PEDV Infection Generates Conformation-Specific Antibodies That Can Be Effectively Detected by a Cell-Based ELISA
Abstract
1. Introduction
2. Materials and Methods
2.1. Cell Lines and Cell Culture
2.2. Plasmid Construction
2.3. Recombinant Baculovirus Preparation
2.4. Virus Titer Determination by TCID50
2.5. Western Blot Analysis
2.6. Detection of Cell Surface Protein by IFA
2.7. Specimen Information
2.8. PEDV-Infected Vero Cells
2.9. Immunocytochemical (ICC) Staining
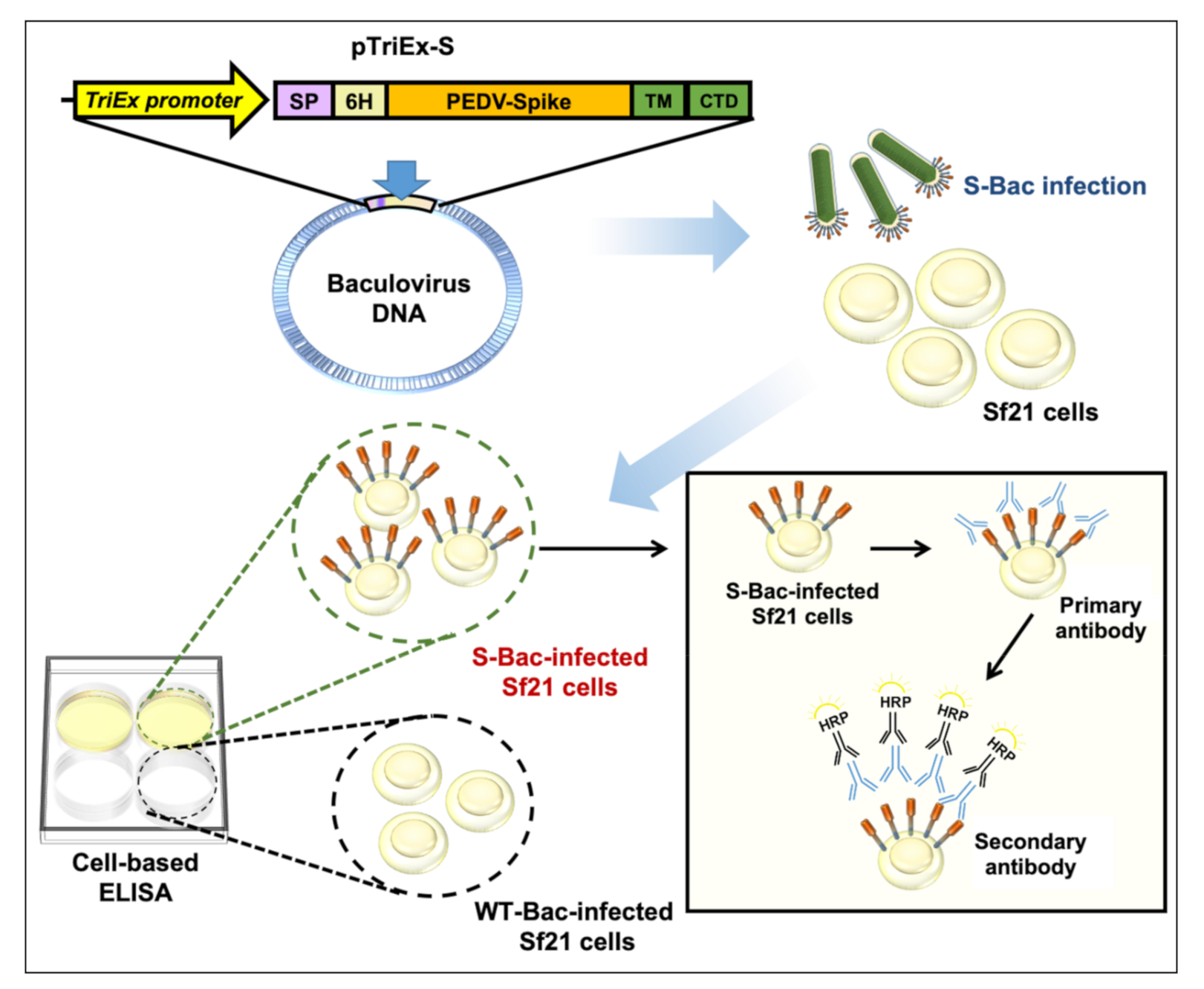
2.10. Establishment of Cell-Based ELISA by S-Bac Infection of Sf21 Cells
2.11. Determination of Cutoff Value, Specificity, and Sensitivity of S-Bac Cell-Based ELISA
2.12. Statistical Analysis
3. Results
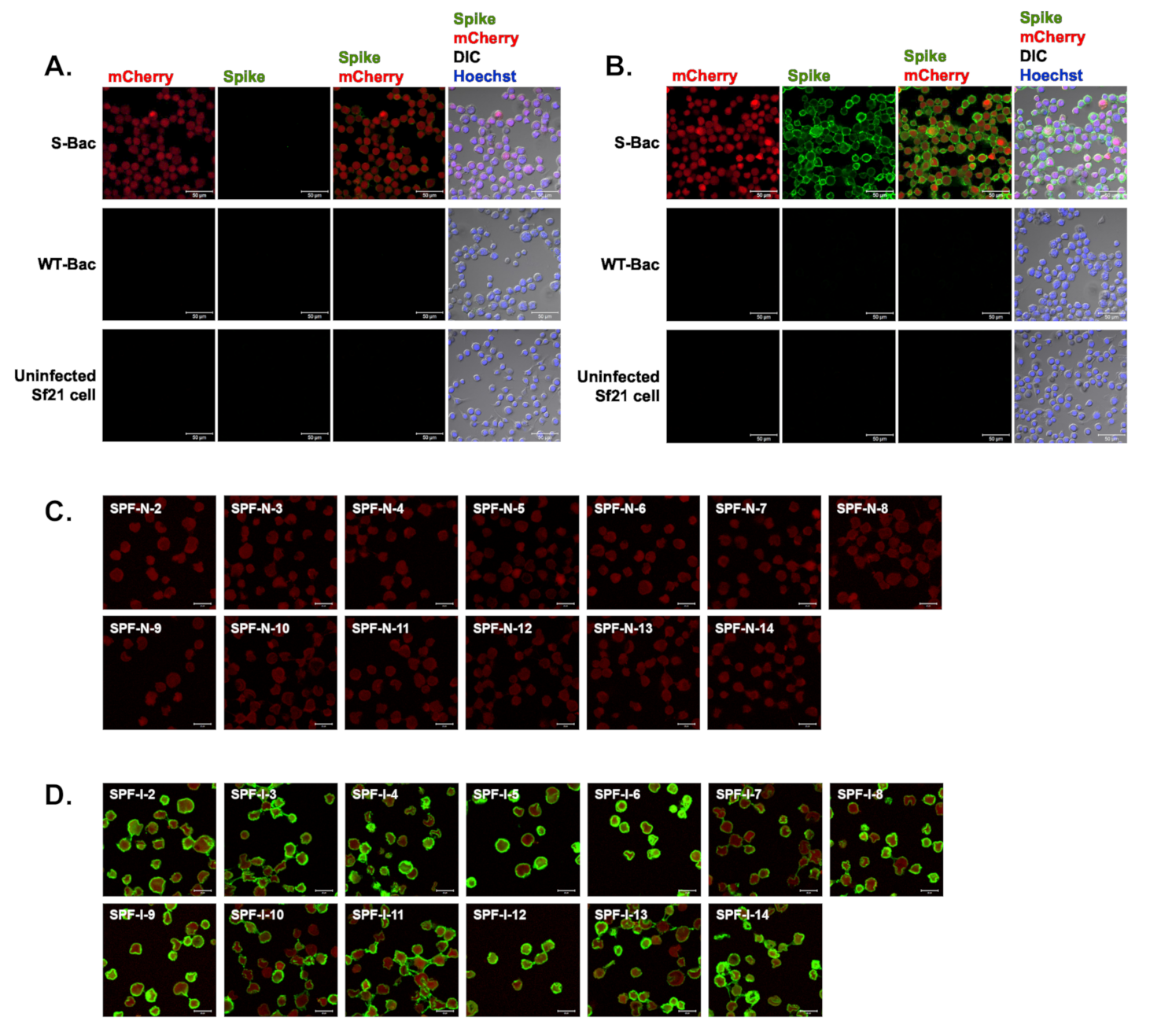
3.1. Construction of S-Bac and Detection of S Protein Expression in Infected Insect Cells
3.2. Spike Protein Displays on Insect Cell Membranes and Is Properly Detected by IFA
3.3. ICC Staining of Antisera against G2b PEDV-PT-Infected Vero Cells
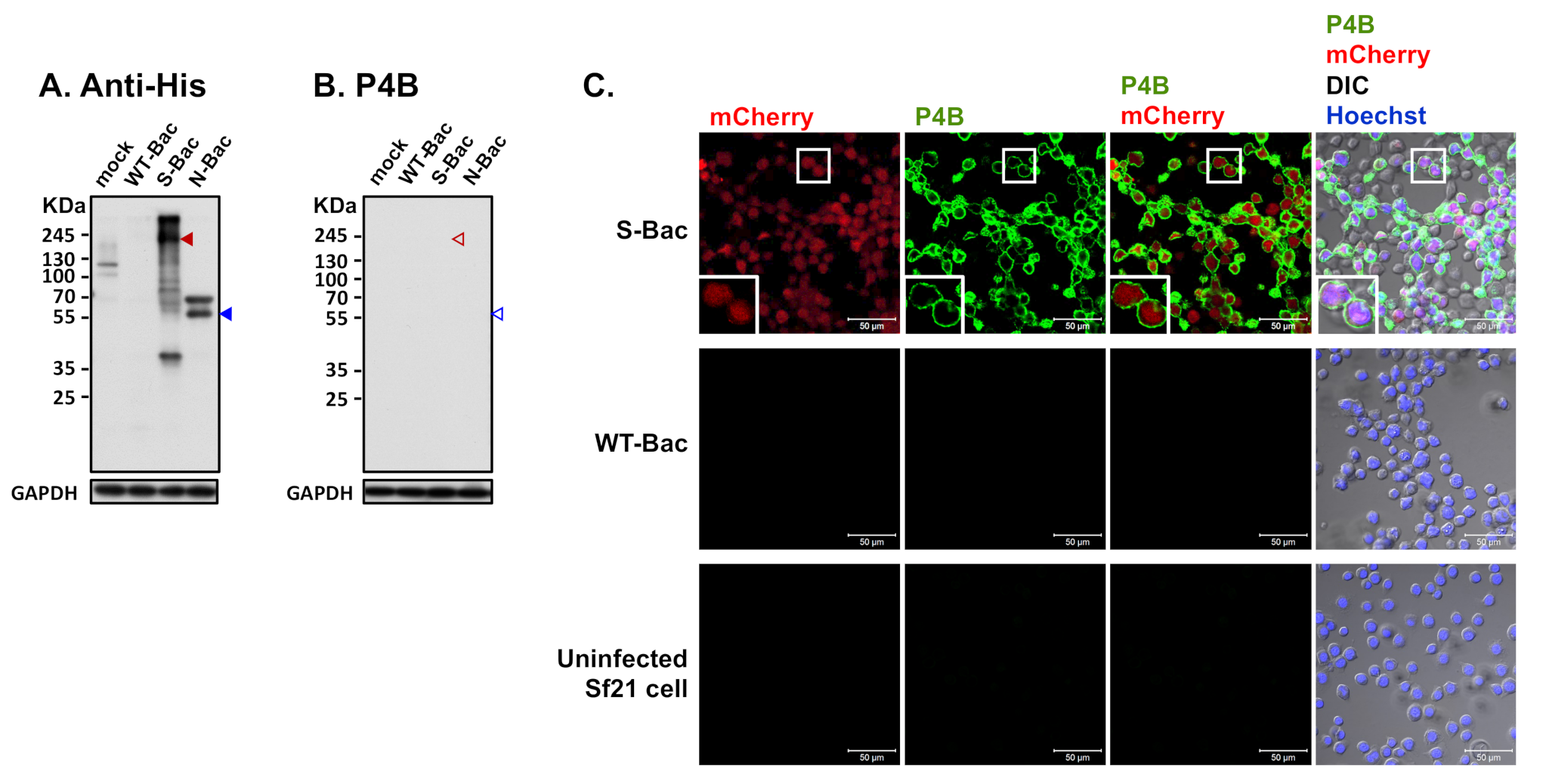
3.4. Western Blot Analysis of Pig Sera Fails to Detect Anti-PEDV S Antisera
3.5. IFA on S-Bac-Infected Cells Identifies Anti-PEDV S Antisera
3.6. A Conformation-Specific Anti-S Antibody May Not Recognize S in a Denatured Western Blot Analysis
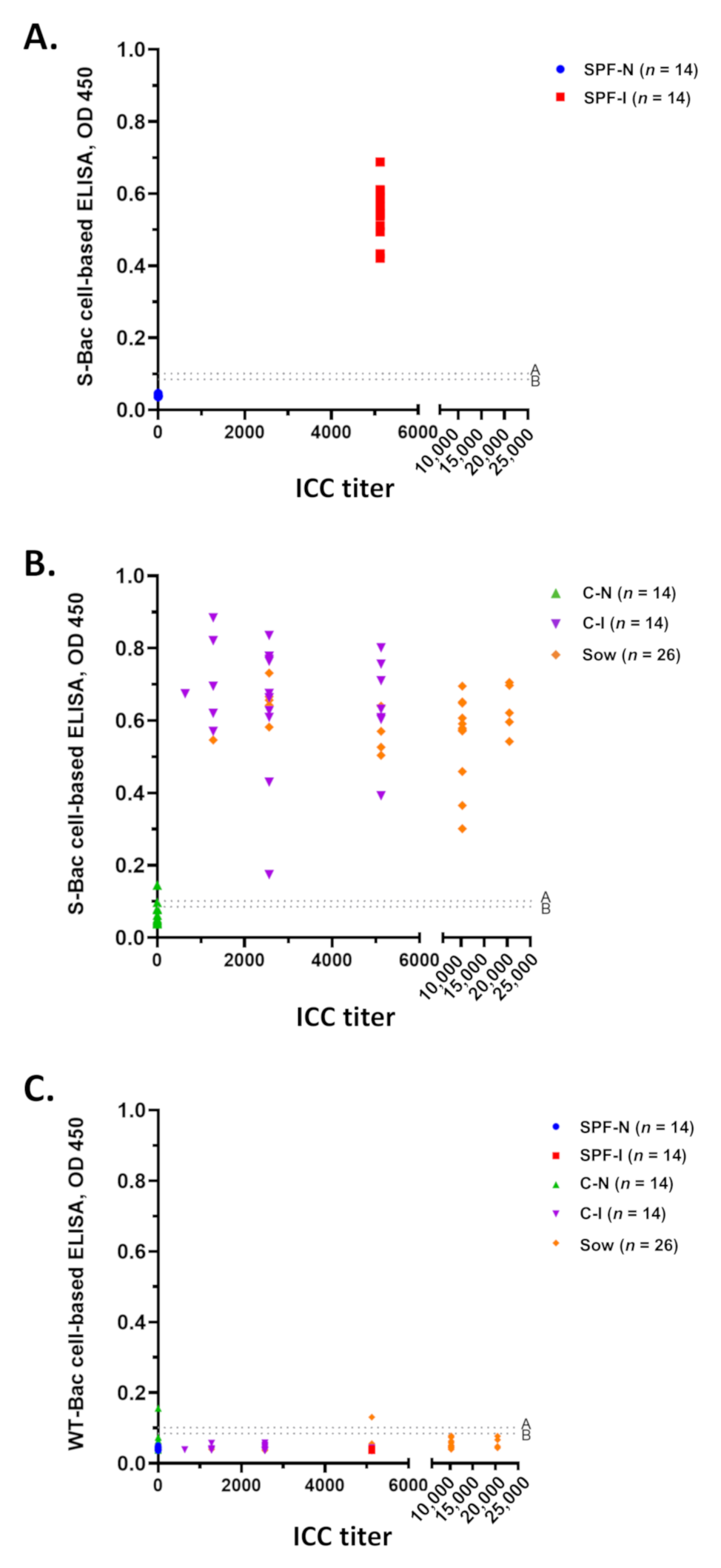
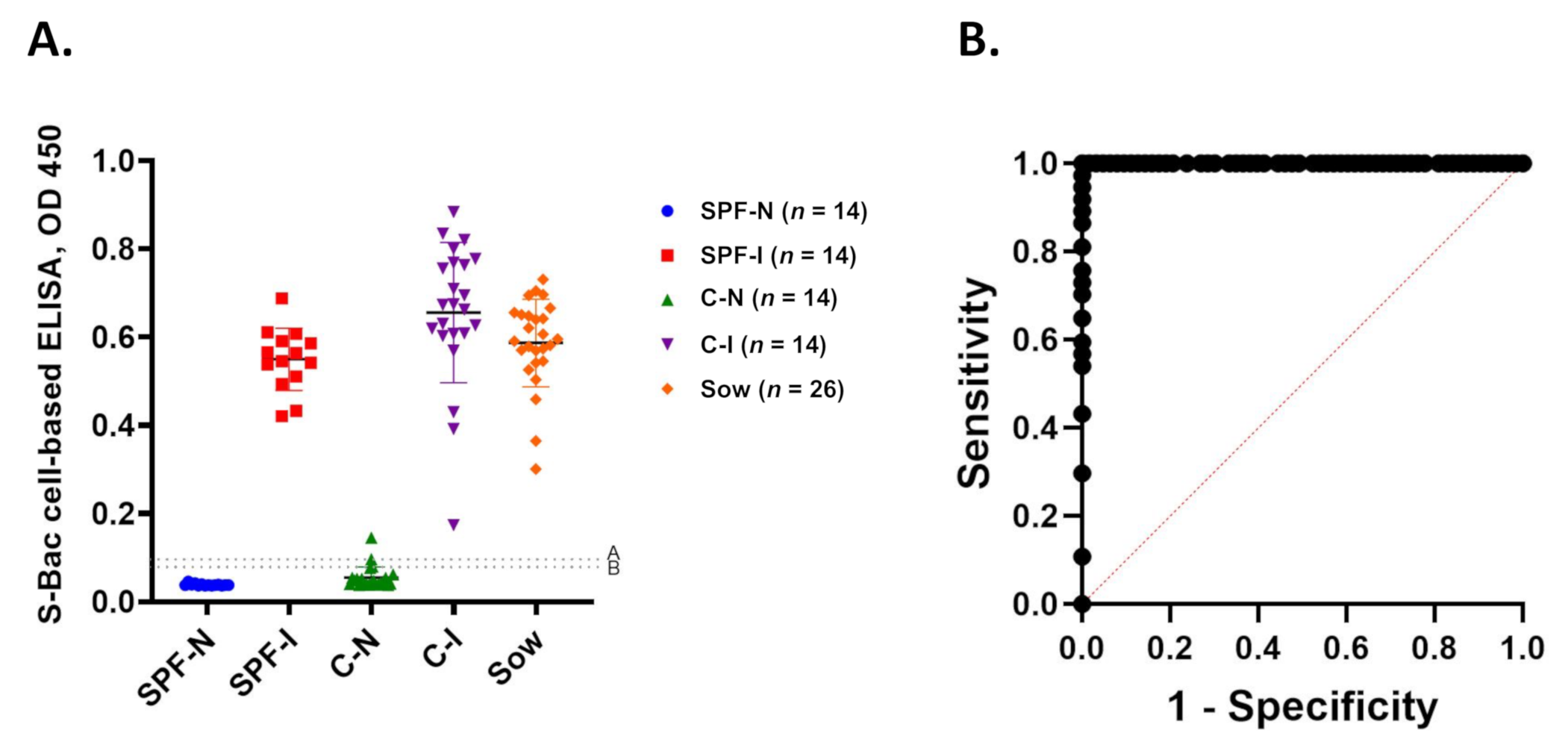
3.7. Establishment of a Novel Cell-Based ELISA Using S-Bac-Infected Insect Cells
4. Discussion
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Jung, K.; Saif, L.J. Porcine epidemic diarrhea virus infection: Etiology, epidemiology, pathogenesis and immunoprophylaxis. Vet. J. 2015, 204, 134–143. [Google Scholar] [CrossRef] [PubMed]
- Lee, C. Porcine epidemic diarrhea virus: An emerging and re-emerging epizootic swine virus. Virol. J. 2015, 12, 193. [Google Scholar] [CrossRef] [PubMed]
