Isolation and Characterisation of Bacteriophages with Activity against Invasive Non-Typhoidal Salmonella Causing Bloodstream Infection in Malawi
Abstract
1. Introduction
2. Materials and Methods
2.1. Cloning Procedures
2.2. Construction of Mutant Hosts
2.3. Phage Isolation
2.4. Strain Selection and Phage Host Range
2.5. RAPD PCR
2.6. Phage DNA Extraction and Sequencing
2.7. Comparative Genomics and Phylogenetic Analyses
2.8. Transmission Electron Microscopy (TEM)
3. Results and Discussion
3.1. Phage Isolation
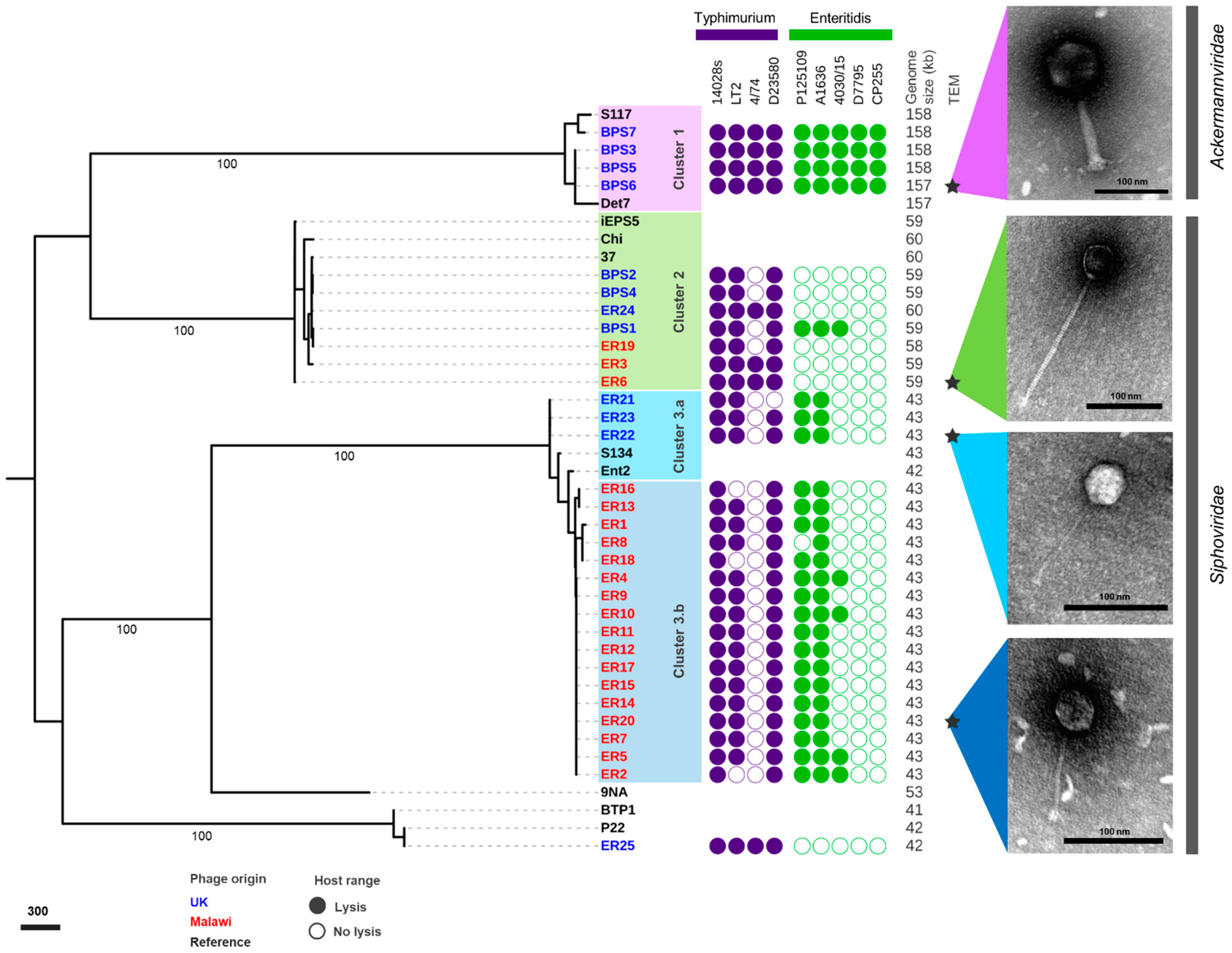
3.2. Host Range Analysis
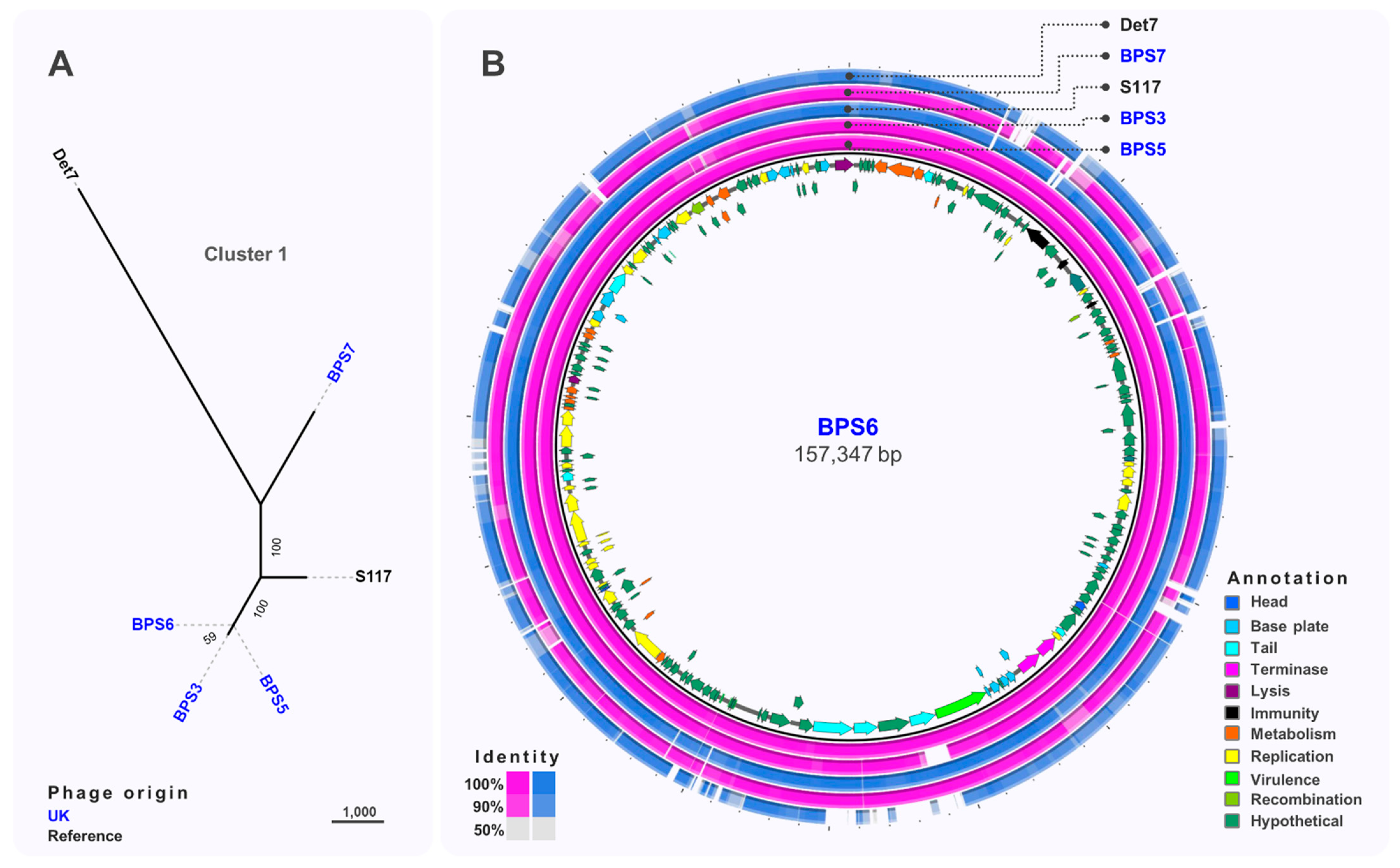
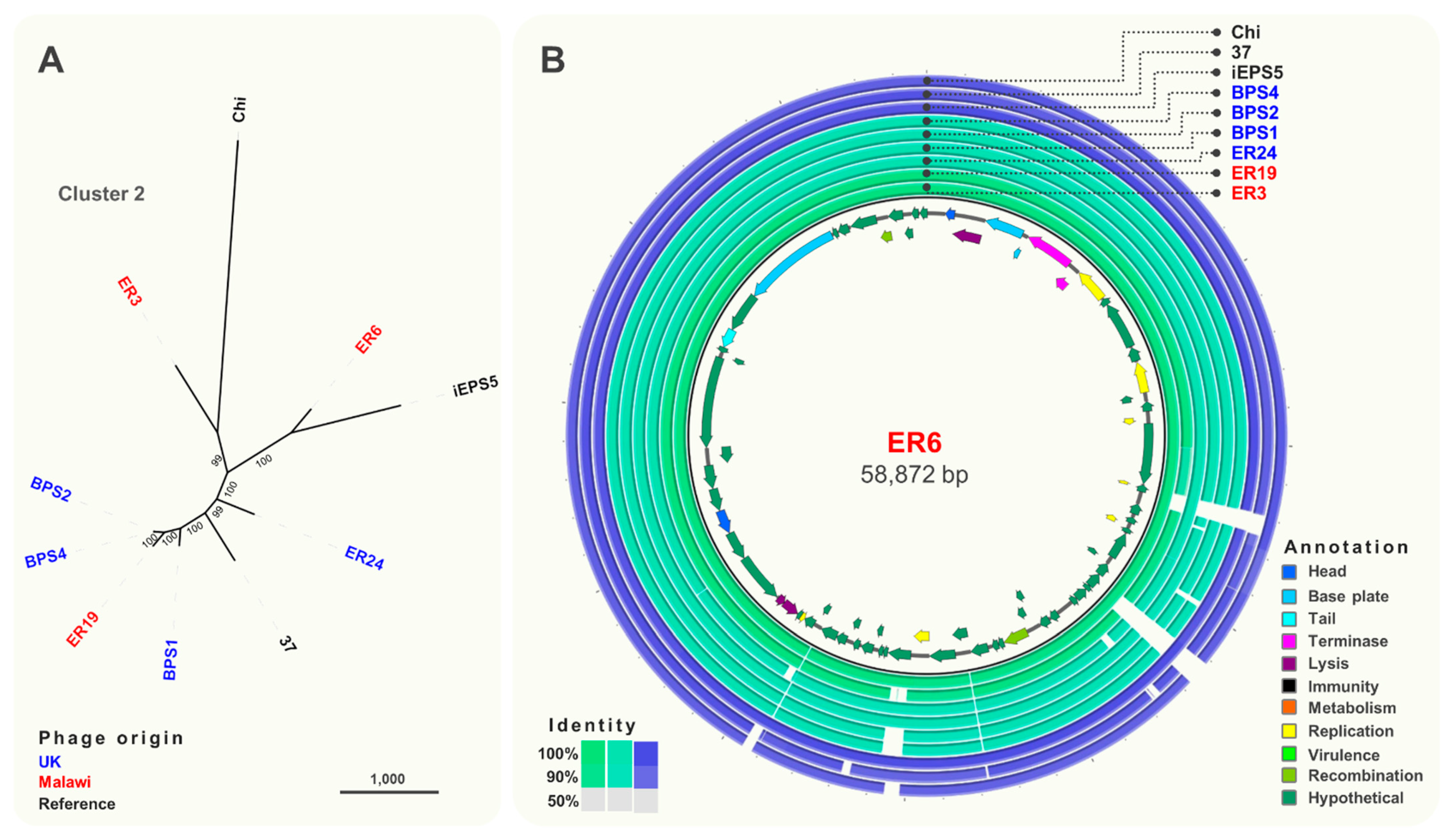
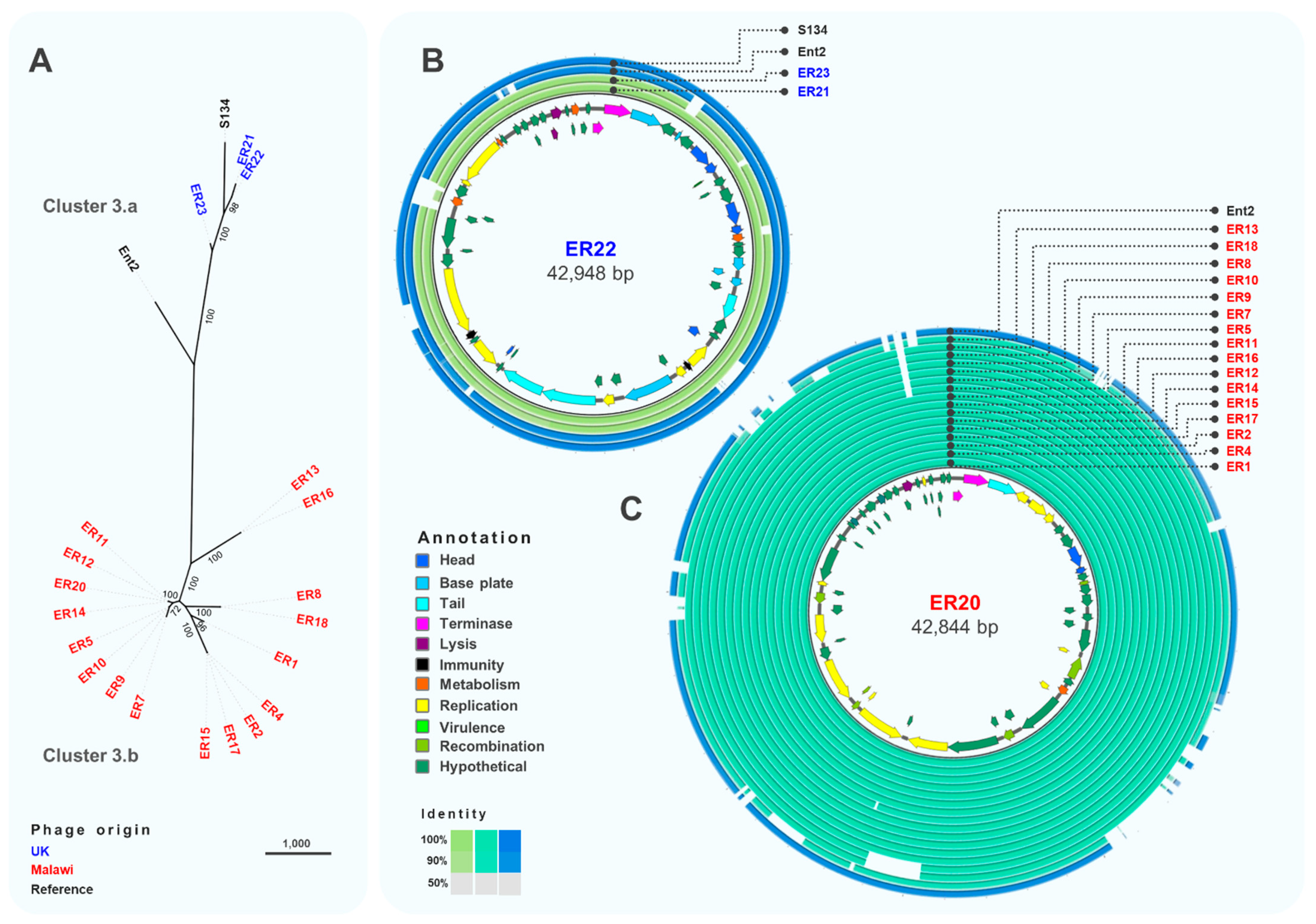
3.3. Comparative Genomics and Phylogeny of Isolated Phages
3.4. Perspective
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Stanaway, J.D.; Parisi, A.; Sarkar, K.; Blacker, B.F.; Reiner, R.C.; Hay, S.I.; Nixon, M.R.; Dolecek, C.; James, S.L.; Mokdad, A.H.; et al. The global burden of non-typhoidal Salmonella invasive disease: A systematic analysis for the Global Burden of Disease Study 2017. Lancet Infect. Dis. 2019, 19, 1312–1324. [Google Scholar] [CrossRef]
- Okoro, C.K.; Kingsley, R.A.; Quail, M.A.; Kankwatira, A.M.; Feasey, N.A.; Parkhill, J.; Dougan, G.; Gordon, M.A. High-resolution single nucleotide polymorphism analysis distinguishes recrudescence and reinfection in recurrent invasive nontyphoidal salmonella typhimurium disease. Clin. Infect. Dis. 2012, 54, 955–963. [Google Scholar] [CrossRef] [PubMed]
- Feasey, N.A.; Hadfield, J.; Keddy, K.H.; Dallman, T.J.; Jacobs, J.; Deng, X.; Wigley, P.; Barquist Barquist, L.; Langridge, G.C.; Feltwell, T.; et al. Distinct Salmonella Enteritidis lineages associated with enterocolitis in high-income settings and invasive disease in low-income settings. Nat. Genet. 2016, 48, 1211–1217. [Google Scholar] [CrossRef]
- Kingsley, R.A.; Msefula, C.L.; Thomson, N.R.; Kariuki, S.; Holt, K.E.; Gordon, M.A.; Harris, D.; Clarke, L.; Whitehead, S.; Sangal, V.; et al. Epidemic multiple drug resistant Salmonella Typhimurium causing invasive disease in sub-Saharan Africa have a distinct genotype. Genome Res. 2009, 19, 2279–2287. [Google Scholar] [CrossRef] [PubMed]
- Okoro, C.K.; Kingsley, R.A.; Connor, T.R.; Harris, S.R.; Parry, C.M.; Al-Mashhadani, M.N.; Kariuki, S.; Msefula, C.L.; Gordon, M.A.; de Pinna, E.; et al. Intracontinental spread of human invasive Salmonella Typhimurium pathovariants in sub-Saharan Africa. Nat. Genet. 2012, 44, 1215–1221. [Google Scholar] [CrossRef]
- Brüssow, H.; Canchaya, C.; Hardt, W.D. Phages and the evolution of bacterial pathogens: From genomic rearrangements to lysogenic conversion. Microbiol. Mol. Biol. Rev. 2004, 68, 560–602. [Google Scholar] [CrossRef] [PubMed]
