Household Cases Suggest That Cats Belonging to Owners with COVID-19 Have a Limited Role in Virus Transmission
Abstract
1. Introduction
2. Materials and Methods
2.1. Owners and Cats
2.2. RT-qPCR from Feline and Environmental Swabs
2.3. Virus Isolation
2.4. ELISA
2.5. Serum Neutralization Assay
2.6. Whole-Genome Sequencing
2.7. Phylogenetic Analysis
3. Results
3.1. Case Description
3.2. Two Cats Out of Five Shed SARS-CoV-2
3.3. One Cat Out of Five Developed Anti-SARS-CoV-2 Antibodies
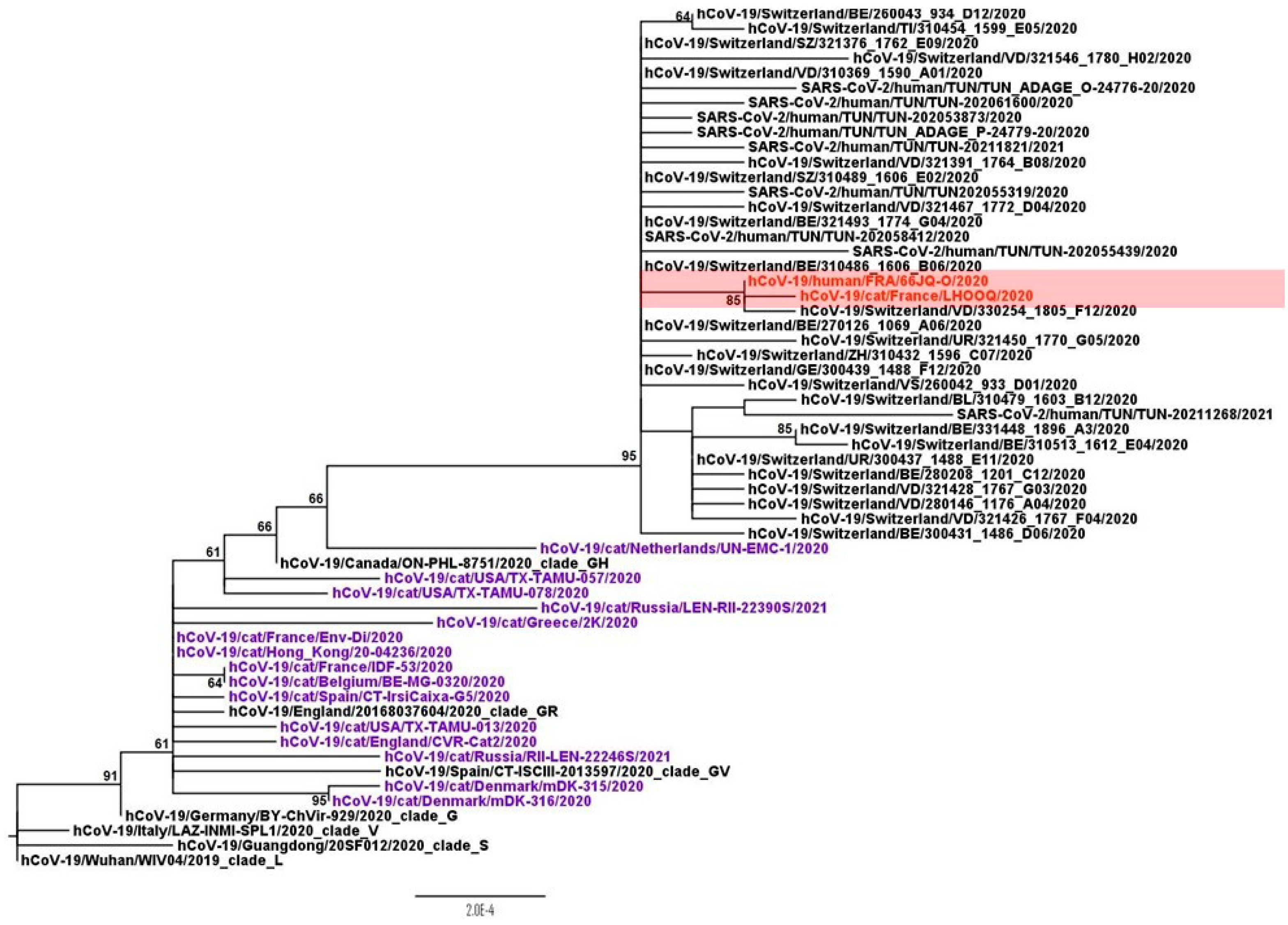
3.4. Phylogenetic Analysis
4. Discussion
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Zhou, P.; Yang, X.-L.; Wang, X.-G.; Hu, B.; Zhang, L.; Zhang, W.; Si, H.-R.; Zhu, Y.; Li, B.; Huang, C.-L.; et al. A pneumonia outbreak associated with a new coronavirus of probable bat origin. Nature 2020, 579, 270–273. [Google Scholar] [CrossRef] [PubMed]
- Musso, N.; Costantino, A.; La Spina, S.; Finocchiaro, A.; Andronico, F.; Stracquadanio, S.; Liotta, L.; Visalli, R.; Emmanuele, G. New SARS-CoV-2 Infection Detected in an Italian Pet Cat by RT-QPCR from Deep Pharyngeal Swab. Pathogens 2020, 9, 746. [Google Scholar] [CrossRef] [PubMed]
- Sailleau, C.; Dumarest, M.; Vanhomwegen, J.; Delaplace, M.; Caro, V.; Kwasiborski, A.; Hourdel, V.; Chevaillier, P.; Barbarino, A.; Comtet, L.; et al. First Detection and Genome Sequencing of SARS-CoV-2 in an Infected Cat in France. Transbound. Emerg. Dis. 2020, 67, 2324–2328. [Google Scholar] [CrossRef] [PubMed]
- Segalés, J.; Puig, M.; Rodon, J.; Avila-Nieto, C.; Carrillo, J.; Cantero, G.; Terron, M.T.; Cruz, S.; Parera, M.; Noguera-Julian, M.; et al. Detection of SARS-CoV-2 in a Cat Owned by a COVID-19−affected Patient in Spain. Proc. Natl. Acad. Sci. USA 2020, 117, 24790–24793. [Google Scholar] [CrossRef]
- Pagani, G.; Lai, A.; Bergna, A.; Rizzo, A.; Stranieri, A.; Giordano, A.; Paltrinieri, S.; Lelli, D.; Decaro, N.; Rusconi, N.; et al. Human-to-Cat SARS-CoV-2 Transmission: Case Report and Full-Genome Sequencing from an Infected Pet and Its Owner in Northern Italy. Pathogens 2021, 10, 252. [Google Scholar] [CrossRef]
