Identification and Distribution of Novel Cressdnaviruses and Circular Molecules in Four Penguin Species in South Georgia and the Antarctic Peninsula
Abstract
1. Introduction
2. Materials and Methods
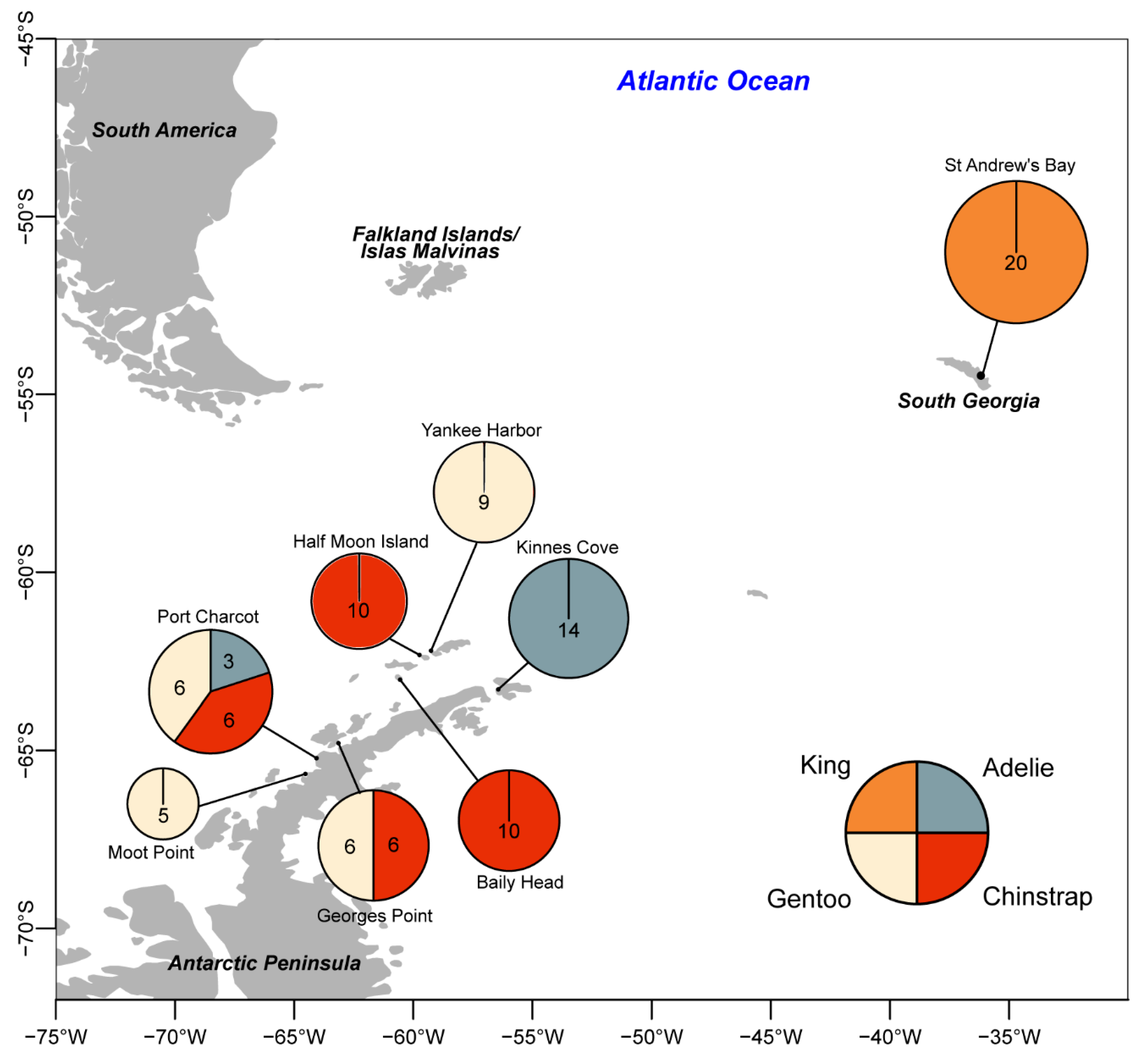
2.1. Field Sampling
2.2. Viral Extraction and High-Throughput Sequencing Analysis
2.3. Recovery and Sequencing of Viral Genomes and Viral-Like Circular Elements
2.4. Bioinformatic Analyses of Recovered Viral Genomes and Viral-Like Elements
3. Results and Discussion
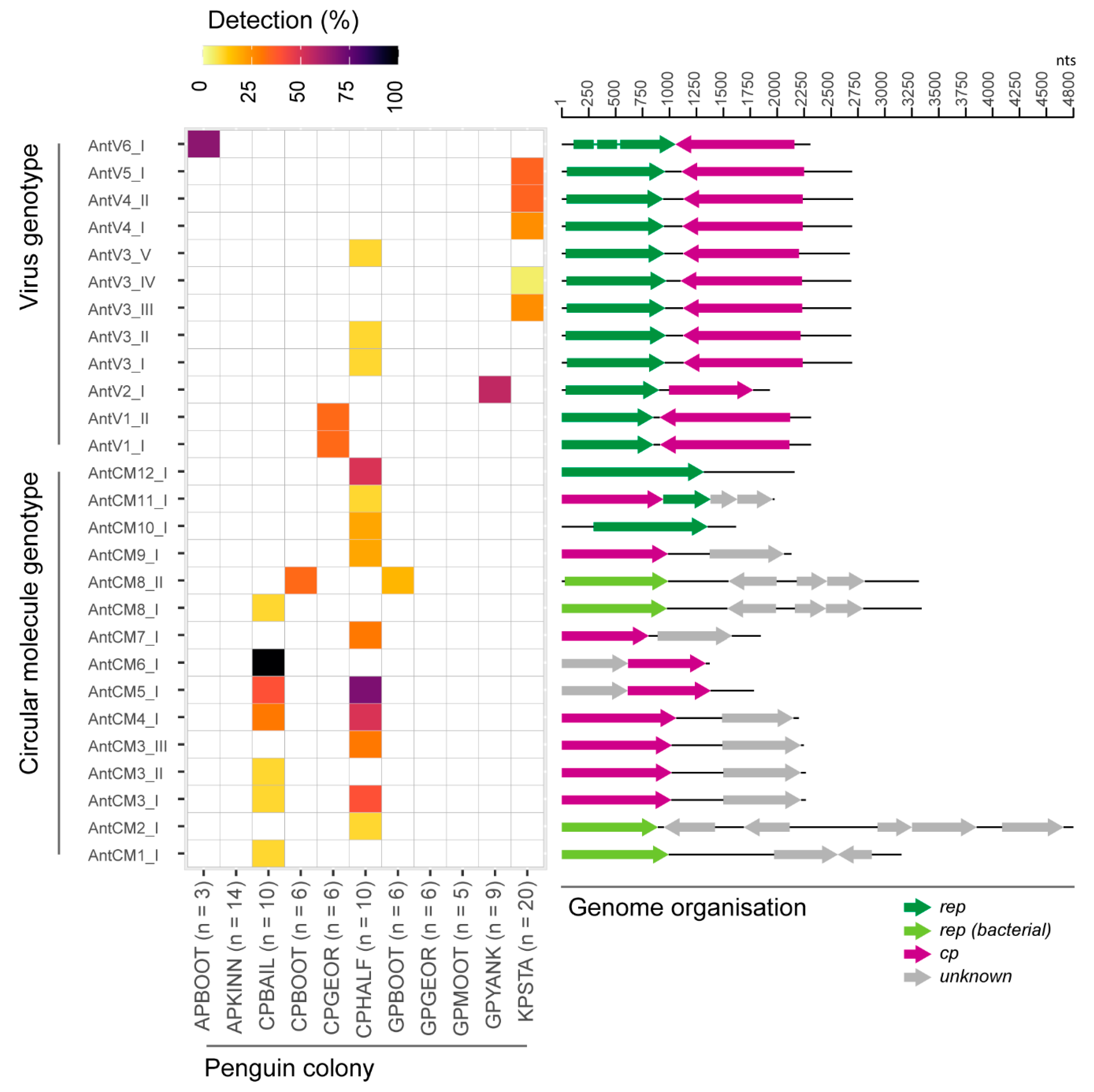
3.1. Identification of Viruses and Viral-Like Molecules from Cloacal Swabs
3.2. Distribution of Viruses and Viral-Like Molecules across Sampling Sites and Penguin Species
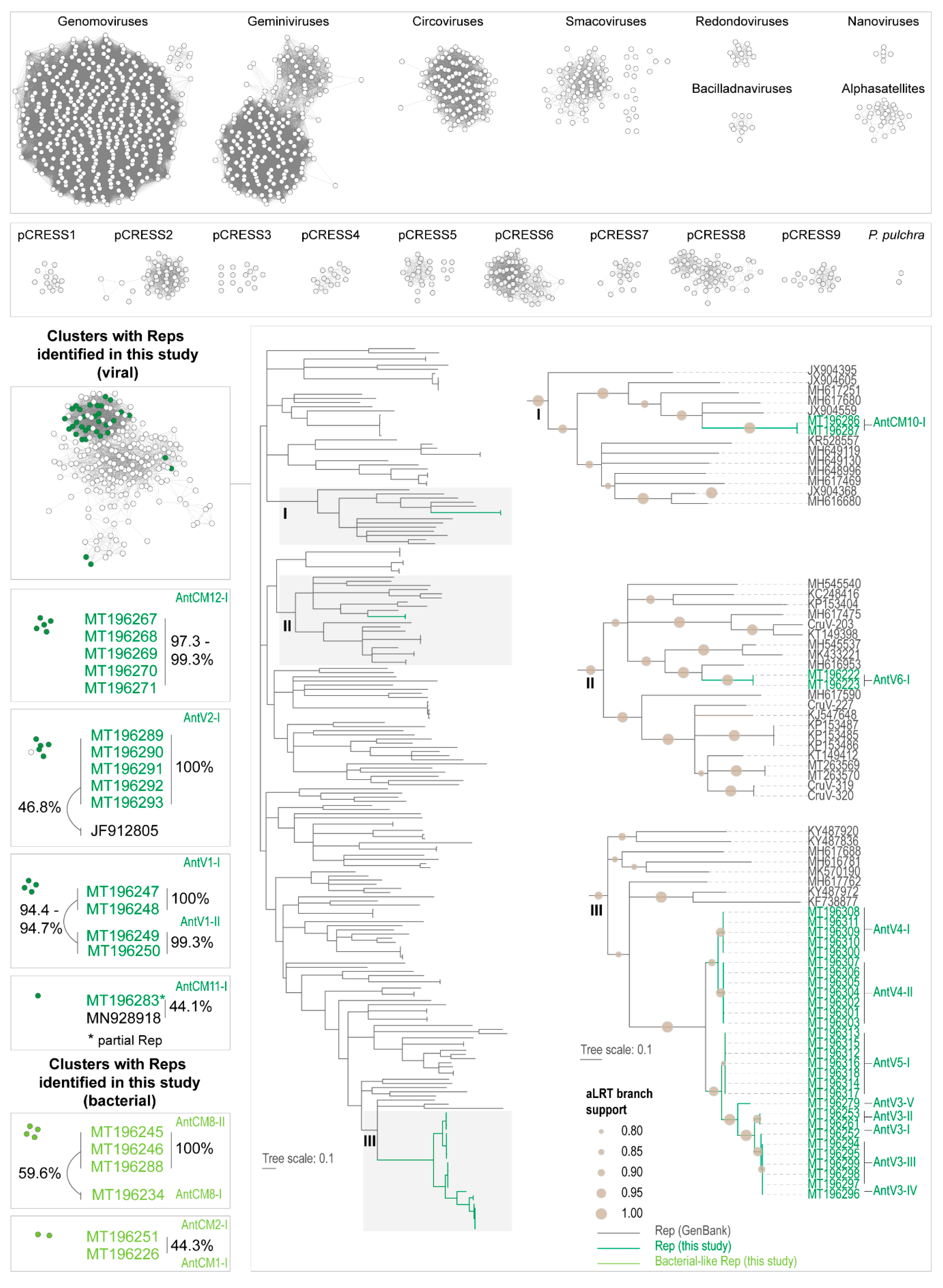
3.3. Analyses of the Replication-Associated Protein
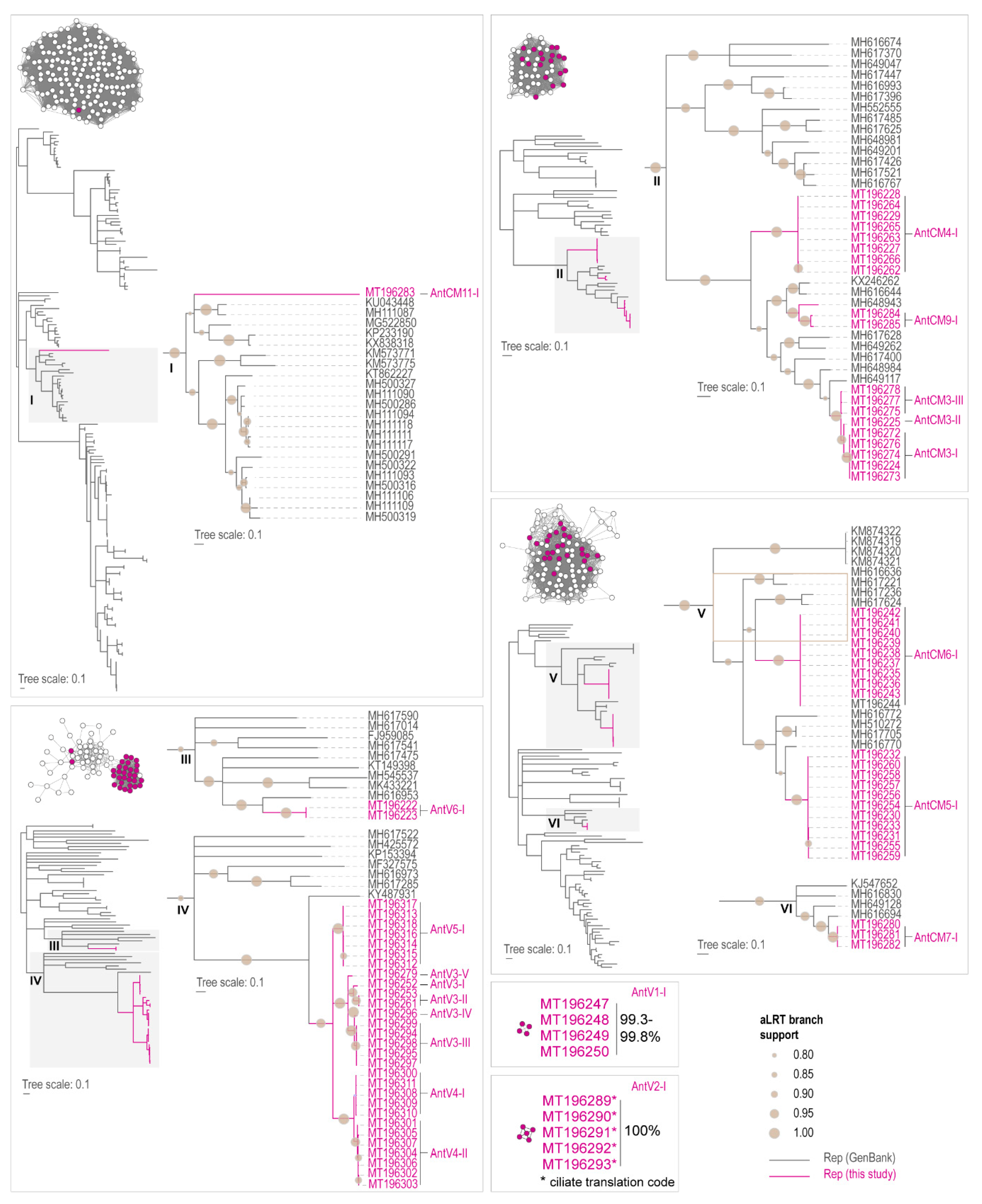
3.4. Analyses of the Capsid Protein
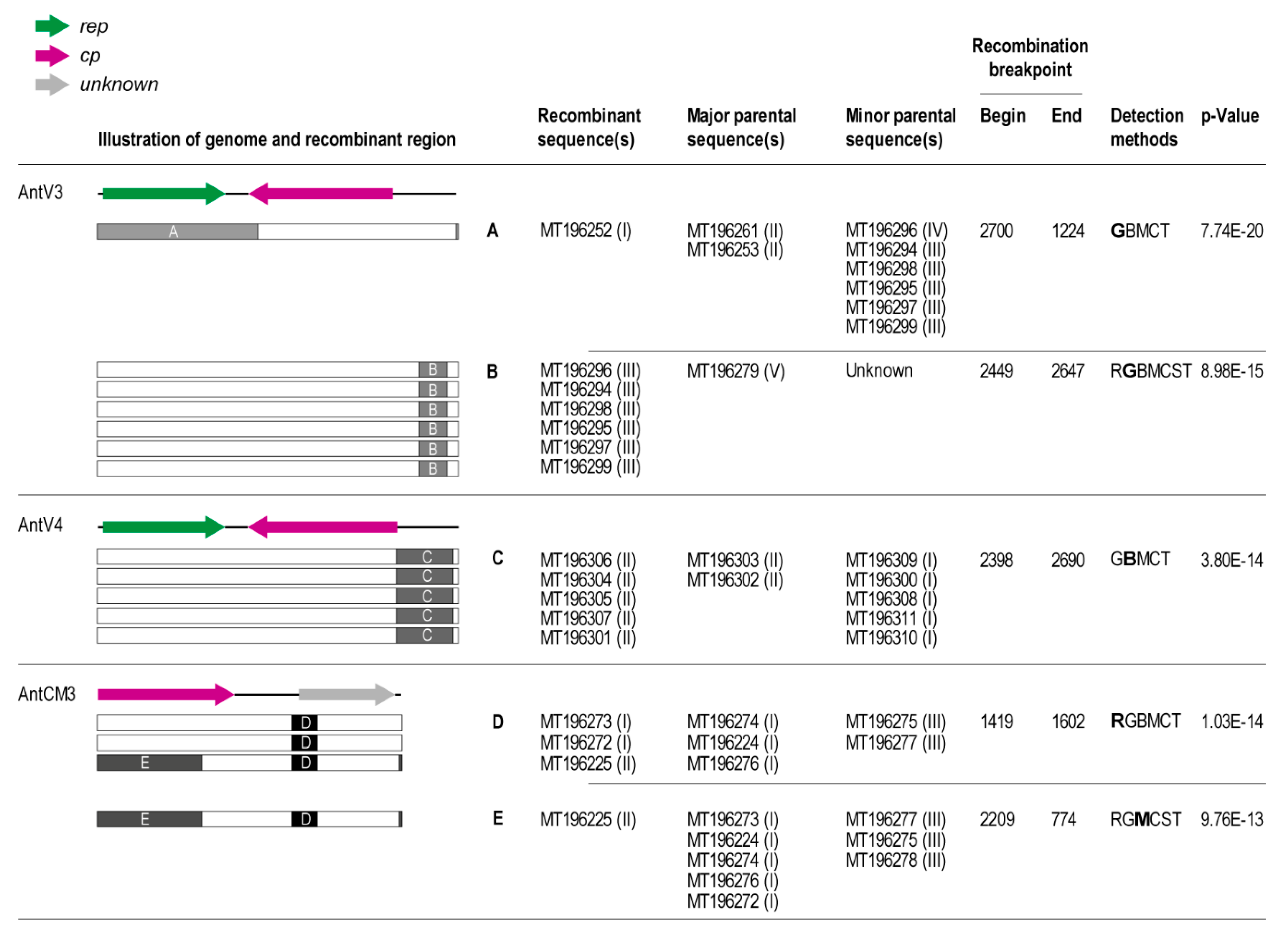
3.5. Recombination Analysis
4. Concluding Remarks
Supplementary Materials
Author Contributions
Funding
Conflicts of Interest
References
- Cole, T.L.; Dutoit, L.; Dussex, N.; Hart, T.; Alexander, A.; Younger, J.L.; Clucas, G.V.; Frugone, M.J.; Cherel, Y.; Cuthbert, R.; et al. Receding ice drove parallel expansions in Southern Ocean penguins. Proc. Natl. Acad. Sci. USA 2019, 166, 26690–26696. [Google Scholar] [CrossRef]
- Gavryushkina, A.; Heath, T.A.; Ksepka, D.T.; Stadler, T.; Welch, D.; Drummond, A.J. Bayesian Total-Evidence Dating Reveals the Recent Crown Radiation of Penguins. Syst. Biol. 2017, 66, 57–73. [Google Scholar] [CrossRef] [PubMed]
- Gardner, H.; Kerry, K.; Riddle, M.; Brouwer, S.; Gleeson, L. Poultry virus infection in Antarctic penguins. Nature 1997, 387, 245. [Google Scholar] [CrossRef] [PubMed]
- Watts, J.M.; Miller, G.D.; Shellam, G.R. Infectious Bursal Disease Virus and Antarctic Birds; Springer: Berlin/Heidelberg, Germany, 2009; pp. 95–105. [Google Scholar]
- Morgan, I.R.; Westbury, H.A. Virological studies of Adelie Penguins (Pygoscelis adeliae) in Antarctica. Avian Dis. 1981, 25, 1019–1026. [Google Scholar] [CrossRef]
- Austin, F.J.; Webster, R.G. Evidence of ortho- and paramyxoviruses in fauna from Antarctica. J. Wildl. Dis. 1993, 29, 568–571. [Google Scholar] [CrossRef] [PubMed]
- Bengtson, J.L.; Boveng, P.; Franzen, U.; Have, P.; Heidejorgensen, M.P.; Harkonen, T.J. Antibodies to Canine-Distemper Virus in Antarctic Seals. Mar. Mammal Sci. 1991, 7, 85–87. [Google Scholar] [CrossRef]
- McFarlane, R.A. Health Assessment and Diseases of the Weddell seal, Leptonochotes weddelli, in Vestfold Hills, East Antarctica. In Health of Antarctic Wildlife; Kerry, K.R., Riddle, M., Eds.; Springer: Berlin/Heidelberg, Germany, 2009. [Google Scholar]
- Mac Donald, J.W.; Conroy, J.W.H. Virus Disease Resembling Puffinosis in the Gentoo Penguin Pygoscelis papua on Signy Island South Orkney Islands. British Antarct. Surv. Bull. 1971, 26, 80–82. [Google Scholar]
- Munro, G. Outbreak of Avian Pox Virus in Gentoo Penguins in the Falklands; Falklands Conservation: Stanley, Falkland Islands, 2006. [Google Scholar]
- Pistorius, P.A. Falkland Island Seabird Monitoring Programme. Annual Report 2008/2009; Falklands Conservation: Stanley, Falkland Islands, 2009. [Google Scholar]
- Smeele, Z.E.; Ainley, D.G.; Varsani, A. Viruses associated with Antarctic wildlife: From serology based detection to identification of genomes using high throughput sequencing. Virus Res. 2018, 243, 91–105. [Google Scholar] [CrossRef]
- Wille, M.; Harvey, E.; Shi, M.; Gonzalez-Acuna, D.; Holmes, E.C.; Hurt, A.C. Sustained RNA virome diversity in Antarctic penguins and their ticks. ISME J. 2020, 14, 1768–1782. [Google Scholar] [CrossRef] [PubMed]
- Adriaenssens, E.M.; Kramer, R.; Van Goethem, M.W.; Makhalanyane, T.P.; Hogg, I.; Cowan, D.A. Environmental drivers of viral community composition in Antarctic soils identified by viromics. Microbiome 2017, 5, 83. [Google Scholar] [CrossRef]