- Fortier, L.-C.; Sekulovic, O. Importance of prophages to evolution and virulence of bacterial pathogens. Virulence 2013, 4, 354–365. [Google Scholar] [CrossRef]
- Owen, S.V.; Wenner, N.; Canals, R.; Makumi, A.; Hammarlöf, D.L.; Gordon, M.A.; Aertsen, A.; Feasey, N.A.; Hinton, J.C.D. Characterization of the prophage repertoire of African Salmonella Typhimurium ST313 reveals high levels of spontaneous induction of novel phage BTP1. Front. Microbiol. 2017, 8, 235. [Google Scholar] [CrossRef]
- Ashton, P.M.; Owen, S.V.; Kaindama, L.; Rowe, W.P.M.; Lane, C.R.; Larkin, L.; Nair, S.; Jenkins, C.; de Pinna, E.M.; Feasey, N.A.; et al. Public health surveillance in the UK revolutionises our understanding of the invasive Salmonella Typhimurium epidemic in Africa. Genome Med. 2017, 9, 1–13. [Google Scholar] [CrossRef]
- Owen, S.V.; Wenner, N.; Dulberger, C.L.; Rodwell, E.V.; Bowers-Barnard, A.; Quinones-Olvera, N.; Rigden, D.J.; Rubin, E.J.; Garner, E.C.; Baym, M.; et al. Prophage-encoded phage defence proteins with cognate self-immunity. bioRxiv 2020. [Google Scholar] [CrossRef]
- Kintz, E.; Davies, M.R.; Hammarlöf, D.L.; Canals, R.; Hinton, J.C.D.; van der Woude, M.W. A BTP1 prophage gene present in invasive non-typhoidal Salmonella determines composition and length of the O-antigen of the lipopolysaccharide. Mol. Microbiol. 2015, 96, 263–275. [Google Scholar] [CrossRef] [PubMed]
- Bernheim, A.; Sorek, R. The pan-immune system of bacteria: Antiviral defence as a community resource. Nat. Rev. Microbiol. 2020, 18, 113–119. [Google Scholar] [CrossRef] [PubMed]
- Rousset, F.; Dowding, J.; Bernheim, A.; Rocha, E.P.C.; Bikard, D. Prophage-encoded hotspots of bacterial immune systems. bioRxiv 2021. [Google Scholar] [CrossRef]
- Sambrook, J.; Russell, D.W. Molecular Cloning: A Laboratory Manual, 3rd ed.; Cold Spring Harbor Laboratory Press: Cold Spring Harbor, NY, USA, 2001. [Google Scholar]
- Pedulla, M.L.; Ford, M.E.; Karthikeyan, T.; Houtz, J.M.; Hendrix, R.W.; Hatfull, G.F.; Poteete, A.R.; Gilcrease, E.B.; Winn-Stapley, D.A.; Casjens, S.R. Corrected sequence of the bacteriophage P22 genome. J. Bacteriol. 2003. [Google Scholar] [CrossRef] [PubMed][Green Version]
- Schmieger, H. Phage P22-mutants with increased or decreased transduction abilities. Mol. Gen. Genet. MGG 1972, 119, 75–88. [Google Scholar] [CrossRef]
- Casjens, S.R.; Jacobs-Sera, D.; Hatfull, G.F.; Hendrix, R.W. Genome sequence of Salmonella enterica phage Det7. Genome Announc. 2015. [Google Scholar] [CrossRef]
- Casjens, S.R.; Leavitt, J.C.; Hatfull, G.F.; Hendrix, R.W. Genome sequence of Salmonella phage 9NA. Genome Announc. 2014. [Google Scholar] [CrossRef]
