Distribution and Genetic Variability of Sapoviruses in Africa
Abstract
1. Introduction
2. Literature
2.1. Search and Selection Strategy
2.2. Data Extraction and Analysis
3. Sapovirus in African Countries
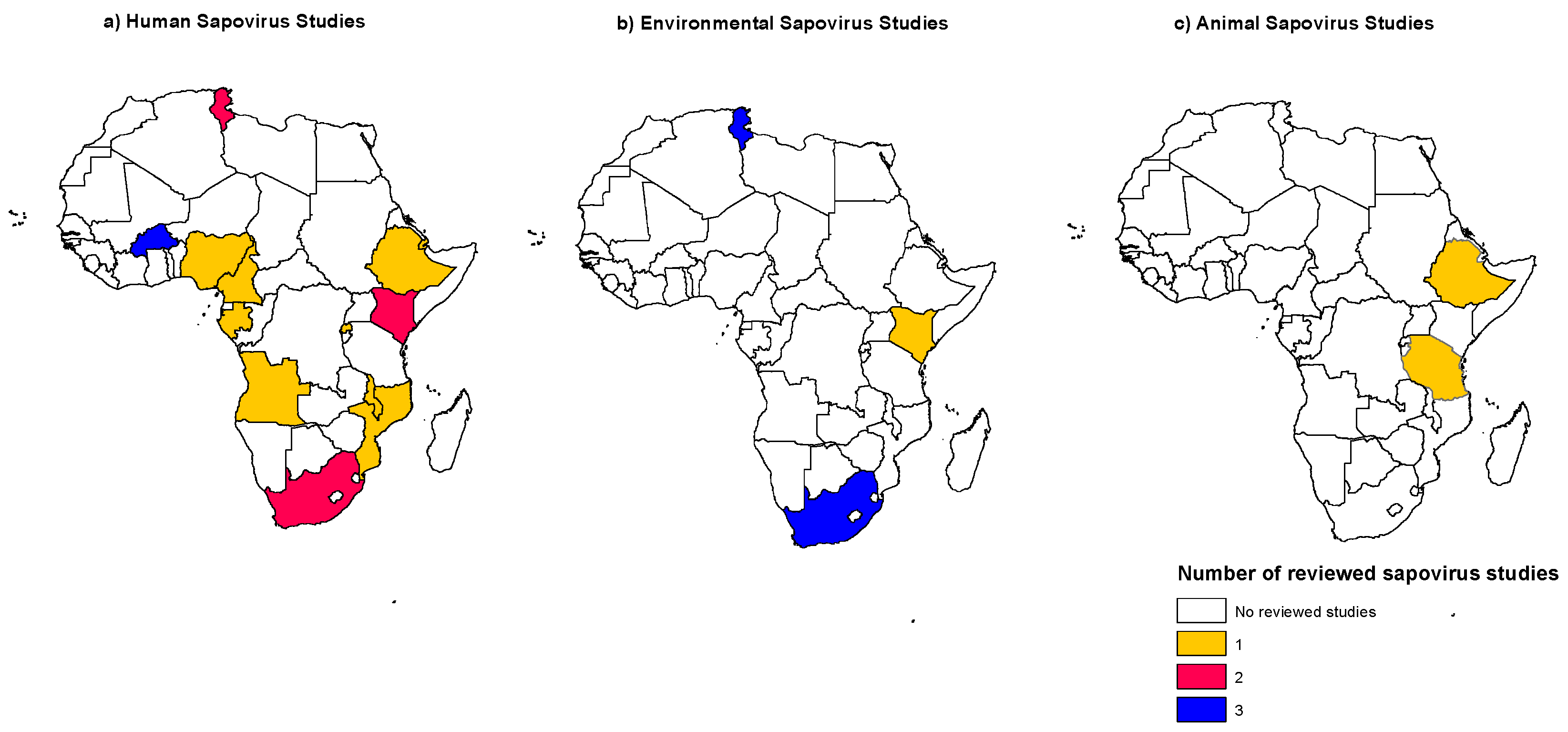
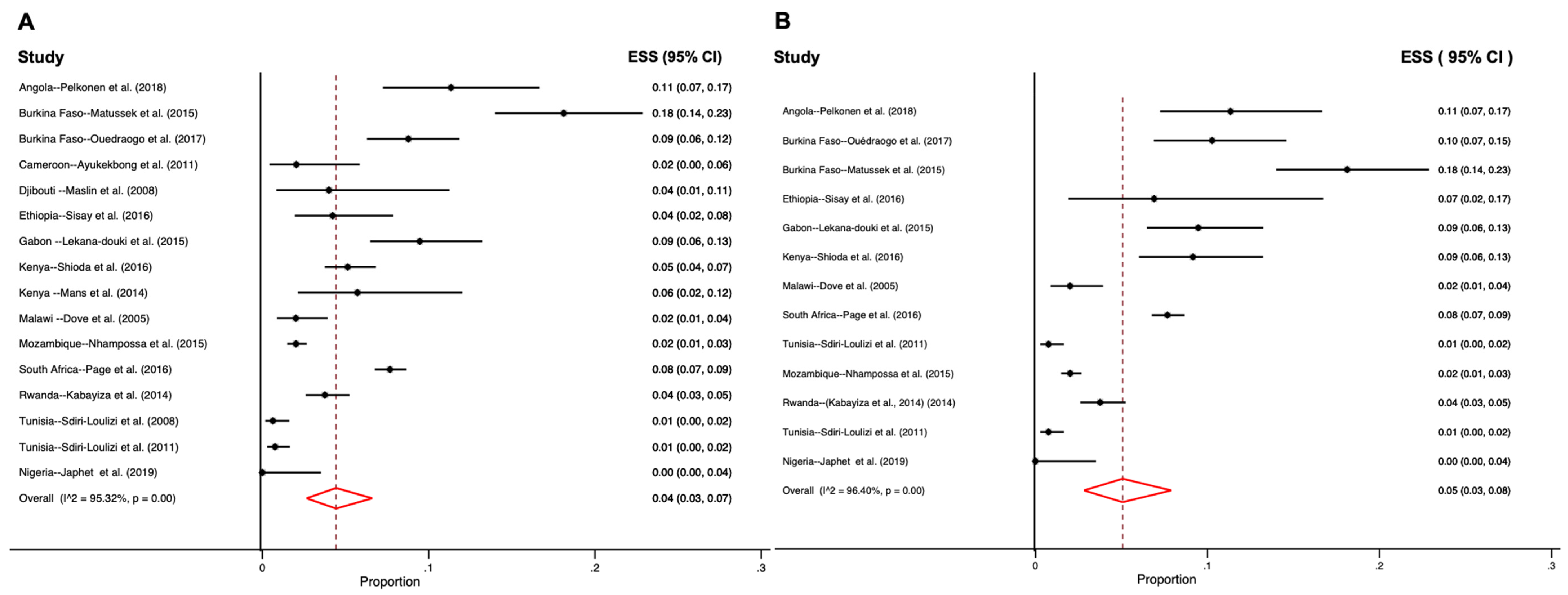
3.1. Distribution of Human Sapovirus Infections
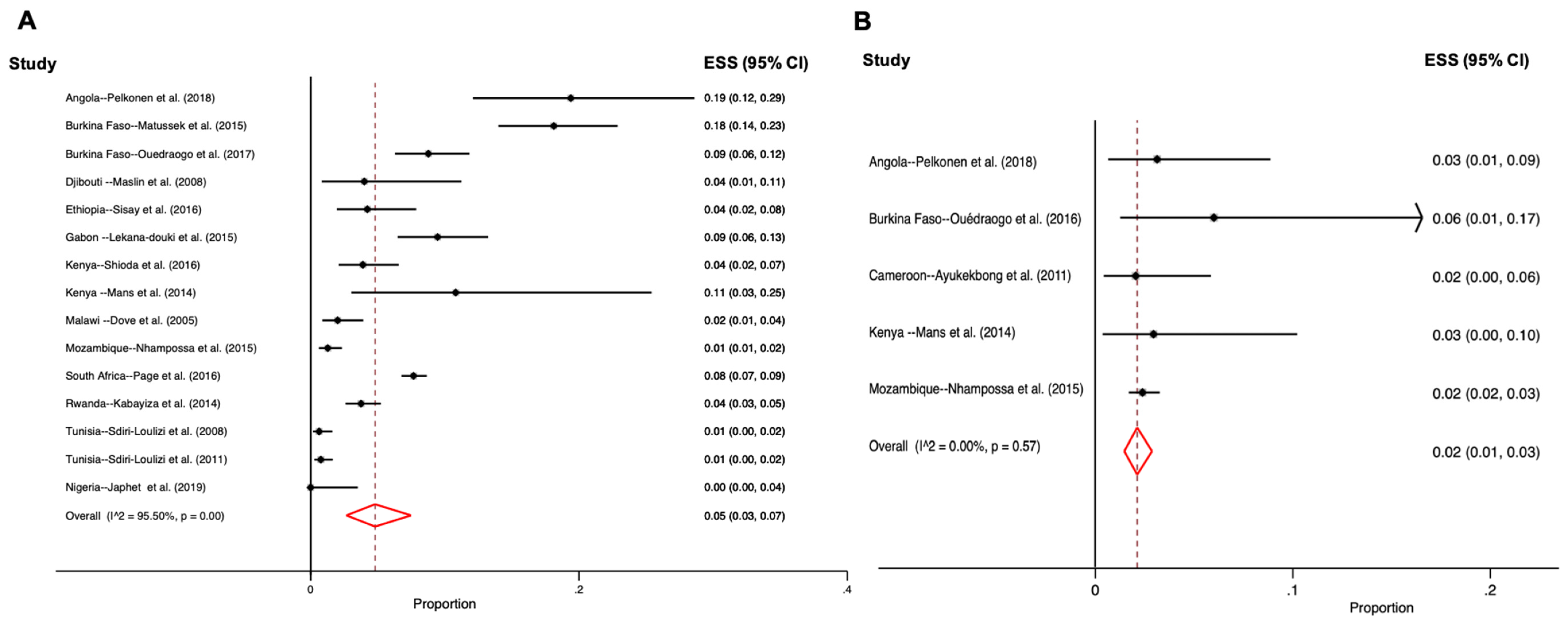
3.2. Age and Sapovirus Distribution
3.3. Sapovirus and Coinfections
3.4. Seasonality of Human Sapovirus Infections
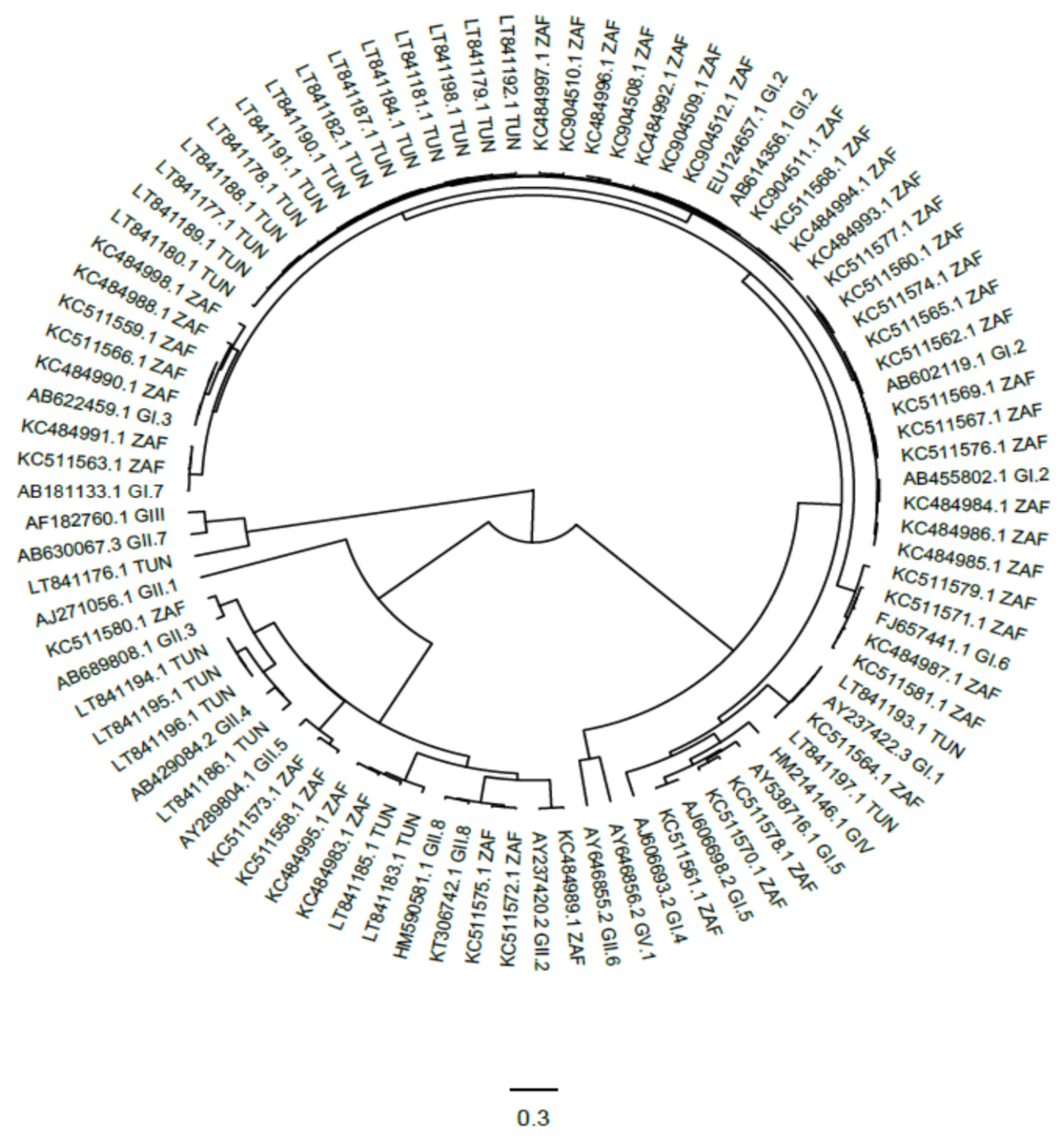
3.5. Genetic Characterization of Human Sapoviruses
3.6. Sapovirus Distribution in the Environment
3.7. Animal Sapovirus Infections
4. Discussion
Supplementary Materials
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
References
- Lauritsen, K.T.; Hansen, M.S.; Johnsen, C.K.; Jungersen, G.; Bottiger, B. Repeated examination of natural sapovirus infections in pig litters raised under experimental conditions. Acta Vet. Scand. 2015, 57, 60. [Google Scholar] [CrossRef] [PubMed]
- Liu, X.; Jahuira, H.; Gilman, R.H.; Alva, A.; Cabrera, L.; Okamoto, M.; Xu, H.; Windle, H.J.; Kelleher, D.; Varela, M.; et al. Etiological Role and Repeated Infections of Sapovirus among Children Aged Less than 2 Years in a Cohort Study in a Peri-urban Community of Peru. J. Clin. Microbiol. 2016, 54, 1598–1604. [Google Scholar] [CrossRef]
- Oka, T.; Wang, Q.; Katayama, K.; Saif, L.J. Comprehensive review of human sapoviruses. Clin. Microbiol. Rev. 2015, 28, 32–53. [Google Scholar] [CrossRef] [PubMed]
- Oka, T.; Lu, Z.; Phan, T.; Delwart, E.L.; Saif, L.J.; Wang, Q. Genetic Characterization and Classification of Human and Animal Sapoviruses. PLoS ONE 2016, 11, e0156373. [Google Scholar] [CrossRef]
- Li, J.; Shen, Q.; Zhang, W.; Zhao, T.; Li, Y.; Jiang, J.; Yu, X.; Guo, Z.; Cui, L.; Hua, X. Genomic organization and recombination analysis of a porcine sapovirus identified from a piglet with diarrhea in China. Virol. J. 2017, 14, 57. [Google Scholar] [CrossRef] [PubMed]
- Vinje, J.; Estes, M.K.; Esteves, P.; Green, K.Y.; Katayama, K.; Knowles, N.J.; L’Homme, Y.; Martella, V.; Vennema, H.; White, P.A.; et al. ICTV Virus Taxonomy Profile: Caliciviridae. J. Gen. Virol. 2019, 100, 1469–1470. [Google Scholar] [CrossRef]
- Hansman, G.S.; Oka, T.; Sakon, N.; Takeda, N. Antigenic diversity of human sapoviruses. Emerg. Infect Dis. 2007, 13, 1519–1525. [Google Scholar] [CrossRef] [PubMed]
- Liu, X.; Yamamoto, D.; Saito, M.; Imagawa, T.; Ablola, A.; Tandoc, A.O., 3rd; Segubre-Mercado, E.; Lupisan, S.P.; Okamoto, M.; Furuse, Y.; et al. Molecular detection and characterization of sapovirus in hospitalized children with acute gastroenteritis in the Philippines. J. Clin. Virol. 2015, 68, 83–88. [Google Scholar] [CrossRef]
- Harada, S.; Tokuoka, E.; Kiyota, N.; Katayama, K.; Oka, T. Phylogenetic analysis of the nonstructural and structural protein encoding region sequences, indicating successive appearance of genomically diverse sapovirus strains from gastroenteritis patients. Jpn. J. Infect. Dis. 2013, 66, 454–457. [Google Scholar] [CrossRef]
- Varela, M.F.; Rivadulla, E.; Lema, A.; Romalde, J.L. Human Sapovirus among Outpatients with Acute Gastroenteritis in Spain: A One-Year Study. Viruses 2019, 11, 144. [Google Scholar] [CrossRef]