- Song, D.; Park, B. Porcine epidemic diarrhoea virus: A comprehensive review of molecular epidemiology, diagnosis, and vaccines. Virus Genes 2012, 44, 167–175. [Google Scholar] [CrossRef]
- Pensaert, M.B.; de Bouck, P. A new coronavirus-like particle associated with diarrhea in swine. Arch. Virol. 1978, 58, 243–247. [Google Scholar] [CrossRef] [PubMed]
- Chen, Q.; Li, G.; Stasko, J.; Thomas, J.T.; Stensland, W.R.; Pillatzki, A.E.; Gauger, P.C.; Schwartz, K.J.; Madson, D.; Yoon, K.J.; et al. Isolation and characterization of porcine epidemic diarrhea viruses associated with the 2013 disease outbreak among swine in the United States. J. Clin. Microbiol. 2014, 52, 234–243. [Google Scholar] [CrossRef]
- Sun, D.; Wang, X.; Wei, S.; Chen, J.; Feng, L. Epidemiology and vaccine of porcine epidemic diarrhea virus in China: A mini-review. J. Vet. Med. Sci. 2016, 78, 355–363. [Google Scholar] [CrossRef]
- Diel, D.G.; Lawson, S.; Okda, F.; Singrey, A.; Clement, T.; Fernandes, M.H.V.; Christopher-Hennings, J.; Nelson, E.A. Porcine epidemic diarrhea virus: An overview of current virological and serological diagnostic methods. Virus Res. 2016, 226, 60–70. [Google Scholar] [CrossRef] [PubMed]
- Cho, Y.Y.; Lim, S.I.; Kim, Y.K.; Song, J.Y.; Lee, J.B.; An, D.J. Complete Genome Sequence of K14JB01, a Novel Variant Strain of Porcine Epidemic Diarrhea Virus in South Korea. Genome. Announc. 2014, 2. [Google Scholar] [CrossRef] [PubMed]
- Lin, C.N.; Chung, W.B.; Chang, S.W.; Wen, C.C.; Liu, H.; Chien, C.H.; Chiou, M.T. US-like strain of porcine epidemic diarrhea virus outbreaks in Taiwan, 2013-2014. J. Vet. Med Sci. 2014, 76, 1297–1299. [Google Scholar] [CrossRef]