- Thomson, N.R.; Clayton, D.J.; Windhorst, D.; Vernikos, G.; Davidson, S.; Churcher, C.; Quail, M.A.; Stevens, M.; Jones, M.A.; Watson, M.; et al. Comparative genome analysis of Salmonella Enteritidis PT4 and Salmonella Gallinarum 287/91 provides insights into evolutionary and host adaptation pathways. Genome Res. 2008, 18, 1624–1637. [Google Scholar] [CrossRef]
- Darboe, S.; Bradbury, R.; Phelan, J.; Kanteh, A.; Muhammad, A.-K.; Worwui, A.; Yang, S.; Nwakanma, D.; Perez-Sepulveda, B.; Kariuki, S.; et al. Genomic diversity and antimicrobial resistance among non-typhoidal Salmonella associated with human disease in The Gambia. bioRxiv 2020. [Google Scholar] [CrossRef]
- Perez-Sepulveda, B.M.; Predeus, A.V.; Fong, W.Y.; Parry, C.M.; Cheesbroughb, J.; Wigley, P.; Feasey, N.A.; Hinton, J.C.D. Complete genome sequences of African Salmonella Enteritidis clinical isolates associated with bloodstream infection. Microbiol. Resour. Announc. 2021, in press. [Google Scholar]
- Fields, P.I.; Swanson, R.V.; Haidaris, C.G.; Heffron, F. Mutants of Salmonella typhimurium that cannot survive within the macrophage are avirulent. Proc. Natl. Acad. Sci. USA 1986, 83, 5189–5193. [Google Scholar] [CrossRef] [PubMed]
- Zinder, N.D.; Lederberg, J. Genetic exchange in Salmonella. J. Bacteriol. 1952, 64, 679–699. [Google Scholar] [CrossRef]
- Rankin, J.D.; Taylor, R.J. The estimation of doses of Salmonella typhimurium suitable for the experimental production of disease in calves. Vet. Rec. 1966, 78, 706–707. [Google Scholar] [CrossRef]
- Simon, R.; Priefer, U.; Pühler, A. A broad host range mobilization system for in vivo genetic engineering: Transposon mutagenesis in gram negative bacteria. Bio/Technology 1983. [Google Scholar] [CrossRef]
- Martínez-García, E.; de Lorenzo, V. Engineering multiple genomic deletions in Gram-negative bacteria: Analysis of the multi-resistant antibiotic profile of Pseudomonas putida KT2440. Environ. Microbiol. 2011. [Google Scholar] [CrossRef]
- Datsenko, K.A.; Wanner, B.L. One-step inactivation of chromosomal genes in Escherichia coli K-12 using PCR products. Proc. Natl. Acad. Sci. USA 2000. [Google Scholar] [CrossRef]
- Koskiniemi, S.; Pränting, M.; Gullberg, E.; Näsvall, J.; Andersson, D.I. Activation of cryptic aminoglycoside resistance in Salmonella enterica. Mol. Microbiol. 2011. [Google Scholar] [CrossRef] [PubMed]
- Doublet, B.; Douard, G.; Targant, H.; Meunier, D.; Madec, J.Y.; Cloeckaert, A. Antibiotic marker modifications of λ Red and FLP helper plasmids, pKD46 and pCP20, for inactivation of chromosomal genes using PCR products in multidrug-resistant strains. J. Microbiol. Methods 2008. [Google Scholar] [CrossRef] [PubMed]
- Heckman, K.L.; Pease, L.R. Gene splicing and mutagenesis by PCR-driven overlap extension. Nat. Protoc. 2007. [Google Scholar] [CrossRef]