- Yinda, C.K.; Conceicao-Neto, N.; Zeller, M.; Heylen, E.; Maes, P.; Ghogomu, S.M.; Van Ranst, M.; Matthijnssens, J. Novel highly divergent sapoviruses detected by metagenomics analysis in straw-colored fruit bats in Cameroon. Emerg. Microbes Infect. 2017, 6, e38. [Google Scholar] [CrossRef]
- Diez-Valcarce, M.; Castro, C.J.; Marine, R.L.; Halasa, N.; Mayta, H.; Saito, M.; Tsaknaridis, L.; Pan, C.Y.; Bucardo, F.; Becker-Dreps, S.; et al. Genetic diversity of human sapovirus across the Americas. J. Clin. Virol. 2018, 104, 65–72. [Google Scholar] [CrossRef]
- Katayama, K.; Miyoshi, T.; Uchino, K.; Oka, T.; Tanaka, T.; Takeda, N.; Hansman, G.S. Novel recombinant sapovirus. Emerg. Infect Dis. 2004, 10, 1874–1876. [Google Scholar] [CrossRef]
- Kuroda, M.; Masuda, T.; Ito, M.; Naoi, Y.; Doan, Y.H.; Haga, K.; Tsuchiaka, S.; Kishimoto, M.; Sano, K.; Omatsu, T.; et al. Genetic diversity and intergenogroup recombination events of sapoviruses detected from feces of pigs in Japan. Infect Genet. Evol. 2017, 55, 209–217. [Google Scholar] [CrossRef]
- Dey, S.K.; Sumiya, M.K.; Shaha, M.; Haque, R.; Okitsu, S.; Ushijima, H. Intragenogroup Recombination in the Complete Genome Sequence of Human Sapovirus Circulating in Bangladesh. Genome Announc. 2018, 6, e00388-18. [Google Scholar] [CrossRef] [PubMed]
- Wang, Q.H.; Han, M.G.; Funk, J.A.; Bowman, G.; Janies, D.A.; Saif, L.J. Genetic diversity and recombination of porcine sapoviruses. J. Clin. Microbiol. 2005, 43, 5963–5972. [Google Scholar] [CrossRef] [PubMed]
- Dos Anjos, K.; Lima, L.M.; Silva, P.A.; Inoue-Nagata, A.K.; Nagata, T. The possible molecular evolution of sapoviruses by inter- and intra-genogroup recombination. Arch. Virol. 2011, 156, 1953–1959. [Google Scholar] [CrossRef] [PubMed]
- Choi, H.L.; Suh, C.I.; Park, S.W.; Jin, J.Y.; Cho, H.G.; Paik, S.Y. Whole-Genome Sequencing Analysis of Sapovirus Detected in South Korea. PLoS ONE 2015, 10, e0132328. [Google Scholar] [CrossRef]
- Pongsuwanna, Y.; Tacharoenmuang, R.; Prapanpoj, M.; Sakon, N.; Komoto, S.; Guntapong, R.; Taniguchi, K. Monthly Distribution of Norovirus and Sapovirus Causing Viral Gastroenteritis in Thailand. Jpn. J. Infect. Dis. 2017, 70, 84–86. [Google Scholar] [CrossRef]
- Torner, N.; Martinez, A.; Broner, S.; Moreno, A.; Camps, N.; Dominguez, A. Working Group for the Study of Acute Viral Gastroenteritis Outbreaks in, C. Epidemiology of Acute Gastroenteritis Outbreaks Caused by Human Calicivirus (Norovirus and Sapovirus) in Catalonia: A Two Year Prospective Study, 2010-2011. PLoS ONE 2016, 11, e0152503. [Google Scholar] [CrossRef]
- Varela, M.F.; Ouardani, I.; Kato, T.; Kadoya, S.; Aouni, M.; Sano, D.; Romalde, J.L. Sapovirus in Wastewater Treatment Plants in Tunisia: Prevalence, Removal, and Genetic Characterization. Appl. Environ. Microbiol. 2018, 84, 324–331. [Google Scholar] [CrossRef] [PubMed]
- Kitajima, M.; Oka, T.; Haramoto, E.; Katayama, H.; Takeda, N.; Katayama, K.; Ohgaki, S. Detection and genetic analysis of human sapoviruses in river water in Japan. Appl. Environ. Microbiol. 2010, 76, 2461–2467. [Google Scholar] [CrossRef] [PubMed]
