Species D Adenoviruses as Oncolytic Viral Vectors
Abstract
1. Introduction
2. Materials and Methods
2.1. Cell Culture
2.2. Recombinant Adenovirus Cloning
2.3. Recombinant Adenovirus Virus Rescue, Purification, and Quantification
2.4. GFP Pictures and Flow Cytometry
2.5. MTT Assay
2.6. qPCR Assay for Viral Replication
2.7. NaI Uptake Assy
2.8. Statistical Analysis
3. Results
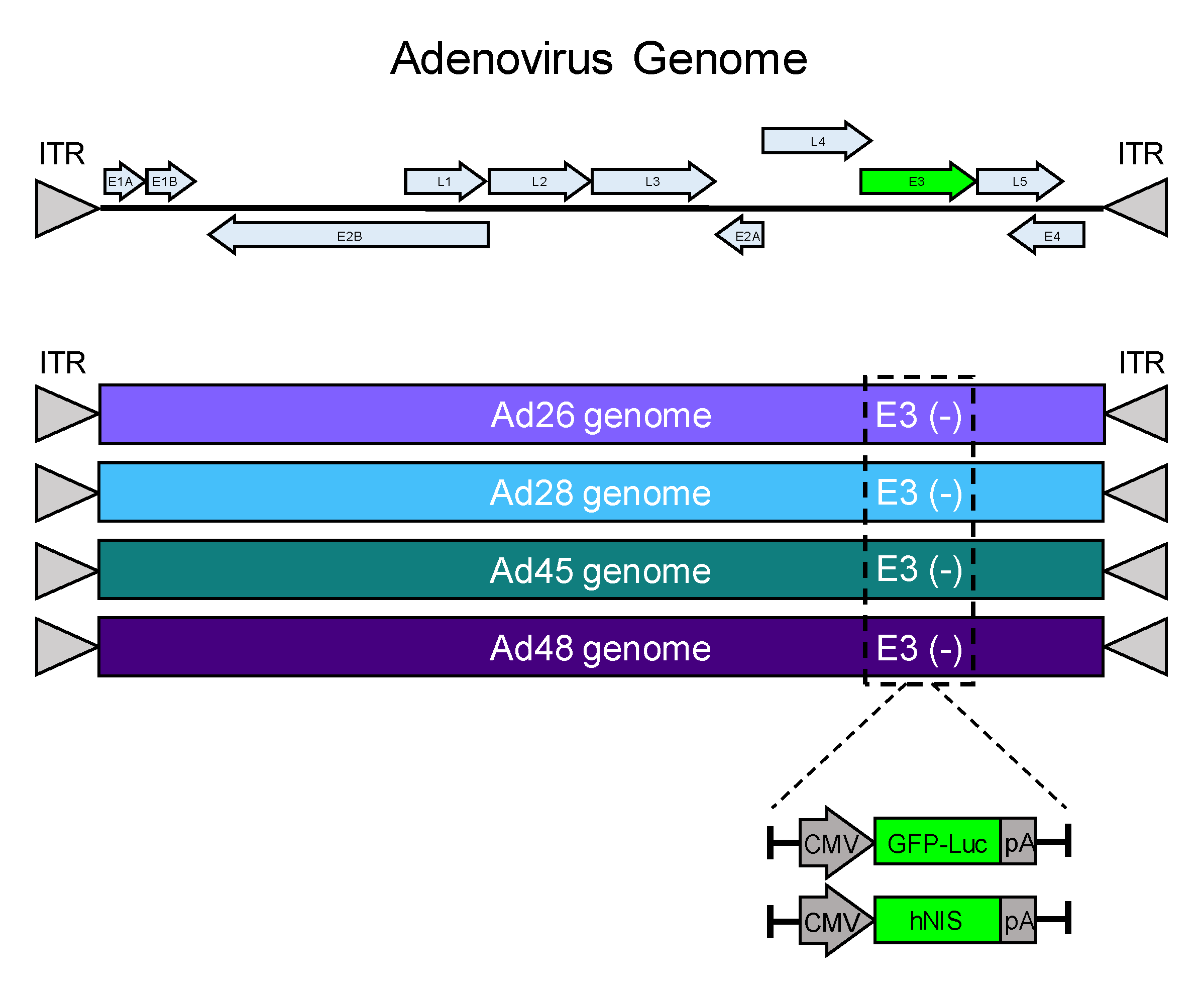
3.1. Development of Replication Competent Species D Adenovirus
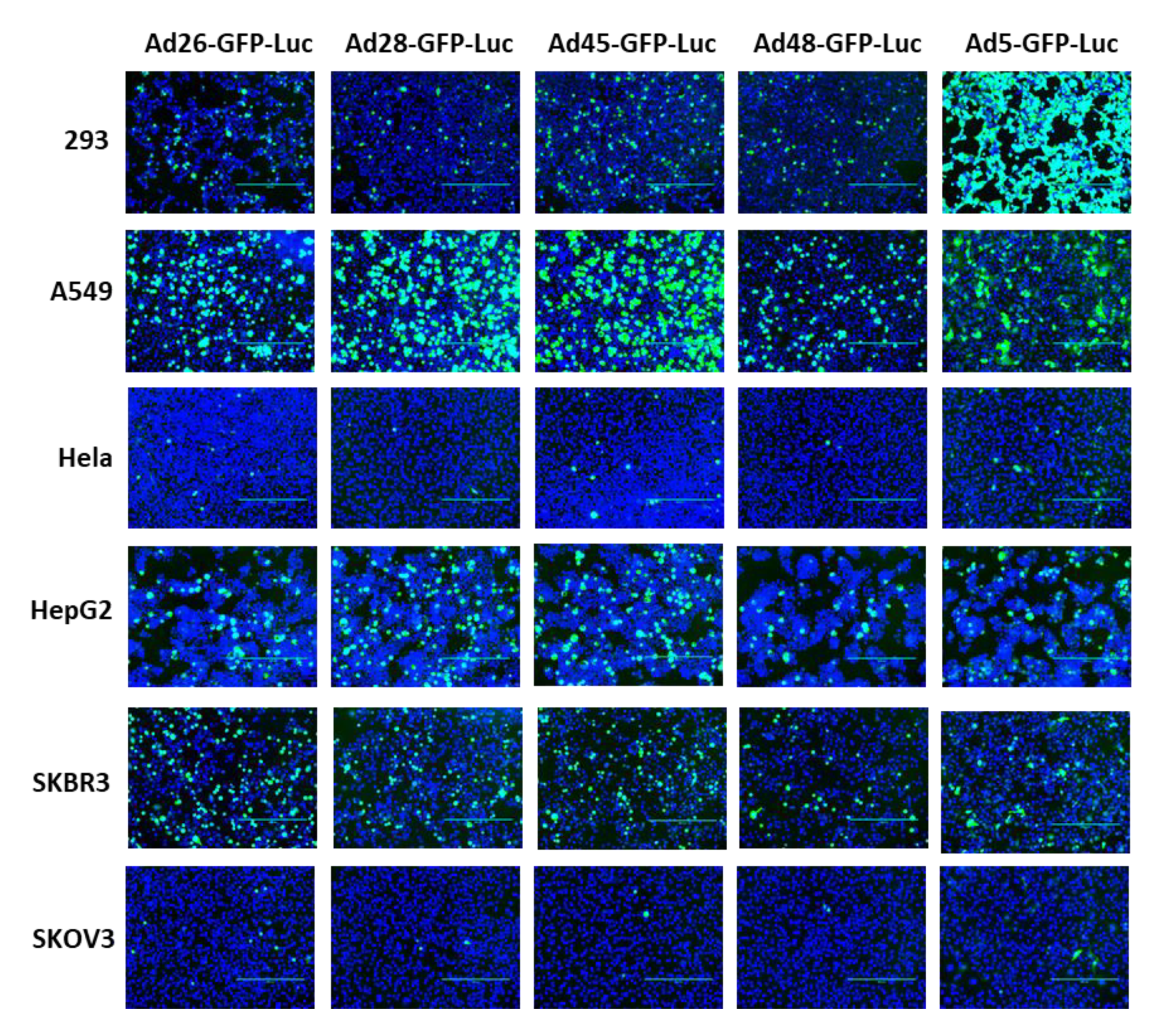
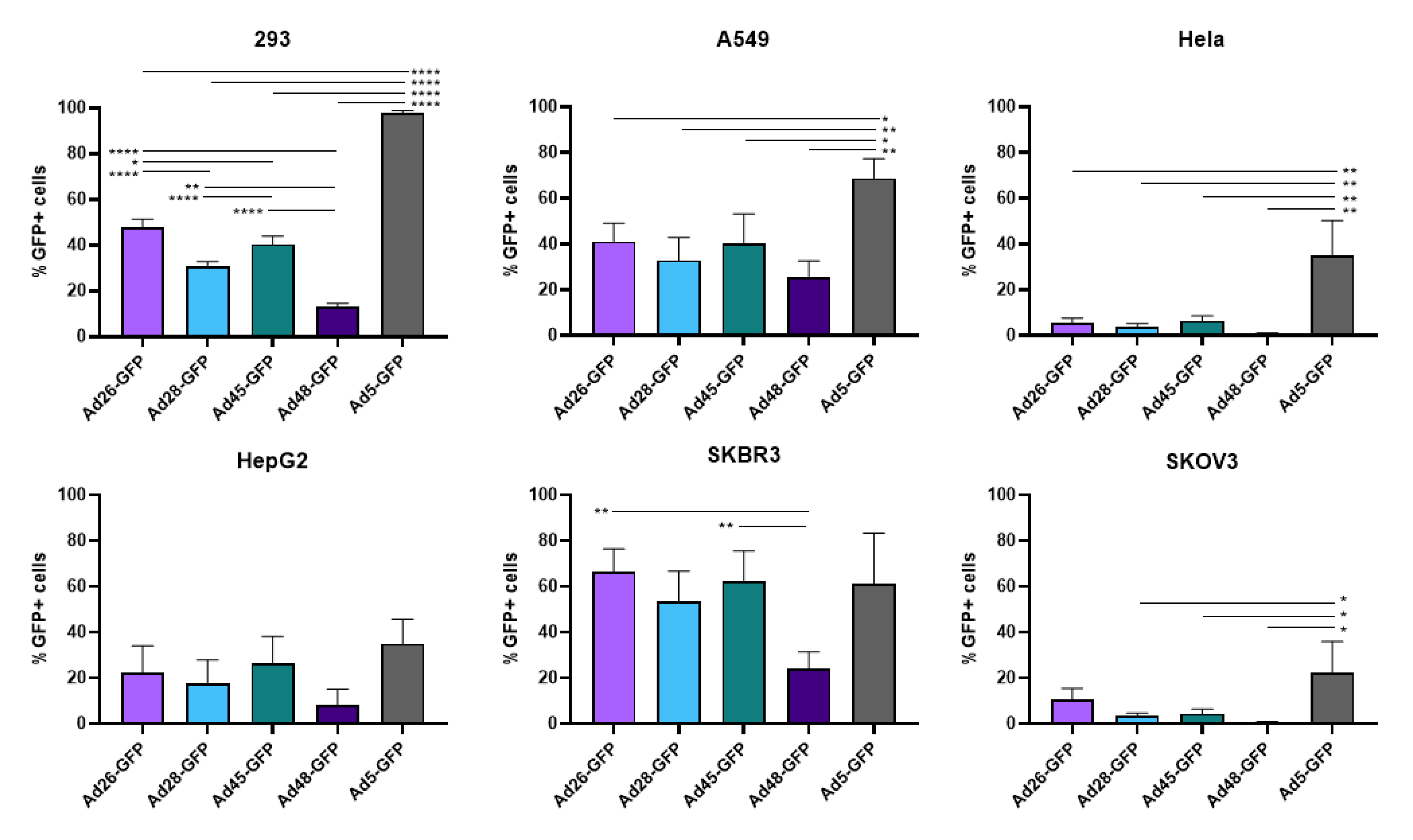
3.2. Viral Transduction and Transgene Expression
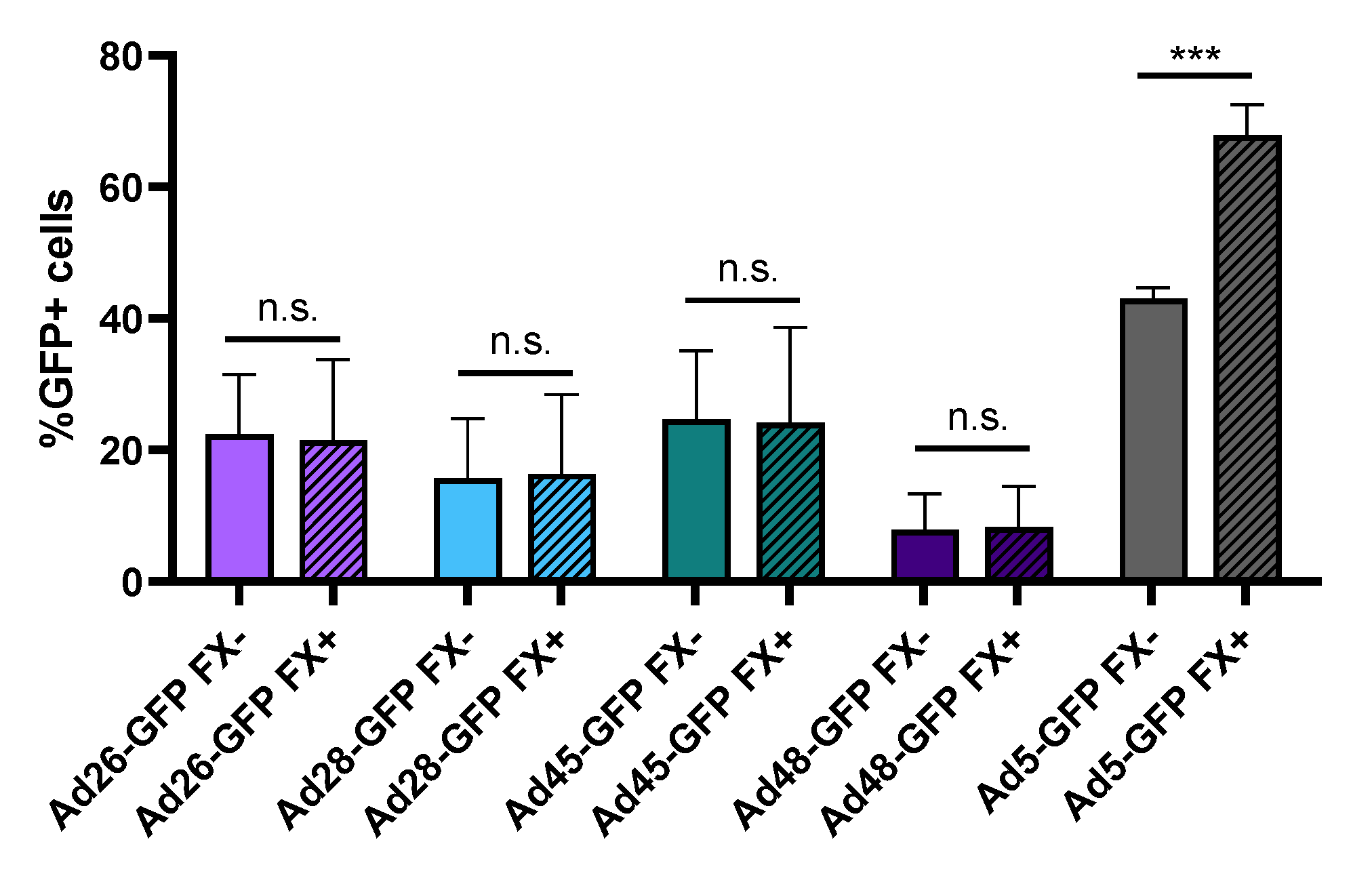
3.3. Binding to Blood Coagulation Factor X
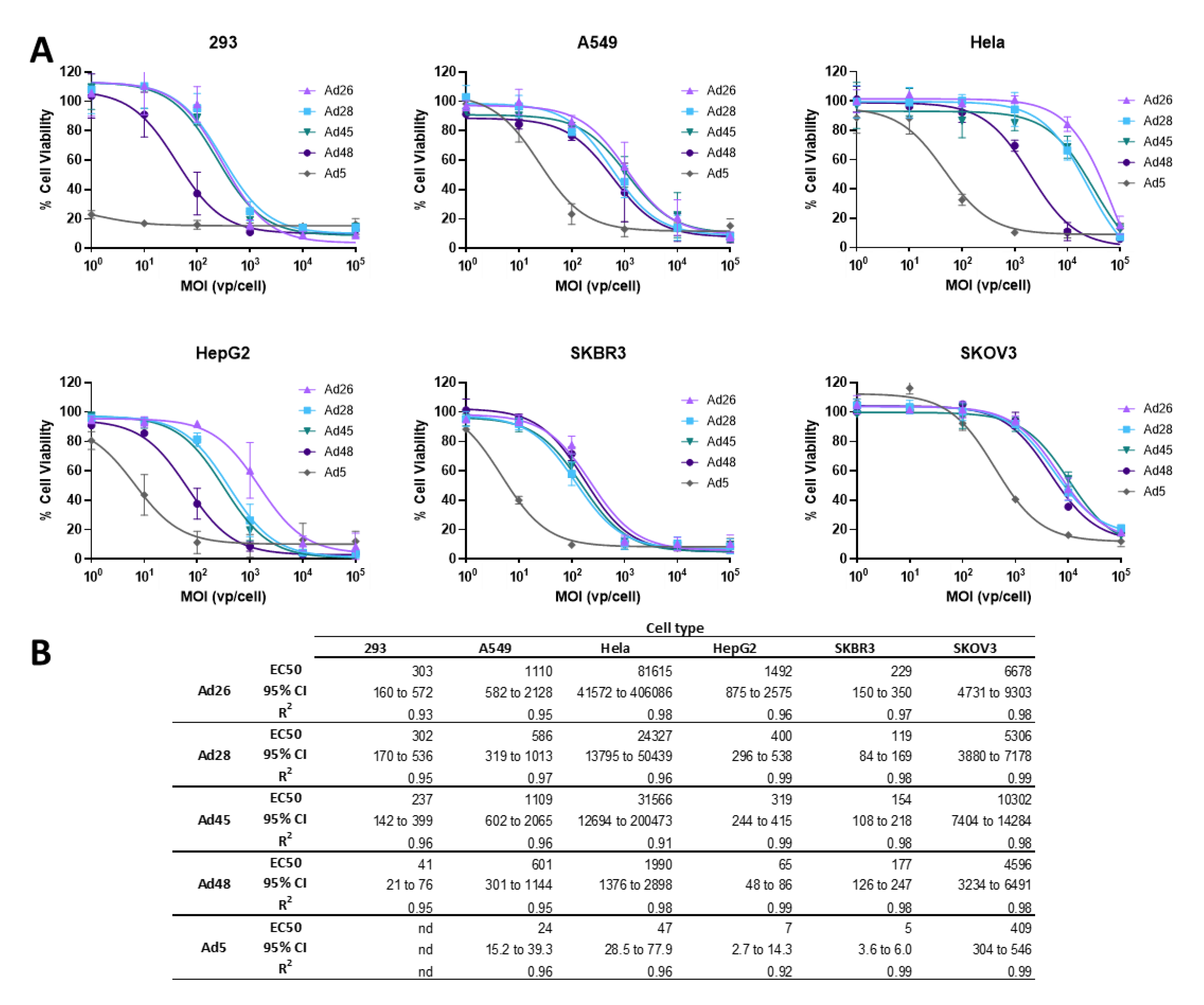
3.4. EC50 of Wild Type Viruses in Cancer Cell Lines
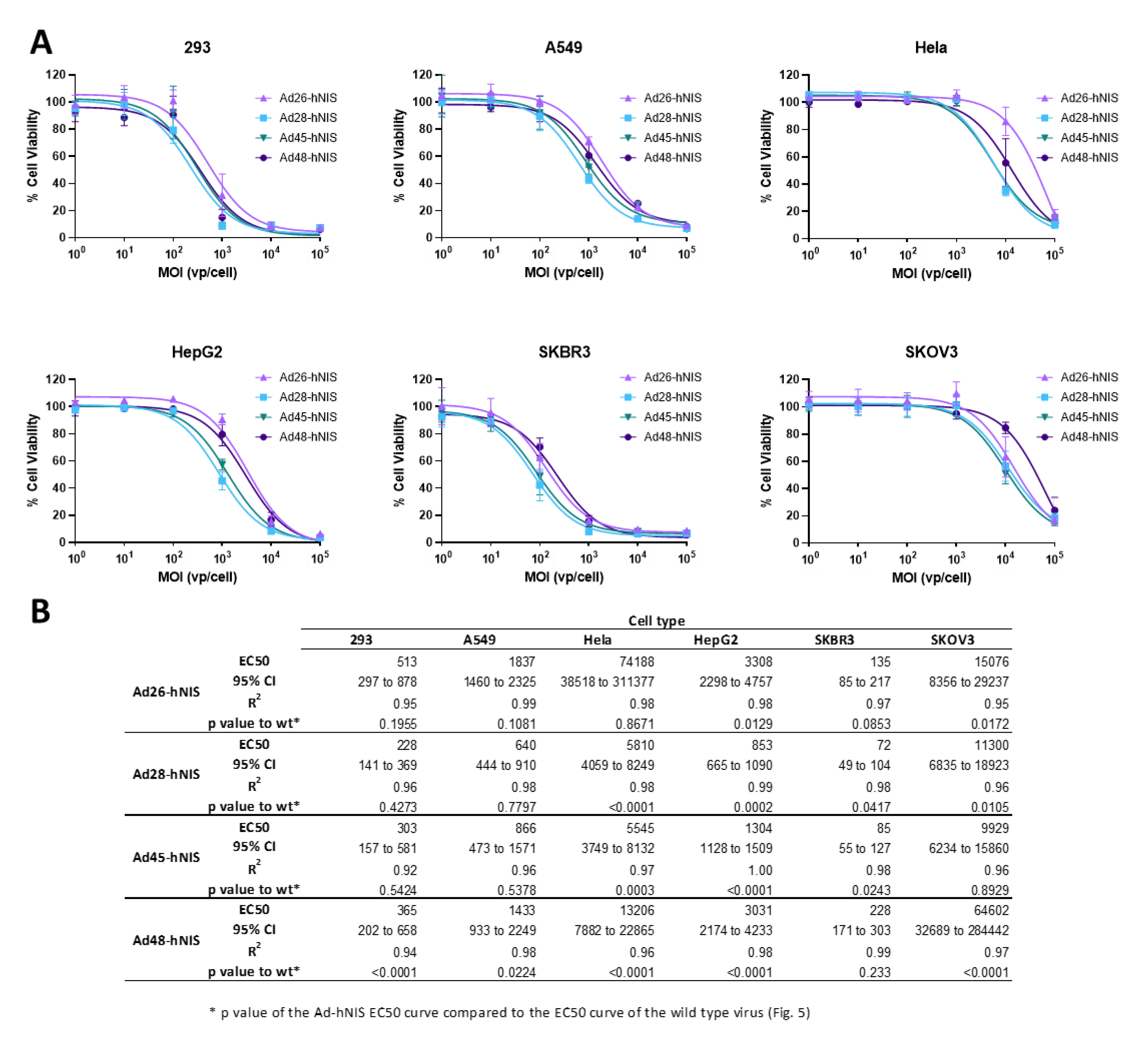
3.5. EC50 of the Recombinant Ad-hNIS Viruses in Cancer Cell Lines
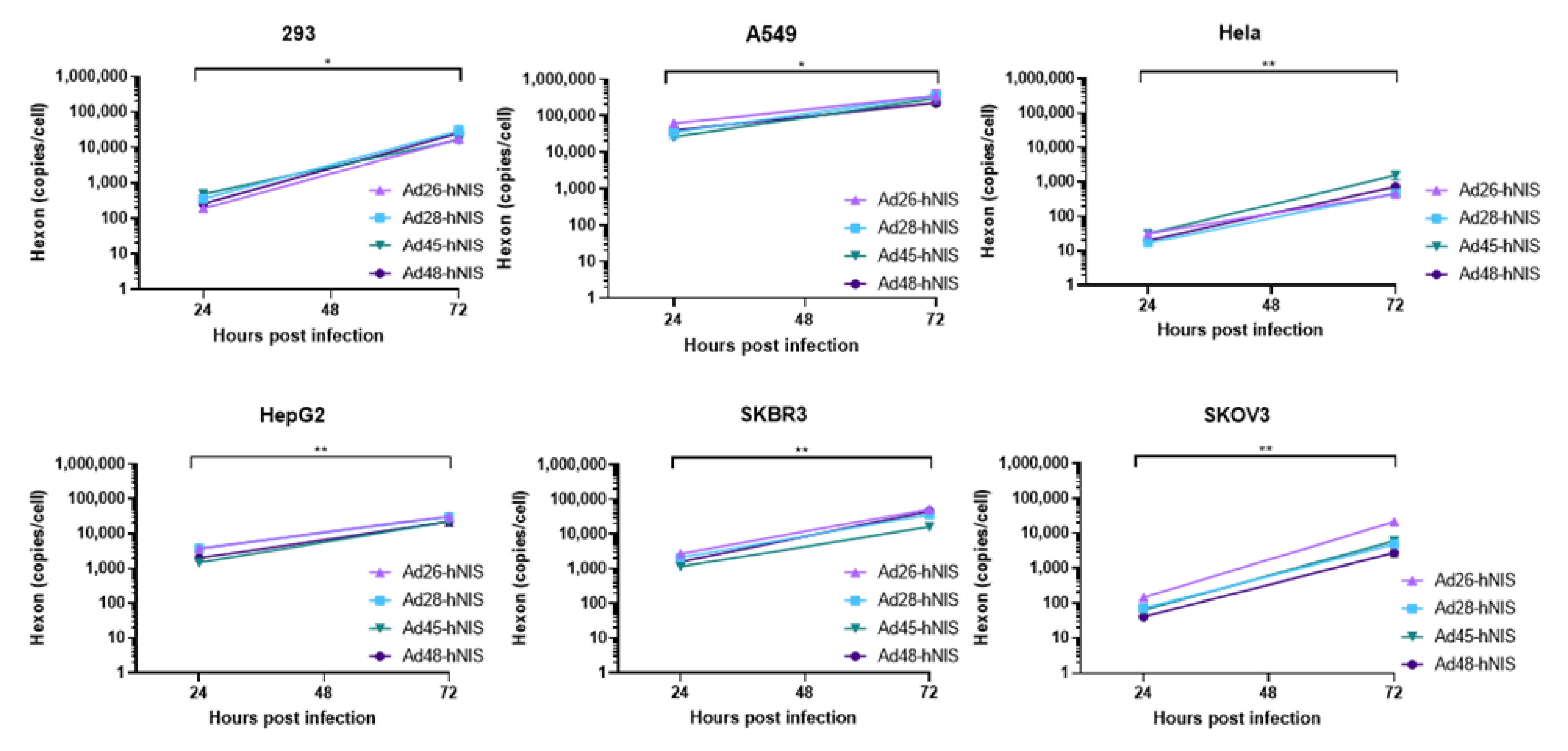
3.6. gDNA Replication Kinetics in Each Cancer Cell Line
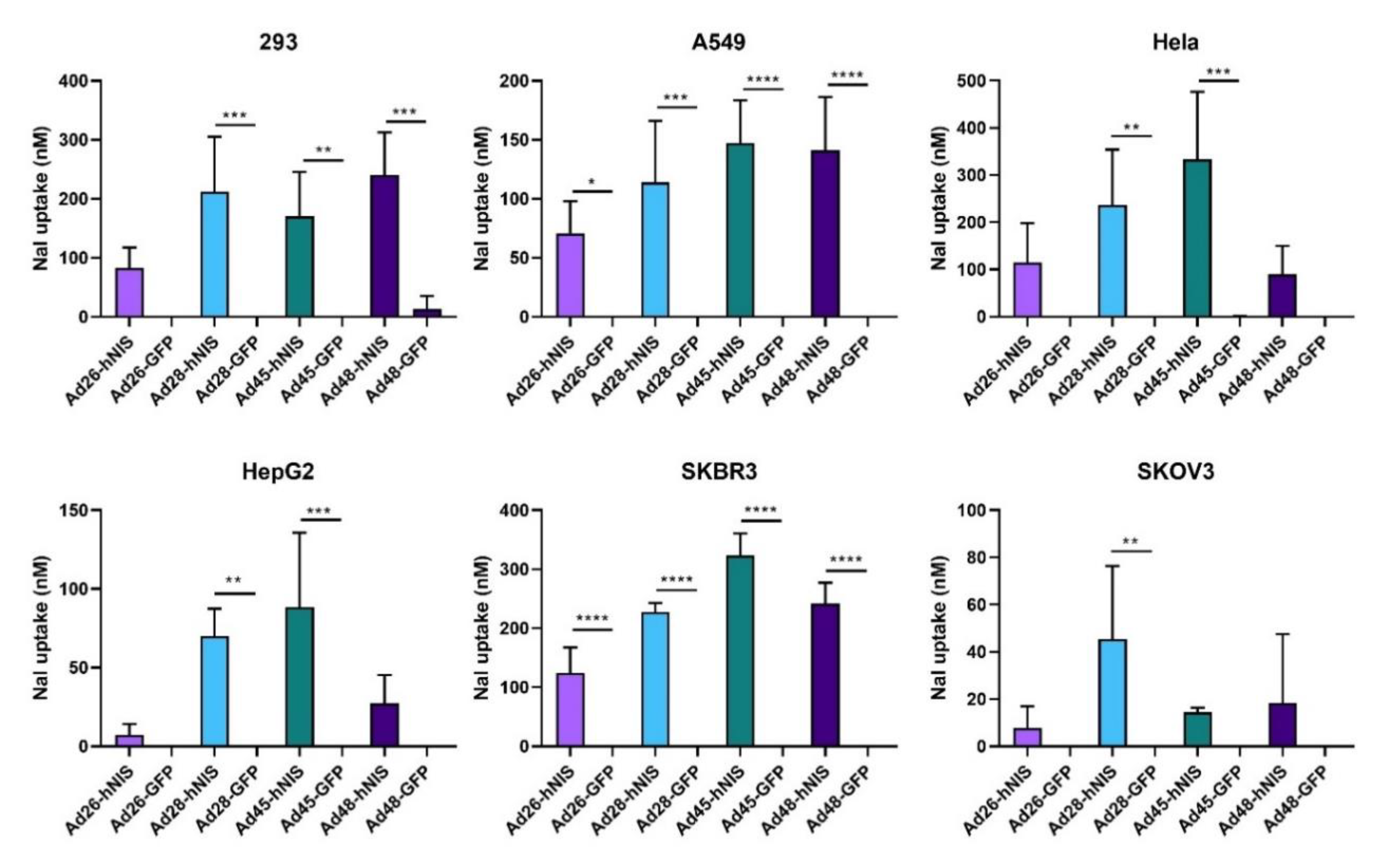
3.7. Increased NaI Uptake after Ad-hNIS Infection
4. Discussion
5. Conclusions
Author Contributions
Funding
Conflicts of Interest
References
- Kaufman, H.L.; Kohlhapp, F.J.; Zloza, A. Oncolytic viruses: A new class of immunotherapy drugs. Nat Rev Drug Discov. 2015, 14, 642–662. [Google Scholar] [CrossRef] [PubMed]
- Uusi-Kerttula, H.; Hulin-Curtis, S.; Davies, J.; Parker, A.L. Oncolytic adenovirus: Strategies and insights for vector design and immuno-oncolytic applications. Viruses 2015, 7, 6009–6042. [Google Scholar] [CrossRef] [PubMed]
