Characterization of Single-Chain Fv Fragments of Neutralizing Antibodies to Rabies Virus Glycoprotein
Abstract
:1. Introduction
2. Materials and Methods
2.1. Construction of Expression Plasmid for 15–13 and 12–22 scFvs
2.2. Expression and Purification of 15–13 and 12–22 scFvs
2.3. Preparation and Purification of 15–13 and 12–22 Fab Fragments
2.4. Preparation of Recombinant RABVG by Silkworm–Baculovirus Expression System
2.5. Thermal Stability Analysis by Differential Scanning Fluorometry (DSF)
2.6. Interaction Analysis Using Biolayer Interferometry (BLI)
3. Results
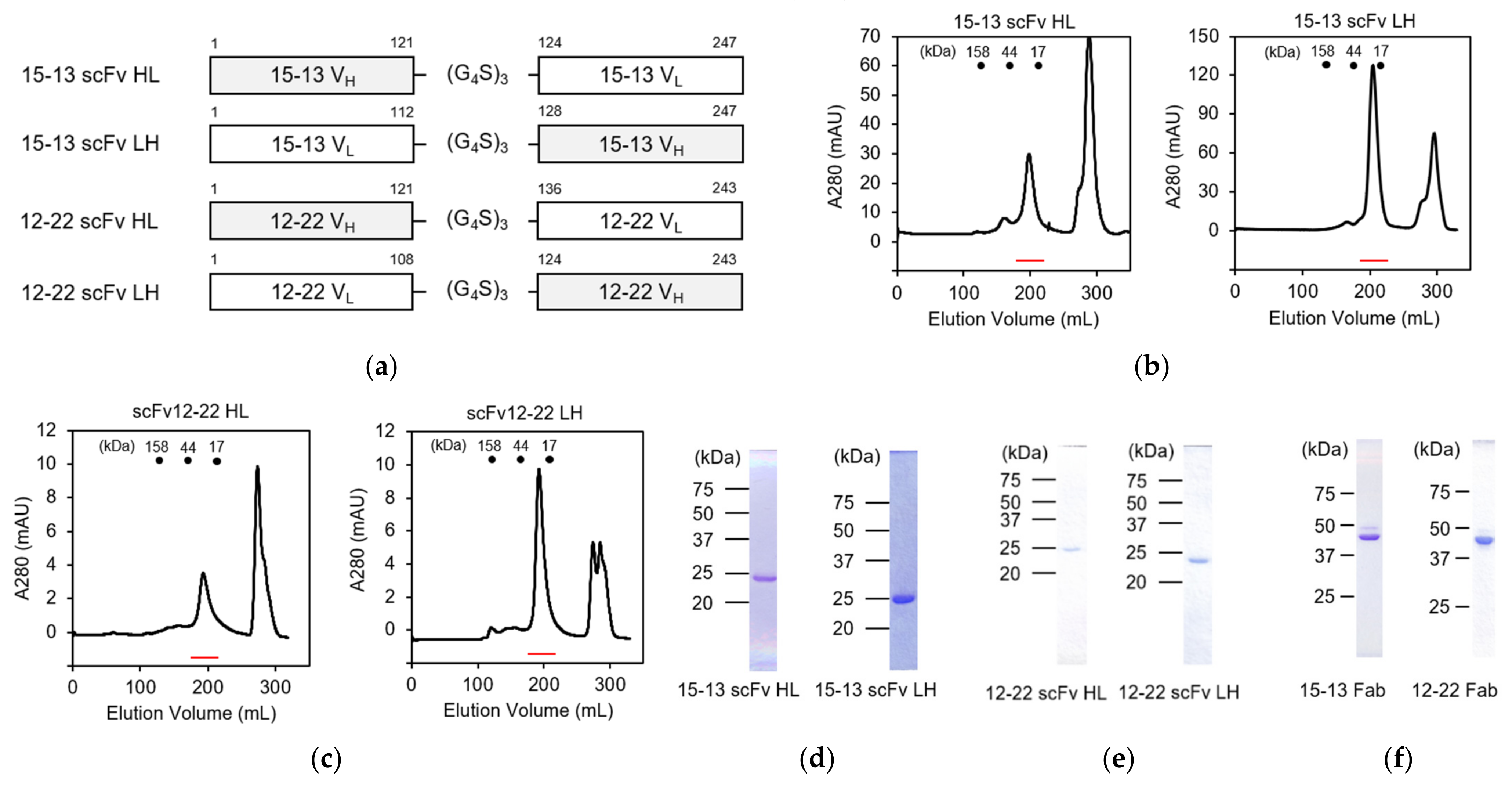
3.1. Cloning of Variable Regions of 15–13 and 12–22 Antibodies and Preparation of Their Single-Chain Fv Fragments
3.2. Preparation of Fab Fragments of 15–13 and 12–22 Antibodies
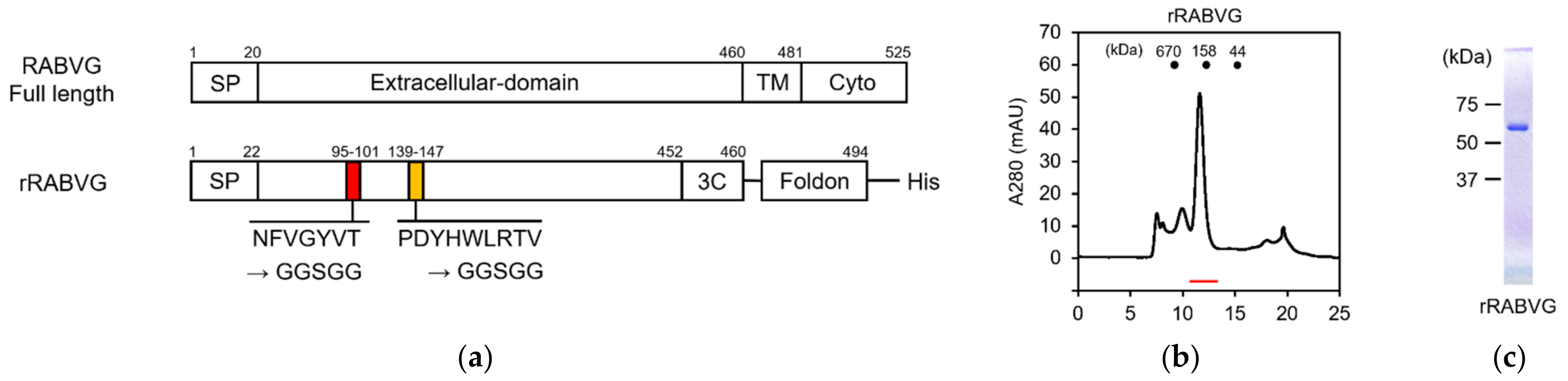
3.3. Construction and Purification of rRABVG
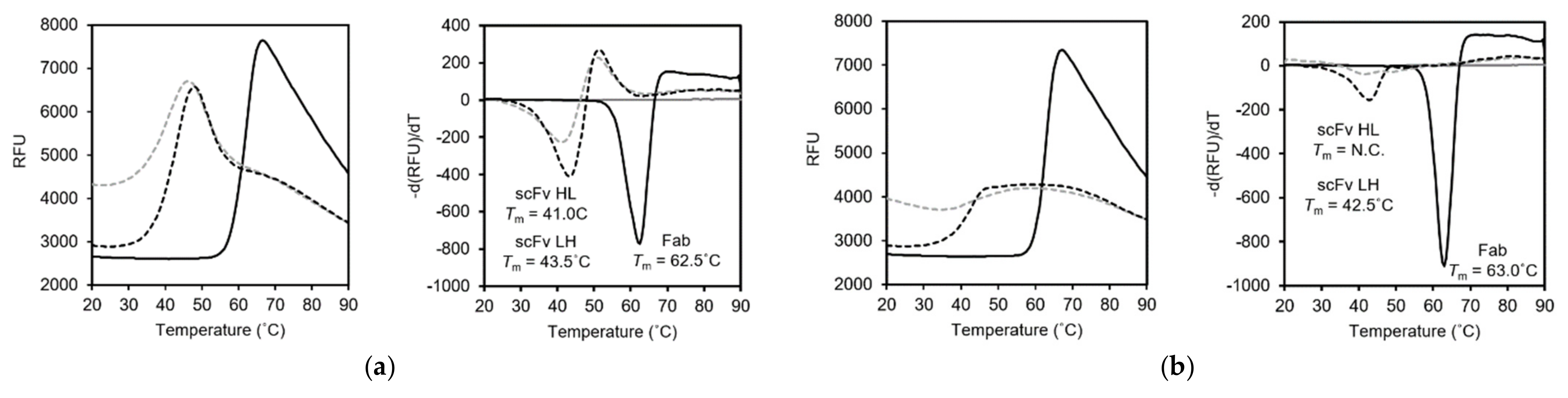
3.4. Differential Scanning Fluorimetry (DSF)
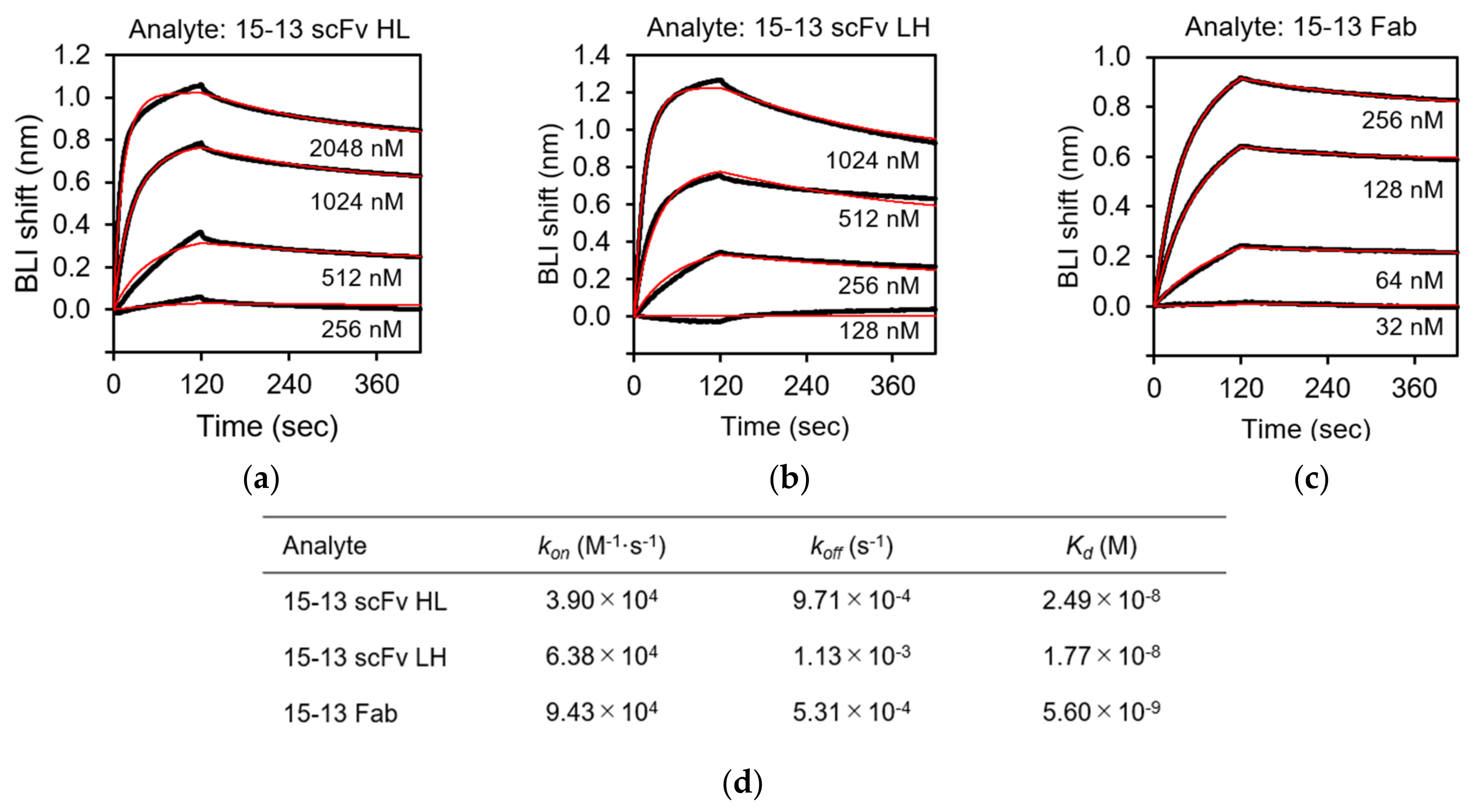
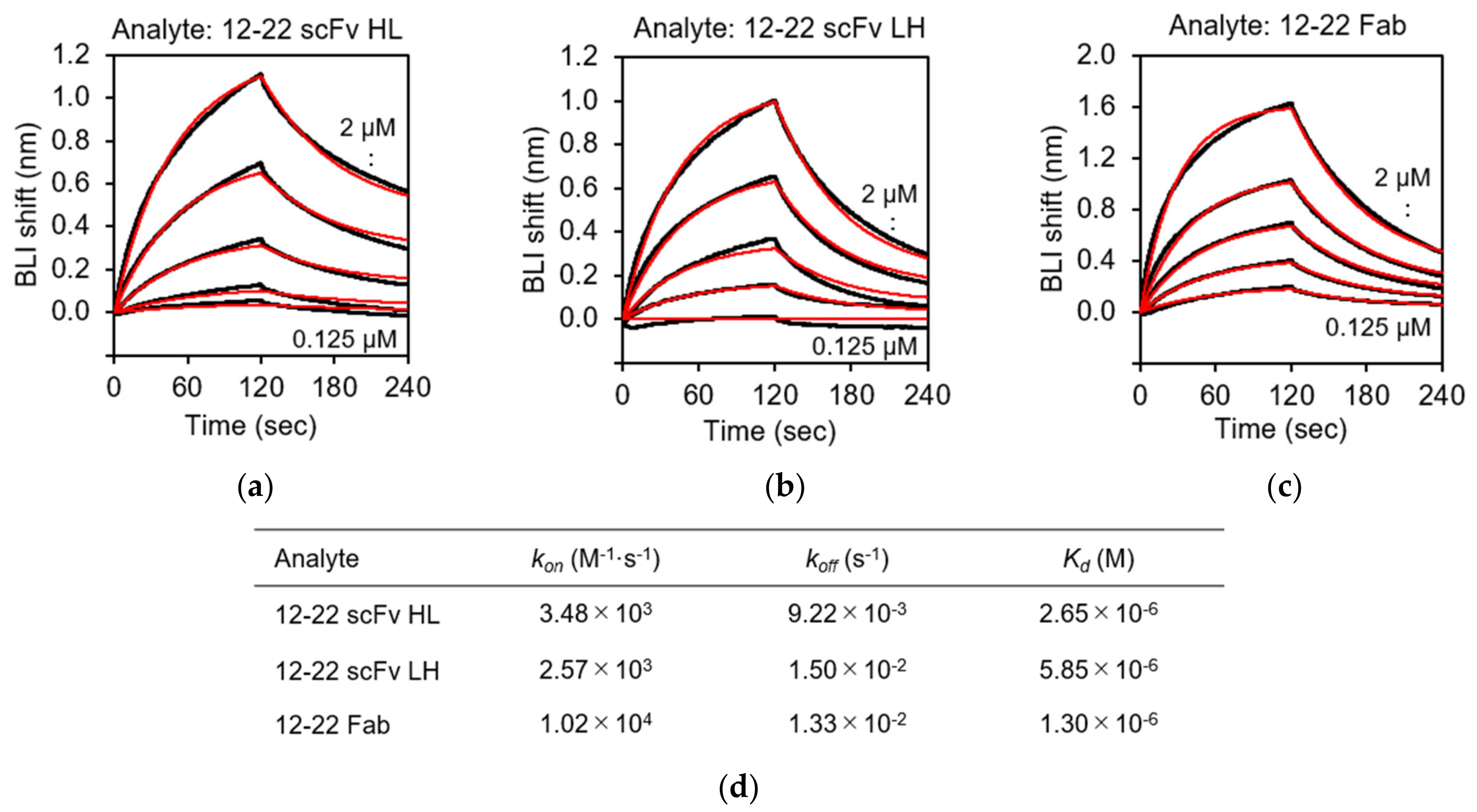
3.5. Biolayer Interferometry (BLI) Analysis of the rRABVG-scFv Interaction
4. Discussion
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Cox, J.H.; Dietzschold, B.; Schneider, L.G. Rabies virus glycoprotein. II. Biological and serological characterization. Infect. Immun. 1997, 16, 754. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Hampson, K.; Coudeville, L.; Lembo, T.; Sambo, M.; Kieffer, A.; Attlan, M.; Barrat, J.; Blanton, J.D.; Briggs, D.J.; Cleaveland, S.; et al. Estimating the Global Burden of Endemic Canine Rabies. PLoS Negl. Trop. Dis. 2015, 9, e0003709. [Google Scholar]
- Rabies. Available online: https://www.who.int/health-topics/rabies#tab=tab_1 (accessed on 8 October 2021).
- Wilde, H.; Tipkong, P.; Khawplod, P. Economic issues in postexposure rabies treatment. J. Travel Med. 1999, 6, 238–242. [Google Scholar] [CrossRef] [Green Version]
- World Health Organization. Rabies Vaccines: WHO Position Paper—April 2018. Wkly. Epidemiol. Rec. Relev. Épidémiologique Hebd. 2018, 93, 201–219. [Google Scholar]
- Tuffereau, C.; Bénéjean, J.; Blondel, D.; Kieffer, B.; Flamand, A. Low-affinity nerve-growth factor receptor (P75NTR) can serve as a receptor for rabies virus. EMBO J. 1998, 17, 7250–7259. [Google Scholar] [CrossRef]
- Thoulouze, M.-I.; Lafage, M.; Schachner, M.; Hartmann, U.; Cremer, H.; Lafon, M. The Neural Cell Adhesion Molecule Is a Receptor for Rabies Virus. J. Virol. 1998, 72, 7181–7190. [Google Scholar] [CrossRef] [Green Version]
- Lentz, T.L.; Burrage, T.G.; Smith, A.L.; Crick, J.; Tignor, G.H. Is the acetylcholine receptor a rabies virus receptor? Science 1982, 215, 182–184. [Google Scholar] [CrossRef]
- Lafon, M. Rabies virus receptors. J. NeuroVirology 2005, 11, 82–87. [Google Scholar] [CrossRef]