- Kebede Fufa, W.; Berhe Gebremedhin, G.; Gebregergs, G.B.; Marama Mokonnon, T. Assessment of Poor Home Management Practice of Diarrhea and Associated Factors among Caregivers of Under-Five Years Children in Urban and Rural Residents of Doba Woreda, Ethiopia: Comparative Cross-Sectional Study. Int. J. Pediatr. 2019, e8345245. [Google Scholar] [CrossRef] [PubMed]
- Groome, M.J.; Madhi, S.A. Five-year cohort study on the burden of hospitalisation for acute diarrhoeal disease in African HIV-infected and HIV-uninfected children: Potential benefits of rotavirus vaccine. Vaccine 2012, 30, A173–A178. [Google Scholar] [CrossRef]
- Cunliffe, N.A.; Gondwe, J.S.; Kirkwood, C.D.; Graham, S.M.; Nhlane, N.M.; Thindwa, B.D.; Dove, W.; Broadhead, R.L.; Molyneux, M.E.; Hart, C.A. Effect of concomitant HIV infection on presentation and outcome of rotavirus gastroenteritis in Malawian children. Lancet 2001, 358, 550–555. [Google Scholar] [CrossRef]
- Rio, C.d. The Global HIV epidemic: What the pathologist needs to know. PMC 2018, 34, 314–317. [Google Scholar] [CrossRef]
- Pietsch, C.; Liebert, U.G. Intrahost viral evolution during chronic sapovirus infections. J. Clin. Virol. 2019, 113, 1–7. [Google Scholar] [CrossRef]
- Guarino, A.; Dupont, C.; Gorelov, A.V.; Gottrand, F.; Lee, J.K.; Lin, Z.; Lo Vecchio, A.; Nguyen, T.D.; Salazar-Lindo, E. The management of acute diarrhea in children in developed and developing areas: From evidence base to clinical practice. Expert Opin. Pharm. 2012, 13, 17–26. [Google Scholar] [CrossRef]
- Bucardo, F.; Reyes, Y.; Svensson, L.; Nordgren, J. Predominance of Norovirus and Sapovirus in Nicaragua after Implementation of Universal Rotavirus Vaccination. PLoS ONE 2014, 9, e98201. [Google Scholar] [CrossRef]
- Mourad Ouzzani, H.H.Z.F.; Ahmed, E. Rayyan a web and mobile app for systematic reviews. Syst. Rev. 2016, 5, 210. [Google Scholar] [CrossRef]
- Drummond, A.J.; Suchard, M.A.; Xie, D.; Rambaut, A. Bayesian phylogenetics with BEAUti and the BEAST 1.7. Mol. Biol. Evol. 2012, 29, 1969–1973. [Google Scholar] [CrossRef] [PubMed]
- Kumar, S.; Stecher, G.; Tamura, K. MEGA7: Molecular Evolutionary Genetics Analysis Version 7.0 for Bigger Datasets. Mol. Biol. Evol. 2016, 33, 1870–1874. [Google Scholar] [CrossRef] [PubMed]
- Ayukekbong, J.; Lindh, M.; Nenonen, N.; Tah, F.; Nkuo-akenji, T.; Bergstro, T. Enteric Viruses in Healthy Children in Cameroon: Viral Load and Genotyping of Norovirus Strains. J. Med. Virol. 2011, 83, 2135–2142. [Google Scholar] [CrossRef] [PubMed]
- Pelkonen, T.; Dos Santos, M.D.; Roine, I.; Dos Anjos, E.; Freitas, C.; Peltola, H.; Laakso, S.; Kirveskari, J. Potential Diarrheal Pathogens Common Also in Healthy Children in Angola. Pediatr. Infect Dis. J. 2018, 37, 424–428. [Google Scholar] [CrossRef] [PubMed]
- Dove, W.; Cunliffe, N.A.; Gondwe, J.S.; Broadhead, R.L.; Molyneux, M.E.; Nakagomi, O.; Hart, C.A. Detection and characterization of human caliciviruses in hospitalized children with acute gastroenteritis in Blantyre, Malawi. J. Med. Virol. 2005, 77, 522–527. [Google Scholar] [CrossRef]
- Nhampossa, T.; Mandomando, I.; Acacio, S.; Quinto, L.; Vubil, D.; Ruiz, J.; Nhalungo, D.; Sacoor, C.; Nhabanga, A.; Nhacolo, A.; et al. Diarrheal Disease in Rural Mozambique: Burden, Risk Factors and Etiology of Diarrheal Disease among Children Aged 0-59 Months Seeking Care at Health Facilities. PLoS ONE 2015, 10, e0119824. [Google Scholar] [CrossRef]
