The First Non-LRV RNA Virus in Leishmania
Abstract
1. Introduction
2. Materials and Methods
2.1. Parasite Culture, DNA Isolation, and Verification of Species Identity
2.2. DsRNA Isolation and Next-Generation Sequencing
2.3. Viral Sequence Assembly
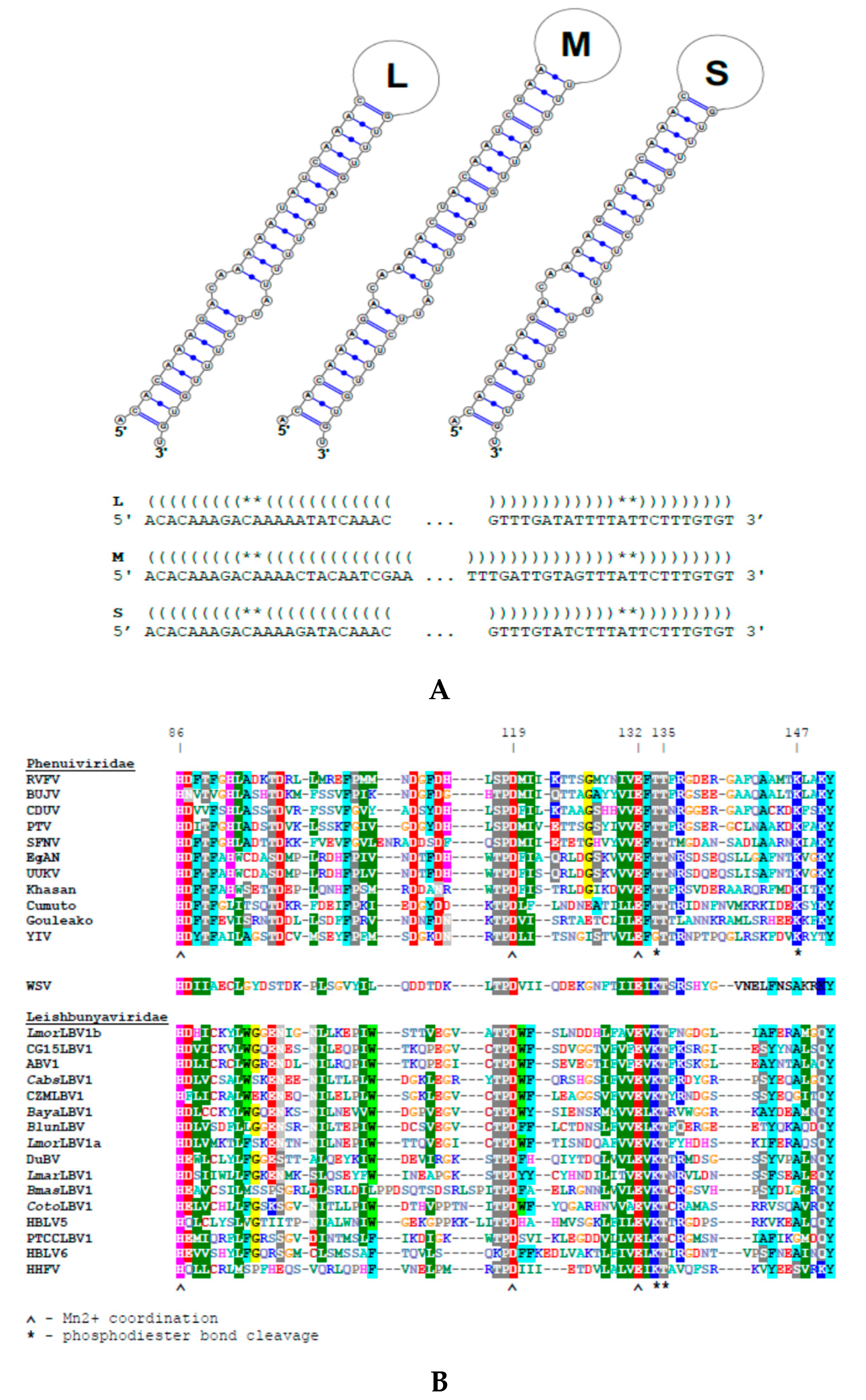
2.4. Prediction of Functional Elements
2.5. Phylogenetic Analyses
2.6. Negative-Stain Transmission Electron Microscopy
2.7. Treatment with Ribavirin
2.8. Macrophage Infection
3. Results
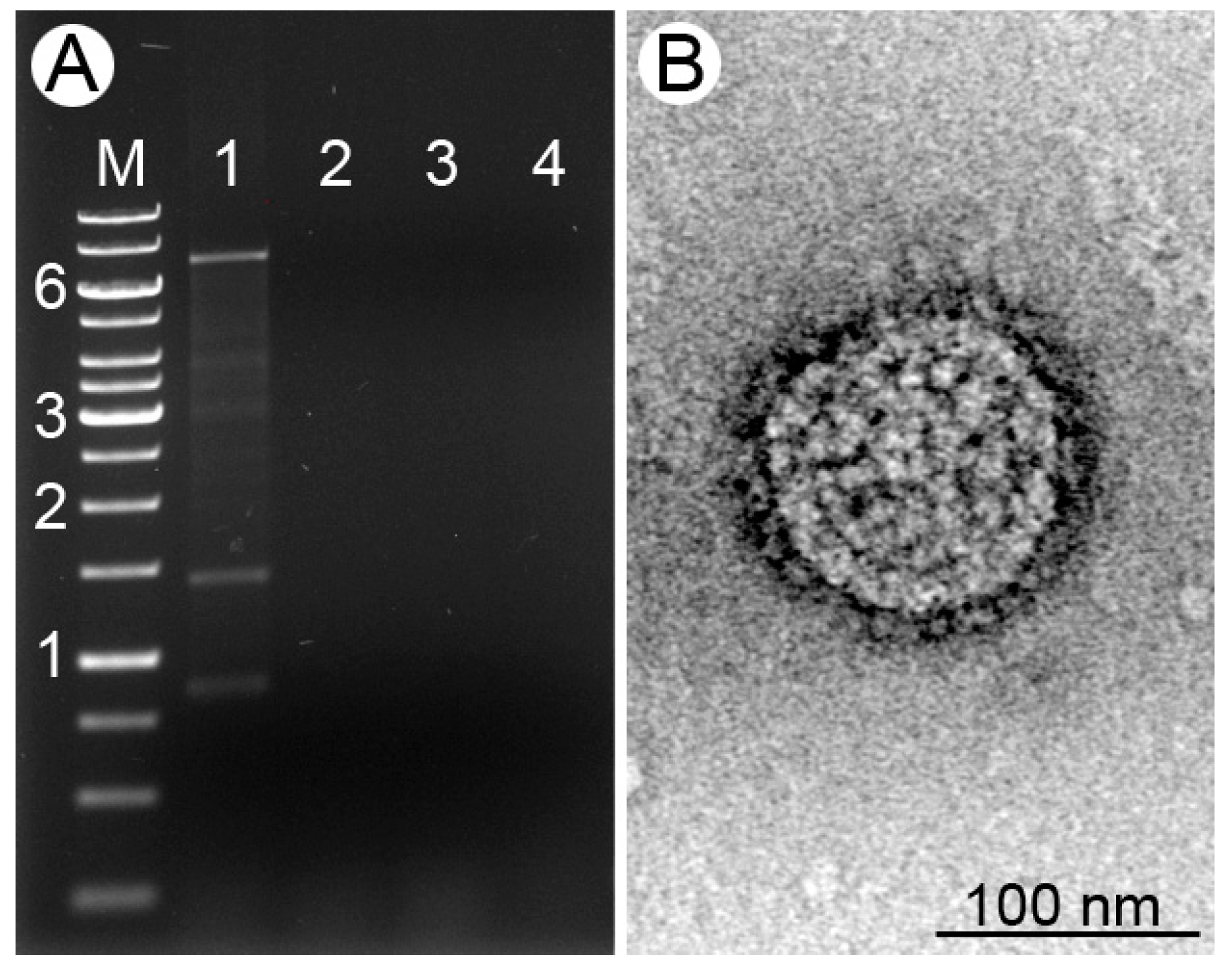
3.1. Viral dsRNA in Leishmania (M.) martiniquensis
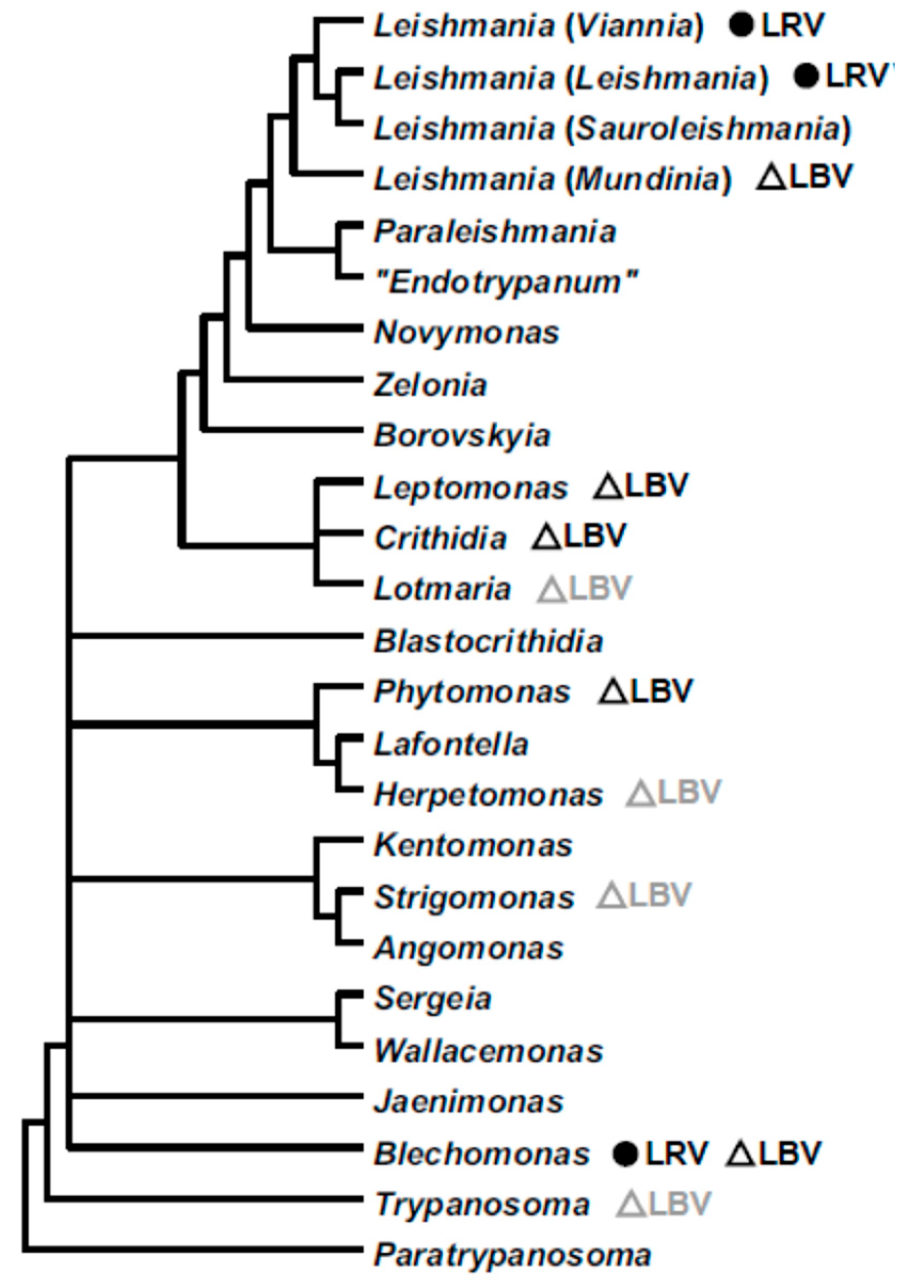
3.2. Phylogeny
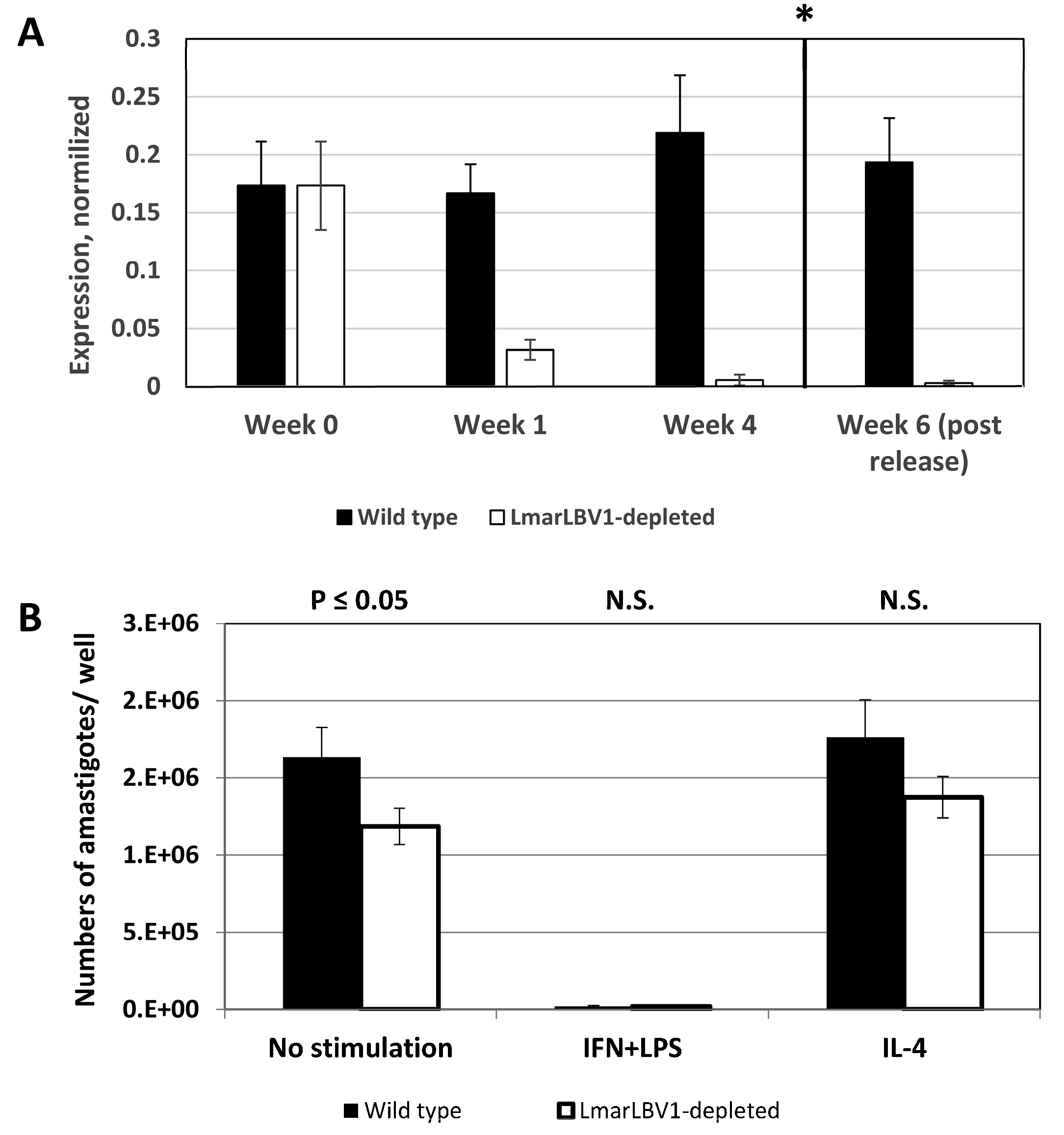
3.3. LmarLBV1 Has Minor Effect on Leishmania Infectivity In Vitro
4. Discussion
Supplementary Materials
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
References
- Lefkowitz, E.J.; Dempsey, D.M.; Hendrickson, R.C.; Orton, R.J.; Siddell, S.G.; Smith, D.B. Virus taxonomy: The database of the International Committee on Taxonomy of Viruses (ICTV). Nucleic Acids Res. 2018, 46, D708–D717. [Google Scholar] [CrossRef] [PubMed]
- Wichgers Schreur, P.J.; Kormelink, R.; Kortekaas, J. Genome packaging of the Bunyavirales. Curr. Opin. Virol. 2018, 33, 151–155. [Google Scholar] [CrossRef] [PubMed]
- Elliott, R.M. Molecular biology of the Bunyaviridae. J. Gen. Virol. 1990, 71, 501–522. [Google Scholar] [CrossRef] [PubMed]
- Sun, Y.; Li, J.; Gao, G.F.; Tien, P.; Liu, W. Bunyavirales ribonucleoproteins: The viral replication and transcription machinery. Crit. Rev. Microbiol. 2018, 44, 522–540. [Google Scholar] [CrossRef]
- Gerlach, P.; Malet, H.; Cusack, S.; Reguera, J. Structural insights into Bunyavirus replication and its regulation by the vRNA promoter. Cell 2015, 161, 1267–1279. [Google Scholar] [CrossRef]
- Junglen, S. Evolutionary origin of pathogenic arthropod-borne viruses - a case study in the family Bunyaviridae. Curr. Opin. Insect Sci. 2016, 16, 81–86. [Google Scholar] [CrossRef]
- Li, C.X.; Shi, M.; Tian, J.H.; Lin, X.D.; Kang, Y.J.; Chen, L.J.; Qin, X.C.; Xu, J.; Holmes, E.C.; Zhang, Y.Z. Unprecedented genomic diversity of RNA viruses in arthropods reveals the ancestry of negative-sense RNA viruses. Elife 2015, 4. [Google Scholar] [CrossRef]
- Shi, M.; Lin, X.D.; Tian, J.H.; Chen, L.J.; Chen, X.; Li, C.X.; Qin, X.C.; Li, J.; Cao, J.P.; Eden, J.S.; et al. Redefining the invertebrate RNA virosphere. Nature 2016, 540, 539–543. [Google Scholar] [CrossRef]
- Shi, M.; Lin, X.D.; Chen, X.; Tian, J.H.; Chen, L.J.; Li, K.; Wang, W.; Eden, J.S.; Shen, J.J.; Liu, L.; et al. The evolutionary history of vertebrate RNA viruses. Nature 2018, 556, 197–202. [Google Scholar] [CrossRef]
- Ullman, D.E.; German, T.L.; Sherwood, J.L.; Westcot, D.M.; Cantone, F.A. Tospovirus replication in insect vector cells - immunocytochemical evidence that the nonstructural protein encoded by the S RNA of Tomato spotted wilt tospovirus is present in thrips vector cells. Phytopathology 1993, 83, 456–463. [Google Scholar] [CrossRef]
- Whitfield, A.E.; Ullman, D.E.; German, T.L. Tospovirus-thrips interactions. Annu. Rev. Phytopathol. 2005, 43, 459–489. [Google Scholar] [CrossRef] [PubMed]
- Grybchuk, D.; Kostygov, A.Y.; Macedo, D.H.; d’Avila-Levy, C.M.; Yurchenko, V. RNA viruses in trypanosomatid parasites: A historical overview. Mem Inst. Oswaldo Cruz 2018, 113, e170487. [Google Scholar] [CrossRef] [PubMed]
