Evaluation of Dried Blood Spot Testing for SARS-CoV-2 Serology Using a Quantitative Commercial Assay
Abstract
1. Introduction
2. Materials and Methods
2.1. Specimens
2.2. Dried Blood Spots
2.3. SARS-CoV-2 Antibody Assay
2.4. Data Analysis
3. Results
3.1. Initial Validation of Quantitative Roche S Assay Using Prepared DBS Specimens
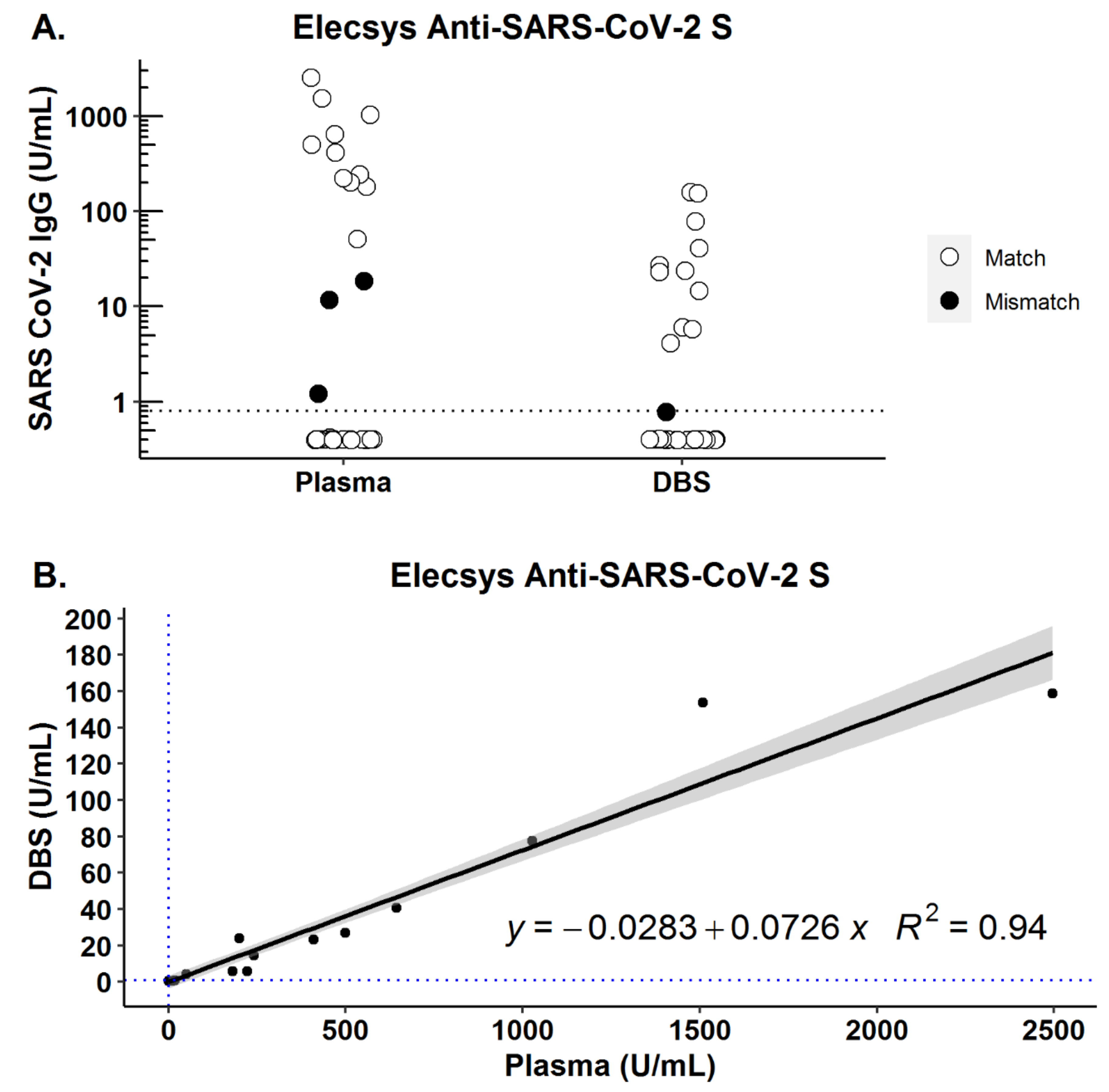
3.2. Performance of Quantitative Roche S Assay in Matched Plasma/DBS Fingerprick Collection
4. Discussion
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Acknowledgments
Conflicts of Interest
References
- Bailey, D.; Konforte, D.; Barakauskas, V.E.; Yip, P.M.; Kulasingam, V.; Abou, E.H.M.; Beach, L.A.; Blasutig, I.M.; Catomeris, P.; Dooley, K.C.; et al. Canadian Society of Clinical Chemists (CSCC) Interim Consensus Guidance for Testing and Reporting of SARS-CoV-2 Serology. Clin. Biochem. 2020, 86, 1–7. [Google Scholar] [CrossRef] [PubMed]
- Krishnamurthy, H.K.; Jayaraman, V.; Krishna, K.; Rajasekaran, K.E.; Wang, T.; Bei, K.; Rajasekaran, J.J.; Yaskin, I.; Rai, A.J.; Choung, R.S.; et al. Antibody Profiling and Prevalence in US Patients during the SARS-CoV2 Pandemic. PLoS ONE 2020, 15, e0242655. [Google Scholar] [CrossRef]
- Protti, M.; Mandrioli, R.; Mercolini, L. Quantitative Microsampling for Bioanalytical Applications Related to the SARS-CoV-2 Pandemic: Usefulness, Benefits and Pitfalls. J. Pharm. Biomed. Anal. 2020, 191, 113597. [Google Scholar] [CrossRef] [PubMed]
- Dimech, W.; Mulders, M.N. A Review of Testing Used in Seroprevalence Studies on Measles and Rubella. Vaccine 2016, 34, 4119–4122. [Google Scholar] [CrossRef]
- Villar, L.M.; Bezerra, C.S.; Cruz, H.M.; Portilho, M.M.; Flores, G.L. Applicability of Oral Fluid and Dried Blood Spot for Hepatitis B Virus Diagnosis. Can. J. Gastroenterol. Hepatol. 2019, 2019, 5672795. [Google Scholar] [CrossRef]
- Villar, L.M.; de Oliveira, J.C.; Cruz, H.M.; Yoshida, C.F.T.; Lampe, E.; Lewis-Ximenez, L.L. Assessment of Dried Blood Spot Samples as a Simple Method for Detection of Hepatitis B Virus Markers. J. Med. Virol. 2011, 83, 1522–1529. [Google Scholar] [CrossRef] [PubMed]
- Flores, G.L.; Cruz, H.M.; Miguel, J.C.; Potsch, D.V.; Pilotto, J.H.; Lewis-Ximenez, L.L.; Lampe, E.; Villar, L.M. Assessing Hepatitis B Immunity Using Dried Blood Spot Samples from HIV+ Individuals. J. Med. Virol. 2018, 90, 1863–1867. [Google Scholar] [CrossRef] [PubMed]
- Croom, H.A.; Richards, K.M.; Best, S.J.; Francis, B.H.; Johnson, E.I.; Dax, E.M.; Wilson, K.M. Commercial Enzyme Immunoassay Adapted for the Detection of Antibodies to Hepatitis C Virus in Dried Blood Spots. J. Clin. Virol. 2006, 36, 68–71. [Google Scholar] [CrossRef]