- Patterson, E.I.; Elia, G.; Grassi, A.; Giordano, A.; Desario, C.; Medardo, M.; Smith, E.R.; Anderson, T.; Prince, G.T.; Patterson, E.; et al. Evidence of Exposure to SARS-CoV-2 in Cats and Dogs from Households in Italy. Nat. Commun. 2020, 11, 6231. [Google Scholar] [CrossRef] [PubMed]
- Sit, T.H.C.; Brackman, C.J.; Ip, S.M.; Tam, K.W.S.; Law, P.Y.T.; To, E.M.W.; Veronica, Y.T.Y.; Sims, L.D.; Tsang, D.N.C.; Chu, D.K.W.; et al. Infection of Dogs With SARS-CoV-2. Nature 2020, 586, 776–778. [Google Scholar] [CrossRef] [PubMed]
- Oreshkova, N.; Molenaar, R.J.; Vreman, S.; Harders, F.; Munnink, B.B.O.; van der Honing, R.W.H.; Gerhards, N.; Tolsma, P.; Bouwstra, R.; Sikkema, R.S.; et al. SARS-CoV-2 Infection in Farmed Minks, The Netherlands, April and May 2020. Eurosurveillance 2020, 25, 2001005. [Google Scholar] [CrossRef] [PubMed]
- Schlottau, K.; Rissmann, M.; Graaf, A.; Schön, J.; Sehl, J.; Wylezich, C.; Höper, D.; Mettenleiter, T.C.; Balkema-Buschmann, A.; Harder, T.; et al. SARS-CoV-2 in Fruit Bats, Ferrets, Pigs, and Chickens: An Experimental Transmission Study. Lancet Microbe 2020, 1, e218–e225. [Google Scholar] [CrossRef]
- Sia, S.F.; Yan, L.-M.; Chin, A.W.H.; Fung, K.; Choy, K.-T.; Wong, A.Y.L.; Kaewpreedee, P.; Parera, R.A.P.M.; Poon, L.L.M.; Nicholls, J.M.; et al. Pathogenesis and Transmission of SARS-CoV-2 in Golden Hamsters. Nature 2020, 583, 834–838. [Google Scholar] [CrossRef]
- Leroy, E.M.; Ar Gouilh, M.; Brugère-Picoux, J. The Risk of SARS-CoV-2 Transmission to Pets and Other Wild and Domestic Animals Strongly Mandates a One-Health Strategy to Control the COVID-19 Pandemic. One Health 2020, 10, 100133. [Google Scholar] [CrossRef] [PubMed]
- Ministère Des Solidarités Et De La Santé. Prise En Charge En Ambulatoire Du Covid-19—Recommandations Aux Professionnels De Santé En Charge Des prélèvements De dépistage Par RT-PCR. 2020. Available online: https://solidarites-sante.gouv.fr/IMG/pdf/covid-19__rt-Pcr-Ambulatoire-Fiche-preleveurs.Pdf (accessed on 20 May 2020).