- Gaudin, Y.; Ruigrok RW, H.; Knossow, M.; Flamand1, A. Low-pH Conformational Changes of Rabies Virus Glycoprotein and Their Role in Membrane Fusion. J. Virol. 1993, 67, 1365–1372. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Yang, F.; Lin, S.; Ye, F.; Yang, J.; Qi, J.; Chen, Z.; Lin, X.; Wang, J.; Yue, D.; Cheng, Y.; et al. Structural Analysis of Rabies Virus Glycoprotein Reveals pH-Dependent Conformational Changes and Interactions with a Neutralizing Antibody. Cell Host Microbe 2020, 27, 441–453.e7. [Google Scholar] [CrossRef]
- Luo, T.R.; Minamoto, N.; Hishida, M.; Yamamoto, K.; Fujise, T.; Hiraga, S.; Ito, N.; Sugiyama, M.; Kinjo, T. Antigenic and Functional Analyses of Glycoprotein of Rabies Virus Using Monoclonal Antibodies. Microbiol. Immunol. 1998, 42, 187–193. [Google Scholar] [CrossRef] [PubMed]
- Imai, S.; Mukai, Y.; Nagano, K.; Shibata, H.; Sugita, T.; Abe, Y.; Nomura, T.; Tsutsumi, Y.; Kamada, H.; Nakagawa, S.; et al. Quality Enhancement of the Non-immune Phage scFv Library to Isolate Effective Antibodies. Biol. Pharm. Bull. 2006, 29, 1325–1330. [Google Scholar] [CrossRef] [Green Version]
- Swindells, M.B.; Porter, C.T.; Couch, M.; Hurst, J.; Abhinandan, K.R.; Nielsen, J.H.; Macindoe, G.; Hetherington, J.; Martin, A.C. abYsis: Integrated Antibody Sequence and Structure—Management, Analysis, and Prediction. J. Mol. Biol. 2017, 429, 356–364. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Al-Lazikani, B.; Lesk, A.M.; Chothia, C. Standard conformations for the canonical structures of immunoglobulins. J. Mol. Biol. 1997, 273, 927–948. [Google Scholar] [CrossRef] [PubMed]