- Mercader, S.; Featherstone, D.; Bellini, W.J. Comparison of Available Methods to Elute Serum from Dried Blood Spot Samples for Measles Serology. J. Virol. Methods 2006, 137, 140–149. [Google Scholar] [CrossRef]
- Cruz, H.M.; Cruz, J.C.M.; Da Silva, E.F.; Portilho, M.M.; Marques, V.A.; Lewis-Ximenez, L.L.; Lampe, E.; Villar, L.M. Comparison of the Performance of Enzyme Immunoassays for Hepatitis B and C Detection in Dried Blood Spot. J. Immunoass. Immunochem. 2018, 39, 228–233. [Google Scholar] [CrossRef]
- Villar, L.M.; Cruz, H.M.; Deodato, R.M.; Miguel, J.C.; Da Silva, E.F.; Flores, G.L.; Lewis-Ximenez, L.L. Usefulness of Automated Assays for Detecting Hepatitis B and C Markers in Dried Blood Spot Samples. BMC Res. Notes 2019, 12, 523. [Google Scholar] [CrossRef] [PubMed]
- Ross, R.S.; Stambouli, O.; Grüner, N.; Marcus, U.; Cai, W.; Zhang, W.; Zimmermann, R.; Roggendorf, M. Detection of Infections with Hepatitis B Virus, Hepatitis C Virus, and Human Immunodeficiency Virus by Analyses of Dried Blood Spots--Performance Characteristics of the ARCHITECT System and Two Commercial Assays for Nucleic Acid Amplification. Virol. J. 2013, 10, 72. [Google Scholar] [CrossRef]
- Lee, C.E.; Ponnampalavanar, S.S.; Omar, S.F.S.; Mahadeva, S.; Ong, L.Y.; Kamarulzaman, A. Evaluation of the Dried Blood Spot (DBS) Collection Method as a Tool for Detection of HIV Ag/Ab, HBsAg, Anti-HBs and Anti-HCV in a Malaysian Tertiary Referral Hospital. Ann. Acad. Med. Singap. 2011, 40, 448–453. [Google Scholar] [PubMed]
- Brandão, C.P.U.; Marques, B.L.C.; Marques, V.A.; Villela-Nogueira, C.A.; Do Ó, K.M.R.; de Paula, M.T.; Lewis-Ximenez, L.L.; Lampe, E.; Sá Ferreira, J.A.; Villar, L.M. Simultaneous Detection of Hepatitis C Virus Antigen and Antibodies in Dried Blood Spots. J. Clin. Virol. 2013, 57, 98–102. [Google Scholar] [CrossRef][Green Version]
- Tuaillon, E.; Kania, D.; Pisoni, A.; Bollore, K.; Taieb, F.; Ngoyi, E.N.O.; Schaub, R.; Plantier, J.-C.; Makinson, A.; van de Perre, P. Dried Blood Spot Tests for the Diagnosis and Therapeutic Monitoring of HIV and Viral Hepatitis B and C. Front. Microbiol. 2020, 11, 373. [Google Scholar] [CrossRef] [PubMed]
- Valentine-Graves, M.; Hall, E.; Guest, J.L.; Adam, E.; Valencia, R.; Shinn, K.; Hardee, I.; Sanchez, T.; Siegler, A.J.; Sullivan, P.S. At-home Self-collection of Saliva, Oropharyngeal Swabs and Dried Blood Spots for SARS-CoV-2 Diagnosis and Serology: Post-collection Acceptability of Specimen Collection Process and Patient Confidence in Specimens. PLoS ONE 2020, 15, e0236775. [Google Scholar] [CrossRef]