- Maslov, D.A.; Opperdoes, F.R.; Kostygov, A.Y.; Hashimi, H.; Lukeš, J.; Yurchenko, V. Recent advances in trypanosomatid research: Genome organization, expression, metabolism, taxonomy and evolution. Parasitology 2019, 146, 1–27. [Google Scholar] [CrossRef] [PubMed]
- Maslov, D.A.; Votýpka, J.; Yurchenko, V.; Lukeš, J. Diversity and phylogeny of insect trypanosomatids: All that is hidden shall be revealed. Trends Parasitol. 2013, 29, 43–52. [Google Scholar] [CrossRef] [PubMed]
- McGhee, R.B.; Cosgrove, W.B. Biology and physiology of the lower Trypanosomatidae. Microbiol Rev. 1980, 44, 140–173. [Google Scholar] [CrossRef]
- Flegontov, P.; Butenko, A.; Firsov, S.; Kraeva, N.; Eliáš, M.; Field, M.C.; Filatov, D.; Flegontova, O.; Gerasimov, E.S.; Hlaváčová, J.; et al. Genome of Leptomonas pyrrhocoris: A high-quality reference for monoxenous trypanosomatids and new insights into evolution of Leishmania. Sci. Rep. 2016, 6, 23704. [Google Scholar] [CrossRef]
- Stevens, J.R.; Gibson, W.C. The evolution of pathogenic trypanosomes. Cad. Saude Publica 1999, 15, 673–684. [Google Scholar] [CrossRef]
- Camargo, E.P. Phytomonas and other trypanosomatid parasites of plants and fruit. Adv. Parasitol. 1999, 42, 29–112. [Google Scholar]
- Lukeš, J.; Skalický, T.; Týč, J.; Votýpka, J.; Yurchenko, V. Evolution of parasitism in kinetoplastid flagellates. Mol. Biochem. Parasitol. 2014, 195, 115–122. [Google Scholar] [CrossRef]
- Dvorák, V.; Shaw, J.J.; Volf, P. Parasite biology: The vectors. In The Leishmaniases: Old Neglected Tropical Diseases; Bruschi, F., Gradoni, L., Eds.; Springer: Cham, Switzerland, 2018; pp. 31–77. [Google Scholar]
- Espinosa, O.A.; Serrano, M.G.; Camargo, E.P.; Teixeira, M.M.; Shaw, J.J. An appraisal of the taxonomy and nomenclature of trypanosomatids presently classified as Leishmania and Endotrypanum. Parasitology 2018, 145, 430–442. [Google Scholar] [CrossRef]
- Kostygov, A.Y.; Yurchenko, V. Revised classification of the subfamily Leishmaniinae (Trypanosomatidae). Folia Parasitol 2017, 64, 020. [Google Scholar] [CrossRef] [PubMed]
- Paranaiba, L.F.; Pinheiro, L.J.; Macedo, D.H.; Menezes-Neto, A.; Torrecilhas, A.C.; Tafuri, W.L.; Soares, R.P. An overview on Leishmania (Mundinia) enriettii: Biology, immunopathology, LRV and extracellular vesicles during the host-parasite interaction. Parasitology 2018, 145, 1265–1273. [Google Scholar] [CrossRef] [PubMed]
- Paranaiba, L.F.; Pinheiro, L.J.; Torrecilhas, A.C.; Macedo, D.H.; Menezes-Neto, A.; Tafuri, W.L.; Soares, R.P. Leishmania enriettii (Muniz & Medina, 1948): A highly diverse parasite is here to stay. PLoS Pathog 2017, 13, e1006303. [Google Scholar] [CrossRef]
- Tarr, P.I.; Aline, R.F., Jr.; Smiley, B.L.; Scholler, J.; Keithly, J.; Stuart, K. LR1: A candidate RNA virus of Leishmania. Proc. Natl. Acad. Sci. USA 1988, 85, 9572–9575. [Google Scholar] [CrossRef]
- Stuart, K.D.; Weeks, R.; Guilbride, L.; Myler, P.J. Molecular organization of Leishmania RNA virus 1. Proc. Natl. Acad. Sci. USA 1992, 89, 8596–8600. [Google Scholar] [CrossRef]
- Widmer, G.; Comeau, A.M.; Furlong, D.B.; Wirth, D.F.; Patterson, J.L. Characterization of a RNA virus from the parasite Leishmania. Proc. Natl. Acad. Sci. USA 1989, 86, 5979–5982. [Google Scholar] [CrossRef]