- Shiroishi, M.; Tsumoto, K.; Amano, K.; Shirakihara, Y.; Colonna, M.; Braud, V.; Allan, D.S.J.; Makadzange, A.; Rowland-Jones, S.; Willcox, B.; et al. Human inhibitory receptors Ig-like transcript 2 (ILT2) and ILT4 compete with CD8 for MHC class I binding and bind preferentially to HLA-G. Proc. Natl. Acad. Sci. USA 2003, 100, 8856–8861. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Sissoëff, L.; Mousli, M.; England, P.; Tuffereau, C. Stable trimerization of recombinant rabies virus glycoprotein ectodomain is required for interaction with the p75NTRreceptor. J. Gen. Virol. 2005, 86, 2543–2552. [Google Scholar] [CrossRef] [PubMed]
- Marasco, W.A.; Sui, J. The growth and potential of human antiviral monoclonal antibody therapeutics. Nat. Biotechnol. 2007, 25, 1421–1434. [Google Scholar] [CrossRef] [PubMed]
- O’Brien, K.L.; Nolan, T. The WHO position on rabies immunization—2018 updates. Vaccine 2019, 37, A85. [Google Scholar] [CrossRef]
- Sparrow, E.; Torvaldsen, S.; Newall, A.; Wood, J.G.; Sheikh, M.; Kieny, M.P.; Abela-Ridder, B. Recent advances in the development of monoclonal antibodies for rabies post exposure prophylaxis: A review of the current status of the clinical development pipeline. Vaccine 2018, 37, A132–A139. [Google Scholar] [CrossRef]