- Morley, G.L.; Taylor, S.; Jossi, S.; Perez-Toledo, M.; Faustini, S.E.; Marcial-Juarez, E.; Shields, A.M.; Goodall, M.; Allen, J.D.; Watanabe, Y.; et al. Sensitive Detection of SARS-CoV-2-Specific Antibodies in Dried Blood Spot Samples. Emerg. Infect. Dis. 2020, 26, 2970–2973. [Google Scholar] [CrossRef] [PubMed]
- Karp, D.G.; Danh, K.; Espinoza, N.F.; Seftel, D.; Robinson, P.V.; Tsai, C.-T. A Serological Assay to Detect SARS-CoV-2 Antibodies in At-home Collected Finger-prick Dried Blood Spots. Sci. Rep. 2020, 10, 20188. [Google Scholar] [CrossRef]
- Sullivan, P.S.; Sailey, C.; Guest, J.L.; Guarner, J.; Kelley, C.; Siegler, A.J.; Valentine-Graves, M.; Gravens, L.; Del Rio, C.; Sanchez, T.H. Detection of SARS-CoV-2 RNA and Antibodies in Diverse Samples: Protocol to Validate the Sufficiency of Provider-Observed, Home-Collected Blood, Saliva, and Oropharyngeal Samples. JMIR Public Health Surveill. 2020, 6, e19054. [Google Scholar] [CrossRef] [PubMed]
- Amendola, A.; Bianchi, S.; Gori, M.; Barcellini, L.; Colzani, D.; Canuti, M.; Giacomet, V.; Fabiano, V.; Folgori, L.; Zuccotti, G.V.; et al. Back to school: Use of Dried Blood Spot for the Detection of SARS-CoV-2-specific Immunoglobulin G (IgG) among Schoolchildren in Milan, Italy. medRxiv 2020. [Google Scholar] [CrossRef]
- Amendola, A.; Bianchi, S.; Gori, M.; Barcellini, L.; Colzani, D.; Canuti, M.; Giacomet, V.; Fabiano, V.; Folgori, L.; Zuccotti, G.V.; et al. Dried Blood Spot as an Alternative to Plasma/Serum for SARS-CoV-2 IgG Detection, an Opportunity to Be Sized to Facilitate COVID-19 Surveillance Among Schoolchildren. Pediatr. Infect. Dis. J. 2021, 40, e46–e47. [Google Scholar] [CrossRef]
- Demonbreun, A.R.; McDade, T.W.; Pesce, L.; Vaught, L.A.; Reiser, N.L.; Bogdanovic, E.; Velez, M.E.; Hsieh, R.R.; Klaisner, C.-N.; Simons, L.M.; et al. Patterns and Persistence of SARS-CoV-2 IgG Antibodies in a US Metropolitan Site. medRxiv 2020. [Google Scholar] [CrossRef]
- Gaugler, S.; Sottas, P.-E.; Blum, K.; Luginbühl, M. Fully Automated Dried Blood Spot Sample Handling and Extraction for Serological Testing of SARS-CoV-2 Antibodies. Drug Test. Anal. 2021, 13, 223–226. [Google Scholar] [CrossRef] [PubMed]
- Iyer, A.S.; Jones, F.K.; Nodoushani, A.; Kelly, M.; Becker, M.; Slater, D.; Mills, R.; Teng, E.; Kamruzzaman, M.; Garcia-Beltran, W.F.; et al. Persistence and Decay of Human Antibody Responses to the Receptor Binding Domain of SARS-CoV-2 Spike Protein in COVID-19 Patients. Sci. Immunol. 2020, 5. [Google Scholar] [CrossRef] [PubMed]