- Pelludat, C.; Mirold, S.; Hardt, W.D. The SopEφ phage integrates into the ssrA gene of Salmonella enterica serovar Typhimurium A36 and is closely related to the Fels-2 prophage. J. Bacteriol. 2003. [Google Scholar] [CrossRef] [PubMed]
- Goldfarb, T.; Sberro, H.; Weinstock, E.; Cohen, O.; Doron, S.; Charpak-Amikam, Y.; Afik, S.; Ofir, G.; Sorek, R. BREX is a novel phage resistance system widespread in microbial genomes. EMBO J. 2015, 34, 169–183. [Google Scholar] [CrossRef]
- Hammarlöf, D.L.; Kröger, C.; Owen, S.V.; Canals, R.; Lacharme-Lora, L.; Wenner, N.; Schager, A.E.; Wells, T.J.; Henderson, I.R.; Wigley, P.; et al. Role of a single noncoding nucleotide in the evolution of an epidemic African clade of Salmonella. Proc. Natl. Acad. Sci. USA 2018, 115, E2614–E2623. [Google Scholar] [CrossRef]
- Canals, R.; Hammarlöf, D.L.; Kröger, C.; Owen, S.V.; Fong, W.Y.; Lacharme-Lora, L.; Zhu, X.; Wenner, N.; Carden, S.E.; Honeycutt, J.; et al. Adding function to the genome of African Salmonella Typhimurium ST313 strain D23580. PLoS Biol. 2019, 17, e3000059. [Google Scholar] [CrossRef]
- Kumwenda, S.; Kambala, C. Determination of Biological, Physical and Chemical Pollutants in Mudi River, Blantyre, Malawi RBF4MNH Evaluation View project. J. Basic Appl. Sci. Res. 2012, 2, 6833–6839. [Google Scholar]
- Kropinski, A.M.; Mazzocco, A.; Waddell, T.E.; Lingohr, E.; Johnson, R. Enumeration of Bacteriophages by Double Agar Overlay Plaque Assay. In Bacteriophages: Methods and Protocols, Volume 1: Isolation, Characterization, and Interactions; Kropinski, A.M., Clokie, M.R.J., Eds.; Humana Press: Totowa, NJ, USA, 2009; pp. 69–76. ISBN 978-1-58829-682-5. [Google Scholar]
- Mazzocco, A.; Waddell, T.E.; Lingohr, E.; Johnson, R.P. Enumeration of bacteriophages using the small drop plaque assay system. In Bacteriophages: Methods in Molecular Biology, Volume 1: Isolation, Characterization, and Interactions; Kropinski, A.M., Clokie, M.R.J., Eds.; Humana Press: Totowa, NJ, USA, 2009; pp. 81–85. ISBN 978-1-58829-682-5. [Google Scholar]
- Gutiérrez, D.; Martín-Platero, A.M.; Rodríguez, A.; Martínez-Bueno, M.; García, P.; Martínez, B. Typing of bacteriophages by randomly amplified polymorphic DNA (RAPD)-PCR to assess genetic diversity. FEMS Microbiol. Lett. 2011, 322, 90–97. [Google Scholar] [CrossRef] [PubMed]
- Wick, R.R.; Judd, L.M.; Gorrie, C.L.; Holt, K.E. Unicycler: Resolving bacterial genome assemblies from short and long sequencing reads. PLoS Comput. Biol. 2017, 13, e1005595. [Google Scholar] [CrossRef] [PubMed]
- Gurevich, A.; Saveliev, V.; Vyahhi, N.; Tesler, G. QUAST: Quality assessment tool for genome assemblies. Bioinformatics 2013, 29, 1072–1075. [Google Scholar] [CrossRef]
- Carver, T.; Harris, S.R.; Berriman, M.; Parkhill, J.; McQuillan, J.A. Artemis: An integrated platform for visualization and analysis of high-throughput sequence-based experimental data. Bioinformatics 2012, 28, 464–469. [Google Scholar] [CrossRef]