- Chao, T.-Y.; Ren, S.; Shen, E.; Moore, S.; Zhang, S.-F.; Chen, L.; Rupprecht, C.E.; Tsao, E. SYN023, a novel humanized monoclonal antibody cocktail, for post-exposure prophylaxis of rabies. PLoS Negl. Trop. Dis. 2017, 11, e0006133. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Bakker, A.; Python, C.; Kissling, C.; Pandya, P.; Marissen, W.; Brink, M.; Lagerwerf, F.; Worst, S.; van Corven, E.; Kostense, S.; et al. First administration to humans of a monoclonal antibody cocktail against rabies virus: Safety, tolerability, and neutralizing activity. Vaccine 2008, 26, 5922–5927. [Google Scholar] [CrossRef] [Green Version]
- Bates, A.; Power, C.A. David vs. Goliath: The Structure, Function, and Clinical Prospects of Antibody Fragments. Antibodies 2019, 8, 28. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Nelson, A.L. Antibody fragments: Hope and hype. MAbs 2010, 2, 77. [Google Scholar] [CrossRef] [Green Version]
- Bühler, P.; Wetterauer, D.; Gierschner, D.; Wetterauer, U.; Beile, U.E.; Wolf, P. Influence of structural variations on biological activity of anti-PSMA scFv and immunotoxins targeting prostate cancer. Anticancer. Res. 2010, 30, 3373–3379. [Google Scholar]
- Kim, Y.-J.; Neelamegam, R.; Heo, M.-A.; Edwardraja, S.; Paik, H.-J.; Lee, S.-G. Improving the productivity of single-chain Fv antibody against c-Met by rearranging the order of its variable domains. J. Microbiol. Biotechnol. 2008, 18, 1186–1190. [Google Scholar]
