First Report on Natural Infection of Nodavirus in an Echinodermata, Sea Cucumber (Apostichopus japonicas)
Abstract
1. Introduction
2. Materials and Methods
2.1. Sample Collection
2.2. Total RNA Extraction
2.3. Reverse Transcription-Nested PCR (RT-nPCR)
2.4. Sequence Alignment and Phylogenetic Tree Analysis
2.5. In Situ Hybridization (ISH) and Histopathological Analysis
2.6. Transmission Electron Microscopy Analysis
3. Results
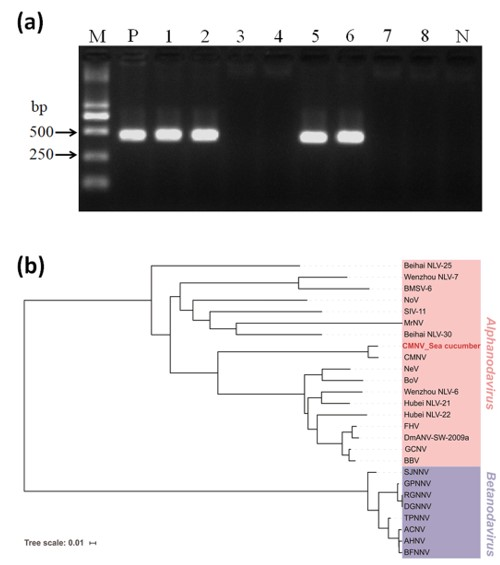
3.1. Detection of CMNV in Sea Cucumber by RT-nPCR
3.2. Phylogenetic Analyses
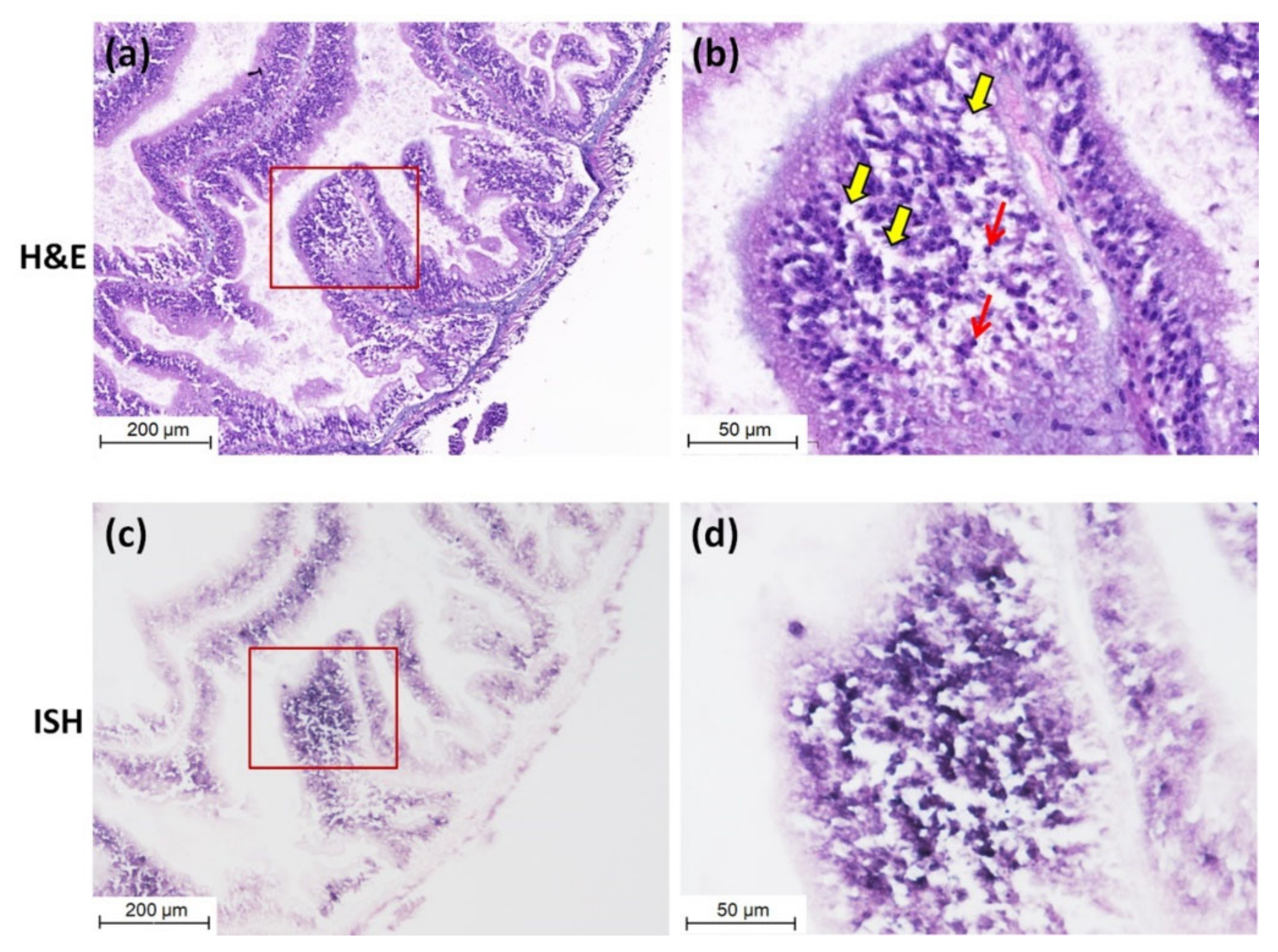
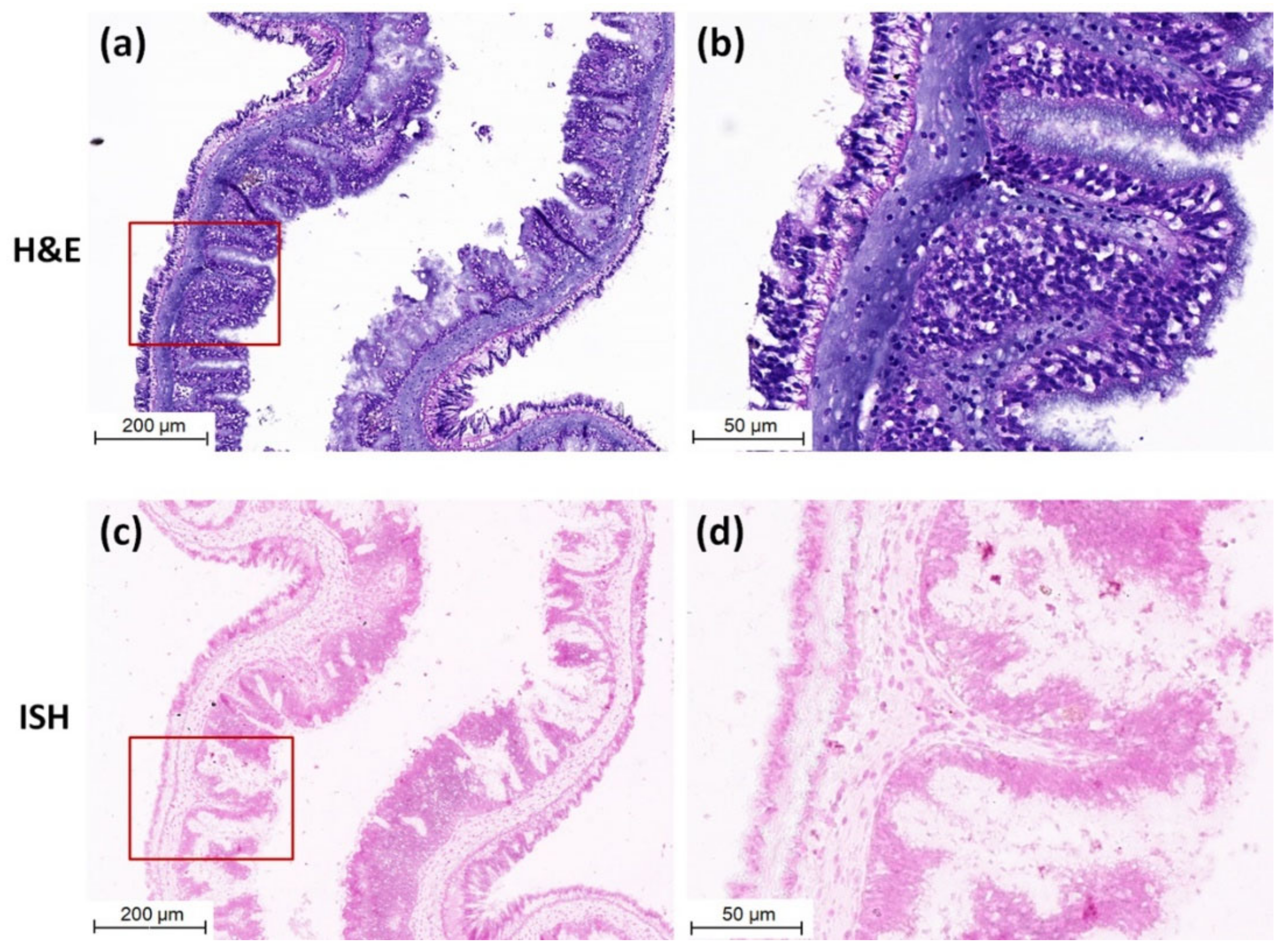
3.3. Detection of CMNV in Sea Cucumber by ISH and Histopathological Analysis
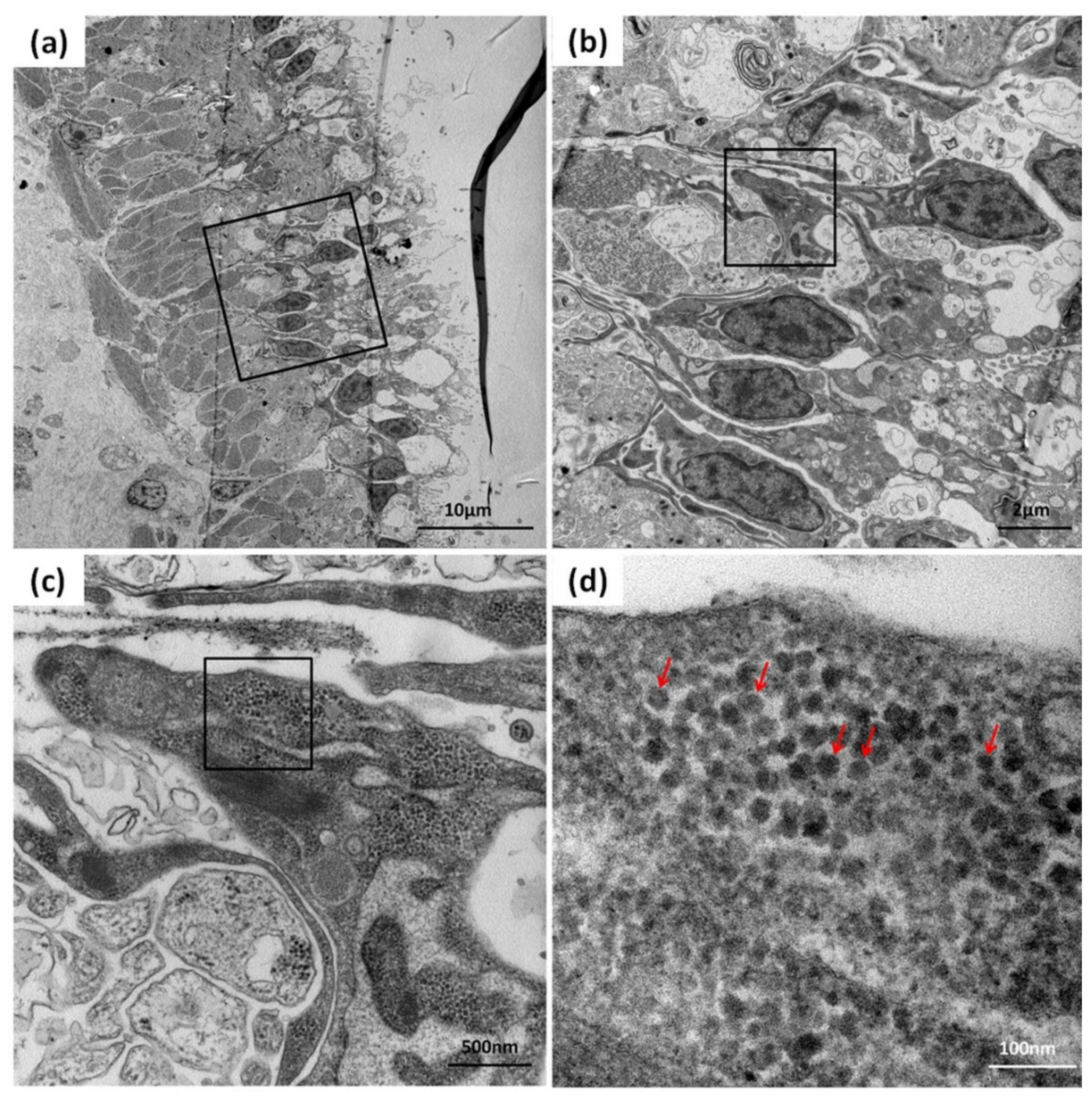
3.4. Detection of CMNV in Sea Cucumber by TEM Analysis
4. Discussion
5. Conclusions
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Conflicts of Interest
References
- Schneider-Schaulies, J. Cellular receptors for viruses: Links to tropism and pathogenesis. J. Gen. Virol. 2000, 81, 1413–1429. [Google Scholar] [CrossRef] [PubMed]
- Kuiken, T.; Holmes, E.C.; McCauley, J.; Rimmelzwaan, G.F.; Williams, C.S.; Grenfell, B.T. Host Species Barriers to Influenza Virus Infections. Science 2006, 312, 394–397. [Google Scholar] [CrossRef] [PubMed]
- Leopardi, S.; Holmes, E.C.; Gastaldelli, M.; Tassoni, L.; Priori, P.; Scaravelli, D.; Zamperin, G.; De Benedictis, P. Interplay between co-divergence and cross-species transmission in the evolutionary history of bat coronaviruses. Infect. Genet. Evol. 2018, 58, 279–289. [Google Scholar] [CrossRef] [PubMed]
- Parrish, C.R.; Holmes, E.C.; Morens, D.M.; Park, E.-C.; Burke, D.S.; Calisher, C.H.; Laughlin, C.A.; Saif, L.J.; Daszak, P. Cross-Species Virus Transmission and the Emergence of New Epidemic Diseases. Microbiol. Mol. Biol. Rev. 2008, 72, 457–470. [Google Scholar] [CrossRef]