- Fields, B.N.; Knipe, D.M.; Howley, P.M. Fields Virology; Wolters Kluwer Health/Lippincott Williams & Wilkins: Philadelphia, PA, USA, 2013. [Google Scholar]
- Nwanegbo, E.; Vardas, E.; Gao, W.; Whittle, H.; Sun, H.; Rowe, D.; Robbins, P.D.; Gambotto, A. Prevalence of neutralizing antibodies to adenoviral serotypes 5 and 35 in the adult populations of the Gambia, South Africa, and the United States. Clin. Diagnost. Lab. Immunol. 2004, 11, 351–357. [Google Scholar] [CrossRef] [PubMed]
- Abbink, P.; Lemckert, A.A.C.; Ewald, B.A.; Lynch, D.M.; Denholtz, M.; Smits, S.; Holterman, L.; Damen, I.; Vogels, R.; Thorner, A.R.; et al. Comparative seroprevalence and immunogenicity of six rare serotype recombinant adenovirus vaccine vectors from subgroups B and D. J. Virol. 2007, 81, 4654–4663. [Google Scholar] [CrossRef] [PubMed]
- Mennechet, F.J.D.; Paris, O.; Ouoba, A.R.; Salazar Arenas, S.; Sirima, S.B.; Takoudjou Dzomo, G.R.; Diarra, A.; Traore, I.T.; Kania, D.; Eichholz, K.; et al. A review of 65 years of human adenovirus seroprevalence. Expert Rev. Vacc. 2019, 18, 597–613. [Google Scholar] [CrossRef] [PubMed]
- Chen, Y.; Yu, D.-C.; Charlton, D.; Henderson, D.R. Pre-existent adenovirus antibody inhibits systemic toxicity and antitumor activity of CN706 in the nude mouse LNCaP xenograft model: Implications and proposals for human therapy. Hum. Gene Ther. 2000, 11, 1553–1567. [Google Scholar] [CrossRef]
- Dhar, D.; Spencer, J.F.; Toth, K.; Wold, W.S.M. Effect of preexisting immunity on oncolytic adenovirus vector INGN 007 antitumor efficacy in immunocompetent and immunosuppressed syrian hamsters. J. Virol. 2009, 83, 2130–2139. [Google Scholar] [CrossRef]