- Zablocki, O.; van Zyl, L.; Adriaenssens, E.M.; Rubagotti, E.; Tuffin, M.; Cary, S.C.; Cowan, D. Niche-dependent genetic diversity in Antarctic metaviromes. Bacteriophage 2014, 4, e980125. [Google Scholar] [CrossRef]
- Zablocki, O.; van Zyl, L.; Adriaenssens, E.M.; Rubagotti, E.; Tuffin, M.; Cary, S.C.; Cowan, D. High-level diversity of tailed phages, eukaryote-associated viruses, and virophage-like elements in the metaviromes of antarctic soils. Appl. Environ. Microbiol. 2014, 80, 6888–6897. [Google Scholar] [CrossRef] [PubMed]
- Lopez-Bueno, A.; Tamames, J.; Velazquez, D.; Moya, A.; Quesada, A.; Alcami, A. High diversity of the viral community from an Antarctic lake. Science 2009, 326, 858–861. [Google Scholar] [CrossRef] [PubMed]
- Yau, S.; Lauro, F.M.; DeMaere, M.Z.; Brown, M.V.; Thomas, T.; Raftery, M.J.; Andrews-Pfannkoch, C.; Lewis, M.; Hoffman, J.M.; Gibson, J.A.; et al. Virophage control of antarctic algal host-virus dynamics. Proc. Natl. Acad. Sci. USA 2011, 108, 6163–6168. [Google Scholar] [CrossRef] [PubMed]
- Rastrojo, A.; Alcamí, A. Chapter Two—Viruses in Polar Lake and Soil Ecosystems. In Advances in Virus Research; Malmstrom, C.M., Ed.; Academic Press: Cambridge, MA, USA, 2018; Volume 101, pp. 39–54. [Google Scholar]
- de Carcer, D.A.; Lopez-Bueno, A.; Alonso-Lobo, J.M.; Quesada, A.; Alcami, A. Metagenomic analysis of lacustrine viral diversity along a latitudinal transect of the Antarctic Peninsula. FEMS Microbiol. Ecol. 2016, 92, fiw074. [Google Scholar] [CrossRef] [PubMed]
- Yau, S.; Seth-Pasricha, M. Viruses of Polar Aquatic Environments. Viruses 2019, 11, 189. [Google Scholar] [CrossRef]
- Zawar-Reza, P.; Arguello-Astorga, G.R.; Kraberger, S.; Julian, L.; Stainton, D.; Broady, P.A.; Varsani, A. Diverse small circular single-stranded DNA viruses identified in a freshwater pond on the McMurdo Ice Shelf (Antarctica). Infect. Genet. Evol. 2014, 26, 132–138. [Google Scholar] [CrossRef]
- Gong, Z.; Liang, Y.; Wang, M.; Jiang, Y.; Yang, Q.; Xia, J.; Zhou, X.; You, S.; Gao, C.; Wang, J.; et al. Viral Diversity and Its Relationship with Environmental Factors at the Surface and Deep Sea of Prydz Bay, Antarctica. Front. Microbiol. 2018, 9, 2981. [Google Scholar] [CrossRef]
- Brum, J.R.; Ignacio-Espinoza, J.C.; Roux, S.; Doulcier, G.; Acinas, S.G.; Alberti, A.; Chaffron, S.; Cruaud, C.; de Vargas, C.; Gasol, J.M.; et al. Ocean plankton. Patterns and ecological drivers of ocean viral communities. Science 2015, 348, 1261498. [Google Scholar] [CrossRef]
- Miranda, J.A.; Culley, A.I.; Schvarcz, C.R.; Steward, G.F. RNA viruses as major contributors to Antarctic virioplankton. Environ. Microbiol. 2016, 18, 3714–3727. [Google Scholar] [CrossRef] [PubMed]
- Sommers, P.; Fontenele, R.S.; Kringen, T.; Kraberger, S.; Porazinska, D.L.; Darcy, J.L.; Schmidt, S.K.; Varsani, A. Single-Stranded DNA Viruses in Antarctic Cryoconite Holes. Viruses 2019, 11, 1022. [Google Scholar] [CrossRef]
- Krupovic, M.; Varsani, A.; Kazlauskas, D.; Breitbart, M.; Delwart, E.; Rosario, K.; Yutin, N.; Wolf, Y.I.; Harrach, B.; Zerbini, F.M.; et al. Cressdnaviricota: A Virus Phylum Unifying Seven Families of Rep-Encoding Viruses with Single-Stranded, Circular DNA Genomes. J. Virol. 2020, 94. [Google Scholar] [CrossRef]
- Bolger, A.M.; Lohse, M.; Usadel, B. Trimmomatic: A flexible trimmer for Illumina sequence data. Bioinformatics 2014, 30, 2114–2120. [Google Scholar] [CrossRef]
- Bankevich, A.; Nurk, S.; Antipov, D.; Gurevich, A.A.; Dvorkin, M.; Kulikov, A.S.; Lesin, V.M.; Nikolenko, S.I.; Pham, S.; Prjibelski, A.D.; et al. SPAdes: A new genome assembly algorithm and its applications to single-cell sequencing. J. Comput. Biol. 2012, 19, 455–477. [Google Scholar] [CrossRef]