- Lekana-Douki, S.E.; Kombila-Koumavor, C.; Nkoghe, D.; Drosten, C.; Drexler, J.F.; Leroy, E.M. Molecular epidemiology of enteric viruses and genotyping of rotavirus A, adenovirus and astrovirus among children under 5 years old in Gabon. Int. J. Infect Dis. 2015, 34, 90–95. [Google Scholar] [CrossRef]
- Matussek, A.; Dienus, O.; Djeneba, O.; Simpore, J.; Nitiema, L.; Nordgren, J. Molecular characterization and genetic susceptibility of sapovirus in children with diarrhea in Burkina Faso. Infect Genet. Evol. 2015, 32, 396–400. [Google Scholar] [CrossRef]
- Ouedraogo, N.; Ngangas, S.M.; Bonkoungou, I.J.; Tiendrebeogo, A.B.; Traore, K.A.; Sanou, I.; Traore, A.S.; Barro, N. Temporal distribution of gastroenteritis viruses in Ouagadougou, Burkina Faso: Seasonality of rotavirus. BMC Public Health 2017, 17, 274. [Google Scholar] [CrossRef]
- Maslin, J.; Nicand, E.; Ambert-Balay, K.; Fouet, C.; Kaplon, J.; Haus, R.; Pothier, P.; Kohli, E. Detection and characterization of human caliciviruses associated with sporadic acute diarrhea in adults in Djibouti (Horn of Africa). Am. J. Trop. Med. Hyg. 2008, 78, 522–526. [Google Scholar] [CrossRef]
- Sisay, Z.; Djikeng, A.; Berhe, N.; Belay, G.; Gebreyes, W.; Abegaz, W.E.; Njahira, M.N.; Wang, Q.H.; Saif, L.J. Prevalence and molecular characterization of human noroviruses and sapoviruses in Ethiopia. Arch. Virol. 2016, 161, 2169–2182. [Google Scholar] [CrossRef]
- Mans, J.; Murray, T.Y.; Kiulia, N.M.; Mwenda, J.M.; Musoke, R.N.; Taylor, M.B. Human caliciviruses detected in HIV-seropositive children in Kenya. J. Med. Virol. 2014, 86, 75–81. [Google Scholar] [CrossRef] [PubMed]
- Shioda, K.; Cosmas, L.; Audi, A.; Gregoricus, N.; Vinje, J.; Parashar, U.D.; Montgomery, J.M.; Feikin, D.R.; Breiman, R.F.; Hall, A.J. Population-Based Incidence Rates of Diarrheal Disease Associated with Norovirus, Sapovirus, and Astrovirus in Kenya. PLoS ONE 2016, 11, e0145943. [Google Scholar] [CrossRef] [PubMed]
- Page, N.; Groome, M.J.; Murray, T.; Nadan, S.; Netshikweta, R.; Keddy, K.H.; Poonsamy, B.; Moyes, J.; Walaza, S.; Kahn, K.; et al. Sapovirus prevalence in children less than five years of age hospitalised for diarrhoeal disease in South Africa, 2009-2013. J. Clin. Virol. 2016, 78, 82–88. [Google Scholar] [CrossRef] [PubMed]
- Sdiri-Loulizi, K.; Hassine, M.; Gharbi-Khelifi, H.; Aouni, Z.; Chouchane, S.; Sakly, N.; Neji-Guediche, M.; Pothier, P.; Ambert-Balay, K.; Aouni, M. Molecular detection of genogroup I sapovirus in Tunisian children suffering from acute gastroenteritis. Virus Genes 2011, 43, 6–12. [Google Scholar] [CrossRef]
- Kabayiza, J.C.; Andersson, M.E.; Nilsson, S.; Baribwira, C.; Muhirwa, G.; Bergstrom, T.; Lindh, M. Diarrhoeagenic microbes by real-time PCR in Rwandan children under 5 years of age with acute gastroenteritis. Clin. Microbiol. Infect 2014, 20, O1128–O1135. [Google Scholar] [CrossRef]
- Sdiri-Loulizi, K.; Gharbi-Khelifi, H.; de Rougemont, A.; Chouchane, S.; Sakly, N.; Ambert-Balay, K.; Hassine, M.; Guediche, M.N.; Aouni, M.; Pothier, P. Acute infantile gastroenteritis associated with human enteric viruses in Tunisia. J. Clin. Microbiol. 2008, 46, 1349–1355. [Google Scholar] [CrossRef]