- Guilbride, L.; Myler, P.J.; Stuart, K. Distribution and sequence divergence of LRV1 viruses among different Leishmania species. Mol. Biochem. Parasitol. 1992, 54, 101–104. [Google Scholar] [CrossRef]
- Scheffter, S.M.; Ro, Y.T.; Chung, I.K.; Patterson, J.L. The complete sequence of Leishmania RNA virus LRV2-1, a virus of an Old World parasite strain. Virology 1995, 212, 84–90. [Google Scholar] [CrossRef]
- Zangger, H.; Hailu, A.; Desponds, C.; Lye, L.F.; Akopyants, N.S.; Dobson, D.E.; Ronet, C.; Ghalib, H.; Beverley, S.M.; Fasel, N. Leishmania aethiopica field isolates bearing an endosymbiontic dsRNA virus induce pro-inflammatory cytokine response. PLoS Negl. Trop. Dis. 2014, 8, e2836. [Google Scholar] [CrossRef]
- Hajjaran, H.; Mahdi, M.; Mohebali, M.; Samimi-Rad, K.; Ataei-Pirkooh, A.; Kazemi-Rad, E.; Naddaf, S.R.; Raoofian, R. Detection and molecular identification of Leishmania RNA virus (LRV) in Iranian Leishmania species. Arch. Virol. 2016, 161, 3385–3390. [Google Scholar] [CrossRef]
- Grybchuk, D.; Kostygov, A.Y.; Macedo, D.H.; Votypka, J.; Lukes, J.; Yurchenko, V. RNA viruses in Blechomonas (Trypanosomatidae) and evolution of Leishmaniavirus. mBio 2018, 9, e01932-01918. [Google Scholar] [CrossRef] [PubMed]
- Ives, A.; Ronet, C.; Prevel, F.; Ruzzante, G.; Fuertes-Marraco, S.; Schutz, F.; Zangger, H.; Revaz-Breton, M.; Lye, L.F.; Hickerson, S.M.; et al. Leishmania RNA virus controls the severity of mucocutaneous leishmaniasis. Science 2011, 331, 775–778. [Google Scholar]
- De Carvalho, R.V.H.; Lima-Junior, D.S.; da Silva, M.V.G.; Dilucca, M.; Rodrigues, T.S.; Horta, C.V.; Silva, A.L.N.; da Silva, P.F.; Frantz, F.G.; Lorenzon, L.B.; et al. Leishmania RNA virus exacerbates leishmaniasis by subverting innate immunity via TLR3-mediated NLRP3 inflammasome inhibition. Nat. Commun. 2019, 10, 5273. [Google Scholar] [CrossRef]
- Hartley, M.A.; Bourreau, E.; Rossi, M.; Castiglioni, P.; Eren, R.O.; Prevel, F.; Couppie, P.; Hickerson, S.M.; Launois, P.; Beverley, S.M.; et al. Leishmaniavirus-dependent metastatic leishmaniasis is prevented by blocking IL-17A. PLoS Pathog 2016, 12, e1005852. [Google Scholar] [CrossRef]
- Rossi, M.; Castiglioni, P.; Hartley, M.A.; Eren, R.O.; Prevel, F.; Desponds, C.; Utzschneider, D.T.; Zehn, D.; Cusi, M.G.; Kuhlmann, F.M.; et al. Type I interferons induced by endogenous or exogenous viral infections promote metastasis and relapse of leishmaniasis. Proc. Natl. Acad. Sci. USA 2017, 114, 4987–4992. [Google Scholar] [CrossRef] [PubMed]
- Widmer, G.; Dooley, S. Phylogenetic analysis of Leishmania RNA virus and Leishmania suggests ancient virus-parasite association. Nucleic Acids Res. 1995, 23, 2300–2304. [Google Scholar] [CrossRef] [PubMed]
- Cantanhêde, L.M.; da Silva Junior, C.F.; Ito, M.M.; Felipin, K.P.; Nicolete, R.; Salcedo, J.M.; Porrozzi, R.; Cupolillo, E.; Ferreira Rde, G. Further evidence of an association between the presence of Leishmania RNA Virus 1 and the mucosal manifestations in tegumentary leishmaniasis patients. PLoS Negl. Trop. Dis. 2015, 9, e0004079. [Google Scholar] [CrossRef]
- Grybchuk, D.; Akopyants, N.S.; Kostygov, A.Y.; Konovalovas, A.; Lye, L.F.; Dobson, D.E.; Zangger, H.; Fasel, N.; Butenko, A.; Frolov, A.O.; et al. Viral discovery and diversity in trypanosomatid protozoa with a focus on relatives of the human parasite Leishmania. Proc. Natl. Acad. Sci. USA 2018, 115, E506–E515. [Google Scholar] [CrossRef]