- O’Leary, N.A.; Wright, M.W.; Brister, J.R.; Ciufo, S.; McVeigh, D.H.R.; Rajput, B.; Robbertse, B.; Smith-White, B.; Ako-Adjei, D.; Astashyn, A.; et al. Reference sequence (RefSeq) database at NCBI: Current status, taxonomic expansion, and functional annotation. Nucleic Acids Res. 2016, 44, D733–D745. [Google Scholar] [CrossRef] [PubMed]
- Altschul, S.F.; Gish, W.; Miller, W.; Myers, E.W.; Lipman, D.J. Basic local alignment search tool. J. Mol. Biol. 1990, 215, 403–410. [Google Scholar] [CrossRef]
- Kearse, M.; Moir, R.; Wilson, A.; Stones-Havas, S.; Cheung, M.; Sturrock, S.; Buxton, S.; Cooper, A.; Markowitz, S.; Duran, C.; et al. Geneious Basic: An integrated and extendable desktop software platform for the organization and analysis of sequence data. Bioinformatics 2012, 28, 1647–1649. [Google Scholar] [CrossRef] [PubMed]
- Muhire, B.M.; Varsani, A.; Martin, D.P. SDT: A Virus Classification Tool Based on Pairwise Sequence Alignment and Identity Calculation. PLoS ONE 2014, 9, e108277. [Google Scholar] [CrossRef]
- Huang, Y.; Niu, B.; Gao, Y.; Fu, L.; Li, W. CD-HIT Suite: A web server for clustering and comparing biological sequences. Bioinformatics 2010, 26, 680–682. [Google Scholar] [CrossRef]
- Kazlauskas, D.; Varsani, A.; Koonin, E.V.; Krupovic, M. Multiple origins of prokaryotic and eukaryotic single-stranded DNA viruses from bacterial and archaeal plasmids. Nat. Commun. 2019, 10, 3425. [Google Scholar] [CrossRef]
- Zallot, R.; Oberg, N.O.; Gerlt, J.A. ‘Democratized’ genomic enzymology web tools for functional assignment. Curr. Opin. Chem. Biol. 2018, 47, 77–85. [Google Scholar] [CrossRef] [PubMed]
- Gerlt, J.A. Genomic Enzymology: Web Tools for Leveraging Protein Family Sequence-Function Space and Genome Context to Discover Novel Functions. Biochemistry 2017, 56, 4293–4308. [Google Scholar] [CrossRef]
- Shannon, P.; Markiel, A.; Ozier, O.; Baliga, N.S.; Wang, J.T.; Ramage, D.; Amin, N.; Schwikowski, B.; Ideker, T. Cytoscape: A software environment for integrated models of biomolecular interaction networks. Genome Res. 2003, 13, 2498–2504. [Google Scholar] [CrossRef]
- Katoh, K.; Standley, D.M. A simple method to control over-alignment in the MAFFT multiple sequence alignment program. Bioinformatics 2016, 32, 1933–1942. [Google Scholar] [CrossRef]
- Katoh, K.; Standley, D.M. MAFFT multiple sequence alignment software version 7: Improvements in performance and usability. Mol. Biol. Evol. 2013, 30, 772–780. [Google Scholar] [CrossRef] [PubMed]
- Capella-Gutierrez, S.; Silla-Martinez, J.M.; Gabaldon, T. trimAl: A tool for automated alignment trimming in large-scale phylogenetic analyses. Bioinformatics 2009, 25, 1972–1973. [Google Scholar] [CrossRef]
- Darriba, D.; Taboada, G.L.; Doallo, R.; Posada, D. ProtTest 3: Fast selection of best-fit models of protein evolution. Bioinformatics 2011, 27, 1164–1165. [Google Scholar] [CrossRef] [PubMed]
- Stover, B.C.; Muller, K.F. TreeGraph 2: Combining and visualizing evidence from different phylogenetic analyses. BMC Bioinform. 2010, 11, 7. [Google Scholar] [CrossRef]
- Letunic, I.; Bork, P. Interactive Tree of Life (iTOL) v4: Recent updates and new developments. Nucleic Acids Res. 2019, 47, W256–W259. [Google Scholar] [CrossRef]
- Martin, D.P.; Murrell, B.; Khoosal, A.; Muhire, B. Detecting and Analyzing Genetic Recombination Using RDP4. Methods Mol. Biol. 2017, 1525, 433–460. [Google Scholar] [CrossRef]
- Levy, H.; Fiddaman, S.R.; Djurhuus, A.; Black, C.E.; Kraberger, S.; Smith, A.L.; Hart, T.; Varsani, A. Identification of Circovirus Genome in a Chinstrap Penguin (Pygoscelis antarcticus) and Adelie Penguin (Pygoscelis adeliae) on the Antarctic Peninsula. Viruses 2020, 12, 858. [Google Scholar] [CrossRef] [PubMed]
- Tisza, M.J.; Pastrana, D.V.; Welch, N.L.; Stewart, B.; Peretti, A.; Starrett, G.J.; Pang, Y.S.; Krishnamurthy, S.R.; Pesavento, P.A.; McDermott, D.H.; et al. Discovery of several thousand highly diverse circular DNA viruses. eLife 2020, 9. [Google Scholar] [CrossRef] [PubMed]