- Murray, T.Y.; Nadan, S.; Page, N.A.; Taylor, M.B. Diverse sapovirus genotypes identified in children hospitalised with gastroenteritis in selected regions of South Africa. J. Clin. Virol. 2016, 76, 24–29. [Google Scholar] [CrossRef]
- Ouedraogo, N.; Kaplon, J.; Bonkoungou, I.J.; Traore, A.S.; Pothier, P.; Barro, N.; Ambert-Balay, K. Prevalence and Genetic Diversity of Enteric Viruses in Children with Diarrhea in Ouagadougou, Burkina Faso. PLoS ONE 2016, 11, e0153652. [Google Scholar] [CrossRef]
- Japhet, M.O.; Famurewa, O.; Adesina, O.A.; Opaleye, O.O.; Wang, B.; Hohne, M.; Bock, C.T.; Mas Marques, A.; Niendorf, S. Viral gastroenteritis among children of 0-5 years in Nigeria: Characterization of the first Nigerian aichivirus, recombinant noroviruses and detection of a zoonotic astrovirus. J. Clin. Virol. 2019, 111, 4–11. [Google Scholar] [CrossRef]
- Brazier, L.; Elguero, E.; Koumavor, C.K.; Renaud, N.; Prugnolle, F.; Thomas, F.; Ategbo, S.; Engoba, M.; Obengui; Leroy, E.M.; et al. Evolution in fecal bacterial/viral composition in infants of two central African countries (Gabon and Republic of the Congo) during their first month of life. PLoS ONE 2017, 12, e0185569. [Google Scholar] [CrossRef] [PubMed]
- Sisay, Z.; Djikeng, A.; Berhe, N.; Belay, G.; Abegaz, W.E.; Wang, Q.H.; Saif, L.J. First detection and molecular characterization of sapoviruses and noroviruses with zoonotic potential in swine in Ethiopia. Arch. Virol. 2016, 161, 2739–2747. [Google Scholar] [CrossRef] [PubMed]
- Kiulia, N.M.; Netshikweta, R.; Page, N.A.; Van Zyl, W.B.; Kiraithe, M.M.; Nyachieo, A.; Mwenda, J.M.; Taylor, M.B. The detection of enteric viruses in selected urban and rural river water and sewage in Kenya, with special reference to rotaviruses. J. Appl. Microbiol. 2010, 109, 818–828. [Google Scholar] [CrossRef] [PubMed]
- Murray, T.Y.; Mans, J.; Taylor, M.B. Human calicivirus diversity in wastewater in South Africa. J. Appl. Microbiol. 2013, 114, 1843–1853. [Google Scholar] [CrossRef] [PubMed]
- Murray, T.Y.; Mans, J.; Taylor, M.B. First detection of human sapoviruses in river water in South Africa. Water Sci. Technol. 2013, 67, 2776–2783. [Google Scholar] [CrossRef] [PubMed]
- Murray, T.Y.; Taylor, M.B. Quantification and molecular characterisation of human sapoviruses in water sources impacted by highly polluted discharged wastewater in South Africa. J. Water Health 2015, 13, 1055–1059. [Google Scholar] [CrossRef]
- Sdiri-Loulizi, K.; Hassine, M.; Aouni, Z.; Gharbi-Khelifi, H.; Chouchane, S.; Sakly, N.; Neji-Guediche, M.; Pothier, P.; Aouni, M.; Ambert-Balay, K. Detection and molecular characterization of enteric viruses in environmental samples in Monastir, Tunisia between January 2003 and April 2007. J. Appl. Microbiol. 2010, 109, 1093–1104. [Google Scholar] [CrossRef]
- Ibrahim, C.; Hammami, S.; Cherif, N.; Mejri, S.; Pothier, P.; Hassen, A. Detection of Sapoviruses in two biological lines of Tunisian hospital wastewater treatment. Int. J. Environ. Health Res. 2019, 29, 400–413. [Google Scholar] [CrossRef]
- Olarte-Castillo, X.A.; Hofer, H.; Goller, K.V.; Martella, V.; Moehlman, P.D.; East, M.L. Divergent Sapovirus Strains and Infection Prevalence in Wild Carnivores in the Serengeti Ecosystem: A Long-Term Study. PLoS ONE 2016, 11, e0163548. [Google Scholar] [CrossRef]