- Chang, Y.C.; Kao, C.F.; Chang, C.Y.; Jeng, C.R.; Tsai, P.S.; Pang, V.F.; Chiou, H.Y.; Peng, J.Y.; Cheng, I.C.; Chang, H.W. Evaluation and Comparison of the Pathogenicity and Host Immune Responses Induced by a G2b Taiwan Porcine Epidemic Diarrhea Virus (Strain Pintung 52) and Its Highly Cell-Culture Passaged Strain in Conventional 5-Week-Old Pigs. Viruses 2017, 9, 121. [Google Scholar] [CrossRef] [PubMed]
- Brian, D.A.; Baric, R.S. Coronavirus genome structure and replication. Curr. Top Microbiol. Immunol. 2005, 287, 1–30. [Google Scholar] [CrossRef]
- Bao, F.; Wang, L.; Zhao, X.; Lu, T.; Na, A.M.; Wang, X.; Cao, J.; Du, Y. Preparation and characterization of a single-domain antibody specific for the porcine epidemic diarrhea virus spike protein. AMB Express 2019, 9, 104. [Google Scholar] [CrossRef]
- Kocherhans, R.; Bridgen, A.; Ackermann, M.; Tobler, K. Completion of the porcine epidemic diarrhoea coronavirus (PEDV) genome sequence. Virus Genes 2001, 23, 137–144. [Google Scholar] [CrossRef] [PubMed]
- Rasmussen, T.B.; Boniotti, M.B.; Papetti, A.; Grasland, B.; Frossard, J.P.; Dastjerdi, A.; Hulst, M.; Hanke, D.; Pohlmann, A.; Blome, S.; et al. Full-length genome sequences of porcine epidemic diarrhoea virus strain CV777; Use of NGS to analyse genomic and sub-genomic RNAs. PLoS ONE 2018, 13, e0193682. [Google Scholar] [CrossRef] [PubMed]
- Lee, D.K.; Park, C.K.; Kim, S.H.; Lee, C. Heterogeneity in spike protein genes of porcine epidemic diarrhea viruses isolated in Korea. Virus Res. 2010, 149, 175–182. [Google Scholar] [CrossRef]
- Gerber, P.F.; Gong, Q.; Huang, Y.W.; Wang, C.; Holtkamp, D.; Opriessnig, T. Detection of antibodies against porcine epidemic diarrhea virus in serum and colostrum by indirect ELISA. Vet. J. 2014, 202, 33–36. [Google Scholar] [CrossRef]
- Oh, J.; Lee, K.W.; Choi, H.W.; Lee, C. Immunogenicity and protective efficacy of recombinant S1 domain of the porcine epidemic diarrhea virus spike protein. Arch. Virol. 2014, 159, 2977–2987. [Google Scholar] [CrossRef]
