1. Introduction
Human astrovirus (HAstV) is an important etiological agent of gastroenteritis, affecting mainly children, the immunocompromised, and the elderly [
1]. Most of these infections are associated with the so-called canonical or classic HAstVs, which comprises eight different serotypes (HAstV-1 to -8), with HAstV-1 being the most prevalent worldwide [
1]. In recent years, new strains have also been reported as causing meningitis and encephalitis in immunosuppressed patients, increasing interest in the study of these viruses [
2].
HAstVs are small, nonenveloped viruses with a single-stranded positive-sense RNA genome of about 6.8 kb. The capsid of the mature, infectious virus has an icosahedral morphology of T = 3 and is composed of two proteins: VP34, which forms the shell of the virus particle (core protein), and VP27, a protein that constitutes the 30 dimeric globular spikes that protrude from the virion (spike protein) [
3]. The spike protein, but not the core protein, has been shown to induce neutralizing antibodies [
4] and several neutralizing antigenic determinants have been mapped on the spike, defined either by X-ray crystallography [
5] or by sequencing HAstV-1, -2, and -8 mutants that escape neutralization by monoclonal neutralizing antibodies [
4]. In addition, the HAstV spike has been shown to specifically bind to the surface of Caco-2 cells and to contain the receptor-binding domain [
5].
HAstV has been shown to enter cells by clathrin-mediated endocytosis [
6]; however, very little is known about the viral receptor or other cell factors that may determine virus entry. In this work we took advantage to the fact that the astrovirus capsid core and spike proteins can be successfully produced in
Escherichia coli while maintaining their native structure [
4,
7,
8] to identify interacting cellular proteins relevant for HAstV infection using a far-Western blotting approach. We found protein disulfide isomerase A4 (PDIA4) is an entry factor associated with genome uncoating of HAstV serotypes 1 and 8.
2. Material and Methods
2.1. Cells and Viruses
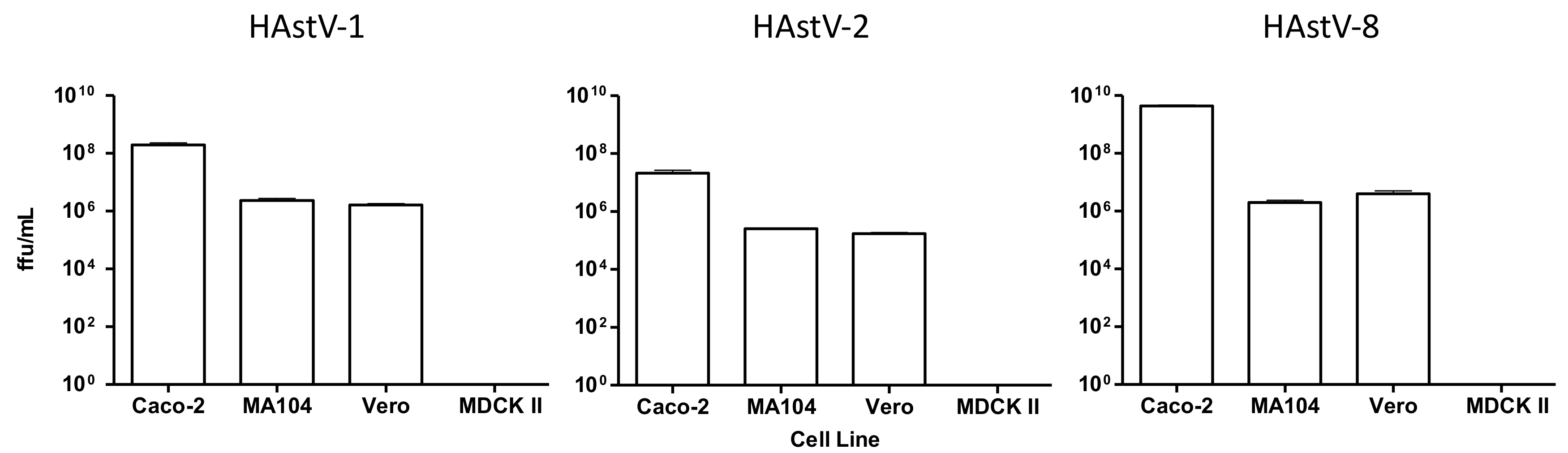
Human colorectal adenocarcinoma cells Caco-2, clone Bbe1 (ATCC, Manassas, VA, USA; CRL-2102), were cultured in high-glucose Dulbecco’s modified Eagle’s medium (DMEM-HG) supplemented with 10% fetal bovine serum (FBS). African green monkey kidney epithelial MA104 (ATCC, Manassas, VA, USA; CRL-2378), Vero (ATCC, Manassas, VA, USA; CCL-81), and Madin-Darby canine kidney (MDCK II) cells (ATCC, Manassas, VA, USA; CRL-2936) were grown in DMEM containing 5% FBS. Human astrovirus serotypes 1 (HAstV-1, strain Oxford), 2 (HAstV-2, strain Oxford) and 8 (HAstV-8, strain Yuc-8) were grown in Caco-2 cells [
4]. Stocks of HAstV-1, HAstV-2 and HAstV-8 were prepared in Caco-2 cells, as previously described [
9]. Briefly, the virus was activated with 200 µg/mL of trypsin for 1 h at 37 °C, and soybean trypsin inhibitor (Sigma, St. Louis, MO, USA) at 200 µg/mL was added immediately before inoculation of cells. The virus was adsorbed for 1 h at 37 °C, and after this time the inoculum was removed and the infection was left to proceed for 16 h at 37 °C.
