Orthobunyaviruses: From Virus Binding to Penetration into Mammalian Host Cells
Abstract
1. Introduction
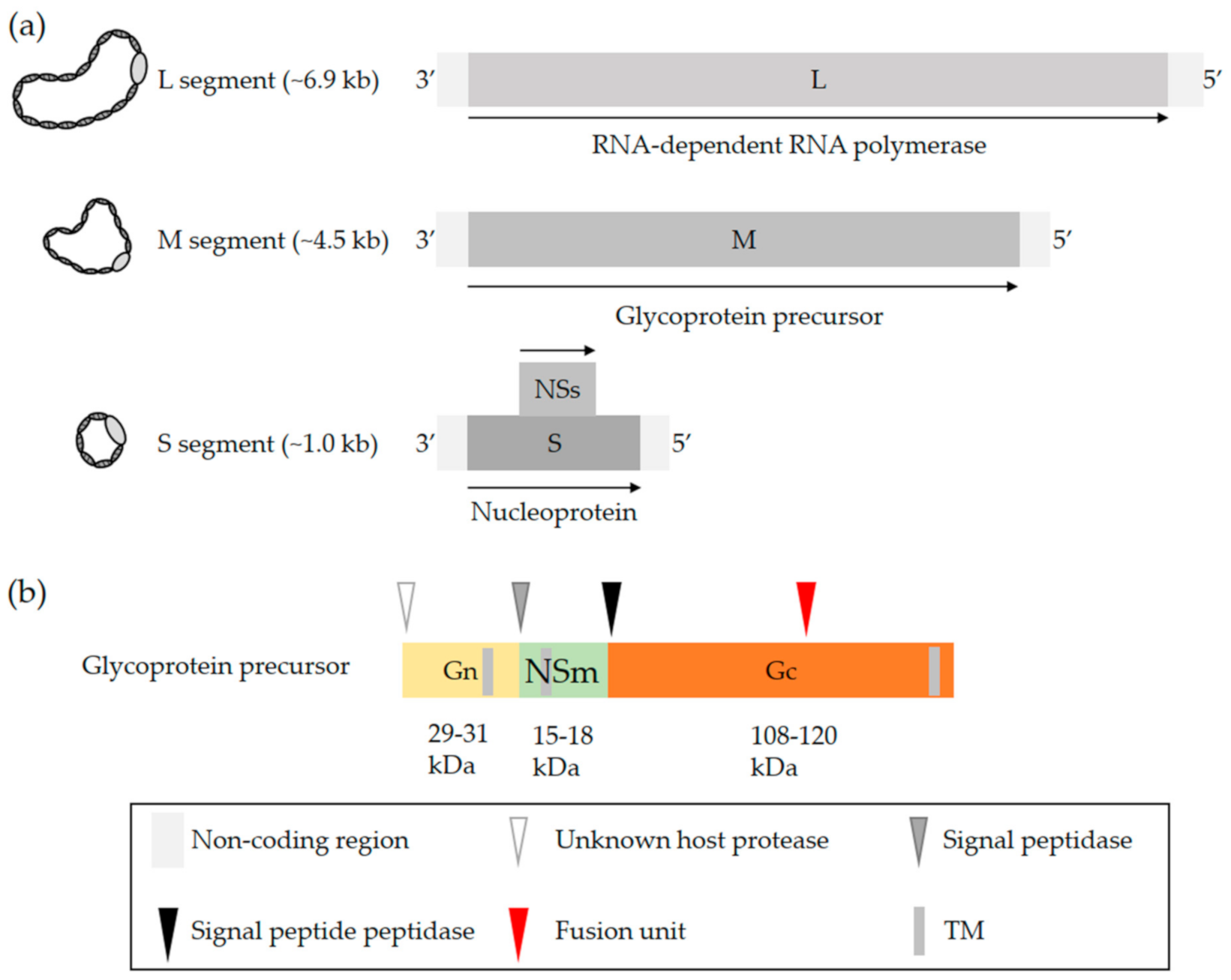
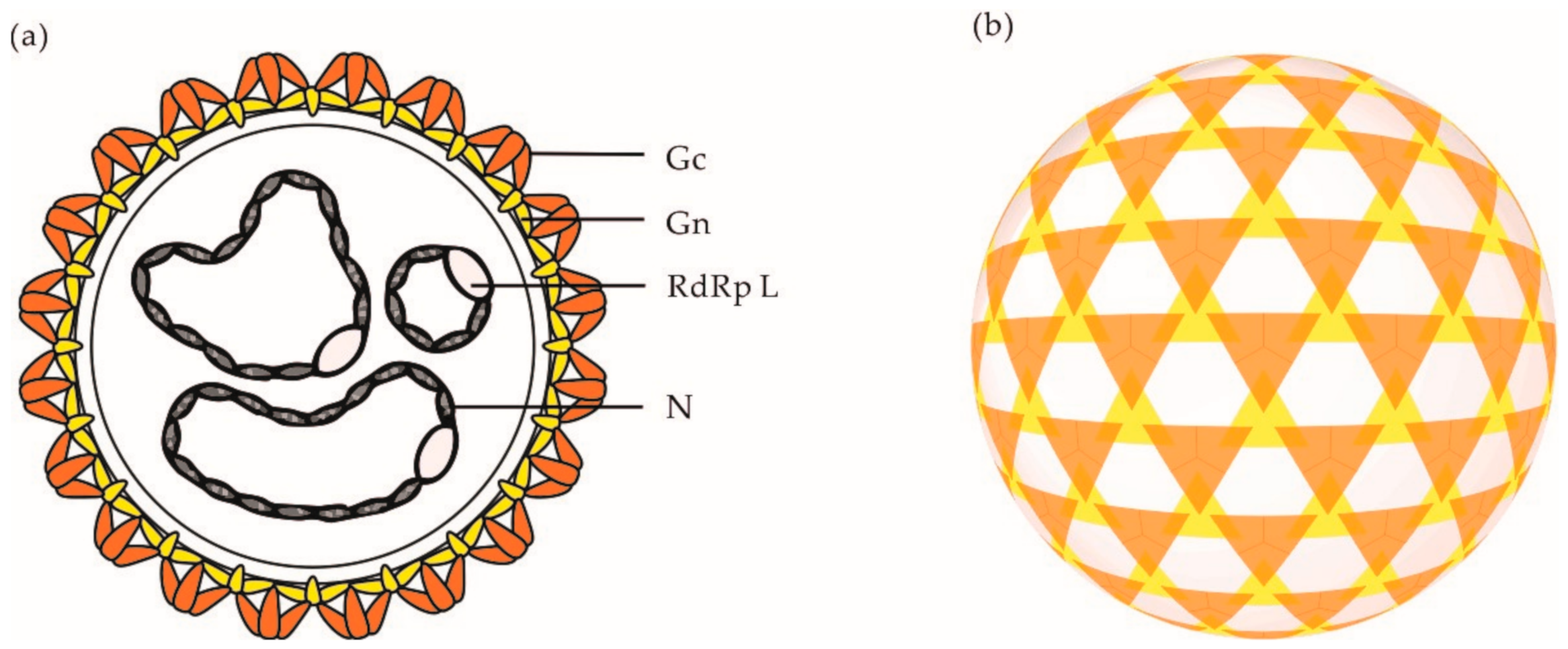
2. OBV Genome and Viral Particles
3. Receptors for OBVs in Mammalian Hosts
4. OBV Uptake
5. OBV Intracellular Trafficking
6. OBV-Cell Membrane Fusion and Penetration
7. Conclusions and Perspectives
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
References
- Abudurexiti, A.; Adkins, S.; Alioto, D.; Alkhovsky, S.V.; Avsic-Zupanc, T.; Ballinger, M.J.; Bente, D.A.; Beer, M.; Bergeron, E.; Blair, C.D.; et al. Taxonomy of the order Bunyavirales: Update 2019. Arch. Virol. 2019, 164, 1949–1965. [Google Scholar] [CrossRef] [PubMed]
- Elliott, R.M. Orthobunyaviruses: Recent genetic and structural insights. Nat. Rev. Microbiol. 2014, 12, 673–685. [Google Scholar] [CrossRef] [PubMed]
- Sakkas, H.; Bozidis, P.; Franks, A.; Papadopoulou, C. Oropouche Fever: A Review. Viruses 2018, 10, 175. [Google Scholar] [CrossRef]
- Romero-Alvarez, D.; Escobar, L.E. Oropouche fever, an emergent disease from the Americas. Microbes Infect. 2018, 20, 135–146. [Google Scholar] [CrossRef] [PubMed]
- Gerrard, S.R.; Li, L.; Barrett, A.D.; Nichol, S.T. Ngari virus is a Bunyamwera virus reassortant that can be associated with large outbreaks of hemorrhagic fever in Africa. J. Virol. 2004, 78, 8922–8926. [Google Scholar] [CrossRef] [PubMed]
- Dutuze, M.F.; Nzayirambaho, M.; Mores, C.N.; Christofferson, R.C. A Review of Bunyamwera, Batai, and Ngari Viruses: Understudied Orthobunyaviruses With Potential One Health Implications. Front. Vet. Sci. 2018, 5, 69. [Google Scholar] [CrossRef] [PubMed]
- Edridge, A.W.D.; van der Hoek, L. Emerging orthobunyaviruses associated with CNS disease. PLoS Negl. Trop. Dis. 2020, 14, e0008856. [Google Scholar] [CrossRef] [PubMed]
- Holden, P.; Hess, A.D. Cache Valley virus, a previously undescribed mosquito-borne agent. Science 1959, 130, 1187–1188. [Google Scholar] [CrossRef] [PubMed]
- Edwards, J.F. Cache Valley virus. Vet. Clin. N. Am. Food Anim. Pract. 1994, 10, 515–524. [Google Scholar] [CrossRef]
- Wernike, K.; Beer, M. Schmallenberg Virus: To Vaccinate, or Not to Vaccinate? Vaccines 2020, 8, 287. [Google Scholar] [CrossRef] [PubMed]
- Haddow, A.D.; Odoi, A. The incidence risk, clustering, and clinical presentation of La Crosse virus infections in the eastern United States, 2003–2007. PLoS ONE 2009, 4, e6145. [Google Scholar] [CrossRef]
- Rodriguez, C.; Gricourt, G.; Ndebi, M.; Demontant, V.; Poiteau, L.; Burrel, S.; Boutolleau, D.; Woerther, P.L.; Calvez, V.; Stroer, S.; et al. Fatal Encephalitis Caused by Cristoli Virus, an Emerging Orthobunyavirus, France. Emerg. Infect. Dis. 2020, 26, 1287–1290. [Google Scholar] [CrossRef]