- Li, W.; van Kuppeveld, F.J.M.; He, Q.; Rottier, P.J.M.; Bosch, B.J. Cellular entry of the porcine epidemic diarrhea virus. Virus Res. 2016, 226, 117–127. [Google Scholar] [CrossRef]
- Deng, F.; Ye, G.; Liu, Q.; Navid, M.T.; Zhong, X.; Li, Y.; Wan, C.; Xiao, S.; He, Q.; Fu, Z.F.; et al. Identification and Comparison of Receptor Binding Characteristics of the Spike Protein of Two Porcine Epidemic Diarrhea Virus Strains. Viruses 2016, 8, 55. [Google Scholar] [CrossRef]
- Yuan, P.; Yang, Z.; Song, H.; Wang, K.; Yang, Y.; Xie, L.; Huang, S.; Liu, J.; Ran, L.; Song, Z. Three Main Inducers of Alphacoronavirus Infection of Enterocytes: Sialic Acid, Proteases, and Low pH. Intervirology 2018, 61, 53–63. [Google Scholar] [CrossRef] [PubMed]
- Li, F. Structure, Function, and Evolution of Coronavirus Spike Proteins. Annu. Rev. Virol. 2016, 3, 237–261. [Google Scholar] [CrossRef] [PubMed]
- Belouzard, S.; Millet, J.K.; Licitra, B.N.; Whittaker, G.R. Mechanisms of coronavirus cell entry mediated by the viral spike protein. Viruses 2012, 4, 1011–1033. [Google Scholar] [CrossRef]
- Chang, C.Y.; Peng, J.Y.; Cheng, Y.H.; Chang, Y.C.; Wu, Y.T.; Tsai, P.S.; Chiou, H.Y.; Jeng, C.R.; Chang, H.W. Development and comparison of enzyme-linked immunosorbent assays based on recombinant trimeric full-length and truncated spike proteins for detecting antibodies against porcine epidemic diarrhea virus. BMC Vet. Res. 2019, 15, 421. [Google Scholar] [CrossRef] [PubMed]
- Chang, C.Y.; Cheng, I.C.; Chang, Y.C.; Tsai, P.S.; Lai, S.Y.; Huang, Y.L.; Jeng, C.R.; Pang, V.F.; Chang, H.W. Identification of Neutralizing Monoclonal Antibodies Targeting Novel Conformational Epitopes of the Porcine Epidemic Diarrhoea Virus Spike Protein. Sci. Rep. 2019, 9, 2529. [Google Scholar] [CrossRef]
