Indigenous versus Lessepsian Hosts: Nervous Necrosis Virus (NNV) in Eastern Mediterranean Sea Fish
Abstract
1. Introduction
2. Materials and Methods
2.1. Fish and Tissue Sampling
2.2. RNA extraction and cDNA synthesis by RT-PCR
2.3. RNA1 and RNA2 PCR amplification
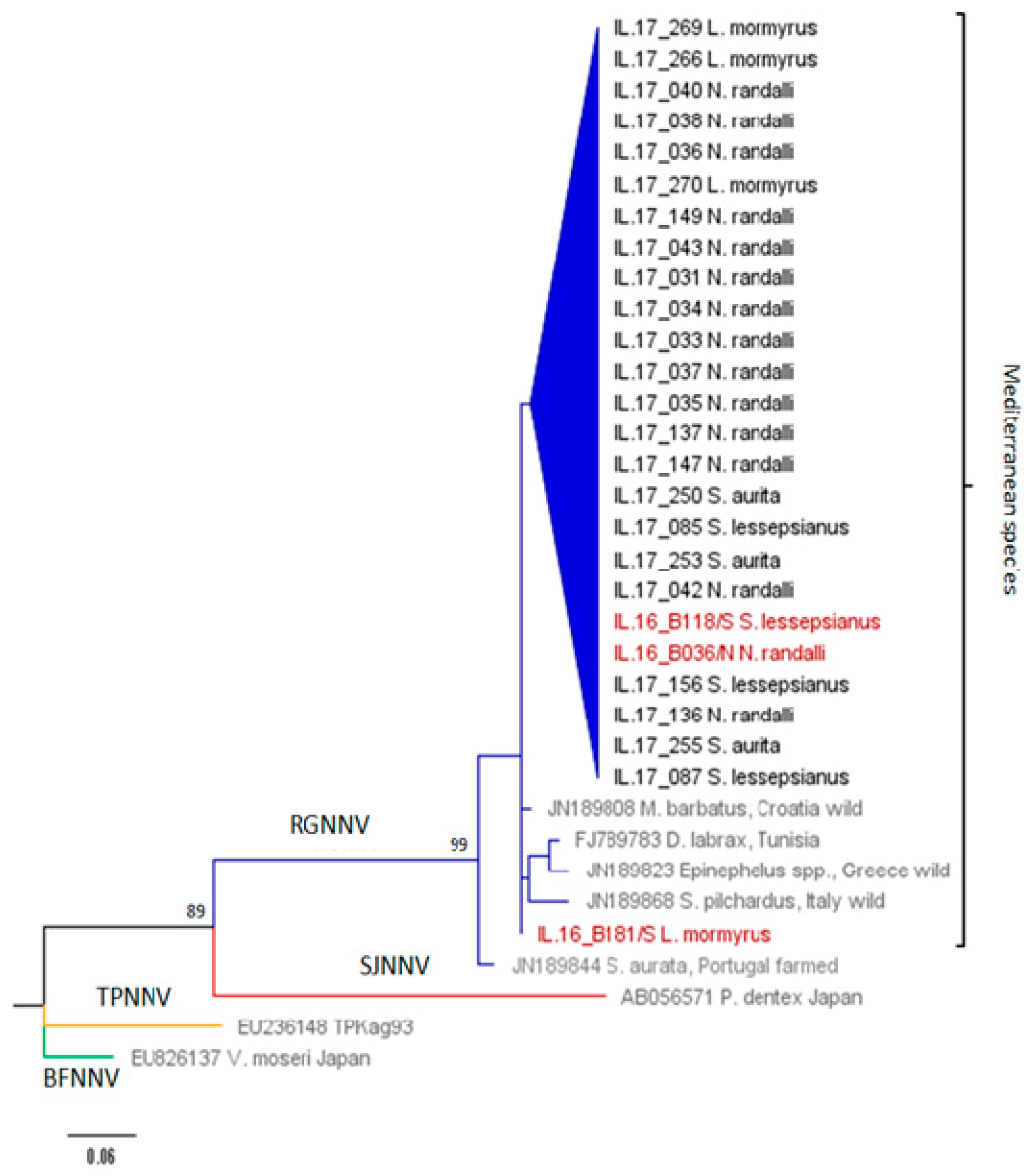
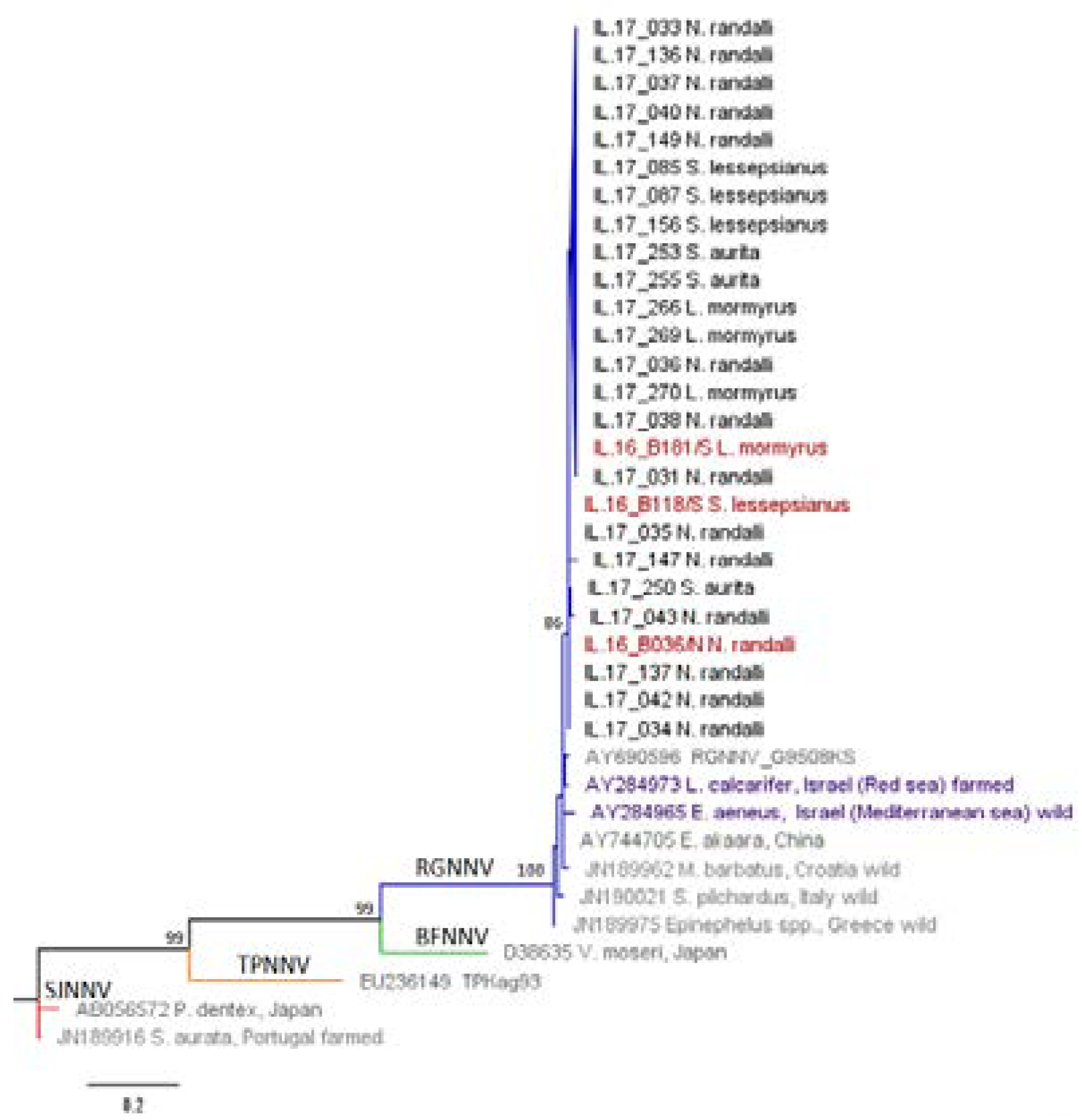
2.4. Sequencing and Phylogenetic Analysis
2.5. Statistical Analysis
3. Results
4. Discussion and Conclusions
Author Contributions
Funding
Conflicts of Interest
References
- Breitbart, M. Marine Viruses: Truth or Dare. Ann Rev. Mar. Sci. 2012, 4, 425–448. [Google Scholar] [CrossRef] [PubMed]
- Corinaldesi, C.; Tangherlini, M.; Dell’Anno, A. From virus isolation to metagenome generation for investigating viral diversity in deep-sea sediments. Sci. Rep. 2017, 7, 8355. [Google Scholar] [CrossRef] [PubMed]
- Laffy, P.W.; Wood-Charlson, E.M.; Turaev, D.; Jutz, S.; Pascelli, C.; Botté, E.S. Reef invertebrate viromics: Diversity, host specificity and functional capacity. Environ. Microbiol. 2018, 20, 2125–2141. [Google Scholar] [CrossRef]
- Coutinho, F.H.; Silveira, C.B.; Gregoracci, G.B.; Thompson, C.C.; Edwards, R.A.; Brussaard, C.P.; Dutilh, B.E.; Thompson, F.L. Marine viruses discovered via metagenomics shed light on viral strategies throughout the oceans. Nat. Commun. 2017, 8, 15955. [Google Scholar] [CrossRef]
- Bandín, I.; Souto, S. Betanodavirus and VER Disease: A 30-year Research Review. Pathogens 2020, 9, 106. [Google Scholar] [CrossRef]
- Vendramin, N.; Zrncic, S.; Padrós, F.; Oraic, D.; Le Breton, A.; Zarza, C.; Olesen, N.J. Fish health in Mediterranean Aquaculture, past mistakes and future challenges. Bull. Eur. Assoc. Fish Pathol. 2016, 36, 38–45. [Google Scholar]