- Akopyants, N.S.; Lye, L.F.; Dobson, D.E.; Lukeš, J.; Beverley, S.M. A novel bunyavirus-like virus of trypanosomatid protist parasites. Genome Announc 2016, 4, e00715-00716. [Google Scholar] [CrossRef]
- Maslov, D.A.; Lukeš, J.; Jirků, M.; Simpson, L. Phylogeny of trypanosomes as inferred from the small and large subunit rRNAs: Implications for the evolution of parasitism in the trypanosomatid protozoa. Mol. Biochem. Parasitol. 1996, 75, 197–205. [Google Scholar] [CrossRef]
- Yurchenko, V.; Kostygov, A.; Havlová, J.; Grybchuk-Ieremenko, A.; Ševčíková, T.; Lukeš, J.; Ševčík, J.; Votýpka, J. Diversity of trypanosomatids in cockroaches and the description of Herpetomonas tarakana sp. n. J. Eukaryot Microbiol 2016, 63, 198–209. [Google Scholar] [CrossRef] [PubMed]
- Gerasimov, E.S.; Kostygov, A.Y.; Yan, S.; Kolesnikov, A.A. From cryptogene to gene? ND8 editing domain reduction in insect trypanosomatids. Eur. J. Protistol. 2012, 48, 185–193. [Google Scholar] [CrossRef] [PubMed]
- Altschul, S.F.; Gish, W.; Miller, W.; Myers, E.W.; Lipman, D.J. Basic local alignment search tool. J. Mol. Biol 1990, 215, 403–410. [Google Scholar] [CrossRef]
- Bolger, A.M.; Lohse, M.; Usadel, B. Trimmomatic: A flexible trimmer for Illumina sequence data. Bioinformatics 2014, 30, 2114–2120. [Google Scholar] [CrossRef] [PubMed]
- Grabherr, M.G.; Haas, B.J.; Yassour, M.; Levin, J.Z.; Thompson, D.A.; Amit, I.; Adiconis, X.; Fan, L.; Raychowdhury, R.; Zeng, Q.; et al. Full-length transcriptome assembly from RNA-Seq data without a reference genome. Nat. Biotechnol. 2011, 29, 644–652. [Google Scholar] [CrossRef]
- Langmead, B.; Salzberg, S.L. Fast gapped-read alignment with Bowtie 2. Nat. Methods 2012, 9, 357–359. [Google Scholar] [CrossRef]
- Li, H.; Handsaker, B.; Wysoker, A.; Fennell, T.; Ruan, J.; Homer, N.; Marth, G.; Abecasis, G.; Durbin, R.; Genome Project Data Processing, S. The Sequence Alignment/Map format and SAMtools. Bioinformatics 2009, 25, 2078–2079. [Google Scholar] [CrossRef]
- Quinlan, A.R.; Hall, I.M. BEDTools: A flexible suite of utilities for comparing genomic features. Bioinformatics 2010, 26, 841–842. [Google Scholar] [CrossRef]
- Wheeler, D.L.; Church, D.M.; Federhen, S.; Lash, A.E.; Madden, T.L.; Pontius, J.U.; Schuler, G.D.; Schriml, L.M.; Sequeira, E.; Tatusova, T.A.; et al. Database resources of the National Center for Biotechnology. Nucleic Acids Res. 2003, 31, 28–33. [Google Scholar] [CrossRef]
- Marchler-Bauer, A.; Derbyshire, M.K.; Gonzales, N.R.; Lu, S.; Chitsaz, F.; Geer, L.Y.; Geer, R.C.; He, J.; Gwadz, M.; Hurwitz, D.I.; et al. CDD: NCBI’s conserved domain database. Nucleic Acids Res. 2015, 43, D222–D226. [Google Scholar] [CrossRef]
- Hofmann, K.; Stoffel, W. TMBase - a database of membrane spanning protein segments. Biol. Chem. Hoppe-Seyler 1993, 374, 166. [Google Scholar]
- Käll, L.; Krogh, A.; Sonnhammer, E.L. Advantages of combined transmembrane topology and signal peptide prediction--the Phobius web server. Nucleic Acids Res. 2007, 35, W429–W432. [Google Scholar] [CrossRef] [PubMed]
- McGuffin, L.J.; Bryson, K.; Jones, D.T. The PSIPRED protein structure prediction server. Bioinformatics 2000, 16, 404–405. [Google Scholar] [CrossRef] [PubMed]
- Petersen, T.N.; Brunak, S.; von Heijne, G.; Nielsen, H. SignalP 4.0: Discriminating signal peptides from transmembrane regions. Nat. Methods 2011, 8, 785–786. [Google Scholar] [CrossRef] [PubMed]