- Liu, J.; Shi, H.; Chen, J.; Zhang, X.; Ji, Z.; Yuan, J.; Shi, D.; Cao, L.; Zhu, X.; Dong, H.; et al. Neutralization of genotype 2 porcine epidemic diarrhea virus strains by a novel monoclonal antibody. Virology 2017, 507, 257–262. [Google Scholar] [CrossRef]
- Gong, L.; Lin, Y.; Qin, J.; Li, Q.; Xue, C.; Cao, Y. Neutralizing antibodies against porcine epidemic diarrhea virus block virus attachment and internalization. Virol. J. 2018, 15, 133. [Google Scholar] [CrossRef]
- Chang, C.Y.; Hsu, W.T.; Chao, Y.C.; Chang, H.W. Display of Porcine Epidemic Diarrhea Virus Spike Protein on Baculovirus to Improve Immunogenicity and Protective Efficacy. Viruses 2018, 10, 346. [Google Scholar] [CrossRef] [PubMed]
- van Oers, M.M.; Pijlman, G.P.; Vlak, J.M. Thirty years of baculovirus-insect cell protein expression: From dark horse to mainstream technology. J. Gen. Virol. 2015, 96, 6–23. [Google Scholar] [CrossRef]
- Tsai, C.H.; Wei, S.C.; Lo, H.R.; Chao, Y.C. Baculovirus as Versatile Vectors for Protein Display and Biotechnological Applications. Curr. Issues Mol. Biol. 2020, 34, 231–256. [Google Scholar] [CrossRef]
- Agathos, S.N. Production scale insect cell culture. Biotechnol. Adv. 1991, 9, 51–68. [Google Scholar] [CrossRef]
- Harrap, K.A. The structure of nuclear polyhedrosis viruses. I. The inclusion body. Virology 1972, 50, 114–123. [Google Scholar]
- Blissard, G.W. Baculovirus--insect cell interactions. Cytotechnology 1996, 20, 73–93. [Google Scholar] [CrossRef]
- Tang, X.C.; Lu, H.R.; Ross, T.M. Hemagglutinin displayed baculovirus protects against highly pathogenic influenza. Vaccine 2010, 28, 6821–6831. [Google Scholar] [CrossRef]
- Wei, S.C.; Tsai, C.H.; Hsu, W.T.; Chao, Y.C. Baculovirus IE2 Interacts with Viral DNA through Daxx To Generate an Organized Nuclear Body Structure for Gene Activation in Vero Cells. J. Virol. 2019, 93. [Google Scholar] [CrossRef]