- Quick, J. NCoV-2019 Sequencing Protocol (single Sample) V1 (protocols.io.Bdbfi2jn). Protocols.io 2020, 13. [Google Scholar] [CrossRef]
- Martin, D.P.; Murrell, B.; Golden, M.; Khoosal, A.; Muhire, B. RDP4: Detection and Analysis of Recombination Patterns in Virus Genomes. Virus Evol. 2015, 1, vev003. [Google Scholar] [CrossRef]
- Hou, Y.J.; Chiba, S.; Halfmann, P.; Here, C.; Kuroda, M.; Dinnon, K.H., III; Leist, S.R.; Schafer, A.; Nakajima, N.; Takahashi, K.; et al. SARS-CoV-2 D614G Variant Exhibits Efficient Replication Ex Vivo and Transmission In Vivo. Science 2020, 370, 1464–1468. [Google Scholar] [CrossRef]
- Gaudreault, N.N.; Trujillo, J.D.; Carossino, M.; Meekins, D.A.; Morozov, I.; Madden, D.W.; Indran, S.V.; Bold, D.; Balaraman, V.; Kwon, T.; et al. SARS-CoV-2 Infection, Disease and Transmission in Domestic Cats. Emerg. Microbes Infect. 2020, 9, 2322–2332. [Google Scholar] [CrossRef] [PubMed]
- Shi, J.; Wen, Z.; Zhong, G.; Yang, H.; Wang, C.; Huang, B.; Liu, R.; He, X.; Shuai, L.; Sun, Z.; et al. Susceptibility of Ferrets, Cats, Dogs, and Other Do-Mesticated Animals to SARS–coronavirus 2. Science 2020, 368, 1016–1020. [Google Scholar] [CrossRef] [PubMed]
- Halfmann, P.J.; Hatta, M.; Chiba, S.; Maemura, T.; Fan, S.; Takeda, M.; Kinoshita, N.; Hattori, S.-i.; Sagai-Tagava, Y.; Iwatsuki-Horimoto, K.; et al. Transmission of SARS-CoV-2 in Do-Mestic Cats. N. Engl. J. Med. 2020, 383, 592–594. [Google Scholar] [CrossRef] [PubMed]
- Neira, V.; Brito, B.; Agüero, B.; Berrios, F.; Valdés, V.; Gutierrez, A.; Aryama, N.; Espinoza, P.; Retamal, P.; Holmes, E.C.; et al. A Household Case Evidences Shorter Shed-Ding of SARS-CoV-2 in Naturally Infected Cats Compared to Their Human Owners. Emerg. Microbes Infect. 2020, 10, 1–22. [Google Scholar]
- Alonso, F.O.M.; Sabino, B.D.; Guimarães, M.A.A.M.; Varella, R.B. Recurrence of SARS-CoV-2 Infection with a More Severe Case After Mild COVID-19, Reversion of RT-QPCR for Positive and Late Antibody Response: Case Report. J. Med. Virol. 2021, 93, 655–656. [Google Scholar] [CrossRef] [PubMed]
- Lin, Y.-C.; Cheng, C.-Y.; Chen, C.-P.; Cheng, S.-H.; Chang, S.-Y.; Hsueh, P.-R. A Case of Transient Existence of SARS-CoV-2 RNA in the Respiratory Tract with the Absence of Anti-SARS-CoV-2 Antibody Response. Int. J. Infect. Dis. 2020, 96, 464–466. [Google Scholar] [CrossRef] [PubMed]
- Temmam, S.; Barbarino, A.; Maso, D.; Behillil, S.; Enouf, V.; Huon, C.; Jarud, A.; Chevallier, L.; Backovic, M.; Perot, P.; et al. Absence of SARS-CoV-2 Infection in Cats and Dogs in Close Contact with a Cluster of COVID-19 Patients in a Veterinary Campus. One Health 2020, 10, 100164. [Google Scholar] [CrossRef] [PubMed]