- Perot, P.; Bielle, F.; Bigot, T.; Foulongne, V.; Bollore, K.; Chretien, D.; Gil, P.; Gutierrez, S.; L’Ambert, G.; Mokhtari, K.; et al. Identification of Umbre Orthobunyavirus as a Novel Zoonotic Virus Responsible for Lethal Encephalitis in 2 French Patients with Hypogammaglobulinemia. Clin. Infect. Dis. 2020. [Google Scholar] [CrossRef]
- Dandawate, C.N.; Rajagopalan, P.K.; Pavri, K.M.; Work, T.H. Virus isolations from mosquitoes collected in North Arcot district, Madras state, and Chittoor district, Andhra Pradesh between November 1955 and October 1957. Indian J. Med. Res. 1969, 57, 1420–1426. [Google Scholar]
- Dutuze, M.F.; Mayton, E.H.; Macaluso, J.D.; Christofferson, R.C. Comparative characterization of the reassortant Orthobunyavirus Ngari with putative parental viruses, Bunyamwera and Batai: In vitro characterization and ex vivo stability. J. Gen. Virol. 2021, 102. [Google Scholar] [CrossRef]
- Briese, T.; Bird, B.; Kapoor, V.; Nichol, S.T.; Lipkin, W.I. Batai and Ngari viruses: M segment reassortment and association with severe febrile disease outbreaks in East Africa. J. Virol. 2006, 80, 5627–5630. [Google Scholar] [CrossRef]
- Endalew, A.D.; Faburay, B.; Wilson, W.C.; Richt, J.A. Schmallenberg Disease-A Newly Emerged Culicoides-borne Viral Disease of Ruminants. Viruses 2019, 11, 1065. [Google Scholar] [CrossRef]
- Waddell, L.; Pachal, N.; Mascarenhas, M.; Greig, J.; Harding, S.; Young, I.; Wilhelm, B. Cache Valley virus: A scoping review of the global evidence. Zoonoses Public Health 2019, 66, 739–758. [Google Scholar] [CrossRef]
- McJunkin, J.E.; Khan, R.R.; Tsai, T.F. California-La Crosse encephalitis. Infect. Dis. Clin. N. Am. 1998, 12, 83–93. [Google Scholar] [CrossRef]
- Carpenter, S.; Groschup, M.H.; Garros, C.; Felippe-Bauer, M.L.; Purse, B.V. Culicoides biting midges, arboviruses and public health in Europe. Antivir. Res. 2013, 100, 102–113. [Google Scholar] [CrossRef]
- da Rosa, J.F.T.; de Souza, W.M.; Pinheiro, F.P.; Figueiredo, M.L.; Cardoso, J.F.; Acrani, G.O.; Nunes, M.R.T. Oropouche Virus: Clinical, Epidemiological, and Molecular Aspects of a Neglected Orthobunyavirus. Am. J. Trop. Med. Hyg. 2017, 96, 1019–1030. [Google Scholar] [CrossRef] [PubMed]
- Barr, J.N.; Elliott, R.M.; Dunn, E.F.; Wertz, G.W. Segment-specific terminal sequences of Bunyamwera bunyavirus regulate genome replication. Virology 2003, 311, 326–338. [Google Scholar] [CrossRef]