- Altschul, S.F.; Gish, W.; Miller, W.; Myers, E.W.; Lipman, D.J. Basic local alignment search tool. J. Mol. Biol. 1990. [Google Scholar] [CrossRef]
- Seemann, T. Prokka: Rapid prokaryotic genome annotation. Bioinformatics 2014, 30, 2068–2069. [Google Scholar] [CrossRef]
- Tynecki, P.; Guziński, A.; Kazimierczak, J.; Jadczuk, M.; Dastych, J.; Onisko, A. PhageAI—Bacteriophage life cycle recognition with machine learning and natural language processing. bioRxiv 2020. [Google Scholar] [CrossRef]
- Adriaenssens, E.; Brister, J.R. How to Name and Classify Your Phage: An Informal Guide. Viruses 2017, 9, 70. [Google Scholar] [CrossRef] [PubMed]
- Sievers, F.; Wilm, A.; Dineen, D.; Gibson, T.J.; Karplus, K.; Li, W.; Lopez, R.; McWilliam, H.; Remmert, M.; Söding, J.; et al. Fast, scalable generation of high-quality protein multiple sequence alignments using Clustal Omega. Mol. Syst. Biol. 2011, 7, 539. [Google Scholar] [CrossRef] [PubMed]
- Stamatakis, A. RAxML version 8: A tool for phylogenetic analysis and post-analysis of large phylogenies. Bioinformatics 2014, 30, 1312–1313. [Google Scholar] [CrossRef]
- Letunic, I.; Bork, P. Interactive Tree of Life (iTOL) v4: Recent updates and new developments. Nucleic Acids Res. 2019, 47, W256–W259. [Google Scholar] [CrossRef]
- Moraru, C.; Varsani, A.; Kropinski, A.M. VIRIDIC—A Novel Tool to Calculate the Intergenomic Similarities of Prokaryote-Infecting Viruses. Viruses 2020, 12, 1268. [Google Scholar] [CrossRef] [PubMed]
- Page, A.J.; Cummins, C.A.; Hunt, M.; Wong, V.K.; Reuter, S.; Holden, M.T.G.; Fookes, M.; Falush, D.; Keane, J.A.; Parkhill, J. Roary: Rapid large-scale prokaryote pan genome analysis. Bioinformatics 2015, 31, 3691–3693. [Google Scholar] [CrossRef] [PubMed]
- Tamura, K.; Stecher, G.; Peterson, D.; Filipski, A.; Kumar, S. MEGA6: Molecular Evolutionary Genetics Analysis Version 6.0. Mol. Biol. Evol. 2013, 30, 2725–2729. [Google Scholar] [CrossRef]
- Alikhan, N.F.; Petty, N.K.; Ben Zakour, N.L.; Beatson, S.A. BLAST Ring Image Generator (BRIG): Simple prokaryote genome comparisons. BMC Genom. 2011, 12, 402. [Google Scholar] [CrossRef] [PubMed]
- Reddy, E.A.; Shaw, A.V.; Crump, J.A. Community-acquired bloodstream infections in Africa: A systematic review and meta-analysis. Lancet Infect. Dis. 2010, 10, 417–432. [Google Scholar] [CrossRef]
- Labrie, S.J.; Samson, J.E.; Moineau, S. Bacteriophage resistance mechanisms. Nat. Rev. Microbiol. 2010, 8, 317–327. [Google Scholar] [CrossRef] [PubMed]
- Broeker, N.K.; Barbirz, S. MicroCommentary Not a barrier but a key: How bacteriophages exploit host’s O-antigen as an essential receptor to initiate infection. Mol. Microbiol. 2017, 105, 353–357. [Google Scholar] [CrossRef] [PubMed]