2.2. Virus Infectivity Assay
The infectious titer of the virus stocks was determined in monolayers of Caco-2 cells grown in 96-well plates. For this, cells were infected as described above with serial two-fold dilutions of the virus stock, and after 16 h of incubation at 37 °C, the cells were fixed with 2% paraformaldehyde in PBS for 20 min at room temperature (RT) and permeabilized by incubation with 2% Triton X-100 in PBS for 15 min at RT. The viral antigen was detected with serotype-specific polyclonal antibodies raised to the capsid spike protein of HAstV-1 (rabbit anti-spike1; dil. 1:1000), -2 (mouse anti-spike2; dil. 1:200) or -8 (rabbit anti-HAstV Yuc8; dil. 1:2000), as described [
4], followed by incubation with the corresponding species-specific peroxidase-conjugated antibodies (KPL, Gaithersburg, MD, USA) diluted 1:3000. The focus forming units (ffus) were visually counted in in a Nikon TMS inverted phase-contrast microscope with a 20× objective.
2.3. Expression and Purification of Recombinant HAstV Capsid Spike Proteins
Expression and purification of recombinant HAstV capsid spike proteins were performed as described previously [
8,
9]. Briefly, an expression plasmid encoding a HAstV spike in-frame with a C-terminal 10-histidine tag was transformed into
E. coli strain BL21(DE3), and protein production was induced with isopropyl-β-D-thiogalactopyranoside.
E. coli cells were lysed by ultrasonication and soluble HAstV spike was purified by metal affinity chromatography. HAstV spike was dialyzed into PBS and further purified by size-exclusion chromatography on a Superdex 200 column in PBS. Synthetic genes codon optimized for
E. coli-encoded HAstV-2-Oxford capsid spike amino acids 431 to 674 (GenBank accession number KY964327), HAstV-1 capsid spike amino acid 429 to 645 (accession number AAC34717.1), or HAstV-8 capsid spike amino acids 429 to 647 (UniProtKB entry Q9IFX1) were used.
2.4. Far-Western Blot Analysis
Cell membrane protein extracts from the different cell lines were prepared using the Qproteome Cell Compartment Kit (Qiagen, Hilden, Germany) according to the manufacturer’s instructions. The proteins in these extracts (around 100 µg per lane) were separated by 10% SDS-polyacrylamide gel electrophoresis (SDS-PAGE) and transferred to nitrocellulose membranes. The nitrocellulose membranes were blocked with 5% skim milk (Carnation, Solon, OH, USA) in PBS-0.1% Tween 20 for 1 h at room temperature, and after this time the membranes were incubated overnight at 4 °C with 100 µg/mL of the corresponding recombinant spike of HAstV serotypes 1, 2 or 8 in 1% skim milk prepared with PBS-0.1% Tween 20, and subsequently washed three times with PBS-0.1% Tween 20. Anti-spike mouse or rabbit polyclonal antibodies generated in our laboratory were added at a 1:1000 dilution and incubated for 1 h at RT. After washing the membranes three times with PBS-0.1% Tween 20, peroxidase-conjugated rabbit or mouse antibodies (KPL, Gaithersburg, MD, USA, dil. 1:3000) were added and incubated for 1 h at RT. Finally, the membranes were washed once and the presence of the astrovirus spike was detected by chemiluminescence using the Western Lightning system (Perkin Elmer, Waltham, MA, USA).
2.5. LC-MS
Protein bands in the 70 to 75 kDa range identified in the far-Western assay were cut out from a parallel SDS gel stained with Coomassie brilliant blue R-250 (Sigma, St. Louis, MO, USA), and the polyacrylamide slices were sent to the Proteomic facility at the Instituto de Biotecnología, UNAM (Cuernavaca, Mexico) for their identification by nano-liquid chromatography (nano-LC)-tandem mass spectrometry (MS/MS).