- Katoh, K.; Standley, D.M. MAFFT multiple sequence alignment software version 7: Improvements in performance and usability. Mol. Biol. Evol. 2013, 30, 772–780. [Google Scholar] [CrossRef] [PubMed]
- Capella-Gutiérrez, S.; Silla-Martinez, J.M.; Gabaldon, T. trimAl: A tool for automated alignment trimming in large-scale phylogenetic analyses. Bioinformatics 2009, 25, 1972–1973. [Google Scholar] [CrossRef]
- Nguyen, L.T.; Schmidt, H.A.; von Haeseler, A.; Minh, B.Q. IQ-TREE: A fast and effective stochastic algorithm for estimating maximum-likelihood phylogenies. Mol. Biol Evol 2015, 32, 268–274. [Google Scholar] [CrossRef]
- Kalyaanamoorthy, S.; Minh, B.Q.; Wong, T.K.F.; von Haeseler, A.; Jermiin, L.S. ModelFinder: Fast model selection for accurate phylogenetic estimates. Nat. Methods 2017, 14, 587–589. [Google Scholar] [CrossRef]
- Ronquist, F.; Teslenko, M.; van der Mark, P.; Ayres, D.L.; Darling, A.; Hohna, S.; Larget, B.; Liu, L.; Suchard, M.A.; Huelsenbeck, J.P. MrBayes 3.2: Efficient Bayesian phylogenetic inference and model choice across a large model space. Syst. Biol. 2012, 61, 539–542. [Google Scholar] [CrossRef]
- Kumar, S.; Stecher, G.; Li, M.; Knyaz, C.; Tamura, K. MEGA X: Molecular Evolutionary Genetics Analysis across computing platforms. Mol. Biol. Evol. 2018, 35, 1547–1549. [Google Scholar] [CrossRef]
- Ishemgulova, A.; Hlavacova, J.; Majerova, K.; Butenko, A.; Lukes, J.; Votypka, J.; Volf, P.; Yurchenko, V. CRISPR/Cas9 in Leishmania mexicana: A case study of LmxBTN1. PLoS ONE 2018, 13, e0192723. [Google Scholar] [CrossRef] [PubMed]
- Ishemgulova, A.; Kraeva, N.; Faktorová, D.; Podešvová, L.; Lukeš, J.; Yurchenko, V. T7 polymerase-driven transcription is downregulated in metacyclic promastigotes and amastigotes of Leishmania mexicana. Folia Parasitol. 2016, 63, 016. [Google Scholar] [CrossRef] [PubMed]
- Garin, Y.J.; Sulahian, A.; Meneceur, P.; Pratlong, F.; Prina, E.; Gangneux, J.; Dedet, J.P.; Derouin, F. Experimental pathogenicity of a presumed monoxenous trypanosomatid isolated from humans in a murine model. J. Eukaryot Microbiol 2001, 48, 170–176. [Google Scholar] [CrossRef] [PubMed]
- Giraud, E.; Leštinová, T.; Derrick, T.; Martin, O.; Dillon, R.J.; Volf, P.; Muller, I.; Bates, P.A.; Rogers, M.E. Leishmania proteophosphoglycans regurgitated from infected sand flies accelerate dermal wound repair and exacerbate leishmaniasis via insulin-like growth factor 1-dependent signalling. PLoS Pathog 2018, 14, e1006794. [Google Scholar] [CrossRef] [PubMed]
- Cifuentes-Muñoz, N.; Salazar-Quiroz, N.; Tischler, N.D. Hantavirus Gn and Gc envelope glycoproteins: Key structural units for virus cell entry and virus assembly. Viruses 2014, 6, 1801–1822. [Google Scholar] [CrossRef] [PubMed]
- Sanchez, A.J.; Vincent, M.J.; Nichol, S.T. Characterization of the glycoproteins of Crimean-Congo hemorrhagic fever virus. J. Virol. 2002, 76, 7263–7275. [Google Scholar] [CrossRef] [PubMed]
- Remnant, E.J.; Shi, M.; Buchmann, G.; Blacquiere, T.; Holmes, E.C.; Beekman, M.; Ashe, A. A diverse range of novel RNA viruses in geographically distinct honey bee populations. J. Virol. 2017, 91, e00158-00117. [Google Scholar] [CrossRef]
- Albornoz, A.; Hoffmann, A.B.; Lozach, P.Y.; Tischler, N.D. Early bunyavirus-host cell interactions. Viruses 2016, 8, 143. [Google Scholar] [CrossRef]