- Johnson, C.K.; Hitchens, P.L.; Evans, T.S.; Goldstein, T.; Thomas, K.; Clements, A.; Joly, D.O.; Wolfe, N.D.; Daszak, P.; Karesh, W.B.; et al. Spillover and pandemic properties of zoonotic viruses with high host plasticity. Sci. Rep. 2015, 5, 14830. [Google Scholar] [CrossRef]
- Holmes, E.C. The Evolution and Emergence of RNA Viruses; Oxford University Press: Oxford, UK, 2009. [Google Scholar]
- Wang, L.-F.; Anderson, D.E. Viruses in bats and potential spillover to animals and humans. Curr. Opin. Virol. 2019, 34, 79–89. [Google Scholar] [CrossRef] [PubMed]
- Bourret, V.; Lyall, J.; Frost, S.D.W.; Teillaud, A.; Smith, C.A.; LeClaire, S.; Fu, J.; Gandon, S.; Guérin, J.-L.; Tiley, L.S. Adaptation of avian influenza virus to a swine host. Virus Evol. 2017, 3, vex007. [Google Scholar] [CrossRef] [PubMed]
- Ding, N.-Z.; Xu, D.-S.; Sun, Y.-Y.; He, H.-B.; He, C.-Q. A permanent host shift of rabies virus from Chiroptera to Carnivora associated with recombination. Sci. Rep. 2017, 7, 1–9. [Google Scholar] [CrossRef]
- Owens, F.; Di Serio, F.; Li, S.; Pallás, V.; Randles, J.; Sano, V. Virus Taxonomy: Ninth Report of the International Committee on Taxonomy of Viruses; Academic Press: San Diego, CA, USA, 2012; pp. 1221–1234. [Google Scholar]
- Johnson, K.L.; Price, B.D.; Ball, L.A. Recovery of Infectivity from cDNA Clones of Nodamura Virus and Identification of Small Nonstructural Proteins. Virology 2003, 305, 436–451. [Google Scholar] [CrossRef]
- Selling, B.H.; Allison, R.F.; Kaesberg, P. Genomic RNA of an insect virus directs synthesis of infectious virions in plants. Proc. Natl. Acad. Sci. USA 1990, 87, 434–438. [Google Scholar] [CrossRef]
- Hurd, H. Review of “Insect Pathogens: Molecular Approaches and Techniques” by S. P. Stock, J. Vandenberg, I. Glazer and N. Boemare. Parasites Vectors 2009, 2, 28. [Google Scholar] [CrossRef]
- Scherer, W.F.; Verna, J.E.; Richter, G.W. Nodamura Virus, an Ether- and Chloroform-Resistant Arbovirus from Japan. Am. J. Trop. Med. Hyg. 1968, 17, 120–128. [Google Scholar] [CrossRef]