- Kohl, A.; Dunn, E.F.; Lowen, A.C.; Elliott, R.M. Complementarity, sequence and structural elements within the 3’ and 5’ non-coding regions of the Bunyamwera orthobunyavirus S segment determine promoter strength. J. Gen. Virol. 2004, 85, 3269–3278. [Google Scholar] [CrossRef] [PubMed]
- Pardigon, N.; Vialat, P.; Girard, M.; Bouloy, M. Panhandles and hairpin structures at the termini of germiston virus RNAs (Bunyavirus). Virology 1982, 122, 191–197. [Google Scholar] [CrossRef]
- Ogg, M.M.; Patterson, J.L. RNA binding domain of Jamestown Canyon virus S segment RNAs. J. Virol. 2007, 81, 13754–13760. [Google Scholar] [CrossRef]
- Kohl, A.; Lowen, A.C.; Leonard, V.H.J.; Elliott, R.M. Genetic elements regulating packaging of the Bunyamwera orthobunyavirus genome. J. Gen. Virol. 2006, 87, 177–187. [Google Scholar] [CrossRef]
- Osborne, J.C.; Elliott, R.M. RNA binding properties of bunyamwera virus nucleocapsid protein and selective binding to an element in the 5’ terminus of the negative-sense S segment. J. Virol. 2000, 74, 9946–9952. [Google Scholar] [CrossRef]
- Dong, H.; Li, P.; Bottcher, B.; Elliott, R.M.; Dong, C. Crystal structure of Schmallenberg orthobunyavirus nucleoprotein-RNA complex reveals a novel RNA sequestration mechanism. RNA 2013, 19, 1129–1136. [Google Scholar] [CrossRef]
- Dong, H.; Li, P.; Elliott, R.M.; Dong, C. Structure of Schmallenberg orthobunyavirus nucleoprotein suggests a novel mechanism of genome encapsidation. J. Virol. 2013, 87, 5593–5601. [Google Scholar] [CrossRef]
- Ariza, A.; Tanner, S.J.; Walter, C.T.; Dent, K.C.; Shepherd, D.A.; Wu, W.; Matthews, S.V.; Hiscox, J.A.; Green, T.J.; Luo, M.; et al. Nucleocapsid protein structures from orthobunyaviruses reveal insight into ribonucleoprotein architecture and RNA polymerization. Nucleic Acids Res. 2013, 41, 5912–5926. [Google Scholar] [CrossRef]
- Li, B.; Wang, Q.; Pan, X.; de Castro, I.F.; Sun, Y.; Guo, Y.; Tao, X.; Risco, C.; Sui, S.F.; Lou, Z. Bunyamwera virus possesses a distinct nucleocapsid protein to facilitate genome encapsidation. Proc. Natl. Acad. Sci. USA 2013, 110, 9048–9053. [Google Scholar] [CrossRef]
- Niu, F.; Shaw, N.; Wang, Y.E.; Jiao, L.; Ding, W.; Li, X.; Zhu, P.; Upur, H.; Ouyang, S.; Cheng, G.; et al. Structure of the Leanyer orthobunyavirus nucleoprotein-RNA complex reveals unique architecture for RNA encapsidation. Proc. Natl. Acad. Sci. USA 2013, 110, 9054–9059. [Google Scholar] [CrossRef]
- Reguera, J.; Cusack, S.; Kolakofsky, D. Segmented negative strand RNA virus nucleoprotein structure. Curr. Opin. Virol. 2014, 5, 7–15. [Google Scholar] [CrossRef]
- Reguera, J.; Malet, H.; Weber, F.; Cusack, S. Structural basis for encapsidation of genomic RNA by La Crosse Orthobunyavirus nucleoprotein. Proc. Natl. Acad. Sci. USA 2013, 110, 7246–7251. [Google Scholar] [CrossRef]