- Liu, C.Y.; Wang, C.H.; Hsiao, W.K.; Lo, H.R.; Wu, C.P.; Chao, Y.C. RING and coiled-coil domains of baculovirus IE2 are critical in strong activation of the cytomegalovirus major immediate-early promoter in mammalian cells. .J. Virol. 2009, 83, 3604–3616. [Google Scholar] [CrossRef] [PubMed]
- Chiou, H.Y.; Huang, Y.L.; Deng, M.C.; Chang, C.Y.; Jeng, C.R.; Tsai, P.S.; Yang, C.; Pang, V.F.; Chang, H.W. Phylogenetic Analysis of the Spike (S) Gene of the New Variants of Porcine Epidemic Diarrhoea Virus in Taiwan. Transbound Emerg. Dis. 2017, 64, 157–166. [Google Scholar] [CrossRef] [PubMed]
- Naik, N.G.; Lo, Y.W.; Wu, T.Y.; Lin, C.C.; Kuo, S.C.; Chao, Y.C. Baculovirus as an efficient vector for gene delivery into mosquitoes. Sci. Rep. 2018, 8, 17778. [Google Scholar] [CrossRef] [PubMed]
- Tsai, C.H.; Wei, S.C.; Jan, J.T.; Liao, L.L.; Chang, C.J.; Chao, Y.C. Generation of Stable Influenza Virus Hemagglutinin through Structure-Guided Recombination. ACS Synth. Biol. 2019, 8, 2472–2482. [Google Scholar] [CrossRef] [PubMed]
- Sun, R.Q.; Cai, R.J.; Chen, Y.Q.; Liang, P.S.; Chen, D.K.; Song, C.X. Outbreak of porcine epidemic diarrhea in suckling piglets, China. Emerg. Infect. Dis. 2012, 18, 161–163. [Google Scholar] [CrossRef] [PubMed]
- Van Diep, N.; Sueyoshi, M.; Norimine, J.; Hirai, T.; Myint, O.; Teh, A.P.P.; Izzati, U.Z.; Fuke, N.; Yamaguchi, R. Molecular characterization of US-like and Asian non-S INDEL strains of porcine epidemic diarrhea virus (PEDV) that circulated in Japan during 2013-2016 and PEDVs collected from recurrent outbreaks. BMC Vet. Res. 2018, 14, 96. [Google Scholar] [CrossRef]
- Guo, J.; Fang, L.; Ye, X.; Chen, J.; Xu, S.; Zhu, X.; Miao, Y.; Wang, D.; Xiao, S. Evolutionary and genotypic analyses of global porcine epidemic diarrhea virus strains. Transbound Emerg. Dis. 2019, 66, 111–118. [Google Scholar] [CrossRef]