- Ball, L.A.; Johnson, K.L. Reverse Genetics of Nodaviruses. Adv. Appl. Microbiol. 1999, 53, 229–244. [Google Scholar] [CrossRef]
- Scotti, P.D.; Dearing, S.; Mossop, D.W. Flock house virus: A Nodavirus isolated fromCostelytra zealandica (White) (Coleoptera: Scarabaeida). Arch. Virol. 1983, 75, 181–189. [Google Scholar] [CrossRef]
- Iwamoto, T.; Okinaka, Y.; Mise, K.; Mori, K.-I.; Arimoto, M.; Okuno, T.; Nakai, T. Identification of Host-Specificity Determinants in Betanodaviruses by Using Reassortants between Striped Jack Nervous Necrosis Virus and Sevenband Grouper Nervous Necrosis Virus. J. Virol. 2004, 78, 1256–1262. [Google Scholar] [CrossRef] [PubMed]
- Furusawa, R.; Okinaka, Y.; Uematsu, K.; Nakai, T. Screening of freshwater fish species for their susceptibility to a betanodavirus. Dis. Aquat. Org. 2007, 77, 119–125. [Google Scholar] [CrossRef] [PubMed]
- Souto, S.; Lopez-Jimena, B.; Alonso, M.; García-Rosado, E.; Bandín, I. Experimental susceptibility of European sea bass and Senegalese sole to different betanodavirus isolates. Veter. Microbiol. 2015, 177, 53–61. [Google Scholar] [CrossRef]
- Zhang, Q.; Liu, Q.; Liu, S.; Yang, H.; Liu, S.; Zhu, L.; Yang, B.; Jin, J.; Ding, L.; Wang, X.; et al. A new nodavirus is associated with covert mortality disease of shrimp. J. Gen. Virol. 2014, 95, 2700–2709. [Google Scholar] [CrossRef]
- Zhang, Q.; Xu, T.; Wan, X.; Liu, S.; Wang, X.; Li, X.; Dong, X.; Yang, B.; Huang, J. Prevalence and distribution of covert mortality nodavirus (CMNV) in cultured crustacean. Virus Res. 2017, 233, 113–119. [Google Scholar] [CrossRef]
- Thitamadee, S.; Prachumwat, A.; Srisala, J.; Jaroenlak, P.; Salachan, P.V.; Sritunyalucksana, K.; Flegel, T.W.; Itsathitphaisarn, O. Review of current disease threats for cultivated penaeid shrimp in Asia. Aquaculture 2016, 452, 69–87. [Google Scholar] [CrossRef]