- Zenetos, A.; Çinar, M.E.; Pancucci-Papadopoulou, M.A.; Harmelin, J.G.; Furnari, G.; Andaloro, F.; Bellou, N.; Streftaris, N.; Zibrowius, H. Annotated list of marine alien species in the Mediterranean with records of the worst invasive species. Mediterr. Mar. Sci. 2005, 6, 63–118. [Google Scholar] [CrossRef]
- Bardsley, D.K.; Edwards-Jones, G. Invasive species policy and climate change: Social perceptions of environmental change in the Mediterranean. Environ. Sci. Policy 2007, 10, 230–242. [Google Scholar] [CrossRef]
- Stern, N.; Levitt, Y.; Galil, B.S.; Diamant, A.; Yokeş, M.B.; Goren, M. Distribution and population structure of the alien Indo-Pacific Randall’s threadfin bream Nemipterus randalli in the eastern Mediterranean Sea. J. Fish Biol. 2014, 85, 394–406. [Google Scholar] [CrossRef]
- Golani, D. Distribution of Lessepsian migrant fish in the Mediterranean. Ital. J. Zool. 1998, 65 (Suppl. 1), 95–99. [Google Scholar] [CrossRef]
- Giusti, A.; Guarducci, M.; Stern, N.; Davidovich, N.; Golani, D.; Armani, A. The importance of distinguishing pufferfish species (Lagocephalus spp.) in the Mediterranean Sea for ensuring public health: Evaluation of the genetic databases reliability in supporting species identification. Fish Res. 2019, 210, 14–21. [Google Scholar] [CrossRef]
- Darwin, C. The Origin of Species; John Murray: London, UK, 1859. [Google Scholar]
- Elton, C.S. The Ecology of Invasion by Animals and Plants; Methuen: London, UK, 1958. [Google Scholar]
- Torchin, M.E.; Lafferty, K.D.; Kuris, A.M. Parasites and marine invasions. Parasitology 2002, 124, 137–151. [Google Scholar] [CrossRef]
- Colautti, R.I.; Ricciardi, A.; Grigorovich, I.A.; MacIsaac, H.J. Is invasion success explained by the enemy release hypothesis? Ecol. Lett. 2004, 7, 721–733. [Google Scholar] [CrossRef]
- Hänfling, B. Understanding the establishment success of non-indigenous fishes: Lessons from population genetics. J. Fish Biol. 2007, 71, 115–135. [Google Scholar] [CrossRef]