- Xi, H.; Yuan, R.; Chen, X.; Gu, T.; Cheng, Y.; Li, Z.; Jiang, C.; Kong, W.; Wu, Y. Purification and on-column refolding of a single-chain antibody fragment against rabies virus glycoprotein expressed in Escherichia coli. Protein Expr. Purif. 2016, 126, 26–32. [Google Scholar] [CrossRef]
- Biggar, K.K.; Dawson, N.J.; Storey, K.B. Real-time protein unfolding: A method for determining the kinetics of native protein denaturation using a quantitative real-time thermocycler. BioTechniques 2012, 53, 231–238. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Bakker, A.B.H.; Marissen, W.E.; Kramer, R.A.; Rice, A.B.; Weldon, W.C.; Niezgoda, M.; Hanlon, C.A.; Thijsse, S.; Backus, H.H.J.; de Kruif, J.; et al. Novel Human Monoclonal Antibody Combination Effectively Neutralizing Natural Rabies Virus Variants and Individual In Vitro Escape Mutants. J. Virol. 2005, 79, 9062–9068. [Google Scholar] [CrossRef] [PubMed] [Green Version]

| Name | Sequence |
|---|---|
| For 15–13 | |
| 15–13 scFv HL_HF | 5′- GGGATTCCATATGGAGGTGAAGGTGGTGGAGTC -3′ |
| 15–13 scFv HL_HR | 5′- CTGCCGCCACCACCAGACCCACCGCCGCCTGAGGAGACGGTGACTGAG -3′ |
| 15–13 scFv HL_LF | 5′- GGTGGTGGCGGCAGTGGAGGTGGTGGATCCGATATTGTTTTGTCACAGTCTCCAGC -3′ |
| 15–13 scFv HL_LR | 5′- GGCGAATTCTCATTTGATTTCCAGCTTTGTCCCAG -3′ |
| 15–13 scFv LH_LF | 5′- GGGATTCCATATGGATATTGTTTTGTCACAGTCTC -3′ |