- Hedley, S.J.; Chen, J.; Mountz, J.D.; Li, J.; Curiel, D.T.; Korokhov, N.; Kovesdi, I. Targeted and shielded adenovectors for cancer therapy. Cancer Immunol. Immunother. 2006, 55, 1412–1419. [Google Scholar] [CrossRef]
- Zhang, Z.; Krimmel, J.; Zhang, Z.; Hu, Z.; Seth, P. Systemic delivery of a novel liver-detargeted oncolytic adenovirus causes reduced liver toxicity but maintains the antitumor response in a breast cancer bone metastasis model. Hum. Gene Ther. 2011, 22, 1137–1142. [Google Scholar] [CrossRef]
- Waddington, S.N.; McVey, J.H.; Bhella, D.; Parker, A.L.; Barker, K.; Atoda, H.; Pink, R.; Buckley, S.M.K.; Greig, J.A.; Denby, L.; et al. Adenovirus serotype 5 hexon mediates liver gene transfer. Cell 2008, 132, 397–409. [Google Scholar] [CrossRef]
- Rosewell Shaw, A.; Suzuki, M. Recent advances in oncolytic adenovirus therapies for cancer. Curr. Opin. Virol. 2016, 21, 9–15. [Google Scholar] [CrossRef] [PubMed]
- Li, Y.; Pong, R.-C.; Bergelson, J.M.; Hall, M.C.; Sagalowsky, A.I.; Tseng, C.-P.; Wang, Z.; Hsieh, J.-T. Loss of adenoviral receptor expression in human bladder cancer cells: A potential impact on the efficacy of gene therapy. Cancer Res. 1999, 59, 325–330. [Google Scholar] [PubMed]
- Fukazawa, T.; Matsuoka, J.; Yamatsuji, T.; Maeda, Y.; Durbin, M.L.; Naomoto, Y. Adenovirus-mediated cancer gene therapy and virotherapy. Int. J. Mol. Med. 2010, 25, 3–10. [Google Scholar] [CrossRef] [PubMed]
- Wunder, T.; Schumacher, U.; Friedrich, R.E. Coxsackie adenovirus receptor expression in carcinomas of the head and neck. Anticancer Res. 2012, 32, 1057–1062. [Google Scholar] [PubMed]
