterial genome assembled de novo using only nanopore sequencing data. Nat. Methods 2015, 12, 733–735. [Google Scholar] [CrossRef] [PubMed]
- Huet, O.; Ramsey, D.; Miljavec, S.; Jenney, A.; Aubron, C.; Aprico, A.; Stefanovic, N.; Balkau, B.; Head, G.A.; de Haan, J.B.; et al. Ensuring animal welfare while meeting scientific aims using a murine pneumonia model of septic shock. Shock 2013, 39, 488–494. [Google Scholar] [CrossRef] [PubMed]
- Hartmann, A.D.; Helps, C.R.; Lappin, M.R.; Werckenthin, C.; Hartmann, K. Efficacy of pradofloxacin in cats with feline upper respiratory tract disease due to Chlamydophila felis or Mycoplasma infections. J. Vet. Intern. Med. 2008, 22, 44–52. [Google Scholar] [CrossRef]
- Steagall, P.V.; Monteiro, B.P. Acute pain in cats: Recent advances in clinical assessment. J. Feline Med. Surg. 2019, 21, 25–34. [Google Scholar] [CrossRef]
- Blair, R.V.; Vaccari, M.; Doyle-Meyers, L.A.; Roy, C.J.; Russell-Lodrigue, K.; Fahlberg, M.; Monjure, C.J.; Beddingfield, B.; Plante, K.S.; Plante, J.A. Acute respiratory distress in aged, SARS-CoV-2–infected African green monkeys but not rhesus macaques. Am. J. Pathol. 2021, 191, 274–282. [Google Scholar] [CrossRef] [PubMed]
- Bodewes, R.; Kreijtz, J.H.; van Amerongen, G.; Fouchier, R.A.; Osterhaus, A.D.; Rimmelzwaan, G.F.; Kuiken, T. Pathogenesis of Influenza A/H5N1 virus infection in ferrets differs between intranasal and intratracheal routes of inoculation. Am. J. Pathol. 2011, 179, 30–36. [Google Scholar] [CrossRef]
- Rimmelzwaan, G.F.; van Riel, D.; Baars, M.; Bestebroer, T.M.; van Amerongen, G.; Fouchier, R.A.; Osterhaus, A.D.; Kuiken, T. Influenza A virus (H5N1) infection in cats causes systemic disease with potential novel routes of virus spread within and between hosts. Am. J. Pathol. 2006, 168, 176–183. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Macdonald, E.S.; Norris, C.R.; Berghaus, R.B.; Griffey, S.M. Clinicopathologic and radiographic features and etiologic agents in cats with histologically confirmed infectious pneumonia: 39 cases (1991–2000). J. Am. Vet. Med. Assoc. 2003, 223, 1142–1150. [Google Scholar] [CrossRef] [PubMed]
- Wiersinga, W.J.; Rhodes, A.; Cheng, A.C.; Peacock, S.J.; Prescott, H.C. Pathophysiology, transmission, diagnosis, and treatment of coronavirus disease 2019 (COVID-19): A Review. JAMA 2020, 324, 782–793. [Google Scholar] [CrossRef] [PubMed]
- Fox, S.E.; Akmatbekov, A.; Harbert, J.L.; Li, G.; Quincy Brown, J.; Vander Heide, R.S. Pulmonary and cardiac pathology in African American patients with COVID-19: An autopsy series from New Orleans. Lancet Respir. Med. 2020, 8, 681–686. [Google Scholar] [CrossRef]
- Borczuk, A.C.; Salvatore, S.P.; Seshan, S.V.; Patel, S.S.; Bussel, J.B.; Mostyka, M.; Elsoukkary, S.; He, B.; Del Vecchio, C.; Fortarezza, F.; et al. COVID-19 pulmonary pathology: A multi-institutional autopsy cohort from Italy and New York City. Mod. Pathol. 2020, 33, 2156–2168. [Google Scholar] [CrossRef]
| Clinical Parameter | 0 (Healthy) | 1 | 2 |
|---|---|---|---|
| Body Weight | No weight loss | 0 to 5% weight loss | >5% weight loss |
| Temperature | 37.2 to 39.1 °C | 39.2 to 39.7 °C | >39.7 °C |
| Respiratory Effort | Normal resting respiratory rate and normal effort | Mild tachypnea, but no overt increase in effort | Marked increase in both respiratory rate and effort; dyspnea |
| Activity | Normal | Reduced activity when disturbed * (lethargy) | Little to no activity disturbed *; reduced activity stimulated ** |
| Behavior | Normal | Reduced interest in food and/or attention | Anorexia and lack of interest |
| Ocular/Nasal Discharge | None | Mild discharge noted | Discharge evident from both nasal and ocular regions |
| Coughing/Wheezing | None | Mild wheezing, but no coughing | Coughing and/or marked wheezing |
| Nucleotide Change | Gene | Amino Acid Shift |
|---|---|---|
| T8782C | ORF1ab | S2839S |
| T18060C | ORF1ab | L5932L |
| G22296A | S gene | R245H |
| C28144T | ORF8 | S251L |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Rudd, J.M.; Tamil Selvan, M.; Cowan, S.; Kao, Y.-F.; Midkiff, C.C.; Narayanan, S.; Ramachandran, A.; Ritchey, J.W.; Miller, C.A. Clinical and Histopathologic Features of a Feline SARS-CoV-2 Infection Model Are Analogous to Acute COVID-19 in Humans. Viruses 2021, 13, 1550. https://doi.org/10.3390/v13081550
Rudd JM, Tamil Selvan M, Cowan S, Kao Y-F, Midkiff CC, Narayanan S, Ramachandran A, Ritchey JW, Miller CA. Clinical and Histopathologic Features of a Feline SARS-CoV-2 Infection Model Are Analogous to Acute COVID-19 in Humans. Viruses. 2021; 13(8):1550. https://doi.org/10.3390/v13081550
Chicago/Turabian StyleRudd, Jennifer M., Miruthula Tamil Selvan, Shannon Cowan, Yun-Fan Kao, Cecily C. Midkiff, Sai Narayanan, Akhilesh Ramachandran, Jerry W. Ritchey, and Craig A. Miller. 2021. "Clinical and Histopathologic Features of a Feline SARS-CoV-2 Infection Model Are Analogous to Acute COVID-19 in Humans" Viruses 13, no. 8: 1550. https://doi.org/10.3390/v13081550
APA StyleRudd, J. M., Tamil Selvan, M., Cowan, S., Kao, Y.-F., Midkiff, C. C., Narayanan, S., Ramachandran, A., Ritchey, J. W., & Miller, C. A. (2021). Clinical and Histopathologic Features of a Feline SARS-CoV-2 Infection Model Are Analogous to Acute COVID-19 in Humans. Viruses, 13(8), 1550. https://doi.org/10.3390/v13081550