| 15–13 scFv LH_LR | 5′- CTGCCGCCACCACCAGACCCACCGCCGCCTTTGATTTCCAGCTTTGTC -3′ |
| 15–13 scFv LH_HF | 5′- GGTGGTGGCGGCAGTGGAGGTGGTGGATCCGAGGTGAAGGTGGTGGAGTC -3′ |
| 15–13 scFv LH_HR | 5′- GGCGAATTCTCATGAGGAGACGGTGACTGAG -3′ |
| For 12–22 | |
| 12–22 scFv HL_HF | 5′- GGGATTCCATATGGAAGTGCAACTGGTGGAGTCTG -3′ |
| 12–22 scFv HL_HR | 5′- CTGCCGCCACCACCAGACCCACCGCCGCCCGAGGAAACGGTGACCG -3′ |
| 12–22 scFv HL_LF | 5′- GGTGGTGGCGGCAGTGGAGGTGGTGGATCCGATATTGTGTTATCACAGTCTCCAGCC -3′ |
| 12–22 scFv HL_LR | 5′- GGCGAATTCTCATTTTATTTCCAGCTTGGTCCCAG -3′ |
| 12–22 scFv LH_LF | 5′- GGGATTCCATATGGATATTGTGTTATCACAGTCTC -3′ |
| 12–22 scFv LH_LR | 5′- CTGCCGCCACCACCAGACCCACCGCCGCCTTTTATTTCCAGCTTGG -3′ |
| 12–22 scFv LH_HF | 5′- GGTGGTGGCGGCAGTGGAGGTGGTGGATCCGAAGTGCAACTGGTGGAGTCTG -3′ |
| 12–22 scFv LH_HR | 5′- GGCGAATTCTCACGAGGAAACGGTGACCGTG -3′ |
| Name | Sequence |
|---|---|
| For RABVG | |
| RABVG_ECD_F | 5′- CTAGCTAGCTTCCCCATTTACACGATACC -3′ |
| RABVG_ECD_R | 5′- CCGCTCGAGTGATTTCGGGAGACC -3′ |
| pFastBac1_F | 5′- CCTATAAATATTCCGGATTA-3′ |
| RABVG_ECDm_FL1F | 5′- GGAGGATCTGGAGGAACCACATTCAAGAG -3′ |
| RABVG_ECDm_FL1R | 5′- TCCTCCAGATCCTCCGGTGTAGGTCTCTGC -3′ |
| RABVG_ECDm_FL2F | 5′- GGTGGTAGTGGTGGTAGAACCACCAAAGAG -3′ |
| RABVG_ECDm_FL2R | 5′- ACCACCACTACCACCGTATGGATTGTGTAG -3′ |
| pFastBac1_R | 5′- CAAATGTGGTATGGCTGATT-3′ |
| Name | Inclusion Body (mg/L) | Refolding Efficiency (%) | Final Yields (mg/L) |