- Chen, C.Y.; Senac, J.S.; Weaver, E.A.; May, S.M.; Jelinek, D.F.; Greipp, P.; Witzig, T.; Barry, M.A. Species D adenoviruses as oncolytics against B-cell cancers. Clin. Cancer Res Offic. J. Am. Assoc. Cancer Res. 2011, 17, 6712–6722. [Google Scholar] [CrossRef]
- Weaver, E.A.; Chen, C.Y.; May, S.M.; Barry, M.E.; Barry, M.A. Comparison of adenoviruses as oncolytics and cancer vaccines in an immunocompetent B cell lymphoma model. Hum. Gene Ther. 2011, 22, 1095–1100. [Google Scholar] [CrossRef]
- Barouch, D.H.; Kik, S.V.; Weverling, G.J.; Dilan, R.; King, S.L.; Maxfield, L.F.; Clark, S.; Ng’ang’a, D.; Brandariz, K.L.; Abbink, P.; et al. International seroepidemiology of adenovirus serotypes 5, 26, 35, and 48 in pediatric and adult populations. Vaccine 2011, 29, 5203–5209. [Google Scholar] [CrossRef]
- Mast, T.C.; Kierstead, L.; Gupta, S.B.; Nikas, A.A.; Kallas, E.G.; Novitsky, V.; Mbewe, B.; Pitisuttithum, P.; Schechter, M.; Vardas, E.; et al. International epidemiology of human pre-existing adenovirus (Ad) type-5, type-6, type-26 and type-36 neutralizing antibodies: Correlates of high Ad5 titers and implications for potential HIV vaccine trials. Vaccine 2010, 28, 950–957. [Google Scholar] [CrossRef]
- Kahl, C.A.; Bonnell, J.; Hiriyanna, S.; Fultz, M.; Nyberg-Hoffman, C.; Chen, P.; King, C.R.; Gall, J.G.D. Potent immune responses and in vitro pro-inflammatory cytokine suppression by a novel adenovirus vaccine vector based on rare human serotype 28. Vaccine 2010, 28, 5691–5702. [Google Scholar] [CrossRef]