- Overath, P.; Stierhof, Y.D.; Wiese, M. Endocytosis and secretion in trypanosomatid parasites - tumultuous traffic in a pocket. Trends Cell Biol 1997, 7, 27–33. [Google Scholar] [CrossRef]
- Landfear, S.M.; Ignatushchenko, M. The flagellum and flagellar pocket of trypanosomatids. Mol. Biochem. Parasitol. 2001, 115, 1–17. [Google Scholar] [CrossRef]
- Szempruch, A.J.; Sykes, S.E.; Kieft, R.; Dennison, L.; Becker, A.C.; Gartrell, A.; Martin, W.J.; Nakayasu, E.S.; Almeida, I.C.; Hajduk, S.L.; et al. Extracellular vesicles from Trypanosoma brucei mediate virulence factor transfer and cause host anemia. Cell 2016, 164, 246–257. [Google Scholar] [CrossRef] [PubMed]
- Allen, C.L.; Goulding, D.; Field, M.C. Clathrin-mediated endocytosis is essential in Trypanosoma brucei. EMBO J. 2003, 22, 4991–5002. [Google Scholar] [CrossRef] [PubMed]
- Hung, C.H.; Qiao, X.; Lee, P.T.; Lee, M.G. Clathrin-dependent targeting of receptors to the flagellar pocket of procyclic-form Trypanosoma brucei. Eukaryot Cell 2004, 3, 1004–1014. [Google Scholar] [CrossRef] [PubMed]
- Léger, P.; Lozach, P.-Y. Bunyaviruses: From transmission by arthropods to virus entry into the mammalian host first-target cells. Future Virol. 2015, 10, 859–881. [Google Scholar] [CrossRef]
- Shi, X.; Elliott, R.M. Analysis of glycoproteins of viruses in the family Bunyaviridae. Methods Mol. Biol. 2007, 379, 137–148. [Google Scholar] [CrossRef] [PubMed]
- Överby, A.K.; Pettersson, R.F.; Neve, E.P. The glycoprotein cytoplasmic tail of Uukuniemi virus (Bunyaviridae) interacts with ribonucleoproteins and is critical for genome packaging. J. Virol. 2007, 81, 3198–3205. [Google Scholar] [CrossRef]
- Strandin, T.; Hepojoki, J.; Vaheri, A. Cytoplasmic tails of bunyavirus Gn glycoproteins – could they act as matrix protein surrogates? Virology 2013, 437, 73–80. [Google Scholar] [CrossRef]
- Wu, Y.; Zhu, Y.; Gao, F.; Jiao, Y.; Oladejo, B.O.; Chai, Y.; Bi, Y.; Lu, S.; Dong, M.; Zhang, C.; et al. Structures of phlebovirus glycoprotein Gn and identification of a neutralizing antibody epitope. Proc. Natl. Acad. Sci. USA 2017, 114, E7564–E7573. [Google Scholar] [CrossRef]
- Guardado-Calvo, P.; Bignon, E.A.; Stettner, E.; Jeffers, S.A.; Perez-Vargas, J.; Pehau-Arnaudet, G.; Tortorici, M.A.; Jestin, J.L.; England, P.; Tischler, N.D.; et al. Mechanistic insight into Bunyavirus-induced membrane fusion from structure-function analyses of the Hantavirus envelope glycoprotein Gc. PLoS Pathog 2016, 12, e1005813. [Google Scholar] [CrossRef]
- Shi, X.; Goli, J.; Clark, G.; Brauburger, K.; Elliott, R.M. Functional analysis of the Bunyamwera orthobunyavirus Gc glycoprotein. J. Gen. Virol. 2009, 90, 2483–2492. [Google Scholar] [CrossRef]
- Spodareva, V.V.; Grybchuk-Ieremenko, A.; Losev, A.; Votýpka, J.; Lukeš, J.; Yurchenko, V.; Kostygov, A.Y. Diversity and evolution of anuran trypanosomes: Insights from the study of European species. Parasit Vectors 2018, 11, 447. [Google Scholar] [CrossRef] [PubMed]
- Kozminsky, E.; Kraeva, N.; Ishemgulova, A.; Dobáková, E.; Lukeš, J.; Kment, P.; Yurchenko, V.; Votýpka, J.; Maslov, D.A. Host-specificity of monoxenous trypanosomatids: Statistical analysis of the distribution and transmission patterns of the parasites from Neotropical Heteroptera. Protist 2015, 166, 551–568. [Google Scholar] [CrossRef] [PubMed]