- Vignon, M.; Sasal, P. Fish introduction and parasites in marine ecosystems: A need for information. Environ. Biol. Fish. 2010, 87, 1–8. [Google Scholar] [CrossRef]
- Møller, A.P.; Rózsa, L. Parasite biodiversity and host defenses: Chewing lice and immune response of their avian hosts. Oecologia 2005, 142, 169–176. [Google Scholar] [CrossRef]
- May, R.M.; Anderson, R.M. Regulation and Stability of Host-Parasite Population Interactions: II. Destabilizing Processes. J. Anim. Ecol. 1978, 47, 249. [Google Scholar] [CrossRef]
- Merella, P.; Pais, A.; Follesa, M.C.; Farjallah, S.; Mele, S.; Piras, M.C.; Garippa, G. Parasites and Lessepsian migration of Fistularia commersonii (Osteichthyes, Fistulariidae): Shadows and light on the enemy release hypothesis. Mar. Biol. 2016, 163, 97. [Google Scholar] [CrossRef]
- Berzak, R.; Scheinin, A.; Davidovich, N.; Regev, Y.; Diga, R.; Tchernov, D.; Morick, D. Prevalence of nervous necrosis virus (NNV) and Streptococcus species in wild marine fish and crustaceans from the Levantine Basin, Mediterranean Sea. Dis. Aquat. Organ. 2019, 133, 7–17. [Google Scholar] [CrossRef]
- Yanong, R.P.E. Necropsy techniques for fish. Semin. Avian Exot. Pet. Med. 2003, 12, 89–105. [Google Scholar] [CrossRef]
- Toffolo, V.; Negrisolo, E.; Maltese, C.; Bovo, G.; Belvedere, P.; Colombo, L.; Dalla Valle, L. Phylogeny of betanodaviruses and molecular evolution of their RNA polymerase and coat proteins. Mol. Phyl. Evol. 2007, 43, 298–308. [Google Scholar] [CrossRef]
- Bovo, G.; Gustinelli, A.; Quaglio, F.; Gobbo, F.; Panzarin, V.; Fusaro, A.; Mutinelli, F.; Caffara, M.; Fioravanti, M.L. Viral encephalopathy and retinopathy outbreak in freshwater fish farmed in Italy. Dis. Aquat. Organ. 2011, 96, 45–54. [Google Scholar] [CrossRef] [PubMed]