- Pooljun, C.; Direkbusarakom, S.; Chotipuntu, P.; Hirono, I.; Wuthisuthimethavee, S. Development of a TaqMan real-time RT-PCR assay for detection of covert mortality nodavirus (CMNV) in penaeid shrimp. Aquaculture 2016, 464, 445–450. [Google Scholar] [CrossRef]
- Liu, S.; Wang, X.; Xu, T.; Li, X.; Du, L.; Zhang, Q. Vectors and reservoir hosts of covert mortality nodavirus (CMNV) in shrimp ponds. J. Invertebr. Pathol. 2018, 154, 29–36. [Google Scholar] [CrossRef] [PubMed]
- Wang, C.; Wang, X.H.; Liu, S.; Sang, S.W.; Zhang, Q.L. Preliminary study on the natural infection of Carassius auratus with covert mortality nodavirus (CMNV). Prog. Fish. Sci. 2019, 40, 25–32. [Google Scholar]
- Zhang, Q.L.; Liu, S.; Li, J.; Xu, T.T.; Wang, X.H.; Fu, G.M.; Li, X.P.; Sang, S.W.; Bian, X.D.; Hao, J.W. Evidence for Cross-Species Transmission of Covert Mortality Nodavirus to New Host of Mugilogobius abei. Front. Microbiol. 2018, 9, 1447. [Google Scholar] [CrossRef]
- Byrne, M.; Rowe, F.; Uthicke, S. Molecular taxonomy, phylogeny and evolution in the family Stichopodidae (Aspidochirotida: Holothuroidea) based on COI and 16S mitochondrial DNA. Mol. Phylogenetics Evol. 2010, 56, 1068–1081. [Google Scholar] [CrossRef]
- Li, J.Y. The Immunological Characters and Pathogenic Study of Cultured Apostichopus japonicus. Ph.D. Thesis, Ocean University of China, Qingdao, China, 2007. [Google Scholar]
- McElroy, S. Beche-de-mer species of commercial value—An update. SPC Beche-De-Mer Info. Bull. 1990, 2, 2–7. [Google Scholar]
- Han, H.; Yi, Y.H.; Li, L.; Liu, B.S.; Pan, M.X.; Yan, B.; Wang, X.H. Triterpene glycosides from sea cucumber Holothuria leucospilota. Chiu J. Nat. Med. 2009, 7, 346–350. [Google Scholar] [CrossRef]
- Tamura, K.; Stecher, G.; Peterson, D.; Filipski, A.; Kumar, S. MEGA6: Molecular Evolutionary Genetics Analysis Version 6.0. Mol. Biol. Evol. 2013, 30, 2725–2729. [Google Scholar] [CrossRef]
- Bell, T.A.; Lightner, D.V. A Handbook of Normal Penaeid Shrimp Histology; World Aquaculture Society: Baton Rouge, LA, USA, 1988. [Google Scholar]