- Kraeva, N.; Butenko, A.; Hlaváčová, J.; Kostygov, A.; Myškova, J.; Grybchuk, D.; Leštinová, T.; Votýpka, J.; Volf, P.; Opperdoes, F.; et al. Leptomonas seymouri: Adaptations to the dixenous life cycle analyzed by genome sequencing, transcriptome profiling and co-infection with Leishmania donovani. PLoS Pathog 2015, 11, e1005127. [Google Scholar] [CrossRef] [PubMed]
- Votýpka, J.; d’Avila-Levy, C.M.; Grellier, P.; Maslov, D.A.; Lukeš, J.; Yurchenko, V. New approaches to systematics of Trypanosomatidae: Criteria for taxonomic (re)description. Trends Parasitol 2015, 31, 460–469. [Google Scholar] [CrossRef]
- Grybchuk-Ieremenko, A.; Losev, A.; Kostygov, A.Y.; Lukeš, J.; Yurchenko, V. High prevalence of trypanosome co-infections in freshwater fishes. Folia Parasitol 2014, 61, 495–504. [Google Scholar] [CrossRef]
- Muniz, J.; Medina, H. [Cutaneous leishmaniasis of the guinea pig, Leishmania enriettii n. sp]. Hospital (Rio J.) 1948, 33, 7–25. [Google Scholar]
- Jariyapan, N.; Daroontum, T.; Jaiwong, K.; Chanmol, W.; Intakhan, N.; Sor-Suwan, S.; Siriyasatien, P.; Somboon, P.; Bates, M.D.; Bates, P.A. Leishmania (Mundinia) orientalis n. sp. (Trypanosomatidae), a parasite from Thailand responsible for localised cutaneous leishmaniasis. Parasit Vectors 2018, 11, 351. [Google Scholar] [CrossRef]
- Dougall, A.; Shilton, C.; Low Choy, J.; Alexander, B.; Walton, S. New reports of Australian cutaneous leishmaniasis in Northern Australian macropods. Epidemiol Infect. 2009, 137, 1516–1520. [Google Scholar] [CrossRef]
- Rose, K.; Curtis, J.; Baldwin, T.; Mathis, A.; Kumar, B.; Sakthianandeswaren, A.; Spurck, T.; Low Choy, J.; Handman, E. Cutaneous leishmaniasis in red kangaroos: Isolation and characterisation of the causative organisms. Int. J. Parasitol. 2004, 34, 655–664. [Google Scholar] [CrossRef]
- Barratt, J.; Kaufer, A.; Peters, B.; Craig, D.; Lawrence, A.; Roberts, T.; Lee, R.; McAuliffe, G.; Stark, D.; Ellis, J. Isolation of novel trypanosomatid, Zelonia australiensis sp. nov. (Kinetoplastida: Trypanosomatidae) provides support for a Gondwanan origin of dixenous parasitism in the Leishmaniinae. PLoS Negl. Trop. Dis. 2017, 11, e0005215. [Google Scholar] [CrossRef]
- Desbois, N.; Pratlong, F.; Quist, D.; Dedet, J.P. Leishmania (Leishmania) martiniquensis n. sp. (Kinetoplastida: Trypanosomatidae), description of the parasite responsible for cutaneous leishmaniasis in Martinique Island (French West Indies). Parasite 2014, 21, 12. [Google Scholar] [CrossRef] [PubMed]
- Seblová, V.; Sádlová, J.; Vojtková, B.; Votýpka, J.; Carpenter, S.; Bates, P.A.; Volf, P. The biting midge Culicoides sonorensis (Diptera: Ceratopogonidae) is capable of developing late stage infections of Leishmania enriettii. PLoS Negl. Trop. Dis. 2015, 9, e0004060. [Google Scholar] [CrossRef]
- Dougall, A.M.; Alexander, B.; Holt, D.C.; Harris, T.; Sultan, A.H.; Bates, P.A.; Rose, K.; Walton, S.F. Evidence incriminating midges (Diptera: Ceratopogonidae) as potential vectors of Leishmania in Australia. Int. J. Parasitol. 2011, 41, 571–579. [Google Scholar] [CrossRef] [PubMed]
- Butenko, A.; Kostygov, A.Y.; Sádlová, J.; Kleschenko, Y.; Bečvář, T.; Podešvová, L.; Macedo, D.H.; Žihala, D.; Lukeš, J.; Bates, P.A.; et al. Comparative genomics of Leishmania (Mundinia). BMC Genomics 2019, 20, 726. [Google Scholar] [CrossRef] [PubMed]