- Liu, F.; Nguyen, M.; Vijayakumar, P.; Kaplan, A.; Meir, A.; Dai, Y.; Wang, E.; Walsh, H.; Ring, A.M.; Omer, S.B.; et al. Newborn Dried Blood Spots for Serologic Surveys of COVID-19. Pediatr. Infect. Dis. J. 2020, 39, e454–e456. [Google Scholar] [CrossRef] [PubMed]
- Moat, S.J.; Zelek, W.M.; Carne, E.; Ponsford, M.J.; Bramhall, K.; Jones, S.; El-Shanawany, T.; Wise, M.P.; Thomas, A.; George, C.; et al. Development of a High-throughput SARS-CoV-2 Antibody Testing Pathway Using Dried Blood Spot Specimens. Ann. Clin. Biochem. 2020, 58. [Google Scholar] [CrossRef]
- Thevis, M.; Knoop, A.; Schaefer, M.S.; Dufaux, B.; Schrader, Y.; Thomas, A.; Geyer, H. Can Dried Blood Spots (DBS) Contribute to Conducting Comprehensive SARS-CoV-2 Antibody Tests? Drug Test. Anal. 2020, 12, 994–997. [Google Scholar] [CrossRef]
- Zava, T.T.; Zava, D.T. Validation of Dried Blood Spot Sample Modifications to Two Commercially Available COVID-19 IgG Antibody Immunoassays. Bioanalysis 2021, 13, 13–28. [Google Scholar] [CrossRef]
- Mulchandani, R.; Brown, B.; Brooks, T.; Semper, A.; Machin, N.; Linley, E.; Borrow, R.; Wyllie, D.; Taylor-Philips, S.; Jones, H.; et al. Use of Dried Blood Spot Samples for SARS-CoV-2 Antibody Detection Using the Roche Elecsys® High throughput Immunoassay. J. Clin. Virol. 2021, 136, 104739. [Google Scholar] [CrossRef] [PubMed]
- McDade, T.W.; McNally, E.M.; Zelikovich, A.S.; D’Aquila, R.; Mustanski, B.; Miller, A.; Vaught, L.A.; Reiser, N.L.; Bogdanovic, E.; Fallon, K.S.; et al. High Seroprevalence for SARS-CoV-2 among Household Members of Essential Workers Detected Using a Dried Blood Spot Assay. PLoS ONE 2020, 15, e0237833. [Google Scholar] [CrossRef]
- McDade, T.W.; McNally, E.M.; D’Aquila, R.; Mustanski, B.; Miller, A.; Vaught, L.A.; Reiser, N.L.; Bogdanovic, E.; Zelikovich, A.S.; Demonbreun, A.R. Enzyme Immunoassay for SARS-CoV-2 Antibodies in Dried Blood Spot Samples: A Minimally-Invasive Approach to Facilitate Community- and Population-based Screening. medRxiv 2020. [Google Scholar] [CrossRef]
- Anderson, M.; Holzmayer, V.; Vallari, A.; Taylor, R.; Moy, J.; Cloherty, G. Expanding Access to SARS-CoV-2 IgG and IgM Serologic Testing Using Fingerstick Whole Blood, Plasma, and Rapid Lateral Flow Assays. J. Clin. Virol. 2021, 104855. [Google Scholar] [CrossRef]
- Feld, J.J.; Kandel, C.; Biondi, M.J.; A Kozak, R.; Zahoor, M.A.; Lemieux, C.; Borgia, S.M.; Boggild, A.K.; Powis, J.; McCready, J.; et al. Peginterferon Lambda for the Treatment of Outpatients with COVID-19: A Phase 2, Placebo-controlled Randomised Trial. Lancet Respir. Med. 2021, 9, 498–510. [Google Scholar] [CrossRef]