- Sanchez, G.J.; Mayta, H.; Pajuelo, M.J.; Neira, K.; Xiaofang, L.; Cabrera, L.; Ballard, S.B.; Crabtree, J.E.; Kelleher, D.; Cama, V.; et al. Epidemiology of Sapovirus Infections in a Birth Cohort in Peru. Clin. Infect. Dis. 2018, 66, 1858–1863. [Google Scholar] [CrossRef]
- Hansman, G.S.; Takeda, N.; Katayama, K.; Tu, E.T.; McIver, C.J.; Rawlinson, W.D.; White, P.A. Genetic diversity of Sapovirus in children, Australia. Emerg. Infect Dis. 2006, 12, 141–143. [Google Scholar] [CrossRef] [PubMed]
- Weldegebriela, G.M.J.M.; Chakauya, J.; Daniel, F.; Masresha, B.; Parasha, D.U.; Tatec, E.J. Impact of rotavirus vaccine on rotavirus diarrhoea in countries of East and Southern Africa. Vaccine 2018, 36, 7124–7130. [Google Scholar] [CrossRef] [PubMed]
- Anya, B.; Okeibunor, J.; Mihigo, R.; Poy, A.; Zawaira, F. Efforts to Reach More Children with Effective Vaccines Through Routine Immunization in The WHO African Region: 2013–2015. J. Immunol. Sci. 2018, 8, 55–62. [Google Scholar] [CrossRef]
- Pang, X.L.; Lee, B.E.; Tyrrell, G.J.; Preiksaitis, J.K. Epidemiology and genotype analysis of sapovirus associated with gastroenteritis outbreaks in Alberta, Canada: 2004-2007. J. Infect. Dis. 2009, 199, 547–551. [Google Scholar] [CrossRef]
- Ayukekbong, J.A.; Andersson, M.E.; Vansarla, G.; Tah, F.; Nkuo-Akenji, T.; Lindh, M.; BergstrÖM, T. Monitoring of seasonality of norovirus and other enteric viruses in Cameroon by real-time PCR: An exploratory study. Epidemiol. Infect. 2013, 142, 1393–1402. [Google Scholar] [CrossRef]
- Sakai, Y.; Nakata, S.; Honma, S.; Tatsumi, M.; Numata-Kinoshita, K.; Chiba, S. Clinical severity of Norwalk virus and Sapporo virus gastroenteritis in children in Hokkaido, Japan. Pediatr. Infect Dis. J. 2001, 20, 849–853. [Google Scholar] [CrossRef]
- Dey, S.K.; Phathammavong, O.; Nguyen, T.D.; Thongprachum, A.; Chan-It, W.; Okitsu, S.; Mizuguchi, M.; Ushijima, H. Seasonal pattern and genotype distribution of sapovirus infection in Japan, 2003–2009. Epidemiol. Infect. 2012, 140, 74–77. [Google Scholar] [CrossRef]
- Parsa-Nahad, M.; Samarbaf-Zadeh, A.R.; Makwandi, M.; Hamid, S.; Mozhgani, R.; Jalilian, S.; Pirmoradi, R.; Shamsi-Zadeh, A. Relative Frequency of Sapovirus Among Children under 5 Years of Age with Acute Gastroenteritis at the Aboozar Hospital, Ahvaz, Iran. Jundishpur J. Microbiol. 2012, 5, 359–361. [Google Scholar] [CrossRef]
- Iwai, M.; Hasegawa, S.; Obara, M.; Nakamura, K.; Horimoto, E.; Takizawa, T.; Kurata, T.; Sogen, S.; Shiraki, K. Continuous presence of noroviruses and sapoviruses in raw sewage reflects infections among inhabitants of Toyama, Japan (2006 to 2008). Appl. Environ. Microbiol. 2009, 75, 1264–1270. [Google Scholar] [CrossRef]
- Farkas, T.; Zhong, W.M.; Jing, Y.; Huang, P.W.; Espinosa, S.M.; Martinez, N.; Morrow, A.L.; Ruiz-Palacios, G.M.; Pickering, L.K.; Jiang, X. Genetic diversity among sapoviruses. Arch. Virol. 2004, 149, 1309–1323. [Google Scholar] [CrossRef]
- Johnson, C.K.; Hitchens, P.L.; Evans, T.S.; Goldstein, T.; Thomas, K.; Clements, A.; Joly, D.O.; Wolfe, N.D.; Daszak, P.; Karesh, W.B.; et al. Spillover and pandemic properties of zoonotic viruses with high host plasticity. Sci. Rep. 2015, 5, 1–8. [Google Scholar] [CrossRef]
| Species | Genogroup | References |
|---|---|---|