- Zhang, Y.; Bergelson, J.M. Adenovirus Receptors. J. Virol. 2005, 79, 12125–12131. [Google Scholar] [CrossRef]
- Li, H.; Rhee, E.G.; Masek-Hammerman, K.; Teigler, J.E.; Abbink, P.; Barouch, D.H. Adenovirus serotype 26 utilizes CD46 as a primary cellular receptor and only transiently activates T lymphocytes following vaccination of rhesus monkeys. J. Virol. 2012, 86, 10862–10865. [Google Scholar] [CrossRef] [PubMed]
- Camacho, Z.T.; Turner, M.A.; Barry, M.A.; Weaver, E.A. CD46-mediated transduction of a species D adenovirus vaccine improves mucosal vaccine efficacy. Hum. Gene Ther. 2014, 25, 364–374. [Google Scholar] [CrossRef] [PubMed]
- Thirion, C.; Lochmüller, H.; Ruzsics, Z.; Boelhauve, M.; König, C.; Thedieck, C.; Kutik, S.; Geiger, C.; Kochanek, S.; Volpers, C.; et al. Adenovirus vectors based on human adenovirus type 19a have high potential for human muscle-directed gene therapy. Hum. Gene Ther. 2006, 17, 193–205. [Google Scholar] [CrossRef] [PubMed]
- Touchefeu, Y.; Franken, P.J.; Harrington, K. Radiovirotherapy: Principles and prospects in oncology. Curr. Pharma. Des. 2012, 18, 3313–3320. [Google Scholar] [CrossRef]
- Miller, A.; Russell, S.J. The use of the NIS reporter gene for optimizing oncolytic virotherapy. Expert Opin. Biol. Ther. 2016, 16, 15–32. [Google Scholar] [CrossRef]
- Peerlinck, I.; Merron, A.; Baril, P.; Conchon, S.; Martin-Duque, P.; Hindorf, C.; Burnet, J.; Quintanilla, M.; Hingorani, M.; Iggo, R.; et al. Targeted Radionuclide Therapy Using a Wnt-Targeted Replicating Adenovirus Encoding the Na/I Symporter. Clin. Cancer Res. 2009, 15, 6595–6601. [Google Scholar] [CrossRef]