- Gencay, Y.E.; Gambino, M.; Prüssing, T.F.; Brøndsted, L. The genera of bacteriophages and their receptors are the major determinants of host range. Environ. Microbiol. 2019, 21, 2095–2111. [Google Scholar] [CrossRef] [PubMed]
- Fong, K.; Tremblay, D.M.; Delaquis, P.; Goodridge, L.; Levesque, R.C.; Moineau, S.; Suttle, C.A.; Wang, S. Diversity and Host Specificity Revealed by Biological Characterization and Whole Genome Sequencing of Bacteriophages Infecting Salmonella enterica. Viruses 2019, 11, 854. [Google Scholar] [CrossRef] [PubMed]
- Choi, Y.; Shin, H.; Lee, J.H.; Ryua, S. Identification and characterization of a novel flagellum-dependent Salmonella-infecting bacteriophage, iEPS5. Appl. Environ. Microbiol. 2013, 79, 4829–4837. [Google Scholar] [CrossRef] [PubMed]
- Meynell, E.W. A phage, øχ, which attacks motile bacteria. J. Gen. Microbiol. 1961, 25, 253–290. [Google Scholar] [CrossRef]
- Karpe, Y.A.; Kanade, G.D.; Pingale, K.D.; Arankalle, V.A.; Banerjee, K. Genomic characterization of Salmonella bacteriophages isolated from India. Virus Genes 2016, 52, 117–126. [Google Scholar] [CrossRef]
- Byl, C.V.; Kropinski, A.M. Sequence of the genome of Salmonella bacteriophage P22. J. Bacteriol. 2000, 182, 6472–6481. [Google Scholar] [CrossRef]
- Grygorcewicz, B.; Grudziński, M.; Wasak, A.; Augustyniak, A.; Pietruszka, A.; Nawrotek, P. Bacteriophage-mediated reduction of Salmonella Enteritidis in swine slurry. Appl. Soil Ecol. 2017, 119, 179–182. [Google Scholar] [CrossRef]
- Woolston, J.; Parks, A.R.; Abuladze, T.; Anderson, B.; Li, M.; Carter, C.; Hanna, L.F.; Heyse, S.; Charbonneau, D.; Sulakvelidze, A. Bacteriophages lytic for Salmonella rapidly reduce Salmonella contamination on glass and stainless steel surfaces. Bacteriophage 2013, 3, e25697. [Google Scholar] [CrossRef]
- Nagel, T.E.; Chan, B.K.; De Vos, D.; El-Shibiny, A.; Kang’ethe, E.K.; Makumi, A.; Pirnay, J.P. The developing world urgently needs phages to combat pathogenic bacteria. Front. Microbiol. 2016, 7, 882. [Google Scholar] [CrossRef] [PubMed]

| Strain | Description a | Reference |
|---|---|---|
| Bacteriophages | ||
| P22 | Podoviridae, temperate, wild type from strain LT2 | [8,15] |
| P22 HT 105/1 int-201 | Podoviridae, virulent transducing phage | [16] |
| BTP1 | Podoviridae, temperate, wild type from strain D23580 | [8] |
| Det7 | Ackermannviridae, virulent, wild type | [17] |
| 9NA | Siphoviridae, virulent, wild type | [18] |
| S. Enteritidis | ||
| P125109 | Wild type | [19] |
| A1636 | Wild type | [3] |
| 4030_15 | Wild type | [20] |
| D7795 | Wild type | [3] |
| CP255 | Wild type | [21] |
| S. Typhimurium | ||
| 14028s | Wild type | [22] |
| LT2 | Wild type | [23] |
| 4/74 derivatives | ||