2.6. Western Blot
Cellular lysates of siRNA-transfected cells or 300 ng of a recombinant PDIA4 protein (Novoprotein, Fremont, CA, USA; CA58) were separated by 10% SDS-PAGE; the proteins were then transferred to nitrocellulose membranes (Millipore, Bedford, MA, USA) as reported [
10]. The membranes were blocked with 5% nonfat milk for 1 h at RT, followed by incubation with anti-PDIA4 monoclonal antibody (ThermoFisher, Rockford, IL, USA; PA1-007; dil. 1:500) for 1 h at RT. The unbound antibody was washed three times with PBS-0.1% Tween 20, and secondary anti-mouse peroxidase-conjugated antibodies, diluted 1:3000 in PBS-0.1% Tween 20, was added and incubated for 1 h at RT. After this time, the membranes were washed three times with PBS-0.1% Tween 20, and the peroxidase activity was detected using the Western Lightning Chemiluminescence Reagent Plus (PerkinElmer Life Sciences, Boston, MA, USA) according to the manufacturer’s instructions.
2.7. RNA Interference
Caco-2 cells were transfected with 5 μM of the siRNA pool of either PDIA4 (Dharmacon M-019249-01-0005) or an irrelevant scrambled control (Dharmacon, Lafayette, CO, USA; D-001206-13-05) using oligofectamine reagent (Invitrogen, Carlsbad, CA, USA), according to the manufacturer´s instructions. Seventy-two hours later, the transfection mixture was removed and the cells were washed twice with minimum essential medium (MEM) and infected with HAstV-2 or -8 at a multiplicity of infection (MOI) of 3, as described above. At 18 hpi the cells were harvested in Laemmli sample buffer and the presence of PDIA4 was assessed by 10% SDS-PAGE and Western blotting.
2.8. Pharmacological Inhibitors
PDI inhibitors 16F16 (Sigma, St. Louis, MO, USA; SML0021), PACMA 31 (Sigma, St. Louis, MO, USA; SML0838), and di-tert-butyl nitroxide (DTBN, Sigma, St. Louis, MO, USA; 300721) were dissolved in DMSO and used at the indicated concentrations. Caco-2 cells grown in 96-well plates were washed once with MEM and then pretreated with the inhibitors, or DMSO as control, for 1 h at RT. After removal of the drugs, the cell monolayers were washed twice with MEM and then infected with the different serotypes of HAstV at an MOI of 0.025 for 1 h at 37 °C. After 16 hpi the cells were fixed and the virus infectivity was determined by a focus-forming unit (ffu) assay, as described above.
2.9. Virus Binding Assay
HAstV-8 virus particles, purified as previously described [
11] were added at an MOI of 30 to Caco-2 cell monolayers in 48-well plates that were pretreated or not with the drugs or with siRNAs as described above, for 1 h on ice. The unbound virus was washed three times with cold PBS. Total RNA was extracted with TRIzol reagent (Invitrogen, Carlsbad, CA, USA) according to the manufacturer´s instructions. cDNA was generated by reverse transcription using M-MLV Reverse Transcriptase (Invitrogen, Carlsbad, CA, USA) and qPCR was performed with Real Q Plus Master Mix Green (Ampliqon, Copenhagen, Denmark) using an ABI Prism 7500 detection system (Applied Biosystems, Foster City, CA, USA). The primers used amplify a region of the ORF 1b of HAstV-8; the amplification conditions were previously reported [
12]. The amount of viral RNA was normalized to that of 18S RNA [
13].
2.10. Viral RNA Transfection
Caco-2 cells were infected with HAstV-8 at an MOI of 2 for 16 h at 37 °C. After this time, total RNA was extracted with TRIzol reagent (Invitrogen, Carlsbad, CA, USA) according to the manufacturer´s instructions. The extracted RNA was used to transfect Caco-2 cells previously transfected with PDIA4 siRNAs or pretreated with the PDI inhibitor 16F16. Briefly, Caco-2 cell monolayers at 60–80% confluence in 48-well plates were incubated for 6 h at 37 °C with a mixture of Lipofectamine 2000 (Invitrogen, Carlsbad, CA, USA) and 200 ng of total RNA extracted from HAstV-8-infected Caco-2 cells. After this time, the transfection mixture was removed, the cells were washed twice with MEM, fresh MEM was added, and the cells were incubated for 24 h before staining for the presence of HAstV-8 focus-forming units (ffu’s) using an immunoperoxidase assay, as described above.