- Garry, C.E.; Garry, R.F. Proteomics computational analyses suggest that the carboxyl terminal glycoproteins of Bunyaviruses are class II viral fusion protein (beta-penetrenes). Theor. Biol. Med. Model. 2004, 1, 1–16. [Google Scholar] [CrossRef]
- Tischler, N.D.; Gonzalez, A.; Perez-Acle, T.; Rosemblatt, M.; Valenzuela, P.D.T. Hantavirus Gc glycoprotein: Evidence for a class II fusion protein. J. Gen. Virol. 2005, 86, 2937–2947. [Google Scholar] [CrossRef]
- Plassmeyer, M.L.; Soldan, S.S.; Stachelek, K.M.; Martin-Garcia, J.; Gonzalez-Scarano, F. California serogroup Gc (G1) glycoprotein is the principal determinant of pH-dependent cell fusion and entry. Virology 2005, 338, 121–132. [Google Scholar] [CrossRef]
- Fazakerley, J.K.; Gonzalez-Scarano, F.; Strickler, J.; Dietzschold, B.; Karush, F.; Nathanson, N. Organization of the middle RNA segment of snowshoe hare Bunyavirus. Virology 1988, 167, 422–432. [Google Scholar] [CrossRef]
- Shi, X.; Botting, C.H.; Li, P.; Niglas, M.; Brennan, B.; Shirran, S.L.; Szemiel, A.M.; Elliott, R.M. Bunyamwera orthobunyavirus glycoprotein precursor is processed by cellular signal peptidase and signal peptide peptidase. Proc. Natl. Acad. Sci. USA 2016, 113, 8825–8830. [Google Scholar] [CrossRef]
- Nakitare, G.W.; Elliott, R.M. Expression of the Bunyamwera virus M genome segment and intracellular localization of NSm. Virology 1993, 195, 511–520. [Google Scholar] [CrossRef]
- Lappin, D.F.; Nakitare, G.W.; Palfreyman, J.W.; Elliott, R.M. Localization of Bunyamwera bunyavirus G1 glycoprotein to the Golgi requires association with G2 but not with NSm. J. Gen. Virol. 1994, 75 Pt 12, 3441–3451. [Google Scholar] [CrossRef]
- Obijeski, J.F.; Bishop, D.H.; Murphy, F.A.; Palmer, E.L. Structural proteins of La Crosse virus. J. Virol. 1976, 19, 985–997. [Google Scholar] [CrossRef]
- Bowden, T.A.; Bitto, D.; McLees, A.; Yeromonahos, C.; Elliott, R.M.; Huiskonen, J.T. Orthobunyavirus ultrastructure and the curious tripodal glycoprotein spike. PLoS Pathog. 2013, 9, e1003374. [Google Scholar] [CrossRef]
- Léger, P.; Lozach, P.-Y. Bunyaviruses: From transmission by arthropods to virus entry into the mammalian host first-target cells. Future Virol. 2015, 10, 859–881. [Google Scholar] [CrossRef]
- Boulant, S.; Stanifer, M.; Lozach, P.Y. Dynamics of virus-receptor interactions in virus binding, signaling, and endocytosis. Viruses 2015, 7, 2794–2815. [Google Scholar] [CrossRef]
- Maginnis, M.S. Virus-Receptor Interactions: The Key to Cellular Invasion. J. Mol. Biol. 2018, 430, 2590–2611. [Google Scholar] [CrossRef]