| 4/74 | Wild type | [24] |
| JH4180 | 4/74 ΔΦ (ΔGifsy-1 ΔGifsy-2 ΔST64B ΔSopEΦ ΔP4-like) | This study |
| D23580 derivatives | ||
| D23580 | Wild type | [4] |
| JH3877 | D23580 ΔBTP1 | [8] |
| JH3949 | D23580 ΔΦ (ΔBTP1 ΔBTP5 ΔGifsy-1 ΔGifsy-2 ΔST64B) | [8] |
| JH4314 | D23580ΔΦ Δbrex::frt (ΔSTMMW_44361- STMMW_44431) | This study |
| JH4655 | D23580ΔΦ Δbrex::frt ΔgalE | This study |
| Escherichia coli | ||
| S17-1 λpir | pro thi hsdR recA chromosome::RP4-2 Tc::Mu Km::Tn7/λpir; Tp R, Sm R | [25] |
| Plasmids | ||
| pEMG | Suicide plasmid; Km R | [26] |
| pNAW1 | pEMG::attBGifsy-2 4/74; Km R | This study |
| pNAW15 | pEMG::attBGifsy-1; Km R | [8] |
| pNAW19 | pEMG::attBSopEΦ&P4-like; Km R | This study |
| pNAW34 | pEMG::∆galE; Km R | This study |
| pNAW42 | pEMG::attBST64-B; Km R | [8] |
| pSW-2 | I-SceI expressing vector, m-toluate inducible; Gm R | [26] |
| pKD4 | frt-aph-frt (Km R cassette flanked with frt) template plasmid; Km R | [27] |
| pSIM5-tet | λ Red recombination plasmid, temperature-inducible; Tc R | [28] |
| pCP20-Gm | Flippase recombinase expression plasmid; Gm R | [29] |
| Common Name | Phage Name | Place of Isolation | Original Host | Cluster a | GenBank Accession Number |
|---|---|---|---|---|---|
| ER1 | vB_SenS_ER1 | Blantyre, Malawi | S. Typhimurium D23580ΔΦbrex | 3.b | MW355461 |
| ER2 | vB_SenS_ER2 | Blantyre, Malawi | S. Typhimurium D23580ΔΦbrex | 3.b | MW355466 |
| ER3 | vB_SenS_ER3 | Blantyre, Malawi | S. Typhimurium D23580ΔΦbrex | 2 | MW355467 |
| ER4 | vB_SenS_ER4 | Blantyre, Malawi | S. Typhimurium D23580ΔΦbrex | 3.b | MW355468 |
| ER5 | vB_SenS_ER5 | Blantyre, Malawi | S. Typhimurium D23580ΔΦbrex | 3.b | MW355469 |
| ER6 | vB_SenS_ER6 | Blantyre, Malawi | S. Typhimurium D23580ΔΦbrex | 2 | MW355470 |
| ER7 | vB_SenS_ER7 | Blantyre, Malawi | S. Typhimurium D23580ΔΦbrex | 3.b | MW355471 |
| ER8 | vB_SenS_ER8 | Blantyre, Malawi | S. Typhimurium D23580ΔΦbrex | 3.b | MW355472 |
| ER9 | vB_SenS_ER9 | Blantyre, Malawi | S. Typhimurium D23580ΔΦbrex | 3.b | MW355473 |
| ER10 | vB_SenS_ER10 | Blantyre, Malawi | S. Typhimurium D23580ΔΦbrex | 3.b | MW355451 |
| ER11 | vB_SenS_ER11 | Blantyre, Malawi | S. Typhimurium D23580ΔΦbrex | 3.b | MW355452 |
| ER12 | vB_SenS_ER12 | Blantyre, Malawi | S. Typhimurium D23580ΔΦbrex | 3.b | MW355453 |
| ER13 | vB_SenS_ER13 | Blantyre, Malawi | S. Typhimurium D23580ΔΦbrex | 3.b | MW355454 |
| ER14 | vB_SenS_ER14 | Blantyre, Malawi | S. Typhimurium D23580ΔΦbrex | 3.b | MW355455 |
| ER15 | vB_SenS_ER15 | Blantyre, Malawi | S. Typhimurium D23580ΔΦbrex | 3.b | MW355456 |
| ER16 | vB_SenS_ER16 | Blantyre, Malawi | S. Typhimurium D23580ΔΦbrex | 3.b | MW355457 |
| ER17 | vB_SenS_ER17 | Blantyre, Malawi | S. Typhimurium D23580ΔΦbrex | 3.b | MW355458 |