- Lightner, D.V. A Handbook of Shrimp Pathology and Diagnostic Procedures for Diseases of Cultured Penaeid Shrimp; World Aquaculture Society: Baton Rouge, LA, USA, 1996. [Google Scholar]
- Nuovo, G.J.; Plaia, T.W.; Belinsky, S.A.; Baylin, S.B.; Herman, J.G. In situ detection of the hypermethylation-induced inactivation of the p16 gene as an early event in oncogenesis. Proc. Natl. Acad. Sci. USA 1999, 96, 12754–12759. [Google Scholar] [CrossRef] [PubMed]
- Fortunato, T.M.; Beltrami, C.; Emanueli, C.; De Bank, P.A.; Pula, G. Platelet lysate gel and endothelial progenitors stimulate microvascular network formation in vitro: Tissue engineering implications. Sci. Rep. 2016, 6, 25326. [Google Scholar] [CrossRef] [PubMed]
- Zechmann, B.; Zellnig, G. Rapid diagnosis of plant virus diseases by transmission electron microscopy. J. Virol. Methods 2009, 162, 163–169. [Google Scholar] [CrossRef]
- Graham, L.; Orenstein, J.M. Processing tissue and cells for transmission electron microscopy in diagnostic pathology and research. Nat. Protoc. 2007, 2, 2439–2450. [Google Scholar] [CrossRef]
- Panphut, W.; Senapin, S.; Sriurairatana, S.; Withyachumnarnkul, B.; Flegel, T.W. A novel integrase-containing element may interact with Laem-Singh virus (LSNV) to cause slow growth in giant tiger shrimp. BMC Veter. Res. 2011, 7, 18. [Google Scholar] [CrossRef] [PubMed]
- Pauly, M.; Snoeck, C.J.; Phoutana, V.; Keosengthong, A.; Sausy, A.; Khenkha, L.; Nouanthong, P.; Samountry, B.; Jutavijittum, P.; Vilivong, K.; et al. Cross-species transmission of poultry pathogens in backyard farms: Ducks as carriers of chicken viruses. Avian Pathol. 2019, 48, 503–511. [Google Scholar] [CrossRef] [PubMed]
- Li, X.; Zai, J.; Zhao, Q.; Nie, Q.; Li, Y.; Foley, B.T.; Chaillon, A. Evolutionary history, potential intermediate animal host, and cross-species analyses of SARS-CoV-2. J. Med Virol. 2020, 92, 602–611. [Google Scholar] [CrossRef] [PubMed]