|---|---|---|---|
| 15–13 scFv HL | 43.6 | 12.2 | 5.3 |
| 15–13 scFv LH | 85.8 | 14.5 | 12.4 |
| 12–22 scFv HL | 57.4 | 1.4 | 0.8 |
| 12–22 scFv LH | 73.6 | 3.0 | 2.2 |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Yumoto, K.; Arisaka, T.; Okada, K.; Aoki, K.; Ose, T.; Masatani, T.; Sugiyama, M.; Ito, N.; Fukuhara, H.; Maenaka, K. Characterization of Single-Chain Fv Fragments of Neutralizing Antibodies to Rabies Virus Glycoprotein. Viruses 2021, 13, 2311. https://doi.org/10.3390/v13112311
Yumoto K, Arisaka T, Okada K, Aoki K, Ose T, Masatani T, Sugiyama M, Ito N, Fukuhara H, Maenaka K. Characterization of Single-Chain Fv Fragments of Neutralizing Antibodies to Rabies Virus Glycoprotein. Viruses. 2021; 13(11):2311. https://doi.org/10.3390/v13112311
Chicago/Turabian StyleYumoto, Kohei, Tomoaki Arisaka, Kazuma Okada, Kyosuke Aoki, Toyoyuki Ose, Tatsunori Masatani, Makoto Sugiyama, Naoto Ito, Hideo Fukuhara, and Katsumi Maenaka. 2021. "Characterization of Single-Chain Fv Fragments of Neutralizing Antibodies to Rabies Virus Glycoprotein" Viruses 13, no. 11: 2311. https://doi.org/10.3390/v13112311
APA StyleYumoto, K., Arisaka, T., Okada, K., Aoki, K., Ose, T., Masatani, T., Sugiyama, M., Ito, N., Fukuhara, H., & Maenaka, K. (2021). Characterization of Single-Chain Fv Fragments of Neutralizing Antibodies to Rabies Virus Glycoprotein. Viruses, 13(11), 2311. https://doi.org/10.3390/v13112311