- Kumar, S.; Stecher, G.; Tamura, K. MEGA7: Molecular Evolutionary Genetics Analysis Version 7.0 for Bigger Datasets. Mol. Biol. Evol. 2016, 33, 1870–1874. [Google Scholar] [CrossRef] [PubMed]
- Guindon, S.; Dufayard, J.F.; Lefort, V.; Anisimova, M.; Hordijk, W.; Gascuel, O. New Algorithms and Methods to Estimate Maximum-Likelihood Phylogenies: Assessing the Performance of PhyML 3.0. Syst. Biol. 2010, 59, 307–321. [Google Scholar] [CrossRef] [PubMed]
- Lefort, V.; Longueville, J.-E.; Gascuel, O. SMS: Smart Model Selection in PhyML. Mol. Biol. Evol. 2017, 34, 2422–2424. [Google Scholar] [CrossRef]
- Ciulli, S.; Galletti, E.; Grodzki, M.; Alessi, A.; Battilani, M.; Prosperi, S. Isolation and Genetic Characterization of Betanodavirus from Wild Marine Fish from the Adriatic Sea. Vet. Res. Commun. 2007, 31 (Suppl. 1), 221–224. [Google Scholar] [CrossRef]
- Moreno, P.; Olveira, J.G.; Labella, A.; Cutrín, J.M.; Baro, J.C.; Borrego, J.J.; Dopazo, C.P. Surveillance of viruses in wild fish populations in areas around the Gulf of Cadiz (South Atlantic Iberian Peninsula). Appl. Environ. Microbiol. 2014, 80, 6560–6571. [Google Scholar] [CrossRef]
- Cherif, N.; Fatma, A. Nodaviruses in Wild Fish Population Collected Around Aquaculture Cage Sites from Coastal Areas of Tunisia. Fish Aquac. J. 2017, 8, 2–7. [Google Scholar] [CrossRef]
- Golani, D.; Sonin, O. The Japanese threadfin bream Nemipterus japonicus, a new Indo-Pacific fish in the Mediterranean Sea. J. Fish Biol. 2006, 68, 940–943. [Google Scholar] [CrossRef]
- Goren, M.; Galil, B.S.; Diamant, A.; Stern, N.; Levitt-Barmats, Y. Invading up the food web? Invasive fish in the southeastern Mediterranean Sea. Mar. Biol. 2016, 163, 180. [Google Scholar] [CrossRef]
- Ben Rais Lasram, F.; Mouillot, D. Increasing southern invasion enhances congruence between endemic and exotic Mediterranean fish fauna. Biol. Invasions 2009, 11, 697–711. [Google Scholar] [CrossRef]