- Zhao, Y.; Guo, M.; Lv, Z.; Zhang, W.; Shao, Y.; Zhao, X.; Li, C. Fas-associated death domain (FADD) in sea cucumber (Apostichopus japonicus): Molecular cloning, characterization and pro-apoptotic function analysis. Dev. Comp. Immunol. 2020, 108, 103673. [Google Scholar] [CrossRef] [PubMed]
- Li, H.; Qiao, G.; Gu, J.; Zhou, W.; Li, Q.; Woo, S.; Xu, D.; Park, S. Phenotypic and genetic characterization of bacteria isolated from diseased cultured sea cucumber Apostichopus japonicus in northeastern China. Dis. Aquat. Org. 2010, 91, 223–235. [Google Scholar] [CrossRef]
- Shao, Y.; Li, C.; Ou, C.; Zhang, P.; Lu, Y.; Su, X.; Li, Y.; Li, T. Divergent Metabolic Responses of Apostichopus japonicus Suffered from Skin Ulceration Syndrome and Pathogen Challenge. J. Agric. Food Chem. 2013, 61, 10766–10771. [Google Scholar] [CrossRef] [PubMed]
- Yang, G.; Tian, X.; Dong, S.; Peng, M.; Wang, D.; Zhang, K. Effects of dietary rhubarb, Bacillus cereus, yeast polysaccharide, and florfenicol supplementation on growth, intestinal morphology, and immune responses of sea cucumber (Apostichopus japonicus). Aquac. Int. 2015, 24, 675–690. [Google Scholar] [CrossRef]
- Song, Z.R. Pathology of Aquatic Animal; Xiamen University Press: Xiamen, China, 2009. [Google Scholar]
- Deng, H.; Zhou, Z.-C.; Wang, N.-B.; Liu, C. The syndrome of sea cucumber (Apostichopus japonicus) infected by virus and bacteria. Virol. Sin. 2008, 23, 63–67. [Google Scholar] [CrossRef]
- Morgan, A. The effect of food availability on early growth, development and survival of the sea cucumber Holothuria scabra (Echinodermata: Holothuroidea). SPC Beche-De-Mer Inf. Bull. 2001, 14, 6–12. [Google Scholar]
- Zhang, C.Y.; Wang, Y.G.; Rong, X.J. Isolation and identification of causative pathogen for skin ulcerative syndrome in Apostichopus japonicus. J. Fish. China 2006, 30, 118–123. [Google Scholar]