- Thamamongood, T.; Aebischer, A.; Wagner, V.; Chang, M.W.; Elling, R.; Benner, C.; Garcia-Sastre, A.; Kochs, G.; Beer, M.; Schwemmle, M. A Genome-Wide CRISPR-Cas9 Screen Reveals the Requirement of Host Cell Sulfation for Schmallenberg Virus Infection. J. Virol. 2020, 94. [Google Scholar] [CrossRef]
- Murakami, S.; Takenaka-Uema, A.; Kobayashi, T.; Kato, K.; Shimojima, M.; Palmarini, M.; Horimoto, T. Heparan Sulfate Proteoglycan Is an Important Attachment Factor for Cell Entry of Akabane and Schmallenberg Viruses. J. Virol. 2017, 91. [Google Scholar] [CrossRef]
- Lozach, P.Y.; Kuhbacher, A.; Meier, R.; Mancini, R.; Bitto, D.; Bouloy, M.; Helenius, A. DC-SIGN as a receptor for phleboviruses. Cell Host Microbe 2011, 10, 75–88. [Google Scholar] [CrossRef]
- Hofmann, H.; Li, X.; Zhang, X.; Liu, W.; Kuhl, A.; Kaup, F.; Soldan, S.S.; Gonzalez-Scarano, F.; Weber, F.; He, Y.; et al. Severe fever with thrombocytopenia virus glycoproteins are targeted by neutralizing antibodies and can use DC-SIGN as a receptor for pH-dependent entry into human and animal cell lines. J. Virol. 2013, 87, 4384–4394. [Google Scholar] [CrossRef]
- Monteiro, J.T.; Schon, K.; Ebbecke, T.; Goethe, R.; Ruland, J.; Baumgartner, W.; Becker, S.C.; Lepenies, B. The CARD9-Associated C-Type Lectin, Mincle, Recognizes La Crosse Virus (LACV) but Plays a Limited Role in Early Antiviral Responses against LACV. Viruses 2019, 11, 303. [Google Scholar] [CrossRef] [PubMed]
- Svajger, U.; Anderluh, M.; Jeras, M.; Obermajer, N. C-type lectin DC-SIGN: An adhesion, signalling and antigen-uptake molecule that guides dendritic cells in immunity. Cell Signal. 2010, 22, 1397–1405. [Google Scholar] [CrossRef] [PubMed]
- Hollidge, B.S.; Nedelsky, N.B.; Salzano, M.V.; Fraser, J.W.; Gonzalez-Scarano, F.; Soldan, S.S. Orthobunyavirus entry into neurons and other mammalian cells occurs via clathrin-mediated endocytosis and requires trafficking into early endosomes. J. Virol. 2012, 86, 7988–8001. [Google Scholar] [CrossRef] [PubMed]
- de Souza Luna, L.K.; Rodrigues, A.H.; Santos, R.I.; Sesti-Costa, R.; Criado, M.F.; Martins, R.B.; Silva, M.L.; Delcaro, L.S.; Proenca-Modena, J.L.; Figueiredo, L.T.; et al. Oropouche virus is detected in peripheral blood leukocytes from patients. J. Med. Virol. 2017, 89, 1108–1111. [Google Scholar] [CrossRef]
- Bangphoomi, N.; Takenaka-Uema, A.; Sugi, T.; Kato, K.; Akashi, H.; Horimoto, T. Akabane virus utilizes alternative endocytic pathways to entry into mammalian cell lines. J. Vet. Med. Sci. 2014, 76, 1471–1478. [Google Scholar] [CrossRef]