- Edelist, D. Fishery Management and Marine Invasion in Israel; University of Haifa: Haifa, Israel, 2013. [Google Scholar]
- Toffan, A.; Panzarin, V.; Toson, M.; Cecchettin, K.; Pascoli, F. Water temperature affects pathogenicity of different betanodavirus genotypes in experimentally challenged Dicentrarchus labrax. Dis. Aquat. Organ. 2016, 119, 231–238. [Google Scholar] [CrossRef] [PubMed]
- Ito, Y.; Okinaka, Y.; Mori, K.I.; Sugaya, T.; Nishioka, T.; Oka, M.; Nakai, T. Variable region of betanodavirus RNA2 is sufficient to determine host specificity. Dis. Aquat. Organ. 2008, 79, 199–205. [Google Scholar] [CrossRef]
- Jimenez, C.; Andreou, V.; Evriviadou, M.; Munkes, B.; Hadjioannou, L.; Petrou, A.; Alhaija, R.A. Epibenthic communities associated with unintentional artificial reefs (modern shipwrecks) under contrasting regimes of nutrients in the Levantine Sea (Cyprus and Lebanon). PLoS ONE 2017, 12, e0182486. [Google Scholar] [CrossRef]
- Rilov, G. Multi-species collapses at the warm edge of a warming sea. Sci. Rep. 2016, 6, 36897. [Google Scholar] [CrossRef] [PubMed]
- Chaidez, V.; Dreano, D.; Agusti, S.; Duarte, C.; Hoteit, I. Decadal trends in Red Sea maximum surface temperature. Sci. Rep. 2017, 7, 1–8. [Google Scholar] [CrossRef] [PubMed]
- Raitsos, D.E.; Hoteit, I.; Prihartato, P.K.; Chronis, T.; Triantafyllou, G.; Abualnaja, Y. Abrupt warming of the Red Sea. Geophys. Res. Lett. 2011, 38, 1–5. [Google Scholar] [CrossRef]
- Chang J-S, Chi S-C. GHSC70 is involved in the cellular entry of nervous necrosis virus. J. Virol. 2015, 89, 61–70. [Google Scholar] [CrossRef]
- Costa, J.Z.; Thompson, K.D. Understanding the interaction between Betanodavirus and its host for the development of prophylactic measures for viral encephalopathy and retinopathy. Fish Shellfish Immunol. 2016, 53, 35–49. [Google Scholar] [CrossRef] [PubMed]