- Zhang, Z.; Xing, R.; Lv, Z.; Shao, Y.; Zhang, W.; Zhao, X.; Li, C. Analysis of gut microbiota revealed Lactococcus garviaeae could be an indicative of skin ulceration syndrome in farmed sea cucumber Apostichopus japonicus. Fish Shellfish Immunol. 2018, 80, 148–154. [Google Scholar] [CrossRef] [PubMed]
- Wang, P.; Chang, Y.; Yu, J.; Li, C.; Xu, G. Acute peristome edema disease in juvenile and adult sea cucumbers Apostichopus japonicus (Selenka) reared in North China. J. Invertebr. Pathol. 2007, 96, 11–17. [Google Scholar] [CrossRef] [PubMed]
- Wang, G.X.; Yuan, J.L.; Zhao, Y.K.; Yuan, M. Isolation, identification and drug sensitivity of the pathogens of the skin ulceration disease in Apostichopus japonicus. J. Northwest A F Univ. (Nat. Sci. Ed.) 2007, 35, 87–90. [Google Scholar]
- Nerva, L.; Forgia, M.; Ciuffo, M.; Chitarra, W.; Chiapello, M.; Vallino, M.; Varese, G.; Turina, M. The mycovirome of a fungal collection from the sea cucumber Holothuria polii. Virus Res. 2019, 273, 197737. [Google Scholar] [CrossRef]
- MacTavish, T.; Stenton-Dozey, J.; Vopel, K.; Savage, C. Deposit-Feeding Sea Cucumbers Enhance Mineralization and Nutrient Cycling in Organically-Enriched Coastal Sediments. PLoS ONE 2012, 7, e50031. [Google Scholar] [CrossRef]
- Li, L.; Li, Q. Effects of stocking density, temperature, and salinity on larval survival and growth of the red race of the sea cucumber Apostichopus japonicus (Selenka). Aquac. Int. 2009, 18, 447–460. [Google Scholar] [CrossRef]
- Liu, X.F.; Xue, C.H.; Wang, Y.M.; Li, H.Y. Comparative analysis of nutritive composition in body wall and internal organs of sea cucumber (Apostichopus japonicus)at Rushan. J. Fish. China 2011, 35, 587–593. [Google Scholar]
- Yang, H.; Hamel, J.F.; Mercier, A. The Sea Cucumber Apostichopus japonicus: History, Biology and Aquaculture; Academic Press: Cambridge, MA, USA, 2015. [Google Scholar]
- Purcell, S.W.; Lovatelli, A.; Vasconcellos, M.; Ye, Y. Managing Sea Cucumber Fisheries with An Ecosystem Approach; FAO Fisheries and Aquaculture Technical Paper 520; FAO: Rome, Italy; p. 157.