- Wen, Z.; Li, J.; Zhang, Y.; Zhou, Q.; Gong, L.; Xue, C.; Cao, Y. Genetic epidemiology of porcine epidemic diarrhoea virus circulating in China in 2012-2017 based on spike gene. Transbound Emerg. Dis. 2018, 65, 883–889. [Google Scholar] [CrossRef]
- Cassar, O.; Gessain, A. Serological and Molecular Methods to Study Epidemiological Aspects of Human T-Cell Lymphotropic Virus Type 1 Infection. Methods Mol. Biol. 2017, 1582, 3–24. [Google Scholar] [CrossRef] [PubMed]
- Okda, F.; Liu, X.; Singrey, A.; Clement, T.; Nelson, J.; Christopher-Hennings, J.; Nelson, E.A.; Lawson, S. Development of an indirect ELISA, blocking ELISA, fluorescent microsphere immunoassay and fluorescent focus neutralization assay for serologic evaluation of exposure to North American strains of Porcine Epidemic Diarrhea Virus. BMC Vet. Res. 2015, 11, 180. [Google Scholar] [CrossRef]
- Ren, X.; Suo, S.; Jang, Y.S. Development of a porcine epidemic diarrhea virus M protein-based ELISA for virus detection. Biotechnol. Lett. 2011, 33, 215–220. [Google Scholar] [CrossRef] [PubMed]
- Hou, X.L.; Yu, L.Y.; Liu, J. Development and evaluation of enzyme-linked immunosorbent assay based on recombinant nucleocapsid protein for detection of porcine epidemic diarrhea (PEDV) antibodies. Vet. Microbiol. 2007, 123, 86–92. [Google Scholar] [CrossRef] [PubMed]
- Shan, Y.; Liu, Y.; Liu, Z.; Li, G.; Chen, C.; Luo, H.; Chen, Y.; Guo, N.; Shi, X.; Zhang, X.; et al. Development and application of an indirect enzyme-linked immunosorbent assay using recombinant S1 for serological testing of porcine epidemic diarrhea virus. Can. J. Microbiol. 2019, 65, 343–352. [Google Scholar] [CrossRef] [PubMed]
- Lin, H.; Zhou, H.; Gao, L.; Li, B.; He, K.; Fan, H. Development and application of an indirect ELISA for the detection of antibodies to porcine epidemic diarrhea virus based on a recombinant spike protein. BMC Vet. Res. 2018, 14, 243. [Google Scholar] [CrossRef]