- Grüner, N.; Stambouli, O.; Ross, R.S. Dried Blood Spots-Preparing and Processing for Use in Immunoassays and in Molecular Techniques. J. Vis. Exp. 2015, e52619. [Google Scholar] [CrossRef]
- Biondi, M.J.; Van Tilborg, M.; Smookler, D.; Heymann, G.; Aquino, A.; Perusini, S.; Mandel, E.; Kozak, R.A.; Cherepanov, V.; Kowgier, M.; et al. Hepatitis C Core-Antigen Testing from Dried Blood Spots. Viruses 2019, 11, 830. [Google Scholar] [CrossRef] [PubMed]
- R Core Team. R: A Language and Environment for Statistical Computing. R Foundation for Statistical Computing; R Core Team: Vienna, Austria, 2013; Available online: https://www.R-project.org/ (accessed on 19 May 2021).
- Wellinghausen, N.; Voss, M.; Ivanova, R.; Deininger, S. Evaluation of the SARS-CoV-2-IgG Response in Outpatients by Five Commercial Immunoassays. GMS Infect. Dis. 2020, 8, Doc22. [Google Scholar] [PubMed]
- Weber, M.C.; Risch, M.; Thiel, S.L.; Grossmann, K.; Nigg, S.; Wohlwend, N.; Lung, T.; Hillmann, D.; Ritzler, M.; Ferrara, F.; et al. Characteristics of Three Different Chemiluminescence Assays for Testing for SARS-CoV-2 Antibodies. Dis. Markers. Res. Artic. Hindawi 2021. [Google Scholar] [CrossRef] [PubMed]
- Catry, E.; Jacqmin, H.; Dodemont, M.; Saad Albichr, I.; Lardinois, B.; de Fays, B.; Delaere, B.; Closset, M.; Laurent, T.; Denis, O.; et al. Analytical and Clinical Evaluation of Four Commercial SARS-CoV-2 Serological Immunoassays in Hospitalized Patients and Ambulatory Individuals. J. Virol. Methods 2021, 289, 114060. [Google Scholar] [CrossRef] [PubMed]
- Turbett, S.E.; Anahtar, M.; Dighe, A.S.; Beltran, W.G.; Miller, T.; Scott, H.; Durbin, S.M.; Bharadwaj, M.; Thomas, J.; Gogakos, T.S.; et al. Evaluation of Three Commercial SARS-CoV-2 Serologic Assays and Their Performance in Two-Test Algorithms. J. Clin. Microbiol. 2020, 59. [Google Scholar] [CrossRef]
- Tan, S.S.; Saw, S.; Chew, K.L.; Wang, C.; Pajarillaga, A.; Khoo, C.; Wang, W.; Ali, Z.M.; Yang, Z.; Chan, Y.H.; et al. Comparative Clinical Evaluation of the Roche Elecsys and Abbott Severe Acute Respiratory Syndrome Coronavirus 2 (SARS-CoV-2) Serology Assays for Coronavirus Disease 2019 (COVID-19). Arch. Pathol. Lab. Med. 2021, 145, 32–38. [Google Scholar] [CrossRef]
- Trabaud, M.-A.; Icard, V.; Milon, M.-P.; Bal, A.; Lina, B.; Escuret, V. Comparison of Eight Commercial, High-throughput, Automated or ELISA Assays Detecting SARS-CoV-2 IgG or Total Antibody. J. Clin. Virol. 2020, 132, 104613. [Google Scholar] [CrossRef]
- Tan, S.S.; Saw, S.; Chew, K.L.; Huak, C.Y.; Khoo, C.; Pajarillaga, A.; Wang, W.; Tambyah, P.; Ong, L.; Jureen, R.; et al. Head-to-head Evaluation on Diagnostic Accuracies of Six SARS-CoV-2 Serological Assays. Pathology 2020, 52, 770–777. [Google Scholar] [CrossRef]