| ER18 | vB_SenS_ER18 | Blantyre, Malawi | S. Typhimurium D23580ΔΦbrex | 3.b | MW355459 |
| ER19 | vB_SenS_ER19 | Blantyre, Malawi | S. Typhimurium D23580ΔΦbrex | 2 | MW355460 |
| ER20 | vB_SenS_ER20 | Blantyre, Malawi | S. Typhimurium D23580ΔΦbrex | 3.b | MW355462 |
| ER21 | vB_SenS_ER21 | Liverpool, UK | S. Typhimurium D23580ΔΦbrex | 3.a | MW355463 |
| ER22 | vB_SenS_ER22 | Liverpool, UK | S. Typhimurium D23580ΔΦbrex | 3.a | MW355464 |
| ER23 | vB_SenS_ER23 | Southport, UK | S. Typhimurium D23580ΔΦbrex | 3.a | MW355465 |
| ER24 | vB_SenS_ER24 | Durham, UK | S. Typhimurium D23580ΔΦbrex | 2 | MW355479 |
| ER25 | vB_SenP_ER25 | Durham, UK | S. Typhimurium D23580ΔΦbrex | ND | MW355480 |
| BPS1 | vB_SenS_BPS1 | Liverpool, UK | S. Typhimurium D23580ΔΦ | 2 | MW355474 |
| BPS2 | vB_SenS_BPS2 | Liverpool, UK | S. Typhimurium D23580ΔΦ | 2 | MW355449 |
| BPS3 | vB_SenAc_BPS3 | Liverpool, UK | S. Typhimurium 4/74 | 1 | MW355475 |
| BPS4 | vB_SenS_BPS4 | Liverpool, UK | S. Typhimurium D23580ΔΦ | 2 | MW355476 |
| BPS5 | vB_SenAc_BPS5 | Liverpool, UK | S. Enteritidis D7795 | 1 | MW355477 |
| BPS6 | vB_SenAc_BPS6 | Liverpool, UK | S. Enteritidis D7795 | 1 | MW355450 |
| BPS7 | vB_SenAc_BPS7 | Liverpool, UK | S. Enteritidis P125109 | 1 | MW355478 |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Rodwell, E.V.; Wenner, N.; Pulford, C.V.; Cai, Y.; Bowers-Barnard, A.; Beckett, A.; Rigby, J.; Picton, D.M.; Blower, T.R.; Feasey, N.A.; et al. Isolation and Characterisation of Bacteriophages with Activity against Invasive Non-Typhoidal Salmonella Causing Bloodstream Infection in Malawi. Viruses 2021, 13, 478. https://doi.org/10.3390/v13030478
Rodwell EV, Wenner N, Pulford CV, Cai Y, Bowers-Barnard A, Beckett A, Rigby J, Picton DM, Blower TR, Feasey NA, et al. Isolation and Characterisation of Bacteriophages with Activity against Invasive Non-Typhoidal Salmonella Causing Bloodstream Infection in Malawi. Viruses. 2021; 13(3):478. https://doi.org/10.3390/v13030478
Chicago/Turabian StyleRodwell, Ella V., Nicolas Wenner, Caisey V. Pulford, Yueyi Cai, Arthur Bowers-Barnard, Alison Beckett, Jonathan Rigby, David M. Picton, Tim R. Blower, Nicholas A. Feasey, and et al. 2021. "Isolation and Characterisation of Bacteriophages with Activity against Invasive Non-Typhoidal Salmonella Causing Bloodstream Infection in Malawi" Viruses 13, no. 3: 478. https://doi.org/10.3390/v13030478
APA StyleRodwell, E. V., Wenner, N., Pulford, C. V., Cai, Y., Bowers-Barnard, A., Beckett, A., Rigby, J., Picton, D. M., Blower, T. R., Feasey, N. A., Hinton, J. C. D., & Perez-Sepulveda, B. M. (2021). Isolation and Characterisation of Bacteriophages with Activity against Invasive Non-Typhoidal Salmonella Causing Bloodstream Infection in Malawi. Viruses, 13(3), 478. https://doi.org/10.3390/v13030478