| Humans | GI, GII, GIV, GV | [2,4,18,19,20] |
| Dogs | GXIII | [4] |
| Pigs | GIII, GV, GVI, GVII, GVIII, GIX, GX, GXI | [1,3,10] |
| Bats | GXIV, (GXVIII, GXVIX-tentative) | [4,11] |
| Mink | GXII | [4] |
| Chimpanzees | GI | [4] |
| Sea lions | GV | [4] |
| Rats | GII, GXV | [4] |
| Reference | Country | Study Period; Duration | Prevalence | Age (Years) | Sapovirus Detection Method | Sequencing Region and Nucleotide Position | Identified Genotypes (Based on Partial VP1 Sequence) | Identified Genotypes (Based on Partial RdRp Sequence) |
|---|---|---|---|---|---|---|---|---|
| [34] | Angola | Dec 2013 to Aug 2014; 8 months May to Oct 2014; 6 months | All 11% (22/194) Symptomatic 19% (19/98) Asymptomatic 3% (3/96) | <5 | Real-time RT PCR a | n/a | n/a | n/a |
| [49] | Burkina Faso | Nov 2011 to Sep 2012; 10 months | Symptomatic 10.3% (27/263) Asymptomatic 6% (3/50) | <5 | Real-time RT-PCR b | Partial RdRp l nt 4366–4884 * | n/a | GII.1(3) GII.2(2) GIV.1(1) Unassigned (3) |
| [38] | Burkina Faso | May 2009 and Mar 2010; 12 months | Symptomatic 18% (56/309) | <5 | Real-Time RT-PCR c | Partial VP1 h nt 5083–5516 * | GI.1(12) GI.2(6) GI.4(4) GII.1(1) GII.2(2) GIV.1(2) | n/a |
| [39] | Burkina Faso | Nov 2011 to Sep 2012; 10 months | Symptomatic 10.3% (27/263) Symptomatic 6.5% (11/170) | <5 5–89 | Real-Time RT-PCR b | n/a | n/a | n/a |
| [33] | Cameroon | Oct to Dec 2009; 2 months | Asymptomatic 2.04% (3/147) | 5–75 | Real-Time RT-PCR c | n/a | n/a | n/a |
| [40] | Djibouti | Sep 2002 to Feb 2004; 15 months | Symptomatic 4% (3/75) | >15 | RT-PCR d | n/a | n/a | |
| [41] | Ethiopia | Jun to Sep 2013; 3 months | Symptomatic 4.2% (9/213) | All | RT-PCR e | Partial RdRp e nt 4327–4656 ** | n/a | GII.1 |
| [37] | Gabon | Mar 2010 to Jun 2011; 15 months | Symptomatic 9.5% (30/317) | <5yrs | multiplex real-time RT-PCR f | n/a | n/a | n/a |
| [43] | Kenya | Jun 2007 to Oct 2008; 16 months Oct 2006 to Feb 2009; 28 months | Symptomatic 4% (13/334) Lwak District 6% (31/524) Kibera District | All | Real-time RT-qPCR c | n/a | n/a | n/a |
| [42] | Kenya | Feb 1999 to Jun 2000; 16 months | Symptomatic 10.8% (4/37) Asymptomatic 2.9% (2/68) | <14 | Real-Time RT-PCR c | Partial VP1 m nt 5098–5878 * | GI.2(2) GI.6(1) GII.1(1) GII.2(2) GII.4(1) | n/a |
| [35] | Malawi | Jul 1998 to Jun 1999; 12 months | Symptomatic 2% (8/398) | <5 | RT-PCR g | Partial RdRp g nt 4354–4684 * | n/a | GII.1 |
| [36] | Mozambique | Dec 2007 to Oct 2011; 46 months | Symptomatic 1.3% (10/784) Asymptomatic 2.4% (38/1595) | <5 | Multiplex RT-PCR h | n/a | n/a | n/a |
| [50] | Nigeria | Aug 2012 to Dec 2013; 16 months | Symptomatic 0% (0/103) | <5 | Real-time RT-qPCR i | n/a | n/a | n/a |
| [44] | South Africa | Apr 2009 to Dec 2013; 44 months | Symptomatic 7.7% (238/3103) | <5 | Real time RT-PCR j | n/a | n/a | n/a |
| [48] | South Africa | Apr 2009 to Dec 2013; 44 months | n/a | <6 | Real time RT-PCR j | Partial VP1 n nt 5159–5498 * | GI.1(27) GI.2(45) GI.3(8) GI.5(8) GI.6(4) GI.7(2) GII.1(29) GII.2 (7) GII.3(14) GII.4(18) GII.5(7) GII.6(2) GII.7(2) GIV.1(49) | n/a |
| [46] | Rwanda | Nov 2009 to Jun 2012; 30 months | Symptomatic 3.8% (33/879) | <5 | Real-time RT-PCR k | n/a | n/a | n/a |
| [45] | Tunisia | Jan 2003 to Apr 2007; 39 months | Symptomatic 0.8% (6/788) | <2 | RT-PCR l | Partial RdRp l nt 4366–4884 * | n/a | GI.1(1) |