- Corman, V.M.; Landt, O.; Kaiser, M.; Molenkamp, R.; Meijer, A.; Chu, D.K.; Drosten, C. Detection of 2019 Novel Corona-Virus (2019-NCoV) by Real-Time RT-PCR. Eurosurveillance 2020, 25, 2000045. [Google Scholar] [CrossRef] [PubMed]
- Penning, L.C.; Vrieling, H.E.; Brinkhof, B.; Riemers, F.M.; Rothuizen, J.; Rutteman, G.R.; Hazewinkel, H.A. A Validation of 10 Feline Reference Genes for Gene Expression Measurements in Snap-Frozen Tissues. Veter. Immunol. Immunopathol. 2007, 120, 212–222. [Google Scholar] [CrossRef] [PubMed]
- De Jonge, H.J.M.; Fehrmann, R.S.N.; De Bont, E.S.J.M.; Hofstra, R.M.W.; Gerbens, F.; Kamps, W.A.; de Vries, E.G.E.; van der Zee, A.G.J.; te Meerman, G.J.; ter Elst, A. Evidence Based Selection of Housekeeping Genes. PLoS ONE 2007, 2, e898. [Google Scholar] [CrossRef]
| Case | Breed | Age (Years) | Gender | Days Between Owners’ Symptoms onset and Cat Swabbing | Positive Swabs | ELISA/ SN (Days 1) | ||
|---|---|---|---|---|---|---|---|---|
| OP | R | Env | ||||||
| 1 | DSH | 1 | Female | 4 | None | None | None | Neg/Neg (54) |
| 2 | DSH | 10 | Male | 6 | None | None | None | Neg/Neg (62) |
| 3 | DSH | 3 | Female | 5 | Day 2: 35.6 | Day 1: 35.7 | Day 1: 33.6 Day 2: 34.2 | Pos/Pos (47) |
| 4 | DSH | 1 | Male | 8 | None | None | None | Neg/Neg (51) |
| 5 | DSH | 6 | Male | 7 | Day 3: 36.7 | None | Day 6: 37.5 | Neg/Neg (174) |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Bessière, P.; Fusade-Boyer, M.; Walch, M.; Lèbre, L.; Brun, J.; Croville, G.; Boullier, S.; Cadiergues, M.-C.; Guérin, J.-L. Household Cases Suggest That Cats Belonging to Owners with COVID-19 Have a Limited Role in Virus Transmission. Viruses 2021, 13, 673. https://doi.org/10.3390/v13040673
Bessière P, Fusade-Boyer M, Walch M, Lèbre L, Brun J, Croville G, Boullier S, Cadiergues M-C, Guérin J-L. Household Cases Suggest That Cats Belonging to Owners with COVID-19 Have a Limited Role in Virus Transmission. Viruses. 2021; 13(4):673. https://doi.org/10.3390/v13040673
Chicago/Turabian StyleBessière, Pierre, Maxime Fusade-Boyer, Mathilda Walch, Laetitia Lèbre, Jessie Brun, Guillaume Croville, Séverine Boullier, Marie-Christine Cadiergues, and Jean-Luc Guérin. 2021. "Household Cases Suggest That Cats Belonging to Owners with COVID-19 Have a Limited Role in Virus Transmission" Viruses 13, no. 4: 673. https://doi.org/10.3390/v13040673
APA StyleBessière, P., Fusade-Boyer, M., Walch, M., Lèbre, L., Brun, J., Croville, G., Boullier, S., Cadiergues, M.-C., & Guérin, J.-L. (2021). Household Cases Suggest That Cats Belonging to Owners with COVID-19 Have a Limited Role in Virus Transmission. Viruses, 13(4), 673. https://doi.org/10.3390/v13040673