- Ng, T.F.; Zhang, W.; Sachsenroder, J.; Kondov, N.O.; da Costa, A.C.; Vega, E.; Holtz, L.R.; Wu, G.; Wang, D.; Stine, C.O.; et al. A diverse group of small circular ssDNA viral genomes in human and non-human primate stools. Virus Evol. 2015, 1, vev017. [Google Scholar] [CrossRef]
- Blinkova, O.; Victoria, J.; Li, Y.; Keele, B.F.; Sanz, C.; Ndjango, J.B.; Peeters, M.; Travis, D.; Lonsdorf, E.V.; Wilson, M.L.; et al. Novel circular DNA viruses in stool samples of wild-living chimpanzees. J. Gen. Virol. 2010, 91, 74–86. [Google Scholar] [CrossRef] [PubMed]
- Steel, O.; Kraberger, S.; Sikorski, A.; Young, L.M.; Catchpole, R.J.; Stevens, A.J.; Ladley, J.J.; Coray, D.S.; Stainton, D.; Dayaram, A.; et al. Circular replication-associated protein encoding DNA viruses identified in the faecal matter of various animals in New Zealand. Infect. Genet. Evol. 2016, 43, 151–164. [Google Scholar] [CrossRef] [PubMed]
- Kraberger, S.; Farkas, K.; Bernardo, P.; Booker, C.; Arguello-Astorga, G.R.; Mesleard, F.; Martin, D.P.; Roumagnac, P.; Varsani, A. Identification of novel Bromus- and Trifolium-associated circular DNA viruses. Arch. Virol. 2015, 160, 1303–1311. [Google Scholar] [CrossRef]
- Reavy, B.; Swanson, M.M.; Cock, P.J.; Dawson, L.; Freitag, T.E.; Singh, B.K.; Torrance, L.; Mushegian, A.R.; Taliansky, M. Distinct circular single-stranded DNA viruses exist in different soil types. Appl Environ. Microbiol. 2015, 81, 3934–3945. [Google Scholar] [CrossRef]
- Phan, T.G.; Mori, D.; Deng, X.; Rajindrajith, S.; Ranawaka, U.; Fan Ng, T.F.; Bucardo-Rivera, F.; Orlandi, P.; Ahmed, K.; Delwart, E. Small circular single stranded DNA viral genomes in unexplained cases of human encephalitis, diarrhea, and in untreated sewage. Virology 2015, 482, 98–104. [Google Scholar] [CrossRef]
- Kraberger, S.; Arguello-Astorga, G.R.; Greenfield, L.G.; Galilee, C.; Law, D.; Martin, D.P.; Varsani, A. Characterisation of a diverse range of circular replication-associated protein encoding DNA viruses recovered from a sewage treatment oxidation pond. Infect. Genet. Evol. 2015, 31, 73–86. [Google Scholar] [CrossRef]
- Pearson, V.M.; Caudle, S.B.; Rokyta, D.R. Viral recombination blurs taxonomic lines: Examination of single-stranded DNA viruses in a wastewater treatment plant. Peer J. 2016, 4, e2585. [Google Scholar] [CrossRef]
- Labonte, J.M.; Suttle, C.A. Previously unknown and highly divergent ssDNA viruses populate the oceans. ISME J. 2013, 7, 2169–2177. [Google Scholar] [CrossRef] [PubMed]
- Dayaram, A.; Galatowitsch, M.L.; Arguello-Astorga, G.R.; van Bysterveldt, K.; Kraberger, S.; Stainton, D.; Harding, J.S.; Roumagnac, P.; Martin, D.P.; Lefeuvre, P.; et al. Diverse circular replication-associated protein encoding viruses circulating in invertebrates within a lake ecosystem. Infect. Genet. Evol. 2016, 39, 304–316. [Google Scholar] [CrossRef] [PubMed]
- Geoghegan, J.L.; Pirotta, V.; Harvey, E.; Smith, A.; Buchmann, J.P.; Ostrowski, M.; Eden, J.S.; Harcourt, R.; Holmes, E.C. Virological Sampling of Inaccessible Wildlife with Drones. Viruses 2018, 10, 300. [Google Scholar] [CrossRef]
- Morandini, V.; Dugger, K.M.; Ballard, G.; Elrod, M.; Schmidt, A.; Ruoppolo, V.; Lescroel, A.; Jongsomjit, D.; Massaro, M.; Pennycook, J.; et al. Identification of a Novel Adelie Penguin Circovirus at Cape Crozier (Ross Island, Antarctica). Viruses 2019, 11, 1088. [Google Scholar] [CrossRef] [PubMed]
- Rosario, K.; Duffy, S.; Breitbart, M. A field guide to eukaryotic circular single-stranded DNA viruses: Insights gained from metagenomics. Arch. Virol. 2012, 157, 1851–1871. [Google Scholar] [CrossRef] [PubMed]
- Orton, J.P.; Morales, M.; Fontenele, R.S.; Schmidlin, K.; Kraberger, S.; Leavitt, D.J.; Webster, T.H.; Wilson, M.A.; Kusumi, K.; Dolby, G.A.; et al. Virus Discovery in Desert Tortoise Fecal Samples: Novel Circular Single-Stranded DNA Viruses. Viruses 2020, 12, 143. [Google Scholar] [CrossRef]