- Charlton, C.L.; Kanji, J.N.; Johal, K.; Bailey, A.; Plitt, S.S.; MacDonald, C.; Kunst, A.; Buss, E.; Burnes, L.E.; Fonseca, K.; et al. Evaluation of Six Commercial Mid- to High-Volume Antibody and Six Point-of-Care Lateral Flow Assays for Detection of SARS-CoV-2 Antibodies. J. Clin. Microbiol. 2020, 58. [Google Scholar] [CrossRef] [PubMed]
- Theel, E.S.; Harring, J.; Hilgart, H.; Granger, D. Performance Characteristics of Four High-Throughput Immunoassays for Detection of IgG Antibodies against SARS-CoV-2. J. Clin. Microbiol. 2020, 58. [Google Scholar] [CrossRef]
- Higgins, V.; Fabros, A.; Wang, X.Y.; Bhandari, M.; Daghfal, D.J.; Kulasingam, V. Anti-SARS-CoV-2 IgM Improves Clinical Sensitivity Early in Disease Course. Clin. Biochem. 2021, 90, 1–7. [Google Scholar] [CrossRef] [PubMed]
- Higgins, V.; Fabros, A.; Kulasingam, V. Quantitative measurement of anti-SARS-CoV-2 antibodies: Analytical and clinical evaluation. J. Clin. Microbiol. 2021, 59. [Google Scholar] [CrossRef] [PubMed]
- Schaffner, A.; Risch, L.; Aeschbacher, S.; Risch, C.; Weber, M.C.; Thiel, S.L.; Jüngert, K.; Pichler, M.; Grossmann, K.; Wohlwend, N.; et al. Characterization of a Pan-Immunoglobulin Assay Quantifying Antibodies Directed against the Receptor Binding Domain of the SARS-CoV-2 S1-Subunit of the Spike Protein: A Population-Based Study. J. Clin. Med. 2020, 9, 3989. [Google Scholar] [CrossRef]
- Mei, J.V.; Alexander, J.R.; Adam, B.W.; Hannon, W.H. Use of Filter Paper for the Collection and Analysis of Human Whole Blood Specimens. J. Nutr. 2001, 131, 1631S–1636S. [Google Scholar] [CrossRef]
- Iankov, I.; Viker, K.; Turgeon, C.; Matern, D.; Galanis, E. Parameters of Immunoglobulin Extraction from Dried Blood Spot Cards and Immunoassays for Detection of Antibody Response to Pathogens Including the Novel SARS-CoV-2. J. Immunol. Methods 2021, 492, 112996. [Google Scholar] [CrossRef]
- Suthar, M.S.; Zimmerman, M.G.; Kauffman, R.C.; Mantus, G.; Linderman, S.L.; Hudson, W.H.; Vanderheiden, A.; Nyhoff, L.; Davis, C.W.; Adekunle, O.; et al. Rapid Generation of Neutralizing Antibody Responses in COVID-19 Patients. Cell Rep. Med. 2020, 1, 100040. [Google Scholar] [CrossRef]
- Suhandynata, R.T.; A Hoffman, M.; Huang, D.; Tran, J.T.; Kelner, M.J.; Reed, S.L.; McLawhon, R.W.; E Voss, J.; Nemazee, D.; Fitzgerald, R.L. Commercial Serology Assays Predict Neutralization Activity Against SARS-CoV-2. Clin. Chem. 2021, 67, 404–414. [Google Scholar] [CrossRef] [PubMed]