| [47] | Tunisia | Jan 2003 to Jun 2005; 29 months | Symptomatic 1% (4/632) | <12 years | RT-PCR l | n/a | n/a | n/a |
| Ref | Country | Study Period; Duration | Prevalence | Specimen Source | Sapovirus Detection Method | Sequencing Primers and Sequenced Region | Identified Genotypes -Based on Partial VP1 Sequence (# of Samples) | Identified Genotypes -Based on Partial RdRp Sequence (# of Samples) |
|---|---|---|---|---|---|---|---|---|
| [53] | Kenya | May 2007 to Feb 2008; 9 months | 90% (9/10) 14.3% (1/7) | urban and rural river water, sewage | Real-time RT-PCR a | n/a | n/a | n/a |
| [56] | South Africa | n/a | 80% (8/10) | wastewater treatment works and affected wastewater bodies | Real-time RT-qPCR b | partial VP1 e nt 5157–5591 and 5159–5498 * | GI.2(6/6) | n/a |
| [54] | South Africa | Aug 2010 to Dec 2011; 16 months | 73% (37/51) | wastewater | Real-time RT-qPCR b | Partial VP1 f nt 5159–5498 * | GI.2 (8) GI.3 (3) GI.6 (1) GI.7 (1) GII.1 (1) GII.2(1) | n/a |
| [55] | South Africa | Jan 2009 to Dec 2010; 23 months | 41% (21/51) in 2009 58% (27/48) in 2010 | river water | Real-time RT-PCR a | Partial VP1 f nt 5159–5498 * | GI.1(1) G1.2(9) GI.3(2) GI.5(3) GI.7(1) GII.3(1) GII.5(1) GII.8(3) | n/a |
| [21] | Tunisia | Dec 2009 to Dec 2010; 12 months | 39.9% (87/218) All 56% (61/109) Untreated 23.9% (26/109) Treated | wastewater treatment plants | Real-time RT-PCR c | Partial VP1 gnt 5159–5591 * | GI.1(2) GI.2(15) GII.1(2) GII.4(4) GII.8(1) | n/a |
| [57] | Tunisia | Jan 2003 and April 2007 | 0% (0/250) sewage | sewage samples | RT-PCR d | n/a | n/a | n/a |
| [58] | Tunisia | Jan-Dec 2011; 12 months | untreated 1 to 17% treated | Waste water | Real-time RT-PCR c | Partial RdRp d nt 4366–4884 * | n/a | GI.3(1) GIV.1(1) Could not assign (2) |
| Reference | Country | Study Period; Duration | Prevalence | Animal Species | Initial Detection | Sequencing Primers and Sequenced Region | Identified Genotypes (Based on Partial RdRp Sequence) |
|---|---|---|---|---|---|---|---|
| [52] | Ethiopia | Jan–Sep 2013; 9 months | 1.7% (2/117) | swine | RT-PCR a,b | Partial RdRp a nt 4327–4656 * Partial RdRp b nt 4568–4884 ** | GIII (1/2) (1/2) could not be assigned |
| [59] | Tanzania | 2001–2012; 11 years | 34.8% (171/514) 33.3% (3/9) 22.2% (2/9) | Spotted hyenas African lion Bat eared fox | RT-PCR c | partial RdRp d | 20/20 clustered together but could not be assigned (potential novel strains) |
© 2020 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Makhaola, K.; Moyo, S.; Kebaabetswe, L.P. Distribution and Genetic Variability of Sapoviruses in Africa. Viruses 2020, 12, 490. https://doi.org/10.3390/v12050490
Makhaola K, Moyo S, Kebaabetswe LP. Distribution and Genetic Variability of Sapoviruses in Africa. Viruses. 2020; 12(5):490. https://doi.org/10.3390/v12050490
Chicago/Turabian StyleMakhaola, Kgomotso, Sikhulile Moyo, and Lemme P. Kebaabetswe. 2020. "Distribution and Genetic Variability of Sapoviruses in Africa" Viruses 12, no. 5: 490. https://doi.org/10.3390/v12050490
APA StyleMakhaola, K., Moyo, S., & Kebaabetswe, L. P. (2020). Distribution and Genetic Variability of Sapoviruses in Africa. Viruses, 12(5), 490. https://doi.org/10.3390/v12050490