- Santos, R.I.; Rodrigues, A.H.; Silva, M.L.; Mortara, R.A.; Rossi, M.A.; Jamur, M.C.; Oliver, C.; Arruda, E. Oropouche virus entry into HeLa cells involves clathrin and requires endosomal acidification. Virus Res. 2008, 138, 139–143. [Google Scholar] [CrossRef]
- Leger, P.; Tetard, M.; Youness, B.; Cordes, N.; Rouxel, R.N.; Flamand, M.; Lozach, P.Y. Differential Use of the C-Type Lectins L-SIGN and DC-SIGN for Phlebovirus Endocytosis. Traffic 2016, 17, 639–656. [Google Scholar] [CrossRef]
- Koch, J.; Xin, Q.; Tischler, N.D.; Lozach, P.Y. Entry of Phenuiviruses into Mammalian Host Cells. Viruses 2021, 13, 299. [Google Scholar] [CrossRef]
- Scott, C.C.; Vacca, F.; Gruenberg, J. Endosome maturation, transport and functions. Semin. Cell Dev. Biol. 2014, 31, 2–10. [Google Scholar] [CrossRef]
- White, J.M.; Whittaker, G.R. Fusion of Enveloped Viruses in Endosomes. Traffic 2016, 17, 593–614. [Google Scholar] [CrossRef]
- Scott, C.C.; Gruenberg, J. Ion flux and the function of endosomes and lysosomes: pH is just the start: The flux of ions across endosomal membranes influences endosome function not only through regulation of the luminal pH. Bioessays 2011, 33, 103–110. [Google Scholar] [CrossRef]
- Albornoz, A.; Hoffmann, A.B.; Lozach, P.Y.; Tischler, N.D. Early Bunyavirus-Host Cell Interactions. Viruses 2016, 8, 143. [Google Scholar] [CrossRef]
- Huotari, J.; Helenius, A. Endosome maturation. EMBO J. 2011, 30, 3481–3500. [Google Scholar] [CrossRef]
- Lozach, P.Y.; Huotari, J.; Helenius, A. Late-penetrating viruses. Curr. Opin. Virol. 2011, 1, 35–43. [Google Scholar] [CrossRef]
- Shtanko, O.; Nikitina, R.A.; Altuntas, C.Z.; Chepurnov, A.A.; Davey, R.A. Crimean-Congo hemorrhagic fever virus entry into host cells occurs through the multivesicular body and requires ESCRT regulators. PLoS Pathog. 2014, 10, e1004390. [Google Scholar] [CrossRef]
- Pekosz, A.; Gonzalez-Scarano, F. The extracellular domain of La Crosse virus G1 forms oligomers and undergoes pH-dependent conformational changes. Virology 1996, 225, 243–247. [Google Scholar] [CrossRef]
- Harrison, S.C. Viral membrane fusion. Virology 2015, 479, 498–507. [Google Scholar] [CrossRef]
- Plassmeyer, M.L.; Soldan, S.S.; Stachelek, K.M.; Roth, S.M.; Martin-Garcia, J.; Gonzalez-Scarano, F. Mutagenesis of the La Crosse Virus glycoprotein supports a role for Gc (1066–1087) as the fusion peptide. Virology 2007, 358, 273–282. [Google Scholar] [CrossRef]
- Soldan, S.S.; Hollidge, B.S.; Wagner, V.; Weber, F.; Gonzalez-Scarano, F. La Crosse virus (LACV) Gc fusion peptide mutants have impaired growth and fusion phenotypes, but remain neurotoxic. Virology 2010, 404, 139–147. [Google Scholar] [CrossRef]