- Fontenele, R.S.; Lacorte, C.; Lamas, N.S.; Schmidlin, K.; Varsani, A.; Ribeiro, S.G. Single Stranded DNA Viruses Associated with Capybara Faeces Sampled in Brazil. Viruses 2019, 11, 710. [Google Scholar] [CrossRef]
- Dunlap, D.S.; Ng, T.F.; Rosario, K.; Barbosa, J.G.; Greco, A.M.; Breitbart, M.; Hewson, I. Molecular and microscopic evidence of viruses in marine copepods. Proc. Natl. Acad. Sci. USA 2013, 110, 1375–1380. [Google Scholar] [CrossRef]
- de la Higuera, I.; Kasun, G.W.; Torrance, E.L.; Pratt, A.A.; Maluenda, A.; Colombet, J.; Bisseux, M.; Ravet, V.; Dayaram, A.; Stainton, D.; et al. Unveiling Crucivirus Diversity by Mining Metagenomic Data. Mbio 2020, 11, e01410–e01420. [Google Scholar] [CrossRef]
- Zhang, W.; Olson, N.H.; Baker, T.S.; Faulkner, L.; Agbandje-McKenna, M.; Boulton, M.I.; Davies, J.W.; McKenna, R. Structure of the Maize streak virus geminate particle. Virology 2001, 279, 471–477. [Google Scholar] [CrossRef]
- Diemer, G.S.; Stedman, K.M. A novel virus genome discovered in an extreme environment suggests recombination between unrelated groups of RNA and DNA viruses. Biol. Direct 2012, 7, 13. [Google Scholar] [CrossRef] [PubMed]
- Krupovic, M.; Zhi, N.; Li, J.; Hu, G.; Koonin, E.V.; Wong, S.; Shevchenko, S.; Zhao, K.; Young, N.S. Multiple layers of chimerism in a single-stranded DNA virus discovered by deep sequencing. Genome Biol. Evol. 2015, 7, 993–1001. [Google Scholar] [CrossRef] [PubMed]
- Roux, S.; Enault, F.; Bronner, G.; Vaulot, D.; Forterre, P.; Krupovic, M. Chimeric viruses blur the borders between the major groups of eukaryotic single-stranded DNA viruses. Nat. Commun. 2013, 4, 2700. [Google Scholar] [CrossRef] [PubMed]
- Jackson, E.W.; Bistolas, K.S.; Button, J.B.; Hewson, I. Novel Circular Single-Stranded DNA Viruses among an Asteroid, Echinoid and Holothurian (Phylum: Echinodermata). PLoS ONE 2016, 11, e0166093. [Google Scholar] [CrossRef] [PubMed]
- Barbosa, A.; Palacios, M.J. Health of Antarctic birds: A review of their parasites, pathogens and diseases. Polar Biol. 2009, 32, 1095. [Google Scholar] [CrossRef] [PubMed]
- Clarke, J.; Kerry, K. Diseases and parasites of penguins. Penguin Conserv. 2000, 13, 5–24. [Google Scholar]
- Woods, R.; Jones, H.I.; Watts, J.; Miller, G.D.; Shellam, G.R. Diseases of Antarctic Seabirds. In Health of Antarctic Wildlife: A Challenge for Science and Policy; Kerry, K.R., Riddle, M., Eds.; Springer: London, UK, 2009. [Google Scholar]
- Grimaldi, W.W.; Seddon, P.J.; Lyver, P.O.; Nakagawa, S.; Tompkins, D.M. Infectious diseases of Antarctic penguins: Current status and future threats. Polar Biol. 2015, 38, 591–606. [Google Scholar] [CrossRef]
- Olivares, F.; Tapia, R.; Galvez, C.; Meza, F.; Barriga, G.P.; Borras-Chavez, R.; Mena-Vasquez, J.; Medina, R.A.; Neira, V. Novel penguin Avian avulaviruses 17, 18 and 19 are widely distributed in the Antarctic Peninsula. Transbound Emerg. Dis. 2019, 66, 2227–2232. [Google Scholar] [CrossRef]
- Lee, S.Y.; Kim, J.H.; Park, Y.M.; Shin, O.S.; Kim, H.; Choi, H.G.; Song, J.W. A novel adenovirus in Chinstrap penguins (Pygoscelis antarctica) in Antarctica. Viruses 2014, 6, 2052–2061. [Google Scholar] [CrossRef]
- Lee, S.Y.; Kim, J.H.; Seo, T.K.; No, J.S.; Kim, H.; Kim, W.K.; Choi, H.G.; Kang, S.H.; Song, J.W. Genetic and Molecular Epidemiological Characterization of a Novel Adenovirus in Antarctic Penguins Collected between 2008 and 2013. PLoS ONE 2016, 11, e0157032. [Google Scholar] [CrossRef]
- Van Doorslaer, K.; Ruoppolo, V.; Schmidt, A.; Lescroel, A.; Jongsomjit, D.; Elrod, M.; Kraberger, S.; Stainton, D.; Dugger, K.M.; Ballard, G.; et al. Unique genome organization of non-mammalian papillomaviruses provides insights into the evolution of viral early proteins. Virus Evol. 2017, 3, vex027. [Google Scholar] [CrossRef] [PubMed]