- Chen, R.F.; Li, Z.H.; Pan, Q.H.; Zhou, J.J.; Tang, Q.B.; Yu, F.Y.; Zhou, Q.B.; Wang, J.; Chen, J.S. In vivo radioiodide imaging and treatment of pancreatic cancer xenografts after MUC1 promoter-driven expression of the human sodium-iodide symporter. Pancreatology 2007, 7, 505–513. [Google Scholar] [CrossRef]
- Faivre, J.; Clerc, J.; Gérolami, R.; Hervé, J.; Longuet, M.; Liu, B.; Roux, J.; Moal, F.; Perricaudet, M.; Bréchot, C. Long-term radioiodine retention and regression of liver cancer after sodium iodide symporter gene transfer in wistar rats. Cancer Res. 2004, 64, 8045–8051. [Google Scholar] [CrossRef][Green Version]
- Hakkarainen, T.; Rajecki, M.; Sarparanta, M.; Tenhunen, M.; Airaksinen, A.J.; Desmond, R.A.; Kairemo, K.; Hemminki, A. Targeted radiotherapy for prostate cancer with an oncolytic adenovirus coding for human sodium iodide symporter. Clin. Cancer Res. 2009, 15, 5396–5403. [Google Scholar] [CrossRef]
- Robinson, C.M.; Seto, D.; Jones, M.S.; Dyer, D.W.; Chodosh, J. Molecular evolution of human species D adenoviruses. Inf. Gen. Evol. J. Mol. Epidemiol. Evolut. Gen. Inf. Dis. 2011, 11, 1208–1217. [Google Scholar] [CrossRef]
- Weaver, E.A. Vaccines within vaccines: The use of adenovirus types 4 and 7 as influenza vaccine vectors. Hum. Vacc.Immunother. 2014, 10, 544–556. [Google Scholar] [CrossRef] [PubMed][Green Version]
- Weaver, E.A.; Barry, M.A. Low seroprevalent species D adenovirus vectors as influenza vaccines. PLoS ONE 2013, 8, e73313. [Google Scholar] [CrossRef] [PubMed]