- Shi, X.; Goli, J.; Clark, G.; Brauburger, K.; Elliott, R.M. Functional analysis of the Bunyamwera orthobunyavirus Gc glycoprotein. J. Gen. Virol. 2009, 90, 2483–2492. [Google Scholar] [CrossRef]
- Shi, X.; van Mierlo, J.T.; French, A.; Elliott, R.M. Visualizing the replication cycle of bunyamwera orthobunyavirus expressing fluorescent protein-tagged Gc glycoprotein. J. Virol. 2010, 84, 8460–8469. [Google Scholar] [CrossRef] [PubMed]
- Hellert, J.; Aebischer, A.; Wernike, K.; Haouz, A.; Brocchi, E.; Reiche, S.; Guardado-Calvo, P.; Beer, M.; Rey, F.A. Orthobunyavirus spike architecture and recognition by neutralizing antibodies. Nat. Commun. 2019, 10, 1–14. [Google Scholar] [CrossRef] [PubMed]
- Carneiro, F.A.; Stauffer, F.; Lima, C.S.; Juliano, M.A.; Juliano, L.; Da Poian, A.T. Membrane fusion induced by vesicular stomatitis virus depends on histidine protonation. J. Biol. Chem. 2003, 278, 13789–13794. [Google Scholar] [CrossRef] [PubMed]
- Kampmann, T.; Mueller, D.S.; Mark, A.E.; Young, P.R.; Kobe, B. The Role of histidine residues in low-pH-mediated viral membrane fusion. Structure 2006, 14, 1481–1487. [Google Scholar] [CrossRef]
- Hacker, J.K.; Hardy, J.L. Adsorptive endocytosis of California encephalitis virus into mosquito and mammalian cells: A role for G1. Virology 1997, 235, 40–47. [Google Scholar] [CrossRef]
- Jacoby, D.R.; Cooke, C.; Prabakaran, I.; Boland, J.; Nathanson, N.; Gonzalez-Scarano, F. Expression of the La Crosse M segment proteins in a recombinant vaccinia expression system mediates pH-dependent cellular fusion. Virology 1993, 193, 993–996. [Google Scholar] [CrossRef]
- Pobjecky, N.; Smith, J.; Gonzalez-Scarano, F. Biological studies of the fusion function of California serogroup Bunyaviruses. Microb. Pathog. 1986, 1, 491–501. [Google Scholar] [CrossRef]
- Stauffer, S.; Feng, Y.; Nebioglu, F.; Heilig, R.; Picotti, P.; Helenius, A. Stepwise priming by acidic pH and a high K+ concentration is required for efficient uncoating of influenza A virus cores after penetration. J. Virol. 2014, 88, 13029–13046. [Google Scholar] [CrossRef]
- Hover, S.; Foster, B.; Fontana, J.; Kohl, A.; Goldstein, S.A.N.; Barr, J.N.; Mankouri, J. Bunyavirus requirement for endosomal K+ reveals new roles of cellular ion channels during infection. PLoS Pathog. 2018, 14, e1006845. [Google Scholar] [CrossRef]
- Hover, S.; King, B.; Hall, B.; Loundras, E.A.; Taqi, H.; Daly, J.; Dallas, M.; Peers, C.; Schnettler, E.; McKimmie, C.; et al. Modulation of Potassium Channels Inhibits Bunyavirus Infection. J. Biol. Chem. 2016, 291, 3411–3422. [Google Scholar] [CrossRef]