- Zídková, L.; Čepička, I.; Votýpka, J.; Svobodová, M. Herpetomonas trimorpha sp. nov. (Trypanosomatidae, Kinetoplastida), a parasite of the biting midge Culicoides truncorum (Ceratopogonidae, Diptera). Int. J. Syst. Evol. Microbiol. 2010, 60, 2236–2246. [Google Scholar] [CrossRef] [PubMed]
- Svobodová, M.; Zídková, L.; Čepička, I.; Oborník, M.; Lukeš, J.; Votýpka, J. Sergeia podlipaevi gen. nov., sp. nov. (Trypanosomatidae, Kinetoplastida), a parasite of biting midges (Ceratopogonidae, Diptera). Int. J. Syst. Evol. Microbiol. 2007, 57, 423–432. [Google Scholar] [CrossRef] [PubMed]
- Podlipaev, S.; Votýpka, J.; Jirků, M.; Svobodová, M.; Lukeš, J. Herpetomonas ztiplika n. sp. (Kinetoplastida: Trypanosomatidae): A parasite of the blood-sucking biting midge Culicoides kibunensis Tokunaga, 1937 (Diptera: Ceratopogonidae). J. Parasitol. 2004, 90, 342–347. [Google Scholar] [CrossRef]
- Bualert, L.; Charungkiattikul, W.; Thongsuksai, P.; Mungthin, M.; Siripattanapipong, S.; Khositnithikul, R.; Naaglor, T.; Ravel, C.; El Baidouri, F.; Leelayoova, S. Autochthonous disseminated dermal and visceral leishmaniasis in an AIDS patient, southern Thailand, caused by Leishmania siamensis. Am. J. Trop Med. Hyg 2012, 86, 821–824. [Google Scholar] [CrossRef]
- Dedet, J.P.; Roche, B.; Pratlong, F.; Cales-Quist, D.; Jouannelle, J.; Benichou, J.C.; Huerre, M. Diffuse cutaneous infection caused by a presumed monoxenous trypanosomatid in a patient infected with HIV. Trans. R. Soc. Trop. Med. Hyg 1995, 89, 644–646. [Google Scholar] [CrossRef]
- Chicharro, C.; Alvar, J. Lower trypanosomatids in HIV/AIDS patients. Ann. Trop. Med. Parasitol. 2003, 97 Suppl. 1, 75–78. [Google Scholar] [CrossRef]
- Dedet, J.P.; Pratlong, F. Leishmania, Trypanosoma and monoxenous trypanosomatids as emerging opportunistic agents. J. Eukaryot Microbiol 2000, 47, 37–39. [Google Scholar] [CrossRef] [PubMed]
- Hartley, M.A.; Ronet, C.; Zangger, H.; Beverley, S.M.; Fasel, N. Leishmania RNA virus: When the host pays the toll. Front. Cell Infect. Microbiol. 2012, 2, 99. [Google Scholar] [CrossRef] [PubMed]

| Viral Sequences | Accession | Length, bp | ORF, AA | RPKM |
|---|---|---|---|---|
| LmarLBV1 S | MK356556 | 721 | 165 | 6,600.13 |
| LmarLBV1 M | MK356555 | 1,244 | 334 | 1,368.42 |
| LmarLBV1 L | MK356554 | 6102 | 2,012 | 1,131.22 |
© 2020 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Grybchuk, D.; Macedo, D.H.; Kleschenko, Y.; Kraeva, N.; Lukashev, A.N.; Bates, P.A.; Kulich, P.; Leštinová, T.; Volf, P.; Kostygov, A.Y.; et al. The First Non-LRV RNA Virus in Leishmania. Viruses 2020, 12, 168. https://doi.org/10.3390/v12020168
Grybchuk D, Macedo DH, Kleschenko Y, Kraeva N, Lukashev AN, Bates PA, Kulich P, Leštinová T, Volf P, Kostygov AY, et al. The First Non-LRV RNA Virus in Leishmania. Viruses. 2020; 12(2):168. https://doi.org/10.3390/v12020168
Chicago/Turabian StyleGrybchuk, Danyil, Diego H. Macedo, Yulia Kleschenko, Natalya Kraeva, Alexander N. Lukashev, Paul A. Bates, Pavel Kulich, Tereza Leštinová, Petr Volf, Alexei Y. Kostygov, and et al. 2020. "The First Non-LRV RNA Virus in Leishmania" Viruses 12, no. 2: 168. https://doi.org/10.3390/v12020168
APA StyleGrybchuk, D., Macedo, D. H., Kleschenko, Y., Kraeva, N., Lukashev, A. N., Bates, P. A., Kulich, P., Leštinová, T., Volf, P., Kostygov, A. Y., & Yurchenko, V. (2020). The First Non-LRV RNA Virus in Leishmania. Viruses, 12(2), 168. https://doi.org/10.3390/v12020168