2.11. Virus Internalization Assay
Confluent Caco-2 cell monolayers in 48-well plates were pretreated with 500 µM of 16F16 for 1 h at RT, and then HAstV-8 at an MOI of 50 was added and the cells were further incubated for 1 h at 4 °C. After this time, the unbound virus was removed by washing the cells three times with PBS and the cell monolayers were switched to 37 °C for 1 h to allow the virus to be internalized. To remove the virus adsorbed to the cell surface that had not been internalized during the incubation period at 37 °C, the cells were incubated with neutralizing monoclonal antibody 2D9 against HAstV-8, diluted 1:20, for 1 h at 4 °C. At this concentration, this antibody is able to remove up to 80% of the cell surface bound virus (unpublished data). Then, the internalized viral RNA was quantified by qRT-PCR as described above.
2.12. Viral Uncoating Assay
To determine the moment at which the virus genome is uncoated and released from the endocytic vesicle, a neutral red assay was performed, as previously described [
6]. Briefly, neutral red-labeled HAstV-8 virus stock was prepared by growing the virus in Caco-2 cells in the presence of 10 µg/mL neutral red for 18 h at 37 °C protected from light. After this time, the virus was harvested and titrated by an immunoperoxidase assay as described above, under light or in a dark room to confirm its photosensitivity. To evaluate the effect of 16F16 on virus uncoating, confluent Caco-2 cell monolayers were pretreated with the drug for 1 h at RT, then the cells were infected with neutral red-labeled HAstV-8 at an MOI of 0.025 and incubated in the dark for 2 h at 37 °C to allow the virus to enter the cells. After this time the cells were exposed to direct white light for 10 min. After light exposure, the infection was continued without protection from light for 16 h at 37 °C; the cells were then fixed and the infectivity determined by an immunoperoxidase assay.
2.13. Statistical Analysis
The statistical significance of the data was evaluated by the Mann–Whitney test using GraphPad Prism 5 (GraphPad Software Inc., San Diego, CA, USA).
4. Discussion
During the process of cell entry, viruses must recognize cellular receptors present on the surface of the host cell but also frequently interact with additional cell molecules that facilitate the internalization of the viral particles and/or the delivery of transcription-competent virus structures, which range from the naked virus genome to subviral particles.
We have previously reported that HAstV-8 enters Caco-2 cells using a clathrin-dependent pathway, and drugs that reduce the cholesterol in the cell membrane or that inhibit actin polymerization decrease astrovirus infectivity [
6]; however, the cellular receptor for these viruses has not yet been identified, although initial binding of the virus to glycans on the cell surface has been suggested [
6,
7]. A virus overlay protein binding assay has been used to characterize putative cellular receptors of different viruses, including adenovirus [
18], Japanese encephalitis virus [
19], nervous necrosis virus [
20], Newcastle disease virus [
21], and rotavirus [
22]. In this work, considering the availability of a recombinant capsid spike protein that mimics the virus domain through which HAstV attaches to the cell surface [
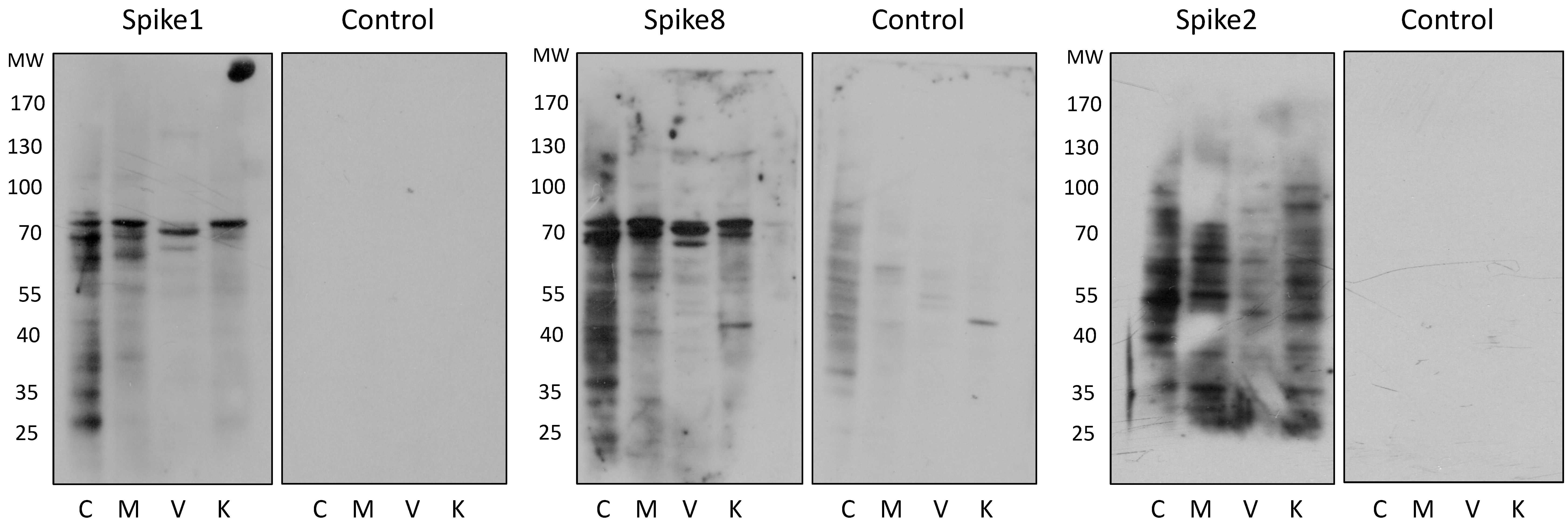
5], we used a far-Western approach to identify cell membrane proteins that could potentially interact with the virus during cell entry. Using this approach we found two distinctive cellular proteins, in the range of 70 to 75 kDa, that were recognized by spikes of HAstV serotypes 1 and 8. Of interest, these two bands, with slight variations in their molecular weight, were recognized in permissive as well as in nonpermissive MDCKII cells, indicating that if this protein is the homolog in the various cell lines tested, this interaction might not be the one that confers specificity of the virus to infect Caco-2 cells.