- Charlton, F.W.; Hover, S.; Fuller, J.; Hewson, R.; Fontana, J.; Barr, J.N.; Mankouri, J. Cellular cholesterol abundance regulates potassium accumulation within endosomes and is an important determinant in bunyavirus entry. J. Biol. Chem. 2019, 294, 7335–7347. [Google Scholar] [CrossRef]
- Sandler, Z.J.; Firpo, M.R.; Omoba, O.S.; Vu, M.N.; Menachery, V.D.; Mounce, B.C. Novel Ionophores Active against La Crosse Virus Identified through Rapid Antiviral Screening. Antimicrob. Agents Chemother. 2020, 64. [Google Scholar] [CrossRef]
- Riblett, A.M.; Blomen, V.A.; Jae, L.T.; Altamura, L.A.; Doms, R.W.; Brummelkamp, T.R.; Wojcechowskyj, J.A. A Haploid Genetic Screen Identifies Heparan Sulfate Proteoglycans Supporting Rift Valley Fever Virus Infection. J. Virol. 2016, 90, 1414–1423. [Google Scholar] [CrossRef]
- Meier, R.; Franceschini, A.; Horvath, P.; Tetard, M.; Mancini, R.; von Mering, C.; Helenius, A.; Lozach, P.Y. Genome-wide small interfering RNA screens reveal VAMP3 as a novel host factor required for Uukuniemi virus late penetration. J. Virol. 2014, 88, 8565–8578. [Google Scholar] [CrossRef]
- Hopkins, K.C.; McLane, L.M.; Maqbool, T.; Panda, D.; Gordesky-Gold, B.; Cherry, S. A genome-wide RNAi screen reveals that mRNA decapping restricts bunyaviral replication by limiting the pools of Dcp2-accessible targets for cap-snatching. Genes Dev. 2013, 27, 1511–1525. [Google Scholar] [CrossRef]
| Genus | Species | Representative Species | Hosts |
|---|---|---|---|
| Herbevirus | 33 | Herbert virus | Invertebrates |
| Orthobunyavirus | 88 | Bunyamwera virus, California encephalitis virus, Germiston virus, La Crosse virus, Oropouche virus, Schmallenberg virus, Umbre virus | Invertebrates, vertebrates |
| Pacuvirus | 5 | Pacui virus | Invertebrates, vertebrates |
| Shangavirus | 1 | Insect virus | Invertebrates |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Windhaber, S.; Xin, Q.; Lozach, P.-Y. Orthobunyaviruses: From Virus Binding to Penetration into Mammalian Host Cells. Viruses 2021, 13, 872. https://doi.org/10.3390/v13050872
Windhaber S, Xin Q, Lozach P-Y. Orthobunyaviruses: From Virus Binding to Penetration into Mammalian Host Cells. Viruses. 2021; 13(5):872. https://doi.org/10.3390/v13050872
Chicago/Turabian StyleWindhaber, Stefan, Qilin Xin, and Pierre-Yves Lozach. 2021. "Orthobunyaviruses: From Virus Binding to Penetration into Mammalian Host Cells" Viruses 13, no. 5: 872. https://doi.org/10.3390/v13050872
APA StyleWindhaber, S., Xin, Q., & Lozach, P.-Y. (2021). Orthobunyaviruses: From Virus Binding to Penetration into Mammalian Host Cells. Viruses, 13(5), 872. https://doi.org/10.3390/v13050872