- Waltz, F.; Pillette, L.; Ambroise, Y. A nonradioactive iodide uptake assay for sodium iodide symporter function. Analyt. Biochem. 2010, 396, 91–95. [Google Scholar] [CrossRef] [PubMed]
- Kalyuzhniy, O.; Di Paolo, N.C.; Silvestry, M.; Hofherr, S.E.; Barry, M.A.; Stewart, P.L.; Shayakhmetov, D.M. Adenovirus serotype 5 hexon is critical for virus infection of hepatocytes in vivo. Proc. Natl. Acad. Sci. USA 2008, 105, 5483–5488. [Google Scholar] [CrossRef] [PubMed]
- Shashkova, E.V.; May, S.M.; Barry, M.A. Characterization of human adenovirus serotypes 5, 6, 11, and 35 as anticancer agents. Virology 2009, 394, 311–320. [Google Scholar] [CrossRef] [PubMed]
- Chen, H.; Xiang, Z.Q.; Li, Y.; Kurupati, R.K.; Jia, B.; Bian, A.; Zhou, D.M.; Hutnick, N.; Yuan, S.; Gray, C.; et al. Adenovirus-based vaccines: Comparison of vectors from three species of adenoviridae. J. Virol. 2010, 84, 10522–10532. [Google Scholar] [CrossRef]
- Baker, A.T.; Greenshields-Watson, A.; Coughlan, L.; Davies, J.A.; Uusi-Kerttula, H.; Cole, D.K.; Rizkallah, P.J.; Parker, A.L. Diversity within the adenovirus fiber knob hypervariable loops influences primary receptor interactions. Nat. Commun. 2019, 10, 741. [Google Scholar] [CrossRef]
- Nestić, D.; Uil, T.G.; Ma, J.; Roy, S.; Vellinga, J.; Baker, A.H.; Custers, J.; Majhen, D. αvβ3 Integrin is required for efficient infection of epithelial cells with human adenovirus type 26. J. Virol. 2018, 93, e01474-18. [Google Scholar] [CrossRef]
- Stichling, N.; Suomalainen, M.; Flatt, J.W.; Schmid, M.; Pacesa, M.; Hemmi, S.; Jungraithmayr, W.; Maler, M.D.; Freudenberg, M.A.; Plückthun, A.; et al. Lung macrophage scavenger receptor SR-A6 (MARCO) is an adenovirus type-specific virus entry receptor. PLoS Path. 2018, 14, e1006914. [Google Scholar] [CrossRef]