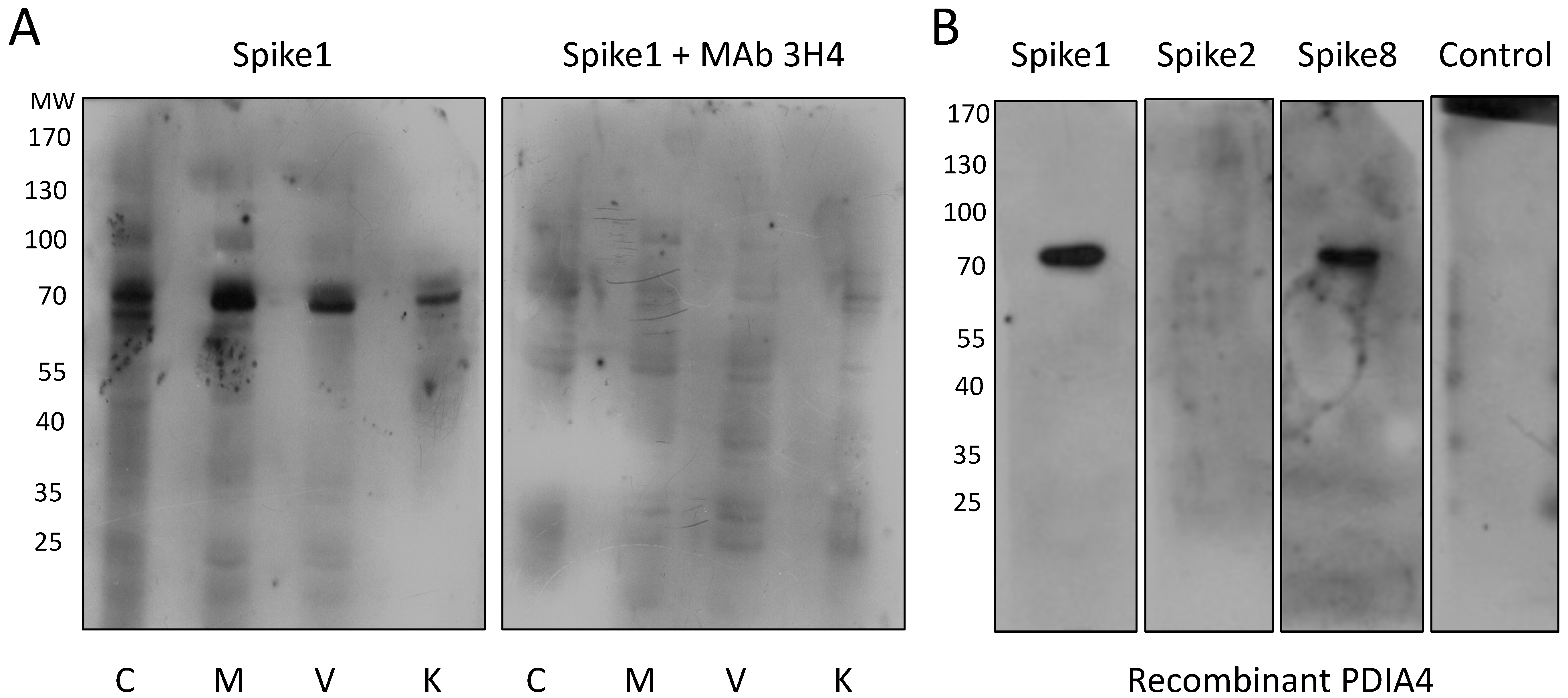
Analysis by LC-MS/MS of proteins in the 70–75 kDa range recovered from preparative SDS-polyacrylamide gels identified several membrane proteins of interest which have been previously reported as cell entry factors for different viruses (
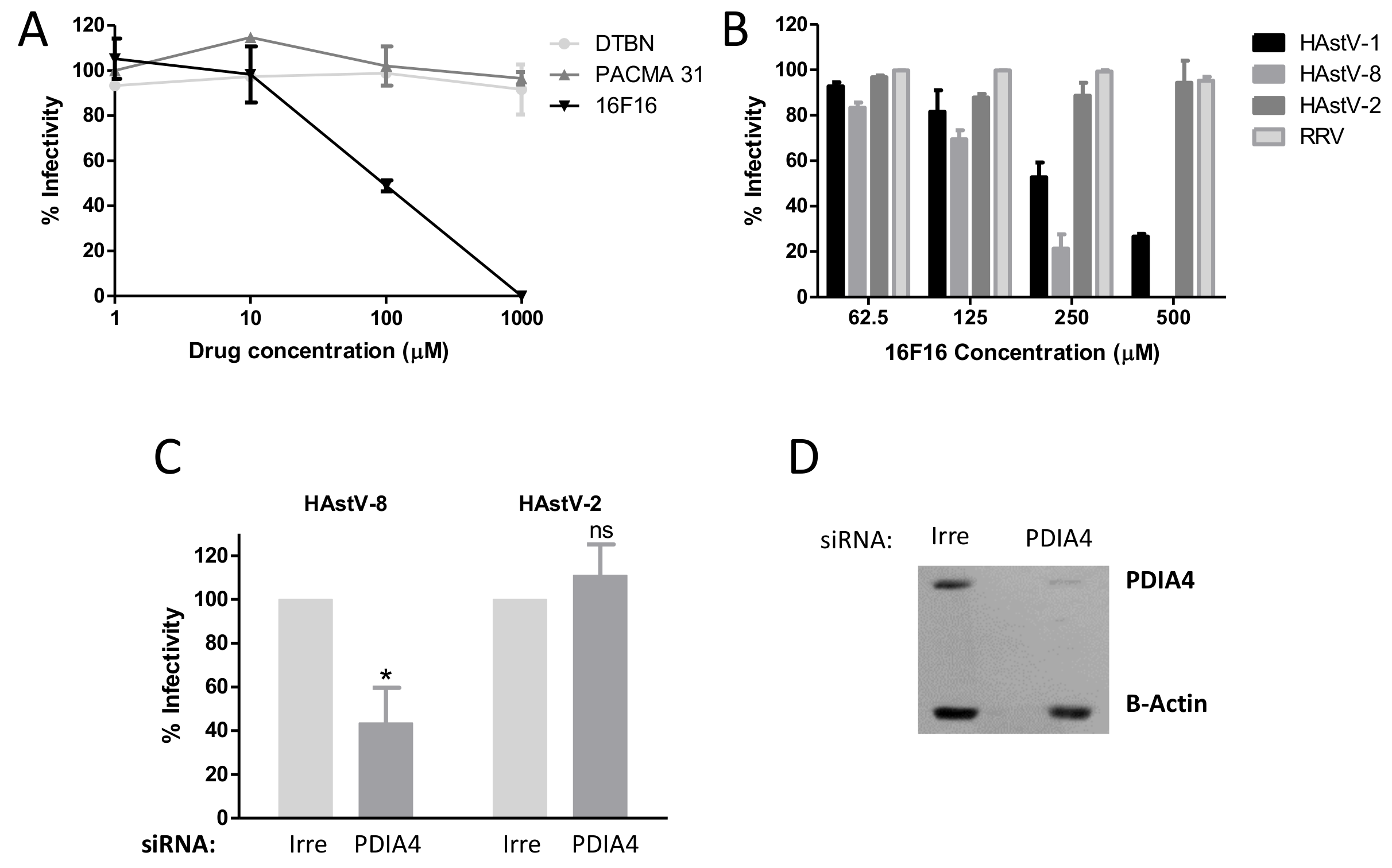
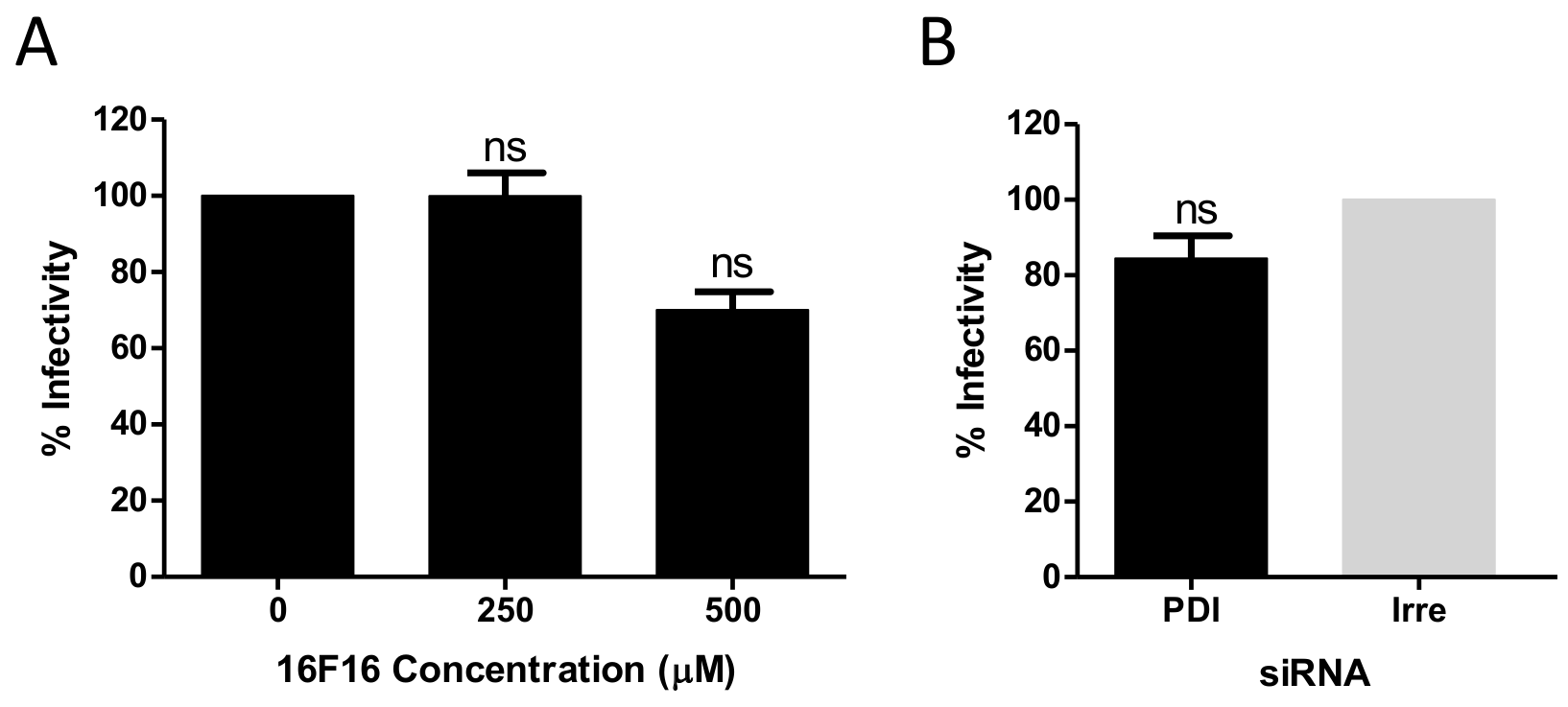
Table S1). Out of these, we chose to further characterize PDIA4 (also known as Erp72), a 72 kDa member of a protein family that catalyzes the formation, breakage, and rearrangement of disulfide bonds within protein molecules. Initially, a pharmacological approach was employed, testing several inhibitors of PDI activity. We found that 16F16 strongly decreased HAstV-1 and -8 infectivity in a dose-dependent