| Groups | No. of Pigs | ICC Staining | S-Bac Infected Cell-Based ELISA | ||||
|---|---|---|---|---|---|---|---|
| (Cutoff Mean+2SD) | (Cutoff Mean+3SD) | ||||||
| + | − | + | − | + | − | ||
| SPF-N | 14 | 0 | 14 | 0 | 14 | 0 | 14 |
| SPF-I | 14 | 14 | 0 | 14 | 0 | 14 | 0 |
| C-N | 23 | 0 | 23 | 2 | 21 | 1 | 22 |
| C-I | 23 | 23 | 0 | 23 | 0 | 23 | 0 |
| Sow | 26 | 26 | 0 | 26 | 0 | 26 | 0 |
| Sensitivity | 100% | 100% | 100% | ||||
| Specificity | 100% | 95% | 97% | ||||
| 𝜅 | 1 | 0.96 | 0.98 | ||||
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Hsu, W.-T.; Chang, C.-Y.; Tsai, C.-H.; Wei, S.-C.; Lo, H.-R.; Lamis, R.J.S.; Chang, H.-W.; Chao, Y.-C. PEDV Infection Generates Conformation-Specific Antibodies That Can Be Effectively Detected by a Cell-Based ELISA. Viruses 2021, 13, 303. https://doi.org/10.3390/v13020303
Hsu W-T, Chang C-Y, Tsai C-H, Wei S-C, Lo H-R, Lamis RJS, Chang H-W, Chao Y-C. PEDV Infection Generates Conformation-Specific Antibodies That Can Be Effectively Detected by a Cell-Based ELISA. Viruses. 2021; 13(2):303. https://doi.org/10.3390/v13020303
Chicago/Turabian StyleHsu, Wei-Ting, Chia-Yu Chang, Chih-Hsuan Tsai, Sung-Chan Wei, Huei-Ru Lo, Robert John S. Lamis, Hui-Wen Chang, and Yu-Chan Chao. 2021. "PEDV Infection Generates Conformation-Specific Antibodies That Can Be Effectively Detected by a Cell-Based ELISA" Viruses 13, no. 2: 303. https://doi.org/10.3390/v13020303
APA StyleHsu, W.-T., Chang, C.-Y., Tsai, C.-H., Wei, S.-C., Lo, H.-R., Lamis, R. J. S., Chang, H.-W., & Chao, Y.-C. (2021). PEDV Infection Generates Conformation-Specific Antibodies That Can Be Effectively Detected by a Cell-Based ELISA. Viruses, 13(2), 303. https://doi.org/10.3390/v13020303