- Varsani, A.; Kraberger, S.; Jennings, S.; Porzig, E.L.; Julian, L.; Massaro, M.; Pollard, A.; Ballard, G.; Ainley, D.G. A novel papillomavirus in Adelie penguin (Pygoscelis adeliae) faeces sampled at the Cape Crozier colony, Antarctica. J. Gen. Virol. 2014, 95, 1352–1365. [Google Scholar] [CrossRef] [PubMed]
- Varsani, A.; Porzig, E.L.; Jennings, S.; Kraberger, S.; Farkas, K.; Julian, L.; Massaro, M.; Ballard, G.; Ainley, D.G. Identification of an avian polyomavirus associated with Adelie penguins (Pygoscelis adeliae). J. Gen. Virol. 2015, 96, 851–857. [Google Scholar] [CrossRef]
- Wille, M.; Aban, M.; Wang, J.; Moore, N.; Shan, S.; Marshall, J.; Gonzalez-Acuna, D.; Vijaykrishna, D.; Butler, J.; Wang, J.; et al. Antarctic Penguins as Reservoirs of Diversity for Avian Avulaviruses. J. Virol. 2019, 93. [Google Scholar] [CrossRef] [PubMed]
- Neira, V.; Tapia, R.; Verdugo, C.; Barriga, G.; Mor, S.; Ng, T.F.F.; Garcia, V.; Del Rio, J.; Rodrigues, P.; Briceno, C.; et al. Novel Avulaviruses in Penguins, Antarctica. Emerg. Infect. Dis. 2017, 23, 1212–1214. [Google Scholar] [CrossRef]
| Penguin Species | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Adélie Penguin | Chinstrap Penguin | Gentoo Penguin | King Penguin | |||||||||
| Site, Region | Kinnes Cove, NEAP | Booth Is., CWAP | Half Moon Is., SH | Baily Head, SH | Georges Point, CWAP | Booth Is., CWAP | Yankee Harbor, SH | Georges Point, CWAP | Booth Is., CWAP | Moot Point, CWAP | St. Andrew’s Bay, SG | |
| Viral Molecule | GenBank Accession(s) | n = 14 Adults = 7 Chicks = 7 | n = 3 Adults = 2 Chicks = 1 | n = 10 Adults=10 Chicks = 0 | n = 10 Adults = 5 Chicks = 5 | n = 6 Adults = 3 Chicks = 3 | n = 6 Adults = 3 Chicks = 3 | n = 9 Adults = 6 Chicks = 3 | n = 6 Adults = 3 Chicks = 3 | n = 6 Adults = 3 Chicks = 3 | n = 5 Adults = 4 Chicks = 1 | n = 20 Adults = 10 Chicks = 10 |
| AntCM_1_I | MT196226 | 1/0 | ||||||||||
| AntCM_2_I | MT196251 | 1/0 | ||||||||||
| AntCM_3_I | MT196224, MT196272–MT196276 | 4/0 | 1/0 | |||||||||
| AntCM_3_II | MT196225 | 1/0 | ||||||||||
| AntCM_3_III | MT196275, MT196277–MT196278 | 3/0 | ||||||||||
| AntCM_4_I | MT196227–MT196229, MT196262–MT196266 | 5/0 | 3/0 | |||||||||
| AntCM_5_I | MT196230–MT196233, MT196254–MT196260 | 7/0 | 4/0 | |||||||||
| AntCM_6_I | MT196235–MT196244 | 5/5 | ||||||||||
| AntCM_7_I | MT196280–MT196282 | 3/0 | ||||||||||
| AntCM_8_I | MT196234 | 1/0 | ||||||||||
| AntCM_8_II | MT196245–MT196246, MT196288 | 2*/0 | 1/0 | |||||||||
| AntCM_9_I | MT196284–MT196285 | 2/0 | ||||||||||
| AntCM_10_I | MT196286–MT196287 | 2/0 | ||||||||||
| AntCM_11_I | MT196283 | 1/0 | ||||||||||
| AntCM_12_I | MT196267–MT196271 | 5/0 | ||||||||||
| AntV_1_I | MT196247–MT196248 | 2/0 | ||||||||||
| AntV_1_II | MT196249–MT196250 | 1/1 | ||||||||||
| AntV_2_I | MT196289–MT196293 | 4/1 | ||||||||||
| AntV_3_I | MT196252 | 1/0 | ||||||||||
| AntV_3_II | MT196253 MT196261 | 1*/0 1*/0 | ||||||||||
| AntV_3_III | MT196294–MT196295, MT196297–MT196299 | 1/4 | ||||||||||
| AntV_3_IV | MT196296 | 0/1 | ||||||||||
| AntV_3_V | MT196279 | 1/0 | ||||||||||
| AntV_4_I | MT1962 | 5/0 | ||||||||||
| AntV_4_II | MT196301–MT196207 | 1/6 | ||||||||||
| AntV_5_I | MT196312–MT196218 | 5/2 | ||||||||||
| AntV_6_I | MT196222–MT196223 | 1/1 | ||||||||||
| Virus/ Circular Molecule | Top BLASTn Hit Accession | Source | Query Cover | E-Value | Identity |
|---|---|---|---|---|---|
| AntCM1 | - | - | - | - | |
| AntCM2 | - | - | - | - | |
| AntCM3 | MH648984 | Seabass tissue | 86% | 4.00 × 10−167 | 73.86% |
| AntCM4 | JX904537 | Sea water (Saanich Inlet) | 48% | 0 | 79.50% |
| AntCM5 | MH510272 | Seabass tissue | 43% | 4.00 × 10−84 | 69.79% |
| AntCM6 | MH616824 | Abalone tissue | 24% | 2.00 × 10−42 | 72.02% |
| AntCM7 | MH616694 | Abalone tissue | 41% | 3.00 × 10−110 | 72.54% |
| AntCM8 | - | - | - | - | |
| AntCM9 | JX904537 | Sea water (Saanich Inlet) | 87% | 0 | 88.53% |
| AntCM10 | JX904559 | Sea water (Saanich Inlet) | 53% | 1.00 × 10−76 | 68.88% |
| AntCM11 | - | - | - | - | |