- Gu, M.; Ma, H.; Mai, K.; Zhang, W.; Ai, Q.; Wang, X.; Bai, N. Immune response of sea cucumber Apostichopus japonicus coelomocytes to several immunostimulants in vitro. Aquaculture 2010, 306, 49–56. [Google Scholar] [CrossRef]
- Liu, H.; Zheng, F.; Sun, X.; Hong, X.; Dong, S.; Wang, B.; Tang, X.; Wang, Y. Identification of the pathogens associated with skin ulceration and peristome tumescence in cultured sea cucumbers Apostichopus japonicus (Selenka). J. Invertebr. Pathol. 2010, 105, 236–242. [Google Scholar] [CrossRef] [PubMed]


| Virus | Abbreviation | GenBank No. * |
|---|---|---|
| Covert mortality nodavirus | CMNV | AIL48199.1 |
| Flock House virus | FHV | AEQ39075.1 |
| Gungahlin Chrysomya noda-like virus | GCNV | QIJ70031.1 |
| Newington virus | NeV | AMO03244.1 |
| Drosophila melanogaster American nodavirus (ANV) strain SW-2009a | DmANV-SW-2009a | ACU32794.1 |
| Black beetle virus | BBV | YP_053043.1 |
| Wenzhou noda-like virus 6 strain | Wenzhou NLV-6 | APG76600.1 |
| Hubei noda-like virus 21 strain | Hubei NLV-21 | APG76486.1 |
| Hubei noda-like virus 22 strain | Hubei NLV-22 | APG76466.1 |
| Boolarra virus | BoV | NP_689439.1 |
| Nodamura virus | NoV | NP_077730.1 |
| Shuangao insect virus 11 strain | SIV-11 | YP_009337806.1 |
| Wenzhou noda-like virus 7 strain | Wenzhou NLV-7 | APG76642.1 |
| Beihai noda-like virus 25 strain | Beihai NLV-25 | APG76164.1 |
| Beihai mantis shrimp virus 6 strain | BMSV-6 | YP_009333376.1 |
| Macrobrachium rosenbergii nodavirus | MrNV | AAQ83832.1 |
| Beihai noda-like virus 30 strain | Beihai NLV-30 | APG76125.1 |
| Striped jack nervous necrosis virus | SJNNV | NP_599247 |
| Golden pompano nervous necrosis virus | GPNNV | ACX54065 |
| Redspotted grouper nervous necrosis virus | RGNNV | ACX69744 |
| Dragon grouper nervous necrosis virus | DGNNV | AAU85148 |
| Tiger puffer nervous necrosis virus | TPNNV | YP_00328875 |
| Atlantic cod nodavirus | ACNV | ABR23192 |
| Atlantic halibut nodavirus | AHNV | AAY34458 |
| Barfin flounder nervous necrosis virus | BFNNV | YP_003288756 |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Wang, C.; Yao, L.; Wang, W.; Sang, S.; Hao, J.; Li, C.; Zhang, Q. First Report on Natural Infection of Nodavirus in an Echinodermata, Sea Cucumber (Apostichopus japonicas). Viruses 2021, 13, 636. https://doi.org/10.3390/v13040636
Wang C, Yao L, Wang W, Sang S, Hao J, Li C, Zhang Q. First Report on Natural Infection of Nodavirus in an Echinodermata, Sea Cucumber (Apostichopus japonicas). Viruses. 2021; 13(4):636. https://doi.org/10.3390/v13040636
Chicago/Turabian StyleWang, Chong, Liang Yao, Wei Wang, Songwen Sang, Jingwei Hao, Chenghua Li, and Qingli Zhang. 2021. "First Report on Natural Infection of Nodavirus in an Echinodermata, Sea Cucumber (Apostichopus japonicas)" Viruses 13, no. 4: 636. https://doi.org/10.3390/v13040636
APA StyleWang, C., Yao, L., Wang, W., Sang, S., Hao, J., Li, C., & Zhang, Q. (2021). First Report on Natural Infection of Nodavirus in an Echinodermata, Sea Cucumber (Apostichopus japonicas). Viruses, 13(4), 636. https://doi.org/10.3390/v13040636