- Baker, A.T.; Mundy, R.M.; Davies, J.A.; Rizkallah, P.J.; Parker, A.L. Human adenovirus type 26 uses sialic acid–bearing glycans as a primary cell entry receptor. Sci. Adv. 2019, 5, eaax3567. [Google Scholar] [CrossRef]
- Uusi-Kerttula, H.; Davies, J.; Coughlan, L.; Hulin-Curtis, S.; Jones, R.; Hanna, L.; Chester, J.D.; Parker, A.L. Pseudotyped αvβ6 integrin-targeted adenovirus vectors for ovarian cancer therapies. Oncotarget 2016, 7. [Google Scholar] [CrossRef] [PubMed]
| Cell Line | Cell Origin |
|---|---|
| 293 | Embryonic kidney |
| A549 | Lung carcinoma |
| Hela | Cervical carcinoma |
| HepG2 | Hepatocellular carcinoma |
| SKBR3 | Breast carcinoma |
| SKOV3 | Ovarian carcinoma |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2020 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Bullard, B.L.; Corder, B.N.; Weaver, E.A. Species D Adenoviruses as Oncolytic Viral Vectors. Viruses 2020, 12, 1399. https://doi.org/10.3390/v12121399
Bullard BL, Corder BN, Weaver EA. Species D Adenoviruses as Oncolytic Viral Vectors. Viruses. 2020; 12(12):1399. https://doi.org/10.3390/v12121399
Chicago/Turabian StyleBullard, Brianna L., Brigette N. Corder, and Eric A. Weaver. 2020. "Species D Adenoviruses as Oncolytic Viral Vectors" Viruses 12, no. 12: 1399. https://doi.org/10.3390/v12121399
APA StyleBullard, B. L., Corder, B. N., & Weaver, E. A. (2020). Species D Adenoviruses as Oncolytic Viral Vectors. Viruses, 12(12), 1399. https://doi.org/10.3390/v12121399