manner, while it did not affect the infectivity of HAstV-2. These findings are consistent with the different binding patterns observed in the far-Western assay for spike 2 as compared to spikes 1 and 8, indicating a serotype-specific effect of the drug on HAstV infection. The specificity of the drug action is supported by the fact that, similarly to HAstV-2, the inhibitor did not affect the infectivity of rotavirus RRV, which belongs to a different virus family. It is interesting that the other PDI inhibitors tested did not inhibit HAstV-1 or -8 infection; probably because the tested drugs have different biochemical properties and mechanisms of action. Importantly, 16F16 has been shown to be permeable to cells, contrasting with DTBN, which is impermeable [
23], suggesting that the internalization of the drug into the cells is important. This is true for influenza infection, where the permeability of PDI inhibitors correlated with their antiviral activity [
24]. Furthermore, the selectivity of each drug for its PDI target has to be considered; while PACMA 31 has been reported to be more selective for PDIA1 (57 kDa) [
25], 16F16 is able to interact with other PDI family proteins as PDIA6 (48 kDa) and PDIA4 (72 kDa; Erp72) [
16]. The effect of silencing the expression of PDIA4 using a pool of siRNAs directed to its mRNA further confirmed the role of this protein in the HAstV infection cycle, although it cannot be discarded that 16F16 may target PDI enzymes in addition to PDIA4 that may play a role in HAstV infection.