- Ucko, M.; Colorni, A.; Diamant, A. Nodavirus infections in Israeli mariculture. J. Fish Dis. 2004, 27, 459–469. [Google Scholar] [CrossRef]
- Panzarin, V.; Fusaro, A.; Monne, I.; Cappellozza, E.; Patarnello, P.; Bovo, G.; Capua, I.; Holmes, E.C.; Cattoli, G. Molecular epidemiology and evolutionary dynamics of betanodavirus in southern Europe. Infect. Genet. Evol. 2012, 12, 63–70. [Google Scholar] [CrossRef] [PubMed]
- Kvitt, H.; Heinisch, G.; Diamant, A. Detection and phylogeny of Lymphocystivirus in sea bream Sparus aurata based on the DNA polymerase gene and major capsid protein sequences. Aquaculture 2008, 275, 58–63. [Google Scholar] [CrossRef]
- Haddad-Boubaker, S.; Bouzgarou, N.; Fakhfakh, E.; Khayech, M.; Mohamed, S.B.; Megdich, A.; Chéhida, N.B. Detection and genetic characterization of lymphocystis disease virus (LCDV) isolated during disease outbreaks in cultured gilt-head sea bream Sparus aurata in Tunisia. Fish Pathol. 2013, 48, 101–104. [Google Scholar] [CrossRef]
| Host Species/Isolate | Origin | NNV Strain | RNA1 | RNA2 |
|---|---|---|---|---|
| Epinephelus akaara | China | RGNNV | - | AY744705 |
| G9508KS | n.a | RGNNV | - | AY690596 |
| Dicentrarchus labrax | Tunisia | RGNNV | FJ789783 | - |
| Sardina pilchardus | Italy, wild | RGNNV | JN189868 | JN190021 |
| Epinephelus spp. | Greece, wild | RGNNV | JN189823 | JN189975 |
| Mullus barbatus | Croatia, wild | RGNNV | JN189808 | JN189962 |
| Epinephelus aeneus | Israel (Med. Sea b), wild | RGNNV | - | AY284965 |
| Lates calcarifer | Israel (Red Sea), farmed | RGNNV | - | AY284973 |
| Sparus aurata | Portugal, farmed | RG/SJ a | JN189844 | JN189916 |
| Pseudocaranx dentex | Japan | SJNNV | AB056571 | AB056572 |