| AntCM12 | MH649197 | Red snapper tissue | 71% | 8.00 × 10−170 | 70.75% |
| AntV1 | - | - | - | - | |
| AntV2 | - | - | - | - | |
| AntV3 | - | - | - | - | |
| AntV4 | - | - | - | - | |
| AntV5 | - | - | - | - | |
| AntV6 | - | - | - | - |
| Virus/ Circular Molecule | Genotype | Isolate | Accession # | Motif I | Motif II | Motif III | Walker A | Walker B | Motif C | Arg Finger |
|---|---|---|---|---|---|---|---|---|---|---|
| AntV1 | I | CPGEORsw001Ad | MT196247 | CVTINN | HVQM | WADQEYCFK | GPAGCGKNKLTT | LMHEY | VFTTN | AFWRRI |
| AntV1 | I | CPGEORsw002Ad | MT196248 | CVTINN | HVQM | WADQEYCFK | GPAGCGKNKLTT | LMHEY | VFTTN | AFWRRI |
| AntV1 | II | CPGEORsw003Ad | MT196249 | CVTINN | HVQM | WADQEYCFK | GPAGCGKNKLTT | LMHEY | VFTTN | AFWRRI |
| AntV1 | II | CPGEORsw003Ch | MT196250 | CVTINN | HVQM | WADQEYCFK | GPAGCGKNKLTT | LMHEY | VFTTN | AFWRRI |
| AntV2 | I | GPYANKsw001Ad | MT196289 | SITYNN | HFQC | MAAFKYCQK | GVPGSGKSYQAR | IIEEM | IVTSN | AIKRRF |
| AntV2 | I | GPYANKsw001Ch | MT196290 | SITYNN | HFQC | MAAFKYCQK | GVPGSGKSYQAR | IIEEM | IVTSN | AIKRRF |
| AntV2 | I | GPYANKsw002Ad | MT196291 | SITYNN | HFQC | MAAFKYCQK | GVPGSGKSYQAR | IIEEM | IVTSN | AIKRRF |
| AntV2 | I | GPYANKsw003Ad | MT196292 | SITYNN | HFQC | MAAFKYCQK | GVPGSGKSYQAR | IIEEM | IVTSN | AIKRRF |
| AntV2 | I | GPYANKsw004Ad | MT196293 | SITYNN | HFQC | MAAFKYCQK | GVPGSGKSYQAR | IIEEM | IVTSN | AIKRRF |
| AntV3 | I | CPHALFsw004Ad | MT196252 | CFTLNN | HWQG | QQAIDYCKK | GETGTGKSRKLW | ALEEW | IVTSN | PLKRRF |
| AntV3 | II | CPHALFsw005Ad | MT196253 | CFTLNN | HWQG | TQAIDYCKK | GETGTGKSRKLW | ALEEW | IVTSN | PLKRRF |
| AntV3 | II | CPHALFsw005Ad | MT196261 | CFTLNN | HWQG | TQAIDYCKK | GETGTGKSRKLW | ALEEW | IVTSN | PLKRRF |
| AntV3 | III | KPSTAsw010Ad | MT196294 | CFTLNN | HWQG | QQAIDYCKK | GETGTGKSRKLW | ALEEW | IVTSN | PLKRRF |
| AntV3 | III | KPSTAsw014Ch | MT196295 | CFTLNN | HWQG | QQAIDYCKK | GETGTGKSRKLW | ALEEW | IVTSN | PLKRRF |
| AntV3 | III | KPSTAsw016Ch | MT196297 | CFTLNN | HWQG | QQAIDYCKK | GETGTGKSRKLW | ALEEW | IVTSN | PLKRRF |
| AntV3 | III | KPSTAsw018Ch | MT196298 | CFTLNN | HWQG | QQAIDYCKK | GETGTGKSRKLW | ALEEW | IVTSN | PLKRRF |
| AntV3 | III | KPSTAsw019Ch | MT196299 | CFTLNN | HWQG | QQAIDYCKK | GETGTGKSRKLW | ALEEW | IVTSN | PLKRRF |
| AntV3 | IV | KPSTAsw015Ch | MT196296 | CFTLNN | HWQG | QQAIDYCKK | GETGTGKSRKLW | ALEEW | IVTSN | PLKRRF |
| AntV3 | V | CPHALFsw003Ad | MT196279 | VFTINN | HWQG | KQAIDYCKK | GATGTGKSRKLW | ALEEW | IVTSN | PLKRRF |
| AntV4 | I | KPSTAsw001Ad | MT196300 | CFTINN | HWQG | QQAIDYCKK | GGTGTGKSRKLW | AIEEW | IVTSN | PLKRRF |
| AntV4 | I | KPSTAsw002Ad | MT196308 | CFTINN | HWQG | QQAIDYCKK | GGTGTGKSRKLW | AIEEW | IVTSN | PLKRRF |
| AntV4 | I | KPSTAsw008Ad | MT196309 | CFTINN | HWQG | QQAIDYCKK | GGTGTGKSRKLW | AIEEW | IVTSN | PLKRRF |
| AntV4 | I | KPSTAsw009Ad | MT196310 | CFTINN | HWQG | QQAIDYCKK | GGTGTGKSRKLW | AIEEW | IVTSN | PLKRRF |
| AntV4 | I | KPSTAsw010Ad | MT196311 | CFTINN | HWQG | QQAIDYCKK | GGTGTGKSRKLW | AIEEW | IVTSN | PLKRRF |
| AntV4 | II | KPSTAsw004Ad | MT196301 | CFTINN | HWQG | QQAIDYCKK | GETGTGKSRKLW | AIEEW | IVTSN | PLKRRF |
| AntV4 | II | KPSTAsw014Ch | MT196302 | CFTINN | HWQG | QQAIDYCKK | GETGTGKSRKLW | AIEEW | IVTSN | PLKRRF |
| AntV4 | II | KPSTAsw015Ch | MT196303 | CFTINN | HWQG | QQAIDYCKK | GETGTGKSRKLW | AIEEW | IVTSN | PLKRRF |
| AntV4 | II | KPSTAsw016Ch | MT196304 | CFTINN | HWQG | QQAIDYCKK | GETGTGKSRKLW | AIEEW | IVTSN | PLKRRF |
| AntV4 | II | KPSTAsw017Ch | MT196305 | CFTINN | HWQG | QQAIDYCKK | GETGTGKSRKLW | AIEEW | IVTSN | PLKRRF |
| AntV4 | II | KPSTAsw018Ch | MT196306 | CFTINN | HWQG | QQAIDYCKK | GETGTGKSRKLW | AIEEW | IVTSN | PLKRRF |
| AntV4 | II | KPSTAsw019Ch | MT196307 | CFTINN | HWQG | QQAIDYCKK | GETGTGKSRKLW | AIEEW | IVTSN | PLKRRF |
| AntV5 | I | KPSTAsw001Ad | MT196312 | CFTLNN | HWQG | KQAIDYCKK | GATGTGKSQKLW | AIEEW | IVTSN | PLKRRF |