Cell surface PDI enzymatic activity has been associated with the cell entry of dengue virus [
26,
27], Sindbis virus [
28], Newcastle disease virus [
29], and human immunodeficiency virus [
30]. PDI may directly interact and modify viral proteins during this process, as reported for human immunodeficiency virus and the Newcastle disease virus, since their fusion proteins undergo a disulfide bond reduction by PDI activity, inducing conformational changes that allow the virus to enter the cell [
29,
30,
31]. The activity of PDI may also indirectly participate in virus entry; in the case of dengue virus it has been reported that PDI inhibitors prevent virus entry into endothelial cells by blocking the activation of integrins, a requirement for virus infection [
28], although the possibility of a direct PDI effect on dengue virus proteins cannot be excluded. Of interest, PDI inhibitors decrease Newcastle disease virus and human immunodeficiency virus cell entry, although binding of these viruses to their receptor is not affected [
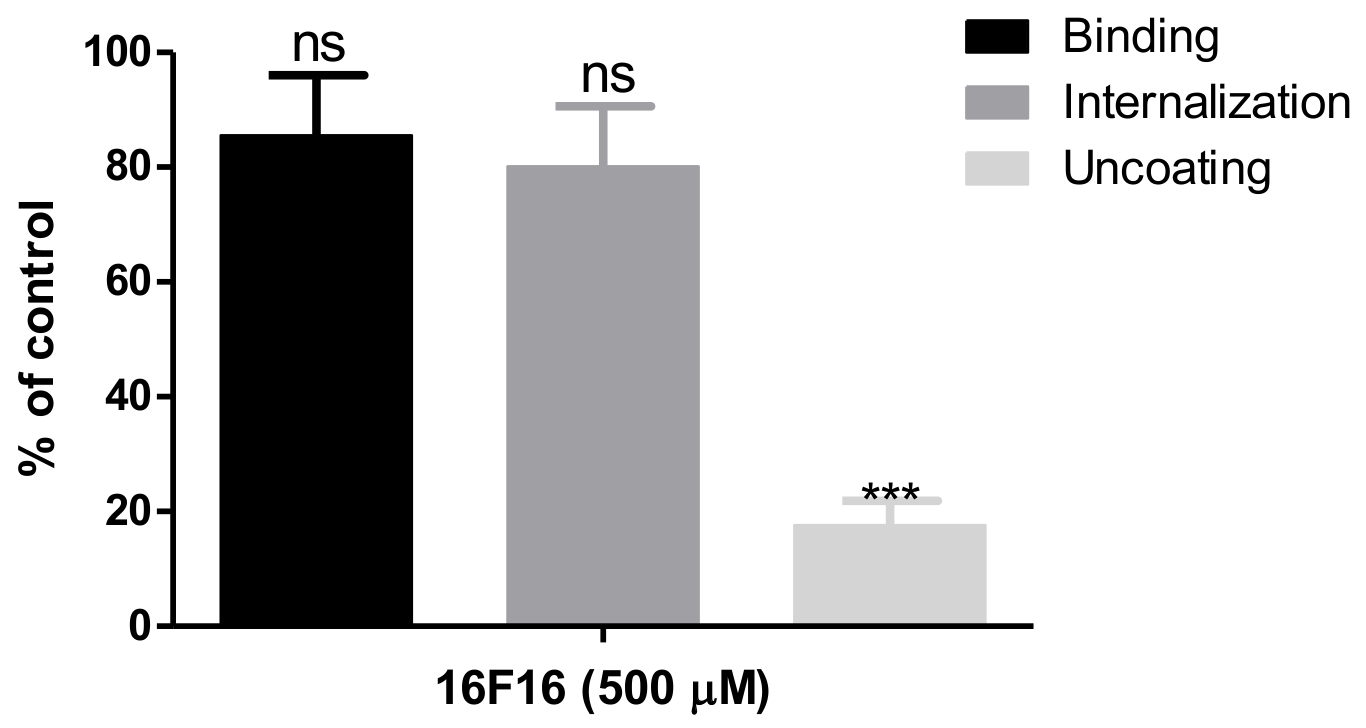
29,
30]. Similar to these observations, in this study we found that the 16F16 PDI inhibitor does not prevent the attachment of the virus to the cell surface or its internalization, but rather it seems to affect virus uncoating. These findings indicate the PDI enzymatic activity in HAstV-1 and -8 infection is required at a post-attachment step.
Whether the HAstV capsid proteins undergo a conformational change due to the enzymatic or chaperone activity of PDI4 that facilitates virus uncoating or whether the PDI activity modifies the oxidation state of a cellular protein that in turn facilitates HAstV uncoating remains to be determined. Future studies should address whether the spike of some serotypes of HAstV is a substrate for PDI activity and how this activity allows the virus genome to be uncoated and be released into the cell’s cytosol.