- Tang, M.S.; Case, J.B.; Franks, C.E.; Chen, R.E.; Anderson, N.W.; Henderson, J.P.; Diamond, M.S.; Gronowski, A.M.; Farnsworth, C.W. Association between SARS-CoV-2 Neutralizing Antibodies and Commercial Serological Assays. Clin. Chem. 2020, 66, 1538–1547. [Google Scholar] [CrossRef] [PubMed]
- Iyer, A.S.; Jones, F.K.; Nodoushani, A.; Kelly, M.; Becker, M.; Slater, D.; Mills, R.; Teng, E.; Kamruzzaman, M.; Garcia-Beltran, W.F.; et al. Dynamics and Significance of the Antibody Response to SARS-CoV-2 Infection. MedRxiv 2020. [Google Scholar] [CrossRef]
- Mattiuzzo, G.; Bentley, E.M.; Hassall, M.; Routley, S.; Richardson, S.; Bernasconi, V.; Kristiansen, P.; Harvala, H.; Roberts, D.; Semple, M.G.; et al. WHO/BS.2020.2403 Establishment of the WHO International Standard and Reference Panel for Anti-SARS-CoV-2 Antibody; World Health Organization: Geneva, Switzerland, 2020. [Google Scholar]

| Comparisons | Threshold (U/mL) | True Negatives | True Positives | False Negatives | False Positives | Sensitivity | Specificity |
|---|---|---|---|---|---|---|---|
| Plasma vs. PCR-confirmed COVID-19 | 0.8 | 11 | 46 | 6 | 0 | 0.88 (95% CI: 0.77, 0.96) | 1 (95% CI: 0.72, 1) |
| DBS vs. PCR-confirmed COVID-19 | 0.8 | 11 | 45 | 7 | 0 | 0.87 (95% CI: 0.74, 0.94) | 1 (95% CI: 0.72, 1) |
| DBS vs. plasma-positive COVID-19 | 0.8 | 17 | 45 | 1 | 0 | 0.98 (95% CI: 0.88, 1) | 1 (95% CI: 0.8, 1) |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Brinc, D.; Biondi, M.J.; Li, D.; Sun, H.; Capraru, C.; Smookler, D.; Zahoor, M.A.; Casey, J.; Kulasingam, V.; Feld, J.J. Evaluation of Dried Blood Spot Testing for SARS-CoV-2 Serology Using a Quantitative Commercial Assay. Viruses 2021, 13, 962. https://doi.org/10.3390/v13060962
Brinc D, Biondi MJ, Li D, Sun H, Capraru C, Smookler D, Zahoor MA, Casey J, Kulasingam V, Feld JJ. Evaluation of Dried Blood Spot Testing for SARS-CoV-2 Serology Using a Quantitative Commercial Assay. Viruses. 2021; 13(6):962. https://doi.org/10.3390/v13060962
Chicago/Turabian StyleBrinc, Davor, Mia J. Biondi, Daniel Li, Heng Sun, Camelia Capraru, David Smookler, Muhammad Atif Zahoor, Julia Casey, Vathany Kulasingam, and Jordan J. Feld. 2021. "Evaluation of Dried Blood Spot Testing for SARS-CoV-2 Serology Using a Quantitative Commercial Assay" Viruses 13, no. 6: 962. https://doi.org/10.3390/v13060962
APA StyleBrinc, D., Biondi, M. J., Li, D., Sun, H., Capraru, C., Smookler, D., Zahoor, M. A., Casey, J., Kulasingam, V., & Feld, J. J. (2021). Evaluation of Dried Blood Spot Testing for SARS-CoV-2 Serology Using a Quantitative Commercial Assay. Viruses, 13(6), 962. https://doi.org/10.3390/v13060962