| Verasper moseri | Japan | BFNNV | EU826137 | D38635 |
| TPKag93 | Japan | TPNNV | EU236148 | EU236149 |
| 2016 | 2017 | Total | |||||||
|---|---|---|---|---|---|---|---|---|---|
| Fish Species | n | Positive | % Prevalence | n | Positive | % Prevalence | n | Positive | % Prevalence |
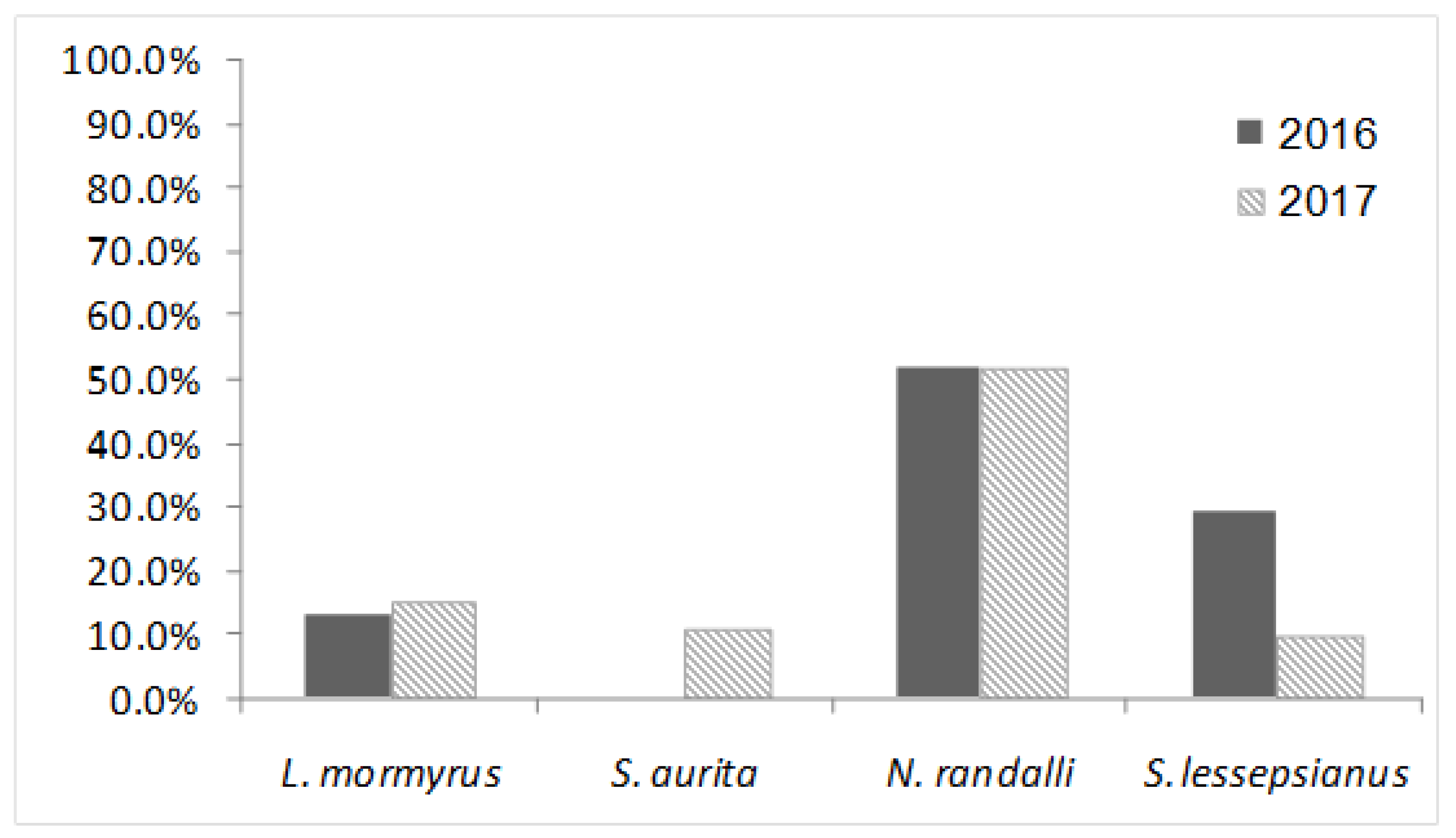
| Lithognathus mormyrus | 30 | 4 | 13.33 | 20 | 3 | 15 | 50 | 7 | 14 |
| Sardinella aurita | 30 | 0 | 0 | 28 | 3 | 10.71 | 58 | 3 | 5.17 |
| Nemipterus randalli * | 29 | 15 | 51.72 | 31 | 16 | 51.61 | 60 | 31 | 51.67 |
| Saurida lessepsianus * | 38 | 11 | 28.95 | 30 | 3 | 10 | 68 | 14 | 20.59 |
| Total | 127 | 30 | 23.62 | 109 | 25 | 22.94 | 236 | 55 | 23.31 |
© 2020 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Lampert, Y.; Berzak, R.; Davidovich, N.; Diamant, A.; Stern, N.; Scheinin, A.P.; Tchernov, D.; Morick, D. Indigenous versus Lessepsian Hosts: Nervous Necrosis Virus (NNV) in Eastern Mediterranean Sea Fish. Viruses 2020, 12, 430. https://doi.org/10.3390/v12040430
Lampert Y, Berzak R, Davidovich N, Diamant A, Stern N, Scheinin AP, Tchernov D, Morick D. Indigenous versus Lessepsian Hosts: Nervous Necrosis Virus (NNV) in Eastern Mediterranean Sea Fish. Viruses. 2020; 12(4):430. https://doi.org/10.3390/v12040430
Chicago/Turabian StyleLampert, Yael, Ran Berzak, Nadav Davidovich, Arik Diamant, Nir Stern, Aviad P. Scheinin, Dan Tchernov, and Danny Morick. 2020. "Indigenous versus Lessepsian Hosts: Nervous Necrosis Virus (NNV) in Eastern Mediterranean Sea Fish" Viruses 12, no. 4: 430. https://doi.org/10.3390/v12040430
APA StyleLampert, Y., Berzak, R., Davidovich, N., Diamant, A., Stern, N., Scheinin, A. P., Tchernov, D., & Morick, D. (2020). Indigenous versus Lessepsian Hosts: Nervous Necrosis Virus (NNV) in Eastern Mediterranean Sea Fish. Viruses, 12(4), 430. https://doi.org/10.3390/v12040430