| AntV5 | I | KPSTAsw002Ad | MT196313 | CFTLNN | HWQG | KQAIDYCKK | GATGTGKSQKLW | AIEEW | IVTSN | PLKRRF |
| AntV5 | I | KPSTAsw004Ad | MT196314 | CFTLNN | HWQG | KQAIDYCKK | GATGTGKSQKLW | AIEEW | IVTSN | PLKRRF |
| AntV5 | I | KPSTAsw008Ad | MT196315 | CFTLNN | HWQG | KQAIDYCKK | GATGTGKSQKLW | AIEEW | IVTSN | PLKRRF |
| AntV5 | I | KPSTAsw009Ad | MT196316 | CFTLNN | HWQG | KQAIDYCKK | GATGTGKSQKLW | AIEEW | IVTSN | PLKRRF |
| AntV5 | I | KPSTAsw018Ch | MT196317 | CFTLNN | HWQG | KQAIDYCKK | GATGTGKSQKLW | AIEEW | IVTSN | PLKRRF |
| AntV5 | I | KPSTAsw020Ch | MT196318 | CFTLNN | HWQG | KQAIDYCKK | GATGTGKSQKLW | AIEEW | IVTSN | PLKRRF |
| AntV6 | I | APBOOTsw001Ad | MT196222 | VFTWNN | HLQG | QECDKYCRK | GESGCGKTRAVH | LLDDI | IVTSQ | ALLRRF |
| AntV6 | I | APBOOTsw001Ch | MT196223 | VFTWNN | HLQG | QECDKYCRK | GESGCGKTRAVH | LLDDI | IVTSQ | ALLRRF |
| AntCM1 | I | CPBAILsw002Ad | MT196226 | IVTLPA | HWQI | QQAYEYVTK | GPTRVGKSRNVV | VLDEF | WVVSN | AFEARF |
| AntCM2 | I | CPHALFsw001Ad | MT196251 | LLTLPA | HWQL | QQVYDYVTK | GPTGVGKTRHIY | VLDEF | WIVSN | AFRRRL |
| AntCM8 | I | CPBAILsw001Ad | MT196234 | FLTISA | HYHA | AKSIKYLKK | GAAGGGKSTLAR | VYDEF | IIVSN | AFKRRI |
| AntCM8 | II | CPBOOTsw002Ad | MT196245 | FITISG | HYHI | HDVLKYIQK | GVPGGGKSTLAR | IFDEF | IIASN | AFKRRI |
| AntCM8 | II | CPBOOTsw002Ad | MT196246 | FITISG | HYHI | HDVLKYIQK | GVPGGGKSTLAR | IFDEF | IIASN | AFKRRI |
| AntCM8 | II | GPBOOTsw003Ad | MT196288 | FITISG | HYHI | HDVLKYIQK | GVPGGGKSTLAR | IFDEF | IIASN | AFKRRI |
| AntCM10 | I | CPHALFsw008Ad | MT196286 | MFTINN | HYQG | KQAVDYVSK | GKPGTGKSHQAR | ILDDL | VVTSN | AINRRF |
| AntCM10 | I | CPHALFsw010Ad | MT196287 | MFTINN | HYQG | KQAVDYVSK | GKPGTGKSHQVR | ILDDL | VVTSN | AINRRF |
| AntCM11 | I | CPHALFsw005Ad | MT196283 | - | - | - | DYEGNKGKSWLSR | IIDVP | MVITN | - |
| AntCM12 | I | CPHALFsw003Ad | MT196267 | FITIPQ | HYHI | NAVITYIKK | GKPNSGKSFMFN | CWDIP | IVFAN | LPQGRT |
| AntCM12 | I | CPHALFsw004Ad | MT196268 | FITIPQ | HYHI | PAVITYIKK | GRPNSGKSFMFN | CWDIP | IVFAN | LPKGRT |
| AntCM12 | I | CPHALFsw008Ad | MT196269 | FITIPQ | HYHI | PAVITYIKK | GRPNSGKSFMFN | CWDIP | IVFAN | LPKGRT |
| AntCM12 | I | CPHALFsw009Ad | MT196270 | FITIPQ | HYHI | PAVITYIKK | GRPNSGKSFMFN | CWDIP | IVFAN | LPKGRT |
| AntCM12 | I | CPHALFsw010Ad | MT196271 | FITIPQ | HYHI | NAVITYIKK | GKPNSGKSFMFN | CWDIP | IVFAN | LPKGRT |
© 2020 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Levy, H.; Fontenele, R.S.; Harding, C.; Suazo, C.; Kraberger, S.; Schmidlin, K.; Djurhuus, A.; Black, C.E.; Hart, T.; Smith, A.L.; et al. Identification and Distribution of Novel Cressdnaviruses and Circular Molecules in Four Penguin Species in South Georgia and the Antarctic Peninsula. Viruses 2020, 12, 1029. https://doi.org/10.3390/v12091029
Levy H, Fontenele RS, Harding C, Suazo C, Kraberger S, Schmidlin K, Djurhuus A, Black CE, Hart T, Smith AL, et al. Identification and Distribution of Novel Cressdnaviruses and Circular Molecules in Four Penguin Species in South Georgia and the Antarctic Peninsula. Viruses. 2020; 12(9):1029. https://doi.org/10.3390/v12091029
Chicago/Turabian StyleLevy, Hila, Rafaela S. Fontenele, Ciara Harding, Crystal Suazo, Simona Kraberger, Kara Schmidlin, Anni Djurhuus, Caitlin E. Black, Tom Hart, Adrian L. Smith, and et al. 2020. "Identification and Distribution of Novel Cressdnaviruses and Circular Molecules in Four Penguin Species in South Georgia and the Antarctic Peninsula" Viruses 12, no. 9: 1029. https://doi.org/10.3390/v12091029
APA StyleLevy, H., Fontenele, R. S., Harding, C., Suazo, C., Kraberger, S., Schmidlin, K., Djurhuus, A., Black, C. E., Hart, T., Smith, A. L., & Varsani, A. (2020). Identification and Distribution of Novel Cressdnaviruses and Circular Molecules in Four Penguin Species in South Georgia and the Antarctic Peninsula. Viruses, 12(9), 1029. https://doi.org/10.3390/v12091029







